Abstract
Based on our continuous effort to investigate chemistry and biology of the plant secondary metabolites, we were able to isolate a glycosidal flavonoid 1 from the Wild Egyptian Artichoke. The activity of dihydromyricetin 3-O-rhamnoside (sin. dihydromyricitrin, ampelopsin 3-O-rhamnoside) (1) against acetylcholinesterase (AChE) and butyrylcholinesterase (BChE); its absolute configuration using X-ray crystallography were determined for the first time. Inhibitory activity of 1 against AChE and BChE enzymes were determined using a slightly modified version of Ellman’s method. Compound 1 was revealed to have a potent inhibition against acetylcholinesterase (AChE) and butyrylcholinesterase (BChE) with IC50 values of 0.070 ± 0.008 and 0.071 ± 0.004 mM, respectively, where IC50 values of the reference drug (galanthamine) were 0.023 ± 0.15 and 0.047 ± 0.91 mM. Compound 1 could be a promising molecule against Alzheimer’s disease.
Similar content being viewed by others
Introduction
In many countries, artichoke is a traditionally consumed vegetable and is used as a leaf plant to produce extracts for dietary and medicinal applications. The artichoke plants and their constituents exhibit diverse and important pharmacological effects and health benefits1,2,3,4. The phytochemical investigation of the artichoke leaf extract resulted in the isolation of compound 1 (Fig. 1). The crystals of 1 were obtained using a mixture of chloroform and methanol and its crystallographic data was obtained which enabled us to determine the absolute configuration. Simulating the chemical molecular interactions formed between ligands and active-site amino acids of proteins brings us to a level of understanding that why ligands would reveal different biochemical characteristics including inhibitory effects on some certain targets and how this would vary between ligands and proteins. With a clear understanding about the atomic interactions triggering compounds to inhibit enzymes would lead us to the development of potential inhibitor compounds. In this study the results of the docking simulations carried out on complexes of AChE and BChE with compound 1 help us determine those amino acids playing a key role inside the ligand-binding domains—and the dominant poses that would be observed.
Results and discussion
The methanolic extract of the leaves of artichoke plant yielded compound 1 after fractionation and repeated chromatographic separations on silica gel column as in references1,5.
Needle crystals of compound 1 were formed from a solvent mixture of chloroform and methanol. The structure of 1 was confirmed by the single-crystal X-ray crystallographic measurements (Fig. 2) and accurate mass measurements (measured m/z 465.1030 (M–H+); calculated 465.1033(M–H+)) and the absolute configuration of the seven chiral centers was determined to be 2R, 3R, 1´´S, 2´´R, 3´´R, 4´´R, and 5´´S.
In the current work, inhibitory effect of compound 1 against AChE and BChE was assayed, which was found to possess a marked inhibition with IC50 values of 0.070 ± 0.008 and 0.071 ± 0.004 mM, respectively. IC50 values of the reference drug (galanthamine) towards AChE and BChE were revealed to be 0.023 ± 0.15 and 0.047 ± 0.91 mM. The chemical interactions of dihydromyricetin 3-O-rhamnoside inside AChE and BChE were simulated using IFD method—by which it can be figured out how the ligand can stick to the binding domains of the proteins and which amino acids have mostly contributed to the interactions. This method is able to generate multiple poses for a single ligand; thus, the ligand-binding energy would vary from pose to pose—following our understanding that ligands inside the binding domains are dynamically stable (Fig. 3).
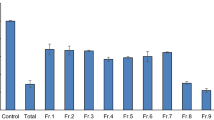
The distributions of the binding energies of the compound inside both AChE and BChE were presented using box plot along with their frequencies in Fig. 4 (the graph in the left panel). The plots present the minimum, maximum, first quartile, median, and third quartile values of the binding energies. The lowest binding energies of the compound were calculated to be around − 12 and − 11 kcal/mol inside AChE and BChE targets, respectively. Both the proteins revealed nearly the same median value, − 8 kcal/mol—by which it would be understand that the compound can sticks to the proteins through favorable interactions. If the binding energies are compared in the two proteins, AChE has been found more accommodating for interaction with the compound. The graph in the right panel in Fig. 4 corresponds to kernel density estimate (KDE) method that measures the probability density function (PDF), yielding the probability that the docking poses fall within specific ranges of binding energies. There are not significant differences between the PDF of the AChE and BChE complexes, but they are not equitable. The portions ranging from − 8 to − 10 kcal/mol have been more converged—however, the distribution belonging to AChE (labeled by red color) has been skewed to the right.
The IFD scores of compound 1 in different poses inside AChE and BChE – the minimum, maximum, first quartile, median, and third quartile values are presented (left panel). Probability density function of the IFD scores between the two systems are compared using KDE method (right panel). The AChE complex was represented with more varieties of the poses which lead to diverse binding energies, compared to the BChE complex. The lowest binding energies were calculated around − 12 and − 10 kcal/mol for the AChE and BChE complexes, respectively, demonstrating that AChE is more favourite for compound 1.
Figure 5 shows the representative poses of the compound inside the AChE and BChE, generated through the top-score poses. What we can understand from the 2D diagrams are the potential interactions (including the polar and nonpolar bonds) formed between the compound and the neighboring amino acids. Additionally, these can create a clearer picture of the roles and level of contribution of each amino acid inside the binding domain, which would lead us to a much deeper understanding of the inhibition mechanism. Regarding the AChE complex, the polar interactions are more dominant, to which amino acids Asp74, Arg296, Tye124, Phe295, Ser293 and Gln291 contributed. Tyr72 has been found the only amino acid forming a Pi–Pi stacking interaction with the aromatic ring of the compound.
2D diagrams of compound 1 inside AChE and BChE. The important amino acids surrounding the ligand are presented with different colors, which correspond to their chemical properties. The polar and nonpolar interactions formed between the atoms of the ligand and the amino acids are shown differently (see the legend). The poses were generated from the systems with the lowest binding energies (top scores), which represent the position of the ligand for each complex—however the interactions would vary between the complexes.
The compound in AChE unlike that in BChE almost equally formed proportion of both hydrogen bonds and hydrophobic interactions with the amino acids. The diagrams suggest that amino acids Ser198 (sidechain) and His438 (backbone) in BChE correspond to amino acids Asp74 (sidechain) and Arg296 (backbone) in BChE, respectively, which have contributed to forming hydrogen bonds with the hydroxyl groups of the ligand. It has been found that both amino acids Phe329 and Trp231 from BChE participated in Pi–Pi stacking interactions with the aromatic ring of the compound—analogous to Tyr72 in AChE.
Crystal data of compound 1
The molecular structure of 1 was crystallized in the Monoclinic, P21, a = 12.2665 (3) Å, b = 6.9450 (2) Å, c = 12.9980 (3) Å, °, β = 97.336 (1)°, V = 1098.25 (5) Å3, Z = 2. In the title molecule, C21H27O15, the crystallographic data and refinement information are presented in the supplementary data (Table s1). Table 1 shows the specified bond lengths and angles. As seen in Fig. 2, the asymmetric unit has one independent molecule in addition to three water molecules. The bonds parameters such as length and angle are all within acceptable limits6. Many intermolecular hydrogen bonds connect molecules in the crystal packing (Fig. 3; Table 2).
Methods
Plant material, extraction, and isolation
See references1,5,6. The plant (Cynara cardunculus L. var. sylvestris (Lam.) Fiori) was identified by Prof. Dr. Ahmed R. Atawia. A voucher specimen with the code “2007-artich-Sinai” was deposited in the Department of Horticulture, Faculty of Agriculture, Benha University, Egypt. The plant name has been checked with http://www.theplantlist.org. The permission to collect artichoke plant and research on it was approved by the Ethical Committee of Research at the Faculty of Pharmacy, Mansoura University, Egypt (code number 2014/71) and was consistent with GCP guidelines and the applicable regulatory requirements.
X-ray crystallography
As in reference7 Molecule 1 was produced as single crystals by gradual evaporation of the pure compound in a solvent combination of chloroform and methanol at ambient temperature. Data were acquired using a Bruker APEX-II D8 Venture area diffractometer with graphite monochromatic Cu K radiation, = 1.54178 at 296 (2) K. Bruker SAINT performed cell refinement and data reduction. The structure was solved using SHELXT9,10. Full-matrix least-squares approaches using anisotropic thermal data for nonhydrogen atoms on F were used for the final refinement. The Cambridge Crystallographic Data Centre’s CCDC 2161839, which provides supplemental crystallographic data for this chemical, may be downloaded for free at https://www.ccdc.cam.ac.uk/datarequest/cif.
Cholinesterase inhibition assays
Inhibitory activities of compound 1 against AChE and BChE enzymes were determined using the modified spectrophotometric method of Ellman procedure8. The enzyme sources were electric eel acetylcholinesterase (Type-VI-S, EC 3.1.1.7, Sigma) and horse serum butyrylcholinesterase (EC 3.1.1.8, Sigma), while the reaction substrates were acetylthiocholine iodide and butyrylthiocholine chloride (Sigma, St. Louis, MO, USA). Our previous paper thoroughly explains the cholinesterase enzyme inhibition experiments we used in the current study9.The IC50 values of compound 1 were calculated via GraphPad Prism 6.01.
Ligand–protein docking simulations
Crystal structures of AChE and BChE were downloaded from Protein Data Bank (PDB)10—their PDB codes are 4EY711 and 5DYW12, respectively. Missing atoms in the backbones and sidechains were added, and the broken and disulfide bonds in the amino acids were fixed by Protein Preparation Wizard of the maestro of Schrödinger software13,14. The PROPKA tool15 was used for predicting the ionization of the systems and evolving it based on biological pH. The energies between the atoms are minimized to remove any potential clashes between the atoms, using OPLS3 force field through the classical minimization. The structure of the ligand was sketched and minimized in biological pH. Protein–ligand simulation was carried out using induced-Fit docking (IFD) method16,17,18, by which more flexibility is given to both ligand and the amino acids in the binding domain to refine themselves during the simulation. The coordinates of the co-crystal ligands inside the proteins were used to understand where to place the docking simulation box.
Conclusion
The plants are prolific producers of natural products. Natural products are lead drugs for treatment of many diseases. Many of the antibiotics are essentially natural products such as erythromycins, aminoglycosides and the antitumor antibiotics. Additionally, there is a huge number of their derivatives which were semisynthesized from the lead drugs of natural products such penicillins and cephalosporins. Compound 1 showed promising activity against acetylcholinesterase (AChE) and butyrylcholinesterase (BChE). These enzymes were associated with pathology of Alzheimer’s disease. Additionally, the absolute configuration of compound 1 were determined for the first time.
Data availability
The datasets generated and/or analyzed during the current study are available in The Cambridge Crystallographic Data Centre's CCDC 2161839; it may be downloaded for free at https://www.ccdc.cam.ac.uk/datarequest/cif. https://www.ccdc.cam.ac.uk/structures/Search?access=referee&ccdc=2161839&Author=Hazem+Ghabbour.
References
Elsebai, M. F., Abass, K., Hakkola, J., Atawia, A. R. & Farag, M. A. The wild Egyptian artichoke as a promising functional food for the treatment of hepatitis C virus as revealed via UPLC-MS and clinical trials. Food Funct. 7, 3006–3016 (2016).
Elsebai, M. F., Mocan, A. & Atanasov, A. Cynaropicrin: A comprehensive research review and therapeutic potential as an anti- hepatitis C virus agent. Front. Pharmacol. https://doi.org/10.3389/fphar.2016.00472 (2016).
BenSalem, M. et al. Pharmacological studies of artichoke leaf extract and their health benefits. Plant Foods Hum. Nutr. 70, 441–453 (2015).
Elsebai, M. F. et al. Pan-genotypic hepatitis C virus inhibition by natural products derived from the wild Egyptian artichoke. J. Virol. 90, 1918–1930 (2016).
Farag, M. A., Elsebai, M. F. & Khattab, A. R. Metabolome based classification of artichoke leaf: A prospect for phyto-equivalency of its different leaf origins and commercial preparations. J. Pharm. Biomed. Anal. 158, 151–159 (2018).
Elsebai, M. F. New pregnane glycoside derivative from Caralluma retrospiciens (Ehrenb). Nat. Prod. Res. 29, 1426–1431 (2015).
Elsebai, M. F. & Ghabbour, H. A. Isocoumarin derivatives from the marine-derived fungus Phoma sp. 135. Tetrahedron Lett. 57, 354–356 (2016).
Ellman, G. L., Courtney, K. D., Andres, V. Jr. & Featherstone, R. M. A new and rapid colorimetric determination of acetylcholinesterase activity. Biochem. Pharmacol. 7, 88–95 (1961).
Polumackanycz, M., Konieczynski, P., Orhan, I. E., Abaci, N. & Viapiana, A. Chemical composition, antioxidant and anti-enzymatic activity of golden root (Rhodiola rosea L.) commercial samples. Antioxidants 11, 919 (2022).
Berman, H. M. et al. The protein data bank. Nucleic Acids Res. 28, 235–242 (2000).
Cheung, J. et al. Structures of human acetylcholinesterase in complex with pharmacologically important ligands. J. Med. Chem. 55, 10282–10286 (2012).
Košak, U. et al. Development of an in-vivo active reversible butyrylcholinesterase inhibitor. Sci. Rep. 6, 39495 (2016).
Sastry, G. M., Adzhigirey, M., Day, T., Annabhimoju, R. & Sherman, W. Protein and ligand preparation: Parameters, protocols, and influence on virtual screening enrichments. J. Comput. Aided Mol. Des. 27, 221–234 (2013).
Maestro, S. Schrödinger Release 2021-2. LLC, New York, NY (2021).
Rostkowski, M., Olsson, M. H. M., Søndergaard, C. R. & Jensen, J. H. Graphical analysis of pH-dependent properties of proteins predicted using PROPKA. BMC Struct. Biol. 11, 6 (2011).
Farid, R., Day, T., Friesner, R. A. & Pearlstein, R. A. New insights about HERG blockade obtained from protein modeling, potential energy mapping, and docking studies. Bioorg. Med. Chem. 14, 3160–3173 (2006).
Sherman, W., Day, T., Jacobson, M. P., Friesner, R. A. & Farid, R. Novel procedure for modeling ligand/receptor induced fit effects. J. Med. Chem. 49, 534–553 (2006).
Sherman, W., Beard, H. S. & Farid, R. Use of an induced fit receptor structure in virtual screening. Chem. Biol. Drug Des. 67, 83–84 (2006).
Funding
Open access funding provided by The Science, Technology & Innovation Funding Authority (STDF) in cooperation with The Egyptian Knowledge Bank (EKB).
Author information
Authors and Affiliations
Contributions
M.F.E. outlined the work, analyzed and interpreted the data and wrote the wrote the draft manuscript and he the correspondence of submission. H.A.G. performed the X-ray experiments. R.E.S. performed the docking studies. I.E.O., and F.S.S.D. performed the enzyme inhibition experiments.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Elsebai, M.F., Ghabbour, H.A., Salmas, R.E. et al. X-ray crystalographic data, absolute configuration, and anticholinesterase effect of dihydromyricitrin 3-O-rhamnoside. Sci Rep 12, 18351 (2022). https://doi.org/10.1038/s41598-022-23240-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-23240-7
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.