Abstract
Real-time monitoring of cellular temperature responses is an important technique in thermal biology and drug development. Recent study identified that Na+/Ca2+ exchanger (NCX)-dependent Ca2+ influx transduces cold signals to circadian clock in mammalian cultured cells. The finding raised an idea that cellular responses to the cold signals can be analyzed by monitoring of clock gene expression. We found that Per1 and Per2 were up-regulated after culture at 27 °C compared to 37 °C in Rat-1 fibroblasts. In order to monitor cold-Ca2+-dependent transcription in living cells, we developed a luciferase-based real-time reporting system by using Per1 promoter, Per2 promoter, Ca2+/cAMP-response elements (CRE) or NFAT-binding elements. We found that benzyloxyphenyl NCX inhibitor KB-R7943 and SN-6, but not SEA-0400 or YM-244769 inhibited the cold induction of Per2. Our study established a real-time monitoring system for cold Ca2+ signaling which can be applied to evaluation of drugs.
Similar content being viewed by others
Introduction
Temperature is one of the most important factors for maintenance of cellular homeostasis1. In response to changes of ambient temperature, gene expression levels are globally influenced to alter various cellular process including metabolic activities and ion transport activities1,2. In molecular mechanism of temperature signaling, temperature-sensitive ion channels expressed in neuronal cells, such as transient receptor potential (TRP) channels, have been intensively studied in animals3. Importantly, even in non-neuronal cells of animals, ambient temperature largely affects cellular physiology3,4. In addition, clear temperature responses were observed among fungi5,6,7, plants8 and bacteria9, while the TRP channels are not present in these organisms. These studies implicate an existence of basic mechanism conserved for temperature response in a wide variety of organisms.
Among various temperature responses, the circadian clock is of particular interest because of its unique property, i.e., temperature-compensation of its period length10,11. Temperature compensation, observed in circadian clock of mammals, insects, fungi, plants and cyanobacteria, is a fundamental property of the cellular clock12,13,14,15. In mammals, the circadian rhythms are generated by transcriptional and translational feedback loops (TTFLs)11. Heterodimers of bHLH transcription factors CLOCK and BMAL1 bind to E-box DNA elements and activates thousands of genes including Per and Cry genes. The translated PERs bind to CRYs to inhibit the transcriptional activity of CLOCK-BMAL1. Thus, gene expression levels of Per genes show clear circadian rhythms16. Because most of the biochemical reactions in the TTFLs are slowed down by temperature decrease, it has been a mystery how circadian clock maintains its stable period length under different temperatures17.
Recently, we found that cytoplasmic Ca2+ signaling is activated by lowering temperature for compensation of the transcriptional oscillation18. In response to temperature decrease, Na+/Ca2+ exchanger (NCX) promotes Ca2+ influx, which activates Ca2+/calmodulin-dependent protein kinase II (CaMKII). CaMKII phosphorylates CLOCK to promote transcription of Per1/219, and the transcriptional activation of clock genes accelerates oscillation of transcriptional feedback loops. The study clarified that the circadian clock is highly responsive to ambient temperature, and Ca2+ signaling transduces the temperature information to the TTFLs. Importantly, NCX-dependent cold Ca2+ signaling is functionally conserved among mammals, Drosophila, Arabidopsis and cyanobacteria, indicating that the cold Ca2+ signaling is a general temperature response inherited from a common ancestor of the organisms18. Based on the study, we thought that development of a real-time monitoring system for the temperature response of the molecular clock is an important technique to study the conserved temperature signaling in living cells.
In the present study, we found that the expression levels of Per1 and Per2 were up-regulated after cold exposure at 27 °C compared to 37 °C. The cold-induced transcriptional changes can be monitored by luciferase reporter driven by Per1 and Per2 promotor in the living Rat-1 fibroblasts. Importantly, we found that a Ca2+-dependent transcriptional reporter including Ca2+/cAMP response elements (CRE)20 or nuclear factor of activated T-cells (NFAT)21 binding elements, also showed temporal increase of bioluminescence levels after temperature shift from 37 to 27 °C. By using the cellular monitoring system, we investigated effects of various NCX inhibitors on the cold induction of Per2. We found that KB-R7943 and SN-6, but not SEA-0400 or YM-244769 dose-dependently inhibited the cold induction of Per2. Importantly, the actions on the Per2 induction correlates with pharmacological actions on amplitude of the circadian rhythm especially at lower temperature. These results demonstrate that the real-time monitoring of Per2 expression levels is a novel method for detecting temperature responses in living cells, and the system enable us to evaluate bioactivities of chemical compounds for drug development.
Results
Real-time monitoring of bioluminescence rhythms by Per1, Per2 or Bmal1 reporter at different temperatures
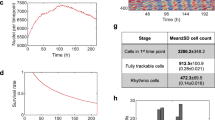
The previous studies revealed that expression levels of Per1 and Per2 are transiently induced by cytoplasmic Ca2+ increase both in the SCN and peripheral cells18,19,22. We found that temperature decrease chronically elevates cytoplasmic Ca2+ levels and Per1 and Per2 transcripts18 (Fig. 1). Based on the results, we thought that the cold responses of Per1 and Per2 genes could be monitored at transcriptional levels. We employed transcriptional reporters that express luciferase under the control of Per1 or Per2 promotor22. In addition, luciferase reporter of Bmal119 promoter was also analyzed as a negative control reporter, because the mRNA levels of Bmal1 showed no increase under lower temperature18. The cells were first synchronized with 1 h-pulse of 0.1 μM dexamethasone, and the medium was replaced with an air culture medium before measuring bioluminescence. In the mouse SCN and liver, Per1 and Per2 show expression rhythms that are anti-phasic to Bmal1 expression rhythms23. Consistent with the in vivo expression, bioluminescence rhythms driven by the luciferase reporter of Per1 or Per2 promoter were nearly anti-phasic to the rhythms by Bmal1 promotor in Rat-1 fibroblasts cultured at 37 °C (Fig. 2a). When the circadian rhythms were analyzed under 27 °C, we found that the clear bioluminescence rhythms driven by the all three reporters were severely damped (Fig. 2b–d). In addition, we found that the bioluminescence levels of Per1 and Per2 were markedly elevated. Then we analyzed bioluminescence levels by quantifying area under the curves (AUC) of the rhythms at both temperatures. Per1 or Per2 reporter showed 3.7-fold or 6.1-fold increase of the expression levels at 27 °C compared to 37 °C, respectively (Fig. 2e). On the other hand, the expression levels of Bmal1 reporter at 27 °C showed no significant change compared to 37 °C, consistent with the mRNA levels reported in our previous study18. These results indicate that Per1 and Per2 luciferase reporters were applicable to monitor cellular response to the cold exposure.
Effects of temperature on transcript levels of clock genes in Rat-1 fibroblasts. Relative mRNA level of Per1 or Per2 after 2-day culture at 37 °C or 27 °C. Mean with s.e.m. from 3 independent samples are shown. The expression levels at 37 °C were set to 100 in vertical axis. ** p < 0.01 compared to the data of 37 °C (Student’s t-test).
Effects of temperature on transcriptional reporters of clock genes. (a) Representative bioluminescence rhythms driven by Per1-luc, Per2-luc, or Bmal1-luc reporter. The first peak is normalized to 1 in the vertical axis. The rhythms were smoothing by 1-h centered moving average for clearness to compare the phase of each reporter. Representative data of bioluminescence rhythms of Per1-luc (b), Per2-luc (c), and Bmal1-luc (d) at 37 °C or 27 °C. (e) Relative bioluminescence levels of Per1, Per2, or Bmal1 calculated by the area under the curve (AUC) from Day 0 to Day 3. The means of AUC at 37 °C were set to 1 in vertical axis. Mean with s.e.m. from 4 independent samples are shown. *** p < 0.001 compared to the data at 37 °C (Student’s t-test).
Ca2+-dependent transcription is activated by cold exposure
The significant cold induction of bioluminescence levels driven by the Per1 and Per2 transcriptional reporters prompted us to investigate acute responses of the reporters to temperature shift from 37 to 27 °C in detail (Fig. 3). The cells were first cultured at 37 °C for 2 days, and then kept at 27 °C in the same luminescence detector for 3 days. For the control, the bioluminescence signals were monitored at 37 °C for 5 days (Fig. S1). We found that the bioluminescence rhythms driven by the Per1, Per2 and Bmal1 reporters observed at 37 °C was acutely damped at 27 °C (Fig. 3a–c), similar to the results of constant condition at 27 °C (Fig. 2). We analyzed effects of temperature on bioluminescence levels by calculating the AUC during Term I and II (as depicted in Fig. 3a). We found that the AUC of Per1 reporter increased to 137% or 162% in Term I or II compared to control area of 37 °C, respectively (Fig. 3a and f). Similarly, the AUC of Per2 reporter was increased to 153% (Term I) or 204% (Term II), whereas that of Bmal1 reporter was decreased to 46% (Term I) or 29% (Term II) (Fig. 3b,c,f). These results indicate that the transcriptional reporters of Per1 and Per2 can be used to monitor temporal responses to the temperature shift.
Effects of temperature shift on transcriptional reporters. Representative data of bioluminescence level of Per1 (a), Per2 (b), Bmal1 (c), CRE (d), or NFAT (e) luciferase reporters in temperature shift experiment. The reporter cells were cultured at 37 °C for 2 days and then temperature was decreased to 27 °C during the rest of the experiment. (f) Effect of temperature decrease from 37 to 27 °C on bioluminescence signals of reporter cell lines. The induction rate was calculated by dividing the AUC during Term I or Term II (27 °C) by the AUC during day 1 to 2 (control, 37 °C) (marked as example in panel a). The means of induction rate at 37 °C was set to 100 in vertical axis. Mean with s.e.m. from 4, 8, 8, 4 or 3 independent samples for Per1, Per2, Bmal1, CRE, or NFAT luciferase reporter cells are shown, respectively. p values: one-way ANOVA among induction rate in Term I, II and control. * p < 0.05, ** p < 0.01, *** p < 0.001, **** p < 0.0001 compared to the data at 37 °C (Dunnett’s test).
We next investigated whether the cold response was observed in a Ca2+-dependent transcriptional reporter including Ca2+/cAMP response elements (CRE) or nuclear factor of activated T cell (NFAT)-binding elements. We found that temperature shift from 37 to 27 °C increased bioluminescence levels of the CRE reporter only during Term I, suggesting that the CRE reporter shows a transient response to the cold exposure (Fig. 3d and f). In contrast, the NFAT reporter, which reflects activation of Ca2+/calcineurin-NFAT signaling, showed a constant increase of the AUC in both Term I and II (Fig. 3e and f). These results demonstrate that CRE and NFAT-dependent transcriptional reporters are applicable to monitor cold Ca2+ signaling in addition to the Per1 and Per2 transcriptional reporters.
Effects of benzyloxyphenyl NCX inhibitors on cold Per2 induction
Based on the results of cold responses observed in the transcriptional reporter cell lines, we thought that the system might be useful to evaluate effects of chemical inhibitors on NCX-dependent cold Ca2+ signaling. First, we tested effects of small molecule NCX inhibitor KB-R794324 (Fig. 4a), which blocks cold induction of cytoplasmic Ca2+ and Per2 mRNA levels18. We found that KB-R7943 blocked the cold induction of bioluminescence levels of Per2 reporter in a dose-dependent manner (Fig. 4b and c). In addition, we investigated the effect of KB-R7943 in cold induction by using Rat-1 Per1-luc reporter cells. Consistent with the results observed in Per2-luc cells, KB-R7943 dose-dependently inhibited the cold induction of Per1 (Fig. S2a and b). Next, we evaluated other benzyloxyphenyl NCX inhibitors SN-6, SEA0400 and YM-24476925,26,27(Fig. 4a). SN-6 dose-dependently inhibited the cold induction of Per2 similar to KB-R7943 (Fig. 4b and c). On the other hand, SEA0400 and YM-244769 showed no significant inhibitory effects on the Per2 induction (Fig. 4b and c). Because KB-R7943 and SN-6 have relatively small functional groups compared to SEA0400 and YM-244769 (Fig. 4a), it is possible that small functional groups on benzyloxyphenyl skeleton are important for the pharmacological activity on the cold induction of Per2 reporter.
Effects of NCX inhibitors on cold Per2 induction. (a) Structures of NCX inhibitors, KB-R7943, SN-6, SEA0400, and YM-244769. Yellow background shows a common structure of the four inhibitors. (b) Effects of NCX inhibitors on cold induction of Per2-luc reporter. (c) Effects of NCX inhibitors on induction rate of Per2-luc reporter. The induction rate was calculated by dividing the AUC during Term II (27 °C) by the AUC during day 1 to 2 (control, 37 °C) (marked as example in Fig. 3a). Because 20 µM YM-244769 showed cell toxicity, the data of that was not determined (n.d.). The means of induction rate of DMSO control group at 37 °C was set to 100 in vertical axis. Mean with s.e.m. from 4 independent samples are shown. * p < 0.05 compared to the DMSO control at 27 °C (Dunnett's test).
Correlation analysis of Per2 induction and circadian rhythm parameters
In order to understand pharmacological actions of the four different benzyloxyphenyl NCX inhibitors on circadian rhythms, we accessed bioluminescence rhythms of Rat-1 Bmal1-luc cells cultured at 32 °C or 37 °C with or without 10 µM NCX inhibitors (Fig. 5a–d and Fig. S3). When the effects of the inhibitors on period length at 37 °C were analyzed, KB-R7943 and SN-6 significantly shortened the period compared to DMSO control group (Fig. 5e). On the other hand, at 32 °C, all the inhibitors showed no significant effect on the period (Fig. 5f). Next, we analyzed the effects of the inhibitors on amplitude of the bioluminescence rhythms. We found that all the inhibitors significantly decreased the amplitude of the circadian rhythms at 37 °C (Fig. 5g). On the other hand, at 32 °C, only KB-R7943 and SN-6 reduced the amplitude (Fig. 5h). These results demonstrate that KB-R7943 and SN-6 have strong effects especially at lower temperature.
Correlation analysis between cold Per2 induction and circadian rhythm parameters. Representative data of the effect of KB-R7943 (a), SN-6 (b), SEA0400 (c), or YM-244769 (d) on relative bioluminescence rhythms of Rat-1 Bmal1-luc cells. Effects of 4 NCX inhibitors on period length of bioluminescence rhythms at 37 °C (e) or 32 °C (f). Effects of 4 NCX inhibitors on amplitude at 37 °C (g) or 32 °C (h). Correlation analysis of cold Per2 induction and period length at 37 °C (i) or 32 °C (j). Correlation analysis of cold Per2 induction and amplitude at 37 °C (k) or 32 °C (l). Mean with s.e.m. from 4 independent samples are shown (e–h). * p < 0.05, ** p < 0.01, *** p < 0.001 compared to the DMSO control (Student’s t-test, with Bonferroni correction). Mean from 4 independent samples are shown (i–l).
Finally, we analyzed the relationships between the circadian rhythm parameters (Fig. 5a–h) and the cold Per2 induction (Fig. 4). Correlation analysis demonstrated that effects on the period length showed a correlation with the cold induction of Per2 (Fig. 5i and j). Importantly, effects on amplitude at 32 °C showed strong correlation with the cold induction of Per2, whereas effects on amplitude at 37 °C showed no correlation (Fig. 5k and l). Previously, we identified that the NCX-dependent Ca2+ signaling is activated at low temperature to increase amplitude of the cellular rhythms18. The inhibitory effects of NCX inhibitors on cold-Per2 induction was strongly correlated with the effects on amplitude of the cellular rhythms especially at lower temperature (Fig. 5i), suggesting that the cold Per2 induction reflects magnitude of the cold Ca2+ response in cells. In conclusion, the real-time monitoring of cold-Per2 induction can be used as a system for detecting cold Ca2+ response.
Discussion
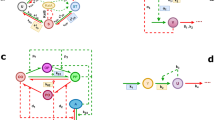
Ca2+ signaling have been intensively studied in input pathway of the circadian clock28,29. In the SCN, light signals are transmitted by glutamate which activates NMDA receptor to promote Ca2+ influx30. The intracellular Ca2+ activates MAPK and CaMKII to phosphorylate transcriptional factor CREB. Because promotor regions of Per1 and Per2 contain CRE22,28,29, Per1 and Per2 are increased immediately after light exposure during subjective night in the SCN31,32,33. Consistent with the photic response, we found that the CRE-dependent transcription is transiently activated by temperature shift from 37 to 27 °C in Rat-1 fibroblasts. In contrast, the Per1 or Per2 reporter showed continuous increase after the cold exposure (Fig. 3). The promoter regions of Per1 and Per2 contain E-box and D-box sequences34, in addition to the CRE. We found that D-box reporter also showed a transient increase of bioluminescence level in response to the temperature decrease (Fig. 6a–c). It is possible that cooperative actions of CRE, E-box and D-box are important for the long-term response of Per1 and Per2 to the cold exposure (Fig. 6d).
Effect of temperature shift on D-box reporter. (a) Representative bioluminescence rhythm of D-box luciferase reporter in temperature shift experiment. (b) Representative rhythms of D-box reporter cell line at 37 °C. (c) Induction rate of D-box reporter cell line. Mean with s.e.m. from 8 independent samples are shown. p values: one-way ANOVA among induction rate in Term I, II and control. ** p < 0.01 compared to the data at 37 °C (Dunnett’s test) (d) Schematic figure of transcriptional regulation of Per1/2. Transcription of Per1/2 is regulated by cAMP response element (CRE), E-box (only one is shown here), and D-box. CRE is essential for various signaling pathways such as cAMP, Ca2+ and Ras, while E-box is the target of CLOCK/BMAL1 regulation, and D-box are regulated by E4BP4 and DBP.
Cold-Ca2+ signaling was originally found in studies of cold tolerance in plants35,36,37. Interestingly, we found that the cold Ca2+ signaling is conserved among bacteria, plants, insects and mammals, indicating that the Ca2+ signal is an ancestral mechanism for cold response. Because the intracellular Ca2+ levels continuously increased during cold exposure in mammalian cells18, the cold Ca2+ signal is unique compared to general Ca2+ signaling that shows a transient increase. Therefore, we think that long term investigation of the cold-Ca2+ signaling in living cells is very important for understanding of fundamental temperature responses. In this study, we attempted to monitor cold Ca2+-dependent transcription by using Per2-luc stable expression cell line. By using this system, we investigated the efficacies of NCX inhibitors. Benzyloxyphenyl NCX inhibitors are thought to interact with a specific receptor site in α-2 loop of NCX1 proteins to block ion transport pore(s)38. KB-R7943 or SN-6, which showed strong effects on Per2 response, has a smaller functional group on the benzene ring compared to SEA-0400 and YM-244769 (Fig. 4a). Therefore, it is possible that bulky functional groups on the benzene ring in SEA-0400 and YM-244769 interfere access of the inhibitor to the putative receptor sites in NCX proteins. In conclusion, we believe that this system can be a powerful tool for investigating temperature response in living cells.
Methods
RT-qPCR experiments by using Rat-1 fibroblasts
The Rat-1 fibroblasts were purchased from American Type Culture Collection (ATCC). The cells were plated on 35-mm dishes (1.0 × 106 cells/dish) and cultured at 37 °C under 5% CO2 in a culture medium [DMEM (Sigma-Aldrich, catalog no. 5796) supplemented with 10% FBS (Biosera), 50 U/ml penicillin and 50 μg/ml streptomycin]. One day after the plating, the medium was changed to air culture medium [DMEM (Sigma-Aldrich, catalog no. D2902) supplemented with 10% FBS, 3.5 mg/ml glucose, 25 U/ml penicillin, 25 μg/ml streptomycin and 10 mM HEPES–NaOH (pH 7.0)], and the cells were cultured at 37 °C or 27 °C under air. The cells were collected on day 2 by using 600 μl TRIzol (Invitrogen). Total RNA was prepared from cultured cells using RNeasy Kit (Qiagen) according to the manufacturer’s protocol. RT-qPCR analysis was performed as described previously18,39.
Real-time monitoring of gene expression rhythms in mammalian cells
Real-time monitoring of gene expression rhythms in mammalian cells was performed by using Rat-1 fibroblasts stably expressing luciferase driven by Per1 promoter, Per2 promoter22, Bmal1 promoter19, D-box elements34, CRE or NFAT binding elements. The CRE reporter or NFAT reporter was constructed by inserting 4 repeats of CREB-binding sequences (AGCCTGACGTCAGAG) or 4 repeats of NFAT-binding sequences (GGAGGAAAAACTGTTTCATACAGAAGGCGT) into pGL3-Basic vector (Promega). The fibroblasts were plated on 35-mm dishes (1.0 × 106 cells/dish) and cultured at 37 °C under 5% CO2 in the culture medium [DMEM (Sigma-Aldrich, catalog no. 5796) supplemented with 10% FBS (Biosera), 50 U/ml penicillin and 50 μg/ml streptomycin]. One day after the plating, the cells were treated with 0.1 μM dexamethasone for 1 h, and the medium was replaced with the air culture medium [DMEM (Sigma-Aldrich, catalog no. D2902) supplemented with 10% FBS, 3.5 mg/ml glucose, 25 U/ml penicillin, 25 μg/ml streptomycin, 0.1 mM luciferin and 10 mM HEPES–NaOH (pH 7.0)]. The bioluminescence signals were continually recorded from the cells cultured under air in a dish-type bioluminescence detector, LumiCycle (Actimetrics). In the temperature shift experiment, the temperature was first set to 37 °C for 2 days and then set to 27 °C for the rest of the measurement.
For normalization of dish-to-dish variation of the bioluminescence levels, the raw data were divided by mean bioluminescence signals recorded for 6 days. The normalized rhythms were detrended by subtracting 24-h centered moving averages, and the average of second, third and fourth peak and trough were used for calculating the amplitudes of the rhythms. Period length was calculated using the average value of peak-to-peak periods and trough-to-trough periods one day after the dexamethasone treatment of cultured cells.
Data availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Storey, K. B. & Storey, J. M. Molecular physiology of freeze tolerance in vertebrates. Physiol. Rev. 97, 623–665 (2017).
Boothby, T. C. Mechanisms and evolution of resistance to environmental extremes in animals. EvoDevo 10, 30 (2019).
Buijs, T. J. & McNaughton, P. A. The role of cold-sensitive ion channels in peripheral thermosensation. Front Cell Neurosci. 14, 262 (2020).
Vriens, J., Nilius, B. & Voets, T. Peripheral thermosensation in mammals. Nat. Rev. Neurosci. 15, 573–589 (2014).
Rensing, L. & Ruoff, P. Temperature effect on entrainment, phase shifting, and amplitude of circadian clocks and its molecular bases. Chronobiol. Int. 19, 807–864 (2002).
Liu, Y. et al. Thermally regulated translational control of FRQ mediates aspects of temperature responses in the neurospora circadian clock. Cell 89, 477–486 (1997).
Liu, Y. et al. How temperature changes reset a circadian oscillator. Science 281, 825–829 (1998).
Yuan, P. G. et al. Calcium signaling-mediated plant response to cold stress. Int. J. Mol. Sci. 19, 1–11 (2018).
Zhang, Y. & Gross, C. A. Cold shock response in bacteria. Annu. Rev. Genet. 55, 377–400 (2021).
Dunlap, J. C. Molecular bases for circadian clocks. Cell 96, 271–290 (1999).
Buhr, E. D. & Takahashi, J. S. Molecular components of the mammalian circadian clock. Handb. Exp. Pharmacol. 217, 3–27 (2013).
Dunlap, J. C. & Loros, J. J. Making time: Conservation of biological clocks from fungi to animals. Microbiol. Spectr. 5, 3 (2017).
Takahashi, J. S. Transcriptional architecture of the mammalian circadian clock. Nat. Rev. Genet. 18, 164–179 (2017).
Nohales, M. A. & Kay, S. A. Molecular mechanisms at the core of the plant circadian oscillator. Nat. Struct. Mol. Biol. 23, 1061–1069 (2016).
Cohen, S. E. & Golden, S. S. Circadian rhythms in cyanobacteria. Microbiol. Mol. Biol. Rev. 79, 373–385 (2015).
Yoo, S. H. et al. PERIOD2::LUCIFERASE real-time reporting of circadian dynamics reveals persistent circadian oscillations in mouse peripheral tissues. Proc. Natl. Acad. Sci. U S A 101, 5339–5346 (2004).
Hastings, J. W. & Sweeney, B. M. On the mechanism of temperature independence in a biological clock. Proc. Natl. Acad. Sci. U S A 43, 804–811 (1957).
Kon, N. et al. Na+/Ca2+ exchanger mediates cold Ca2+ signaling conserved for temperature-compensated circadian rhythms. Sci. Adv. 7, eabe8132 (2021).
Kon, N. et al. CaMKII is essential for the cellular clock and coupling between morning and evening behavioral rhythms. Genes Dev. 28, 1101–1110 (2014).
Dash, P. K. et al. cAMP response element-binding protein is activated by Ca2+/Calmodulin as well as cAMP-dependent protein kinase. Proc. Natl. Acad. Sci. U S A 88, 5061–5065 (1991).
Park, Y. J. et al. The role of calcium-calcineurin-NFAT signaling pathway in health and autoimmune diseases. Front. Immunol. 11, 1–14 (2020).
Travnickova-Bendova, Z., Cermakian, N., Reppert, S. M. & Sassone-Corsi, P. Bimodal regulation of mPeriod promoters by CREB-dependent signaling and CLOCK/BMAL1 activity. Proc. Natl. Acad. Sci. U S A 99, 7728–7733 (2002).
Ueda, H. R. et al. System-level identification of transcriptional circuits underlying mammalian circadian clocks. Nat. Genet. 37, 187–192 (2005).
Iwamoto, T. & Shigekawa, M. Differential inhibition of Na+/Ca2+ exchanger isoforms by divalent cations and isothiourea derivative. Am. J. Physiol. 275, C423–C430 (1998).
Iwamoto, T. et al. Molecular determinant of Na+/Ca2+ exchange (NCX1) inhibition by SEA0400. J. Biol. Chem. 279, 7544–7553 (2004).
Iwamoto, T. et al. The exchanger inhibitory peptide region-dependent inhibition of Na+/Ca2+ exchange by SN-6 [2-[4-(4-nitrobenzyloxy) benzyl] thiazolidine-4-carboxylic acid ethyl ester], a novel benzyloxyphenyl derivative. Mol. Pharmacol. 66, 45–55 (2004).
Iwamoto, T. & Kita, S. YM-244769, a novel Na+/Ca2+ exchange inhibitor that preferentially inhibits NCX3, efficiently protects against hypoxia/reoxygenation-induced SH-SY5Y neuronal cell damage. Mol. Pharmacol. 70, 2075–2083 (2006).
Tischkau, S. A. et al. Ca2+/cAMP response element-binding protein (CREB)-dependent activation of Per1 is required for light-induced signaling in the suprachiasmatic nucleus circadian clock. J. Biol. Chem. 278, 718–723 (2003).
Shigeyoshi, Y. et al. Light-induced resetting of a mammalian circadian clock is associated with rapid induction of the mPer1 transcript. Cell 91, 1043–1053 (1997).
Gooley, J. J., Lu, J., Chou, T. C., Scammell, T. E. & Saper, C. B. Melanopsin in cells of origin of the retinohypothalamic tract. Nat. Neurosci. 4, 1165 (2001).
Albrecht, U., Sun, Z. S., Eichele, G. & Lee, C. C. A differential response of two putative mammalian circadian regulators, mPer1 and mPer2, to light. Cell 91, 1055–1064 (1997).
Shearman, L. P., Zylka, M. J., Weaver, D. R., Kolakowski, L. F. Jr. & Reppert, S. M. Two period homologs: Circadian expression and photic regulation in the suprachiasmatic nuclei. Neuron 19, 1261–1269 (1997).
Wilsbacher, L. D. et al. Photic and circadian expression of luciferase in mPeriod1-luc transgenic mice in vivo. Proc. Natl. Acad. Sci. U S A 99, 489–494 (2002).
Yoshitane, H. et al. Functional D-box sequences reset the circadian clock and drive mRNA rhythms. Commun. Biol. 2, 300 (2019).
Pomeroy, M. K. & Chris, J. A. Effect of low temperature and calcium on survival and membrane properties of isolated winter wheat cells. Plant Physiol. 78, 484–488 (1985).
Mori, K. et al. Ca2+-permeable mechanosensitive channels MCA1 and MCA2 mediate cold-induced cytosolic Ca2+ increase and cold tolerance in Arabidopsis. Sci. Rep. 8, 550 (2018).
Wilkins, K. A., Matthus, E., Swarbreck, S. M. & Davies, J. M. Calcium-mediated abiotic stress signaling in roots. Front. Plant Sci. 7, 1296 (2016).
Iwamoto, T. Sodium-calcium exchange inhibitors: Therapeutic potential in cardiovascular diseases. Future Cardiol. 1, 519–529 (2005).
Kon, N. et al. Activation of TGF-β/activin signaling resets the circadian clock through rapid induction of Dec1 transcripts. Nat. Cell Biol. 10, 1463–1469 (2008).
Acknowledgements
We are grateful to members of laboratory of integrative animal physiology and members of Transformative Research Area “Hibernation biology” for helpful discussion. This work is supported in part by Japanese Society for the Promotion of Science (JSPS) Grants-in-Aid for Scientific Research (KAKENHI) to N. K. (18H06066, 20H03292 and 20H05796) and T. Y. (19H05643). N. K. was supported by Suntory Rising Stars Encouragement Program in life Sciences (SunRiSE), Tomizawa Jun-ichi & Keiko Fund of Molecular Biology Society of Japan for Young Scientist, DAIKO FOUNDATION, Research Foundation for Opto-Science and Technology, Kato Memorial Bioscience Foundation, Pharmacodynamics Research Foundation and Academic Research & Industry-Academia-Government Collaboration of Nagoya University. H. W. is supported by JSPS Research Fellowship for Young Scientists.
Author information
Authors and Affiliations
Contributions
N.K. and H.W. conceived the experiments. H.W conducted all of the cell experiments. S.M., M.K. and K.I. conducted part of the temperature shift experiments in cells. Y.A. gave technical advice for gene expression analysis. H.W. performed statistical analysis and figure generation. T.Y. provided a part of laboratory equipment and financial supports. N.K. and H.W. wrote the manuscript, and all authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, Ht., Miyairi, S., Kitamura, M. et al. Real time monitoring of cold Ca2+ dependent transcription and its modulation by NCX inhibitors. Sci Rep 12, 17325 (2022). https://doi.org/10.1038/s41598-022-22166-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-22166-4
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.