Abstract
Extracellular polysaccharide (EPS) produced by a deep-sea, psychrotolerant Bacillus altitudinis SORB11 was evaluated by considering physiochemical nature and structural constituents. The productivity of crude EPS was measured ~ 13.17 g L−1. The surface topography of the crude EPS showed a porous, webbed structure along with a branched coil-like configuration. The crystalline crude EPS contained a high amount of sulfur. Further, the crude EPS was subjected for purification. The molecular weight of purified EPS was determined ~ 9.8 × 104 Da. The purified EPS was appeared to show glucomannan-like configuration that is composed of → 4)-β-Manp-(1 → and → 4)-β-Glcp-(1 → residues. So, this polysaccharide was comparable to the structure of plant-derived glucomannan. Subsequently, EPS biosynthesis protein clusters like EpsC, EpsD, EpsE, and glycosyltransferase family proteins were predicted from the genome of strain SORB11, which may provide an insight into the production of glucomannan-type of polysaccharide. This low molecular weight linear form of glucomannan-type EPS might be involved to form a network-like unattached aggregation, and helps in cell-to-cell interaction in deep-sea microbial species.
Similar content being viewed by others
Introduction
Microbes inhabiting extreme environments often secret extracellular polysaccharides with distinctive features and these polysaccharides eventually protects microorganism from life-threatening conditions1,2. Extracellular polysaccharides with diverse functions participate in biofilm formation to construct a comfortable extracellular environment, adhesion, molecular recognition, intracellular signal transduction, and sometimes pathogenesis3,4. As a substantial component and a major part of reduced carbon reservoir in the ocean, extracellular polysaccharides influenced by altering the physical and biogeochemical microenvironment around the cells. For unattached aggregation in a marine environment, polysaccharides with proteins, lipids, and nucleic acids form an architectural matrix4,5,6,7,8,9,10. In an aqueous solution, exopolysaccharides showed their hydrophilic nature due to the presence of hydroxyl and carboxyl groups that confer a net negative charge, thus providing acidic properties4,5. Bacterial exopolysaccharides are found in different forms such as biofilms, aggregated in a matrix or in dissolved forms by forming weak interactions with other organic and inorganic materials that provide hydrogen bonding, electrostatic interactions, dispersion as well as cohesive forces. However, these weak interactions depend on the polymer size and frequency of the functional groups. Furthermore, cross-linking within adjacent polymer chains influenced by electrolyte concentration enables permanent attachment11,12. The hydrated extracellular polysaccharide matrix creates a buffer zone to provide stability in adjacent changing environments and helps in localization of secreted exoenzymes for the cycling of organic and inorganic materials12,13. Highly hydrated porous polysaccharide matrix acts as a sponge that traps nutrients in flowing liquids. High polyhydroxyl containing extracellular polysaccharides provide buffering in low-temperature, high-salinity4,14,15.
In the Southern Ocean region, highly concentrated and varied microbial communities amalgamate to form transparent exopolysaccharide particles (TEPs) that aggregate with other organic debris and produce larger particles known as “marine snow” that serves as a vertical exporter of fixed carbons from the euphotic zone to deep waters4,16. Bacterial extracellular polysaccharides in marine environment act as glue and form a fibrillar framework that provides opportunities for interactions with different free-living bacterial cells, as well as nutrient uptake. Extracellular polysaccharides produced by bacterial populations from particulate material of Southern Ocean are often decorated with sulfate, acetyl, and succinyl groups, all of these provide polyanionic quality due to the ionization of acidic groups at pH of seawater environment4,17,18,19.
The investigation of extracellular polysaccharides produced by bacterial communities from deep-sea ecosystems provided a glimpse of how biomolecules were stabilized in harsh environmental conditions in spite of their biotechnological interest13,14,20,21,22. Therefore, in this present study, we explored the morphological, and physical properties of crude extracellular polysaccharide (EPS) and the structural configuration of purified EPS produced by a deep-sea, psychrotolerant bacterium Bacillus altitudinis SORB11, which was previously reported from 3800 m deep Indian sector of the Southern Ocean23. Moreover, In-silico analysis of coding gene clusters responsible for EPS biosynthesis was also performed using the genome of strain SORB11.
Results and discussions
Production and characterization of EPS
Free-living marine bacterial cells usually get aggregated with each other. Consequently, a fibrillar network is formed by secreting extracellular polysaccharides, and this ultrafine structure is assumed for cell-to-cell communication and to uptake nutrition from the environment4,24. A peculiar network formation (Fig. 1a) was observed between the colonies of B. altitudinis SORB11. Cells were rod-shaped bacilli, elongated with a diameter of ~ 1.0 µm and length of ~ 2.5 µm (Fig. 1b). Deposition of polysaccharide-like substances surrounding the cells (Fig. 1b) as well as polysaccharide precipitation was observed in the cell culture of strain SORB11 (Fig. 1c).
The extracted lyophilized EPS appeared as creamish white in color (Fig. 1d), it was soluble in polar solvents like water and dimethyl sulfoxide: dissolved ~ 1.0 g of crude EPS in 100 mL of water at 25 °C. Under optimum growth conditions (20 °C, pH level 8.00), ~ 13.17 g L−1 dry weight of crude EPS (Fig. 1e) was obtained using ethanol as a solvent. A total of 89.23% sugar component was present in the polysaccharide, with a 1.79% protein, and was devoid of glucuronic acid (Fig. 1f).
A heterogeneous type of surface morphology was exhibited by the crude EPS. The dried EPS powder showed porous surface topology in scanning electron micrographs (Fig. 2a–c). Whereas, a webbed configuration was mostly detected along with a long-branching patterns as well as, formation of coil-like structures of the EPS in solution at high magnification in Field emission scanning electron micrographs (Fig. 2d–i). In Atomic force micrographs (Fig. 2j–l), EPS showed ~ 4.3 µm long branching patterns at 50.0 µg/mL solution. But at 10.0 µg/mL and 5.0 µg/mL concentrations, the branched EPS was scattered and formed chain-like molecular structures (~ 200 nm long and ~ 10–25 nm diameter). Scattered molecules showed both impregnated as well as spherical surface topologies. Polysaccharides usually form disordered random coils with flexible chain-like configurations in an aqueous solution25,26. In marine environment, B. altitudinis SORB11 secrets a porous polysaccharide matrix that may helps to trap nutrients from flowing liquids. Moreover, adjacent polymer chains may cross-link between them to enable permanent attachment with the so-called “marine snow”4.
SEM images at magnifications 1.00 k × (a), 5.00 k × (b), and 10.00 k × (c) of lyophilized crude EPS; FE-SEM images at magnifications 500 × (d), 2.00 k × (e–f), 2.50 k × (g), 4.00 k × (h), and 10.00 k × (i) of 1.0 mg/mL crude EPS in solution; planar and 3-D AFM images representing 50.0 µg/mL (j), 10.0 µg/mL (k), and 5.0 µg/mL (l) concentrations of crude EPS in solution.
The key non-metal elements of crude EPS were carbon, oxygen, and sulfur as revealed from EDX spectrum (Fig. 3a). The high content of sulfate was attributed to the peak at 168 eV which may responsible for the polyanionic nature of crude EPS4,15. That result was further confirmed by elemental analysis where with 25.19 w% carbon was the dominant element followed by 5.95 w% hydrogen. The presence of 2.95 w% nitrogen indicated the protein contamination which was confirmed earlier by the protein assay. The carbon/nitrogen ratio was 8.54. A fair amount of sulfur (9.36 w%) was also detected in the crude form of EPS (Fig. 3b).
X-ray diffraction pattern of crude EPS showed a major crystalline region from 15° to 40° (Fig. 3c). The diffraction pattern exhibited several intense narrow crystalline diffraction peaks at 16°, 21°, 29°, and 32° in the 2θ region that revealing the crystalline property of crude EPS27.
Thermogravimetric analysis was conducted dynamically among weight loss versus temperature (Fig. 3d). The EPS was stable over a wide range of temperatures. A major weight loss was recorded between 250 to 300 °C. So, the crude form of EPS was thermally stable up to ≥ 250 °C.
A standard plant-derived glucomannan (Fig. 4a) and B. altitudinis SORB11 derived crude EPS (SORB-EPS) (Fig. 4b) were analyzed by IR spectra to detect functional groups and band assignments. For both polysaccharides, the absorption band that appeared in the range of 1000 − 1200 cm−1 indicated the presence of β-(1 → 4) linkage, but an adjacent peak in the range of 1031–1093 cm−1 that appeared only for glucomannan was due to the presence of (1 → 3) linked β-glucan25,28,29. Peaks at 3274 cm−1 and 2921 cm−1 were assigned to the O–H and C–H stretching and bending vibrations, respectively. The peak at 1638 cm−1 appeared due to the C=O stretching vibration of the protonated carboxylic acid or N-acetyl bonding27,30. Only in SORB-EPS, typical peaks that appeared at around 1220 cm−1 and 827 cm−1 were due to the stretching vibration of asymmetric S=O and symmetric C–O–S, respectively17,31,32,33. The presence of a significant amount of sulfur in SORB-EPS was previously confirmed by elemental and EDX analysis.
Crude SORB-EPS and standard glucomannan were purified with Gel-chromatographic technique. Fractions from major peaks were collected and lyophilized to get homogeneous polysaccharides in pure form (Fig. 5a, b). The molecular weight (Mw) of standard glucomannan was calculated ~ 1.50 × 106 Da. Species of Amorphophallus konjac derived k-glucomannan generally represented with a Mw ~ 1.32 × 106 Da34. Whereas, Mw of the major fraction i.e., fraction-I of SORB-EPS was calculated to be ~ 9.8 × 104 Da using dextrans as standard and was subjected for further characterization.
Monosaccharide analysis of standard glucomannan reflected the presence of mannose and glucose in a molar ratio of 1.64:1.00. Similarly, SORB-EPS showed a similar monosaccharide profile like glucomannan i.e., a heteropolysaccharide composed of mannose and glucose but with a different molar ratio of 8.86:1.00 (Fig. 5c). Therefore, the above results indicated the SORB-EPS was somewhat similar to that of plant-derived glucomannan35. However, it is an established fact that the ratio of mannose and glucose may vary depending on the origin of glucomannan36.
Methylation analysis of glucomannan represented 4-linked-D-mannopyranosyl and 4-linked-D-glucopyranosyl as the most abundant residues (Sup file 1). The presence of 3-linked-D-glucopyranosyl, as well as terminal residues, was also detected (Table 1). K-glucomannan derived from Amorphophallus konjac is a straight-chain polymer composed of β-(1 → 4)-linked D-mannose and D-glucose with a slide branching through β-(1 → 3)-glucosyl linkages34.
Likewise in the purified SORB-EPS, the most abundant residues were terminal and 4-linked-D-mannopyranosyl (Table 1). A 4-linked-D-glucopyranosyl residue was also identified but no such 3-linked branching pattern like plant-derived glucomannan was detected. In this way, the EPS produced by B. altitudinis SORB11 was linear and composed of β-(1 → 4)-linked D-mannose and β-(1 → 4)-linked D-glucose that described a simplified structure of glucomannan.
The purified SORB-EPS was further subjected to proton and carbon NMR spectroscopy and the signals of mannose and glucose units according to their structures as well as the sequence of sugar units were compared with glucomannan stated in the literature25,37,38,39,40,41,42,43,44,45,46,47,48. Signals at 1.956 ppm, 1.990 ppm, and 2.197 ppm were assigned as methyl protons (CH3) of the acetyl group. Other signals like 5.090 ppm and 5.023 ppm were assigned as C-1-linked hydrogens of mannose and glucose, respectively. Overlapped signals from 3.302 to 4.098 ppm attributed to the hydrogens linked to C-2 to C-6 of both mannose and glucose. Signals at 4.194 ppm and 5.269 ppm indicated the position of acetyl groups at C-2, C-3, and C-6 like acetyl polysaccharides (Fig. 6a).
Carbon signals measurement was carried out with an overnight run, but S/N ratio was not that high may be due to the low concentration of the solution. The resonances at around 20.0 ppm to 25.0 ppm revealed the abundance of O-acetyl groups in SORB-EPS. The shifts at 103.0 ppm and 104.0 ppm were due to the C-1 anomeric resonance of 4-linked glucopyranosyl and at 101.0 ppm due to the C-1 anomeric resonance of 4-linked mannopyranosyl residues. The C-4 down-field shifts of 4-linked glucopyranosyl appeared at 77.0 ppm and 4-linked mannopyranosyl units at 79.0 ppm were due to the involvement of glycosidic linkages. The typical resonances of C-5, C-3, and C-2 of β-1, 4-linked glucopyranosyl residues were observed at 76.0 ppm, 75.0 ppm, and 74.0 ppm, respectively. The signals for C-5, C-3, and C-2 of 4-linked mannopyranosyl residues were observed at 76.0 ppm, 72.0 ppm, and 71.0 ppm, respectively. The resonances at C-6 of 4-linked glucopyranosyl and 4-linked mannopyranosyl residues were observed at 61.0 ppm and 62.0 ppm, respectively (Fig. 6b). So, the proton, as well as carbon signals of purified SORB-EPS, were similar to that of plant-derived glucomannan. However, for more structural clarification, homo and heteronuclear 2-D NMR spectroscopy are needed to perform further.
Predicted EPS biosynthesis pathway
A total of 143 cell wall and capsule-related proteins were predicted in the genome of B. altitudinis SORB11, where 44 proteins belong to the capsular and extracellular polysaccharide part, 30 proteins belong to Gram-Positive cell wall components, and 69 uncategorized proteins. Under the capsular and extracellular polysaccharide part, genes were present that encode proteins for dTDP-rhamnose synthesis (4), CMP-N-acetylneuraminate biosynthesis (3), polysaccharide deacetylases (5), rhamnose containing glycans (10), exopolysaccharide biosynthesis (8), and sialic acid metabolism (14). For EPS biosynthesis, tyrosine kinase transmembrane modulator (EpsC), tyrosine kinase (EpsD) and phosphogalactosyl transferase (EpsE) proteins were predicted. An operon yqxM-sipW-tasA that positively regulates extracellular matrix biosynthesis was predicted. Moreover, a total of 26 glycosyltransferase family proteins were found in the genome of strain SORB11, viz., multispecies glycosyltransferase family 1 (1), bifunctional and multispecies glycosyltransferase family 2 (7), glycosyltransferase family 4 (2), glycosyltransferase family 39 (2), multispecies WecB/TagA/CpsF family glycosyltransferase (1), and multispecies glycosyltransferase (13) proteins.
Usually, for the regulation of EPS biosynthesis, interactions occur with the gene products homologous to epsCDE. In the initial step of EPS biosynthesis, EpsE is required. But for the activation of EpsE, functional EpsC and EpsD proteins are necessary where EpsC is required for the phosphorylation of EpsD. So, the activity of EpsE regulation by EpsC and EpsD possibly interacts in a complex to enable the biosynthesis of repeat units49,50.
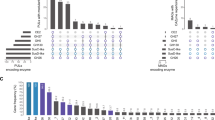
Heteropolysaccharide biosynthesis starts with the formation of intracellular EPS precursors that serve as the donor monomers for most of the repeating unit biosynthesis. The monomeric units of the EPS backbone are transferred by glycosyltransferases and facilitate the formation of glycosidic linkages between them51,52,53. Two types of glycosyltransferase enzyme activity are reported for the biosynthesis of glucomannan, mannosyltransferase which utilizes GDP-D-mannose and glucosyltransferase which utilizes GDP-D-Glucose as a substrate54,55. Here, a well-studied OsCSLA1 protein sequence that basically incorporates glucose into glucomannan from the donor substrate GDP-glucose54,56 was compared with the glycosyltransferase family protein clusters of strain SORB11 (Fig. 7). OsCSLA1 protein clustered with glycosyltransferase family 39 (functions as 4-amino-4-deoxy-L-arabinose transferase or related glycosyltransferases of PMT family); glycosyltransferase family 4 (functions as GDP-mannose-dependent α-1-2-phosphatidylinositol mannosyltransferase); bifunctional glycosyltransferase family 2 (functions as polypeptide N-acetyl galactosaminyltransferase); glycosyltransferase (functions as UDP-n-acetylglucosamine-n-acetylmuramyl-(pentapeptide) pyrophosphoryl-undecaprenoln-acetylglucosamine transferase); and multispecies of glycosyltransferase family protein (functions as ATP synthase subunits region ORF 6, the UDP complex structure of the sixth gene product of the F1-ATPase operon). So, this set of proteins along with other glycosyltransferases (Sup file 2.) may responsible for the biosynthesis of glucomannan type of extracellular polysaccharide in strain SORB11.
As per the previous report, there are four types of generalized transport mechanisms for the biosynthesis of polymers in bacterial system. However, ABC transporter dependent as well as, Wzx/Wzy dependent pathway which assembled by various glycosyltransferases are mainly responsible for the biosynthesis of heteropolysaccharides57,58. The different steps of a probable EPS biosynthesis pathway were stated according to the coding genes present in the genome of strain SORB11 (Fig. 8).
Discussion
Marine bacteria generally produce linear heteropolysaccharides that are arranged in clusters of 10 or fewer repeating units of pentoses (arabinose, ribose, xylose), hexoses (glucose, mannose, galactose, allose, rhamnose, fucose), amino sugars (glucosamine, galactosamine), and uronic acids (glucuronic acid, galacturonic acid) with an average Mw ranging from 1 × 105 to 3 × 105 Da4,5,6. Species of Vibrio, a heterotrophic facultative anaerobe, isolated from extreme deep-sea of East Pacific Rise produced novel extracellular polymers that composed of heteropolysaccharides with uronic acids and amino sugars, and trace neutral sugars. The species of Pseudoalteromonas reported from particulates of Antarctica and Southern Ocean produced sulfated heteropolysaccharides with uronic acids, acetyl, and succinyl groups; whereas, Pseudoalteromonas species isolated from the deep-sea of Guaymas Basin produced sulfated (6–13%) heteropolysaccharides with high uronic acids, pyruvate, and acetate groups13. The species of Alteromonas isolated from 2600 m depth of East Pacific Rise produced high molecular weight deepsane type of extracellular polysaccharide11,59.
Here, B. altitudinis SORB11 produced low molecular weight thermally stable sulfur containing heteropolysaccharides. The capsular polysaccharide (CPS) of strain SORB11 was also composed of d-mannose, d-glucose, d-galactose60. The sulfur content within both extracellular and capsular polysaccharides of strain SORB11 is the indication of its marine origin4. The EPS was a linear and simplified structure of plant-derived glucomannan. The genome of strain SORB11 contained protein coding genes corresponding to EPS production. So, these gene clusters are likely to interact in a complex, enabling the biosynthesis of polysaccharide repeat unit.
Other way, biofilms are multicellular aggregates composed of various combinations of DNA, protein, and polysaccharides and are stabilized by an extracellular matrix. In the marine ecosystems, biofilm formation takes place when planktonic bacteria start to secret exopolysaccharide matrix particularly to attach themselves with a surface61,62. The transition of biofilm formation is governed by a series of complex regulatory proteins. In the genome of strain SORB11, some of the genes were significantly predicted for exopolysaccharide matrix formation: for example, remA gene encoding multispecies of extracellular matrix/biofilm regulator protein RemA, ahpA gene encoding multispecies of biofilm-specific peroxidase enzyme AhpA, and multispecies of SipW-dependent-type signal peptide-containing protein. In Bacillus subtilis, two small proteins RemA and RemB were appeared to act in parallel like other biofilm regulators, SinR, AbrB, DegU were found to involve for the activation of matrix biosynthesis operons. Exopolysaccharide and TasA protein are the two primary structural components of an extracellular matrix assembly. The signal peptidase SipW encoded by yqxM operon is required for TasA secretion and localization to the matrix. Moreover, AhpA is a peroxidase enzyme that is expressed during biofilm formation, protecting cells within biofilms by scavenging free radicals62,63. The capsular polysaccharide of strain SORB11 showed a significant level of free radical scavenging activities60. So, the extracellular part i.e., EPS would expect to exhibit a satisfactory level of scavenging potentialities.
Conclusion
In this study, the nature and chemical structure of EPS produced by a deep-sea inhabiting B. altitudinis SORB11 were explicitly stated. It was a novel low molecular weight glucomannan type of EPS having mannose and glucose. The genome also represented the presence of glucosyltransferase and the other needful family proteins of glycosyltransferase for the biosynthesis of glucomannan. This low molecular weight microbial glucomannan act as a connector by forming networks surrounding the cells and moreover, it was hypothesized that this structure might help in cell-to-cell communication and supply nutrition under stressful conditions. However, investigations on the microbial glucomannan containing sulfur would expect to unfold the complete nature and functions of the unusual network formed by marine, free-living bacteria for survival under floating conditions.
Material and methods
Isolation of strain SORB11
The water sample was collected aseptically using a rosette conductivity temperature depth (CTD) instrument (Sea-Bird Electronics, Inc. Florida, USA) from the Indian Sector of Southern Ocean (50°S, 47°E; Station No. 22) in austral summer (January–March, 2011) during the 5th Southern Ocean Expedition conducted by National Centre for Antarctic and Ocean Research (NCAOR) and Ministry of Earth Sciences (MoES), Government of India. Water sample was collected from 3800 m depth. During the sampling period, sea surface temperature was 5.5 °C, depth chlorophyll maxima was 55 m and atmospheric pressure was 1019 hPa. Strain SORB11 was isolated, cultivated in Zobell Marine Broth 2216 (ZMB, HiMedia) medium at 20 °C and identified as Bacillus altitudinis23.
Production and optimization of crude EPS
Freshly prepared ~ 10.0 µL culture from exponential growth phase (Log CFU/mL = 6.60) of strain SORB11 was inoculated and cultivated on 1.0 L ZMB medium at 20 °C for 7 days at 150 rpm to increase the cell mass. The cell culture was then centrifuged at 5000×g for 20 min at 4 °C. For the removal of protein part, 5% (w/v) trichloroacetic acid (Sigma-Aldrich) was added with the cell-free supernatant and incubated overnight at 4 °C. The suspension was then centrifuged at 12,000×g for 20 min at 4 °C. An equal volume of 90% chilled ethanol (Sigma-Aldrich) was added with the supernatant and incubated overnight at 4 °C. The precipitated EPS part was collected by centrifugation at 12,000×g for 20 min at 4 °C. The pellet was washed with acetone two times and freeze-dried. The dry weight was calculated of that lyophilized EPS powder and stored at 4 °C for further analysis30. In order to get maximum EPS production under optimum growth conditions, in the precipitation step, different solvents like methanol (Sigma-Aldrich), isopropanol (Sigma-Aldrich), and ethyl acetate (Sigma-Aldrich) were used instead of ethanol.
Characterization of EPS
Chemical analysis
Carbohydrate estimation was performed to determine the total sugar content using glucose (Sigma-Aldrich) as standard64. Lowry method using Bovine Serum Albumin (BSA, Sigma-Aldrich) as a standard65 was performed to detect the total protein content. The Carbazole method was performed to determine glucuronic acid66.
Solubility test
For the dissolution of EPS, 1.0 mg powder was mixed with 3.0 mL of various non-polar and polar solvents like chloroform (Sigma-Aldrich), trifluoroethanol (Sigma-Aldrich), dimethyl sulfoxide (Sigma-Aldrich), acetic acid (Sigma-Aldrich), and water. The mixtures were properly vortexed and solubilities were checked15.
Surface topography study
Scanning electron microscopy (SEM, S530 Hitachi) was performed with lyophilized EPS. Then, 1.0 mg/mL EPS solution using Milli-Q was prepared, drop casted on the stab and air-dried. Field emission scanning electron microscopy (FE-SEM, Zeiss Gemini 300, Germany) of that dried, gold-coated (IB2 ion coater) EPS sample was performed with an accelerating voltage of 5.00 kV. Further, 50.0, 10.0, and 5.0 µg/mL of crude EPS solutions were prepared using Milli-Q and 1.0 µL of each EPS solutions were drop casted onto a freshly cleaved mica surface. The sample was then air dried at ambient humidity and temperature for 2 hours and the micrographs were taken with an Atomic force microscope (AFM, 5500 Agilent Technologies, USA) at tapping mode. The cell morphology of strain SORB11 was also observed under the AFM25.
Elemental analysis
The quantification of C, H, N, S, and C/N ratio was determined with a CHNS analyzer (Vario EL III, M/s Elementar, Germany). A ~ 5.0 mg of crude EPS powder was mixed with vanadium pentoxide oxidizer in a tin capsule (1000 °C reactor) to perform the analysis. Further, elements like C, H, N, and S content of the crude EPS were measured with an Energy-dispersive X-ray spectroscope (EDS, INCA 250, Oxford Instruments, UK) of EPS was performed using a microprobe.
Physical property study
The physical structural property of crude EPS powder was measured with an X-ray diffractometer (XRD, PANalytical X'pert PRO 2200, Malvern Instruments, UK) equipped with Cu radiation at 40 mA and 40 kV. The X-ray diffraction pattern was recorded from 5.00° to 80.00° at a 2θ angle.
Thermal stability analysis
Thermogravimetric analysis with ~ 5.0 mg of crude EPS powder was carried out using TG/ DTA (Pyris Diamond, Parkin Elmer, USA). The sample weights and the signals were recorded from 10 to 800 °C temperature range under a nitrogen atmosphere using alumina as a control with a heating rate of 10 °C/min.
Infrared spectroscopy
Fourier transform infrared spectroscopy (FT-IR, IR-Prestige 21, Shimadzu, Japan) of strain SORB11-derived EPS, i.e., represented as SORB-EPS and a plant-derived glucomannan (Now Foods) as a standard were carried out for the detection of functional groups. The IR spectra were recorded at room temperature with a resolution of 1 cm−1 in 400–4000 cm−1 using KBr pellet method.
Purification of crude EPS and molecular weight determination
Size-exclusion based gel permeation chromatography (GPC) was performed using Sepharose-6B (GE Healthcare) column (90 cm × 2.1 cm) to purify both SORB-EPS and glucomannan. A ~ 25 mg polysaccharide sample was dissolved in 5.0 mL deionized water and passed through a 0.22 µ filter. The total volume of the filtrate was loaded on the top of the column. Deionized water was used as eluent with a flow rate of 0.1 mL/s. In each tube, 4.0 mL of eluent was collected and measured spectrophotometrically (UV-2600, Shimadzu, Japan) by using phenol-H2SO4 method at 490 nm. Finally, the fractions containing sugar were collected and lyophilized to get the purified polysaccharides. Molecular weight of SORB-EPS and glucomannan were also determined by these Gel-chromatographic techniques using T-40, T-60, T-70, T-200, and T-250 species of dextran (Sigma-Aldrich) as standard67,68.
Monosaccharide composition analysis of purified EPS
For the analysis of monosaccharide component, alditol acetate derivatization was performed using 1.0 mg of lyophilized polysaccharide hydrolyzed with 2 M trifluoracetic acid (Sigma-Aldrich) at 120 °C for 2 hours followed by a further reduction using Sodium Borohydride (NaBD4, Sigma-Aldrich), and acetylation was performed with acetic anhydride (SD Fine chemicals) and pyridine (Sigma-Aldrich) (1:1) at 80 °C for 20 minutes69,70. 1.0 µL derivatized monosaccharides were injected into the Gas Chromatography coupled with a mass spectrometer analyzer (Agilent Technologies 7890, USA) using helium as carrier gas (temperature gradient was 80 °C for 2 min, 80–170 °C at 30 °C/min, 170–240 °C at 4 °C/min, 240 °C for 30 min and ionized by electrons impact at 70 eV).
Glycosyl linkage analysis of purified EPS
The composition of glycosyl linkage of SORB-EPS and standard glucomannan were analyzed by the generation of partially methylated alditol acetate derivatives71,72,73. ~ 2–3 mg samples were dissolved in 2.0 mL Dimethyl Sulfoxide Anhydrous (DMSO, Sigma-Aldrich) and ~ 15.0 mg freshly prepared Sodium Hydroxide (NaOH, Sigma-Aldrich) was added and the reaction mixtures were allowed to stand for 15 minutes at room temperature. 1.0 mL iodomethane (Sigma-Aldrich) was added to each sample and stirred for 30 minutes at room temperature. After removing excess iodomethane, washed and dried methylated samples were further derivatized into partially methylated alditol acetate and analyzed by Gas Chromatography coupled with a mass spectrometer analyzer (Agilent Technologies 7890, USA) as described in “monosaccharide composition analysis” section. The GC EI-MS spectra of the partially methylated alditol acetate (PMAA) derived from a glycosidically linked sugar residue were identified using The CCRC Spectral Database for PMAA’s.
Nuclear magnetic resonance spectroscopy of purified SORB-EPS
For 1H and 13C NMR spectroscopy, ~ 15.0 mg purified SORB-EPS was dissolved in 0.5 mL D2O (99.9%, Sigma-Aldrich). The spectra were recorded on a 500 MHz NMR spectrometer (Bruker, USA) with a 5 mm inverse probe at 25 °C sample temperature. Acetone (H 2.225 ppm, C 30.4 ppm) was used as an internal standard71,72.
Prediction of probable EPS biosynthesis pathway
The coding gene clusters responsible for EPS biosynthesis were annotated from the draft genome of strain SORB1123 from NCBI-PGAP74 and RAST75. OsCSLA1 protein54 (sp|Q7PC76|CSLA1_ORYSJ) was retrieved from NCBI-GenBank database. Glycosyltransferase family proteins responsible for EPS biosynthesis were analyzed by building homology model using SWISS-MODEL76 and functions from CAZy database77. Phylogenetic relationships and similarity index were studied by constructing a tree using minimum evolution with the close neighbor interchange inference method and 1000 bootstrap replicons in MEGA 778.
Data availability
Whole-genome shotgun project of SORB11 was deposited at DDBJ/ENA/GenBank under the accession number MEHW00000000; BioProject number PRJNA341563; BioSample number SAMN05726035 (https://www.ncbi.nlm.nih.gov/assembly/GCA_002042895.1). OsCSLA1 protein sequence is available in the NCBI-GenBank database (https://www.ncbi.nlm.nih.gov/gene/4328585).
References
Wang, X. et al. Marine polysaccharides attenuate metabolic syndrome by fermentation products and altering gut microbiota: An overview. Carbohyd. Polym. 195, 601–612 (2018).
Nicolaus, B. et al. Exopolysaccharides from extremophiles: from fundamentals to biotechnology. Environ. Technol. 31(10), 1145–1158 (2010).
Nichols, C. M. et al. Effects of incubation temperature on growth and production of exopolysaccharides by an Antarctic sea ice bacterium grown in batch culture. Appl. Environ. Microbiol. 71(7), 3519–3523 (2005).
Nichols, C. M. et al. Bacterial exopolysaccharides from extreme marine environments with special consideration of the Southern Ocean, sea ice, and deep-sea hydrothermal vents: A review. Mar. Biotechnol. 7(4), 253–271 (2005).
Nichols, C. M. et al. Chemical characterization of exopolysaccharides from Antarctic marine bacteria. Microb. Ecol. 49(4), 578–589 (2005).
Nichols, C. M. et al. Production of exopolysaccharides by Antarctic marine bacterial isolates. J. Appl. Microbiol. 96(5), 1057–1066 (2004).
Nicolaus, B. et al. Polysaccharides from extremophilic microorganisms. Orig. Life Evol. Biosph. 34(1), 159–169 (2004).
Nicolaus, B. et al. A thermophilic Bacillus isolated from an Eolian shallow hydrothermal vent able to produce exopolysaccharides. Syst. Appl. Microbiol. 23(3), 426–432 (2000).
Nicolaus, B. et al. Production and characterization of exopolysaccharides excreted by thermophilic bacteria from shallow, marine hydrothermal vents of flegreanares (Italy). Syst. Appl. Microbiol. 25(3), 319–325 (2002).
Nicolaus, B. et al. Haloarcula spp able to biosynthesize exo- and endopolymers. J. Ind. Microbiol. Biotechnol. 23(6), 489–496 (1999).
Le Costaouëc, T. et al. Structural data on a bacterial exopolysaccharide produced by a deep-sea Alteromonas macleodii strain. Carbohyd. Polym. 90(1), 49–59 (2012).
Casillo, A. et al. Structure-activity relationship of the exopolysaccharide from a psychrophilic bacterium: a strategy for cryoprotection. Carbohyd. Polym. 156, 364–371 (2017).
Casillo, A. et al. Exopolysaccharides from marine and marine extremophilic bacteria: Structures, properties, ecological roles and applications. Mar. Drugs 16(2), 69 (2018).
Kennedy, F. et al. Short note: extracellular export and consumption of glucose in Antarctic sea ice. Polar Biol. 45, 763–768 (2022).
Raveendran, S. et al. Biocompatible nanofibers based on extremophilic bacterial polysaccharide, Mauran from Halomonas maura. Carbohydr. Polym. 92(2), 1225–1233 (2013).
Passow, U. & Alldredge, A. Distribution, size and bacterial colonization of transparent exopolymer particles (TEP) in the ocean. Mar. Ecol. Prog. Ser. 113, 185–198 (1994).
Kokoulin, M. S. et al. Sulfated O-polysaccharide with anticancer activity from the marine bacterium Poseidonocella sedimentorum KMM 9023T. Carbohyd. Polym. 202, 157–163 (2018).
Tziveleka, L. A., Ioannou, E. & Roussis, V. Ulvan, a bioactive marine sulphated polysaccharide as a key constituent of hybrid biomaterials: A review. Carbohyd. Polym. 218, 355–370 (2019).
Martin-Pastor, M. et al. Structure, rheology, and copper-complexation of a hyaluronan-like exopolysaccharide from Vibrio. Carbohyd. Polym. 222, 114999 (2019).
Chen, G. et al. Exopolysaccharide of Antarctic bacterium Pseudoaltermonas sp. S-5 induces apoptosis in K562 cells. Carbohyd. Polym. 121, 107–114 (2015).
Kokoulin, M. S. et al. Structure and in vitro antiproliferative activity of the acidic capsular polysaccharide from the deep-sea bacterium Psychrobacter submarinus KMM 225T. Carbohyd. Polym. 262, 117941 (2021).
Zhang, S. et al. Cocktail polysaccharides isolated from Eckloniakurome against the SARS-CoV-2 infection. Carbohyd. Polym. 275, 118779 (2022).
Halder, U. et al. Draft Genome Report of Bacillus altitudinis SORB11 isolated from the Indian Sector of the Southern Ocean. Genome Announc. 5(23), e00339-e417 (2017).
Flemming, H. C. & Wingender, J. Relevance of microbial extracellular polymeric substances (EPSs)—Part I: Structural and ecological aspects. Water Sci. Technol. 43(6), 1–8 (2001).
Luo, Q. L. et al. Chemical properties and antioxidant activity of a water-soluble polysaccharide from Dendrobium officinale. Int. J. Biol. Macromol. 89, 219–227 (2016).
Shao, L. et al. Molecular characteristics of an exopolysaccharide from Lactobacillus rhamnosus KF5 in solution. Int. J. Biol. Macromol. 72, 1429–1434 (2014).
Singh, R. P. et al. Isolation and characterization of exopolysaccharides from seaweed associated bacteria Bacillus licheniformis. Carbohyd. Polym. 84, 1019–1026 (2011).
Yu, L. et al. Preparation and partial structural characterization of the exopolysaccharide from Bacillus mucilaginosus SM-01. Carbohyd. Polym. 146, 217–223 (2016).
Ahmad, A. et al. International Journal of Biological Macromolecules Extraction and characterization of β- d-glucan from oat for industrial utilization. Int. J. Biol. Macromol. 46, 304–309 (2010).
Sardaria, R. R. R. et al. Evaluation of the production of exopolysaccharides by two strains of the thermophilic bacterium Rhodothermus marinus. Carbohyd. Polym. 156, 1–8 (2017).
Vessella, G. et al. Exploiting diol reactivity for the access to unprecedented low molecular weight curdlan sulfate polysaccharides. Carbohyd. Polym. 269, 118324 (2021).
Seedevi, P. et al. Bioactive potential and structural chracterization of sulfated polysaccharide from seaweed (Gracilari acorticata). Carbohyd. Polym. 155, 516–524 (2017).
Liu, X. et al. A rhamnan-type sulfated polysaccharide with novel structure from Monostroma angicava Kjellm (Chlorophyta) and its bioactivity. Carbohyd. Polym. 173, 732–748 (2017).
Lafarge, C. & Cayot, N. Potential use of mixed gels from konjac glucomannan and native starch for encapsulation and delivery of aroma compounds: A review. Starch-Stärke 70(9–10), 1700159 (2018).
Parente, J. P. et al. Structural characterization of an acetylated glucomannan with anti-inflammatory activity and gastroprotective property from Cyrtopodium andersonii. Carbohyd. Res. 391, 16–21 (2014).
Alonso-Sande, M. et al. Glucomannan, a promising polysaccharide for biopharmaceutical purposes. Eur. J. Pharm. Biopharm. 72, 453–462 (2009).
Mao, Y. H. et al. Effects of konjac glucomannan with different molecular weights on gut microflora with antibiotic perturbance in in vitro fecal fermentation. Carbohyd. Polym. 273, 118546 (2021).
Xueqian, W. et al. An O-acetyl-glucomannan from the rhizomes of Curculigoorchioides: Structural characterization and anti-osteoporosis activity in vitro. Carbohyd. Polym. 174, 48–56 (2017).
Nguyen, A. T. et al. Characterization of glucomannan from Amorphophallus panomensis in Vietnam. Vietnam J. Sci. Technol. 54(2), 224–224 (2016).
Campestrini, L. H. et al. NMR and rheological study of Aloe barbadensis partially acetylated glucomannan. Carbohyd. Polym. 94(1), 511–519 (2013).
De Souza, A. C. et al. A robust and universal NMR method for the compositional analysis of polysaccharides. Carbohyd. Polym. 95(2), 657–663 (2013).
Enomoto-Rogers, Y., Ohmomo, Y. & Iwata, T. Syntheses and characterization of konjac glucomannan acetate and their thermal and mechanical properties. Carbohyd. Polym. 92(2), 1827–1834 (2013).
An, N. T. et al. Characterization of glucomannan from some Amorphophallus species in Vietnam. Carbohyd. Polym. 80, 308–311 (2010).
Ishrud, O. et al. Isolation and structure analysis of a glucomannan from the seeds of Libyan dates. J. Agric. Food Chem. 49(8), 3772–3774 (2001).
Ishrud, O. et al. Structural of a glucomannan from Lupinus varius seed. Carbohyd. Polym. 65(4), 410–413 (2006).
Teleman, A. et al. Isolation and characterization of O-acetylated glucomannans from aspen and birch wood. Carbohyd. Res. 338(6), 525–534 (2003).
Crescenzi, V. et al. A high field NMR study of the products ensuing from konjak glucomannan C (6)-oxidation followed by enzymatic C (5)-epimerization. Biomacromol 3(6), 1343–1352 (2002).
Cescutti, P. et al. Structure of the oligomers obtained by enzymatic hydrolysis of the glucomannan produced by the plant Amorphophallus konjac. Carbohyd. Res. 337(24), 2505–2511 (2002).
Kang, H. J. & LaPointe, G. Production of Exopolysaccharides by Lactococcus lactis sub sp. cremoris MG1363 expressing the eps gene clusters from two strains of Lactobacillus rhamnosus. Microbiol. Biotechnol. Lett. 46(2), 91–101 (2018).
Minic, Z. et al. Control of EpsE, the phosphoglycosyltransferase initiating exopolysaccharide synthesis in Streptococcus thermophilus, by EpsD tyrosine kinase. J. Bacteriol. 189, 1351–1357 (2007).
Soumya, M. P. & Nampoothiri, K. M. An overview of functional genomics and relevance of glycosyltransferases in exopolysaccharide production by lactic acid bacteria. Int. J. Biol. Macromol. 184, 1014–1025 (2021).
Lairson, L. L. et al. Glycosyltransferases: structures, functions, and mechanisms. Annu. Rev. Biochem. 77, 521–555 (2008).
Péant, B. et al. Comparative analysis of the exopolysaccharide biosynthesis gene clusters from four strains of Lactobacillus rhamnosus. Microbiology 151(6), 1839–1851 (2005).
Gille, S. et al. Deep sequencing of voodoo lily (Amorphophallus konjac): An approach to identify relevant genes involved in the synthesis of the hemicellulose glucomannan. Planta 234, 515–526 (2011).
Heller, J. S. & Villemez, C. L. Interaction of Soluble Glucosyl- and Mannosyl-Transferase enzyme activities in the synthesis of a Glucomannan. Biochem. J. 129, 645–655 (1972).
Liepman, A. H. et al. Functional genomic analysis supports conservation of function among cellulose synthase-like a gene family members and suggests diverse roles of mannans in plants. Plant Physiol. 143(4), 1881–1893 (2007).
Schmid, J. Recent insights in microbial exopolysaccharide biosynthesis and engineering strategies. Curr. Opin. Biotechnol. 53, 130–136 (2018).
Schmid, J., Sieber, V. & Rehm, B. Bacterial exopolysaccharides: biosynthesis pathways and engineering strategies. Front. Microbiol. 6, 496 (2015).
Raguénès, G. H. C. et al. Alteromonas infernus sp. nov., a new polysaccharide-producing bacterium isolated from a deep-sea hydrothermal vent. J. Appl. Microbiol. 82(4), 422–430 (1997).
Halder, U. et al. Synthesis of copper oxide nanoparticles using capsular polymeric substances produced by Bacillus altitudinis and investigation of its efficacy to kill pathogenic Pseudomonas aeruginosa. Chem. Eng. J. Adv. 11, 100294 (2022).
Harrison-Balestra, C. et al. A wound-isolated Pseudomonas aeruginosa grows a biofilm in vitro within 10 hours and is visualized by light microscopy. Dermatol. Surg. 29(6), 631–635 (2003).
Winkelman, J. T. et al. RemA (YlzA) and RemB (YaaB) regulate extracellular matrix operon expression and biofilm formation in Bacillus subtilis. J. Bacteriol. 191(12), 3981–3991 (2009).
Zwick, J. V. et al. AhpA is a peroxidase expressed during biofilm formation in Bacillus subtilis. Microbiologyopen 6(1), e00403 (2017).
Dubois, M. et al. Colorimetric method for determination of sugars and related substances. Anal. Chem. 28, 350–356 (1956).
Lowry, O. H. et al. Protein measurement with the Folin phenol reagent. J. Biol. Chem. 193(1), 265–275 (1951).
Knutson, C. A. & Jeanes, A. A new modification of the carbazole analysis: Application to heteropolysaccharides. Anal. Biochem. 24, 470–481 (1968).
Varma, C. A. K. & Kumar, K. J. Structural, functional and pH sensitive release characteristics of water-soluble polysaccharide from the seeds of Albizia lebbeck L.. Carbohyd. Polym. 175, 502–508 (2017).
Nandan, C. K. et al. Isolation and characterization of polysaccharides of a hybrid mushroom (backcross mating between PfloVv12 and Volvariella volvacea). Carbohyd. Res. 346(15), 2451–2456 (2011).
Fox, A., Morgan, S. L. & Gilbart, J. Preparation of alditol acetates and their analysis by gas chromatography (GC) and mass spectrometry (MS). In Analysis of Carbohydrates by GLC and MS, 87–117 (1989).
Ali, U. et al. Effect of β-glucanfatty acid esters on microstructure and physical properties of wheat straw arabinoxylan films. Carbohyd. Polym. 161, 90–98 (2017).
Ali, U. et al. Effect of arabinoxylan and β-glucan stearic acid ester coatings on post-harvest quality of apple (Royal Delicious). Carbohyd. Polym. 209, 338–349 (2019).
Mazumder, K. & York, W. S. Structural analysis of arabinoxylans isolated from ball-milled switchgrass biomass. Carbohyd. Res. 345, 2183–2219 (2010).
Usuldin, S. R. A. et al. In-depth spectral characterization of antioxidative (1, 3)-β-D-glucan from the mycelium of an identified tiger milk mushroom Lignosus rhinocerus strain ABI in a stirred-tank bioreactor. Biocatal. Agric. Biotechnol. 23, 101455 (2020).
Tatusova, T. et al. NCBI prokaryotic genome annotation pipeline. Nucleic Acids Res. 44(14), 6614–6624 (2016).
Aziz, R. K. et al. The RAST server: rapid annotations using subsystems technology. BMC Genomics 9, 75–89 (2008).
Waterhouse, A. et al. SWISS-MODEL: homology modelling of protein structures and complexes. Nucleic Acids Res. 46(W1), W296–W303 (2018).
Terrapon, N. et al. Automatic prediction of polysaccharide utilization loci in Bacteroidetes species. Bioinformatics (Oxford, England) 31(5), 647–655 (2015).
Kumar, S., Stecher, G. & Tamura, K. MEGA7: molecular evolutionary genetics analysis version 70 for bigger datasets. Mol. Biol. Evol. 33(7), 1870–1874 (2016).
Acknowledgements
We respectfully acknowledge UGC–Centre for Advanced Study, Department of Botany, The University of Burdwan, and DST FIST II for providing research facilities. We are also thankful to The University Science Instrumentation Centre (USIC), The University of Burdwan for SEM, FE-SEM, XRD, and EDX studies; Central Research Facility (CRF), IIT, Kharagpur for AFM study; Advanced Instrument Research Facilities (AIRF), JNU, New Delhi for NMR study; Central Instrumental Facilities (CIF), BIT, Mesra for elemental analysis; and Instrumentation Facilities, NABI, Mohali for GC-MS and TG analysis. We are very much thankful to Dr. Usman Ali, Dr. Swati Kanwar, Mr. Rohit Maurya, and Mr. Atul Kumar Kesarwani from NABI, Mohali and Mr. Saptarshi Samajdhar from BIT, Mesra for their kind help and hospitality during the experiments. We are extremely thankful to Dr. Joy K Roy, Scientist, NABI, Mohali for his kind help and support.
Funding
MoES and NCAOR. The bacterial strain was collected during the 5th Indian Southern Ocean Expedition (No. MoES/NCAOR/SOS/1/2007-PC-I; dated Jan 04, 2011), fully supported by Ministry of Earth Sciences (MoES), New Delhi and National Center for Polar and Ocean Research (NCAOR), Goa. UH is thankful to SRF (State Fund) fellowship [Fc (Sc.)/RS/SF/ BOT./2017-18/22] to conduct this research work.
Author information
Authors and Affiliations
Contributions
Conceptualization and Methodology, U.H., R.B.; Experiments, U.H., K.M., K.J.K.; Writing and Editing, U.H., R.B., K.M.; All authors have agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Halder, U., Mazumder, K., Kumar, K.J. et al. Structural insight into a glucomannan-type extracellular polysaccharide produced by a marine Bacillus altitudinis SORB11 from Southern Ocean. Sci Rep 12, 16322 (2022). https://doi.org/10.1038/s41598-022-20822-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-20822-3
This article is cited by
-
Genome-based characterization of the deep-sea psychrotolerant bacterium Bacillus altitudinis SORB11 isolated from the Indian Sector of the Southern Ocean
Polar Biology (2024)
-
Production and characterization of exopolysaccharide (EPS) from marine Bacillus halotolerans and its antibacterial activity against clinical pathogens
Biologia (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.