Abstract
Picocyanobacteria (< 2 µm in diameter) are significant contributors to total phytoplankton biomass. Due to the high diversity within this group, their seasonal dynamics and relationship with environmental parameters, especially in brackish waters, are largely unknown. In this study, the abundance and community composition of phycoerythrin rich picocyanobacteria (PE-SYN) and phycocyanin rich picocyanobacteria (PC-SYN) were monitored at a coastal (K-station) and at an offshore station (LMO; ~ 10 km from land) in the Baltic Sea over three years (2018–2020). Cell abundances of picocyanobacteria correlated positively to temperature and negatively to nitrate (NO3) concentration. While PE-SYN abundance correlated to the presence of nitrogen fixers, PC-SYN abundance was linked to stratification/shallow waters. The picocyanobacterial targeted amplicon sequencing revealed an unprecedented diversity of 2169 picocyanobacterial amplicons sequence variants (ASVs). A unique assemblage of distinct picocyanobacterial clades across seasons was identified. Clade A/B dominated the picocyanobacterial community, except during summer when low NO3, high phosphate (PO4) concentrations and warm temperatures promoted S5.2 dominance. This study, providing multiyear data, links picocyanobacterial populations to environmental parameters. The difference in the response of the two functional groups and clades underscore the need for further high-resolution studies to understand their role in the ecosystem.
Similar content being viewed by others
Introduction
Unicellular picocyanobacteria (< 2 µm in diameter) belonging to the Synechococcus/Cyanobium/Synechocystis genus (SYN) are highly adaptable and widespread in aquatic ecosystems1. The genus is polyphyletic2,3 and is characterized by a high diversity in terms of pigment content4, taxonomy5 and physiology6. SYN populations are a major contributor to carbon flux in oligotrophic oceans7,8 as well as coastal, estuarine and freshwaters9,10,11. However, there are relatively few studies that focus on SYN dynamics and community composition in freshwater12,13,14 and estuaries15,16,17 despite showcasing higher diversity than their marine counterparts 14. Under climate change scenarios, the abundance of SYN cells is expected to increase1,18 while the rest of the phytoplankton community biomass is expected to decrease19,20. Understanding the distribution and dynamics of SYN ecotypes is therefore of high interest for evaluating ecosystem consequences.
Distribution and seasonal occurrence of marine phylogenetic clades have been linked to environmental factors, increasing our understanding on marine SYN dynamics and distribution21,22,23. SYN populations are composed of unicellular phycoerythrin (PE)-rich cells (PE-SYN, adapted to blue and green light) and phycocyanin (PC)-rich cells (PC-SYN, adapted to red light). It has been observed that PE-SYN are more prevalent in low turbidity waters (generally open waters), while PC-SYN are better adapted to turbid waters (generally coastal waters)24,25,26,27,28. Other factors, such as water column stability and stratification can also have a significant effect on the PE:PC ratios11,29. Peak SYN cell abundances are normally recorded during periods of warm temperatures and low nutrient concentrations8,30. SYN cells may use ammonium (NH4) from nitrogen (N2)-fixers or from regeneration through ammonification as main source of nitrogen31,32,33,34 or can take up dissolved organic nitrogen (DON) directly35,36,37. Other nutrients such as phosphate (PO4) have also been observed to have an effect on picocyanobacterial dynamics and distribution38.
Pigment related genes cpcBA, mpeBA, and mpeW have been used to describe SYN diversity39. Studies based on 16S rRNA, 16S-23S internally transcribed spacer (ITS), petB, psbB, rpoC1, narB genes as well as full genome analysis have provided more detailed phylogeny despite not being directly related to the pigment content or other morphological characteristics40,41,42,43,44,45,46,47. Phylogenetically, SYN populations can be divided into three subclusters: S5.1, composed strictly of marine strains, S5.2, containing both marine and brackish adapted strains and S5.3 containing mainly oceanic strains from surface waters21,48,49,50,51. These subclusters can be further divided into 20 well defined clades, which are closely related to the physio-ecological characteristics of SYN52,53. In addition, there are a number of brackish and freshwater clades that are not taxonomically assigned to known subclusters12,14.
The Baltic Sea is characterized by a salinity gradient from 2.9 PSU in the Bothnian Bay to 22 PSU in the Kattegat. In the Baltic Proper, picocyanobacterial blooms take place during the summer at temperatures up to 20–24 °C and nitrogen limited conditions54,55,56. Peak abundances during this period reach up to 105 cells mL−155,57 corresponding to 21% contribution to the total phytoplankton community in terms of Chl a58 and 56% in terms of carbon biomass52. The diversity of the picocyanobacterial community in the Baltic Sea has been characterized by amplification and sequencing of the V4-V5 16S rRNA gene hypervariable region (e.g.6,15,17,59). SYN sequences can dominate the sequence libraries6,17. During summer, the SYN populations are dominated by brackish strains from the S5.2 in the North, and there is a transition towards a community dominated by S5.1 at 13–16 PSU in the South17. Amplification and sequencing of the pigment genes for PE (cpeBA) and PC (cpcBA) showed that the Baltic Sea is equally populated by PE-SYN and PC-SYN in the upper 30 m of the water column29. In the summer, the SYN population is dominatied by an ecotype with a novel pigment gene which was recently characterized by a metagenome-assembled genome (MAG) reconstructed from the Baltic Proper (BACL30)16. The discovery of this unique brackish strain is linked to the existence of a global brackish microbiome60,61. The current data suggests that the high physiological diversity of the picocyanobacterial community shows adaptation to different temperatures and nutrient regimes6 but few annual studies have been conducted which limits the understanding of seasonal SYN dynamics.
Recently, Huber et al.62 showed that the hypervariable region V5-V7 of the 16S rRNA gene provided high resolution of picocyanobacterial diversity in both marine and freshwater environments. An accurate phylogenetic clade classification over multiple years in the dynamic Baltic Sea can help to disentangle the drivers of SYN and fill knowledge gaps of brackish and freshwater strains. Here we monitored the abundance of SYN cells with flow cytometry and studied its clade community composition at high-resolution during three consecutive years at a coastal and an offshore station. The goal was to study the relationship between SYN pigment ecotypes and phylogenetic clades with environmental parameters in a brackish environment.
Results
Environmental conditions
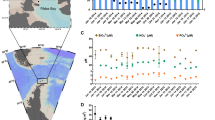
Sampling was carried out every second week at the Linnaeus Microbial Observatory (LMO), an offshore station and weekly at the K-station, a coastal station during 2018–2020. Seawater temperature ranged from 0 to 5 °C during winter at both stations and increased during spring (~ 3–15 °C) and summer (~ 15–20 °C; Fig. 1A). In 2018, there was an earlier increase of temperature during summer compared to 2019 and 2020 (average 3 °C higher in the beginning of June) and the maximum temperatures during the study period were 24 and 22 °C at the K-station and LMO respectively. At LMO, the increase of temperature during summer resulted in stratification (mixed layer approx. depth: 20 m) that lasted until early to mid-autumn (stratification index (N2): 0.2–0.66 × 10–3 s−2; Supplementary Fig. S2A and B). In 2018, early signs of stratification were observed during spring due to the exceptionally high temperatures. Salinity ranged between 6.5 and 8 PSU (Fig. 1B). At the K-station, salinity increased during the spring to summer period while no clear seasonality was observed at the LMO. Inorganic nitrogen (NO3) concentrations ranged between < 0.06 (detection limit) and 5 µM, with the highest concentrations observed at the K-station during winter, which sharply declined upon temperature increase during spring (Fig. 1C). Phosphate concentrations (PO4) decreased during spring (from 1 to 0.2 µM) and remained low throughout the summer with occasional peaks up to 1 µM at the K-station (Fig. 1D). At the K-station, silicate (SiO2) showed a strong seasonality ranging from 4 to 25 µM with peaks in summer and early autumn (Fig. 1E). Silicate did not show a clear seasonality at LMO. Concentrations ranged between 10 and 20 µM during 2018, sharply declined during spring in 2019 and remained stable around 5 µM throughout 2020.
Phytoplankton dynamics
Chlorophyll a (Chl a) concentration was generally higher at the K-station (1–7 µg L-1) compared to LMO (1–4.5 µg L−1; Fig. 2A). At the K-station, phytoplankton carbon biomass increased from 0.4 mg C mL−1 in the autumn to winter period up to 50.0 mg C mL−1 in the spring and 147.7 mg C mL−1 in the summer (Fig. 2B). The phytoplankton community was dominated by diatoms during autumn, winter and first half of the spring (Fig. 2C). During the second half of the spring the community transitioned to ciliates and dinoflagellates dominance. N2-fixers (filamentous cyanobacteria) were only observed during the summer 2018, reaching 111.3 mg C mL−1 in July. At the LMO, the maximum carbon biomass increased from 5.9 mg C mL−1 in the autumn and winter up to 597.4 mg C mL−1 in the spring and 167.7 mg C mL−1 in the summer (Fig. 2B). During the autumn, winter and spring, the community was dominated by ciliates and dinoflagellates (Fig. 2D), and occasionally by diatoms. During the summer the phytoplankton community was dominated by N2-fixers, reaching 111.3 mg C mL−1 total carbon biomass.
K-station and LMO measurements for (a) Chl a (µg L−1), (b) total phytoplankton (> 5 µm in diameter) carbon biomass concentration (mg C mL−1) based on microscopy, (c) K-station and (d) LMO relative contribution of phytoplankton divisions (> 5 µm in diameter) based on microscopy, (e) PE-SYN (cells mL−1) and (f) PC-SYN (cells mL−1).
Picocyanobacterial dynamics
PE-SYN concentration was low in the winter (LMO: 2.7 × 102 cells mL−1, K-station: 8.2 × 102 cells mL−1) and had peak abundances in the period from spring to summer (up to 3.8 × 105 cells mL−1) and summer to autumn (up to 2.6 × 105 cells mL−1) at the K-station and LMO respectively (Fig. 2E). PC-SYN increased from low concentrations in the winter (K-station: 72 cells mL−1, LMO: 8 cells mL−1) to peak concentrations during spring and summer (K-station: 2.1 × 105 cells mL−1, LMO: 4.8 × 103 cells mL−1; Fig. 2F).
To analyze the effect of each independent variable on PE-SYN and PC-SYN abundance, we used the partial least square regression (PLS) method to establish a regression equation. The number of components utilized for each equation was 2 and 6 for PE-SYN and PC-SYN respectively. Only the independent variables with variable importance in projection (VIP) > 1 were considered relevant for the final equation (Table 1). The model for PE-SYN explained 38% of the cell abundance seasonal changes with 52% variation of the independent variables. The regression equation between PE-SYN cell abundance and the independent variables was (1):
The model for PC-SYN explained 40% of the cell abundance seasonal changes with 46% variation of the independent variables. The regression equation between PC-SYN cell abundance and the independent variables was (2):
Picocyanobacteria community composition
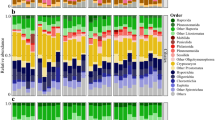
In total, there were 2169 ASVs identified as SYN in the amplicon libraries. At the LMO, the number of ASVs remained more stable throughout the seasons (Fig. 3). The ASVs were affiliated with 13 different subgroups and clades: Bornholm, subalpine II, S5.1, S5.2, Ib and the newly defined A/B, KS1, KS2, KS3, KS4, KS5 and KS6. Clade A/B was defined as a combination between clade A and clade B, since the V5-V7 region of the 16S rRNA gene targeted in this study did not allow for a clear separation between these two clades. 39 ASVs represented > 1% of picocyanobacterial sequences in at least one sample, of which only one ASV belonged to the marine S5.1 clade (Supplementary Fig. S3), eight belonged to the S5.2 clade, 15 belonged to clade A/B and the rest were divided among freshwater clades (H, Subalpine I, Bornholm, Ib, KS1, KS2, KS3, KS4, KS5 and KS6). ASVs within clade A/B dominated at both stations during autumn (maximum contribution at the K-station 2018-11-10: 92% and LMO 2020-11-26: 92%), winter (maximum contribution at the K-station 2020-11-10: 90% and LMO 2019-02-19: 96%) and spring (maximum contribution at the K-station 2019-05-14: 98% and LMO 2020-05-26: 98%). During summer, the K-station libraries were dominated by S5.2 (maximum contribution 2019-08-13: 91%), followed by an increase in KS3 at the end of the summer to beginning of autumn (maximum contribution 2019-10-08: 68%). At the LMO the period of late summer to early autumn was occasionally dominated by S5.2 (maximum contribution 2018-09-11: 68%). The picocyanobacterial community was more diverse at the coastal K-station compared to the offshore LMO (K-station registered 1026 ASVs and LMO registered 195 ASVs). The phylogenetic classification (which only includes ASVs that represented > 1% of the sequences for at least one sampling point) showed that some sequences that belong to the clades KS2 (ASV00070), KS5 (ASV07850) KS6 (ASV00066) and S5.2 (ASV07854 and ASV07849) were only present at the coastal K-station. The number of ASVs was consistently higher at the K-station, with the highest numbers of ASVs present during autumn to early spring (Fig. 3).
Relative abundance of ASVs with contribution > 1% to the total Synechococcus community in the K-station and LMO. The line plot on top represents total picocyanobacterial abundance for each sampling point. The heatmap represents the relative abundance of each ASV for each sampling point. The color code on the left indicates the clade of each ASV represented on the heatmap. The bottom barplot indicates the relative abundance of each subgroup/clade to the total Synechococcus community. The bottom dot plot indicates the number of ASVs for each sampling point.
Correlations between environmental and biological variables with picocyanobacterial composition
The test on Procrustes analysis and the CAs revealed a significant association between the distribution of the phylogenetic clades and the ASVs (squared m12 P value < 0.001, Supplementary Fig. S4). The stepwise CCA showed a significant relationship between clade composition and eight environmental and biological variables: NO3 (µM), SiO2 (µM), PO4 (µM), temperature (°C), salinity (PSU), N2-fixers biomass (mg C mL−1), PE-SYN (cells mL−1) and PC-SYN (cells mL−1; Table 2). The first two axis explained 49% of the taxonomic composition (Fig. 4). The first axis indicated a clear separation of summer samples. Summer samples were associated with high abundance of PE-SYN and PC-SYN, increase of temperature and high PO4. Autumn, winter and spring sites from the model clustered together and were associated with high NO3. PC-SYN abundance and temperature were the most important variables as indicated by the length of the variable arrows (Fig. 4).
Discussion
This study shows that temperature was one of the main parameters driving picocyanobacterial abundance. The correlation between temperature and increase in picocyanobacterial abundance is well known in marine8,9, estuarine63,64 and freshwater systems65,66. Picocyanobacterial abundance started increasing at > 10 °C, in line with other temperate ecosystems (e.g.67) and previous records in the Baltic Sea9. Peak abundances at the coastal K-station and offshore LMO during 2019 and 2020 were in the same range, with the exception of the summer of 2018, when K-station peak abundances reached 4.7 × 105 cells mL−1. These numbers are comparable to other observations in the Baltic Proper during summer (105 cells mL−1), and confirm previous observations that picocyanobacterial abundances are as high at coastal and offshore locations55,57,68. It is also important to note that the picocyanobacterial abundances at < 10 °C reported in this study were notably higher than previous reports in the Gulf of Finland and in other temperate ecosystems8,9. The thermal bound for SYN has recently been defined as > 5 °C18. However, in this study abundances of > 104 cells mL−1 were recorded during winter time at both K-station (0–5.7 °C) and LMO (2.7–5.8 °C), suggesting that the strains present in the Baltic Sea are well adapted to low temperatures, in line with previous observations in marine69 and freshwater environments70.
According to the PLS models independent variables only explained 38 and 40% of the total variation of PE-SYN and PC-SYN respectively. This indicates that other important controllers such as light quality4, grazing by ciliates and flagellates71 or viral lysis72 may be important to include in future models. SYN was divided into PE-SYN and PC-SYN depending of the pigment content. PE-SYN and PC-SYN coexisted at similar abundances during the summer, confirming previous observations based on cpcBA and cpeBA libraries29. PE-SYN are better adapted to blue light which can penetrate deeper in the mixing layer, while PC-SYN are adapted to red light, which is dissipated at the surface 4. As a result, PE-SYN showed no correlation with the stratification index (N2) and was equally prevalent at both the K-station and the LMO. On the other hand, PC-SYN abundance variation was strongly linked to N2 and was more prevalent at the coastal K-station compared to the offshore LMO, in line with observations in other estuaries and freshwater lakes11,73. These results suggest a horizontal gradient in the Baltic Sea, where PC-SYN is more prevalent in coastal shallow areas while in offshore areas abundances are lower and tightly joined to stratification. In the future, an increase in the stratification periods as a result of global warming could reinforce PC-SYN dominance on the picocyanobacterial community. PC-SYN have been observed to avoid predation, viral lysis and to have a negative effect on co-occurring filter-feeders, which can affect the energy flow to upper trophic layers74,75,76. Thus, understanding the physiology and ecology of PC-SYN, a generally understudied group of SYN, is of high importance for the understanding of current and future climate scenarios.
Nutrient availability, particularly nitrogen species, was correlated to picocyanobacterial dynamics according to the PLS models. Both PE-SYN and PC-SYN showed moderate negative correlation with NO3 concentration. Some studies have suggested preference of picocyanobacteria for NH4 over NO3 at high temperature conditions (> 15 °C)55,77. Thus, newly fixed nitrogen in the form of NH4 from N2-fixers or regeneration during blooms may be an important driver supporting picocyanobacterial growth during summer31,32,33, in line with the positive correlation between N2-fixers and PE-SYN. However, PE-SYN peak abundances at the K-station and LMO were in the same range, although N2-fixers were only observed at the K-station during 2018. This suggests that PE-SYN can benefit from newly fixed nitrogen, but the presence of N2-fixers is not a requirement to achieve peak abundances and thus other nitrogen pools should also be considered. The main nitrogen source for picocyanobacteria during the summer may be regeneration through ammonification34. This may be especially important in coastal and shallow water areas (< 50 m depth) where studies have estimated that it can represent up to 97% of the nitrogen requirements33,78. In addition, peak abundances at the LMO were sustained during the first half of the autumn at < 10 °C, which indicates that picocyanobacteria can uptake NO3 at low concentrations efficiently52. Direct uptake of DON may also benefit SYN over other phytoplankton35,36,37.
The community composition was studied using 16S rRNA gene sequences amplified using specific primers that target almost exclusively picocyanobacteria62. The results corroborate that the V5-V7 region of the 16S rRNA gene showcases higher variability in picocyanobacteria than the V3-V4 region, revealing an unprecendented high strain diversity in the Baltic Sea with a particularly high number of ASVs at the coastal K-station. Most of the previously defined clades and clusters were described62, but some clades were not well resolved. For example, clades A and B clustered together (clade A/B) contrasting with phylogeny based on other regions of the 16S rRNA gene12,79. All ASVs in clade A/B displayed similar seasonal variation in relative abundance, suggesting a similar ecophysiology whithin the clade. The increase in contribution of S5.2 took place in June-July, when temperature was > 18 °C and NO3 concentration was low. This result suggests that S5.2 affiliated picocyanobacterial strains are adapted to high temperatures and may use NH4 as a primary nitrogen source33,55,77. On the other hand, clade A/B dominated during the colder months, indicating that picocyanobacteria strains in this clade have a preference for low temperatures and high NO3 concentration.
This study indicates a coastal to offshore differentiation in picocyanobacterial community composition. The coastal K-station presented higher ASV diversity than the offshore LMO. Moreover, the clades KS2, KS5 and KS6 were only present at the coastal K-station, which suggest that some clades are only present in the coastal region. These results contradict previous observations in the Baltic Proper where no significant differences in community composition were observed in coastal offshore gradients6. One explanation is that the higher resolution achieved with the primers in this study have revealed differences that could not be detected with less specific primers. Community composition seasonal dynamics in the coastal and offshore stations also showed major differences. At the K-station picocyanobacterial peak abundances took place when S5.2 was dominating the community while at the LMO, peak abundances took place under clade A/B dominance. The composition at the coastal compared to the offshore station could be driven by low PO4 levels as S5.2 was positively linked to PO4 concentration. At the LMO, PO4 was low during summer (mean summer concentration: 0.17 µM) while it remained high at the K-station (mean summer concentration: 0.63 µM), explaining the lower contribution of S5.2 in the LMO. The potential PO4 limitation could also explain why at the LMO S5.2 has lower contributions during summer to the community composition compared to the K-station. However, to fully understand picocyanobacterial dynamics, other parameters such as light hours80, NH4 recycling rates33 or nutrient competition with specific phytoplankton groups77 should also be considered.
The most abundant ASV in the dataset, ASV_00001, was identical to the V5-V7 region of a metagenome-assembled genome (MAG) reconstructed from the Baltic Proper (BACL30) and has been identified as dominant in the Baltic Sea16,60. Phylogenetic classification based on six ribosomal proteins included BACL30 in the S5.2 clade16; however this classification may have been biased by the lack of genome sequenced freshwater SYN strains. In this study, ASV_00001 had 99% identity with the strain MW73D5, a freshwater strain included in clade A/B12. Most of the ASVs of the picocyanobacterial community in the Baltic Sea were more similar to freshwater strains rather than estuarine and marine strains. Hugerth et al.60 studied bacterial MAG phylogeny (including BACL30 as the only representative for SYN) from brackish environments and observed that a significant proportion of the MAGs had closest affiliation to strains of brackish origin. The affiliation of ASV_00001 to clade A/B (freshwater) suggests that Baltic Sea picocyanobacteria have evolved from freshwater strains. ASV_00001 showed high contributions during the summer, particularly at the LMO (maximum relative contribution: 78%), in line with observations in other offshore locations15,16. However, the highest contributions took place in the cold months, indicating that BACL30 is well adapted to low temperature conditions.
This study provides a detailed description of picocyanobacterial seasonal abundance and community composition during three years at a coastal and an offshore station in the Baltic Proper, showing that SYN are highly adaptable and diverse. The results link SYN populations to environmental parameters. The Baltic Sea is warming up faster than other oceans81, and it is predicted that metereological summer in the Baltic Proper will take place 20 days earlier by 210082. Longer and warmer summers could result in earlier picocyanobacterial blooms in the coast as a consequence of achieving optimal temperatures for S5.2 ecotypes earlier in the spring/summer season. This effect could be further magnified by earlier and more extensive blooms of N2-fixing cyanobacteria resulting in higher NH4 availability, which are projected as a consequence of global warming82,83. However, at offshore locations in the Baltic Proper, the picocyanobacterial summer bloom could be delayed since optimal temperature for clade A/B would take place later in the year. In future climate conditions, the contribution of picocyanobacteria to the carbon pools is expected to increase while bigger phytoplankton (PPE and other protists) are expected to decrease84,85. These changes in community composition together with potential changes in SYN peak abundances timing could result in significant alterations of picophytoplankton grazers, and consequently the rest of the trophic chain86. The results in this study highlight that besides temperature, water column stratification and nutrient availability also play an important role in picocyanobacterial dynamics and community composition.
Methods.
Field sampling
Water was sampled every second week at the Linnaeus Microbial Observatory (LMO, 56° 55′ 51.24″ N, 17° 3′ 38.52″ E, 2 m sampling depth), an offshore station located 10 km off the east coast of Öland (Sweden) and weekly at the K-station, a coastal station located in the city of Kalmar, Sweden (56° 39′ 25.4″ N 16° 21′ 36.6″ E, 1 m sampling depth) during 2018–2020. The temperature and salinity were measured using a conductivity/temperature/depth sensor CTD® Castaway at the K-station and a CTD probe (AAQ 1186-H, Alec Electronics, Japan) at the LMO. To remove large particles, samples were filtered through a 200 µm mesh.
Nutrients and Chl a
Water for measuring dissolved inorganic nutrients (NH4, NO2 + NO3 (referred to as NO3), PO4 and SiO2,) was sampled and frozen at − 20 °C until analysis using standard protocols (UV-Spectrophotometer87). For measuring Chl a, 50–200 mL seawater was filtered through duplicate 25 mm A/E glass fiber filters (∼1 μm pore size, Pall life Sciences, Ann Arbor, MI, USA). Filters were incubated overnight in darkness in 5 mL of ethanol (96%) and fluorescence was measured the following day using a fluorometer (Turner design Model #040, Tucson, USA) following the Jespersen & Christoffersen88 protocol.
Phytoplankton abundance and community composition
Samples for phytoplankton identification (> 5 µm diameter) were fixed in acid lugol (1% final concentration) and counted according to the Utermöhl method89 using an inverted microscope (Nikon TMS, Tokyo, Japan). The phytoplankton carbon biomass concentration (mg C mL−1) was derived from the cell abundance and carbon biomass89,90. In addition to the traditional taxonomy the genus Aphanizomenon, Nodularia and Dolichospermum were included in the group defined as N2-fixers.
Picophytoplankton abundance
Samples to determine picophytoplankton abundance were fixed with glutaraldehyde solution Grade I 25% in H2O (Sigma-Aldrich, Missouri, USA; 1% final concentration) and stored at − 80 °C until analysis. K-station samples until 12th May 2020 and LMO samples until 10th July 2019 were analyzed using a Cyflow® Cube8 flow cytometer (Partec®, Germany) at 10 µL s−1 while after that date a BD FACSverse (BD Biosciences) was used instead. Picophytoplankton were counted as three populations: photosynthetic picoeukaryotes (PPE), phycoerythrin rich (PE-SYN) SYN cells and phycocyanin rich (PC-SYN) SYN based on the gating described in Alegria Zufia et al.55. The observations for PPE abundance variation are presented in Supplementary Figure S1. Gating and visualization of the flow cytometric data were carried out using FCSalyzer ver. 0.9.22-alpha91.
DNA extraction, PCR amplification and sequencing
Samples for DNA analysis were collected by filtering 400 mL of water through a 0.2 µm Supor®-200 filter (Pall Corporation, USA). The filters were stored at − 80 °C until extraction. DNA was extracted using the FastDNA™ SPIN Kit for Soil from MP Biomedicals Inc. according to manufacturer´s instructions with the addition of an incubation with proteinase-K (0.02 µg/µL, final concentration) at 55 °C for one hour. Sample concentration was measured using an Invitrogen Qubit 2.0 fluorometer (Thermo Fisher Scientific Inc.). Sample purity was assessed using a Thermo Scientific™ NanoDrop 2000 spectrophotometer (Thermo Fisher Scientific Inc.). The V5-V7 hypervariable region of the 16 rRNA gene was amplified using the primers Cya-771F (5’-AGGGGAGCGAAAGGGATTA-3’) and Cya-1294R (5’-GCCTACGATCTGAACTGAGC-3’) described in62. The PCR reaction was prepared in duplicates for each sample using the Thermo Scientific Phusion High-Fidelity PCR Master Mix according to the manufacturer’s instructions with a reaction volume of 25 µL. The PCR was performed on a T100™ Thermal Cycler (BIO RAD, USA) with an initial denaturation at 98 °C for 30 s; 20 cycles of denaturation at 98 °C for 10 s, annealing at 55 °C for 1 min and extension at 72 °C for 5 s; and a final extension step at 72 °C for 2 min. Amplicon sequences were purified with AMPure XP (Beckman Coulter, USA) according to manufacturer’s instructions prior to performing the index PCR. Indexes were attached to individual samples using NEXTERA Dual indexes (Illumina Inc.) in a PCR with an initial denaturation at 98 °C for 30 s; 12 cycles of denaturation at 98 °C for 10 s, annealing at 62 °C for 30 s and extension at 72 °C for 5 s; and a final extension step at 72 °C for 2 min. Amplicon sequences were purified again after index PCR with AMPure XP (Beckman Coulter, USA). The purified amplicons were quantified with Invitrogen Qubit 2.0 fluorometer (Thermo Fisher Scientific Inc.) and pooled at equimolar concentrations. Indexed samples were sequenced with Illumina MiSeq (Illumina Inc, USA) with 300 cycle paired-end sequencing.
Bioinformatics processing
The resulting reads were denoised and screened for chimera removal with ampliseq (v1.1, https://github.com/nf-core/ampliseq) which runs on QIIME2 (2019.10)92 and DADA2 (1.10.0)93 and taxonomy assignment of the resulting amplicon sequencing variants (ASVs) were done using the SILVA 132 database with a 90% identity threshold. Among all sequences in the libraries, 95% belonged to cyanobacteria, of which 97% were classified as Synechococcales. Then, a phylogenetic tree was constructed with the aligned sequences to cross check that all of them indeed affiliated with Synechococcales; sequences that did not affiliate were excluded from further analyses. The number of sequences for each sample after each step of the quality control pipeline are specified in Supplementary Table S1.
Phylogenetic analysis
The ASVs that represented > 1% of the sequences in at least one sampling point were selected. To determine the phylogenetic clade affiliation, the closest representative sequences were identified and retrieved using the BLASTn-Search engine in the NCBI database. Sequences were aligned using MAFFT v.794 and a phylogenetic tree was constructed by the maximum likelihood method (ML) in MEGAX software95 following the GTR + G + I model (bootstrap values inferred from 1000 replicates). ASVs were associated to a specific clade or subgroup when they were located in a (mostly) monophyletic branch that contains a well defined reference sequence associated to that clade. References for clades were obtained from Huber et al.62, Crosbie et al.96, Silva et al.97, Marsan et al.98 and Choi et al.45. Branches that did not associate with previously known clades were given new clade names. The resulting tree was edited using the interactive tree of Life (iTOL, http://itol.embl.de).
Statistical analysis
Datasets were rarefied to a sequence depth of 27,819 sequences per sample. All statistical analyses were performed using R version 3.6.199 and the vegan package100. All plots were produced using the ggplot2 (3.3.6)101, and ComplexHeatmap (3.15)102.
Partial least square regression
Partial least square regression (PLS) was used to evaluate the picocyanobacterial relationship with the following independent variables: in situ nutrients (NO3 (µM), PO4 (µM), SiO2 (µM)), temperature (°C), salinity (PSU), stratification index (N2; 10–3 s−2), presence of N2-fixers (mg C mL−1) and total phytoplankton biomass (mg C mL-1). The model was performed separately for PE-SYN (cells mL−1) and PC-SYN (cells mL−1) separately and by pooling together the data from K-station and LMO. All variables were log10(x + 1) transformed for standardization. The stratification index was calculated following the Brunt-Väisälä frequency (3)103:
where g is the gravitational acceleration (m s−2), ρ is the average density of seawater (1.025 g cm−3), ρs is the surface density (g cm−3) and ρb is the density at the bottom (g cm−3). The K-station is 3 m deep, and thus all the water column is permanently in the photic zone. However, the calculated N2 was ~ 0, indicating an unstable water column. Because of this, 1 × 10–3 s−2 values were assigned to all sampling dates in the K-station, indicating a strong stratification. The variable importance in projection (VIP) was used to determine the relative importance of the independent variables considered104,105. Variables were considered significant when VIP > 1.
Procrustes analysis
Procrustes analysis was used to compare correspondence analysis (CA) based on phylogenetic clades and ASVs. The goal of this analysis was to test whether the distribution of the phylogenetic clades was congruent with the distribution of ASVs. The significance of the association of the two datasets was later explored with a squared m12 test (999 permutations, P value < 0.05).
Canonical correspondence analysis
A step-wise canonical correspondence analysis (CCA) (P value = 0.05) was performed to identify the relationship of environmental and biotic variables to SYN community clade composition. The variables considered were: NO3 (µM), PO4 (µM), SiO2 (µM), temperature (°C), salinity (PSU), stratification index (10–3 s−2), phytoplankton biomass (mg C mL−1), N2- fixers biomass (mg C mL−1), PE-SYN (cells mL−1) and PC-SYN (cells mL−1). To avoid type I errors, results were considered significant when P value after Holm correction resulted in P value < 0.05. All the independent variables were log10(x + 1) transformed for standardization.
Data availability
The datasets generated and/or analysed during the current study are available in the NCBI repository, BioProject PRJNA810944.
References
Flombaum, P. et al. Present and future global distributions of the marine Cyanobacteria Prochlorococcus and Synechococcus. PNAS 110, 9824–9829 (2013).
Honda, D. & Yokota, A. Detection of seven major evolutionary lineages in cyanobacteria based on the 165 rRNA gene sequence analysis with new sequences of five marine Synechococcus strains. J Mol Evol 48, 723–739 (1999).
Robertson, B. R., Tezuka, N. & Watanabe, M. M. Phylogenetic analyses of Synechococcus strains (cyanobacteria) using sequences of 16S rDNA and part of the phycocyanin operon reveal multiple evolutionary lines and reflect phycobilin content. Int. J. Syst. Evol. Microbiol. 51, 861–871 (2001).
Stomp, M. et al. Adaptive divergence in pigment composition promotes phytoplankton biodiversity. Nature 432, 104–107 (2004).
Albrecht, M., Pröschold, T. & Schumann, R. Identification of Cyanobacteria in a eutrophic coastal lagoon on the Southern Baltic Coast. Front. Microbiol. 8, 1–16 (2017).
Bertos-Fortis, M. et al. Unscrambling cyanobacteria community dynamics related to environmental factors. Front. Microbiol. 7, 625 (2016).
Guidi, L. et al. Plankton networks driving carbon export in the oligotrophic ocean. Nature 532, 465–470 (2016).
Hunter-Cevera, K. R. et al. Seasons of syn. Limnol. Oceanogr. 65, 1–18 (2019).
Kuosa, H. Picoplanktonic algae in the northern Baltic Sea: Seasonal dynamics and flagellate grazing. Mar. Ecol. Prog. Ser. 73, 269–276 (1991).
Sathicq, M. B., Unrein, F. & Gómez, N. Recurrent pattern of picophytoplankton dynamics in estuaries around the world: The case of Río de la Plata. Mar. Environ. Res. 161, 105136 (2020).
Rajaneesh, K. M. & Mitbavkar, S. Factors controlling the temporal and spatial variations in Synechococcus abundance in a monsoonal estuary. Mar. Environ. Res. 92, 133–143 (2013).
Crosbie, N. D., Pöckl, M. & Weisse, T. Dispersal and phylogenetic diversity of nonmarine picocyanobacteria, inferred from 16S rRNA gene and cpcBA-intergenic spacer sequence analyses. Appl. Environ. Microbiol. 69, 5716–5721 (2003).
Ernst, A., Becker, S., Wollenzien, U. I. A. & Postius, C. Ecosystem-dependent adaptive radiations ofpicocyanobacteria inferred from 16S rRNA and ITS-1 sequence analysis. Microbiology 149, 217–228 (2003).
Sánchez-Baracaldo, P., Handley, B. A. & Hayest, P. K. Picocyanobacterial community structure of freshwater lakes and the Baltic Sea revealed by phylogenetic analyses and clade-specific quantitative PCR. Microbiology 154, 3347–3357 (2008).
Hu, Y. O. O., Karlson, B., Charvet, S. & Andersson, A. F. Diversity of pico- to mesoplankton along the 2000 km salinity gradient of the Baltic Sea. Front. Microbiol. 7, 679 (2016).
Larsson, J. et al. Picocyanobacteria containing a novel pigment gene cluster dominate the brackish water Baltic Sea. ISME J. 8, 1892–1903 (2014).
Celepli, N. et al. Meta-omic analyses of Baltic Sea cyanobacteria: Diversity, community structure and salt acclimation. Environ. Microbiol. 19, 673–686 (2017).
Flombaum, P., Wang, W. L., Primeau, F. W. & Martiny, A. C. Global picophytoplankton niche partitioning predicts overall positive response to ocean warming. Nat. Geosci. 13, 116–120 (2020).
Bopp, L. et al. Multiple stressors of ocean ecosystems in the 21st century: Projections with CMIP5 models. Biogeosciences 10, 6225–6245 (2013).
Cabré, A., Marinov, I. & Leung, S. Consistent global responses of marine ecosystems to future climate change across the IPCC AR5 earth system models. Clim. Dyn. 45, 1253–1280 (2015).
Wang, T., Chen, X., Qin, S. & Li, J. Phylogenetic and phenogenetic diversity of Synechococcus along a yellow sea section reveal its environmental dependent distribution and co-occurrence microbial pattern. J. Mar. Sci. Eng. 9, 1018 (2021).
Tai, V. & Palenik, B. Temporal variation of Synechococcus clades at a coastal Pacific Ocean monitoring site. ISME J. 3, 903–915 (2009).
Ahlgren, N. A. & Rocap, G. Diversity and distribution of marine Synechococcus: Multiple gene phylogenies for consensus classification and development of qPCR assays for sensitive measurement of clades in the ocean. Front. Microbiol. 3, 1–24 (2012).
Rajaneesh, K. M., Mitbavkar, S., Anil, A. C. & Sawant, S. S. Synechococcus as an indicator of trophic status in the Cochin backwaters, west coast of India. Ecol. Indic. 55, 118–130 (2015).
Campbell, L. & Carpenter, E. J. Characterization of phycoerythrin-containing Synechococcus spp. populations by immunofluorescence. J. Plankton Res. 9, 1167–1181 (1987).
Stomp, M. et al. Colourful coexistence of red and green picocyanobacteria in lakes and seas. Ecol. Lett. 10, 290–298 (2007).
Callieri, C. & Stockner, J. G. Freshwater autotrophic picoplankton: A review. J. Limnol. 61, 1–14 (2002).
Liu, H., Jing, H., Wong, T. H. C. & Chen, B. Co-occurrence of phycocyanin- and phycoerythrin-rich Synechococcus in subtropical estuarine and coastal waters of Hong Kong. Environ. Microbiol. Rep. 6, 90–99 (2014).
Haverkamp, T. et al. Diversity and phylogeny of Baltic Sea picocyanobacteria inferred from their ITS and phycobiliprotein operons. Environ. Microbiol. 10, 174–188 (2008).
Otero-Ferrer, J. L. et al. Factors controlling the community structure of picoplankton in contrasting marine environments. Biogeosciences 15, 6199–6220 (2018).
Ploug, H. et al. Carbon, nitrogen and O2 fluxes associated with the cyanobacterium Nodularia spumigena in the Baltic Sea. ISME J. 5, 1549–1558 (2011).
Ohlendieck, U., Stuhr, A. & Siegmund, H. Nitrogen fixation by diazotrophic cyanobacteria in the Baltic Sea and transfer of the newly fixed nitrogen to picoplankton organisms. J. Mar. Syst. 25, 213–219 (2000).
Klawonn, I. et al. Untangling hidden nutrient dynamics: Rapid ammonium cycling and single-cell ammonium assimilation in marine plankton communities. ISME J. 13, 1960–1974 (2019).
Lin, Y., Wang, L., Xu, K., Huang, H. & Ren, H. Algae biofilm reduces microbe-derived dissolved organic nitrogen discharges: Performance and mechanisms. Environ. Sci. Technol. 55, 6227–6238 (2021).
Berthelot, H., Bonnet, S., Camps, M., Grosso, O. & Moutin, T. Assessment of the dinitrogen released as ammonium and dissolved organic nitrogen by unicellular and filamentous marine diazotrophic cyanobacteria grown in culture. Front. Mar. Sci. https://doi.org/10.3389/fmars.2015.00080 (2015).
Loick-Wilde, N. et al. De novo amino acid synthesis and turnover during N2 fixation. Limnol. Ocean. 63, 1076–1092 (2018).
Glibert, P. M. & Bronk, D. A. Release of dissolved organic nitrogen by marine diazotrophic cyanobacteria Trichodesmium spp.. Appl. Environ. Microbiol. 60, 3996–4000 (1994).
Kuo, J. et al. Picoplankton dynamics and picoeukaryote diversity in a hyper-eutrophic subtropical lagoon. J. Environ. Sci. Heal. 4, 521–523 (2014).
Grébert, T. et al. Light color acclimation is a key process in the global ocean distribution of Synechococcus cyanobacteria. PNAS 115, E2010–E2019 (2018).
Urbach, E., Scanlan, D., Distel, D., Waterbury, J. & Chisholm, S. Rapid diversification of marine picophytoplankton with dissimilar light-harvesting structures inferred from sequences of Prochlorococcus and Synechococcus (cyanobacteria). J. Mol. Biol. 46, 188–201 (1998).
Farrant, G. K. et al. Delineating ecologically significant taxonomic units from global patterns of marine picocyanobacteria. PNAS 113, E3365–E3374 (2016).
Rocap, G., Distel, D. L., Waterbury, J. B. & Chisholm, S. W. Resolution of Prochlorococcus and Synechococcus ecotypes by using 16S–23S ribosomal DNA internal transcribed spacer sequences. Appl. Environ. Microbiol. 68, 1180–1191 (2002).
Mazard, S., Ostrowski, M., Partensky, F. & Scanlan, D. J. Multi-locus sequence analysis, taxonomic resolution and biogeography of marine Synechococcus. Environ. Microbiol. 14, 372–386 (2012).
Huang, S. et al. Novel lineages of Prochlorococcus and Synechococcus in the global oceans. ISME J. 6, 285–297 (2011).
Choi, D. H. & Noh, J. H. Phylogenetic diversity of Synechococcus strains isolated from the East China Sea and the East Sea. FEMS Microbiol. Ecol. 69, 439–448 (2009).
Lee, M. D. et al. Marine Synechococcus isolates representing globally abundant genomic lineages demonstrate a unique evolutionary path of genome reduction without a decrease in GC content. Environ. Microbiol. 21, 1677–1686 (2019).
Paerl, R., Foster, R., Jenkins, B., Montoya, J. & Zehr, J. Phylogenetic diversity of cyanobacterial narB genes from various marine habitats. Environ. Microbiol. 10, 3377–3387 (2008).
Fuller, N. et al. Clade-specific 16S ribosomal DNA oligonucleotides reveal the predominance of a single marine Synechococcus clade throughout a stratified water column in the Red sea. Appl. Environ. Microbiol. 69, 2430–2443 (2003).
Scanlan, D. J. et al. Ecological genomics of marine Picocyanobacteria. Microbiol. Mol. Biol. Rev. 73, 249–299 (2009).
Mazard, S., Wilson, W. H. & Scanlan, D. J. Dissecting the physiological response to phosphorus stress in marine Synechococcus isolates (cyanophyceae). J. Phycol. 48, 94–105 (2012).
Li, J. et al. Synechococcus bloom in the Pearl River Estuary and adjacent coastal area –With special focus on flooding during wet seasons. Sci. Total Environ. 692, 769–783 (2019).
Zwirglmaier, K. et al. Global phylogeography of marine Synechococcus and Prochlorococcus reveals a distinct partitioning of lineages among oceanic biomes. Environ. Microbiol. 10, 147–161 (2008).
Sohm, J. A. et al. Co-occurring Synechococcus ecotypes occupy four major oceanic regimes defined by temperature, macronutrients and iron. ISME J. 10, 333–345 (2016).
Bunse, C. et al. High frequency multi-year variability in Baltic Sea microbial plankton stocks and activities. Front. Microbiol. 10, 1–18 (2019).
Alegria Zufia, J., Farnelid, H. & Legrand, C. Seasonality of coastal picophytoplankton growth, nutrient limitation and biomass contribution. Front. Microbiol. 12, 1–13 (2021).
Granéli, E., Wallström, K., Larsson, U., Granéli, W. & Elmgren, R. Nutrient limitation of primary production in the Baltic Sea Area. Ambio 19, 142–151 (1990).
Mazur-Marzec, H. et al. Occurrence of cyanobacteria and cyanotoxin in the Southern Baltic Proper. Filamentous cyanobacteria versus single-celled picocyanobacteria. Hydrobiologia 701, 235–252 (2013).
Stal, L. et al. BASIC: Baltic Sea cyanobacteria. An investigation of the structure and dynamics of water blooms of cyanobacteria in the Baltic Sea—Responses to a changing environment. Cont. Shelf Res. 23, 1695–1714 (2003).
Herlemann, D. P. et al. Transitions in bacterial communities along the 2000 km salinity gradient of the Baltic Sea. ISME J. 5, 1571–1579 (2011).
Hugerth, L. W. et al. Metagenome-assembled genomes uncover a global brackish microbiome. Genome Biol. 16, 1–18 (2015).
Walve, J. & Larsson, U. Seasonal changes in Baltic Sea seston stoichiometry: The influence of diazotrophic cyanobacteria. Mar. Ecol. Prog. Ser. 407, 13–25 (2010).
Huber, P. et al. Primer design for an accurate view of picocyanobacterial community structure by using high-throughput sequencing. Appl. Environ. Microbiol. 85, 1–17 (2019).
Jiang, T. et al. Temporal and spatial variations of abundance of phycocyanin- and phycoerythrin-rich Synechococcus in Pearl River Estuary and adjacent coastal area. J. Ocean Univ. China 15, 897–904 (2016).
Li, S. et al. Unexpected predominance of photosynthetic picoeukaryotes in shallow eutrophic lakes. J. Plankton Res. 38, 830–842 (2016).
Collos, Y. et al. Oligotrophication and emergence of picocyanobacteria and a toxic dinoflagellate in Thau lagoon, southern France. J. Sea Res. 61, 68–75 (2009).
Bec, B., Husseini-Ratrema, J., Collos, Y., Souchu, P. & Vaquer, A. Phytoplankton seasonal dynamics in a Mediterranean coastal lagoon: Emphasis on the picoeukaryote community. J. Plankton Res. 27, 881–894 (2005).
Hunter-Cevera, K. R. et al. Physiological and ecological drivers of early spring blooms of coastal phytoplankter. Science 354, 326–329 (2016).
Albertano, P., Di Somma, D. & Capucci, E. Cyanobacterial picoplankton from the central Baltic Sea: Cell size classification by image analyzed fluorescence microscopy. J. Plankton Res. 19, 1405–1416 (1997).
Paulsen, M. L. et al. Synechococcus in the Atlantic gateway to the Arctic Ocean. Front. Mar. Sci. https://doi.org/10.3389/fmars.2016.00191 (2016).
Felföldi, T. et al. Diversity and seasonal dynamics of the photoautotrophic picoplankton in Lake Balaton (Hungary). Aquat. Microb. Ecol. 63, 273–287 (2011).
Grinienė, E., Šulčius, S. & Kuosa, H. Size-selective microzooplankton grazing on the phytoplankton in the Curonian Lagoon (SE Baltic Sea). Oceanologia 58, 292–301 (2016).
Tsai, A. Y., Gong, G. C., Huang, Y. W. & Chao, C. F. Estimates of bacterioplankton and Synechococcus spp. mortality from nanoflagellate grazing and viral lysis in the subtropical Danshui River estuary. Estuar. Coast. Shelf Sci. 153, 54–61 (2015).
Camacho, A., Miracle, M. R. & Vicente, E. Which factors determine the abundance and distribution of picocyanobacteria in inland waters? A comparison among different types of lakes and ponds. Arch. Hydrobiol. 157(321), 338 (2003).
Berry, D. L. et al. Shifts in cyanobacterial strain dominance during the onset of harmful algal blooms in Florida Bay, USA. Microb. Ecol. 70, 361–371 (2015).
Zborowsky, S. & Lindell, D. Resistance in marine cyanobacteria differs against specialist and generalist cyanophages. PNAS 116, 16899–16908 (2019).
Wall, C. C., Rodgers, B. S., Gobler, C. J. & Peterson, B. J. Responses of loggerhead sponges Spechiospongia vesparium during harmful cyanobacterial blooms in a sub-tropical lagoon. Mar. Ecol. Prog. Ser. 451, 31–43 (2012).
Glibert, P. M. et al. Pluses and minuses of ammonium and nitrate uptake and assimilation by phytoplankton and implications for productivity and community composition, with emphasis on nitrogen-enriched conditions. Limnol. Oceanogr. 61, 165–197 (2016).
Herbert, R. A. Nitrogen cycling in coastal marine ecosystems. FEMS Microbiol. Rev. 23, 563–590 (1999).
Cai, J., Hodoki, Y. & Nakano, S. I. Phylogenetic diversity of the picocyanobacterial community from a novel winter bloom in Lake Biwa. Limnology 22, 161–167 (2021).
Guyet, U. et al. Synergic effects of temperature and irradiance on the physiology of the marine Synechococcus strain WH7803. Front. Microbiol. 11, 1707 (2020).
Meier, H. E. M. et al. Ensemble modeling of the Baltic Sea ecosystem to provide scenarios for management. Ambio 43, 37–48 (2014).
Neumann, T. et al. Extremes of temperature, oxygen and blooms in the baltic sea in a changing climate. Ambio 41, 574–585 (2012).
Andersson, A. et al. Projected future climate change and Baltic Sea ecosystem management. Ambio 44, 345–356 (2015).
Schmidt, K. et al. Increasing picocyanobacteria success in shelf waters contributes to long-term food web degradation. Glob. Change Biol. 26, 5574–5587 (2020).
Legrand, C. et al. Interannual variability of phyto-bacterioplankton biomass and production in coastal and offshore waters of the Baltic Sea. Ambio 44, 427–438 (2015).
Capuzzo, E. et al. A decline in primary production in the North Sea over 25 years, associated with reductions in zooplankton abundance and fish stock recruitment. Glob. Change Biol. 24, e352–e364 (2017).
Valderrama, J. C. Methods of nutrient analysis. In Manual on Harmful Marine Microalgae (eds Hallagraeff, G. M. et al.) 251–268 (IOC Manuals and Guides, 1995).
Jespersen, A. M. & Christoffersen, K. Measurements of chlorophyll-a from phytoplankton using ethanol as extraction solvent. Arch. Hydrobiol. 109, 445–454 (1987).
Edler, L. Recommendations on methods for marine biological studies in the Baltic Sea. Phytoplankton and chlorophyll (Baltic Marine Biologists BMB (Sweden), 1979).
HELCOM Phytoplankton Expert Group. Phytoplankton biovolume and carbon content. https://www.ices.dk/data/Documents/ENV/PEG_BVOL.zip (2013).
Mostböck, S. FCSalyzer (2021).
Gregory Caporaso, J. et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods https://doi.org/10.1038/nmeth.f.303 (2010).
Callahan, B. et al. DADA2: High-resolution sample inference from Illumina amplicon data. Nat. Methods 13, 581–583 (2016).
Katoh, K., Misawa, K., Kuma, K. I. & Miyata, T. MAFFT: A novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res. 30, 3059–3066 (2002).
Kumar, S., Stecher, G., Li, M., Knyaz, C. & Tamura, K. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 35, 1547–1549 (2018).
Crosbie, N. D., Pöckl, M. & Weisse, T. Rapid establishment of clonal isolates of freshwater autotrophic picoplankton by single-cell and single-colony sorting. J. Microbiol. Methods 55, 361–370 (2003).
Silva, C. S. P., Genuário, D. B., Vaz, M. G. M. V. & Fiore, M. F. Phylogeny of culturable cyanobacteria from Brazilian mangroves. Syst. Appl. Microbiol. 37, 100–112 (2014).
Marsan, D., Wommack, K. E. & Ravel, J. Draft genome sequence of Synechococcus sp. strain CB0101, isolated from the Chesapeake Bay estuary. Genome Announc. 2, e01111 (2014).
R Core Team. R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing, Vienna, Austria. R version 3.5.1. https://www.r-project.org/ (2019).
Oksanen, J. et al. Package ‘vegan’ (2020).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis (Springer-Verlag, New York, 2016) (ISBN 978-3-319-24277-4).
Gu, Z., Eils, R. & Schlesner, M. Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics 32, 2847–2849 (2016).
Gill, A. E. Atmosphere-Ocean Dynamics (Academic Press, USA, 1982).
Li, X., Wang, Y., Li, J. & Lei, B. Physical and socioeconomic driving forces of land-use and land-cover changes: A Case Study of Wuhan City, China. Discret Dyn. Nat. Soc. 2016 (2016).
Paliy, O. & Shankar, V. Application of multivariate statistical techniques in microbial ecology. Mol. Ecol. 25, 1032–1057 (2016).
Acknowledgements
We would like to thank Anders Månsson, Soran Mahmoudi, Maurice Hirwa, Christien Laber and Laura Bas Conn for assistance with the K-station sampling, Kalmar Kommun for access to Kallbadhuset where the K-station is located, Clara Pérez Martínez, Kristofer Bergström, Sabina Arnautovic, Camilla Karlsson, Emil Fridolfsson and Benjamin Pontiller for sampling at the LMO, EON, Northern Offshore Services (NOS), and the Provider crew for technical assistance during sampling at the LMO, Sonia Brugel from Umeå University for flow cytometry counts, Jasmine Weber-Pierson and Thi Quyen Nham for the nutrient analysis, Justyna Kobos and Lidia Nawrocka from University of Gdansk for the microscopic phytoplankton counts, Anabella Aguilera for assistance with the phylogenetic trees construction and Daniel Lundin and Emelie Nilsson for assistance on the genetic data analysis. Sequencing was performed by the SNP&SEQ Technology Platform in Uppsala. The facility is part of the National Genomics Infrastructure (NGI) Sweden and Science for Life Laboratory. The SNP&SEQ Platform is also supported by the Swedish Research Council and the Knut and Alice Wallenberg Foundation.This work was supported by The Swedish Research Council FORMAS (2017-00468 to HF and the Swedish Strategic programme Ecochange-Climate change and the Baltic Sea ecosystem to CL) and Anna-Greta and Holger Crafoord Foundation (CR2019-0012) to HF.
Funding
Open access funding provided by Linnaeus University.
Author information
Authors and Affiliations
Contributions
H.F., C.L., and J.A.Z. conceived the study. J.A.Z. and H.F. conducted the field sampling. J.A.Z. performed the laboratory and data analysis and plotted the figures. J.A.Z., C.L., and H.F. wrote and edited the manuscript. All authors contributed to the article and approved the submitted version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zufia, J.A., Legrand, C. & Farnelid, H. Seasonal dynamics in picocyanobacterial abundance and clade composition at coastal and offshore stations in the Baltic Sea. Sci Rep 12, 14330 (2022). https://doi.org/10.1038/s41598-022-18454-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-18454-8
This article is cited by
-
Multiyear analysis uncovers coordinated seasonality in stocks and composition of the planktonic food web in the Baltic Sea proper
Scientific Reports (2023)
-
Seasonality and Strain Specificity Drive Rapid Co-evolution in an Ostreococcus-Virus System from the Western Baltic Sea
Microbial Ecology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.