Abstract
Exposure to community reservoirs of gram-negative antibiotic-resistant bacteria (GN-ARB) genes poses substantial health risks to individuals, complicating potential infections. Transmission networks and population dynamics remain unclear, particularly in resource-poor communities. We use a dynamic compartment model to assess GN-ARB transmission quantitatively, including the susceptible, colonised, infected, and removed populations at the community-hospital interface. We used two side streams to distinguish between individuals at high- and low-risk exposure to community ARB reservoirs. The model was calibrated using data from a cross-sectional cohort study (N = 357) in Chile and supplemented by existing literature. Most individuals acquired ARB from the community reservoirs (98%) rather than the hospital. High exposure to GN-ARB reservoirs was associated with 17% and 16% greater prevalence for GN-ARB carriage in the hospital and community settings, respectively. The higher exposure has led to 16% more infections and attributed mortality. Our results highlight the need for early-stage identification and testing capability of bloodstream infections caused by GN-ARB through a faster response at the community level, where most GN-ARB are likely to be acquired. Increasing treatment rates for individuals colonised or infected by GN-ARB and controlling the exposure to antibiotic consumption and GN-ARB reservoirs, is crucial to curve GN-ABR transmission.
Similar content being viewed by others
Introduction
The emergence and spread of antimicrobial-resistant microorganisms, particularly gram-negative (GN) antibiotic-resistant bacteria (ARB), affect population health globally1,2,3. Infections due to ARB are associated with significant disease burden, including higher mortality, longer hospitalisations, and increased health cost4,5. The worldwide emergence of resistance has occurred primarily due to antibiotic misuse and overconsumption6. The situation is even more critical in low-and middle-income countries (LMICs), where often no prescription is needed, and lack of access to novel compounds is common7,8,9. Moreover, the increase in demand for livestock products in LMICs has resulted in intensive use of antimicrobials in animal production, leading to an increase in ARB10,11. According to the Organization of Economic Cooperation and Development (OECD), the higher burden of GN-ARB faced by LMICs is associated with various factors including weaker regulations on the food industry and antibiotic use in humans or animals, inadequate water, sanitation, and hygiene infrastructure, and high contamination and concentration of environmental pollutants12,13,14.
GN bacteria impose a higher risk to public health than gram-positive pathogens, as they develop ARB faster, and there are fewer therapeutic alternatives available to manage these infections15,16,17. GN bacteria harbour a myriad of mechanisms to avoid antibiotics' action on their cell structure, such as the presence of degradational enzymes, efflux pumps, and membrane permeability18. The danger is imminent when facing infections caused by these pathogens, and particularly when dealing with bloodstream infections (BSI). Estimates suggest that BSI from ARB causes 33,000 annual deaths worldwide, the majority of which are caused by GN bacteria19. Most BSIs occur in the healthcare settings, particularly in intensive care units (ICUs)2,20. However, ARB infections, including community-onset BSIs21, are becoming increasingly relevant at the community level due to various factors, including inadequate antibiotic use, clonal dissemination, and community transmission networks and reservoirs, such as crowded households and workplaces, and educational facilities22.
Using a One Health approach, researchers in the past decade have drawn attention to the elevated exposure to environmental sources of transmission as one of the most relevant factors for the emergence and spread of GN-ARB in the community11,23,24,25,26,27,28,29,30,31,32,33,34. There is an ever-more evident connection between ARB in humans and environmental risk factors such as the interaction with animals (pets and livestock), food production and pesticides, poor waste management, contaminated water, and living conditions including crowding, pollution, and inadequate water, hygiene, and sanitation infrastructure13,35. All of these pose a high risk of transmission and disease burden, especially amongst the most disadvantaged populations where daily life might be easily jeopardised36,37. Furthermore, the existing disparate within-country health and sociodemographic inequalities in LMICs aggravate the situation. Indeed, there is a sizeable variation in ARB levels due to the context-specific high (hotspots) or moderate/low transmission risks within rural and urban settlements38.
The transmission of ARB and its human population dynamics have been modelled at the community level via SEIHS-adjusted (Susceptible, Exposed or colonised, Infected, and Hospitalised populations) compartmental models looking at in-human colonisation and infection24,25,27,39,40,41,42,43,44,45. Most existing literature considers antibiotic consumption as the primary driver of ARB39,40,41. However, recent population-based studies have examined interactions between healthcare settings and communities, suggesting that ARB are primarily acquired in the community when there is a significant presence of ARB reservoirs and, therefore, high transmission risks25,27. To the best of our knowledge, no modelling framework has considered disadvantaged populations, which account for about 80% of the global population, and this study aims to do that.
Specifically, we used a compartmental-based dynamic mathematical model to quantitatively characterise the dynamics of the transmission of GN-ARB in a rural community by looking at high and low risks of exposure to ARB reservoirs. We focused on Molina, a resource-poor peri-urban agricultural community in the south of Chile (Fig. 1). Previous research has shown a high prevalence of GN-ARB, specifically 40%, 29%, and 21% for quinolones-, third-generation cephalosporins-resistant, and carbapenem-resistant GN bacteria, respectively46. These estimated prevalence are substantially above OECD’s estimates for Chile (32%, 28%, and 17%, respectively)14. Consequently, we examined low and high GN-ARB transmission risk scenarios to help inform interventions to reduce the emergence of GN-ARB in a resource-limited community setting.
Results
Human population analysis
Figure 2 shows the population dynamics for susceptible and colonised individuals in the community in the low-risk (panel A) and the high-risk (panel B) scenarios, and individuals colonised at the hospital, or infected in the community or the hospital in the low risk (Panel C) and high-risk (Panel D) scenarios. Our results suggest that ARB was primarily acquired in the community with a deficient proportion of inpatients with hospital-acquired GN-ARB colonisation and infection (Fig. 2, and Figs. D1–D5, supplementary material). However, the numbers increase under high exposure to ARB reservoirs within the community and hospitalised individuals. Even though transmission rates within the community are substantially lower than in the hospital, the entry rate of individuals into the hospital is meaner. Therefore, the model suggests that the dissemination of pathogens occurs vastly in the community posing a greater number of colonised individuals and disease burden due to the higher population size and direct/indirect person-to-ARB reservoirs contact. Our model suggests that 98% of the total GN-ARB acquisition (colonisation) occurs in the community, compared to 1% in the hospital, for the high- and low-risk scenarios. Even though transmission constitutes greater absolute numbers in the community, the percentage of people having hospital-acquired BSIs (HHH) from those colonised by GN-ARB within the hospital (ZH) was 3.36%, compared to 0.06% for the same definitions within the community (i.e., [IC + IHC]/ZC). Finally, community-acquired infections (IC and IHC) represented 97.6% of the total population of individuals having BSIs; however, the community mortality burden was low (7.8%), compared to the hospital (92.2%) where most individuals are treated for BSIs.
Population dynamics for individuals colonised or infected in the community or the hospital, by risk scenario. Notes: Panels (A) and (B) show the susceptible and colonised individuals in the community in the low-risk and high-risk scenarios, respectively. Panels (C) and (D) show individuals colonised at the hospital, or infected in the community or the hospital in the low risk and high-risk scenarios, respectively. ✵ stands for IHH(t). Complete results of the population dynamics by compartment with their respective 95% CIs are presented in the supplementary material, Figs. D2-5.
Population dynamics by risk-scenario
Figure 3 depicts the proportion of the total population (N) over time, by compartment and risk scenario (see Fig. D5, supplementary material for model specific results at the end of the study period). Compared to the low-risk scenario, the proportion of susceptible population rapidly decreased under the high-risk scenario posing a higher disease burden (Fig. 3, panel A). Higher exposure to ARB reservoirs (27% greater derived from the \(\rho\) coefficient) translates into an increased proportion of ZC under the high-risk scenario, which exceeded the low-risk comparison group by 10 raw percentage points (16% higher population size; see Fig. 3 panel C). Consequently, the susceptible population was reduced by 21% after contrasting both groups (e.g., {ZC High-risk − ZC Low-risk}/ZC Low-risk). The rest of the compartments presented less than 2% of the total population in the system modelled (Fig. 3, panel B, D, E and F). Compared to the low-risk scenario, the results show a significant variation over the number of hospitalised individuals colonised by GN-ARB (ZH was 17.4% higher, Fig. 3 panel D), hospital- and community-acquired infections (IHC and IHH were 16.1% and 16.0% higher, respectively), and deaths (removed) (15.9% greater) in the high-risk scenario (Fig. 3, panel B).
Proportion of the population (N) per compartment and overtime, by risk group. Notes: All groups coloured in light blue (or dark) sum up 100% of the population (N) including those removed. The proportions were calculated from the main model over the end of the study period (also see Fig. D5, supplementary material). S: Susceptible population, ZC: Colonised individuals by a GN-ARB in the community; RC: Individuals removed (dead due to GN-ARB BSI in the community); RH: Individuals removed (dead due to GN-ARB BSI in the hospital). IH: Individuals with a GN-ARB BSI in the hospital, that is comprised of IHH: Individuals with a hospital-acquired GN-ARB BSI; IHC: Individuals with a community-acquired GN-ARB BSI; IH: Individuals with a GN-ARB BSI in the community; ZH: Colonised individuals by a GN-ARB in the hospital. Figure D5.2 (supplementary material) shows the proportion of the total population by compartment and risk scenario at the end period. Y-axes stand for each specific compartment proportion to the total population (e.g., the y-axis in panel A stands for the proportion of the total population consisting of S(t)).
Sensitivity analyses
Our sensitivity analyses showed that the most influential parameters increasing GN-ARB transmission were hospitalisation rate (\({\delta }_{I}\)), treatment rate for hospitalised individuals having BSIs (\({\omega }_{H}\)), and the probability of a having a GN-ARB BSI in the community (\({\xi }_{C}\)). Diversely, the probability of having a GN-ARB BSI in the hospital (\({\xi }_{H}\)) was one of the less influential parameters determining GN-ARB transmission.
In univariate analyses, we observed that a 0.1 increase over the hospitalisation rate (\({\delta }_{I}\)=0.9, compared to baseline \({\delta }_{I}\)=0.8) produced 9% and 10% fewer GN-ARB BSIs within the community under high and low-risk scenarios, and 1.6- and 0.6-times fewer deaths attributed to GN-ARB BSIs, respectively (Fig. D6.1–2, supplementary material). Moreover, varying the probability of having a BSI in the community (\({\xi }_{C}\)) had a directly proportional effect (linear) on the number of BSIs in the community and hospital settings (i.e., two times higher \({\xi }_{C}\) is translated into two times greater number of individuals having BSIs) and the attributable number of deaths, regardless of the risk scenario (Fig. D7.1–2, supplementary material).
Conversely, increasing (\({\xi }_{H}\)) had a negligible effect on the number of hospital-acquired infections and no effect on the total number of infections within the hospital (including community- and hospital-acquired infections; Fig. D8, supplementary material). Similarly, improving treatment rates by 25% (\({\omega }_{H}\)) within the hospital setting reduced the prevalence of hospital-acquired infections by 19% under both risk scenarios (Fig. D9, supplementary material). Our results also showed that larger clearance rates (\(\gamma\)) support bacterial decolonisation, highly favouring the low-risk scenario (e.g., a 25% improvement over \(\gamma\) produced a 15.5% decrease in BSIs in the community, compared to 13.5% under the high-risk scenario: Fig. D10, supplementary material).
Finally, we analysed different hypothetical cases for the transmission parameter (\(\beta\)C) which highly impacts the susceptible and colonised populations (Fig. D11, supplementary material). Greater transmission rates had a non-substantial effect on the crude numbers of individuals having BSIs (six to eight more individuals with BSIs observed if \(\beta\)C is ten times greater). Even though these crude numbers are not significant in magnitude, we noted that higher \(\beta\)C is associated with a higher marginal increase of the number of BSIs under the low risk than the high-risk scenario. For example, a 25% higher \(\beta\)C is translated into a 16.6% and 10% greater number of individuals with BSIs in the hospital, respectively.
In the bivariate analyses, we found that individuals having community-acquired BSIs would be reduced to the minimum (15 cases on average) if hospitalisation rates for people with BSIs are improved or in the range of 60% < \({\delta }_{I}\)<100% and \({\xi }_{C}\) is diminished or between 0.2*\({\xi }_{C}\) and 1.5*\({\xi }_{C}\) (Fig. 4, panels A and B). \({\xi }_{C}\) is directly related to the greater number of community-acquired even if profound changes are seen over the spontaneous clearance or treatment rate received for BSIs within the community (\({\omega }_{C}\)) (Fig. 4, panels C and D). However, when it comes to hospitalised patients, the treatment rate for BSIs (\({\omega }_{H}\)) directly affect the number of individuals with BSIs (greater impact over community-acquired), despite of \({\xi }_{H}\), and displaying the highest burden for those individuals at high-risk (Fig. 4, panels E and F).
Bivariate analyses result of the main parameters on the number of infections in the community (panel A–D) and hospital (panel E–F), by risk group. Notes: A and B show a bivariate analysis of the hospitalisation rate for individuals with a GN-ARB BSI and the probability of developing GN-ARB BSI, both in the community, on the number of individuals having GN-ARB BSI in the community. Figures (C) and (D) depict the variation of the probability of GN-ARB BSI and the spontaneous clearance (or treatment rate), both in the community. Figures (E) and (F) display the bivariate relationship between the probability of GN-ARB and treatment rate, both in the hospitals, on the number of people having GN-ARB in the same setting. Lighter (yellow) colours mean a higher number of people infected by GN-ARB, whereas darker (blue) colours mean lower.
In antibiotic-specific models, we found a 9.9% higher percentage of people colonised and infected by GN quinolone-resistant bacteria in the community over time, compared to our base GN-ARB model for the high-risk group (Fig. E5, bottom panel). Whereas it was 1.6% greater for cephalosporins-resistant GN bacteria, and 13.4% lower for carbapenem-resistant GN bacteria. Among the different bacterium transmission parameters surveyed from the literature, we identified minor variations after accounting for the Enterobacterales family and other than Enterobacterales, including Acinetobacter baumanii, specific transmission dynamics (observed variation < 0.0017% compared to our base GN-ARB model; supplementary material, Fig. E6).
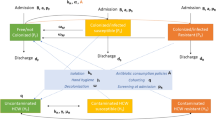
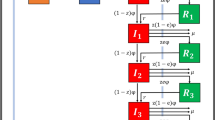
Compartment model for GN-ARB community transmission. Notes: Tables 1 and 2 show baseline conditions and parameter specifications of the compartment model. Subscript “H” stands for hospital population whereas “C” for community. IH is divided into IHH and IHC for hospital- and community-acquired BSIs. The compartment S indicates that the whole cohort is susceptible to antibiotics because gram-negative bacteria are an essential part of human gut microbiota and other mucosal surfaces. Infectiousness (I) occurs after being colonised (Z) by resistant bacteria. Individuals are immediately transferred to the susceptible S(t) disease-free population compartment after full clearance of infection. The model assumes that people can be infected only by one type of resistant bacterium simultaneously, which cannot evolve. (R) compartments are for removed individuals (death).
Discussion
Our results suggest that most GN-ARB acquisition (colonisation) is observed in the community, causing a substantial impact on the number of infections and mortality, particularly among those individuals at high-risk exposure to ARB reservoirs. This result is consistent with the literature on transmission dynamics in disadvantaged and rural communities, where inadequate access to clean water, sanitation, and hygiene infrastructure and higher exposure to pollution and GN-ARB reservoirs impose a considerable risk to the health system and community health47,48,49,50.
Similar to a recent study25, most of the human acquisition of ARB is related to the large number of people agglomerated in community settings (~ 98% of the total population), where close contact is common, including crowded households, educational facilities, and agricultural workplaces. For instance, the turnover of the hospital population in Molina was about 0.000054 per day, according to the Chilean Ministry of Health51, which evidences the reduced patients’ inflow. Our study expands from previous modelling studies24,27,45,52,53 in two ways. Firstly, it includes the acquisition of ARB using a broader population perspective while incorporating the hospital and community populations from a recently studied rural middle-income community. This is in contrast to most of the existing mathematical models, which are largely focused on high-income urban areas. Secondly, we generate two risk scenarios for community reservoirs and analyse the contribution of ARB carriage and burden associated, which differs from previous efforts that mainly analyse hospital population dynamics and transmission risks in that particular setting26,27,42,43,44.
Why does the high-risk scenario pose a greater exposure to ARB reservoirs?
Compared to the low-risk scenario, the model results showed that the high-risk scenario resulted in 1.27 times higher ARB rate due to higher exposure to GN-ARB reservoirs or risk factors in the community, consistent with the literature30,54,55,56,57,58,59,60,61. We found that high-risk exposure exacerbated the disease burden produced by GN-ARB in the community and hospital. The main risk factors in the community were previous antibiotic consumption and overcrowded spaces. Increased community-level antibiotic consumption has been associated with inappropriate antibiotic dispensing and misuse54, specifically in rural and most deprived areas where poor health determinants exacerbate the development and acquisition of resistant strains55. Crowded households and higher animal contact increase the exposure of susceptible populations to colonisation with ARB56. However, other local factors previously documented in Molina might influence these figures secondarily60. For instance, antibiotic consumption in food-producing animals in Chile has been estimated at 77 mg per animal population unit (cattle, pigs, or chicken)11 with 24.8% ARB prevalence among them58, posing greater risks at the human-animal interface and specifically to rural agricultural areas such as Molina57. Also, the high mass of agriculture-related workers who are frequently exposed to animal contact and chemical hazards (e.g., toxic substances, fertilisers, and pesticides) may result in selective pressure that promotes the development of antimicrobial resistance among specific bacterial populations59.
Sensitivity analyses
Our model's key parameters that modify ARB acquisition and infections are the transmission parameter, hospitalisation rates, exposure to ARB reservoirs, and bacterial spontaneous clearance rates. Whereas higher treatment rates within hospitalised individuals and probability of developing community-acquired BSIs increased considerably the number of infections and mortality burden in the hospital. ARB carriage could be mitigated controlling transmission rates (e.g., 25% increase in β indicates 15.5% higher number of individuals colonised by GN-ARB), which affect the number of infections in the community (12% increase) and mortality.
If appropriate healthcare for GN-ARB BSIs was improved at the hospital, the number of subsequent infections would be substantially lower regardless of the risk scenario. A two-fold improvement in targeting bacterial clearance and treatment in the community would reduce colonisation and infection rates in the community and hospital settings by decreasing the prevalence of BSIs by 50–75%, with a higher impact in the low-risk scenario (75% decrease).
Quinolone-resistant GN bacteria posed the greatest burden, according to our study results. A 9.9% higher prevalence of colonised and infected people was observed compared to our base GN-ARB model results. Quinolones were introduced in Chile between 1998 and 2015, primarily for veterinary usage, including aquaculture and agriculture62. Recent reports showed excessive use and resistance genes among residues in marine sediments, which has contributed to a larger number of human pathogens isolated in those areas63.
Interventions and policy
Firstly, if reducing ARB colonisation is our goal, interventions should target exposure in the community. Previous studies have described interventions that may be useful locally to reduce transmission and carriage64,65,66. Through awareness campaigns by employing large-scale educational and stewardship programs, changes in human behaviour may decrease the spread of GN-ARB in the community64. In particular, the development and guidance of prescription standards to avoid unnecessary consumption (overconsumption and misuse) and over-the-counter sales of antibiotics is paramount27.
It is essential to prevent new cases through active community surveillance programs and mitigate the potential risks of environmental and hospital exposures, including antibiotic use in food-producing animals and contaminated food61,65,66,67. Applying a One Health approach may help control the transfer of ARB genes at the animal-human interface (direct or indirect contact with livestock or food chain), because clonal dissemination plays an important role in the spread of ARB pathogens. Research on the specific transmission mechanisms is urgently needed to design more effective interventions to reduce ARB carriage in humans68. Less acquisition of ARB in the community and hospital settings is also facilitated with higher rates of clearance, suggesting that community-based (or post-discharge) decolonisation schemes would bottom down ARB prevalence shifting the transmission dynamics over time. For instance, improving the adherence to hand hygiene has been cost-effective in reducing the acquisition of GN-ARB (carriage) by 10% in a 6-month period69. Additionally, universal screening for incoming patients has been cost saving (compared to doing nothing) to reduce ABR carriage in hospitals where ARB incidence levels are above 0.3%70, which represents the case of the community hospital examined in the present study.
Secondly, if reducing mortality attributed to GN-ARB BSI is the focus, increasing healthcare access and quality (enhanced accessibility and active surveillance) and improving treatment success rates for BSIs within the hospital are critical. In particular, early-stage identification (and testing capability) through a rapid response from healthcare facilities might favour the early prediction of the clinical progression of BSIs in communities at high risk right before the disease burden is aggravated71,72. The risk of BSIs is notably greater in low-and-middle-income communities due to under-resourced infrastructure to diagnose and quantify these complications. Current standards to detect GN-ARB BSIs in hospitals are limited to automated blood culture systems within laboratories having reduced capacity, which may take up to 2–4 days73. New rapid diagnostic assays are available for identifying and detecting the causative organism and its susceptibility, producing earlier outcomes than conventional phenotypic susceptibility and subculture testing74. However, their high cost is frequently a major hurdle to their implementation in low-resource settings such as Molina.
Limitations, assumptions, and strengths of the study
The article has some limitations. First, there is a coexistence of drug-susceptible and drug-resistant GN bacteria calibrated to existing data and literature parameters that are pathogen-specific, which might differ from less transmissible pathogens and organisms’ aetiology. However, no other community-based study provides information on ARB for other pathogens-ATB combination pairs in Chile. Second, our model was calibrated based on a small sample of study participants. This may affect the model results because of the high variability in the computation of some of the parameters, potentially introducing biases. Also, age-specific or sociodemographic stratification was not included due to sample constraints. Nevertheless, this is the first study in a middle-income community where ARB was tested by collecting and analysing faecal samples for GN-ARB colonisation in humans. Third, literature is absent on GN-ARB in most disadvantaged communities, so the main parameters were primarily obtained from studies focusing on middle and high-income countries. This may have underestimated the effects over population dynamics, but no further evidence is available, and data were provided from high-quality and reliable sources.
We assumed spontaneous clearance of ABR was equivalent among individuals in the community and the hospital. However, clearance rates might be higher in the community because hospitalized patients are ill or may have weaker immune systems.
One critical parameter is the hospitalisation rate for people with GN-ARB BSI in a rural community because it is highly sensitive to the prevalence of untreated BSI. We assumed this to be 80% considering that BSI required immediate attention75. This parameter could be lower in low-income settings with limited access to healthcare, but we did not have data to test this assumption. Our estimates should be considered conservative; if hospitalization rates were lower, the disease burden would be even higher than estimated.
We assumed that transmission rates in the community are lower than in the hospital, but that ARB reservoirs exposure is higher. For instance, our study suggests that greater consumption of β-lactam antibiotics, which have been incrementally introduced in the community46, might be associated with a higher prevalence of community-acquired ARB acquisition. This may change if other pathogens are introduced into the community. For example, co-colonization with clinically relevant ARB-pathogens might occur, favouring horizontal gene transfer and further dissemination of ARB. However, we focused on the most likely scenarios to occur. Also, we could not perform pathogen-specific models because gram staining was primarily used in the parallel study. Nevertheless, we sourced transmission parameters for different GN-ARB types and did not observe meaningful variations over time.
Our study's main strengths include using a novel approach to account for the high risk of exposure to GN-ARB reservoirs in the community by considering their indirect interaction with the environment, hospitals, and animals as risk variables. We used an existing and transparent quantitative framework, including extensive sensitivity analyses, to cover a broader range of scenarios where people may acquire GN-ARB. Interventions are also suggested depending on the problem to be targeted. Specifically, antibiotic decolonisation in extensive therapies might be highly cost-effective in high-risk and limited-resourced scenarios to account for a reduced transmission and burden of disease in the future76. This study forms part of the evidence base required to prioritise new strategies to ease GN-ARB transmission and its associated burden, including using vaccines and diagnostics rollout as cost-effectiveness analyses to combat the short and long term spread27,77. Likewise, modelling approaches should consider stochastic effects in the modelling structure as most disadvantaged communities face constant economic and social fluctuations or a higher exposure to natural disasters (e.g., earthquakes, floods).
Rethinking the surveillance system
Our study model incentives an integrated One-Health approach within the local surveillance program to tackle antimicrobial use and resistance in humans and animals, particularly in communities at higher risks of antibiotic exposure68. Multisectoral synergetic efforts between the plant and food safety, environmental sources and wildlife, and human and animal health departments could be promoted by the authorities and national ARB action plan to control the presence of ARB with active surveillance. Identifying and testing individuals in communities of highly endemic risk exposure should be prioritised by developing locality-specific evaluation protocols to target ARB dissemination and associated disease burden rapidly and effectively.
Our study lays out a different structure for ARB quantification and burden attributed by considering the impact of the community exposure to ARB reservoirs. We do not need to emphasize whether ARB is detected in the hospital setting, but rather target the community where ARB is acquired. Our results suggest that including GN-ARB control in the community will help stop further propagation, specifically for those individuals at high risk of ARB exposure scenarios. ARB in the communities is increasing; understanding and quantifying transmission is essential to curb this emerging public health problem.
Methods
Mathematical model and setting description
We developed a dynamic compartmental model for GN-ARB transmission in the rural town of Molina. Molina has 46,000 inhabitants78, a poverty rate of about 13.5%, and one of the highest age-standardized mortality rates for chronic diseases in Chile79. High income inequality, low economic growth, and a substantial proportion of the population with inadequate water, sanitation, and hygiene infrastructure, make Molina a heterogeneous and useful case study to understand GN-ARB transmission beyond western high-income countries (supplementary material, sections A and B).
Figure 5 shows the flow diagram of the compartmental model. We considered Molina’s population in time “t” (N(t)), and demography dynamics to be regulated by a constant birth rate (Λ) and a natural mortality rate given by (\(\varnothing\)). Henceforth, “bacteria” refers to GN bacteria. The model has eight compartments accounting for susceptible population S(t); population colonised by a GN-ARB either at the community ZC(t) or hospital ZH(t), which can be denoted as exposed population with no BSI; and population with bloodstream infections caused by resistant bacteria either at the community IC(t) or hospital setting IH(t). The latter is divided into those with hospital-acquired infections (IHH(t)) and community-acquired (IHC(t)). Finally, individuals might die due to GN-ARB BSIs in the hospital or community settings (RH and RC, respectively).
We assumed that the population is constant N(t) = S(t) + ZC(t) + ZH(t) + IC(t) + IHC(t) + IHH(t) + RC(t) + RH(t) and the presence of GN-ARB determines colonisation. The disease is defined as BSI caused by GN-ARB. We employed the same model for high and low exposure to ARB reservoirs (risk groups/scenarios) in the community. Our model was initiated using baseline data from late 2018 and early 2019. Differential equations are shown in the supplementary material, section C.
Data collection and measurement specifications
Baseline conditions within the compartments and the risk coefficient for ARB reservoirs exposure were extracted from a previous study80. The study provides the prevalence of GN-ARB and the exposure to ARB reservoirs in the Chilean community between 12/2018 until 5/2019. GN-ARB included any quinolone-resistant, extended-spectrum cephalosporin-resistant, or carbapenem-resistant GN bacteria. Gram staining was used to differentiate GN bacteria. The prevalence of GN-ARB in the hospital was also incorporated, based on the same antibiotic-bacterium pairs. Section A of the supplementary material contains further details on the study used to extract our data and parameters’ information.
Model parametrisation
Tables 1 and 2 show the initial conditions and the description of the main variables in the model. We calibrated the transmission parameter (\(\beta\)) by matching the model projected incidence to the overall incidence of GN-ARB colonisation over time provided from a longitudinal study of bacterial susceptibility levels (incidence) to ATB within Chilean hospitals81 using an Approximate Bayesian Computation Markov chain Monte Carlo simulation82. To compute \(\beta\), we tested different regression specifications (e.g., linear, Gaussian, and polynomial) to get the best goodness-of-fit based on R2. Consequently, as \(\beta\) was calibrated to hospital data, we used a population-based ratio obtained from the literature25 to adjust the parameter to the community. Our risk coefficient (\(\rho\)) indicates a high-risk scenario of exposure to ARB reservoirs, compared to a low-risk scenario. It was computed following a two-stage protocol. Firstly, we employed a logistic regression to capture the adjusted ARB rates, obtained from the parallel study on colonisation of GN-ARB in the community. Then, we divided the predicted ARB rates from the previous step into two groups: high and low ARB (calculated as predicted adjusted rates above or below the median values, respectively). Secondly, we calculated the risk ratios using a 2 × 2 matrix between high/low ARB groups and the variable of high/low exposure to ARB reservoirs in the community (ϱ). Community ARB reservoirs considered control variables for the computation of ρ were antibiotic consumption, animal proximity, and contact, household overcrowding, animal products consumption, agricultural occupation for exposure to pesticides, previous hospitalisation, and other sociodemographic variables. All details on the computation of these parameters are found in supplementary material, section B.
We utilised Monte Carlo simulations to estimate 95% confidence intervals (CI) for each compartment to account for uncertainty in decision-making and quantitative risks. We estimated the 95% CI by recalculating \(\beta\) based on a random normal distribution and using 1000 replications.
Additionally, we carried out sensitivity analyses over the main parameters to ensure the results did not hinge on parameter assumptions. GN-ARB BSI probability and hospitalisation rate in the community are the primary sources of uncertainty over the disease dynamics of GN-ARB. We employed univariate and bivariate sensitivity analyses over the main parameters associated with higher GN-ARB burden to account for this variability. One way sensitivity analyses included the variation over the probability of BSI in the community \({(\xi }_{C})\) and the hospital (\({\xi }_{H})\), and the rate of hospitalization for GN-ARB infections \({(\delta }_{I})\) as it is unclear how many people get hospitalised for BSI in a rural community. We used a two-way sensitivity analysis for the GN-ARB BSI probability and hospitalisation rate in the community and GN-ARB BSI probability and treatment rate in the hospital setting due to the higher risk of developing hospital-acquired BSIs (e.g., from the use of intravenous devices, therapeutic interventions). Furthermore, we divided our model into antibiotic-specific resistance types, including GN resistant to carbapenems, cephalosporins, and quinolones. We adjusted our \(\beta\) and ϱ parameters to the detailed antibiotics (supplementary material, section E). Also, we surveyed the literature to understand the magnitude of the transmission parameters following different GN bacteria type (supplementary material, Section E).
The complete analysis was computed on MATLAB ® version R2019b (MathWorks Inc., Natick, MA, USA www.mathworks.com) and Python programming language version 3.9 (Python Software Foundation, https://www.python.org/). The complete script is available on Python Jupyter notebook at https://bit.ly/2ZpucKh.
Data availability
Data is publicly available or within the manuscript. The complete script is available on Python Jupyter notebook at https://bit.ly/2ZpucKh. No ethics approval required.
Abbreviations
- ARB:
-
Antibiotic-resistant bacteria
- ATB:
-
Antibiotic
- BSI:
-
Bloodstream Infections
- GN:
-
Gram-negative
- HICs:
-
High-income countries
- ICU:
-
Intensive care units
- LMICs:
-
Low- and middle-income countries
- MDR:
-
Multidrug resistance
- OECD:
-
Organisation of Economic Cooperation and Development
- WHO:
-
World Health Organisation
References
Jee, Y. et al. Antimicrobial resistance: A threat to global health. Lancet Infect. Dis. 18, 939–940 (2018).
Peleg, A. Y. & Hooper, D. C. Hospital-acquired infections due to gram-negative bacteria. N. Engl. J. Med. 362, 1804–1813 (2010).
Hofer, U. The cost of antimicrobial resistance. Nat. Rev. Microbiol. 17, 3–3 (2019).
Hawkey, P. The growing burden of antimicrobial resistance. J. Antimicrob. Chemother. 62, i1–i9 (2008).
Maragakis, L. L., Perencevich, E. N. & Cosgrove, S. E. Clinical and economic burden of antimicrobial resistance. Expert Rev. Anti-infect. Ther. 6, 751–763 (2008).
Holmes, A. H. et al. Understanding the mechanisms and drivers of antimicrobial resistance. Lancet 387, 176–187 (2016).
Okeke, I. N., Lamikanra, A. & Edelman, R. Socioeconomic and behavioral factors leading to acquired bacterial resistance to antibiotics in developing countries. Emerg. Infect. Dis. 5, 18 (1999).
Istúriz, R. E. & Carbon, C. Antibiotic use in developing countries. Infect. Control Hosp. Epidemiol. 21, 394–397 (2000).
Malik, B. & Bhattacharyya, S. Antibiotic drug-resistance as a complex system driven by socio-economic growth and antibiotic misuse. Sci. Rep. 9, 1–12 (2019).
Silbergeld, E. K., Graham, J. & Price, L. B. Industrial food animal production, antimicrobial resistance, and human health. Annu. Rev. Public Health 29, 151–169 (2008).
Van Boeckel, T. P. et al. Global trends in antimicrobial resistance in animals in low-and middle-income countries. Science 365, 6459 (2019).
Klein, E. Y. et al. Assessment of WHO antibiotic consumption and access targets in 76 countries, 2000–15: An analysis of pharmaceutical sales data. Lancet Infect. Dis. 21, 107–115 (2021).
Rousham, E. K., Unicomb, L. & Islam, M. A. Human, animal and environmental contributors to antibiotic resistance in low-resource settings: Integrating behavioural, epidemiological and One Health approaches. Proc. R. Soc. B Biol. Sci. 285, 20180332 (2018).
Organisation for Economic Co-operation Development. Stemming the Superbug Tide: Just a Few Dollars More (OECD Publishing, 2018).
Kumarasamy, K. K. et al. Emergence of a new antibiotic resistance mechanism in India, Pakistan, and the UK: A molecular, biological, and epidemiological study. Lancet Infect. Dis. 10, 597–602 (2010).
Boucher, H. W. et al. Bad bugs, no drugs: No ESKAPE! An update from the Infectious Diseases Society of America. Clin. Infect. Dis. 48, 1–12 (2009).
Cornaglia, G. & Rossolini, G. M. Forthcoming therapeutic perspectives for infections due to multidrug-resistant Gram-positive pathogens. Clinical Microbiology and infection 15(3), 218–223 (2009).
Ruppé, É., Woerther, P.-L. & Barbier, F. Mechanisms of antimicrobial resistance in Gram-negative bacilli. Ann. Intensive Care 5, 21 (2015).
Cassini, A. et al. Attributable deaths and disability-adjusted life-years caused by infections with antibiotic-resistant bacteria in the EU and the European Economic Area in 2015: A population-level modelling analysis. Lancet Infect. Dis. 19, 56–66 (2019).
Tacconelli, E. et al. ESCMID guidelines for the management of the infection control measures to reduce transmission of multidrug-resistant Gram-negative bacteria in hospitalized patients. Clin. Microbiol. Infect. 20, 1–55 (2014).
Laupland, K. B. & Church, D. L. Population-based epidemiology and microbiology of community-onset bloodstream infections. Clin. Microbiol. Rev. 27, 647–664 (2014).
Furuya, E. Y. & Lowy, F. D. Antimicrobial-resistant bacteria in the community setting. Nat. Rev. Microbiol. 4, 36–45 (2006).
D’Costa, V. M. et al. Antibiotic resistance is ancient. Nature 477, 457 (2011).
Austin, D. & Anderson, R. Studies of antibiotic resistance within the patient, hospitals and the community using simple mathematical models. Philos. Trans. R. Soc. Lond. Ser. B Biol. Sci. 354, 721–738 (1999).
Knight, G. M. et al. Quantifying where human acquisition of antibiotic resistance occurs: A mathematical modelling study. BMC Med. 16, 137 (2018).
Levin, B. et al. The population genetics of antibiotic resistance. Clin. Infect. Dis. 24, S9–S16 (1997).
Niewiadomska, A. M. et al. Population-level mathematical modeling of antimicrobial resistance: A systematic review. BMC Med. 17, 1–20 (2019).
Wang, Y. et al. Comprehensive resistome analysis reveals the prevalence of NDM and MCR-1 in Chinese poultry production. Nat. Microbiol. 2, 1–7 (2017).
Liu, Y.-Y. et al. Emergence of plasmid-mediated colistin resistance mechanism MCR-1 in animals and human beings in China: A microbiological and molecular biological study. Lancet Infect. Dis. 16, 161–168 (2016).
Manaia, C. M. Assessing the risk of antibiotic resistance transmission from the environment to humans: Non-direct proportionality between abundance and risk. Trends Microbiol. 25, 173–181 (2017).
Agga, G. E., Arthur, T. M., Durso, L. M., Harhay, D. M. & Schmidt, J. W. Antimicrobial-resistant bacterial populations and antimicrobial resistance genes obtained from environments impacted by livestock and municipal waste. PLoS ONE 10, e0132586 (2015).
Seiler, C. & Berendonk, T. U. Heavy metal driven co-selection of antibiotic resistance in soil and water bodies impacted by agriculture and aquaculture. Front. Microbiol. 3, 399 (2012).
Wellington, E. M. et al. The role of the natural environment in the emergence of antibiotic resistance in Gram-negative bacteria. Lancet Infect. Dis. 13, 155–165 (2013).
Mughini-Gras, L. et al. Attributable sources of community-acquired carriage of Escherichia coli containing β-lactam antibiotic resistance genes: A population-based modelling study. Lancet Planet. Health 3, e357–e369 (2019).
McEwen, S. A. & Collignon, P. J. Antimicrobial resistance: A one health perspective. Microbiol. Spectrum 6(2), 10 (2018).
Omulo, S. et al. Carriage of antimicrobial-resistant bacteria in a high-density informal settlement in Kenya is associated with environmental risk-factors. Antimicrob. Resist. Infect. Control 10, 1–12 (2021).
Ramay, B. M. et al. Antibiotic use and hygiene interact to influence the distribution of antimicrobial-resistant bacteria in low-income communities in Guatemala. Sci. Rep. 10, 1–10 (2020).
Reid, L. Ethics and Drug Resistance: Collective Responsibility for Global Public Health 257–278 (Springer, 2020).
Levin, B. R., Baquero, F. & Johnsen, P. J. A model-guided analysis and perspective on the evolution and epidemiology of antibiotic resistance and its future. Curr. Opin. Microbiol. 19, 83–89 (2014).
Austin, D., Kakehashi, M. & Anderson, R. The transmission dynamics of antibiotic–resistant bacteria: The relationship between resistance in commensal organisms and antibiotic consumption. Proc. R. Soc. Lond. Ser. B Biol. Sci. 264, 1629–1638 (1997).
Blanquart, F., Lehtinen, S., Lipsitch, M. & Fraser, C. The evolution of antibiotic resistance in a structured host population. J. R. Soc. Interface 15, 20180040 (2018).
Elettreby, M. F. & Ahmed, E. Multi-drug antimicrobial resistance model. Math. Methods Appl. Sci. 43, 10462–10473 (2020).
Lipsitch, M. The rise and fall of antimicrobial resistance. Trends Microbiol. 9, 438–444 (2001).
Lipsitch, M., Bergstrom, C. T. & Levin, B. R. The epidemiology of antibiotic resistance in hospitals: Paradoxes and prescriptions. Proc. Natl. Acad. Sci. 97, 1938–1943 (2000).
Kouyos, R. D., zur Wiesch, P. A. & Bonhoeffer, S. On being the right size: The impact of population size and stochastic effects on the evolution of drug resistance in hospitals and the community. PLoS Pathog. 7, e1001334 (2011).
Bralic, R. A. et al. Colonization with antibiotic-resistant gram-negative bacteria in population-based hospital and community settings in Chile. Infect. Control Hosp. Epidemiol. 41, s175–s176 (2020).
Roug, A., Byrne, B. A., Conrad, P. A. & Miller, W. Zoonotic fecal pathogens and antimicrobial resistance in county fair animals. Comp. Immunol. Microbiol. Infect. Dis. 36, 303–308 (2013).
Wegener, H. C. Improving Food Safety Through a One Health Approach: Workshop Summary. 331 (National Academies Press) (2012).
Pokharel, S., Raut, S. & Adhikari, B. (BMJ Specialist Journals, 2019).
Chopra, S. & Rizvi, M. Antimicrobial resistance in low-and middle-income countries. Clin. Infect. Dis. 69(7), 1264–1265 (2019).
Chilean Ministry of Health. Deparment of Statistics and Health Information, https://deis.minsal.cl/ (2021).
Kardaś-Słoma, L. et al. Impact of antibiotic exposure patterns on selection of community-associated methicillin-resistant Staphylococcus aureus in hospital settings. Antimicrob. Agents Chemother. 55, 4888–4895 (2011).
Cooper, B. et al. Methicillin-resistant Staphylococcus aureus in hospitals and the community: Stealth dynamics and control catastrophes. Proc. Natl. Acad. Sci. 101, 10223–10228 (2004).
Nguyen, N. V. et al. Community-level consumption of antibiotics according to the AWaRe (Access, Watch, Reserve) classification in rural Vietnam. JAC-Antimicrob. resist. 2, dlaa048 (2020).
Schmiege, D., Evers, M., Kistemann, T. & Falkenberg, T. What drives antibiotic use in the community? A systematic review of determinants in the human outpatient sector. Int. J. Hygiene Environ. Health 226, 113497 (2020).
Alividza, V. et al. Investigating the impact of poverty on colonization and infection with drug-resistant organisms in humans: A systematic review. Infect. Dis. Poverty 7, 1–11 (2018).
Iramiot, J. S., Kajumbula, H., Bazira, J., Kansiime, C. & Asiimwe, B. B. Antimicrobial resistance at the human–animal interface in the Pastoralist Communities of Kasese District, South Western Uganda. Sci. Rep. 10, 1–15 (2020).
Van Boeckel, T. P. et al. Reducing antimicrobial use in food animals. Science 357, 1350–1352 (2017).
Russell, A. Bacterial outer membrane and cell wall penetration and cell destruction by polluting chemical agents and physical conditions. Sci. Progr. 86, 283–312 (2003).
Cortes, S. et al. First measurement of human exposure to current use pesticides (CUPs) in the atmosphere of central Chile: The case study of Mauco cohort. Atmos. Pollut. Res. 11, 776–784 (2020).
Byarugaba, D. Antimicrobial resistance in developing countries and responsible risk factors. Int. J. Antimicrob. Agents 24, 105–110 (2004).
Millanao, A. et al. Antimicrobial resistance in Chile and The One Health paradigm: Dealing with threats to human and veterinary health resulting from antimicrobial use in salmon aquaculture and the clinic. Revista Chilena de Infectologia 35, 299 (2018).
Miranda, C. D., Godoy, F. A. & Lee, M. R. Current status of the use of antibiotics and the antimicrobial resistance in the Chilean salmon farms. Front. Microbiol. 9, 1284 (2018).
Finch, R. G., Metlay, J. P., Davey, P. G. & Baker, L. J. Educational interventions to improve antibiotic use in the community: Report from the International Forum on Antibiotic Resistance (IFAR) colloquium, 2002. Lancet Infect. Dis. 4, 44–53 (2004).
Roca, I. et al. The global threat of antimicrobial resistance: Science for intervention. New Microbes New Infect. 6, 22–29 (2015).
Llor, C. & Bjerrum, L. Antimicrobial resistance: Risk associated with antibiotic overuse and initiatives to reduce the problem. Therap. Adv. Drug Saf. 5, 229–241 (2014).
Ayukekbong, J. A., Ntemgwa, M. & Atabe, A. N. The threat of antimicrobial resistance in developing countries: Causes and control strategies. Antimicrob. Resist. Infect. Control 6, 47 (2017).
Aenishaenslin, C. et al. Evaluating the integration of one health in surveillance systems for antimicrobial use and resistance: A conceptual framework. Front. Vet. Sci. 8, 169 (2021).
Talaminos, A. et al. Modelling the epidemiology of Escherichia coli ST131 and the impact of interventions on the community and healthcare centres. Epidemiol. Infect. 144, 1974–1982 (2016).
Lapointe-Shaw, L. et al. Cost-effectiveness analysis of universal screening for carbapenemase-producing Enterobacteriaceae in hospital inpatients. Eur. J. Clin. Microbiol. Infect. Dis. 36, 1047–1055 (2017).
Inglis, T. J. & Urosevic, N. Where sepsis and antimicrobial resistance countermeasures converge. Front. Public Health 5, 6 (2017).
Perez, K. K. et al. Integrating rapid diagnostics and antimicrobial stewardship improves outcomes in patients with antibiotic-resistant Gram-negative bacteremia. J. Infect. 69, 216–225 (2014).
Opota, O., Croxatto, A., Prod’hom, G. & Greub, G. Blood culture-based diagnosis of bacteraemia: State of the art. Clin. Microbiol. Infect. 21, 313–322 (2015).
Stevenson, L. G., Drake, S. K. & Murray, P. R. Rapid identification of bacteria in positive blood culture broths by matrix-assisted laser desorption ionization-time of flight mass spectrometry. J. Clin. Microbiol. 48, 444–447 (2010).
Huttunen, R. & Aittoniemi, J. New concepts in the pathogenesis, diagnosis and treatment of bacteremia and sepsis. J. Infect. 63, 407–419 (2011).
Toth, D. J., Samore, M. H. & Nelson, R. E. Economic evaluations of new antibiotics: The high potential value of reducing healthcare transmission through decolonization. Clin. Infect. Dis. 72, S34–S41 (2021).
O'Neill, J. Review on antimicrobial resistance: tackling drug-resistant infections globally: Final report and recommendations (2016).
Instituto Nacional de Estadísticas. (INE, 2018).
Ferreccio, C. et al. Study protocol for the Maule Cohort (MAUCO) of chronic diseases, Chile 2014–2024. BMC Public Health 16, 122 (2015).
Araos, R. et al. Society for Healthcare Epidemiology of America (SHEA) Vol. 6 (Atlanta, EEUU, 2020).
Allel, K. et al. Ch. 4, 115–151 (Centro de Políticas Públicas UC. Propuestas para Chile. Concurso de Políticas Públicas 2019. Pontificia Universidad Católica, 2020).
Sadegh, M. & Vrugt, J. A. Approximate bayesian computation using markov chain monte carlo simulation: Dream (abc). Water Resour. Res. 50, 6767–6787 (2014).
Jernigan, J. A. et al. Multidrug-resistant bacterial infections in US hospitalized patients, 2012–2017. N. Engl. J. Med. 382, 1309–1319 (2020).
Gambra, M. et al. Incidence and mortality of bacteremia in a public hospital in Santiago. Rev. Med. Chil. 140, 859–866 (2012).
Melzer, M. & Petersen, I. Mortality following bacteraemic infection caused by extended spectrum beta-lactamase (ESBL) producing E. coli compared to non-ESBL producing E. coli. J. Infect. 55, 254–259 (2007).
Haverkate, M. R., Derde, L. P., Brun-Buisson, C., Bonten, M. J. & Bootsma, M. C. Duration of colonization with antimicrobial-resistant bacteria after ICU discharge. Intensive Care Med. 40, 564–571 (2014).
Allel, K. et al. Socioeconomic factors associated with antimicrobial resistance of Pseudomonas aeruginosa, Staphylococcus aureus, and Escherichia coli in Chilean hospitals (2008–2017). Rev Panam Salud Publica 44, 1 (2020).
Søgaard, M., Thomsen, R. W., Bang, R. B., Schønheyder, H. C. & Nørgaard, M. Trends in length of stay, mortality and readmission among patients with community-acquired bacteraemia. Clin. Microbiol. Infect. 21, e781-789 (2015).
O’Brien, J. M. Jr., Ali, N. A., Aberegg, S. K. & Abraham, E. Sepsis. Am. J. Med. 120, 1012–1022 (2007).
Wirtz, V. J., Dreser, A. & Gonzales, R. Trends in antibiotic utilization in eight Latin American countries, 1997–2007. Rev. Panam. Salud Publica 27, 219–225 (2010).
Acknowledgements
All authors attest they meet the ICMJE criteria for authorship and have reviewed and approved the final article. This article was supported by a full scholarship provided by the Chilean Government “Beca de Magíster en el Extranjero Becas Chile, Convocatoria 2019, N° 73200098, Asociación Nacional de Investigación y Desarrollo (ANID)”. This research was also supported by the Agencia Nacional de Investigación y Desarrollo (ANID) Millennium Science Initiative Program MICROB-R [grant NCN17_081 to RA, JM, EU], Fondo de Financiamiento de Centros de Investigación en Áreas Prioritarias FONDAP (grant 15110017 to EU), and Fondo Nacional de Desarrollo Científico y Tecnológico FONDECYT (grant 1211933 to KA, JM, and EU). We thank the Advanced Center for Chronic Diseases Chile and the people from the MAUCO project for facilitating, collecting and gather the parallel study from which the main parameters were computed.
Funding
The funders of the study had no role in study design, data collection, analysis, interpretation, writing of the report, or in the decision to submit the paper for publication.
Author information
Authors and Affiliations
Contributions
Study design and analytical methods: K.A., J.P.G. Data provision and accessibility for the MDRGN and MAUCO studies: C.F., R.A. Data analyses: K.A., J.P.G., L.G., D.T., E.U. Manuscript writing: K.A. Data interpretation, critical manuscript review, edition, final approval: C.F., K.A., J.P.G., E.U., L.G., J.M., D.T., R.A.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Allel, K., Goscé, L., Araos, R. et al. Transmission of gram-negative antibiotic-resistant bacteria following differing exposure to antibiotic-resistance reservoirs in a rural community: a modelling study for bloodstream infections. Sci Rep 12, 13488 (2022). https://doi.org/10.1038/s41598-022-17598-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-17598-x
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.