Abstract
Lignans are widely distributed plant secondary metabolites that have received attention for their benefits to human health. Sesamin is a furofran lignan that is conventionally extracted from Sesamum seeds and shows anti-oxidant and anti-inflammatory activities in the human liver. Sesamin is biosynthesized by the Sesamum-specific enzyme CYP81Q1, and the natural sources of sesamin are annual plants that are at risk from climate change. In contrast, Forsythia species are widely distributed perennial woody plants that highly accumulate the precursor lignan pinoresinol. To sustainably supply sesamin, we developed a transformation method for Forsythia leaf explants and generated transgenic Forsythia plants that heterologously expressed the CYP81Q1 gene. High-performance liquid chromatography (HPLC) and LC-mass spectrometry analyses detected sesamin and its intermediate piperitol in the leaves of two independent transgenic lines of F. intermedia and F. koreana. We also detected the accumulation of sesamin and piperitol in their vegetatively propagated descendants, demonstrating the stable and efficient production of these lignans. These results indicate that CYP81Q1-transgenic Forsythia plants are promising prototypes to produce diverse lignans and provide an important strategy for the cost-effective and scalable production of lignans.
Similar content being viewed by others
Introduction
The aging of world populations highlights the importance of plant secondary metabolites such as alkaloids, flavonoids, terpenoids, and lignans with benefits for human health1,2. Lignans are phenylpropanoid dimers with diverse functions, and dietary lignans have attracted attention as food nutrients3,4. (+)-Sesamin is a furofuran lignan that is commercially available as a health-promoting supplement5. In mammals, (+)-Sesamin metabolites attenuate oxidation and inflammation for the protection of the liver6,7. (+)-Sesamin also shows anti-cancer properties8. (+)-Sesamin is commercially available via extraction at concentrations (4–6 mg/g) from Sesamum indicum (sesame) seed oil5,9,10. Sesame plants, the strongest known synthesizers of (+)-sesamin, are annuals that are threatened by climate change11,12. Thus, new plant sources are required for the efficient and stable production of (+)-sesamin.
In land plants, the lignan metabolic pathway branches from that of lignin at the coupling of monolignols such as coniferyl alcohol, which is synthesized from phenylalanine (Fig. 1)13,14. Dirigent protein metabolizes coniferyl alcohol and specifically synthesizes the precursor lignan pinoresinol (Fig. 1)15,16,17,18. The lignan metabolic pathway further diverges into structurally and functionally diverse lignans by plant species-specific enzymes (Fig. 1)5,14,19. In sesame plants, cytochrome P450 81Q1 (CYP81Q1) catalyzes the sequential conversion of (+)-pinoresinol to (+)-piperitol and (+)-sesamin (Fig. 1)9.
Biosynthesis of (+)-piperitol and (+)-sesamin in Forsythia plants by heterologous expression of the sesame CYP81Q1 gene. Lignan biosynthesis is initiated from phenylalanine. Two coniferyl alcohols are coupled to the precursor lignan (+)-pinoresinol in land plants (green). In sesame plants, CYP81Q1 sequentially synthesizes (+)-piperitol and (+)-sesamin (red).
Forsythia species such as Forsythia intermedia (Fi) and F. koreana (Fk) are widely distributed perennial woody plants. Extracts of Forsythia plants have been empirically used in traditional medicines20. Forsythia species produce various lignans and other polyphenols21,22,23, and their biosynthesis and regulation have been analyzed18,20,21,24,25. Forsythia leaves accumulate the precursor lignan (+)-pinoresinol at high levels but lack (+)-sesamin biosynthesis (Fig. 1)21. To sustainably supply beneficial lignans including (+)-sesamin, heterologous expression of the lignan-biosynthetic enzyme genes in Forsythia plants is promising.
Previously, we demonstrated the ectopic accumulation of (+)-sesamin in cultured CYP81Q1-transgenic Fk cells26,27 and showed their ability to produce (+)-sesamin. However, the mass production of (+)-sesamin using transgenic cells is not practical in light of the cost of large-scale cell culture. In contrast, given the large biomass generated by Forsythia leaves, CYP81Q-transgenic Forsythia plants could efficiently and stably produce (+)-sesamin. In this study, we generated CYP81Q1-transgenic Forsythia plants that stably produce the intermediate (+)-piperitol and the product (+)-sesamin.
Results
Initially, we established a practical method for the transformation of Forsythia plants (Fig. 2A). Fi leaf explants (n = 451) were soaked in a suspension of Agrobacterium tumefaciens cells harboring the Pro35S:nGFP28 plasmid and regenerated shoots and roots during culture for two years (see “Materials and methods”, Fig. 2B–G, Table 1). Eight of ten independent kanamycin-resistant lines exhibited signals for nGFP presence and expression when examined by fluorescence microscopy (Fig. 2H) and genomic (Fig. 2I) and RT-PCR analyses (Fig. 2J); no nGFP presence or expression was seen in wild-type (WT) plants. Two kanamycin-resistant plants did not show GFP presence or expression, probably due to somaclonal variation that eliminated the transgene (Fig. 2I,J). Even after vegetative propagation for four years, newly generated FiPro35S:nGFP leaves maintained GFP fluorescence, demonstrating stable Pro35S:nGFP transformation of Forsythia plants.
Generation of transgenic Pro35S:nGFP plants. (A) A scheme of Agrobacterium-mediated transformation of Forsythia leaf explants followed by the regeneration of the whole plant body. CIM callus-inducing medium, SIM shoot-inducing medium. (B) The Pro35S:nGFP plasmid. nptII kanamycin resistance gene, 35S cauliflower mosaic virus 35S promoter, NLS nuclear localization signal, GFP green florescence protein, nos nos terminator. (C) Leaf explants co-cultured with Agrobacterium cells. (D) Leaf explants dedifferentiated into calluses and regenerating adventitious shoots. (E) An adventitious shoot before separation from calluses. (F) GFP fluorescence in a FiPro35S:nGFP shoot. (G) A FiPro35S:nGFP plant transferred into soil. (H) GFP fluorescence in a FiPro35S:nGFP leaf (nGFP; right) in contrast with the lack of fluorescence in a non-transformed wild-type (WT) leaf (left). Dotted white line marks the WT leaf margin. (I, J) Genomic (I) and reverse transcription-polymerase chain reaction (RT-PCR) (J) analyses of GFP gene. Cyt (cytochrome) served as an internal control. The numbers above the gel images indicate the plant lines regenerated from independent calluses. Scale bars = 1 cm in (C) to (H).
We also introduced the 35S promoter-regulated sesame CYP81Q1 gene (Pro35S:CYP81Q1)29 into Fi and Fk. After co-culture of 956 Fi and 273 Fk leaf explants with Agrobacterium cells harboring the Pro35S:CYP81Q1 plasmid, the resulting transgenic Forsythia plants were propagated vegetatively in soil through repeated rounds of cutting and growth in our plant culture room—conditions under which Forsythia plants continuously developed their leaves without flowering (Table 1). Eventually, two independent transgenic lines developed normally on soil (Fig. 3A) and showed CYP81Q1 gene presence (Fig. 3B) and expression (Fig. 3C).
Generation of transgenic Pro35S:CYP81Q1 plants. (A) Control and Pro35S:CYP81Q1 plants. Scale bars = 10 cm. (B, C) Genomic (B) and reverse transcribed (RT)-PCR (C) analyses of CYP81Q1 gene. Cyt (cytochrome) served as an internal control. The numbers above the gel images indicate the plant lines regenerated from independent calluses.
To detect the accumulation of the products of CYP81Q1, we subjected the leaves of FiPro35S:CYP81Q1, FkPro35S:CYP81Q1, control FiPro35S:nGFP, and FkWT plants to high-performance liquid chromatography (HPLC) and LC-mass spectrometry (MS) analyses (Fig. 4). HPLC and LC–MS analyses indicated that the heterologous expression of the CYP81Q1 gene in Forsythia leaves resulted in the production of (+)-piperitol and (+)-sesamin (Fig. 4), and that the leaves of the primary transformants FiPro35S:CYP81Q1 and FkPro35S:CYP81Q1 accumulated (+)-piperitol (14.23 and 39.45 µg/g DW, respectively) and (+)-sesamin (5.57 and 27.21 µg/g DW), while control FiPro35S:nGFP and FkWT plants did not (Table 2).
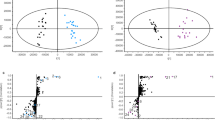
HPLC (A) and LC–MS (B) analysis of control and Pro35S:CYP81Q1 plants. Chromatograms showing specific accumulation of (+)-piperitol (P1) and (+)-sesamin (P2) in the Pro35S:CYP81Q1 leaves. In (B), pinoresinol (P3); m/z = 381.12, piperitol; m/z = 379.11, sesamin; m/z = 377.09, 3′-ethoxysesamin; m/z = 421.11, TIC total ion current.
We further examined whether the ability to biosynthesize piperitol and sesamin was inherited by descendant transgenic plants. After two and three rounds of vegetative propagation, descendant Pro35S:CYP81Q1 plants produced similar amounts of (+)-piperitol and (+)-sesamin (Table 2). The total amounts of (+)-piperitol and (+)-sesamin (Table 2) exceeded those in CYP81Q1-Fk cells (10 µg/g DW)5. In contrast, the LC–MS analysis detected pinoresinol (P3 in Fig. 4B) and the content of pinoresinol was not changed in the control and the transgenic Pro35S:CYP81Q1 plants [(FiPro35S:nGFP and FiPro35S:CYP81Q1 (9.34 ± 3.64 and 11.68 ± 5.44 mg/g DW, respectively) and FkWT and FkPro35S:CYP81Q1 (24.24 ± 10.06 and 25.35 ± 7.63 mg/g DW, respectively)]; Supplemental Fig. S1), indicating that the biosynthesis of pinoresinol is intact in the presence of CYP81Q1. Altogether, we demonstrated that the Pro35S:CYP81Q1 plants efficiently produced (+)-piperitol and (+)-sesamin.
Discussion
In this study, we provide evidence that transgenic Forsythia plants produce ectopic lignans, (+)-piperitol and (+)-sesamin. These results suggest that sesamolin and sesaminol, which are antioxidant sesame lignans metabolized from (+)-sesamin by CYP92B1430, is expected to be produced via additional introduction of the CYP92B14 gene into Pro35S:CYP81Q1 plants. Another lignan, podophyllotoxin, may also be produced in transgenic Forsythia plants. Podophyllotoxin is at present extracted from the rhizomes of Podophyllum species and clinically utilized in cancer therapy31. In Podophyllum podophyllotoxin biosynthesis, matairesinol, which Forsythia species also accumulate in the pathway downstream of pinoresinol, is metabolized to pluviatolide by CYP719A2332. Additional enzymes (CYP71CU1, 2-oxoglutarate/Fe(II)-dependent dioxygenase, and O-methyltransferases) convert pluviatolide to the proposed precursor deoxypodophyllotoxin33. Thus, methods for introducing multiple genes into Forsythia plants will pave the way for the generation of podophyllotoxin and its related compounds by transgenic plants. In a previous study, we generated triple-transgenic Forsythia cultured cells26, suggesting the possibility of multigene transformation of Forsythia plants.
The production of various specialized plant metabolites has been attempted in transgenic and synthetic biology-based microorganisms1,34,35,36,37,38,39,40. However, most microorganisms appear to lack (+)-sesamin and its precursor (+)-pinoresinol9,41. Moreover, tremendous bioinformatic and screening processes are often required for the generation of genetically-engineered microorganisms that produce plant lignans. Additionally, mass production using such engineered microorganisms could be limited by genetic instability, infectious contamination, unexpected product toxicity, and low fermentation performance38,40. In contrast, Forsythia species are perennial shrubs, and thus Pro35S:CYP81Q1 plants are easily cultivated via explant, a marked advantage for stable growth and mass propagation in a plant factory. Moreover, a plant factory provides plasticity in place and time of the production of sesamin in plants42, unlike agricultural production that is limited by climates, seasons, and farmlands. Thus, we will be able to produce sesamin using our transgenic Forsythia plants in a plant factory.
Also of significance is the reproducible generation of transgenic Forsythia plants expressing the CYP81Q1 gene. The lignan biosynthetic pathway has previously been modified in cultured plants, cells, and hairy roots of Carthamus, Linum, Hyptis, Juniperus, Podophyllum, Sesamum, and Forsythia species43,44,45,46,47,48,49,50,51,52,53,54,55,56,57,58,59, but the transgenic plants have not been reported. For the scalable production of lignans, our transgenic Forsythia plant method provides an important strategy for future metabolic engineering of such lignan-producing plants.
The transgenic Forsythia plants have limitation in the content of sesamin as compared with sesame seeds. To increase the content of sesamin in the transgenic Forsythia plants, we will be able to apply multigene transformation strategy. Previously, triple-transgenic Forsythia cells with the RNAi construct for endogenous pinoresinol-lariciresinol reductase, and the overexpression-construct of pinoresinol glucosylating enzyme, which increase the level of the precursor pinoresinol, as well as CYP81Q1 gene, produced higher level of sesamin than the single-transgenic CYP81Q1 cells26. The same strategy of multigene transformation of Forsythia plants is expected to increase the content of sesamin. Moreover, overexpression of enzymes upstream of pinoresinol stimulated accumulation of podophyllotoxin-related lignans60. Thus, overexpression of the upstream enzymes may increase the content of sesamin in transgenic Forsythia plants. Because the content of pinoresinol was not changed in the control and the Pro35S:CYP81Q1 plants (Supplemental Fig. S1), the moderate activity of CYP81Q1 seems to limit the rate of production of sesamin from pinoresinol. As many cytochrome P450 enzymes function in complex with their native oxidoreductases30, co-expression of CYP81Q1 and S. indicum oxidoreductase genes in transgenic Forsythia plants is a future option to strengthen the activity of CYP81Q1 for higher production of sesamin. In addition, the red light condition increased the content of sesamin in the transgenic Forsythia cells26, suggesting that irradiation of red light to transgenic Forsythia plant may increase the content of sesamin.
In conclusion, we have generated Forsythia plants as promising prototypes for the efficient and sustainable heterologous production of beneficial lignans.
Materials and methods
Plant growth conditions
F. intermedia and F. koreana plants were obtained from Niigata Prefectural Botanical Garden and Dr. Toshiaki Umezawa (Kyoto University), respectively. The use of these plants for this study has been approved by the providers. All the experimental work on plant material described in this study complies with the relevant institutional, national, and international guidelines and legislation. Both of Forsythia plants were maintained in pots filled with soil inside a plant culture room under a cycle of 16-h-light (photosynthetic photon flux density of 50 to 75 µmol m−2 s−1)/8-h-dark at 22 °C unless otherwise indicated. After surface-sterilization of 10 cm length Forsythia shoots using 70% ethanol for 30 s and 1% sodium hypochlorite solution for 15 min, the Forsythia plants were maintained in vitro in a culturing box on modified MS medium (MS salts, MS vitamin, 30 g/L sucrose, 22.5 mg/L CuSO45H2O, 5 g/L Agargel [Sigma-Aldrich, MO]) and transferred to fresh medium every two to three months.
Transformation and preparation of Agrobacterium cells
For construction of Pro35S:nGFP28, the NLS was inserted upstream of GFP and the resulting fusion NLS-GFP genes was inserted into pBI101. For construction of Pro35S:CYP81Q129, the coding sequence of CYP81Q1 (accession number AB194714) was inserted downstream of the Pro35S promoter in pBINplus. Agrobacterium tumefaciens GV3101 cells were individually transformed with Pro35S:nGFP28 and Pro35S:CYP81Q129 plasmids using a Gene Pulser II electroporator (BioRad, CA). The resultant Agrobacterium cells were cultured in LB liquid medium at 27 °C until the optical density at 600 nm (OD600) reached 1.5 to 2.0. The Agrobacterium cells were collected by centrifugation of the medium (HP25; Beckman, CA) at 6000×g for 15 min at room temperature, resuspended in the transformation solution (Gamborg’s B5 salts, MS vitamins, 30 g/L sucrose, 2.0 mg/L indole-3-acetic acid [IAA; Nacalai tesque, Japan], 0.5 mg/L 6-benzyladenopurine [BA; Nacalai tesque, Japan], 20 mg/L acetosyringone [Tokyo Chemical Industry, Japan], 0.02% 500 W Additive [equivalent to Silwet L-77; DOW CORNING, MI]), and further diluted with additional transformation solution to a concentration of OD600 = 0.7.
Transformation and culture of Forsythia leaf explants
To prepare leaf explants, the third to sixth leaves from the top of one-month-cultured Forsythia plants were detached at their petioles and cut into 10 × 5 mm squares using a surgical knife. For the co-culture, the leaf explants were submerged in the Agrobacterium cell-suspended transformation solution for 2 min at room temperature, the transformation solution was wiped off using sterilized paper towels, and the explants were transferred onto sterilized filter papers (No.1 70 mm; ADVANTEC, Japan) over callus-inducing medium (CIM; Gamborg’s B5 salts, B5 vitamin, 30 g/L sucrose, 2.0 mg/L IAA, 0.5 mg/L BA, 20 mg/L acetosyringone, 3 g/L gellan gum [Kanto Chemical, Japan]), and maintained at 22 °C in the dark for three days. To suppress the overgrowth of the Agrobacterium cells, the leaf explants were transferred onto shoot-inducing medium (SIM; MS salts, MS vitamin, 30 g/L sucrose, 0.5 mg/L IAA, 2.0 mg/L BA, 3 g/L gellan gum) supplemented with 10 mg/L meropenem (Tokyo Chemical Industry, Japan) for four days in the dark. To isolate kanamycin-resistant plants, leaf explants were cultured on SIM supplemented with kanamycin (75 mg/L for F. intermedia and 50 mg/L for F. koreana; Nacalai tesque, Japan) and 10 mg/L meropenem under 16-h-light/8-h-dark conditions and transferred onto fresh medium every two weeks up to six months. Adventitious shoots regenerated at the periphery of the dedifferentiated explants were transferred onto shoot-elongating medium (MS salts, MS vitamin, 30 g/L sucrose, 22.5 mg/L CuSO45H2O, 2.0 mg/L BA, 5 g/L Agargel) supplemented with kanamycin and meropenem in sterilized glass tubes until the shoots elongated to 5 cm height. Shoots were transferred and maintained on hormone-free medium (MS salts, MS vitamin, 30 g/L sucrose, 22.5 mg/L CuSO4·5H2O, 5 g/L Agargel) until their rooting.
Microscopy
GFP fluorescence was observed using a M205 fluorescence stereomicroscope (Leica Microsystems, Germany) using the GFP3 filter and recorded by LAS AF software (Leica Microsystems, Germany).
Extraction of nucleotides and detection of genes
The genomic DNAs of Forsythia leaves were prepared using a Nucleon PhytoPure Genomic DNA Extraction kit (GE Healthcare, Sweden) according to the manufacturer’s instruction. The total RNAs of Forsythia leaves were prepared using an RNAeasy Plant Mini Kit (Qiagen, Germany) according to the manufacturer’s instruction, cleared by precipitation in 4 M lithium chloride solution (final concentration), treated with DNase I (Qiagen, Germany), and further subjected to reverse transcription using SuperScript III (Thermo Fisher Scientific, MA) with an Oligo(dT) primer (Thermo Fisher Scientific, MA). For the detection of genes of interest, the genomic and complementary DNAs were individually subjected to PCR analysis using appropriate sets of primers (Supplementary Table S1), electrophoresed, and stained by ethidium bromide solution (Supplementary Fig. S2).
Measurement of lignans
Cultured transgenic and control Forsythia plants were transferred into soil in pots, acclimatized for three weeks until rooting, and grown for two months until the plants reached 15 to 20 cm height. The third to fifth leaves from the top of each plant were pooled, frozen in liquid nitrogen, lyophilized to permit measurement of dry weight (DW) using an FDU-2110 device (EYELA, Japan), extracted with 50% methanol (v/v) containing 2.25 µM (final concentration) 2′-ethoxysesamin (Supplementary Fig. S3 and Table S2) as the internal standard, and processed as described previously29. The leaf extracts were subjected into reverse-phase HPLC (Alliance 2960, Waters Corporation, MA) using a Develosil C30-UG-5 column (4.6 × 150 mm; Nomura Chemical, Japan) under conditions described previously26. Lignans were monitored by UV absorption at 283 nm, and their concentrations were calculated by Empower2 software (https://www.waters.com/waters/library.htm?locale=en_US&lid=1529008,Waters Corporation, MA) according to the areas of the peaks in the chromatograms while referencing standard curves of authentic piperitol and sesamin with technical duplicates or triplicates. For LC–MS analysis, the leaf extracts were subjected to LC–MS-IT-TOF (Shimazu, Japan) and analyzed as described previously30. Lignans were detected using a photodiode array detector and and analyzed by LabSolutions LCMS version 3.8.1 (https://www.an.shimadzu.co.jp/lcms/support/download/index.htm, Shimazu, Japan).
Statistical analysis
The lignan contents were measured using five or six biological replicates and statistically analyzed by two-tailed Student’s t tests (Table 2 and Supplemental Fig. S1).
References
Wurtzel, E. T. & Kutchan, T. M. Plant metabolism, the diverse chemistry set of the future. Science 353, 1232–1236 (2016).
Jacobowitz, J. R. & Weng, J.-K. Exploring uncharted territories of plant specialized metabolism in the postgenomic era. Ann. Rev. Plant Biol. 71, 631–658 (2020).
Durazzo, A. et al. Dietary lignans: Definition, description and research trends in databases development. Molecules 23, 3251 (2018).
Rodríguez-García, C., Sánchez-Quesada, C., Toledo, E., Delgado-Rodríguez, M. & Gaforio, J. J. Naturally lignan-rich foods: A dietary tool for health promotion?. Molecules 24, 917 (2019).
Satake, H. et al. Essences in metabolic engineering of lignan biosynthesis. Metabolites 5, 270–290 (2015).
Kabe, Y. et al. Annexin A1 accounts for an anti-inflammatory binding target of sesamin metabolites. NPJ Sci. Food 4, 4 (2020).
Nakai, M. et al. Novel antioxidative metabolites in rat liver with ingested sesamin. J. Agric. Food Chem. 51, 1666–1670 (2003).
Majdalawieh, A. F., Massri, M. & Nasrallah, G. K. A comprehensive review on the anti-cancer properties and mechanisms of action of sesamin, a lignan in sesame seeds (Sesamum indicum). Eur. J. Pharm. 815, 512–521 (2017).
Ono, E. et al. Formation of two methylenedioxy bridges by a Sesamum CYP81Q protein yielding a furofuranlignan, (+)-sesamin. Proc. Natl. Acad. Sci. USA 103, 10116–10121 (2006).
Andargie, M., Vinas, M., Rathgeb, A., Möller, E. & Karlovsky, P. Lignans of sesame (Sesamum indicum): A comprehensive review. Molecules 26, 883 (2021).
Dossa, K. et al. The emerging oilseed crop sesamum indicum enters the “Omics” era. Front. Plant Sci. 8, 1154 (2017).
Dossa, K. et al. The genetic basis of drought tolerance in the high oil crop Sesamum indicum. Plant Biotechnol. J. 17, 1788–1803 (2019).
Davin, L. B. & Lewis, N. G. Dirigent proteins and dirigent sites explain the mystery of specificity of radical precursor coupling in lignan and lignin biosynthesis. Plant Physiol. 123, 453–461 (2000).
Suzuki, S. & Umezawa, T. Biosynthesis of lignans and norlignans. J. Wood Sci. 53, 273–284 (2007).
Kim, K. W. et al. Trimeric structure of (+)-pinoresinol-forming dirigent protein at 195 A° resolution with three isolated active sites. J. Biol. Chem. 290, 1308–1318 (2015).
Yonekura-Sakakibara, K. et al. Seed-coat protective neolignans are produced by the dirigent protein AtDP1 and the laccase AtLAC5 in Arabidopsis. Plant Cell 33, 129–152 (2021).
Gasper, R. et al. Dirigent protein mode of action revealed by the crystal structure of AtDIR6. Plant Physiol. 172, 2165–2175 (2016).
Davin, L. B. et al. Stereoselective bimolecular phenoxy radical coupling by an auxiliary (dirigent) protein without an active center. Science 275, 362–366 (1997).
Teponno, R. B., Kusari, S. & Spiteller, M. Recent advances in research on lignans and neolignans. Nat. Prod. Rep. 33, 1044–1092 (2016).
Rosati, C., Cadic, A., Duron, M. & Simoneau, P. Forsythia. Biotechnology in Agriculture and Forestry. In Transgenic Crops VI (eds Pua, E. C. & Davey, M. R.) 299–318 (Springer, 2007).
Umezawa, T. Diversity in lignan biosynthesis. Phytochem. Rev. 2, 371–390 (2003).
Guo, H., Liu, A. H., Ye, M., Yang, M. & Guo, D. A. Characterization of phenolic compounds in the fruits of Forsythia suspensa by high-performance liquid chromatography coupled with electrospray ionization tandem mass spectrometry. Rap. Commun. Mass Spectrom. 21, 715–729 (2007).
Dong, Z. et al. Forsythiae Fructus: A review on its phytochemistry, quality control, pharmacology and pharmacokinetics. Molecules 22, 1466 (2017).
Shiraishi, A. et al. De novo transcriptomes of Forsythia koreana using a novel assembly method: Insight into tissue- and species-specific expression of lignan biosynthesis-related gene. PLoS ONE 11, e0164805 (2016).
Sun, L. et al. Comparative transcriptome analyses of three medicinal Forsythia species and prediction of candidate genes involved in secondary metabolisms. J. Nat. Med. 72, 867–881 (2018).
Murata, J., Matsumoto, E., Morimoto, K., Koyama, T. & Satake, H. Generation of triple-transgenic Forsythia cell cultures as a platform for the efficient, stable, and sustainable production of lignans. PLoS ONE 10, e0144519 (2015).
Kim, H. J. et al. Metabolic engineering of lignan biosynthesis in forsythia cell culture. Plant Cell Physiol. 50, 2200–2209 (2009).
Koyama, T. et al. A regulatory cascade involving class II ethylene response factor transcriptional repressors operates in the progression of leaf senescence. Plant Physiol. 162, 991–1005 (2013).
Tera, M. et al. Identification of a binding protein for sesamin and characterization of its roles in plant growth. Sci. Rep. 9, 8631 (2019).
Murata, J. et al. Oxidative rearrangement of (+)-sesamin by CYP92B14 co-generates twin dietary lignans in sesame. Nat. Commun. 8, 2155 (2017).
Gordaliza, M. Natural products as leads to anticancer drugs. Clin. Transl. Oncol. 9, 767–776 (2007).
Marques, J. V. et al. Next generation sequencing in predicting gene function in podophyllotoxin biosynthesis. J. Biol. Chem. 288, 466–479 (2013).
Lau, W. & Sattely, E. S. Six enzymes from mayapple that complete the biosynthetic pathway to the etoposide aglycone. Science 349, 1224–1228 (2015).
Zhu, Q. et al. Plant synthetic metabolic engineering for enhancing crop nutritional quality. Plant Commun. 1, 100017 (2020).
Sato, F. & Kumagai, H. Microbial production of isoquinoline alkaloids as plant secondary metabolites based on metabolic engineering research. Proc. Jpn. Acad. B. 89, 165–182 (2013).
Kotopka, B. J., Li, Y. & Smolke, C. D. Synthetic biology strategies toward heterologous phytochemical production. Nat. Prod. Rep. 35, 902–920 (2018).
Pyne, M. E., Narcross, L. & Martin, V. J. J. Engineering plant secondary metabolism in microbial systems. Plant Physiol. 179, 844–861 (2019).
Choi, K. R. et al. Systems metabolic engineering strategies: integrating systems and synthetic biology with metabolic engineering. Trends Biotechnol. 37, 817–837 (2019).
Birchfield, A. S. & McIntosh, C. A. Metabolic engineering and synthetic biology of plant natural products: A minireview. Curr. Plant Biol. 24, 100163 (2020).
Arya, S. S., Rookes, J. E., Cahill, D. M. & Lenka, S. K. Next-generation metabolic engineering approaches towards development of plant cell suspension cultures as specialized metabolite producing biofactories. Biotech. Adv. 45, 107635 (2020).
Hano, C. F., Dinkova-Kostova, A. T., Davin, L. B., Cort, J. R. & Lewis, N. G. Lignans: Insights into their biosynthesis, metabolic engineering, analytical methods and health benefits. Front. Plant Sci. 11, 630327 (2021).
De Martinis, D. et al. Plant molecular farming: Fast, scalable, cheap, sustainable. Front. Plant Sci. 7, 1148 (2016).
Fuss, E. Lignans in plant cell and organ cultures: An overview. Phytochem. Rev. 2, 307–320 (2003).
Bayindir, Ü., Alfermann, A. W. & Fuss, E. Hinokinin biosynthesis in Linum corymbulosum Reichenb. Plant J. 55, 810–820 (2008).
Ragamustari, S. K. et al. Substrate-enantiomer selectivity of matairesinol O-methyltransferases. Plant Biotechnol. 31, 257–267 (2014).
Lalaleo, L. et al. Effect of in vitro morphogenesis on the production of podophyllotoxin derivatives in callus cultures of Linum album. J. Plant Physiol. 228, 47–58 (2018).
Renouard, S. et al. Investigation of Linum flavum (L.) hairy root cultures for the production of anticancer aryltetralin lignans. Int. J. Mol. Sci. 19, 990 (2018).
Bazaldúa, C. et al. Improving the production of podophyllotoxin in hairy roots of Hyptis suaveolens induced from regenerated plantlets. PLoS ONE 14, e0222464 (2019).
Mikac, S. et al. Bioproduction of anticancer podophyllotoxin and related aryltretralin-lignans in hairy root cultures of Linum flavum L. In Reference Series in Phytochemistry. Plant Cell and Tissue Differentiation and Secondary Metabolites (eds Ramawat, K. G. et al.) 503–540 (Springer, 2021).
Markulin, L. et al. Pinoresinol–lariciresinol reductases, key to the lignan synthesis in plants. Planta 249, 1695–1714 (2019).
Muranaka, T., Miyata, K., Ito, K. & Tachibana, S. Production of podophyllotoxin in Juniperus chinensis callus cultures treated with oligosaccharides and a biogenetic precursor. Phytochemistry 49, 491–496 (1998).
Esmaeilzadeh Bahabadi, S. et al. Time-course changes in fungal elicitor-induced lignan synthesis and expression of the relevant genes in cell cultures of Linum album. J. Plant Physiol. 169, 487–491 (2012).
Bahabadi, S. E. et al. Increased lignan biosynthesis in the suspension cultures of Linum album by fungal extracts. Plant Biotechnol. Rep. 5, 367–373 (2011).
Bahabadi, E. S. The effect of chitosan and chitin oligomers on gene expression and lignans production in Linum album cell cultures. J. Med. Plants 13, 46–53 (2014).
Hazra, S., Bhattacharyya, D. & Chattopadhyay, S. Methyl jasmonate regulates podophyllotoxin accumulation in Podophyllum hexandrum by altering the ROS-responsive podophyllotoxin pathway gene expression additionally through the down regulation of few interfering miRNAs. Front. Plant Sci. 8, 164 (2017).
Corbin, C. et al. Functional characterization of the pinoresinol–lariciresinol reductase-2 gene reveals its roles in yatein biosynthesis and flax defense response. Planta 246, 405–420 (2017).
Yousefzadi, M. et al. The effect of light on gene expression and podophyllotoxin biosynthesis in Linum album cell culture. Plant Physiol. Biochem. 56, 41–46 (2012).
Hata, N. et al. Differences in plant growth and leaf sesamin content of the lignan-rich sesame variety “Gomazou” under continuous light of different wavelengths. Plant Biotechnol. 30, 1–8 (2013).
Morimoto, K. et al. Effects of light on production of endogenous and exogenous lignans by Forsythia koreana wildtype and transgenic cells. Plant Biotechnol. 28, 331–337 (2011).
Shultz, B. J., Kim, S. Y., Lau, W. & Sattely, E. S. Total biosynthesis for milligram-scale production of etoposide intermediates in a plant chassis. J. Am. Chem. Soc. 141, 19231–19235 (2019).
Acknowledgements
We thank the staff of the Niigata Prefectural Botanical Gardens and Dr. Umezawa for providing shoots of Forsythia plants. We thank Drs. Shinzo Tsuda, Yukihisa Katsumoto, and Yoshihiko Nanasato for helpful discussion and the Ministry of Economy, Trade and Industry for funding.
Author information
Authors and Affiliations
Contributions
T.K., E.M., T.O., J.M., M.H., N.H., A.O., and E.O. performed experiments, and T.K. and H.S. designed the research and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Koyama, T., Matsumoto, E., Okuda, T. et al. Transgenic Forsythia plants expressing sesame cytochrome P450 produce beneficial lignans. Sci Rep 12, 10152 (2022). https://doi.org/10.1038/s41598-022-14401-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-14401-9
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.