Abstract
X-autosome translocation (XY1Y2) has been reported in distinct groups of vertebrates suggesting that the rise of a multiple sex system within a species may act as a reproductive barrier and lead to speciation. The viability of this system has been linked with repetitive sequences located between sex and autosomal portions of the translocation. Herein, we investigate Oecomys auyantepui, using chromosome banding and Fluorescence In Situ Hybridization with telomeric and Hylaeamys megacephalus whole-chromosome probes, and phylogenetic reconstruction using mtDNA and nuDNA sequences. We describe an amended karyotype for O. auyantepui (2n = 64♀65♂/FNa = 84) and report for the first time a multiple sex system (XX/XY1Y2) in Oryzomyini rodents. Molecular data recovered O. auyantepui as a monophyletic taxon with high support and cytogenetic data indicate that O. auyantepui may exist in two lineages recognized by distinct sex systems. The Neo-X exhibits repetitive sequences located between sex and autosomal portions, which would act as a boundary between these two segments. The G-banding comparisons of the Neo-X chromosomes of other Sigmodontinae taxa revealed a similar banding pattern, suggesting that the autosomal segment in the Neo-X can be shared among the Sigmodontinae lineages with a XY1Y2 sex system.
Similar content being viewed by others
Introduction
Chromosomal rearrangements are drivers in karyotypic evolution and are often associated with speciation1,2,3,4,5. Mammals are known to exhibit a stable sex determination system, but distinct sex-autosome translocations may have triggered the separation of Theria and Prototheria (monotremes) (190 MYA) and between Eutheria (placental mammals) and Metatheria (marsupials) (166 MYA)6. Although the euchromatic region of the X chromosome is considered conserved among highly rearranged karyotypes of placental mammals6, recent investigations on Arvicolinae (Myomorpha) rodents have shown that the X chromosome has undergone several intrachromosomal rearrangements, such as centromere shifts, peri- and paracentric inversions, that were also accompanied by repetitive sequences7. Regardless of whether chromosomal rearrangements are the primary cause of speciation2,8, or whether karyotypic divergence between closely related species are a casualty of the speciation process9,10, the most deleterious among the speciation-linked rearrangements11,12 are tandem translocations, reciprocal translocations13,14 and X-autosome translocations15,16.
The rise of an X-autosome translocation is subordinated to the same epigenetic mechanism that guarantees dosage compensation between normal females (XX) and males (XY) by silencing one of the Xs in females17. In this type of event, the inactivation progress in one of the X chromosomes of females18 spreads to the autosomal segment translocated to the X, silencing genes in the autosomal portion19 generating deletion/duplications with deleterious effects20.
Although deleterious effects of sex-autosome translocations have been described in the literature for humans and mice (e.g., male sterility; embryonic lethality)15,21,22, this type of chromosomal rearrangement has been reported in natural populations of distinct groups of vertebrates, such as fish23,24, anurans25 and mammals17,26. The presence of intercalary heterochromatic blocks between autosomal and ancestral X chromosome segments could suppress the X-inactivation progress in the autosomal segment, allowing viability in this system16,27,28,29. Several studies have shown the presence of heterochromatic blocks, telomeric repeats and/or rDNA (ribosomal DNA) clusters in different mammalian lineages that exhibit X-autosome translocation, for example in bats (Chiroptera) of genera Artibeus, Carollia, and Uroderma29,30,31,32; in rodents (Rodentia) of genera Nannomys (Muridae)17, Proechimys (Echimyidae)33,34,35, and Taterillus (Muridae)16; in diprotodont marsupials (Diprotodontia) of genus Wallabia36; and in ruminants (Artiodactyla) of genera Antilope and Gazella37.
In rodents from the Brazilian Amazon, the XX/XY1Y2 multiple sex system has been reported only in two genera from the Echimyidae family: Lonchothrix38 and Proechimys33,34,35. In Lonchothrix emiliae, the multiple sex system was identified based on classic banding38, while in the Proechimys taxa it was detected by FISH (Fluorescence In Situ Hybridization) with whole chromosome probes (chromosome painting) from P. roberti and P. goeldii35. In Sigmodontinae rodents (Rodentia, Cricetidae), the Oryzomyini tribe currently comprises 29 genera and is the most diverse of the 11 tribes within the subfamily39,40,41, but multiple sex systems are acknowledged solely in representatives of the Akodontini, Phyllotini and Reithrodontini tribes: Deltamys kempi (Akodontini) exhibits a X1X1X2X2/X1X2Y sex system due to a translocation involving chromosomes 2 and Y42; Salinomys delicatus (Phyllotini) shows a XY1Y2 system43; and Reithrodon (Reithrodontini) exhibits a XY1Y2 system (Uruguay population) and a Neo-XY system (Brazil population)44.
In Oryzomyini, the genus Oecomys has been particularly challenging in taxonomy, distribution patterns and speciation mechanisms. Comprising 19 species to date, Oecomys has been investigated using several approaches, such as morphology, nuclear DNA (nuDNA), mitochondrial DNA (mtDNA), and cytogenetics, which have shown that some lineages correspond to species complexes39,45,46,47,48,49,50,51. Oecomys auyantepui has been recognized as a monophyletic lineage and a monotypic taxon48,52. The species is distributed from southeastern Venezuela to north-central Brazil, in the Guiana subregion of Amazonia39, and exhibits two sympatric populations with distinct diploid numbers (2n) of 64 and 66 and autosomal fundamental numbers (FNa) of 110 and 114, respectively52. A third karyotype of 2n = 72/FNa = 80 was described53. In addition, an interstitial telomeric sequence (ITS) was identified at the centromeric region of the bi-armed X chromosomes in karyotypes with 2n = 64 and 66, which suggests that chromosomal rearrangements have driven the evolution of this chromosome in O. auyantepui52.
It is noteworthy that cytogenetics studies with Oecomys have shown a substantial diversity in 2n and FNa, ranging from 54 to 86 and from 62 to 140, respectively39,45,47,48,50,52,53,54,55. However, specific events that shaped extant karyotypes remain unclear for most species, except for O. catherinae from Pará (OCA-PA; 2n = 62/FNa = 62), O. catherinae from Rio de Janeiro (OCA-RJ; 2n = 60/FNa = 62), O. paricola cytotype A (OPA-A; 2n = 72/FNa = 75), O. paricola cytotype B (OPA-B; 2n = 70/FNa = 75), and O. paricola cytotype C (OPA-C; 2n = 70/FNa = 72) that were investigated by chromosome painting with Hylaeamys megacephalus whole chromosome probes (HME; Oryzomyini)47,50. In addition to elucidating the chromosomal rearrangements that occurred in these species, the chromosome painting analysis helped to delineate taxonomic limits, as the authors47,50 were able to identify a hidden diversity and proposed that O. catherinae and O. paricola “eastern clade” were composed of two and three species, respectively.
Considering the evolutionary force of chromosomal rearrangements regarding speciation and diversification of species, we set out to investigate if the emergence of a new sex-system triggered the speciation process in the monotypic taxon Oecomys auyantepui.
In order to achieve this goal, we used classic cytogenetics, telomeric and HME whole chromosome probes54, mtDNA (mitochondrial DNA) and nuDNA (nuclear DNA) sequences. Here we discuss the chromosomal evolution of the genus, and report for the first time a multiple sex system (XX/XY1Y2) in Oryzomyini rodents. We also compared the taxa from the present study with other species analyzed elsewhere using the same set of probes47,50,54,55,56,57,58,59.
Results
Classic and molecular cytogenetics
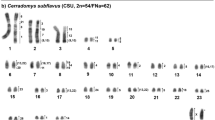
Oecomys auyantepui (OAU) has a 2n = 64♀65♂/FNa = 84 karyotype, with a multiple sex system (XX/XY1Y2). The autosomal set consists of 20 acrocentric pairs (1–20) and 11 meta/submetacentric pairs (21–31). In females sex chromosomes were recognized as two medium-sized submetacentric Neo-X chromosomes; in males sex chromosomes were identified as one Neo-X and two Ys: Y1 chromosome was a medium submetacentric (original Y) and Y2 was a small acrocentric (Xp homologue) (Fig. 1a). The constitutive heterochromatin (CH) is distributed in the centromeric regions of almost all autosomes, the Neo-X and Y2 chromosomes. The CH is a small region in most of the autosomes and the Y1 chromosome has a large heterochromatic block in the long arm (Fig. 1b).
Oecomys auyantepui (2n = 65♂/FNa = 84) (a) G-banded karyotype with chromosome painting revealed by Hylaeamys megacephalus (HME) whole chromosome probes54, and (b) C-banded karyotype. An asterisk indicates a centromere.
Cross-species FISH with HME probes yielded 42 signals on the OAU chromosomes (Fig. 1a, Table 1, see Supplementary Figs. 1 and 2). Ten autosomal probes are conserved; of them, four (HME 8, 15, 24 and 25) hybridize to whole chromosomes of OAU (1, 10, 11 and 28, respectively) and six (HME 7, 12, 18, 20, 21 and 26) hybridized with portions of chromosomes of OAU (21q, 23p, 25q, 25p proximal, 8q distal and 25p distal, respectively). Twelve autosomal probes show multiple signals in OAU: HME 1 hybridize to OAU 2, 19 and 23q; HME 2 hybridize to OAU 4 and 22q; HME 4 hybridize to OAU 14 and 24; HME 5 hybridize to OAU 7, 18 and 31; HME 6 hybridize to OAU 21p and 30; HME (9,10) hybridize to OAU 5, 22p and 26p; HME 11 hybridize to OAU 9 and 20; HME (13,22) hybridize to OAU 8q proximal, 17 and 26q; HME 14 hybridize to OAU 13q distal and 15; HME (16,17) hybridize to OAU 6 and 27; HME 19 hybridize to OAU 12 and 13q proximal; HME 23 hybridize to OAU 16 and 29.
The HME 3 probe hybridizes to the short arm of the Neo-X chromosome (OAU Xp), and also hybridizes to OAU 3, and Y2; the HME X chromosome hybridizes to the long arm of the Neo-X (OAU Xq).
Seven OAU autosomal pairs show hybridization signals to multiple HME probes: OAU 8 (HME (13,22)/21); OAU 13 (HME 19/14); OAU 21 (HME 6/7); OAU 22 (HME (9,10)/2); OAU 23 (HME 12/1); OAU 25 (HME 26/20/18); OAU 26 (HME (9,10)/(13,22)) (Fig. 2).
FISH results obtained from O. auyantepui (OAU; 2n = 64♀65♂/FNa = 84), using HME whole chromosome probes54. Each box corresponds to an OAU pair shown in Fig. 1a that corresponded to more than one HME homologue. Single or multiple images are presented to exhibit full coverage with HME probes on OAU chromosomes. OAU chromosomal pairs identification are shown below the chromosomes, while HME probes are shown beside the chromosomes. An asterisk indicates a centromere. HME whole chromosome probes are shown in green (FITC) and red (CY3); the counterstaining is blue (DAPI).
FISH with telomeric probes showed hybridization signals at the distal regions of all chromosomes, plus a large interstitial telomeric sequence (ITS) at the centromere of the Neo-X chromosome (Fig. 3).
Phylogenetic analysis
A more detailed phylogenetic analysis of the genus Oecomys was already proposed48. Thus, in this work we focused on O. auyantepui and representatives of each Oecomys species/clade recognized in the literature48,49,51 (Supplementary Table 1). The genus Oecomys was recovered as monophyletic in the topologies obtained with the Cytochrome b (Cytb) dataset, the Cytochrome C Oxidase Subunit I (COI) dataset, and with the concatenated dataset (Cytb + beta-fibrinogen intron 7 [FGB-I7]), with high support values recorded only in the Bayesian Inference (BI) analyses (Figs. 4, 5, 6). In the Cytb topology, lineages of O. bicolor and O. cleberi were not recovered as reciprocally monophyletic, as well as lineages of O. mamorae and O. franciscorum (Fig. 5).
Bayesian Inference topology based on Cytochrome C Oxidase Subunit I. The numbers above branches indicate posterior probability values for Bayesian Inference analysis (only values > 0.80 are shown) and bootstrap values for Maximum Likelihood analysis (only values > 65% are shown). Bold numbers indicate the samples from this study. Sample data are provided in Supplementary Table 1.
Bayesian Inference topology based on Cytochrome b. The numbers above branches indicate posterior probability values for Bayesian Inference analysis (only values > 0.60 are shown) and bootstrap values for Maximum Likelihood analysis (only values > 60% are shown). Bold numbers indicate the samples from this study. Percentage values are mean genetic distances (p-distances) between high supported selected lineages of O. auyantepui. Representatives of different lineages within O. bicolor, O. cleberi, O. roberti, O. mamorae, O. paricola, and O. catherinae complexes are indicated by the letters C (central clade), E (eastern clade), N (northern clade), NW (northwestern clade), S (southern clade), W (western clade), and Wm (westernmost clade), following48. Subscribed letters b and c indicate lineages attributed by48 to O. bicolor and O. cleberi, respectively. Subscribed letters f and m indicate lineages attributed by48 to O. franciscorum and O. mamorae, respectively. Cross (+) denotes karyotype information not obtained from the specimen included in the phylogenetic analysis. Sample data are provided in Supplementary Table 1.
Bayesian Inference topology based on mitochondrial Cytochrome b and nuclear beta-fibrinogen intron 7 concatenated. The numbers above branches indicate posterior probability values for Bayesian Inference analysis (only values > 0.90 are shown) and bootstrap values for Maximum Likelihood analysis (only values > 60% are shown). Bold numbers indicate the samples from this study. Representatives of different lineages within O. bicolor, O. cleberi, O. roberti, O. paricola, and O. catherinae complexes are indicated by the letters C (central clade), E (eastern clade), N (northern clade), NW (northwestern clade), S (southern clade), W (western clade), and Wm (westernmost clade), following48. Asterisk (*) denotes specimens identified as O. bicolor by48. Cross (+) denotes karyotype information not obtained from the specimen included in the phylogenetic analysis. Sample data are provided in Supplementary Table 1.
We found mean interspecific p-distances ranging from 6.47% (between O. concolor and Oecomys sp.2) to 14.80% (between O. franciscorum/O. mamorae and O. matogrossensis), and mean intraspecific p-distances varying from zero in several species to 6.2% in O. bicolor/O. cleberi (2.15% in O. auyantepui, particularly; Table 2).
In all three datasets, O. auyantepui was recovered as a monophyletic taxon with high support. The COI phylogeny was the only one that included all O. auyantepui karyotyped samples from this work and from the literature51,52, as well as most sequences available on GenBank (Supplementary Table 1). In the COI topology, specimens of O. auyantepui formed a polytomy, with no resolution among most specimens, including those with similar karyotypes (Fig. 4). In the Cytb topology, the specimen N228 was recovered as the most divergent within O. auyantepui, with 5.20% of mean genetic divergence from its conspecifics. The remaining specimens formed a subclade with neither resolution nor support among most of the specimens. As in the COI topology, specimens with similar karyotypes did not nest in a subclade (Fig. 5). Finally, in the concatenated data topology, the specimen ROM 114,316 was the first one to diverge, and the specimen ROM 114,059 was recovered as sister to the 2n = 65♂/FNa = 84 specimens included in this analysis. Although these latter specimens appeared as a subclade, there was no support for that (Fig. 6).
Discussion
Chromosomal evolution and signatures in Oecomys (Rodentia, Sigmodontinae)
As mentioned above, there is a large variation in 2n from 54 to 86 and in FNa from 62 to 140 among the Oecomys species, with karyotypes mainly composed of one-armed chromosomes45,47,48,50,52,53,60,61. The variation in 2n and FNa occur both within and between species, indicating that fusions/fissions, pericentric inversions (or centromeric repositioning), translocations, and addition/deletion of constitutive heterochromatin are the main forces acting in the chromosomal evolution of this group of Sigmodontinae rodents. Thus, we used chromosome painting to make a comparative analysis of taxa in the present study (O. auyantepui) and in the literature (O. catherinae and O. paricola) to precisely identify the rearrangements among them47,50.
As a result, chromosome painting analysis in the karyotypes of O. auyantepui (OAU), O. paricola (OPA-A, OPA-B and OPA-C) and O. catherinae (OCA-PA and OCA-RJ) (Supplementary Table 2) showed that the chromosomal variation in 2n from 60 to 72 and in FNa from 62 to 84 are due to 23 fusion/fission events, four translocations, seven pericentric inversions and amplification/deletion of constitutive heterochromatin on two autosomal syntenic blocks plus the X chromosome (Supplementary Fig. 3), with only seven syntenic blocks conserved without detectable rearrangements. Remarkably, we observed that the rearrangements that differentiate OPA cytotypes (OPA-A, OPA-B, and OPA-C) from each other are different from those responsible for the variability between OCA cytotypes (OCA-PA and OCA-RJ) and OAU. This suggests that the rearrangements mainly occurred in distinct syntenic blocks among these species (Supplementary Fig. 3). Consequently, we propose that each of these three species has evolved independently and has not followed the same path of rearrangements or the same chromosomes.
Moreover, by detecting an elevated number of chromosomal rearrangements among three taxa (O. auyantepui, O. catherinae, and O. paricola) with not-so-distant 2n (from 60 to 72), we assume that the chromosomal evolution in Oecomys is more complex than previously thought. In this sense, the use of HME whole chromosome probes in representatives of the main lineages of Oecomys provides a more accurate view of chromosomal evolution, and associated with detailed phylogeographic studies, is a key factor in understanding the speciation processes of this diverse and speciose group of Sigmodontinae rodents.
By comparing the OAU karyotype with the other Sigmodontinae species investigated by HME whole chromosome probes (Supplementary Table 2), we identified that OAU exhibited the chromosomal signatures proposed for the Oecomys genus, namely the fragmentation of HME 1 into three blocks and the syntenic block HME (13,22)/2150. We also detected an exclusive trait for OAU, the syntenic block HME 26/20/18, which is different from those described for OCA (HME (9,10)/14/5, 23/19/11 and 26/11) and OPA (HME 4/19); OAU also shared and the fragmentation of HME 3 into two blocks with OPA, previously described as exclusive for this species50. Among the eleven chromosomal signatures proposed for the Sigmodontinae subfamily (HME 7/(9,10), 8, 1/12, 6/21, 11/(16,17), 5/(16,17), 20/(13,22), 15, 19/14/19, 24, and 26)47,50,61,69,70,71,72,73, OAU exhibits only five: HME 8, 15, 24 and 1/12, plus the chromosomal signature HME 19/14/19, which is present as a derived character due to a fission that generates the HME 19/14 (OAU 13) and 19 (OAU 12).
New sex system in Oecomys auyantepui (XX/XY1Y2)
In the genus Oecomys, sex chromosomes variation in size and morphology are frequently related to the addition/deletion of CH, which is common in rodents62; the one-armed X chromosome often exhibits CH at the centromeric region, while the bi-armed form presents a heterochromatic block in the short arm and the variation in length of the Y is often due to the size of the heterochromatic block in the long arm45,47,48,50,52. However, the bi-armed X chromosome of O. auyantepui from the present study exhibits an euchromatic short arm, with CH at the centromere (Fig. 1b). This indicates that events other than the general addition/deletion of CH were involved in this variation, and this is supported by the HME 3 hybridization signal in the short arm of OAU Neo-X (Xp) and Y2 (see Supplementary Fig. 1), validating the emergence of a new sex system (XX/XY1Y2) in O. auyantepui (2n = 64♀65♂/FNa = 84). Regarding Sigmodontinae rodents, this is the first record of a multiple sex system in Oryzomyini, since this type of sex determination system has previously been reported only in representatives of Akodontini (X1X1X2X2/X1X2Y)42, Phyllotini (XY1Y2)43 and Reithrodontini (XY1Y2 and Neo-XY)44 tribes.
In order to understand the Neo-X origin in O. auyantepui, we constructed a dendrogram that shows a hypothetical scenario with the chromosomes involved. We made a comparative analysis among the other Sigmodontinae taxa studied with the same set of probes (Supplementary Table 2) and considered as outgroup the 16 karyotypes from the genera Akodon, Blarinomys, Necromys, Oxymycterus, Thaptomys (Akodontini), Cerradomys, Hylaeamys, and Neacomys (Oryzomyini), while as ingroup we considered the karyotypes of Oecomys catherinae, O. paricola and O. auyantepui (Oryzomyini). Except for Cerradomys, all karyotypes from the outgroup showed a HME 3 hybridization signal in one large acrocentric chromosome pair. The X was acrocentric, or bi-armed with a heterochromatic block in the short arm. Thus, we propose that: (1) the ancestral forms of the HME 3 and X chromosomes were medium acrocentrics; (2) O. catherinae maintained the autosomal ancestral form, while the X exhibited an addition of CH in the short arm; (3) the HME 3 divided by fission into two blocks of unequal size (one large and one small) before the diversification events that led to the O. paricola and O. auyantepui species; (4) in O. paricola, the HME 3 small block remained as an independent chromosome, and the X exhibited an addition of CH in the short arm; (5) in O. auyantepui, a Robertsonian translocation between the HME 3 small block with the acrocentric X formed the Neo-X chromosome (Fig. 7).
Idiograms of the chromosomes involved in the emergence of a new sex-system (XX/XY1Y2) in Oecomys auyantepui (OAU; 2n = 64♀65♂/FNa = 84), as assessed based on HME probes54. Idiograms in black (Y and Y1) were evaluated only by classical cytogenetics. Arrows show direction of chromosome change. We used O. paricola-cytotype A50 and O. catherinae-Pará47 as representatives of their respective species. Numbers below idiograms correspond to the identification of the chromosomal pair; numbers beside idiograms correspond to the HME probes. The bottom box encompasses an idiogram of HME karyotype previously elaborated58 and adapted in the present study. Each HME chromosome is shown with a single color, except the pairs (9,10), (13,22) and (16,17), which have one color each. (H) Indicates large block of constitutive heterochromatin.
The proposal that intercalary heterochromatic blocks, telomeric repeats and/or rDNA clusters between the ancestral X and the translocated autosome served as a boundary that suppresses the X-inactivation process in the autosomal portion16,17,28,29,30 is in accordance with our results, which show a centromeric heterochromatic block (Fig. 1b) and a large block of ITS in the Neo-X chromosome of O. auyantepui (Fig. 3). In the rodent Mus minutoides (Muridae) immunofluorescence techniques demonstrated patterns of histone modification in three types of sex chromosomes (Y, X and a mutant X) that confirmed that the X-inactivation does not spread into the translocated autosomal portion17. This feature is of prime importance and guarantees the viability of this multiple sex system. In fact, several studies have described natural populations of vertebrates (e.g., bats, rodents, marsupials and ruminants) that possess this type of rearrangement16,17,26,30,31,32,35,36,37,63; in cases where such repetitive sequences are absent, deleterious effects such as poor viability and infertility are observed15,16. The contribution of repetitive sequences in the evolution of the X chromosome was also proposed7. The authors microdissected the Terricola savii X chromosome into five specific-region probes, hybridized in 20 species of Arvicolinae rodents (Myomorpha), and identified multiple intrachromosomal rearrangements, such as centromere shifts, peri- and paracentric inversions, which were related to the amplification and distribution of repetitive sequences among the Xs of distinct Arvicolinae species7.
Exceptions from this proposal are documented in the common ancestor of eutherian mammals, since a sex-autosome translocation occurred and it may have triggered the separation between Eutheria (placental mammals) and Metatheria (marsupials) (166 MYA)6, with no intercalary heterochromatic block found between the X and autosomal ancestral segments. Distinct processes from the intercalary heterochromatic block would be involved in the regulation of X-autosome viability and in the suppression of deleterious effects16.
It is noteworthy that telomeric repeats occur at the ends of chromosomes where they provide stability, while ITS are relics of chromosomal fusions that arose during karyotype evolution64. Thus, although we have identified other chromosomes resulting from Robertsonian translocations in O. auyantepui (OAU 21–23, 25; Figs. 2 and 3), the ITS was found only in the Neo-X. An investigation of the effects of these telomeric sequences on gene expression, recombination and rearrangements, was made by introducing 800 bp of the telomere repeat (TTAGGG) in the adenosine phosphoribosyltransferase (APRT) gene in Chinese hamster ovary cells65. The main result was that gene rearrangements were greatly increased65. This type of chromosome instability is in accordance with the proposal that het-ITS (heterochromatic-ITS) seem to be intrinsically prone to breakage66, and that ITSs are hotspots for chromosomal rearrangements64. Therefore, the elimination of this sequence during chromosomal evolution could be a mechanism that provides karyotypic stability64,65,66,67 and might explain its absence in the rearranged autosomes of O. auyantepui, while its presence in the Neo-X makes the latter prone to other chromosomal rearrangements, despite providing stability against X-inactivation of autosomal segments16,17.
We noticed that the other cytotypes found in O. auyantepui (2n = 64, 66 and 72) from the Jatapú and Jari Rivers (Fig. 8, localities 5 and 4, respectively) exhibited distinct morphologies of the X chromosomes (medium submetacentric, large metacentric, and large submetacentric, respectively). Although they are found within a simple sex determination system (XX/XY), the X chromosomes had euchromatic short arms and repetitive sequences at the centromere, such as an ITS in the karyotypes with 2n = 64, 6652. Perhaps these differences in size and morphology, plus the presence of ITS, could be a clue to a more complex rearrangement of the X chromosome during its evolution.
Map showing the distribution, study area and sampling points for Oecomys auyantepui specimens analysed in the present study and from literature. Available karyotypic information is shown with description of 2n (diploid number), FNa (autosomal fundamental number), and if is associated with simple (XX/XY) or multiple (XX/XY1Y2) sex system. Each symbol indicates the karyotype obtained from the locality; more than one symbol indicates that distinct karyotypes were collected at the same locality. Geographic limits of O. auyantepui are based on sample points provided in the present study (localities 1–3), and from literature: locality 452, locality 553, localities 6–1439, and from GenBank (localities 15–35). The localities mentioned are: Brazil: Pará, Óbidos, forest fragment 7 km distant from the town’s center (locality 1); Trombetas State Forest (locality 2); Grão-Pará Ecological Station (locality 3); Jari River (locality 4); Amazonas, Jatapú River (locality 5); 80 km North of Manaus (locality 6); Amapá, Serra do Navio (locality 7). French Guiana: Trois Sauts (locality 8); Paracou (locality 9). Suriname: Brokopondo, Brownsberg Nature Park (locality 10); Sipaliwini, Avanavero Falls, Kabalebo River (locality 11). Guyana: Upper Takutu–Upper Essequibo (locality 12), Cuyuni-Mazaruni, Kartabo, Cuyuni River (locality 13). Venezuela: Bolívar, Auyan-tepuí, South slope, Río Caroni (locality 14). Samples obtained from GenBank (localities 15–35) are detailed in Supplementary Table 1.
Investigations carried out in some groups with multiple sex systems show that the chromosomal evolution in the Neo-X and/or Neo-Y gives rise to other derived systems. This is described in Stenodermatinae bats68, in which chromosome painting revealed that a XY1Y2 system originated a Neo-XY system, due to a translocation between Y1 and Y2; this Neo-XY has then diverged into two more derivate systems: in one branch, a fission in the Neo-X created a X1X1X2X2/X1X2Y; while in the other branch, a fusion between an autosome and the Neo-Y originated a Neo-X1X1X2X2/X1X2Y. In rodents from the genus Reithrodon (Sigmodontinae, Reithrodontini), the Uruguay population exhibits a XY1Y2 system, while the Brazil population shows a Neo-XY system44; a hypothetical intermediate Neo-X/Y1Y2 formula was the ancestor for the Uruguayan form, while the Brazilian form originated through a fusion between the Y1 and Y2, that gave rise to the Neo-XY system44. The evolutionary process and specific events responsible for the variation in size and morphology of the X chromosomes in the O. auyantepui cytotypes (2n = 64, 66 and 72)52,53 will be elucidated only after the employment of HME whole chromosome probes.
Taking into consideration the phylogenetic relationships and karyotypic data within O. auyantepui, the COI phylogeny is the only one that includes all karyotyped samples for this species from both the present study (2n = 64♀65♂) and the literature (2n = 64, 66 and 72)52,53. Despite being considered as a valuable tool for highlighting cases requiring systematic attention among small mammal species69, our COI sequences of O. auyantepui formed a polytomy with only a few subclades mostly with no satisfactory support (Fig. 4). In fact, specimens of O. auyantepui with the multiple sex system grouped in a separated branch only in the concatenated Cytb + FGB-I7 topologies, but with low support in both BI and ML analyses (Fig. 6). Although our molecular analyses do not exhibit a better resolution in the terminal branches, the karyotypic information indicates that we are dealing with at least two distinct lineages. We have three karyotypes with a simple sex system XX/XY (2n = 64, 66 and 72) that could represent variation within a single lineage, while the other lineage corresponds to specimens with a multiple sex system XX/XY1Y2 (2n = 64♀65♂).
The fact that four potentially karyotypic variants are in a small distribution area (Localities 1–4, Fig. 8) indicates that isolating mechanisms are operating, and the rise of a multiple sex determination system may be acting as a post-zygotic barrier between these apparently sympatric populations, since the difference in sex determination systems will generate aberrations in meiotic synapsis. Taking into consideration the low level of genetic divergence within O. auyantepui (mean p-distance 2.15%; Table 2) and the impossibility of interbreeding between these two sex systems, it is most likely that a speciation process is already on course. Similar results were observed in Deltamys rodents (Sigmodontinae, Akodontini) where the two distinct sex determination systems (XX/XY and X1X1X2X2/X1X2Y) act as a reproductive barrier that is reflected in the phylogenetic divergence of 11.13–12.14% between the two divergent lineages42. Despite the low level of genetic divergence in O. auyantepui mentioned above, it is worthy of note that the specimen N228 exhibited 5.20% of mean genetic divergence from other conspecifics included in the Cytb topology (Fig. 5). Further studies are necessary to evaluate if this individual represents a new species, which may be either cytogenetically or morphologically distinct from its closely related O. auyantepui individuals.
Some authors have proposed that changes in the sex determination system can alter behavior patterns and contribute to pre-zygotic isolation mechanisms, as described in threespine stickleback fish Gasterosteus aculeatus70, in which populations with XX/XY and X1X1X2X2/X1X2Y systems exhibit different courtship behavior. In the rodent Mus minutoides (Muridae), it was observed that XY females have more reproductive success than XX females and are more aggressive and have a stronger bite than XY males71. Thus, differences in the X chromosomes of O. auyantepui could lead to behavioral modifications and act in pre-zygotic isolation mechanisms between these two taxonomic entities.
An evaluation of the genetic (Cytb) structure of Oecomys aff. roberti (= O. tapajinus) populations observed isolated and stable populations, but with no influence from the mid-Araguaia River opposite banks72. Two sympatric O. paricola “eastern clade” populations (OPA-A and OPA-B) from the Belém area of endemism exhibited distinct karyotypes50. Both works reached similar conclusions for their respective analyzed species, proposing that Oecomys taxa can exhibit isolated populations in the absence of strong geographic barriers.
As discussed above, the fact that four potentially karyotypic variants of O. auyantepui are in a small distribution area (Localities 1–4, Fig. 8) could be explained by this type of populational structure for Oecomys taxa50,72. In this scenario, the rise of a rearranged chromosomal form within an isolated population could be stablished in a few generations, as rodents exhibit a tendency to be organized in demes73,74 with low interbreeding among distinct populations35. The reproductive barrier would be more intense between the ancestral (XX/XY) and derived (XX/XY1Y2) systems, as the hybrid offspring would be infertile6. We propose that we are dealing with a case of chromosomal speciation, triggered by the emergence of a new sex system (XX/XY1Y2) in isolated O. auyantepui populations. Consequently, O. auyantepui is a species complex with at least two distinct lineages, which disagrees with the literature data that recover this taxon as a monotypic species sensu48.
A comparative analysis of the G-banding patterns among the Sigmodontinae Neo-X chromosomes of O. auyantepui (present work), Reithrodon (Reithrodontini44), and Salinomys delicatus (Phyllotini43) was performed and this revealed that the autosomal portion translocated in the Neo-X exhibits similar G-banding patterns in the three taxa. This suggests that the same autosomal segment is shared among these distinct lineages. Literature data show that distinct genera within groups that have multiple sex systems may also share the same autosome in the sex-autosome translocation, as in primates from genera Alouatta75 and Aotus76 and in bats from genera Artibeus, and Uroderma77, Chiroderma, Mesophylla, and Vampyriscus68. However, it has been shown that, in rodents from genera Proechimys (Echimyidae) and Nannomys (Muridae), species within a genus can have different autosomes translocated to the X chromosome26,35.
We suggest that the employment of HME whole chromosome probes in Reithrodon, Salinomys delicatus and Oecomys auyantepui with 2n = 64, 66 and 72 will elucidate the origin of Neo-X chromosomes in Sigmodontinae rodents; also, this will shed light on the evolutionary relationships among the four karyotypes of O. auyantepui, and clarify if we are dealing with simple (XX/XY) and multiple (XX/XY1Y2) sex determination systems; or with derived lineages from the XY1Y2 system.
Material and methods
Ethics
The specimens were captured using pitfall traps78 and kept stress-free with full access to food and water until their necessary euthanasia that was performed in accordance with animal welfare guidelines established by Brazilian resolution CFMV n.1000/2012. The necessary euthanasia occurred in accordance with animal welfare guidelines established by the Animal Ethics Committee (Comitê de Ética Animal) from Universidade Federal do Pará (Permit 68-2015), which also approved all experimental protocols of this research. The captures were authorized by the Brazilian Environment Department under license (IBAMA 02047.000384/2007-34). JCP has a permanent field permit (number 13248) from “Instituto Chico Mendes de Conservação da Biodiversidade”. The Cytogenetics Laboratory from UFPa has a special permit number 19/2003 from the Ministry of Environment for samples transport and 52/2003 for using the samples for research. All methods are reported in accordance with ARRIVE guidelines (https://arriveguidelines.org/).
Samples
We studied the karyotypes of three adult samples of Oecomys auyantepui from distinct locations in Óbidos municipality, Pará state, Brazil. The wildlife samples were collected according to the following: one male specimen (UFPAM 2027) was collected in a forest 7 km distant from the town’s center (01° 51′ 15″ S, 55° 32′ 53.4″ W), one female sample (MPEG 39,927) was collected at the Trombetas State Forest (00° 57′ 45.97″ S, 55° 31′ 20.28″ W), and one male sample (MPEG 40,457) was collected at the Grão-Pará Ecological Station (00° 37′ 49.01″ N, 55° 43′ 42.60″ W). The specimens were deposited at the zoological collections of Museu Paraense Emílio Goeldii (MPEG) and the Museu de Zoologia da Universidade Federal do Pará (MUFPA). Both institutions are in Belém, Pará state, Brazil.
Cytogenetics
The metaphase chromosomal preparations were obtained from bone marrow extraction79. The slides containing chromosomal preparations underwent C-Banding80, G-Banding81 and FISH82 techniques. The FISH experiments were performed with human telomeric probes (All Telomere, ONCOR), and with 24 whole chromosome painting probes from a female of Hylaeamys megacephalus (HME; 2n = 54)54; from the 24 HME whole chromosome probes, 21 correspond to one chromosome each (including the X chromosome), while three corresponded to two pairs of chromosomes each (HME (9,10), (13,22) and (16,17).
Image capture and analysis
We used an Olympus BX41 microscope and a CCD 1300QDS digital camera to obtain digital images from G-banded and C-banded karyotypes, which were analyzed using the GenASIs software v. 7.2.7.34276. The Nikon H550S microscope with a DS-Qi1Mc digital camera captured the FISH images, which were analyzed using the Nis-Elements software. The karyotypes were organized according to the literature83. The final images were edited using Adobe Photoshop CS6 software.
Molecular analysis
We obtained sequences from Oecomys auyantepui samples UFPAM 2027 (Field number LTO05) and MPEG 40,457 (field number CN285). We used partial nucleotide sequences of the mitochondrial genes Cytochrome b (Cytb; 801 base pairs) and Cytochrome C Oxidase Subunit I (COI; 657 base pairs), and sequence data from nuclear beta-fibrinogen intron 7 (FGB-I7; 727 base pairs). We followed the protocols described in49 for the extraction, amplification, and sequencing of Cytb and FGB-I7 genes; we also followed the same protocols for the COI gene, with the primers Fish F2 (TCGACTAATCATAAAGATATCGGCAC) and Fish R2 (ACTTCAGGGTGACCGAAGAATCAGAA)84. The sequences obtained in the present study are available on GenBank under accession numbers OM927735, OM927739, OM927737 (CN285); OM927736, OM927740, OM927738 (LTO05) (see Supplementary Table 1).
We also used the sequences available on Genbank (http://www.ncbi.nlm.nih.gov/genbank) from the three genes in order to include all COI sequences available in this database for O. auyantepui (48 sequences) plus representative COI sequences of Oecomys species (14 sequences), totaling 64 sequences in our COI analysis; all Cytb sequences available for O. auyantepui (nine sequences) plus representative Cytb sequences of each Oecomys species/clade previously recognized48,49,51 (43 sequences), totaling 52 sequences in our Cytb analysis; and all FGB-I7 sequences available for O. auyantepui (two sequences) plus representative FGB-I7 sequences of each Oecomys species/clade recognized48,51) (15 sequences), totaling 19 sequences in our concatenated data analysis (Cytb + FGB-I7) (Supplementary Table 1). Euryoryzomys nitidus, Hylaeamys megacephalus and Oligoryzomys utiaritensis were used as outgroup for Cytb and concatenated analyses, and one sequence of H. megacephalus for COI analysis (Supplementary Table 1).
The alignment and editing of the Cytb, COI and concatenated (Cytb + FGB-I7) sequences were conducted using the BioEdit Sequence Alignment Editor program, version 7.0.5.385 with ClustalW tool. A search for the best nucleotide substitution model was made using jModeltest 2.1.6 software86 on CIPRES platform87, which selected TIM2 + I + G for Cytb, GTR + I + G for COI, and TIM2 + I + G and TPM2uf + G for the concatenated Cytb and FGB-I7, respectively.
The phylogenetic reconstructions were made using Bayesian Inference (BI) and Maximum Likelihood (ML) methods. BI run in MrBayes 3.2.788 with four chains. The algorithm was based on 50 million generations, sampled every 1,000 generations. ML reconstruction was obtained using Garli–2.089 with 1,000 bootstrap replicates and majority consensus tree. The tree was displayed and edited in Figtree v. 1.4.390. Branches supports were evaluated on Bayesian posterior probability in BI and by bootstrap in ML. Cytb genetic divergence values (p-distances) for O. auyantepui clades (obtained on Cytb analysis) were calculated with MEGA791.
Map
The map was made using QGIS v. 3.10.7. The shapefiles containing geographic data (elevation, hydrography, and country limits) were obtained from DIVA-GIS92, in the link https://www.diva-gis.org/gdata. Geographic limits of Oecomys auyantepui are based on sample points provided in the present study (localities 1–3), by Lira53 (locality 4), Gomes Junior et al.52 (locality 5), Patton et al.39 (localities 6–14), and from GenBank (localities 15–35). More detailed information is available in Supplementary Table 1.
Data availability
The datasets generated and/or analysed during the current study are available in the GenBank repository (https://www.ncbi.nlm.nih.gov/genbank/). All accession numbers supporting the results reported in this article are found in the main text and in the supplementary files.
Abbreviations
- mtDNA:
-
Mitochondrial DNA
- nuDNA:
-
Nuclear DNA
- Cytb:
-
Cytochrome b
- COI:
-
Cytochrome C Oxidase Subunit I
- FGB-I7:
-
Beta-fibrinogen intron 7
- BI:
-
Bayesian Inference
- ML:
-
Maximum Likelihood
- FISH:
-
Fluorescence In Situ Hybridization
- ITS:
-
Interstitial telomeric sequence
- 2n:
-
Diploid number
- FNa:
-
Autosomal fundamental number
- CH:
-
Constitutive heterochromatin
- het-ITS:
-
Heterochromatic-ITS
- APRT:
-
Adenosine phosphoribosyltransferase
- HSA:
-
Human whole chromosome probes
- MMU:
-
Mus musculus
- HME:
-
Hylaeamys megacephalus
- OAU:
-
Oecomys auyantepui
- OCA-PA:
-
Oecomys catherinae From Pará
- OCA-RJ:
-
Oecomys catherinae From Rio de Janeiro
- OPA-A:
-
Oecomys paricola Cytotype A
- OPA-B:
-
Oecomys paricola Cytotype B
- OPA-C:
-
Oecomys paricola Cytotype C
- CLA:
-
Cerradomys langguthi
- NVO:
-
Neacomys vossi
- NEL:
-
Neacomys elieceri
- NXI:
-
Neacomys xingu
- NMA:
-
Neacomys marajoara
- NPA:
-
Neacomys paracou
- NSP-E:
-
Neacomys Sp. E
- NAM:
-
Neacomys amoenus
- TNI:
-
Thaptomys nigrita
- AMO:
-
Akodon montensis
- ASP:
-
Akodon Sp.
- NLA:
-
Necromys lasiurus
- OAM:
-
Oxymycterus amazonicus
- BBR:
-
Blarinomys breviceps
References
GL Bush SM Case AC Wilson JL Patton 1977 Rapid speciation and chromosomal evolution in mammals Proc. Natl. Acad. Sci. 74 9 3942 3946 https://doi.org/10.1073/pnas.74.9.3942
M King 1993 Species Evolution Cambridge University Press
LH Rieseberg 2001 Chromosomal rearrangements and speciation Trends Ecol. Evol. 16 7 351 358 https://doi.org/10.1016/s0169-5347(01)02187-5
R Castiglia 2013 Sympatric sister species in rodents are more chromosomally differentiated than allopatric ones: Implications for the role of chromosomal rearrangements in speciation Mammal Rev. 44 1 4 https://doi.org/10.1111/mam.12009
P Franchini 2020 Reconstructing the evolutionary history of chromosomal races on islands: A Genome-Wide Analysis of Natural House Mouse Populations Mol. Biol. Evol. 37 10 2825 2837 https://doi.org/10.1093/molbev/msaa118
JAM Graves 2016 Did sex chromosome turnover promote divergence of the major mammal groups? De novo sex chromosomes and drastic rearrangements may have posed reproductive barriers between monotremes, marsupials and placental mammals BioEssays 38 8 734 743 https://doi.org/10.1002/bies.201600019
SA Romanenko 2020 Evolutionary rearrangements of X chromosomes in voles (Arvicolinae, Rodentia) Sci. Rep. 10 1 1 11 https://doi.org/10.1038/s41598-020-70226-4
MJD White 1978 Modes of Speciation W.H. Freeman & Co
JW Sites C Moritz 1987 Chromosomal evolution and speciation revisited Syst. Zool. 36 2 153 174 https://doi.org/10.2307/2413266
JA Coyne HA Orr 1998 The evolutionary genetics of speciation Philos. Trans. R. Soc. Lond. B. 353 287 305 https://doi.org/10.1098/rstb.1998.0210
JB Searle 1998 Speciation, chromosomes, and genomes Genome Res. 8 1 1 3 https://doi.org/10.1101/gr.8.1.1
SA Romanenko V Volobouev 2012 Non-Sciuromorph rodent karyotypes in evolution Cytogenet. Genome Res. 137 233 245 https://doi.org/10.1159/000339294
MJD White 1973 Animal Cytology and Evolution 3 Cambridge University
B Wang Y Xia J Song W Wang Y Tang 2013 Case report: Potential speciation in humans involving robertsonian translocations Biom. Res. 24 1 171 174
T Ashley 2002 X-Autosome translocations, meiotic synapsis, chromosome evolution and speciation Cytogenet. Genome Res. 96 33 39 https://doi.org/10.1159/000063030
G Dobigny C Ozouf-Costaz C Bonillo V Volobouev 2004 Viability of X-autosome translocations in mammals: An epigenomic hypothesis from a rodent case-study Chromosoma 113 34 41 https://doi.org/10.1007/s00412-004-0292-6
F Veyrunes J Perez 2018 X inactivation in a mammal species with three sex chromosomes Chromosoma 127 261 267 https://doi.org/10.1007/s00412-017-0657-2
JAM Graves 2006 Sex chromosome specialization and degeneration in mammals Cell 124 901 914 https://doi.org/10.1016/j.cell.2006.02.024
E Lifschytz DL Lindsey 1972 The role of X-chromosome inactivation during spermatogenesis PNAS 69 1 182 186 https://doi.org/10.1073/pnas.69.1.182
AJ Sharp HT Spotswood DO Robinson BM Turner PA Jacobs 2002 Molecular and cytogenetic analysis of the spreading of X inactivation in X;autosome translocations Hum. Mol. Genet. 1 25 3145 3156 https://doi.org/10.1093/hmg/11.25.3145
AG Searle CV Beechey EP Evans M Kirk 1983 Two new X-autosome translocations in the mouse Cytogenet. Cell Genet. 35 279 292 https://doi.org/10.1159/000131880
JW Kim 2012 Molecular and clinical characteristics of 26 cases with structural Y chromosome aberrations Cytogenet. Genome Res. 136 270 277 https://doi.org/10.1159/000338413
L Centofante LAC Bertollo O Moreira-Filho 2006 Cytogenetic characterization and description of an XX/XY1Y2 sex chromosome system in catfish Harttia carvalhoi (Siluriformes, Loricariidae) Cytogenet. Genome Res. 112 320 324 https://doi.org/10.1159/000089887
EA Oliveira 2018 Tracking the evolutionary pathway of sex chromosome among fishes: Characterizing the unique XX/XY1Y2 system in Hoplias malabaricus (Teleostei, Characiformes) Chromosoma 127 115 128 https://doi.org/10.1007/s00412-017-0648-3
RCR Noronha 2020 Meiotic analyses show adaptations to maintenance of fertility in X1Y1X2Y2X3Y3X4Y4X5Y5 system of amazon frog Leptodactylus pentadactylus (Laurenti, 1768) Sci. Rep. 10 16327 https://doi.org/10.1038/s41598-020-72867-x
F Veyrunes J Catalan B Sicard TJ Robinson JM Duplantier L Granjon 2004 Autosome and sex chromosome diversity among the African pygmy mice, subgenus Nannomys (Murinae; Mus) Chromosome Res. 12 4 369 382 https://doi.org/10.1023/B:CHRO.0000034098.09885.e6
CI Aquino VV Abril JMB Duarte 2013 Meiotic pairing of B chromosomes, multiple sexual system, and Robertsonian fusion in the red brocket deer Mazama americana (Mammalia, Cervidae) Genet. Mol. Res. 12 3 3566 3574 https://doi.org/10.4238/2013.September.13.1
S Kasahara B Dutrillaux 1983 Chromosome banding patterns of four species of bats, with special reference to a case of X-autosome translocation Ann. Genet. 26 197 201
AJ Solari MI Pigozzi 1994 Fine structure of the XY body in the XY1Y2 trivalent of the bat Artibeus lituratus Chromosome Res. 2 53 58
RCR Noronha 2001 Sex-autosome translocations: Meiotic behavior suggests an inactivation block with permanence of autosomal gene activity in Phyllostomid bats Caryologia 54 267 277 https://doi.org/10.1080/00087114.2001.10589235
RCR Noronha 2004 Meiotic analyses of the sex chromosomes in Carolliinae-Phyllostomidae (Chiroptera): NOR separates the XY1Y2 into two independent parts Caryologia 57 1 9 https://doi.org/10.1080/00087114.2004.10589365
DA Parish P Vise HA Whman JJ Bull RJ Baker 2002 Distribution of LINEs and other repetitive elements in the karyotype of the bat Carollia: Implications for X-chromosome inactivation Cytogenet. Genome Res. 96 191 197 https://doi.org/10.1159/000063038
PJS Amaral 2013 Proechimys (Rodentia, Echimyidae): Characterization and taxonomic considerations of a form with a very low diploid number and a multiple sex chromosome system BMC Genet. 14 21 27 https://doi.org/10.1186/1471-2156-14-21
MJ Rodrigues da Costa 2016 Cryptic species in Proechimys goeldii (Rodentia, Echimyidae)? A case of molecular and chromosomal differentiation in allopatric populations Cytogenet. Genome Res. 148 199 210 https://doi.org/10.1159/000446562
W Oliveira Da Silva 2019 Identification of two independent X-autosome translocations in closely related mammalian (Proechimys) species Sci. Rep. 9 4047 https://doi.org/10.1038/s41598-019-40593-8
R Toder 1997 Comparative chromosome painting between two marsupials: Origins of an XX/XY1Y2 sex chromosome system Mamm. Genome. 8 6 418 422 https://doi.org/10.1007/s003359900459
M Vassart A Séguéla H Hayes 1995 Chromosomal evolution in gazelles J. Hered. 86 3 216 227 https://doi.org/10.1093/oxfordjournals.jhered.a111565
L Dias de Oliveira 2019 First cytogenetic information for Lonchothrix emiliae and taxonomic implications for the genus taxa Lonchothrix + Mesomys (Rodentia, Echimyidae, Eumysopinae) PLoS ONE 14 4 e0215239 https://doi.org/10.1371/journal.pone.0215239
JL Patton UFJ Pardiñas G D’Elía 2015 Mammals of South America: Rodents 2 University of Chicago Press
UFJ Pardiñas P Teta J Salazar-Bravo 2015 A new tribe of sigmodontinae rodents (Cricetidae) Mastoz Neot. 22 1 171 186
PR Gonçalves 2020 Unraveling deep branches of the sigmodontinae tree (Rodentia: Cricetidae) in Eastern South America J. Mamm. Evol. 27 139 160 https://doi.org/10.1007/s10914-018-9444-y
K Ventura V Fagundes G D’Elía AU Christoff Y Yonenaga-Yassuda 2011 A new allopatric lineage of the rodent Deltamys (Rodentia: Sigmodontinae) and the chromosomal evolution in Deltamys kempi and Deltamys sp. Cytogenet. Genome Res. 135 126 134 https://doi.org/10.1159/000331584
C Lanzone 2011 XY1Y2 chromosome system in Salinomys delicatus (Rodentia, Cricetidae) Genetica 139 1143 1147 https://doi.org/10.1007/s10709-011-9616-7
MO Ortells AO Reig N Brum-Zorrilla AO Scaglia 1988 Cytogenetics and karyosystematics of phyllotine rodents (Cricetidae, Sigmodontinae) I. Chromosome multiformity and gonosomai-autosomai transiocation in Reithrodon Genetica 77 1 53 63 https://doi.org/10.1007/BF00058549
CC Rosa 2012 Genetic and morphological variability in South American rodent Oecomys (Sigmodontinae, Rodentia): Evidence for a complex of species J. Genet. 91 265 277 https://doi.org/10.1007/s12041-012-0182-2
UFJ Pardiñas P Teta J Salazar-Bravo P Myers CA Galliari 2016 A new species of arboreal rat, genus Oecomys (Rodentia, Cricetidae) from Chaco J. Mamm. 97 1177 1196 https://doi.org/10.1093/jmammal/gyw070
SM Malcher 2017 Oecomys catherinae (Sigmodontinae, Cricetidae): Evidence for chromosomal speciation? PLoS ONE 12 7 e0181434 https://doi.org/10.1371/journal.pone.0181434
EY Suárez-Villota AP Carmignotto MV Brandão AR Percequillo MJDJ Silva 2018 Systematics of the genus Oecomys (Sigmodontinae: Oryzomyini): Molecular phylogenetic, cytogenetic and morphological approaches reveal cryptic species Zool. J. Linn. Soc. 184 1 182 210 https://doi.org/10.1093/zoolinnean/zlx095
J Saldanha 2019 Genetic diversity of Oecomys (Rodentia, Sigmodontinae) from the Tapajós River basin and the role of rivers as barriers for the genus in the region Mamm. Biol. 97 41 49 https://doi.org/10.1016/j.mambio.2019.04.009
W Oliveira Da Silva 2020 Karyotypic divergence reveals that diversity in the Oecomys paricola complex (Rodentia, Sigmodontinae) from eastern Amazonia is higher than previously thought PLoS ONE 15 10 e0241495 https://doi.org/10.1371/journal.pone.0241495
J Saldanha RV Rossi 2021 Integrative analysis supports a new species of the Oecomys catherinae complex (Rodentia, Cricetidae) from Amazonia J. Mamm. 102 69 89 https://doi.org/10.1093/jmammal/gyaa145
RG Gomes Júnior 2016 Intense genomic reorganization in the genus Oecomys (Rodentia, Sigmodontinae): Comparison between DNA barcoding and mapping of repetitive elements in three species of the Brazilian Amazon Comp. Cytogenet. 10 3 401 426 https://doi.org/10.3897/CompCytogen.v10i3.8306
Lira, T. Citogenética clássica e molecular de alguns representantes da tribo Oryzomyini (Rodentia, Cricetidae) da Amazônia Central. Ph.D dissertation, Universidade Federal do Amazonas. Manaus, Amazonas, Brazil (2012).
CY Nagamachi 2013 FISH with whole chromosome and telomeric probes demonstrates huge karyotypic reorganization with ITS between two species of Oryzomyini (Sigmodontinae, Rodentia): Hylaeamys megacephalus probes on Cerradomys langguthi karyotype Chromosome Res. 21 107 119 https://doi.org/10.1007/s10577-013-9341-4
P Suárez 2015 Clues on syntenic relationship among some species of Oryzomyini and Akodontini Tribes (Rodentia: Sigmodontinae) PLoS ONE 10 12 e0143482https://doi.org/10.1371/journal.pone.0143482
AL Pereira 2016 Extensive chromosomal reorganization among species of New World muroid rodents (Cricetidae, Sigmodontinae): Searching for phylogenetic ancestral traits PLoS ONE 11 1 e0146179https://doi.org/10.1371/journal.pone.0146179
W Oliveira Da Silva 2017 Chromosomal diversity and molecular divergence among three undescribed species of Neacomys (Rodentia, Sigmodontinae) separated by Amazonian rivers PLoS ONE 12 8 e0182218 https://doi.org/10.1371/journal.pone.0182218
W Oliveira Da Silva 2019 Chromosomal phylogeny and comparative chromosome painting among Neacomys species (Rodentia, Sigmodontinae) from eastern Amazonia BMC Evol. Biol. 19 184 https://doi.org/10.1186/s12862-019-1515-z
W Oliveira Da Silva 2020 Chromosomal signatures corroborate the phylogenetic relationships within Akodontini (Rodentia, Sigmodontinae) Int. J. Mol. Sci. 21 2415 https://doi.org/10.3390/ijms21072415
JL Patton MN Silva JR Malcolm 2000 Mammals of the Rio Juruá and the Evolutionary and ecological diversification of Amazonia B Am. Mus. Nat. Hist. 244 202 292
A Langguth V Maia M Mattevi 2005 Karyology of large size Brazilian species of the genus Oecomys Thomas, 1906 (Rodentia, Muridae, Sigmodontinae) Arq. Mus. Nac. Rio de Janeiro 63 183 190
SA Romanenko PL Perelman VA Trifonov AS Graphodatsky 2012 Chromosomal evolution in Rodentia Heredity 108 4 16 https://doi.org/10.1038/hdy.2011.110
MI Rahn 2016 Protein markers of synaptic behavior and chromatin remodeling of the neo-XY body in phyllostomid bats Chromosoma 125 4 701 708 https://doi.org/10.1007/s00412-015-0566-1
AY Aksenova SM Mirkin 2019 At the beginning of the end and in the middle of the beginning: Structure and maintenance of telomeric DNA repeats and interstitial telomeric sequences Genes 10 2 118 https://doi.org/10.3390/genes10020118
AE Kilburn MJ Shea RG Sargent JH Wilson 2001 Insertion of a telomere repeat sequence into a mammalian gene causes chromosome instability Mol. Cell Biol. 21 1 126 135 https://doi.org/10.1128/MCB.21.1.126-135.2001
A Ruiz-Herrera SG Nergadze M Santagostino E Giulotto 2008 Telomeric repeats far from the ends: Mechanisms of origin and role in evolution Cytogenet. Genome Res. 122 219 228 https://doi.org/10.1159/000167807
J Meyne 1990 Distribution of non-telomeric sites of the (TTAGGG)n telomeric sequence in vertebrate chromosomes Chromosoma 99 3 10 https://doi.org/10.1007/BF01737283
AJB Gomes 2016 Chromosomal phylogeny of Vampyressine bats (Chiroptera, Phyllostomidae) with description of two new sex chromosome systems BMC Evol. Biol. 16 119 https://doi.org/10.1186/s12862-016-0689-x
AV Borisenko BK Lim NV Ivanova RH Hanner PD Hebert 2008 DNA barcoding in surveys of small mammal communities: A field study in Suriname Mol. Ecol. Resour. 8 3 471 479 https://doi.org/10.1111/j.1471-8286.2007.01998.x
J Kitano 2009 A role for a neo-sex chromosome in stickleback speciation Nature 461 1079 1083 https://doi.org/10.1038/nature08441
S Ginot J Claude J Perez F Veyrunes 2017 Sex reversal induces size and performance differences among females of the African pygmy mouse, Mus minutoides J. Exp. Biol. 220 1947 1951 https://doi.org/10.1242/jeb.157552
RG Rocha 2014 Seasonal flooding regime and ecological traits influence genetic structure of two small rodents Ecol. Evol. 4 4598 4608 https://doi.org/10.1002/ece3.1336
J Gilmour J Gregor 1939 Demes: A suggested new terminology Nature 144 333 https://doi.org/10.1038/144333a0
JL Patton SW Sherwood 1983 Chromosome evolution and speciation in rodents Ann. Rev. Ecol. Syst. 14 139 158 https://doi.org/10.1146/annurev.es.14.110183.001035
EHC Oliveira de 2002 The phylogeny of howler monkeys (Alouatta, Platyrrhini): Reconstruction by multicolor cross-species chromosome painting Chromosome Res. 10 669 683 https://doi.org/10.1023/A:1021520529952
NP Araújo R Stanyon VS Pereira M Svartman 2019 Interspecific chromosome painting provides clues to the ancestral karyotype of the New World monkey genus Aotus J. Mamm. Evol. 26 283 290 https://doi.org/10.1007/s10914-017-9403-z
JC Pieczarka 2013 A phylogenetic analysis using multidirectional chromosome painting of three species (Uroderma magnirostrum, U. bilobatum and Artibeus obscurus) of subfamily Stenodermatinae (Chiroptera-Phyllostomidae) Chromosome Res. 21 4 383 392 https://doi.org/10.1007/s10577-013-9365-9
PS Corn 1994 Straight-line drift fences and pitfall traps WR Heyer MA Donnelly RW McDiarmid LC Hayek MS Foster Eds Measuring and Monitoring Biological Standard Methods for Amphibians Smithsonian Institution Press 109 117
CE Ford JL Hamerton 1956 A colchicine, hypotonic-citrate, squash sequence for mammalian chromosomes Stain Technol. 31 247 251 https://doi.org/10.3109/10520295609113814
AT Sumner 1972 A simple technique for demonstrating centromeric heterochromatin Exp. Cell Res. 75 304 306 https://doi.org/10.1016/0014-4827(72)90558-7
AT Sumner HJ Evans RA Buckland 1971 New technique for distinguishing between human chromosomes Nat. Lond. New Biol. 31 282 https://doi.org/10.1038/newbio232031a0
F Yang NP Carter L Shi MA Ferguson-Smith 1995 A comparative study of karyotypes of muntjacs by chromosome painting Chromosoma 103 642 652 https://doi.org/10.1007/BF00357691
A Levan K Fredga AA Sandberg 1964 Nomenclature for centromeric position on chromosomes Hereditas 52 201 220
RD Ward TS Zemlak BH Innes PR Last PDN Hebert 2005 DNA barcoding Australia’s fish species Philos. Trans. R. Soc. Lond. B Biol. Sci. 360 1847 1857 https://doi.org/10.1098/rstb.2005.1716
TA Hall 1999 BioEdit: A user-friendly biological sequence alignment editor and analysis program for Windows 85/98/NT Nucleic Acids Symp. Ser. 41 95 98
D Darriba GL Taboada R Doallo D Posada 2012 jModelTest 2: More models, new heuristics and parallel computing Nat. Methods. 9 8 772 https://doi.org/10.1038/nmeth.2109
Miller, M. A., Pfeiffer, W. & Schwartz, T. Creating the CIPRES Science Gateway for inference of large phylogenetic trees. Proceedings of the Gateway Computing Environments Workshop (GCE) IEE, New Orleans, LA, 1–8 (2010). https://doi.org/10.1109/GCE.2010.5676129
F Ronquist 2012 MrBayes 3.2: Efficient Bayesian phylogenetic inference and model choice across a large model space Syst. Biol. 61 3 539 542 https://doi.org/10.1093/sysbio/sys029
Zwickl, D. J. Genetic algorithm approaches for the phylogenetic analysis of large biological sequence datasets under the maximum likelihood criterion. Ph.D. dissertation, The University of Texas at Austin. Austin, Texas (2016).
Rambaut, A. FigTree v1.4.3 2006–2016. http://tree.bio.ed.ac.uk/software/figtree/ (2016).
S Kumar G Stecher K Tamura 2016 MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets Mol. Biol. Evol. 33 7 1870 1874 https://doi.org/10.1093/molbev/msw054
Hijmans, R. J. et al. DIVA-GIS. Vsn 5.0. A geographic information system for the analysis of species distribution data. http://www.diva-gis.org (2004).
Acknowledgements
The authors are grateful to MSc. Jorge Rissino, to MSc Shirley Nascimento and Maria da Conceição for assistance in laboratory work. The authors thank Instituto Chico Mendes de Conservação da Biodiversidade (ICMBio) and Secretaria de Estado de Meio Ambiente do Pará (SEMA-PA) for the authorization of sample collections. This study is part of the Thesis of CCR in Neuroscience and Biology Cellular, under a CAPES Scholarship. CYN (305880/2017-9) and JCP (305876/2017-1) are grateful to CNPq for Productivity Grants. WOS is grateful for PNPD/CAPES Scholarship (88882.315778/2019-01).
Funding
This research was supported by Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq), the Fundação Amazônia Paraense de Amparo à Pesquisa (FAPESPA) and the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES) on projects coordinated by CY Nagamachi (Edital BIONORTE-CNPq, Proc 552032/2010-7; Edital BIONORTE-FAPESPA, ICAAF 007/2011; Edital Pró-Amazônia Proc 047/2012); the FAPESPA (Edital Vale – Proc 2010/110447) and Banco Nacional de Desenvolvimento Econômico e Social – BNDES (Operação 2.318.697.0001) on a project coordinated by JC Pieczarka; CNPq for Productivity Grants to CYN (305880/2017-9) and to JCP (305876/2017-1); PNPD/CAPES Scholarship (88882.315778/2019-01) to WOS. The article processing charge was paid by PROPESP-UFPA (Edital 06/2021 – PAPQ/PROPESP).
Author information
Authors and Affiliations
Contributions
W.O.S. worked on chromosome painting and molecular analysis, made the conceptualization, the data curation, the formal analysis and wrote the original draft. C.C.R. and J.S. developed laboratory techniques, revised the manuscript and edited it. M.A.F.-S., P.C.M.O’.B., R.V.R., J.C.P. and C.Y.N. contribute with resources. W.O.S., J.C.P. and C.Y.N. obtained funding for this research. P.C.M.O’.B. and M.A.F.-S. also made the formal analysis of all data, revised the manuscript and edited it. R.V.R. and J.S. worked on the molecular analysis and its interpretation. J.C.P. worked on chromosome painting and formal analysis of all data. C.Y.N. worked on data curation, project administration and supervision of W.O.S. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Oliveira da Silva, W., Rosa, C.C., Ferguson-Smith, M.A. et al. The emergence of a new sex-system (XX/XY1Y2) suggests a species complex in the “monotypic” rodent Oecomys auyantepui (Rodentia, Sigmodontinae). Sci Rep 12, 8690 (2022). https://doi.org/10.1038/s41598-022-12706-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-12706-3
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.