Abstract
Fast and accurate identification of phage virion proteins (PVPs) would greatly aid facilitation of antibacterial drug discovery and development. Although, several research efforts based on machine learning (ML) methods have been made for in silico identification of PVPs, these methods have certain limitations. Therefore, in this study, we propose a new computational approach, termed SCORPION, (StaCking-based Predictior fOR Phage VIrion PrOteiNs), to accurately identify PVPs using only protein primary sequences. Specifically, we explored comprehensive 13 different feature descriptors from different aspects (i.e., compositional information, composition-transition-distribution information, position-specific information and physicochemical properties) with 10 popular ML algorithms to construct a pool of optimal baseline models. These optimal baseline models were then used to generate probabilistic features (PFs) and considered as a new feature vector. Finally, we utilized a two-step feature selection strategy to determine the optimal PF feature vector and used this feature vector to develop a stacked model (SCORPION). Both tenfold cross-validation and independent test results indicate that SCORPION achieves superior predictive performance than its constitute baseline models and existing methods. We anticipate SCORPION will serve as a useful tool for the cost-effective and large-scale screening of new PVPs. The source codes and datasets for this work are available for downloading in the GitHub repository (https://github.com/saeed344/SCORPION).
Similar content being viewed by others
Introduction
Bacteriophages are viruses that can infect and thrive in bacteria. It can be found in several environments including soil, freshwater and marine. The infectious phage particle is essentially comprised of a nucleic acid component (i.e. either DNA or RNA) in which they are encapsulated in a coat of protein known as capsids1. Individual types of bacteriophage can display an extremely high specificity towards a particular susceptible bacterial host species. The surface of which they will typically attach themselves irreversibly to and inject their genetic materials to the cellular interior. They are able to persist in the host by using one of two major strategies that are termed lytic and lysogenic life cycle2. Bacteriophages may represent a promising alternative to antiobiotics owing to the following properties: a lack of toxicity toward human cells, lack of harm caused to normal flora and ability to target antibiotic-resistant bacteria3. Phage structural proteins (PVPs) consists of capsid proteins, tail proteins and phage particle enzymes. PVPs are mainly responsible for orchestrating bacteriophage interaction with their specific bacterial hosts so their manipulation may represent an avenue to generate novel classes of antimicrobial agents4. Current experimental approaches for the identification of PVPs from non-PVPs include many scientific instruments and methodologies such as mass spectrometry, sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-GE) based proteomic methods and protein analysis arrays5,6,7. While these methods represent gold standard approaches for PVP identification, they are difficult to employ for the analysis of PVPs at large scale as they are laborious and costly methods. Thus, researchers have invested much in efforts to develop computational models for predicting PVPs directly from their sequence information as a useful alternative.
To date, a variety of machine learning (ML)-based methods, including iVIREONS8, Feng et al.’s method9, PVPred10, PVP-SVM11, PhagePred12, Tan et al.’s method13, Ru et al.’s method14, Pred-BVP-Unb15 and PVPred-SCM16, Zhang et al.’s method17, Meta-iPVP18, iPVP-MCV19 and VirionFinder20 have all been developed and proposed for PVP identification. Table 1 provides a summary of these machine learning-based methods along with their employed ML algorithms, feature descriptors and evaluation strategies. In 2013, Seguritan et al. developed the first PVP predictor called iVIREONS8 based on ANN algorithm trained with AAC and PIP to predict viral structural proteins. Shortly afterward, Feng et al. created a high-quality dataset consisting of 99 PVPs and 208 non-PVPs, and also developed a NB-based predictor9 cooperating with AAC and DPC. Most recently, Han et al. developed an ensemble-based model named iPVP-MCV19 by combing three types of PSSM descriptors (i.e. PSSM-AAC, PSSM-composition and DP-PSSM). Until now, iPVP-MCV have represented a state-of-the-art predictor for PVP identification. More detail information for all of the existing methods is summarized in an article by Kabir et al.21. Although above mentioned methods do efficiently facilitate the prediction of PVPs, there are some issues that still need to be addressed. First, the training dataset used by several existing methods in PVP identification was relatively small. This is an important consideration, as several previous studies have demonstrated that training with a large number of datasets is crucial for building a comprehensive predictive model18,22,23,24. Second, almost all of the existing methods were developed by employing single ML methods to train the model. Therefore, their performance might not be optimal in some cases. However, ensemble models are capable to provide a greatly improved performance compared to baseline models22,24,25,26,27. Finally, the prediction performance for these existing methods is still not satisfactory for many real therapeutic applications.
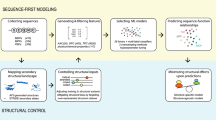
To address these limitations, we present a novel approach, termed SCORPION (StaCking-based Predictior fOR Phage VIrion PrOteiNs) to improve the accurate prediction of PVPs. The overall procedure for the development of SCORPION is illustrated in Fig. 1. Notably, SCORPION employs 13 different sequence-based feature descriptors from multiple perspectives (i.e., compositional information, composition–transition–distribution information, position-specific information and physicochemical properties) to extract the key pattern of PVPs. These feature descriptors were used to train a total of 130 baseline models by using 10 popular ML algorithms. Probabilistic features (PFs) were then generated by using these 130 baseline models, and considered as a new feature vector. To improve the predictive performance, a two-step feature selection strategy was applied to identify m out of 130 PFs. Finally, the optimal PF feature vector were used to develop an effective stacked model (SCORPION) by using the stacked ensemble learning strategy. Our comparative results base on cross-validation and independent tests indicate that SCORPION outperformed its baseline models. Moreover, SCORPION achieved a better performance than several existing methods for PVP prediction in terms of in terms of ACC (0.873), Sp (0.905), MCC (0.748) and AUC (0.891) on the independent dataset. These comparative results highlight the effectiveness and generalizability of SCORPION.
Materials and methods
Overall framework of SCORPION
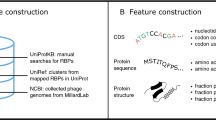
As can be seen in Fig. 1, there exist four major steps, including dataset construction, baseline models construction, new feature representations and the stacked model development. First, The same benchmark dataset derived from Charoenkwan et al.18 were used to train and optimized baseline models and SCORPION. Second, 13 different feature descriptors were individually fed to 10 different ML algorithms to build the 130 baseline models using tenfold cross-validation. In addition, we comprehensively compared 13 different feature descriptors to determine the feature descriptors that are beneficial to PVP identification. Third, we constructed variant stacked models by using different sets of feature vectors. Forth, the optimal PF vector was determined and fed to RF algorithm in order to construct the final stacked model (SCORPION) by using the stacked ensemble learning strategy. Finally, we compared the predictive performance of SCORPION against its constitute baseline models and existing methods.
Dataset collection
As described in an article by Kabir et al.21, there are three well-known benchmark datasets (i.e. Feng20139, Manavalan201811 and Charoenkwan2020_2.018) that have been established for developing existing PVP predictors. In this study, we utilized the Charoenkwan2020_2.0 dataset established by Charoenkwan et al.18 as the benchmark dataset to assess the performance of SCORPION. Below, we provided two main reasons why we used the Charoenkwan2020_2.0 dataset. First, the Charoenkwan2020_2.0 dataset contained a larger number of PVPs and non-PVPs than other datasets. Specifically, the Charoenkwan2020_2.0 dataset combined Feng20139 and Manavalan201811 datasets along with novel PVPs and non-PVPs obtained from the UniProt database (release 2019_11)28. Second, a lower CD-HIT threshold of 0.4 was used to exclude more redundant sequences in the Charoenkwan2020_2.0 dataset. As a result, the Charoenkwan2020_2.0 dataset contained of 313 PVPs and 313 non-PVPs. In the Charoenkwan2020_2.0 dataset, the training and independent datasets (PVPs, non-PVPs) consisted of (250, 250) and (63, 63), respectively. All datasets used in this study are available on https://github.com/saeed344/SCORPION.
Feature encodings
In this study, we used 13 different sequence-based feature descriptors containing amino acid composition (AAC), amino acid index (AAI), amphiphilic pseudo-amino acid composition (APAAC), composition in CTD (CTDC), distribution part of CTD (CTDD), transition in CTD (CTDT), dipeptide deviation from expected mean (DDE), dipeptide composition (DPC), enhance Amino Acid composition (EAAC), pseudo amino acid composition (PAAC), PSSM_AAC, PSSM_DP and PSSM_COM to extract the key information of PVPs and non-PVPs. These sequence-based feature descriptors provides us four different aspects consisting of compositional information, composition-transition-distribution information, position-specific information and physicochemical properties having sufficient information to develop a comprehensive predictive model. Details of all 13 feature descriptors are provided in Table 2. Here, the iFeature Python package29 was utilized to calculate all the 13 feature descriptors.
Stacking ensemble learning framework of SCORPION
In this study, the stacked ensemble learning strategy was utilized to develop SCORPION for improving the prediction of PVPs. Unlike other ensemble learning strategies, this strategy enables an automatic integration of different ML classifiers in order to construct a single robust prediction model23. The stacked strategy has successfully achieve better performance as compared with its constituent baseline models23,24,27,30,31. The stacking strategy consists of two main steps, while the corresponding models at each step are referred to as baseline and meta models, respectively.
In the first step, the PVPs and non-PVPs in the training dataset were extracted by using 13 different feature encoding schemes from four different perspectives containing AAC, AAI, APAAC, CTDC, CTDD, CTDT, DDE, DPC, EAAC, PAAC, PSSM_AAC, PSSM_DP and PSSM_COM with corresponding dimensions of 20, 11, 22, 39, 39, 195, 400, 400, 20, 21, 20, 400 and 400, respectively32,33,34,35. Herein, we used the default iFeature parameter settings29 to generate APAAC and PAAC descriptors. Then, each feature descriptor was individually employ to train 10 different ML algorithms (KNN, RF, SVM, decision tree (DT), extremely randomized trees (ET), logistic regression (LR), multi-layer perceptron (MLP), naive Bayes (NB), partial least squares regression (PLS) and extreme gradient boosting (XGB)). To enhance the predictive performance, all ML classifiers were trained and optimized using the scikit-learn package in Python (version 0.22)36. Specifically, the optimal parameters of ET, LR, MLP, RF, SVM and XGB classifiers were carefully determined under the tenfold cross-validation procedure on the training dataset, where the search range is shown in Supplementary Table S1. In the case of the remaining ML classifiers, they were constructed by using their default parameters. Therefore, we obtained a total of 130 baseline models (10 MLs × 13 encodings).
In the second step, each baseline model provided us three types of features from three perspectives containing PF, class feature (CF) and the combination of PF and CF (PCF). The PF is based on the predicted probability scores to be PVPs which is in the range of 0–1. In case of the CF, the protein sequence P is labeled as 1 if its predicted probability scores is greater than 0.5, otherwise the protein sequence P is labeled as 0. As a result, the protein sequence P was represented to 130-D, 130-D and 260-D feature vectors for PF, CF and PCF, respectively. The PF, CF and PCF were considered as new feature vectors. RF algorithm was employed as the meta model (called mRF) to train the stacked model. As result, we obtained three different stacked models based on three new feature vectors (i.e. PF, CF and PCF). To improve the discriminative ability of the new feature vectors, we used a two-step feature selection strategy to optimize PF, CF and PCF feature vectors. At the first step, we used XGB classifier to rank the features in PF, CF and PCF. The XGB classifier is widely used in the feature importance analysis23,37. Using the XGB classifier, we constructed a ranking list of features with respect to their importance scores. Higher ranked features in this list are the most important features. At the second step, we constructed n different feature subsets containing the top ranked features ranging from top 5 to top 100 features with an interval of 5. Then, we inputted all feature subsets into mRF models and optimized the mRF models’ parameters using tenfold cross-validation scheme. The feature subset achieving the highest Matthews correlation coefficient (MCC) was considered as the optimal feature subset. The implementation of these classifiers in the two-step feature selection strategy is the same as used in our previous studies18,31,38,39,40,41
Performance evaluation strategies
In order to examine the performance of our proposed predictor, we used five common statistical metrics including ACC, MCC, sensitivity (Sn) and specificity (Sp)24,42 as described follows:
where TP, TN, FP and FN represent the number of true positives, true negatives, false positive and false negatives, respectively. In addition, the area under the receiver operating characteristic (AUC) was employed as another statistical metric39,40,41,43.
Results and discussion
Performance evaluation between different classifiers and feature encodings
In this section, we investigated the effect of individual feature descriptor for PVP identification. Specifically, 13 different feature encoding from multiple perspectives (i.e. compositional information (AAC, APAAC, DDE, DPC, EAAC and PAAC), composition-transition-distribution information (CTDC, CTDD and CTDT), position-specific information (PSSM_AAC, PSSM_DP and PSSM_COM) and physicochemical properties (AAI)) were inputted to 10 different ML algorithms (DT, ET, KNN, LR, MLP, NB, PLS, RF, SVM and XGB) for developing 130 baseline models. We evaluated the predictive performance of the 130 baseline models with a default threshold of 0.5 by performing tenfold cross-validation and independent tests on the training and independent datasets, respectively. For convenience of discussion, Fig. 2 shows the performance of the 30 best-performing baseline models in the term of cross-validation MCC. In addition, the performance results for all the 130 baseline models are provided in Supplementary Tables S2 and S3.
From Fig. 2, Supplementary Tables S2 and S3, several observations can be made. First, PSSM_AAC descriptor was the most powerful one for PVP identification with average cross-validation AAC and MCC of 0.802 and 0.610, respectively. In the meanwhile, PSSM_COM and AAC descriptors performed well with the second and third highest average cross-validation MCC of 0.582 and 0.556, respectively. Second, SVM-based and LR-based classifiers outperformed other ML-based classifiers in terms of ACC (0.782–0.784), Sp (0.780–0.788), MCC (0.570–0.576) and AUC (0.849–0.850). Third, among all the 130 baseline models, the baseline model trained with MLP algorithm in conjunction with PSSM_COM descriptor (MLP-PSSM_COM) attained the best performance with cross-validation AAC and MCC of 0.840 and 0.684, while its ACC, MCC and AUC were 0.778, 0.556 and 0.859, respectively, as evaluated by the independent test. Taken together, the single feature-based models were not effective enough for PVP identification. On the other hand, the integration of variant ML classifier for constructing a single meta-predictor might improve the model’s performance.
Performance evaluation of different stacked models
As mentioned in the “Materials and methods” section, we designed and developed three different stacked models based on three types of new feature representations consisting of PF (130D), CF (130D) and PCF (260D). Specifically, these three new feature representations were inputted to RF algorithm for developing three different mRF models. The performance comparison results amongst the three mRF models are provided in Tables 3 and 4. As can be seen, it is clear that PF and PCF feature vectors achieved better performance in terms of all performance metrics based on both tenfold cross-validation and independent tests. To further improve the discriminative ability of our new features, we utilized the two-step feature selection scheme to optimize PF, CF and PCF feature vectors. Herein, the feature selection scheme identified 50, 5 and 5 informative PFs, CFs and PCFs, respectively, for generating three optimal feature sets. Tables 3 and 4 shows that the three optimal feature sets attained a similar performance based on tenfold cross-validation test. In case of the independent test results, optimal PF feature vector outperformed other feature sets in terms of four out of five performance metrics (i.e. ACC, Sp, MCC and AUC). Particularly, ACC, Sp, MCC and AUC of optimal PF feature vector were 0.881, 0.952, 0.770 and 0.922, respectively (Table 4). The optimal PF feature vector consisted of the 50 informative features of PF. More details of the 50 informative features of PF were reported in Supplementary Table S4. Overall, we observed that the optimal PF feature vector was the most powerful feature for effectively capturing the key pattern of PVPs. For convenience of discussion, the mRF model trained with the optimal PF feature vector is referred herein as SCORPION.
New feature representations improve the predictive performance
To investigate whether the optimal PF feature vector is effective in improving the predictive performance, we investigate and performed three sets of comparative experiments as follows. First, we compared the performance of SCORPION (50D) with the model without the optimal PF feature vector (80D). Second, the performance of the optimal PF feature vector was compared with 13 different feature descriptors. Finally, we compared the performance of SCORPION with its constituent baseline models. The performance comparison results between SCORPION and other methods are provided in Figs. 3, 4, Supplementary Tables S5 and S6.
Performance comparison of the optimal PFs with the top five commonly used feature descriptors on the training (A,B) and independent tests (C,D). Prediction results of the optimal PFs with the top five commonly used feature descriptors in terms of ACC, Sn, Sp and MCC (A,C). ROC curves and AUC values of the optimal PFs with the top five commonly used feature descriptors (B,D).
From Fig. 3, we observe that SCORPION performed better than that of the model without the optimal PF feature vector in terms of all the five performance metrics on both the training and independent datasets. Impressively, ACC, Sn, Sp, MCC and AUC of SCORPION were 10.40%, 7.55%, 8.54%, 20.78% and 4.61%, respectively, higher than that of the model without the optimal PF feature vector on the independent dataset. After that, we compared the optimal PF feature vector with 13 different feature descriptors. As can be seen from Supplementary Tables S5 and S6, amongst 13 different feature descriptors, the five best-performing descriptors in terms of cross-validation MCC contained PSSM_COM, PSSM_AAC, AAC, PSSM_DP and EAAC. Here, we built RF classifiers with the five best-performing descriptors and evaluate the RF classifiers’ performance based on the tenfold cross-validation and independent tests. The performance comparison results between the optimal PF feature vector and these five best-performing descriptors are depicted in Fig. 4. In the meanwhile, Supplementary Table S5 shows that the highest cross-validation ACC and MCC of 0.868 and 0.743, respectively, were achieved by using the optimal PF feature vector, while PSSM_COM performed well with the second highest cross-validation ACC and MCC of 0.814 and 0.633, respectively. In case of the independent test results, the optimal PF feature vector significantly outperformed the second-best descriptor in terms of four out of five performance metrics (i.e. ACC, Sp, MCC and AUC). Specifically, the optimal PF feature vector’s ACC, Sp, MCC and AUC were 12.70%, 25.40%, 25.87% and 12.22%, respectively, higher than the second-best descriptor. In addition, we compared the distribution of the feature space of the optimal PF feature vector and the five best-performing descriptors on the training dataset by using the t-distributed stochastic neighbor embedding (t-SNE) based on the scikit-learn (version 0.22)44,45. Figure 5 shows six t-SNE plots representing their distributions between positive (red spots) and negative (green spots) samples in a 2D feature space. As can be seen, we notice that a clear separation between red and green spots was achieved in the feature space of the optimal PF feature vector. Finally, we compared the predictive performance of SCORPION against its constituent baseline models. Figure 2 shows that MLP-PSSM_COM performed well with the highest cross-validation ACC and MCC. As can be seen from Fig. 6, SCORPION attained the overall best performance as compared with MLP-PSSM_COM in terms of all performance metrics on both training and independent datasets. Remarkably, SCORPION’s ACC, Sp, MCC and AUC were 10.32%, 19.05%, 21.40% and 6.35%, respectively, higher than MLP-PSSM_COM. This confirmed that the optimal PF feature vector derived from the integration of variant ML classifier were beneficial for PVP identification and could improve the model’s predictive performance.
Performance comparison of SCORPION with the top five baseline models on the training (A,B) and independent tests (C,D). Prediction results of SCORPION and the top five baseline models in terms of ACC, Sn, Sp and MCC (A,C). ROC curves and AUC values of SCORPION with the top five baseline models (B,D).
Model interpretation
In this section, we utilized the SHAP approach to analyze feature importance for SCORPION and three selected baseline models (i.e. RF-AAC, XGB-DPC and LR-XGB) for providing better understanding of these five models to generate their prediction outcomes. The impact of each feature on these three models’ prediction outcomes is illustrated in Fig. 7. To be specific, Fig. 7A–D show the top 20 PFs, top 20 amino acids and top 20 dipeptides respectively, based on SHAP values along with its directionality for each model, where the top 20 PFs were obtained from 20 top-ranked important features having the highest XGB classifier’s feature importance scores. Details of the top 20 PFs along with their feature importance scores are provided in Supplementary Table S7. It should be noted that negative and positive SHAP values drive the predictions as PVP and non-PVP classes, respectively, while the feature with the largest SHAP values is the most important. As seen in Fig. 7A, it is apparent that when the top five PFs of the five baseline models of MLP-PSSM_DP, NB-PSSM_AAC, MLP-PSSM_AAC, XGB-DPC and NB-PAAC had low SHAP values. Among these five baseline models, MLP-PSSM_AAC achieved the best performance in terms of cross-validation MCC (0.864). For a given unknown protein sequence P, it is predicted as PVP class if MLP-PSSM_AAC provides a low prediction probability, otherwise it is predicted as non-PVP class. From Fig. 7B, the five top-ranked informative amino acids based on SHAP values are Cys, His, Gly, Lys and Thr. Amongst these five top-ranked informative amino acids, Cys His and Lys exhibited low SHAP values, while Gly and Thr exhibited high SHAP values, suggesting that Cys His and Lys. From Fig. 7C,D, the seven top-ranked informative dipeptide based on SHAP values are TD, YT, HL, SE, MK, TG and SN.
Comparison of SCORPION with conventional ML classifiers and existing methods
In this section, the same training and independent datasets established by Charoenkwan et al.18 were employed to assess and compare the predictive performance of SCORPION against and existing methods (i.e. PVPred, PVP-SVM, PVPred-SCM, Meta-iPVP and iPVP-MCV). The performance comparison results are shown in Tables 5 and 6. In case of the tenfold cross-validation results, SCORPION and iPVP-MCV achieved better performances than Meta-iPVP in terms of all performance metrics (Table 5). In addition, SCORPION secured the best predictive performance on the independent dataset, while iPVP-MCV attained the second-best performance value. Specifically, SCORPION significantly outperformed the compared existing method in terms of ACC, Sp and MCC, while iPVP-MCV achieved the best Sn (Table 6). In the meanwhile, SCORPION’s ACC, Sp and MCC were 4.80%, 17.44% and 9.88%, respectively, higher than iPVP-MCV. Altogether, our comparative results indicate that our predictor was able to attain the best predictive performance of PVP identification as compared to the existing methods.
The significant improvement of our predictor SCORPION can be characterized to three major reasons. First, our predictor was trained and optimized using an up-to-date dataset established by Charoenkwan et al.18 containing a larger number of PVPs and non-PVPs than other datasets. Second, our predictor combined variant sequence-based feature descriptors from different perspectives consisting of compositional information, composition-transition-distribution information, position-specific information and physicochemical properties. Third, the two-step feature selection scheme was utilized for identifying the most informative features that can help to precisely discriminate PVPs from non-PVPs.
Conclusions
In this study, we introduced SCORPION, a novel, stacked, machine learning-based approach for accurate identification of PVPs. Specifically, SCORPION employed 13 different feature encoding schemes (categorized into four main groups) to encode PVPs and non-PVPs sequences and used 10 popular ML algorithms to build a pool of baseline models. These baseline models were then used to generate and construct the PF feature vector, which were considered as new feature representations. Finally, the optimal PF feature vector was optimized by using a two-step feature selection strategy and used this feature vector to develop the stacked model (SCORPION). Extensive benchmarking experiments show that SCORPION was effective and outperformed its constitute baseline models. In addition, when compared with five well-known existing methods (i.e. PVPred, PVP-SVM, PVPred-SCM, Meta-iPVP and iPVP-MCV) on the independent dataset, SCORPION achieved a superior predictive performance as compared the compared methods for PVP identification in terms of ACC (0.873), Sp (0.905), MCC (0.748) and AUC (0.891), thereby highlighting its effectiveness and generalizability. We anticipate that SCORPION will be a valuable tool for facilitating antibacterial drug discovery and development.
Data availability
All the data used in this study are available at https://github.com/saeed344/SCORPION.
References
Clark, J. R. & March, J. B. Bacteriophages and biotechnology: Vaccines, gene therapy and antibacterials. Trends Biotechnol. 24(5), 212–218 (2006).
Samson, J. E., Magadan, A. H., Sabri, M. & Moineau, S. Revenge of the phages: Defeating bacterial defences. Nat. Rev. Microbiol. 11(10), 675–687 (2013).
Lyon, J. Phage therapy’s role in combating antibiotic-resistant pathogens. JAMA 318(18), 1746–1748 (2017).
Lekunberri, I., Subirats, J., Borrego, C. M. & Balcazar, J. L. Exploring the contribution of bacteriophages to antibiotic resistance. Environ. Pollut. 220(Pt B), 981–984 (2017).
Jara-Acevedo, R. et al. Screening phage-display antibody libraries using protein arrays. In Phage Display (eds Hust, M. & Lim, T. S.) 365–380 (Springer, 2018).
Lavigne, R., Ceyssens, P.-J. & Robben, J. Phage proteomics: Applications of mass spectrometry. In Bacteriophages (eds Clokie, M. R. J. & Kropinski, A. M.) 239–251 (Springer, 2009).
Yuan, Y. & Gao, M. Proteomic analysis of a novel bacillus jumbo phage revealing glycoside hydrolase as structural component. Front. Microbiol. 7, 745 (2016).
Seguritan, V. et al. Artificial neural networks trained to detect viral and phage structural proteins. PLoS Comput. Biol. 8, e1002657 (2012).
Feng, P.-M., Ding, H., Chen, W. & Lin, H. Naive bayes classifier with feature selection to identify phage virion proteins. Comput. Math. Methods Med. 2013, 1–6 (2013).
Ding, H., Feng, P.-M., Chen, W. & Lin, H. Identification of bacteriophage virion proteins by the ANOVA feature selection and analysis. Mol. BioSyst. 10(8), 2229–2235 (2014).
Manavalan, B., Shin, T. H. & Lee, G. PVP-SVM: Sequence-based prediction of phage virion proteins using a support vector machine. Front. Microbiol. 9, 476 (2018).
Pan, Y. et al. Identification of bacteriophage virion proteins using multinomial naive bayes with g-gap feature tree. Int. J. Mol. Sci. 19(6), 1779 (2018).
Tan, J.-X., Dao, F.-Y., Lv, H., Feng, P.-M. & Ding, H. Identifying phage virion proteins by using two-step feature selection methods. Molecules 23(8), 2000 (2018).
Ru, X., Li, L. & Wang, C. Identification of phage viral proteins with hybrid sequence features. Front. Microbiol. 10, 507 (2019).
Arif, M. et al. Pred-BVP-Unb: Fast prediction of bacteriophage virion proteins using un-biased multi-perspective properties with recursive feature elimination. Genomics 112, 1565 (2019).
Charoenkwan, P., Kanthawong, S., Schaduangrat, N., Yana, J. & Shoombuatong, W. PVPred-SCM: Improved prediction and analysis of phage virion proteins using a scoring card method. Cells 9(2), 353 (2020).
Zhang, L., Zhang, C., Gao, R. & Yang, R. An ensemble method to distinguish bacteriophage virion from non-virion proteins based on protein sequence characteristics. Int. J. Mol. Sci. 16(9), 21734–21758 (2015).
Charoenkwan, P., Nantasenamat, C., Hasan, M. M. & Shoombuatong, W. Meta-iPVP: A sequence-based meta-predictor for improving the prediction of phage virion proteins using effective feature representation. J. Comput. Aided Mol. Des. 34(10), 1105–1116 (2020).
Han, H., Zhu, W., Ding, C. & Liu, T. J. S. iPVP-MCV: A multi-classifier voting model for the accurate identification of phage virion proteins. Symmetry 13(8), 1506 (2021).
Fang, Z. & Zhou, H. VirionFinder: Identification of complete and partial prokaryote virus virion protein from virome data using the sequence and biochemical properties of amino acids. Front. Microbiol. 12, 9 (2021).
Kabir, M., Nantasenamat, C., Kanthawong, S., Charoenkwan, P. & Shoombuatong, W. Large-scale comparative review and assessment of computational methods for phage virion proteins identification. EXCLI J. 21, 11–29 (2022).
Wei, L., Zhou, C., Chen, H., Song, J. & Su, R. ACPred-FL: A sequence-based predictor using effective feature representation to improve the prediction of anti-cancer peptides. Bioinformatics 34(23), 4007–4016 (2018).
Xie, R. et al. DeepVF: A deep learning-based hybrid framework for identifying virulence factors using the stacking strategy. Brief. Bioinform. 22(3), 125 (2021).
Charoenkwan, P. et al. StackIL6: A stacking ensemble model for improving the prediction of IL-6 inducing peptides. Brief. Bioinform. 22(6), 172 (2021).
Rao, B., Zhou, C., Zhang, G., Su, R. & Wei, L. ACPred-fuse: Fusing multi-view information improves the prediction of anticancer peptides. Brief. Bioinform. 21(5), 1846–1855 (2020).
Wei, L., Ye, X., Xue, Y., Sakurai, T. & Wei, L. ATSE: A peptide toxicity predictor by exploiting structural and evolutionary information based on graph neural network and attention mechanism. Brief. Bioinform. https://doi.org/10.1093/bib/bbab041 (2021).
Basith, S., Lee, G. & Manavalan, B. STALLION: A stacking-based ensemble learning framework for prokaryotic lysine acetylation site prediction. Brief. Bioinform. https://doi.org/10.1093/bib/bbab376 (2021).
UniProt Consortium. UniProt: A worldwide hub of protein knowledge. Nucleic Acids Res. 47(D1), D506–D515 (2019).
Chen, Z. et al. iFeature: A python package and web server for features extraction and selection from protein and peptide sequences. Bioinformatics 34(14), 2499–2502 (2018).
Charoenkwan, P. et al. StackDPPIV: A novel computational approach for accurate prediction of dipeptidyl peptidase IV (DPP-IV) inhibitory peptides. Methods. https://doi.org/10.1016/j.ymeth.2021.12.001 (2021).
Malik, A. A. et al. StackHCV: A web-based integrative machine-learning framework for large-scale identification of hepatitis C virus NS5B inhibitors. J. Comput. Aided Mol. Des. 35(10), 1037–1053 (2021).
Feng, C.-Q. et al. iTerm-PseKNC: A sequence-based tool for predicting bacterial transcriptional terminators. Bioinformatics 35(9), 1469–1477 (2019).
Lai, H.-Y. et al. iProEP: A computational predictor for predicting promoter. Mol. Therapy-Nucleic Acids 17, 337–346 (2019).
Lv, H. et al. Evaluation of different computational methods on 5-methylcytosine sites identification. Briefi. Bioinform. 21, 982 (2019).
Su, Z.-D. et al. iLoc-lncRNA: Predict the subcellular location of lncRNAs by incorporating octamer composition into general PseKNC. Bioinformatics 34(24), 4196–4204 (2018).
Pedregosa, F. et al. Scikit-learn: Machine learning in python. J. Mach. Learn. Res. 12, 2825–2830 (2011).
Hasan, M. M. et al. Meta-i6mA: An interspecies predictor for identifying DNA N 6-methyladenine sites of plant genomes by exploiting informative features in an integrative machine-learning framework. Brief. Bioinform. 22(3), 202 (2021).
Ahmed, S., Kabir, M., Arif, M., Ali, Z. & Swati, Z. N. K. Prediction of human phosphorylated proteins by extracting multi-perspective discriminative features from the evolutionary profile and physicochemical properties through LFDA. Chemometr. Intell. Lab. Syst. 203, 104066 (2020).
Shoombuatong, W. et al. Towards the revival of interpretable QSAR models. In Advances in QSAR Modelling (ed. Roy, K.) 3–55 (Springer, 2017).
Shoombuatong, W. et al. Towards predicting the cytochrome P450 modulation: From QSAR to proteochemometric modeling. Curr. Drug Metab. 18(6), 540–555 (2017).
Shoombuatong, W., Schaduangrat, N. & Nantasenamat, C. Towards understanding aromatase inhibitory activity via QSAR modeling. EXCLI J. 17, 688 (2018).
Azadpour, M., McKay, C. M. & Smith, R. L. Estimating confidence intervals for information transfer analysis of confusion matrices. J. Acoust. Soc. Am. 135(3), 140–146 (2014).
Shoombuatong, W., Schaduangrat, N. & Nantasenamat, C. Unraveling the bioactivity of anticancer peptides as deduced from machine learning. EXCLI J. 17, 734 (2018).
Van Der Maaten, L. Accelerating t-SNE using tree-based algorithms. J. Mach. Learn. Res. 15(1), 3221–3245 (2014).
Van der Maaten, L. & Hinton, G. Visualizing data using t-SNE. J. Mach. Learn. Res. 9(11), 2579 (2008).
Li, Z.-R. et al. PROFEAT: A web server for computing structural and physicochemical features of proteins and peptides from amino acid sequence. Nucleic Acids Res. 34, W32–W37 (2006).
Charoenkwan, P., Kanthawong, S., Nantasenamat, C., Hasan, M. M. & Shoombuatong, W. iDPPIV-SCM: A sequence-based predictor for identifying and analyzing dipeptidyl peptidase IV (DPP-IV) inhibitory peptides using a scoring card method. J. Proteome Res. 19(10), 4125–4136 (2020).
Kawashima, S. & Kanehisa, M. AAindex: Amino acid index database. Nucleic Acids Res. 28(1), 374–374 (2000).
Chou, K.-C. Using amphiphilic pseudo amino acid composition to predict enzyme subfamily classes. Bioinformatics 21(1), 10–19 (2005).
Tomii, K. & Kanehisa, M. Analysis of amino acid indices and mutation matrices for sequence comparison and structure prediction of proteins. Protein Eng. Des. Sel. 9(1), 27–36 (1996).
Dubchak, I., Muchnik, I., Holbrook, S. R. & Kim, S.-H. Prediction of protein folding class using global description of amino acid sequence. Proc. Natl. Acad. Sci. 92(19), 8700–8704 (1995).
Wang, M. et al. DeepMal: Accurate prediction of protein malonylation sites by deep neural networks. Chemometr. Intell. Lab. Syst. 207, 104175 (2020).
Charoenkwan, P. et al. Improved prediction and characterization of anticancer activities of peptides using a novel flexible scoring card method. Sci. Rep. 11(1), 1–13 (2021).
Charoenkwan, P., Yana, J., Nantasenamat, C., Hasan, M. M. & Shoombuatong, W. iUmami-SCM: A novel sequence-based predictor for prediction and analysis of umami peptides using a scoring card method with propensity scores of dipeptides. J. Chem. Inf. Model. 60(12), 6666–6678 (2020).
Wang, J. et al. POSSUM: A bioinformatics toolkit for generating numerical sequence feature descriptors based on PSSM profiles. Bioinformatics 33(17), 2756–2758 (2017).
Acknowledgements
This work was fully supported by College of Arts, Media and Technology, Chiang Mai University and partially supported by Chiang Mai University and Mahidol University. In addition, computational resources were supported by Information Technology Service Center (ITSC) of Chiang Mai University.
Author information
Authors and Affiliations
Contributions
Conceptualization, project administration, supervision, methodology and visualization: W.S.; methodology, validation, visualization and software: S.A.; writing—original draft: W.S., M.M.H., P.C., P.L., and M.A.M.; writing—review and editing: J.M.W.Q. and W.S. All authors reviewed and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ahmad, S., Charoenkwan, P., Quinn, J.M.W. et al. SCORPION is a stacking-based ensemble learning framework for accurate prediction of phage virion proteins. Sci Rep 12, 4106 (2022). https://doi.org/10.1038/s41598-022-08173-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-08173-5
This article is cited by
-
Deepstacked-AVPs: predicting antiviral peptides using tri-segment evolutionary profile and word embedding based multi-perspective features with deep stacking model
BMC Bioinformatics (2024)
-
StackedEnC-AOP: prediction of antioxidant proteins using transform evolutionary and sequential features based multi-scale vector with stacked ensemble learning
BMC Bioinformatics (2024)
-
Antimicrobial resistance crisis: could artificial intelligence be the solution?
Military Medical Research (2024)
-
Leveraging a meta-learning approach to advance the accuracy of Nav blocking peptides prediction
Scientific Reports (2024)
-
In silico drug discovery: a machine learning-driven systematic review
Medicinal Chemistry Research (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.