Abstract
Environmental heterogeneity resulting from human-modified landscapes can increase intraspecific trait variation. However, less known is whether such phenotypic variation is driven by plastic or adaptive responses to local environments. Here, we study five bumble bee (Apidae: Bombus) species across an urban gradient in the greater Saint Louis, Missouri region in the North American Midwest and ask: (1) Can urban environments induce intraspecific spatial structuring of body size, an ecologically consequential functional trait? And, if so, (2) is this body size structure the result of plasticity or adaptation? We additionally estimate genetic diversity, inbreeding, and colony density of these species—three factors that affect extinction risk. Using ≥ 10 polymorphic microsatellite loci per species and measurements of body size, we find that two of these species (Bombus impatiens, Bombus pensylvanicus) exhibit body size clines across the urban gradient, despite a lack of population genetic structure. We also reaffirm reports of low genetic diversity in B. pensylvanicus and find evidence that Bombus griseocollis, a species thought to be thriving in North America, is inbred in the greater Saint Louis region. Collectively, our results have implications for conservation in urban environments and suggest that plasticity can cause phenotypic clines across human-modified landscapes.
Similar content being viewed by others
Introduction
In the Anthropocene, we have witnessed precipitous declines of biodiversity1, with approximately 1 million species currently in threat of extinction2. Anthropogenic effects on the globe are widely recognized as the primary drivers of this biodiversity loss2. Humans have transformed up to one-half of global land surfaces3, thereby fragmenting previously continuous habitat and presenting many species with environments unencountered in their evolutionary past4. Anthropogenic change may increase extinction risk by inducing mismatch between functional traits and the environment, if such traits are not sufficiently plastic5. Additionally, by creating barriers to dispersal, such habitat fragmentation may induce genetic differentiation among isolated subpopulations and loss of genetic diversity within them, which can further exacerbate population declines6. As functional traits mediate population performance via effects on fitness7, while population genetics indicate long-term population stability8, effective conservation efforts are strengthened by integrative assessments of population genetics and how functional traits are distributed in human-modified environments.
Biodiversity loss among pollinating insects is particularly important for empirical inquiry, as insects are primarily responsible for the pollination of wild plants and agricultural crops9. Of the pollinating insects, the most comprehensive estimates of decline are for bees (Anthophila)10 and butterflies (Rhopalocera)11. Various bee taxa have experienced range contractions12, abundance declines12, and local extinctions13,14, thereby resulting in species richness losses. Among these taxa are the bumble bees (Hymenoptera: Apidae: Bombus), a monophyletic group of eusocial bees primarily native to temperate and subpolar regions of the Northern Hemisphere15. Bumble bees have undergone precipitous declines throughout their native range16,17, with estimates suggesting that approximately one-third of bumble bee species are in decline18. Anthropogenic habitat modification is widely recognized as a predominant driver of these declines10, with habitat loss reducing the availability of forage and nesting sites10, fragmentation inducing heterogeneity in species occurrences19, and population success differing between rural and urban areas20.
Previous studies have demonstrated that human-modified environments can structure bee communities interspecifically, based on the matching of functional traits to local environments21,22. However, less known is whether human-modified environments can structure bee functional traits intraspecifically. The literature is increasingly documenting that species may respond to human-induced environmental heterogeneity by increasing intraspecific trait variation23. In bees, body size is one functional trait that has considerable ecological consequences. At the community-level, body size influences pollination system connectivity by dictating the floral species from which a bee can forage24. This is predominantly due to allometric scaling between bee body size and tongue length, and the functional match between tongue length and corolla tube length24. At the individual-level, body size influences a suite of characteristics, including dispersal distance25, foraging efficiency26, and resistance to starvation27. In bumble bees, body size is developmentally plastic, with higher rates of larval feeding yielding larger adult workers28,29. This plasticity can result in up to tenfold differences in worker body size within colonies, despite workers from monogamous queens being highly related (r = 0.75)15,30. Furthermore, body size may influence bumble bees’ susceptibility to decline; species with larger average body size31 or lower variation in body size32 appear more susceptible to negative effects of human activity. Despite the known ecological implications of bumble bee body size, we lack a comprehensive understanding of how body size can be structured within-species across human-modified environments.
Conservation efforts are strengthened by considering how functional traits are distributed across heterogeneous landscapes23. Understanding the link between environment and phenotype is critical for habitat restoration33 and species relocations34. Additionally, phenotypic divergence between subpopulations may indicate variance in environmental quality and differential extinction risk among subpopulations35. Coupling functional trait investigations with population genetics can elucidate whether phenotypic divergence mirrors patterns of population genetic structuring36. If these mirror one another, phenotypic divergence may indicate divergent selection between subpopulations, while phenotypic divergence without genetic structure may indicate plasticity in local environments despite high rates of gene flow36. This is important as divergent selection can alter the delineation of evolutionarily significant units37 and the degree to which functional traits are plastic can affect range shifts, extinction, and persistence of threatened species5,38. Conservation efforts can be further strengthened by population genetics by estimating factors that may contribute to extinction risk, including inbreeding, reduced genetic diversity, and low effective population size39. Various conservation-genetic techniques have been developed to study bee ecology and evolution40. Genotyping of microsatellites has proven particularly versatile6,41 and is a robust method for detecting genetic effects of recent habitat fragmentation, even in species with high gene flow42.
Here, we investigate body size spatial structuring and population genetics in five bumble bee species across the greater Saint Louis, Missouri region: Bombus auricomus, Bombus bimaculatus, Bombus griseocollis, Bombus impatiens, and Bombus pensylvanicus. These species have experienced divergent population trends over the past two centuries in North America; B. auricomus and B. pensylvanicus have decreased relative abundance, while B. impatiens, B. bimaculatus, and B. griseocollis have experienced abundance increases17. The International Union for Conservation of Nature (IUCN) Red List categorizes all of these species as “Least Concern” with stable population trends, except for B. pensylvanicus, which is listed as “Vulnerable” with a declining population trend43. Recent data suggest a listing of “Critically Endangered” for B. pensylvanicus in Canada, following IUCN Red List criteria44. In addition to body size being intraspecifically variable, these species also exhibit marked interspecific differences in worker body size32. Bumble bees are typically regarded as dietary generalists15; however, given that body size influences the floral species that bees can forage from24, these interspecific size differences may result in a degree of resource partitioning between species. By estimating population genetics using microsatellites and analyzing intraspecific spatial structure of body size, we provide an integrative, comparative assessment of conservation genetics and trait variation in a group of at-risk pollinating insects. We ask the following questions: (1) do these species exhibit intraspecific spatial structure in body size across an urban gradient and, if so, (2) is this body size structure the result of plastic or adaptive responses to local environments? We additionally estimate genetic diversity, inbreeding, and colony density for these species throughout the greater Saint Louis region, as these factors can help inform conservation efforts. As anthropogenic changes to the biosphere continue to drive biodiversity loss, it is of paramount importance to understand functional trait variability and conservation genetics of groups at risk of extinction.
Methods
Study sites and sampling
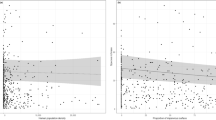
We sampled bumble bees in the greater Saint Louis, Missouri region in 2018, throughout the entire period of colony activity for each species. The five focal bumble bee species in this study (B. auricomus, B. bimaculatus, B. griseocollis, B. impatiens, B. pensylvanicus) can all be reliably found throughout this area45. We sampled bumble bees weekly from each of four sites: Calvary Cemetery (CC), EarthDance Farms (ED), Castlewood State Park (CW) (permission by Missouri Department of Natural Resources, Application for Research in Missouri State Parks 2018; Christopher Crabtree personal communication), and Shaw Nature Reserve (SNR) (Fig. 1A). These sites occur along a gradient from Saint Louis city to an area west of Saint Louis, which follows a trend of decreasing human population density (number of people km−2) with increased distance from Saint Louis (Fig. 1B). To calculate human population density, we used data on cities and towns from the United States Census Bureau46,47. We used population estimates for July 1st, 201847 as measures of human population size per locality and land area (converted to km246) as measures of total area per locality that a human population may occupy. We calculated human population density as the average number of people km−2, by dividing population estimates by land area. Human population density is a commonly used metric for anthropogenic influence on the environment48,49; therefore, we consider our sites as occurring along an urban gradient, where sites occurring in localities with greater human population density are considered more urban (Fig. 1B; see Supplemental Materials for site descriptions). As the minimum distance separating any two of these sites is greater than the typical dispersal distance of queen bumble bees50, we treat all conspecific bees per individual site as a putative subpopulation.
(A) Map of sampling locations. CC Calvary Cemetery, CW Castlewood State Park, ED EarthDance Farms, SNR Shaw Nature Reserve. Map generated with Google Earth 9.154.0.1 (https://earth.google.com). (B) Human population density per locality. Left panel: Urban gradient depicted by human population density per locality from CC (Saint Louis City, MO) to SNR (Pacific, MO). Distance from CC is the distance from CC to the approximate midpoint of a locality that occurs along the trajectory from CC to SNR. Right panel: Human population density of each locality where a site is located.
We opportunistically collected bees by hand-netting and immediately transferred them to individual ventilated vials. For all bees collected while actively foraging on a flower, we recorded the floral genus the bee was foraging on. We employed non-lethal sampling51 and released bees following data collection. Before release, we identified bees to species and sex following color patterning and morphological descriptions of Williams et al.52, removed a mid-leg tarsus from each bee and immediately stored it below 0 °C in 100% ethanol for microsatellite genotyping. For a subset of bees, we also measured thorax width using digital calipers (standard practice for measurements of bee body size15,53) prior to release.
Microsatellite genotyping
We performed DNA extraction and PCR amplification at the University of Missouri—St. Louis. Immediately prior to DNA extraction, we dried mid-leg tarsus samples and transferred each sample to a 96 well plate. In between samples, we immersed the forceps used for this work in 95% ethanol to prevent cross contamination. We followed a Chelex-based DNA extraction protocol54, whereby we added 150 μL Chelex 100 (Bio-Rad) and 5 μL Proteinase K (Apex Bioresearch) to each sample, and subsequently incubated samples in a Bio-Rad T100 Thermal Cycler with the following conditions: (1) 55 °C for 1 h, (2) 99 °C for 15 min, (3) 37 °C for 1 min, and (4) 99 °C for 15 min. Prior to PCR amplification, we stored extracted DNA samples at − 20 °C.
We genotyped each sample at 18 dye-labeled microsatellite loci55,56,57,58. Not all loci were successfully amplified or reliably scored within each species, so each species had its own complement of loci used for analyses (Table S1). We ran two multiplex PCRs per sample (i.e., plexes A and B), with six to nine microsatellite primers in each multiplex. Each multiplex reaction mixture contained 1 μL Chelex DNA extraction supernatant, 2 μL Promega 5 × buffer, 0.56 μL MgCl2 25 mM (Promega), 0.6 μL dNTP (Promega), 0.2 μL bovine serum albumin (Promega), 0.08 μL Taq polymerase (Promega), 2.28–3.08 μL H2O, and 0.045–0.400 μL of each primer (forward labelled with VIC, NED, 6-FAM, or PET dyes; Applied Biosystems). Each sample had a total reaction mixture volume of 10 μL, contained in a new well of a 96 well plate. We performed each PCR using a Bio-Rad T100 Thermal Cycler with the following conditions: (1) 95 °C hot start, (2) initial denaturation at 95 °C for 3.5 min, (3) 31 cycles of 95 °C for 30 s, 55 °C (plex A) or 58 °C (plex B) for 1.25 min, 72 °C for 45 s, and (4) final extension of 72 °C for 15 min. Subsequently, we sent 2 μL of each PCR product to the University of Missouri DNA Core for fragment analysis, where DNA Core staff added formamide and an internal size standard (600 LIZ). We scored alleles using Geneious 11.0.4 with the Microsatellite Plugin59. Following microsatellite genotyping, we verified species identifications by confirming that each individual’s alleles fell within their species-specific allele bins. Furthermore, we discarded from downstream genetic analyses all individuals and loci with 20% or greater genotyping failure per species.
Colony density
Measuring effective population size (Ne) can be problematic in eusocial insects, as non-reproductive worker abundances can inflate Ne, unless colony relationships are controlled for60. Therefore, we used colony density (Nc) (i.e., effective colony number) as a measure of Ne, which estimates the number of colonies at a site after controlling for colony relationships among workers6,60,61. We calculated Nc solely with female genotypes. Prior to estimating Nc, we removed loci per species that had ≥ 25% null allele frequency following Chakraborty et al.62, using the R package PopGenReport 2.063. We estimated Nc per subpopulation by first reconstructing female sibships in Colony 2.064 using a 5% genotyping error rate and a 95% probability of females being full siblings. Following sibship reconstructions, we calculated Nc following Geib et al.61. To do so, we first determined the number of sampled females (Ni), the number of successfully genotyped females (Ng), and the number of colonies detected by Colony (Nnr). We then calculated the number of colonies detected standardized for genotyping success as Nns = (Nnr/Ng) × Ni. Finally, we calculated Nc according to the Crozier model for effective population size of eusocial haplodiploid species that estimates detected colonies plus colonies not detected by sampling: Nc = (4.5Nnm)/(1 + 2m); N is detected colony number, n is queen number per colony, and m is mating frequency65. Accordingly, for species like bumble bees, that are characterized by monogyny and monoandry15, this calculation simplifies to Nc = 1.5 × Nns6. We did not calculate Nc for any subpopulation with 15 or fewer successfully genotyped females (i.e., Ng ≤ 15).
Population genetic analyses
We included only one randomly chosen sister per colony for population genetic analyses. After retaining one sister per colony, we checked loci for linkage disequilibrium (LD) using the R package Genepop ‘007 1.1.466. If we found two or more loci to be in significant LD (p < 0.05), we retained only one of these loci for further genetic analyses. We tested individual loci for Hardy–Weinberg equilibrium (HWE) using the R package PopGenReport 2.063.
Following these quality control measures, we calculated allelic richness (i.e., mean allele number per locus; AR) per subpopulation and global AR per species (i.e., species-level AR grouping samples across sites). As AR can be sensitive to variances in sample size, sample size rarefaction is the preferred method of standardizing AR for comparative studies67. Prior to calculating AR values, we rarefied subpopulation sample sizes to the lowest subpopulation sample size across all five species, using the R package hierfstat 0.04-2268. For global measures of AR, we rarefied each species’ sample size to the sample size of the species with the lowest overall sample size.
To assess genetic differentiation among intraspecific subpopulations, we calculated FST across all loci per species69 in FSTAT 2.9.4. To ensure that our data had sufficient statistical power to detect true genetic differentiation, we performed a power simulation per species with the program POWSIM 4.1, which tests the null hypothesis of no genetic differentiation between subpopulations, given different combinations of sample size, loci, and alleles70. See Supplemental Materials for full power analysis methods.
We tested each species for possible inbreeding by (1) calculating the inbreeding coefficient, FIS, across all loci per species69 in FSTAT 2.9.4, and (2) inspecting males for diploidy. In bee populations, diploid male frequency increases with inbreeding due to increased rates of homozygosity at the complementary sex determination locus71. To assess male diploidy, for each male bee we recorded whether each successfully genotyped locus was scored as homozygous or heterozygous. Following Darvill et al.72, we then recorded a male as diploid if three or more of his loci were scored as heterozygous. For calculations of FST, FIS, and subpopulation AR, we removed all individuals from populations with < 25 samples following our quality control measures73. However, we did not remove individuals from populations with a low sample size for our calculations of global AR, while still ensuring that only one randomly chosen sister per colony was included in these calculations.
Body size variation analyses
For all body size variation analyses, we included only one randomly chosen sister per colony and excluded all subpopulations that included ≤ 15 workers with thorax width measurements. Given our weekly sampling protocol across sites, these measurements collectively represent body size variation across each species’ entire period of colony activity. To determine whether our focal bumble bee species exhibit intraspecific spatial structure in body size, we compared intraspecific subpopulations for significantly different average body sizes. We first ran an analysis of variance (ANOVA) with thorax width as the response variable, and site and species as categorical predictors. Subsequently, we ran contrasts between least squares means for each unique pairing of intraspecific subpopulations. We used a Bonferroni corrected α-value to determine statistical significance of these contrasts. To compute these contrasts, we used the R package lsmeans 2.3074.
Results
Sampling and genotyping
Across all species and sites, we collected 839 bees; 774 females and 65 males. Sample sizes were variable across species and sites (Tables 1 and 2), ranging from conspecific bees being absent or found in low abundance to upwards of 70 conspecific bees collected at a site. Following all genotyping quality control measures, each species had a minimum of 10 loci used in population genetic analyses (Fig. S1; Table S1). A description of these quality control results and loci retained per species can be found in the Supplemental Materials.
Colony density
Each species had variable colony densities across sites. Nc ranges from a minimum of 19.6 (B. pensylvanicus at ED) to a maximum of 98.7 (B. bimaculatus at CW) (Table 1). We could not calculate Nc for B. auricomus, B. griseocollis, or B. pensylvanicus at CW, and for B. bimaculatus at CC, due to fewer than 15 females having been successfully genotyped for these subpopulations (i.e., Ng < 15) (Table 1).
Population genetic analyses
Throughout the greater Saint Louis region, genetic differentiation between intraspecific subpopulations was low to absent in each species, with FST ≤ 0.002 in each species and all 95% CIs including zero (Table 3). Each power simulation revealed statistical power > 0.99 for detecting an FST = 0.05 using both Chi-square and Fisher’s exact tests. Accordingly, our sampling protocol had a > 99% probability of detecting true FST values of 0.05. FIS values were more variable, ranging from a minimum of 0.023 (B. bimaculatus) to a maximum of 0.151 (B. griseocollis) (Table 3). Zero is only included in the FIS 95% CI of B. bimaculatus. All males collected were haploid, except in B. griseocollis for which 21 of 25 collected males (84%) were diploid (i.e., ≥ 3 loci scored as heterozygous) (Table 2). Global AR calculations were rarefied to a sample size of 88 per species, following B. pensylvanicus having the lowest overall sample size (i.e., 88 female genotypes retained × 2 alleles/female = 176 alleles). Subpopulation AR calculations were rarefied to a subpopulation size of 28, as the subpopulation included in genetic analyses with the lowest sample size was B. pensylvanicus at CC (i.e., 28 female genotypes retained × 2 alleles/female = 56 alleles). AR varies interspecifically (i.e., between species’ global AR values) and between intraspecific loci (Table 3; Fig. S1). Bombus pensylvanicus had the lowest AR across all species (global AR = 6.29 ± 1.42 SE) and B. impatiens had the highest AR (global AR = 10.24 ± 2.21 SE). We could not calculate FST, FIS, and site-specific AR for B. auricomus, B. griseocollis, or B. pensylvanicus at CW, B. bimaculatus at CC, and B. pensylvanicus at ED due to < 25 genotypes remaining in each of these subpopulations following our quality control measures.
Body size variation analyses
We find evidence for spatial structuring of intraspecific body size for bumble bees in the greater Saint Louis region. Our full ANOVA revealed significant effects of species, site, and their interaction on worker thorax width (species, site, and species × site all p < 0.0001). After Bonferroni correction, average body size significantly differs between intraspecific subpopulations of B. impatiens and B. pensylvanicus. Specifically, for B. impatiens, worker body size was larger on average at the urban CC site than at the more rural CW site (contrast of least square means p < 0.0001) (Fig. 2). For B. pensylvanicus worker body size was larger on average at the rural SNR site than at the more urban CC and ED sites (both contrasts of least square means p < 0.0001) (Fig. 2). No other species showed significant spatial structuring of average body size (all contrasts of least square means p > 0.006) (Table S2). The Bonferroni adjusted α-value used for determining statistical significance between average body size contrasts is α = 0.00278 (i.e., 0.05/18 contrasts) (Table S2). We did not include B. auricomus, B. griseocollis, or B. pensylvanicus at CW, and B. bimaculatus at CC in these analyses due to ≤ 15 workers having thorax width measurements at these subpopulations. See Table S3 for all worker thorax width sample sizes and body size means per subpopulation.
Thorax widths of worker bumble bees (Bombus spp.) across the urban gradient in the greater Saint Louis region. Sites are arranged from left to right by increasing distance from Saint Louis City. Asterisks (*) indicate statistically significant differences between means of intraspecific subpopulations following Bonferroni correction (i.e., p < 0.00278). CC Calvary Cemetery, CW Castlewood State Park, ED EarthDance Farms, SNR Shaw Nature Reserve.
Discussion
Studying five bumble bee species across an urban gradient in the greater Saint Louis region, we find evidence for intraspecific spatial structuring of body size, despite genetic homogeneity among subpopulations. Specifically, two species, B. impatiens and B. pensylvanicus, exhibited body size clines across the urban gradient; however, the direction of these clines were not consistent between species (i.e., sites with increased urbanization were associated with larger B. impatiens and smaller B. pensylvanicus). As our study sites occur along a gradient from the city of Saint Louis to a rural area west of the city (Fig. 1), these results suggest that human-modified environments can drive body size differences between intraspecific subpopulations of pollinating insects. Notably, we also reaffirm previous reports of low genetic diversity in B. pensylvanicus12,41 and find evidence that B. griseocollis, a species thought to be thriving in North America, may be inbred in the greater Saint Louis region.
Two non-mutually exclusive explanations may account for the observed intraspecific body size clines across the urban gradient: phenotypic plasticity or local adaptation. We argue that our results suggest plasticity underlies the observed phenotypic clines for two primary reasons. First, we did not find evidence for genetic structure in any of our studied species; i.e., all FST values are low (all FST ≤ 0.002; Table 3) and our power analyses indicate that our data had sufficient statistical power to detect true genetic differentiation, if it were present. This suggests high rates of intraspecific gene flow, either currently or in the recent past, throughout the greater Saint Louis region. High rates of gene flow often limit subpopulations from adapting to their local environments, by homogenizing traits throughout a metapopulation75. Second, body size is an exceptionally plastic trait in bumble bees, with tenfold differences in body size occurring among highly related intra-colony workers (r = 0.75)15,30. Plasticity can shield a population from local adaptation by moving the population toward an adaptive peak, thus enabling persistence in a changed environment without adaptive genetic change76. Accordingly, the lack of genetic structure, coupled with the known plasticity of bumble bee body size, supports the observed body size spatial structuring being a result of plastic responses to local environments, as opposed to adaptive genetic divergence. However, we cannot definitively rule out the possibility of local adaptation; in rare cases, subpopulations can become locally adapted even while gene flow is maintained77. It is possible that recent habitat fragmentation has induced strong differential selection between subpopulations, though sufficient time has not passed for population genetics to reflect this. However, this may be an unlikely explanation of our results, as microsatellites can document genetic effects of recent fragmentation in species of pollinating insects with high gene flow42.
Several environmental factors may drive this observed spatial structuring of body size. In bumble bees, worker larvae fed a higher quality diet or at higher rates develop into larger adults28,29. It is possible that body size clines in urban environments result from differences in nutritional quality and/or quantity among sites, whereby large size is promoted by high nutritional quality/quantity (or small size results from a constraint of low nutritional quality/quantity). While we did not directly quantify nutrition in this study, our data suggest this may be a likely explanation of our results. First, in all cases where average body size significantly differed between intraspecific sites (i.e., between CC and CW for B. impatiens and between SNR and both CC and ED for B. pensylvanicus; Fig. 2), conspecific females were observed foraging from a higher richness of floral genera at the sites where body size was larger (Table S4). As bees often optimize nutritional intake by foraging from a variety of floral species78, this may correspond to bees having more balanced diets at sites with a higher richness of exploitable floral genera. Second, at all sites where average body size was larger intraspecifically, not only were more floral genera exploited, but colony density was higher as well (Table 1). Numerous studies indicate that colony success is dependent on nutritional availability at a site78,79. Thus, the higher colony density observed at sites with a greater richness of exploited floral genera supports the idea that these sites conferred greater nutritional quality and/or quantity. This interpretation—i.e., that the observed body size clines are driven by differences in nutritional quality—would account for the opposite direction of clines found for B. impatiens and B. pensylvanicus, as long as nutritional quality is greater in urban environments for B. impatiens, but is lower in urban environments for B. pensylvanicus. Future research should explore the degree of resource partitioning between these two species and whether such partitioning results in contrasting resource quality clines across urban gradients.
It is notable that the greatest magnitude of body size spatial structuring was observed in B. pensylvanicus. Numerous reports have suggested B. pensylvanicus is the species most at risk of extinction among those studied43,44 and our finding of B. pensylvanicus having the lowest genetic diversity among bumble bee species throughout the greater Saint Louis region (lowest AR in both 2018 and 2017; see Supplemental Materials for description of 2017 population genetics; Table 3; Table S5) reaffirms these reports. Interestingly, however, at SNR—the site where B. pensylvanicus was largest intraspecifically and was found feeding from a comparatively high number of floral genera—B. pensylvanicus colony density was highest both intraspecifically (i.e., across sites) and interspecifically (i.e., highest interspecific colony density at SNR in both 2018 and 2017) (Table 1 and Table S5). This may suggest that sites with high floral species richness provide robust support to B. pensylvanicus populations, thus indicating the potential role that floral enrichment across urbanized landscapes can play in supporting populations of threatened bumble bee species.
The importance of investigating functional trait diversity of threatened species has been increasingly recognized as conservation program efficacy depends on environmental effects on the development and expression of phenotype33,80 and plasticity is a primary response of species to global change4. The body size clines we observed suggest that human-modified environments can induce landscape-level structuring of developmentally plastic functional traits. Conservation programs should be cognizant of when traits are developmentally, but irreversibly, plastic, as is the case for body size in bumble bees29,30. For example, Lema and Nevitt document that pupfish (Cyprinodon spp.) exhibit a developmentally plastic small body size as a result of high water temperature and low food availability35. They suggest that management programs should consider this by captively breeding pupfish in similar conditions to the population they will be reintroduced to, so that large individuals with high dietary requirements are not reintroduced into a food-limited environment35. Similarly, if the body size clines we observed resulted from nutritional differences among sites, this may suggest that spatial structuring of bumble bee body size can be used to indicate variance in environmental quality, with subpopulations with relatively smaller average body sizes being targeted for floral enrichment. However, alternative explanations may underlie the observed body size clines. For example, spatial heterogeneity in environmental contaminants could differentially expose subpopulations to pollutants, which may have downstream effects on foraging behavior81 and the development of adult body size82. Alternatively, in urban areas, increased metabolic demands imposed by the urban-heat-island (UHI) effect are expected to drive shifts toward smaller body size in certain taxa83. The direction of the B. pensylvanicus body size cline across the urban gradient follows the predicted direction under the UHI effect, analogous to the Brazilian stingless bee, Melipona fasciculata84; however, the B. impatiens cline follows the opposite pattern. Furthermore, in taxa where body size positively correlates with dispersal distance, habitat fragmentation may drive increased body size to promote movement of individuals between habitat patches83,85. Interestingly, Theodorou et al.86 found evidence for this trend in Bombus terrestris in Central Europe, where workers were larger in cities compared to nearby rural sites, potentially due to urban fragmentation. However, similar to the UHI effect, the contrasting directions of the body size clines found for B. impatiens and B. pensylvanicus complicate this as a likely explanation for our results. To gain a comprehensive understanding of this system, future studies should directly quantify nutrition, environmental factors, and fragmentation across subpopulations.
Our results exemplify the importance of simultaneously investigating functional trait distributions and conservation genetics of species in urban environments. While B. griseocollis does not exhibit spatial structuring of body size throughout the greater Saint Louis area, we find evidence that B. griseocollis is potentially inbred in this region. Bombus griseocollis had the highest inbreeding coefficient (FIS) and the second lowest global AR among the studied species (Table 3) and 84% of sampled B. griseocollis males were diploid in 2018 (Table 2). In haplodiploid bees, males develop via either (1) parthenogenesis, in which hemizygosity at the sex-determining locus produces a viable, haploid male, or (2) a fertilized egg, in which homozygosity at the sex-determining locus produces a sterile, diploid male71. As inbreeding promotes an increased proportion of homozygosity80, diploid males may occur at higher frequencies in inbred haplodiploid populations. While additional sampling in the Midwest is needed, in replicate years and populations, these results suggest relatively high rates of inbreeding in Saint Louis B. griseocollis populations, despite B. griseocollis being broadly distributed and abundant throughout much of the United States87 and listed as “Least Concern” by the IUCN Red List43. Indeed, future research on B. griseocollis populations is needed, as understanding why the observed rates of B. griseocollis male diploidy are so high will be critical to implementing effective conservation programs. Collectively, our results indicate the utility of simultaneously investigating phenotypic and genetic variation of threatened species, as phenotypic and genetic signatures of population stability can occur independently of one another and together provide a more complete understanding of population stability across heterogeneous landscapes.
Conclusions
The conservation of threatened species is strengthened by integrative assessments of functional trait variability and population genetics. We document that bumble bees can exhibit intraspecific body size clines in human-modified environments, despite subpopulations being genetically homogenous. These results suggest that urbanization can induce landscape-level structuring of functional traits that are developmentally plastic, potentially due to nutritional differences across sites. We additionally find evidence that (1) B. pensylvanicus has comparatively low genetic diversity, reaffirming findings from previous studies12,41 and (2) B. griseocollis is inbred in the greater Saint Louis region. Collectively, these results are informative for the development of bumble bee conservation programs and add to a growing body of literature on how threatened species are affected by human-modified environments.
Data availability
All data used for analyses can be found in the Supplementary Information.
References
Corlett, R. T. The Anthropocene concept in ecology and conservation. Trends Ecol. Evol. 30, 36–41 (2015).
IPBES. Summary for Policymakers of the Global Assessment Report on Biodiversity and Ecosystem Services of the Intergovernmental Science-Policy Platform on Biodiversity and Ecosystem Services (IPBES Secretariat, 2019).
Vitousek, P. M. Human domination of Earth’s ecosystems. Science 277, 494–499 (1997).
Wong, B. B. M. & Candolin, U. Behavioral responses to changing environments. Behav. Ecol. 26, 665–673 (2015).
Hale, R. & Swearer, S. E. Ecological traps: Current evidence and future directions. Proc. R. Soc. B Biol. Sci. 283, 1–8 (2016).
Charman, T. G., Sears, J., Green, R. E. & Bourke, A. F. G. Conservation genetics, foraging distance and nest density of the scarce Great Yellow Bumblebee (Bombus distinguendus). Mol. Ecol. 19, 2661–2674 (2010).
Violle, C. et al. Let the concept of trait be functional!. Oikos 116, 882–892 (2007).
Husemann, M., Zachos, F. E., Paxton, R. J. & Habel, J. C. Effective population size in ecology and evolution. Heredity 117, 191–192 (2016).
Wagner, D. L. Insect declines in the Anthropocene. Annu. Rev. Entomol. 65, 457–480 (2020).
Goulson, D., Nicholls, E., Botías, C. & Rotheray, E. L. Bee declines driven by combined stress from parasites, pesticides, and lack of flowers. Science 347, 1255957 (2015).
Thogmartin, W. E. et al. Monarch butterfly population decline in North America: Identifying the threatening processes. R. Soc. Open Sci. 4, 170760 (2017).
Cameron, S. A. et al. Patterns of widespread decline in North American bumble bees. Proc. Natl. Acad. Sci. U.S.A. 108, 662–667 (2011).
Burkle, L. A., Marlin, J. C. & Knight, T. M. Plant-pollinator interactions over 120 years: Loss of species, co-occurrence, and function. Science 340, 1611–1615 (2013).
Grixti, J. C., Wong, L. T., Cameron, S. A. & Favret, C. Decline of bumble bees (Bombus) in the North American Midwest. Biol. Conserv. 142, 75–84 (2009).
Goulson, D. Bumblebees: Behaviour, Ecology, and Conservation (Oxford University Press, Oxford, 2010).
Colla, S. R., Gadallah, F., Richardson, L., Wagner, D. & Gall, L. Assessing declines of North American bumble bees (Bombus spp.) using museum specimens. Biodivers. Conserv. 21, 3585–3595 (2012).
Hatfield, R. et al. IUCN assessments of North American Bombus spp. http://www.xerces.org/ (2015).
Arbetman, M. P., Gleiser, G., Morales, C. L., Williams, P. & Aizen, M. A. Global decline of bumblebees is phylogenetically structured and inversely related to species range size and pathogen incidence. Proc. R. Soc. B Biol. Sci. 284, 20170204 (2017).
Bommarco, R. et al. Dispersal capacity and diet breadth modify the response of wild bees to habitat loss. Proc. R. Soc. B Biol. Sci. 277, 2075–2082 (2010).
Hall, D. M. et al. The city as a refuge for insect pollinators. Conserv. Biol. 31, 24–29 (2017).
Banaszak-Cibicka, W. & Żmihorski, M. Wild bees along an urban gradient: Winners and losers. J. Insect Conserv. 16, 331–343 (2012).
Wilson, C. J. & Jamieson, M. A. The effects of urbanization on bee communities depends on floral resource availability and bee functional traits. PLoS One 14, e0225852 (2019).
Thompson, M.J., Capilla-Lasheras, P.C., Dominoni, D.M., Réale, D. & Charmantier, A. Phenotypic variation in urban environments: mechanisms and implications. Trends Ecol. Evol. 37, 171–182 (2022).
Peat, J., Tucker, J. & Goulson, D. Does intraspecific size variation in bumblebees allow colonies to efficiently exploit different flowers?. Ecol. Entomol. 30, 176–181 (2005).
Greenleaf, S. S., Williams, N. M., Winfree, R. & Kremen, C. Bee foraging ranges and their relationship to body size. Oecologia 153, 589–596 (2007).
Spaethe, J. & Weidenmüller, A. Size variation and foraging rate in bumblebees (Bombus terrestris). Insectes Soc. 49, 142–146 (2002).
Couvillon, M. J. & Dornhaus, A. Small worker bumble bees (Bombus impatiens) are hardier against starvation than their larger sisters. Insectes Soc. 57, 193–197 (2010).
Pendrel, B. A. & Plowright, R. C. Larval feeding by adult bumble bee workers (Hymenoptera: Apidae). Behav. Ecol. Sociobiol. 8, 71–76 (1981).
Sutcliffe, G. H. & Plowright, R. C. The effects of food supply on adult size in the bumble bee Bombus terricola Kirby (Hymenoptera: Apidae). Can. Entomol. 120, 1051–1058 (1988).
Couvillon, M. J. & Dornhaus, A. Location, location, location: Larvae position inside the nest is correlated with adult body size in worker bumble-bees (Bombus impatiens). Proc. R. Soc. B Biol. Sci. 276, 2411–2418 (2009).
Bartomeus, I. et al. Historical changes in northeastern US bee pollinators related to shared ecological traits. Proc. Natl. Acad. Sci. U.S.A. 110, 4656–4660 (2013).
Austin, M. W. & Dunlap, A. S. Intraspecific variation in worker body size makes North American bumble bees (Bombus spp.) less susceptible to decline. Am. Nat. 194, 381–394 (2019).
Watters, J. V., Lema, S. C. & Nevitt, G. A. Phenotype management: A new approach to habitat restoration. Biol. Conserv. 112, 435–445 (2003).
Haddaway, N. R., Mortimer, R. J. G., Christmas, M., Grahame, J. W. & Dunn, A. M. Morphological diversity and phenotypic plasticity in the threatened British white-clawed crayfish (Austropotamobius pallipes). Aquat. Conserv. Mar. Freshw. Ecosyst. 22, 220–231 (2012).
Lema, S. C. & Nevitt, G. A. Testing an ecophysiological mechanism of morphological plasticity in pupfish and its relevance to conservation efforts for endangered Devils Hole pupfish. J. Exp. Biol. 209, 3499–3509 (2006).
Crispo, E. Modifying effects of phenotypic plasticity on interactions among natural selection, adaptation and gene flow. J. Evol. Biol. 21, 1460–1469 (2008).
Fraser, D. J. & Bernatchez, L. Adaptive evolutionary conservation: Towards a unified concept for defining conservation units. Mol. Ecol. 10, 2741–2752 (2001).
Nicotra, A. B. et al. Plant phenotypic plasticity in a changing climate. Trends Plant Sci. 15, 684–692 (2010).
Spielman, D., Brook, B. W. & Frankham, R. Most species are not driven to extinction before genetic factors impact them. Proc. Natl. Acad. Sci. U.S.A. 101, 15261–15264 (2004).
Woodard, S. H. et al. Molecular tools and bumble bees: Revealing hidden details of ecology and evolution in a model system. Mol. Ecol. 24, 2916–2936 (2015).
Lozier, J. D., Strange, J. P., Stewart, I. J. & Cameron, S. A. Patterns of range-wide genetic variation in six North American bumble bee (Apidae: Bombus) species. Mol. Ecol. 20, 4870–4888 (2011).
Williams, B. L., Brawn, J. D. & Paige, K. N. Landscape scale genetic effects of habitat fragmentation on a high gene flow species: Speyeria idalia (Nymphalidae). Mol. Ecol. 12, 11–20 (2003).
IUCN. The IUCN Red List of Threatened Species. https://www.iucnredlist.org. Accessed 18 Dec 2019 (2019).
MacPhail, V. J., Richardson, L. L. & Colla, S. R. Incorporating citizen science, museum specimens, and field work into the assessment of extinction risk of the American Bumble bee (Bombus pensylvanicus De Geer 1773) in Canada. J. Insect Conserv. 23, 597–611 (2019).
Camilo, G. R., Muñiz, P. A., Arduser, M. S. & Spevak, E. M. A checklist of the bees (Hymenoptera: Apoidea) of St. Louis, Missouri, USA. J. Kansas Entomol. Soc. 90, 175–188 (2018).
United States Census Bureau. Land Area and Persons Per Square Mile. https://www.census.gov/quickfacts/fact/note/US/LND110210. Accessed 26 March 2020 (2010).
United States Census Bureau. City and Town Population Totals: 2010–2018. https://www.census.gov/data/tables/time-series/demo/popest/2010s-total-cities-and-towns.html. Accessed 26 March 2020 (2020).
Thompson, K. & Jones, A. Human population density and prediction of local plant extinction in Britain. Conserv. Biol. 13, 185–189 (1999).
Fontana, C. S., Burger, M. I. & Magnusson, W. E. Bird diversity in a subtropical South-American City: Effects of noise levels, arborisation and human population density. Urban Ecosyst. 14, 341–360 (2011).
Lepais, O. et al. Estimation of bumblebee queen dispersal distances using sibship reconstruction method. Mol. Ecol. 19, 819–831 (2010).
Holehouse, K. A., Hammond, R. L. & Bourke, A. F. G. Non-lethal sampling of DNA from bumble bees for conservation genetics. Insectes Soc. 50, 277–285 (2003).
Williams, P. H., Thorp, R., Richardson, L. & Colla, S. R. Bumble Bees of North America (Princeton University Press, 2014).
Cane, J. H. Estimation of bee size using intertegular span (Apoidea). J. Kansas Entomol. Soc. 60, 145–147 (1987).
Walsh, P. S., Metzger, D. A. & Higuchi, R. Chelex 100 as a medium for simple extraction of DNA for PCR-based typing from forensic material. Biotechniques 10, 506–513 (1991).
Estoup, A., Scholl, A., Pouvreau, A. & Solignac, M. Monoandry and polyandry in bumble bees (Hymenoptera; Bombinae) as evidenced by highly variable microsatellites. Mol. Ecol. 4, 89–94 (1995).
Estoup, A., Solignac, M., Cornuet, J. M., Goudet, J. & Scholl, A. Genetic differentiation of continental and island populations of Bombus terrestris (Hymenoptera: Apidae) in Europe. Mol. Ecol. 5, 19–31 (1996).
Funk, C. R., Schmid-Hempel, R. & Schmid-Hempel, P. Microsatellite loci for Bombus spp. Mol. Ecol. Notes 6, 83–86 (2006).
Stolle, E. et al. Novel microsatellite DNA loci for Bombus terrestris (Linnaeus, 1758). Mol. Ecol. Resour. 9, 1345–1352 (2009).
Kearse, M. et al. Geneious Basic: An integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28, 1647–1649 (2012).
Chapman, R. E. & Bourke, A. F. G. The influence of sociality on the conservation biology of social insects. Ecol. Lett. 4, 650–662 (2001).
Geib, J. C., Strange, J. P. & Galen, A. Bumble bee nest abundance, foraging distance, and host-plant reproduction: Implications for management and conservation. Ecol. Appl. 25, 768–778 (2015).
Chakraborty, R., Andrade, M. D. E., Daiger, S. P. & Budowle, B. Apparent heterozygote deficiencies observed in DNA typing data and their implications in forensic applications. Ann. Hum. Genet. 56, 45–57 (1992).
Gruber, B. & Adamack, A. T. PopGenReport: Simplifying basic population genetic analyses in R. Methods Ecol. Evol. 5, 384–387 (2014).
Wang, J. Sibship reconstruction from genetic data with typing errors. Genetics 166, 1963–1979 (2004).
Crozier, R. H. Genetics of sociality. In Social Insects Vol. I (ed. Hermann, H. R.) 223–286 (Academic Press, 1979).
Rousset, F. genepop’007: A complete re-implementation of the genepop software for Windows and Linux. Mol. Ecol. Resour. 8, 103–106 (2008).
Leberg, P. L. Estimating allelic richness: Effects of sample size and bottlenecks. Mol. Ecol. 11, 2445–2449 (2002).
Goudet, J. hierfstat, a package for r to compute and test hierarchical F-statistics. Mol. Ecol. Notes 5, 184–186 (2005).
Weir, B. S. & Cockerham, C. C. Estimating F-statistics for the analysis of population structure. Evolution 38, 1358–1370 (1984).
Ryman, N. & Palm, S. POWSIM: A computer program for assessing statistical power when testing for genetic differentiation. Mol. Ecol. Notes 6, 600–602 (2006).
Zayed, A. & Packer, L. High levels of diploid male production in a primitively eusocial bee (Hymenoptera: Halictidae). Heredity 87, 631–636 (2001).
Darvill, B., Ellis, J. S., Lye, G. C. & Goulson, D. Population structure and inbreeding in a rare and declining bumblebee, Bombus muscorum (Hymenoptera: Apidae). Mol. Ecol. 15, 601–611 (2006).
Hale, M. L., Burg, T. M. & Steeves, T. E. Sampling for microsatellite-based population genetic studies: 25 to 30 Individuals per population is enough to accurately estimate allele frequencies. PLoS One 7, e45170 (2012).
Lenth, R. V. Least-squares means: The R package lsmeans. J. Stat. Softw. 69, 1–33 (2016).
Fitzpatrick, S. W. et al. Gene flow constrains and facilitates genetically based divergence in quantitative traits. Copeia 105, 462–474 (2017).
Price, T. D., Qvarnström, A. & Irwin, D. E. The role of phenotypic plasticity in driving genetic evolution. Proc. R. Soc. B Biol. Sci. 270, 1433–1440 (2003).
Liu, B.-J., Zhang, B.-D., Xue, D.-X., Gao, T.-X. & Liu, J.-X. Population structure and adaptive divergence in a high gene flow marine fish: The small yellow croaker (Larimichthys polyactis). PLoS One 11, e0154020 (2016).
Vaudo, A. D., Tooker, J. F., Grozinger, C. M. & Patch, H. M. Bee nutrition and floral resource restoration. Curr. Opin. Insect Sci. 10, 133–141 (2015).
Woodard, S. H. & Jha, S. Wild bee nutritional ecology: Predicting pollinator population dynamics, movement, and services from floral resources. Curr. Opin. Insect Sci. 21, 83–90 (2017).
Keller, L. F. & Waller, D. M. Inbreeding effects in wild populations. Trends Ecol. Evol. 17, 230–241 (2002).
Sivakoff, F. S. & Gardiner, M. M. Soil lead contamination decreases bee visit duration at sunflowers. Urban Ecosyst. 20, 1221–1228 (2017).
Whitehorn, P. R., Norville, G., Gilburn, A. & Goulson, D. Larval exposure to neonicotinoid imidacloprid impacts adult size in the farmland butterfly Pieris brassicae. PeerJ 6, e4772 (2018).
Merckx, T., Kaiser, A. & Van Dyck, H. Increased body size along urbanization gradients at both community and intraspecific level in macro-moths. Glob. Change Biol. 24, 3837–3848 (2018).
Oliveira, M. O., Brito, T. F., Campbell, A. J. & Contrera, F. A. L. Body size and corbiculae area variation of the stingless bee Melipona fasciculata Smith, 1854 (Apidae, Meliponini) under different levels of habitat quality in the eastern Amazon. Entomol. Gen. 39, 45–52 (2019).
Warzecha, D., Diekötter, T., Wolters, V. & Jauker, F. Intraspecific body size increases with habitat fragmentation in wild bee pollinators. Landsc. Ecol. 31, 1449–1455 (2016).
Theodorou, P., Baltz, L. M., Paxton, R. J. & Soro, A. Urbanization is associated with shifts in bumblebee body size, with cascading effects on pollination. Evol. Appl. 14, 53–68 (2021).
Strange, J. P. & Tripodi, A. D. Characterizing bumble bee (Bombus) communities in the United States and assessing a conservation monitoring method. Ecol. Evol. 9, 1061–1069 (2019).
Acknowledgements
We thank two anonymous reviewers for their constructive feedback, which improved our manuscript’s quality. Our gratitude is extended to Joyce Knoblett of the USDA-ARS Pollinating Insect—Biology, Management, Systematics Research Unit for assisting with the molecular work of our 2017 samples. We thank Shaw Nature Reserve, EarthDance Farms, Calvary Cemetery, Castlewood State Park, and their employees for access to these sites, which made this work possible. In particular, we acknowledge James Trager (Shaw Nature Reserve) and Eric Stevens (EarthDance) who provided helpful insight to learning about the biota of their respective sites. We thank the Missouri Department of Natural Resources for granting us permission to sample at Castlewood State Park. We thank Patricia Parker, Kenneth Olsen, and various members of ASD’s laboratory for their valuable feedback and discussion on this project.
Funding
This work was funded by grants to MWA from the Whitney R. Harris World Ecology Center, Trans World Airlines, and the Webster Groves Nature Study Society. MWA was partially funded by the Raven Fellowship (spring 2019) and the Graduate Dissertation Fellowship (fall 2019, spring 2020) through the University of Missouri—St. Louis.
Author information
Authors and Affiliations
Contributions
M.W.A. conceptualized the study, acquired funding, wrote the main manuscript text, and performed field work and molecular analysis. A.D.T. and J.P.S. performed molecular analysis. A.S.D. provided supervision for the study. All authors edited and reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Austin, M.W., Tripodi, A.D., Strange, J.P. et al. Bumble bees exhibit body size clines across an urban gradient despite low genetic differentiation. Sci Rep 12, 4166 (2022). https://doi.org/10.1038/s41598-022-08093-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-08093-4
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.