Abstract
Colorectal cancer (CRC) is one of the most prevalent cancers worldwide, which after breast, lung and, prostate cancers, is the fourth prevalent cancer in the United States. Long non-coding RNAs (lncRNAs) have an essential role in the pathogenesis of CRC. Therefore, bioinformatics studies on lncRNAs and their target genes have potential importance as novel biomarkers. In the current study, publicly available microarray gene expression data of colorectal cancer (GSE106582) was analyzed with the Limma, Geoquery, Biobase package. Afterward, identified differentially expressed lncRNAs and their target genes were inserted into Weighted correlation network analysis (WGCNA) to obtain modules and hub genes. A total of nine differentially expressed lncRNAs (LINC01018, ITCH-IT, ITPK1-AS1, FOXP1-IT1, FAM238B, PAXIP1-AS1, ATP2B1-AS1, MIR29B2CHG, and SNHG32) were identified using microarray data analysis. The WGCNA has identified several hub genes for black (LMOD3, CDKN2AIPNL, EXO5, ZNF69, BMS1P5, METTL21A, IL17RD, MIGA1, CEP19, FKBP14), blue (CLCA1, GUCA2A, UGT2B17, DSC2, CA1, AQP8, ITLN1, BEST4, KLF4, IQCF6) and turquoise (PAFAH1B1, LMNB1, CACYBP, GLO1, PUM3, POC1A, ASF1B, SDCCAG3, ASNS, PDCD2L) modules. The findings of the current study will help to improve our understanding of CRC. Moreover, the hub genes that we have identified could be considered as possible prognostic/diagnostic biomarkers. This study led to the determination of nine lncRNAs with no previous association with CRC development.
Similar content being viewed by others
Introduction
Colorectal cancer (CRC) is one of the most prevalent digestive system cancers worldwide. After breast, lung, and prostate cancers, it is the fourth prevalent cancer in the United States1. Nearly two million new CRC cases and 700,000 deaths from this cancer are reported every year2. Its incidence and mortality rate is 25% higher in men than women, and most of the reported CRC cases are in the colon, while fewer cases are in the rectum3. Thanks to novel diagnostic tools and therapeutic interventions, the mortality rate has decreased significantly from 1999 to 20154. Although most reported cases of CRC are sporadic (70%), a significant part of the cases occur in patients with familial history of CRC (25%) or hereditary colorectal cancer syndromes (~ 10%)5. Therefore, further studies are still required to clarify the involved molecular mechanisms in CRC.
Non-coding RNAs (ncRNAs) are small molecules that are not translated to proteins and participate in various regulatory functions of the cell. They consist of different families including, microRNAs (miRNAs), small nuclear RNAs (snRNAs), PIWI-interacting RNAs (piRNAs), and long non-coding RNAs (lncRNAs)6. LncRNAs were recently identified as regulatory molecules with a length of more than 200 nucleotides. They bear some resemblances with mRNAs, including a cap at the 5′ end, having more than one exon, being transcribed by RNA polymerase II (RNA pol II), and being located in the cytoplasm or the nucleus. However, this class has some dissimilarities with mRNAs, including lower expression level, poorer conservation among other species, inability to be transcribed to a protein, and tissue/stage-specific expression7. It has been reported that lncRNAs may be up/down-regulated in cancerous cells compared to healthy ones, indicating the possible role of these molecules as an oncogene or a tumor suppressor8.
LncRNAs are involved in carcinogenesis and progression of CRC9,10,11. It has been demonstrated that lncRNAs regulate various cellular functions related to CRC pathogenesis, including cell proliferation, apoptosis, migration, invasion, metastasis, differentiation, DNA damage, drug resistance, epithelial-mesenchymal transition (EMT), development, controlling cancer stem cells, and cell cycle12. Thanks to high-throughput methods and novel bioinformatics approaches, such as microarray and RNA-seq, many lncRNAs with altered expression in CRC cells have been identified13. Forrest et al. identified more than 200 differentially expressed lncRNAs by analyzing RNA sequencing data from The Cancer Genome Atlas (TCGA) dataset. Moreover, they concluded these lncRNAs regulate cell cycle genes and increase resistance to apoptosis14. Studies have reported that some lncRNAs are significantly overexpressed in the CRC cells and tissues, correlating with metastasis and weak patient prognosis15. In addition to CRC cells, lncRNAs with altered expression level has been reported in peripheral blood components such as serum or plasma16.
Weighted gene co-expression network analysis (WGCNA) is an in-silico system biology tool to analyze gene expression in a complex network of regulatory genes. This tool based on R programming can identify clusters of highly correlated genes (modules) based on genetic correlations17. Therefore, it is helpful for identifing novel diagnostic and prognostic biomarkers for cancer. Zhou et al. reported a number of hub genes and miRNAs which was associated with stages of CRC18. In the current study, the WGCNA algorithm was employed to construct a co-expression network of lncRNAs associated with CRC and their target genes. This study would help identify possible new biomarkers for CRC and reach a better understanding of the molecular pathways contributing to this disease.
Methods
Data acquisition and processing of lncRNA expression profiles
Microarray gene expression data of colorectal cancer with the series number GSE106582 was obtained from the publicly available Gene Expression Omnibus (GEO) database to identify lncRNA candidates (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=gse106582) .GSE106582 was provided by the University clinics Freiburg (Freiburg, Germany), and CRC patients were recruited at the University Hospital of Heidelberg. Total RNA from 77 tumors and 117 mucosa samples were analyzed, including 68 tumor-mucosa pairs using Illumina HumanHT12v4 gene chips. Next, downloading and reading expression profile data was conducted in R environment using GEOquery package19. Differential expression genes (DEGs) analysis was conveyed via Limma (linear models for microarray data) package20,21.
Weighted gene correlation network analysis and the identification of modules
Considering the fact that the functions of most lncRNAs are unknown, the prediction of their functions principally depends on the examination of their co-expressed genes. Network analysis was conducted using the WGCNA package in R22 to evaluate the relative significance of lncRNAs and their module membership. Briefly, WGCNA was performed on the GSE106582 dataset obtained from 77 CRC and 117 mucosa tissues. To distinguish modules with different expression patterns, a soft threshold power was selected to create co-expression networks. Next, the pearson correlation coefficient was used to evaluate the weighted co-expression relationships in the adjacency matrix. Then, a topological overlap matrix (TOM) similarity function was applied to transform the matrix into a TOM, which was used to estimate the co-expression relationships between genes. The networks were established by merging genes with extremely similar co-expression patterns into modules. Consequently, the module with the key lncRNAs and their co-expressed genes was achieved. The reconstructed co-expression network was visualized using the Cytoscape software (version 3.7.0) and Cytohubba plugin (version 0.1)23.
Functional annotation of the co-expressed genes in the module
Gene Ontology (GO) is a simple technique applied for annotating large numbers of genes to define attributes of gene products in three non-overlapping domains of molecular biology, including Molecular Function (MF), Biological Process (BP), and Cellular Component (CC)24. To identify genes and their corresponding functionalities, the Kyoto Encyclopedia of Genes and Genomes (KEGG) was employed to systematically analyze gene functions (www.kegg.jp/kegg/kegg1.html)25. To determine the potential functions of novel lncRNAs and their associated biological pathways, a functional enrichment analysis of their co-expressed genes was performed using the Fun Rich software (version 3.1.3).
Statistical analysis
GSE106582 downloading and reading expression profile data was conducted in an R environment using the GEOquery package18. Differential expression genes (DEGs) were assessed between CRC and mucosal samples by empirical Bayesian method via t-test. DEGs analysis was conveyed via Limma (linear models for microarray data) package19,20. The cutoff criteria of the adjusted p-value (FDR) < 0.05 and | logFC | ≥ 0.5 were considered the threshold for significance to extract DEGs and DELs among 48.107 Probe sets. The top 2450 genes were evaluated with the critical value of the adjusted P-value < 0.05, and logFC ≥ |0.5|, were selected for further analysis. Subsequently, the DEGs were filtered by 5023 lncRNAs were retrieved from HGNC BioMart (https://biomart.genenames.org/) to detect differentially expressed lncRNAs (DELs).
Results
Identification of LncRNA candidates associated with colorectal cancer
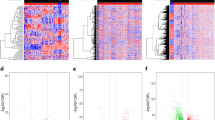
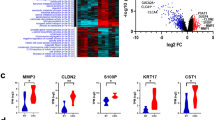
GSE106582 gene expression profiles were selected in this study. This dataset contained 117 normal samples and 77 CRC samples. Following analysis of the dataset with the Limma package, the difference between CRC and normal tissues was presented in volcano plots (Fig. 1). Based on the criteria of adjusted P-value < 0.05 and logFC ≥ |0.5|, a total of 2449 DEGs were screened from GSE106582, including 1170 upregulated genes and 1279 downregulated genes. LncRNA expression data analysis of the GSE106582 dataset resulted in the identification of 32 DELs (Supplementary File 1), among which nine lncRNAs are detected as novel lncRNAs with no previous association with CRC development, including LINC01018, SNHG32, ITCH-IT1, ITPK1-AS1, FOXP1-IT1, FAM238B, PAXIP1-DT, ATP2B1-AS1, and MIR29B2CHG (Table 1). Of these, 8 lncRNAs were down-regulated (p-value < 0.05) and one lncRNA (SNHG32) was up-regulated (p-value < 0.05) in CRC tissues compared to normal tissues.
Identifying differently expressed genes between colorectal cancer and normal tissues. Heatmap of the difference between tumoral and normal samples of the GSE106582 dataset with R. Box-Scatter plot of the expression data of the lncRNAs in tumor tissues vs normal of the GSE106582 dataset for LINC01018.
Construction of weighted gene co-expression network analysis
In this study, a co-expression network was constructed using GSE106582, the expression amounts of 2449 DEGs were analyzed for the co-expression network constructing the “WGCNA” package. Primarily, the outlier cases were displaced, and the hierarchical clustering analysis was accomplished with the “hclust” R function (Supplementary File 2). Meanwhile, the pickSoftThreshold function was used to determine scale independence and mean connectivity analysis of modules with several power values. Afterward, to guarantee a scale-free network, we picked β = 14 as the soft-thresholding power (Fig. 2A), to double-check the scale-free topology R2 with a linear regression plot (scale-free R2 = 0.93) (Fig. 2B). Therefore, β = 14 was selected to produce a hierarchical clustering tree with different colors signifying diverse modules. As demonstrated in Fig. 2C, six co-expressed gene modules were identified with gray modules representing non-co-expressed genes, and each module was marked by a color. The green module is the smaller module with 128 genes. At the same time, the blue module is the largest module with 780 genes. Additionally, the background color is grey and represents the 289 genes not attributed to any module (Table 2). Ultimately, we examined the interactive connections amongst the six modules, plotted the heatmap of the network, and showed the relative independence of each module in Fig. 2D and the multi-dimensional scaling (MDS) plot presented in Fig. 2E.
Network visualization plots. (A) Scale independence and mean connectivity analysis. The proper soft threshold power = 14 was selected. (B) The histogram of connectivity distribution and the scale-free topology panels. (C) Clustering dendrogram of genes, with dissimilarity based on the topological overlap. (D) Heatmap plot to represent the TOM among the genes in different modules. (E) Multidimensional scaling (MDS) plots to describe the entire gene expression network.
Identification of novel lncRNA modules
We investigated the modules of novel lncRNAs to predict their functions through their co-expressed genes and the construction of a regulatory network between lncRNAs and protein-coding genes. We found LINC01018, ITCH-IT, ITPK1-AS1, FOXP1-IT1, FAM238B, and PAXIP1-AS1 in the black module, ATP2B1-AS1, and MIR29B2CHG in the turquoise module, and SNHG32 in the blue module. The list of genes for each module is detailed in Supplementary File 3.
Gene co-expression modules correspond to CRC
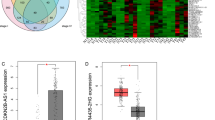
In addition, we examined the associations of gene modules and cancer phenotype, which was based on the correlation between module eigengenes and clinical traits. The results revealed two of the total six gene modules were strongly correlated with tumoral status, including turquoise (R = 0.91, P = 3E−69), and grey (R = 0.96, P = 1E−99), while the grey module is non-co-expressed genes and not considered for further studies (Fig. 3A). In addition, the blue gene modules (R = 0.91, P = 3E−69) negatively correlate with tumoral status in a significant manner. However, other clinical traits, including age and gender, are not correlated with gene modules. Furthermore, the eigengene dendrogram and heatmap were designed to distinguish groups of correlated eigengenes associated with tumoral status. The results re-validated the correlations of the turquoise and blue gene modules with tumoral status (Fig. 3B). Finally, the plots of module membership in different gene modules vs. gene significance determined that the turquoise and blue gene modules have significant correlations with CRC and demonstrate these gene modules are associated with CRC (Fig. 3C, D).
Gene co-expression modules correlated with colorectal cancer. (A) Module–trait relationships. Each cell includes the corresponding correlation and P-value. (B) The eigengene dendrogram and heatmap classify groups of correlated eigengenes. A scatter plot of the gene significance for Tumoral versus the module membership in the Turquoise (C) and Blue (D) modules.
Construction of PPI network
CytoHubba plugin was used to construct interaction networks and identify hub genes in each module. Networks for black, blue, and turquoise modules are depicted in Figs. 4, 5 and 6, respectively. Moreover, the plugin has identified thirty hub genes based on their degrees, ranks, and scores that are summarized in Table 3.
Turquoise module network. The top 10 genes in this module all have the same score and are equally involved in the respective pathways listed in Table 6.
Functional enrichment analysis
The GO enrichment and KEGG pathway analyses were carried out to understand the biological characteristics of all modules. The involved cellular components, molecular functions, biological processes, and biological pathways for each module are summarized in Tables 4, 5 and 6.
Discussion
CRC is a global concern due to its high mortality and morbidity rates. Medical systems worldwide have been endeavored to reduce CRC rate using novel diagnostic and prognostic methods26. However, it is still one of the leading medical burdens over the globe27. High throughput technologies like microarray have been a valuable tool to compare the expression profile of normal and tumor cells. The omitted data has been precious to better understand expression alterations in cancers28. Using tools like WGCNA, we can study the interconnections between genes and obtain differentially expressed genes29.
In recent years, WGCNA has been used to comprehend lncRNAs role in cancers. Giulietti et al. identified eleven lncRNAs using this method as key regulators in pancreatic cancer, which could be used as novel diagnostic/prognostic markers30. Jiang et al. used the WGCNA method and found four lncRNAs associated with the carcinogenesis and progression of colon adenocarcinoma. In the current study, nine differentially expressed lncRNAs (LINC01018, ITCH-IT, ITPK1-AS1, FOXP1-IT1, FAM238B, PAXIP1-AS1, ATP2B1-AS1, MIR29B2CHG, and SNHG32) were identified using microarray data analysis (GSE106582). Afterward, WGCNA was performed on the lncRNAs and their target genes, which resulted in three significant modules. Further bioinformatics studies on hub genes of every module showed that they are involved in concrete pathways and biological processes.
The role of some of identified DELs in cancer pathogenesis has been previously reported in the literature. Miao et al. identified LINC01018 as a prognostic marker for gastric cancer31. It also has a tumor suppressor role in hepatocellular carcinoma that upregulates FOXO1 by sponging miR-182-5p32. In a recent study, Liting Wang et al. (2021) found that LINC01018 / hsa-miR-182-5p / ADH4 were strongly correlated. Moreover, the regulatory axis of ceRNA in the human body, by regulating the expression of key proteins in important signaling pathways can become a checkpoint inhibitor and regulate the incidence of liver cancer33.
In the study of Hu et al., a combination of five lncRNAs, including ITPK1-AS1, was introduced as a useful prognostic marker for gastric cancer. A 2020 study also found that an ITPK1-AS1 anti-sensory ncRNA with 0.56-fold induction was the highest gene regulated by e-cigarettes compared to traditional cigarettes in active bronchial epithelial cells in smokers34,35. It is also a potential prognostic biomarker of colon adenocarcinoma36. Using bioinformatics tools, FOXP1-IT1 and other lncRNAs have been recognized as a useful prognostic marker for colon adenocarcinoma37. A study examining the expression pattern of different lncRNAs induced by TGF β1 predicts that FOXP1-IT1 is highly regulated by RAD21, possibly involved in oncogenic conversion and tumorigenesis in response to DNA repair and induction of genomic instability38. PAXIP1-AS1 is located in the glioma cell nucleus, and its overexpression increases migration, invasion, and angiogenesis of human umbilical vein endothelial cells in glioma39. Zhou et al., in a bioinformatics study, identified MIR29B2CHG as a useful prognostic marker for adrenocortical carcinoma40. A significant decrease of MIR29B2CHG was observed in the triple negative types of breast cancer. It is a host gene for producing of miR-29b2 and miR-29c, which plays a suppressive role in the progression of breast cancer41.
A co-expression network analysis using WGCNA between lncRNAs and their target genes resulted in black, blue, and turquoise modules. Enrichment and functional analysis using Cytoscape plugin Cytohubba indicated that the modules involved fundamental cellular pathways. Results showed black module was involved in the metabolism of RNA, FGF signaling pathway, and regulation of gene expression in beta cells; the blue module was mainly engaged in mesenchymal to epithelial transition, fatty acid, triglyceride, and fatty acid beta-oxidation, and the turquoise model played a critical role in mitotic cell cycle, DNA replication, and mitotic M-M G1 phase. Gene ontology study revealed modules participate in essential biological processes and molecular functions. For example, the genes of black and turquoise modules participate in regulating nucleic acid metabolisms. Moreover, they act in transcription regulation activity and DNA/RNA binding molecular functions. In The blue module, genes are involved in metabolism, energy pathway, and transportation biological processes. In addition, they function in catalytic, transporter, and hormone activities. The genes of black and blue modules are operating in the lysosomes and exosomes. Also, the turquoise module genes are located in the nucleus and mitochondrion.
Based on identified biological pathways, modules and hub genes might have an essential role in developing and malignancy of CRC. In the black module, Otte et al. detected the elevated expression level of several self-renewal and stemness-associated genes in cultures with active FGF2 signaling42. The p38 MAPKs are a family of serine/threonine kinases that mainly respond to external stresses43. They participate in significant cancer progression-related mechanisms, including cell metabolism, invasion, inflammation, and angiogenesis44. Glucagon-like peptide 1 (GLP-1) is secreted from intestinal L-cells and participates in insulin secretion and β-cell growth. It has been suggested that sustained activation of the GLP-1 receptor may indirectly result in colon cancer by hyperinsulinemia45. As other recognized pathways in the black module regulate gene expression in pancreatic beta cells and the synthesis/secretion of Incretin, there may be an association between insulin secretion, colon cancer, and genes in this module which needs to be further studied.
The genes in the blue module participate in the EMT process. The cells of the colon lose their epithelial trait and gain some mesenchymal characteristics that help them migrate to other parts of the body. This process is the main reason for liver metastasis that occurs in CRC patients46. In addition, it has been reported that lipid is required for cancer cells to proliferate. Due to this fact, pathways that participate in lipid synthesis would be proper targets to design therapeutic agents47. As summarized in Table 4, genes in the blue module participate in fatty acid, triacylglycerol, and ketone body metabolism. Fatty acid beta-oxidation is another identified pathway that cancer cells rely on for survival, stemness, metastasis, immune suppression, and drug resistance48. Considering all the above-mentioned discoveries, further study would elucidate the exact role of this module and lipid involvement in cancer.
The cytoHubba plugin has pointed out several hub genes for black (LMOD3, CDKN2AIPNL, EXO5, ZNF69, BMS1P5, METTL21A, IL17RD, MIGA1, CEP19, FKBP14), blue (CLCA1, GUCA2A, UGT2B17, DSC2, CA1, AQP8, ITLN1, BEST4, KLF4, IQCF6) and turquoise (PAFAH1B1, LMNB1, CACYBP, GLO1, PUM3, POC1A, ASF1B, SDCCAG3, ASNS, PDCD2L) modules. The number of identified hub genes has been previously reported to participate in colorectal cancer pathogenesis, downregulation of miR-193a-3p results in upregulation of IL17RD, which promotes colon cancer through inflammation49. Also, this protein promotes cancer by concealing cancer cells from immune surveillance50. As a pro-proliferation factor, Yang et al. indicated FKBP14 was upregulated in CRC tissues, which were associated with the poor prognosis of CRC patients51. CLCA1 has a tumor suppressor role by inhibiting the Wnt/beta-catenin signaling pathway and the EMT process. The study on human CRC samples indicates its expression has been significantly decreased52. Polymorphisms in the UGT2B17 gene have been associated with CRC risk in the Caucasian population53. Dsc2 is the only expressed member of the desmocollins family in the normal colorectal cell. A study on CRC cells shows Dsc2 is switched to Dsc1 and Dsc3 during cancer development54. Overexpression of AQP8 has significantly decreased CRC cell growth and metastasis55. Aleksandrova et al. indicated circulating ITLN1 concentration has been correlated with CRC risk56. KLF4 as a tumor suppressor inhibits colorectal cancer cell growth and is associated with poor overall survival57. A component of the ubiquitin pathway, CACYBP, is overexpressed in CRC patients and has increased cancer proliferation58.
Conclusion
In the current study, with the help of bioinformatics tools, black, blue, and turquoise modules were regarded as the most critical modules in the progression and development of CRC. Moreover, thirty genes were recognized as hub genes that could be possible biomarkers for the diagnosis and prognosis of CRC. In addition, nine lncRNAs including, LINC01018, SNHG32, ITCH-IT1, ITPK1-AS1, FOXP1-IT1, FAM238B, PAXIP1-DT, ATP2B1-AS1, and MIR29B2CHG were identified with no previous association with CRC development which may serve important roles in the pathogenesis of CRC.
Data availability
All data generated or analysed during this study are included in this published article [and its supplementary information files].
References
Siegel, R. L., Miller, K. D. & Jemal, A. Cancer statistics, 2020. CA Cancer J. Clin. 70(1), 7–30 (2020).
Stewart, B. W., Kleihues, P. World cancer report (2003).
Ahmed, M. Colon cancer: a clinician’s perspective in 2019. Gastroenterol. Res. 13(1), 1–10 (2020).
Cronin, K. A. et al. Annual Report to the Nation on the Status of Cancer, part I: National cancer statistics. Cancer 124(13), 2785–2800 (2018).
Hadjipetrou, A., Anyfantakis, D., Galanakis, C. G., Kastanakis, M. & Kastanakis, S. Colorectal cancer, screening and primary care: a mini literature review. World J. Gastroenterol. 23(33), 6049–6058 (2017).
Wright, M. W. & Bruford, E. A. Naming “junk”: human non-protein coding RNA (ncRNA) gene nomenclature. Hum. Genomics 5(2), 90–98 (2011).
Cruz-Miranda, G. M. et al. Long non-coding RNA and acute leukemia. Int. J. Mol. Sci. 20(3), 735 (2019).
Qi, P. & Du, X. The long non-coding RNAs, a new cancer diagnostic and therapeutic gold mine. Mod. Pathol. 26(2), 155–165 (2013).
Okayama, H., Schetter, A. J. & Harris, C. C. MicroRNAs and inflammation in the pathogenesis and progression of colon cancer. Dig. Dis. 30(suppl 2), 9–15 (2012).
Tang, X. et al. Regulation mechanism of long noncoding RNAs in colon cancer development and progression. Yonsei Med. J. 60(4), 319–325 (2019).
Carlevaro-Fita, J. et al. Cancer LncRNA census reveals evidence for deep functional conservation of long noncoding RNAs in tumorigenesis. Commun. Biol. 3(1), 56 (2020).
Chen, S. & Shen, X. Long noncoding RNAs: functions and mechanisms in colon cancer. Mol. Cancer 19(1), 167 (2020).
Wang, J. et al. Regulatory roles of non-coding RNAs in colorectal cancer. Int. J. Mol. Sci. 16(8), 19886–19919 (2015).
Forrest, M. E. et al. Colon cancer-upregulated long non-coding RNA lincDUSP regulates cell cycle genes and potentiates resistance to apoptosis. Sci. Rep. 8(1), 7324 (2018).
Kam, Y. et al. Detection of a long non-coding RNA (CCAT1) in living cells and human adenocarcinoma of colon tissues using FIT-PNA molecular beacons. Cancer Lett. 352(1), 90–96 (2014).
Xie, X. et al. Long non-coding RNAs in colorectal cancer. Oncotarget 7(5), 5226–5239 (2016).
Langfelder, P. & Horvath, S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinform. 9, 559 (2008).
Zhou, X. G. et al. Identifying miRNA and gene modules of colon cancer associated with pathological stage by weighted gene co-expression network analysis. OncoTargets Ther. 11, 2815–2830 (2018).
Davis, S. & Meltzer, P. S. GEOquery: a bridge between the Gene Expression Omnibus (GEO) and bioconductor. Bioinformatics 23(14), 1846–1847 (2007).
Smyth, G. K. Limma: linear models for microarray data. In Bioinformatics and Computational Biology Solutions Using R and Bioconductor (eds Gentleman, R. et al.) 397–420 (Springer, 2005).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucl. Acids Res. 43(7), e47 (2015).
Langfelder, P. & Horvath, S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinform. 9(1), 559 (2008).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13(11), 2498–2504 (2003).
Harris, M. et al. The Gene Oncology (GO) database and informatics resource. Nucl. Acids Res. 32, 258–261 (2004).
Ogata, H. et al. KEGG: Kyoto Encyclopedia of Genes and Genomes. Nucl. Acids Res. 27(1), 29–34 (1999).
Siegel, R. L., Miller, K. D. & Jemal, A. Cancer statistics, 2015. CA Cancer J. Clin. 65(1), 5–29 (2015).
Schmoll, H. J. et al. ESMO Consensus Guidelines for management of patients with colon and rectal cancer. A personalized approach to clinical decision making. Ann. Oncol. Off. J. Eur. Soc. Med. Oncol. 23(10), 2479–2516 (2012).
Choi, J. K., Yu, U., Yoo, O. J. & Kim, S. Differential coexpression analysis using microarray data and its application to human cancer. Bioinformatics (Oxford, England) 21(24), 4348–4355 (2005).
Qiu, X. et al. Weighted gene co-expression network analysis identified MYL9 and CNN1 are associated with recurrence in colorectal cancer. J. Cancer 11(8), 2348–2359 (2020).
Giulietti, M., Righetti, A., Principato, G. & Piva, F. LncRNA co-expression network analysis reveals novel biomarkers for pancreatic cancer. Carcinogenesis 39(8), 1016–1025 (2018).
Miao, Y. et al. Comprehensive analysis of a novel four-lncRNA signature as a prognostic biomarker for human gastric cancer. Oncotarget 8(43), 75007–75024 (2017).
Wang, S. et al. LINC01018 confers a novel tumor suppressor role in hepatocellular carcinoma through sponging microRNA-182-5p. Am. J. Physiol. Gastrointest. Liver Physiol. 317(2), G116–G126 (2019).
Wang, L., Chen, Z. Y., Wang, X. L., Zeng, Q. L., Jiang, S. L. & Li, X. L. Linc01018/hsa-mir-182-5p/ADH4 axis study on the role and mechanism in the occurrence and development of liver cancer (2021).
Hu, Z. et al. Five-long non-coding RNA risk score system for the effective prediction of gastric cancer patient survival. Oncol. Lett. 17(5), 4474–4486 (2019).
Herr, C. et al. Cigarette smoke and electronic cigarettes differentially activate bronchial epithelial cells. Respir. Res. 21(1), 1–13 (2020).
Wang, W.-J. et al. A competing endogenous RNA network reveals novel potential lncRNA, miRNA, and mRNA biomarkers in the prognosis of human colon adenocarcinoma. J. Surg. Res. 235, 22–33 (2019).
Jin, L., Li, C., Liu, T. & Wang, L. A potential prognostic prediction model of colon adenocarcinoma with recurrence based on prognostic lncRNA signatures. Hum. Genomics 14(1), 24 (2020).
Hu, W. et al. Microarray profiling of TGF-β1-induced long non-coding RNA expression patterns in human lung bronchial epithelial BEAS-2B cells. Cell. Physiol. Biochem. 50(6), 2071–2085 (2018).
Xu, H. et al. Long non-coding RNA PAXIP1-AS1 facilitates cell invasion and angiogenesis of glioma by recruiting transcription factor ETS1 to upregulate KIF14 expression. J. Exp. Clin. Cancer Res. CR 38(1), 486 (2019).
Zhou, Y. et al. Bioinformatic analysis of long non-coding RNA-associated competing endogenous RNA network in adrenocortical carcinoma. Transl. Cancer Res. 8(5), 2175–2186 (2019).
Shafaroudi, A. M. et al. Expression and function of C1orf132 long-noncoding RNA in breast cancer cell lines and tissues. Int. J. Mol. Sci. 22(13), 6768 (2021).
Otte, J. et al. FGF signalling in the self-renewal of colon cancer organoids. Sci. Rep. 9(1), 17365 (2019).
Rouse, J. et al. A novel kinase cascade triggered by stress and heat shock that stimulates MAPKAP kinase-2 and phosphorylation of the small heat shock proteins. Cell 78(6), 1027–1037 (1994).
Grossi, V., Peserico, A., Tezil, T. & Simone, C. p38α MAPK pathway: a key factor in colorectal cancer therapy and chemoresistance. World J. Gastroenterol. 20(29), 9744–9758 (2014).
Gallagher, E. J. & LeRoith, D. The proliferating role of insulin and insulin-like growth factors in cancer. Trends Endocrinol. Metab. TEM. 21(10), 610–618 (2010).
Xu, H. et al. The mechanisms of colorectal cancer cell mesenchymal–epithelial transition induced by hepatocyte exosome-derived miR-203a-3p. J. Bio-X Res. 1(2), 62–72 (2018).
Currie, E., Schulze, A., Zechner, R., Walther, T. C. & Farese, R. V. Jr. Cellular fatty acid metabolism and cancer. Cell Metab. 18(2), 153–161 (2013).
Ma, Y. et al. Fatty acid oxidation: an emerging facet of metabolic transformation in cancer. Cancer Lett. 435, 92–100 (2018).
Pekow, J. et al. miR-193a-3p is a key tumor suppressor in ulcerative colitis-associated colon cancer and promotes carcinogenesis through upregulation of IL17RD. Clin. Cancer Res. Off. J. Am. Assoc. Cancer Res. 23(17), 5281–5291 (2017).
Wu, D. et al. Interleukin-17: a promoter in colorectal cancer progression. Clin. Dev. Immunol. 2013, 436307 (2013).
Yang, J. et al. Targeting PI3K in cancer: mechanisms and advances in clinical trials. Mol. Cancer 18(1), 1–28 (2019).
Li, X. et al. CLCA1 suppresses colorectal cancer aggressiveness via inhibition of the Wnt/beta-catenin signaling pathway. Cell Commun. Signal. CCS. 15(1), 38 (2017).
Angstadt, A. Y. et al. The effect of copy number variation in the phase II detoxification genes UGT2B17 and UGT2B28 on colorectal cancer risk. Cancer 119(13), 2477–2485 (2013).
Khan, K. et al. Desmocollin switching in colorectal cancer. Br. J. Cancer 95(10), 1367–1370 (2006).
Wu, D. Q. et al. AQP8 inhibits colorectal cancer growth and metastasis by down-regulating PI3K/AKT signaling and PCDH7 expression. Am. J. Cancer Res. 8(2), 266–279 (2018).
Aleksandrova, K. et al. Circulating omentin as a novel biomarker for colorectal cancer risk: data from the EPIC-potsdam cohort study. Cancer Res. 76(13), 3862–3871 (2016).
Ma, Y. et al. KLF4 inhibits colorectal cancer cell proliferation dependent on NDRG2 signaling. Oncol. Rep. 38(2), 975–984 (2017).
Zhai, H. et al. CacyBP/SIP promotes the proliferation of colon cancer cells. PLoS ONE 12(2), e0169959 (2017).
Funding
Dr Sara Shamdani was supported by a grant via “la Fondation de la Maison de la Chimie” for the project R20150LL with the Grant Number: RAK20150LLA.
Author information
Authors and Affiliations
Contributions
S.C.K, S.R and S.N conceived the study. S.C.K, S.R and S.SH performed the experiments and analyzed the data. S.C.K, S.R, S.SH, G.U and S.N wrote the manuscript; G.U, S.SH and S.N critically reviewed the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
Dr. Sina NASERIAN is the deputy Chief Scientific and Technical Officer at ScreenCell Company. Dr. Sara SHAMDANI is the General Director of the CellMedEx Company. The rest of the authors declare that they have no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chodary Khameneh, S., Razi, S., Shamdani, S. et al. Weighted correlation network analysis revealed novel long non-coding RNAs for colorectal cancer. Sci Rep 12, 2990 (2022). https://doi.org/10.1038/s41598-022-06934-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-06934-w
This article is cited by
-
Regulation of overexpression lncRNA ATP2B1-AS1 on lung adenocarcinoma progression
Journal of Cardiothoracic Surgery (2024)
-
MiR-380 inhibits the proliferation and invasion of cholangiocarcinoma cells by silencing LIS1
Cancer Cell International (2024)
-
Weighted gene co-expression network analysis for hub genes in colorectal cancer
Pharmacological Reports (2024)
-
Identification of genes related to ribosomal proteins in colorectal cancer: exploring their potential as biomarkers, prognostic indicators, and therapeutic targets
Molecular Biology Reports (2024)
-
Identification and functional analysis of LncRNA-XIST ceRNA network in prostate cancer
BMC Cancer (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.