Abstract
The proto-oncogene MYC is important for development and cell growth, however, its abnormal regulation causes cancer. Recent studies identified distinct enhancers of MYC in various cancers, but any MYC enhancer(s) in hepatocellular carcinoma (HCC) remain(s) elusive. By analyzing H3K27ac enrichment and enhancer RNA (eRNA) expression in cultured HCC cells, we identified six putative MYC enhancer regions. Amongst these, two highly active enhancers, located ~ 800 kb downstream of the MYC gene, were identified by qRT-PCR and reporter assays. We functionally confirmed these enhancers by demonstrating a significantly reduced MYC expression and cell proliferation upon CRISPR/Cas9-based deletion and/or antisense oligonucleotide (ASO)-mediated inhibition. In conclusion, we identified potential MYC enhancers of HCC and propose that the associated eRNAs may be suitable targets for HCC treatment.
Similar content being viewed by others
Introduction
The transcription factor (TF) MYC is essential for various cellular processes, including cell growth, proliferation, differentiation, and apoptosis1. MYC tightly regulates the normal state, but dysregulation of MYC is prevalent in cancer. MYC expression is upregulated in 50–60% of all cancers2,3. Accordingly, studies on the enhancers of MYC were conducted, and it was found that their location is cancer-specific4. For example, the MYC enhancer is located approximately 0.7 Mb downstream of the gene in prostate cancer, 70 kb upstream in pancreatic cancer, and 1.9 Mb upstream in glioma5,6,7. However, studies on MYC enhancers in HCC have been lacking.
HCC is the most common type of primary liver cancer. It is prevalent globally and a leading cause of cancer-related death8,9. Significant genetic and epigenetic alterations exist in HCC. Their accumulation in key genes involved in cell survival, proliferation, apoptosis, and metastasis leads to carcinogenesis10. The dysregulation of MYC plays a vital role in proliferation and invasion, including tumor initiation and progression in HCC11,12.
Generally, both genetic and epigenetic changes could lead to altered MYC expression13. In the present study, we started with an epigenetic approach. Based on published genome maps featuring various epigenetic parameters, we identified novel MYC enhancers in HCC cell lines and then confirmed them with functional, genetic, and gene expression analyses. Specifically, we demonstrate that, at least in part, these novel enhancers operate through the expression of eRNAs.
eRNAs are noncoding transcripts generated by most, if not all, active enhancers and have been shown to play a central role in regulating gene expression14. They are often expressed cell type-specifically to control cell fate, and in cancer, they can be used as new diagnostic markers and drug targets15,16. The present study suggests that the same approach may also be applicable to HCC.
Results
Effects of BET inhibition on HCC cell proliferation and MYC expression
The drugs, JQ1 and OTX015, inhibit the Bromodomain and Extra-Terminal motif (BET) protein family member BRD4 that is required to maintain super-enhancer (SE) activation17 by interacting with TFs and chromatin remodeling proteins18. In addition, BET inhibitors interfere with BRD4-associated eRNA elongation19. Therefore, to study the effects of enhancer inhibition on HCC cells, we incubated the Huh7 and HepG2 cell lines for up to 72 h with JQ1 or OTX015. Compared to controls, the numbers of Huh7 and HepG2 cells were significantly reduced after 48 h and 24 h, respectively, of inhibitor treatment (Fig. 1A). In both cell lines, inhibition for 24 h significantly decreased the proportion of 5-ethynyl-2’-deoxyuridine (EdU)-incorporating cells, indicating reduced proliferation (Fig. 1B). Next, in the HepG2 cells, we determined the effect of the BET inhibitors on the mRNA levels of MYC and VEGFA (encoding vascular endothelial growth factor A). The expression of VEGF is elevated in various cancers, including HCC, and is one of the targets of BET inhibitors20,21,22. Each inhibitor reduced both mRNA levels to approximately 70% (Fig. 1C). Furthermore, we measured the expression of these genes in response to two transcription inhibitors, p300/cAMP response element-binding (CREB)-binding protein (CBP) inhibitor C646 (50 µM, 24 h) and RNAPII transcription elongation inhibitor 5,6-dichloro-1-β-d-ribofuranosylbenzimidazole (DRB) (50 µM, 24 h). C646 treatment significantly reduced the expression of MYC but not of VEGFA, while DRB significantly reduced the expression of both genes (Fig. 1D).
BET inhibitor suppresses cell viability, cell proliferation, MYC expression, and MYC target gene expression in Huh7 and HepG2 cells. (A) Cell viability was determined by the WST1 assay and represented by the relative absorbance at 450 nm. Huh7 and HepG2 cells were treated with 5 µM BET inhibitors (JQ1 or OTX015) for different durations (4 h, 24 h, 48 h, and 72 h). (B) The proliferation of Huh7 and HepG2 cells treated with 5 µM BET inhibitors for 24 h was assessed by EdU assay. Representative images and the number of EdU-positive cells (%) are shown. Original magnification, × 400. The data represent three biologically independent experiments. **p < 0.01. (C) qRT-PCR analysis of MYC mRNA and MYC target genes and VEGFA mRNA in BET inhibitor-treated HepG2 cells (5 µM, 24 h). (D) qRT-PCR analysis of MYC mRNA and MYC target genes, VEGFA mRNA in p300/CBP inhibitor, C646 or RNA polymerase II (RNAPII) transcription elongation inhibitor, DRB-treated HepG2 cells (50 µM, 6 h). The values are presented as the mean ± SD from triplicate well measurements. *p < 0.05 and **p < 0.01.
Thus, proliferation of the cultured HCC cells was inhibited by the BET inhibitors along with a reduced MYC and VEGFA expression. Furthermore, the effect of BET inhibitors on reducing MYC expression was closely related to RNA transcription inhibition.
Identification of putative MYC enhancers in HCC cell lines
Next, we examined ENCODE ChIP-seq and global run-on sequencing (GRO-seq) data of the HepG2 cell line in order to localize the potential MYC enhancers (Fig. 2). We found that the downstream regions of MYC were more enriched for H3K27ac, a histone mark for an active enhancer. Using the HepG2 cells GRO-seq peaks (GSE92375)23,24, H3K27ac ChIP-seq peaks (GSE29611)25, and p300 ChIP-seq (GSE32465)26 at the UCSC Genome Browser (http://genome.ucsc.edu), we identified six putative enhancer loci of the MYC gene (R1-6) (Fig. 2A, Supplementary Table S4). We did not observe the same peaks in iPSC-induced hepatocyte-like cells (GSM3271003), primary hepatocytes (GSM3271012)27, and liver tissue (GSM1112809)28 (Fig. 2A).
Identification of MYC enhancer in HepG2 cells. (A) USCS genome browser view of the GRO-seq peak, H3K27ac enrichment, and p300 binding sites along the MYC locus. Potential MYC enhancer is indicated + 500 kb downstream (red boxes). An enlarged display of H3K27ac-enriched reads aligned to the putative enhancer regions (GRCh38/Chr 8: 127,712,576–128,633,744). Six putative enhancer regions (R1, R2, R3, R4, R5, and R6) located downstream of the MYC gene are numbered. Hepatocyte-like cells, HLCs; primary human hepatocytes, PHHs; lung adenocarcinoma, LUAD. (B) Luciferase assay was used to identify the regions of active enhancers for MYC expression. These six putative enhancer regions were cloned upstream of the firefly luciferase reporter (Luc2 gene). (C) The fragments of the R2 region were placed upstream of a luciferase reporter. (D) Fragments of the R3 region were placed upstream of a luciferase reporter. For each transfection, the firefly luciferase activity was normalized to that of the Renilla reniformis luciferase activity. The data are normalized to the pGL4.26 construct. The data represent three independent experiments. **p < 0.01.
To seek direct functional evidence for the suspected enhancer activities in HCC cell lines, we examined the activity of a luciferase reporter in transiently transfected Huh7 cells. Each enhancer region was cloned into the minimal promoter vector pGL4.26 immediately upstream of the luciferase gene. For comparison, we included lung adenocarcinoma (LUAD)-R3 and LUAD-R4, two SE of MYC known to exhibit high and low activity, respectively, in lung adenocarcinoma cells29. As expected, transfection of R2 and R3 increased luciferase activity by approximately tenfold. In contrast, LUAD-R3 and LUAD-R4 did not show an enhanced activity (Fig. 2B). Next, we analyzed 500 bp fragments from within the R2 and R3 regions. Of the R2 fragments, the R2-3-containing plasmid showed the highest enhancer activity (Fig. 2C), while of the R3 fragments, R3-2- and R3-3-containing plasmid were most active (Fig. 2D). These results suggest R2 (R2-3) and R3 (R3-2 and R3-3) as candidate regulators of the transcriptional activation of MYC in HCC cells.
eRNA of putative MYC enhancers in HCC cells
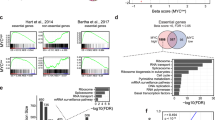
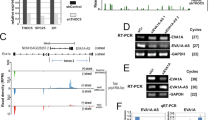
Next, we analyzed the R2 and R3 regions of the HepG2 cells for eRNA expression using qRT-PCR (Supplementary Table S1). Based on the GRO-seq data, six different sets of primers were designed (Fig. 3A). We found that the RNAPII transcription elongation inhibitor DRB caused a significant reduction in sense eRNA expression in regions R2, R3, R4, and R6 but not R1 and R5 (Fig. 3B). R2 and R3 were, therefore, further studied for expression changes through treatment with BET inhibitors. As expected, eRNA expression in regions R2 and R3 was significantly decreased (Fig. 3C). Additionally, R1, R4, R5, and R6 eRNAs were decreased in BET inhibitor-treated HCC cells (Fig. S1). Together, these results indicate that there is a correlation between the activity of enhancers and eRNA transcription. From this, we hypothesized that the BET inhibitors suppressed MYC expression through the regulation of eRNA expression.
The eRNA expression by MYC enhancer regions in HepG2 cells. (A) USCS genome browser view of the GRO-seq peak, H3K27ac enrichment, and p300 binding sites along the MYC locus of HepG2 cells. An enlarged display of GRO-seq peak (+ strand and − strand) and H3K27ac enriched reads aligned to the putative enhancer regions (GRCh38/Chr 8: 128,158,808–128,626,390). eRNA primers were designed for six putative enhancer regions (R1, R2, R3, R4, R5, and R6). R1-S, R2-S, etc., stand for the sense strand eRNAs of the R1, R2, etc. regions. The red line above the GRO-seq (+) strand indicates target eRNA. (B) qRT-PCR of eRNA transcription levels in HepG2 cells treated with different concentrations of DRB (10 µM, 25 µM, and 50 µM; 6 h). (C) qRT-PCR of eRNA transcription levels in BET inhibitor-treated HepG2 cells (5 µM, 24 h). The values are the mean ± SD from triplicate well measurements. *p < 0.05 and **p < 0.01.
Disruption of MYC enhancers affects MYC-related gene expression and cell growth in HCC cells
Since our eRNA expression experiments showed enhancer activity of R3, we tested whether its deletion affects MYC gene expression. We performed the deletion in the Huh7 cell line using the CRISPR/Cas9 system. The targeted sequences are located on chromosome 8 (Chr 8: 128,556,059–128,557,653) downstream of the MYC gene (Supplementary Table S3). After plating the transfected suspension at limiting dilution density, genomic PCR revealed a deletion of the R3 region in one of the wells showing growth. DNA sequencing revealed a 357 bp deletion on Chr 8:128,556,457–128,556,814 (Fig. 4A). Agarose gel electrophoresis confirmed the R3 region deletion (Fig. S2). Note that we did not rule out that due to either delayed Cas9 activity or contamination with wild type cells, the expanded gene-manipulated cultures (which showed a reduced growth rate) used for analysis were not genetically homogeneous. Hence, the effects of R3 deletion reported below are minimum estimates.
Generation of MYC enhancer region-deleted Huh7 cells via the CRISPR/Cas9 system. (A) Schematic illustration of the MYC enhancer locus structure and the wild-type (WT) and mutant (ΔR3) allele sequences around the R3 target region of Huh7 cells. (B) qRT-PCR results showing the levels of MYC mRNA, (C) R2 and R3-associated eRNAs in WT and R3-edited Huh7 cells. (D) TAD of MYC from Hi-C matrix of heatmap on chr8:126,840,000–130,080,000 in HepG2 cells (http://3dgenome.fsm.northwestern.edu/view.php). (E) qRT-PCR results showing the levels of MYC-related lncRNA and (F) MYC-related gene mRNAs in WT and R3-edited Huh7 cells. The values are the mean ± SD from triplicate well measurements. *p < 0.05 and **p < 0.01. (G) Cell proliferation was determined using a WST-1 assay and represented by the relative absorbance at 450 nm. WT and edited Huh7 cells were cultured in a growth medium for 96 h. The obtained absorbance was normalized to each 0 h absorbance. The data represent three biologically independent experiments. **p < 0.01. (H) Colony formation ability of WT and R3-edited Huh7 cells. Cells were grown for 10 days and stained with Crystal Violet. The relative colony formation efficiency was measured as a percentage of the area covered by the colonies. The data represent three biologically independent experiments. **p < 0.01. (I) WT and R3-edited Huh7 cells were cultured in cancer stem cell (CSC) growth media for a spheroid formation assay under ultralow adherence conditions. Cells grown for 7 days and 14 days in spheroid-forming conditions are shown in bright-field images taken with a 4X objective. The number of large spheres (over 100 µm) was counted; scale bar = 100 µm. The data represent three biologically independent experiments. **p < 0.01.
The R3-deleted cells showed reduced MYC gene expression relative to wild-type cells (Fig. 4B). Using qRT-PCR analysis, we found that R2 eRNA (R2-S) and R3 eRNA (R3-S) expression were significantly decreased (Fig. 4C), and so was the expression of lncRNA CCAT1 (Chr 8: 127,207,382–127,219,268), a gene that is not part of the same TAD (Topologically associated domains) as MYC but is known to be regulated by MYC in HCC30. By contrast, the expression of the neighboring lncRNA gene, PVT1, which is in the same TAD as MYC, was unaffected (Chr 8: 127,795,799–128,101,256). Similarly, FAM49B, which is not in the same TAD, was scarcely affected (Chr 8: 129,841,470–130,016,672)31 (Fig. 4D,E). We then analyzed the expression of two more MYC-related genes, IRF2 and TERT, which are known to be repressed and activated by MYC, respectively32,33. In agreement, we found that IRF2 was upregulated in R3-deleted cells; however, TERT was downregulated (Fig. 4F).
Next, we performed cell proliferation and colony formation assays. The R3-deleted cells exhibited a decreased proliferation and colony formation ability (Fig. 4G,H). Similarly, the R3 deletion reduced the spheroid formation by Huh7 cells (Fig. 4I).
It is known that MYC deletion is incompatible with proliferation34. It was therefore not surprising that our attempts to delete R2 and R3 did not result in well growing homogeneous cell lines. In fact, we could not recover R2-deleted cell cultures, suggesting a strong positive effect of R2 on MYC expression (see also further below the section on ASO inhibition). In contrast, we obtained passagable cultures that showed at least a significant proportion of R3-deleted cells (Fig. S2). These cultures exhibited a reduction of both the expression of MYC (Fig. 4B) and of their proliferation (Fig. 4G).
These results confirm that enhancer deletion influences cancer cell growth by reducing MYC expression35.
Inhibition of MYC eRNA causes an effect equivalent to MYC enhancer disruption
To further confirm the functional role of MYC eRNA, the antisense oligonucleotides, ASO R2-P1 and ASO R3-P2, were designed to bind the sense eRNAs at MYC-R2 (R2-S) and MYC-R3 (R3-S), respectively (Fig. 5A; Supplementary Table S1), and transfected into Huh7 cells. As expected, both eRNAs were specifically decreased by the corresponding ASOs (Fig. 5B,C). Importantly, both transfections also resulted in a significantly reduced MYC gene expression relative to the control (ASO NC) (Fig. 5D). Furthermore, when R2-S was targeted, the expression levels of the lncRNAs, PVT1, CCAT1, and FAM49B, were altered similarly as in the MYC-R3 deletion (Fig. 5E, compare with Fig. 4E). In addition, the expression of IRF2 and ICAM1, another gene known to be repressed by MYC36, was increased by both ASOs (Fig. 5F, also compared with Fig. 4F).We next analyzed the proliferation and spheroid-forming abilities of the ASO-transfected Huh7 cells. Both ASO R2-P1 and ASO R3-P2 significantly reduced cell proliferation compared to ASO NC (Fig. 5G), again mimicking the R3 deletion (compare with Fig. 4G). In addition, the spheroid-forming assay showed that inhibition of R2- and R3-induced eRNAs by ASO treatment negatively affected growth (Fig. 5H). These results indicate that inhibition of eRNA can mimic a direct deletion of the corresponding chromosomal DNA (compare with Fig. 4).
Effects of ASOs targeting MYC eRNAs in Huh7 cells. (A) Schematic illustration of the MYC enhancer locus structure and USCS genome browser view of the GRO-seq peak, H3K27ac enrichment, and p300 binding sites along the R2 and R3 regions. The targeting ASOs, ASO R2-P1 and ASO R3-P2, were designed to bind to the R2-S and R3-S eRNAs, respectively. (B,C) Huh7 cells were treated with 125 pmol of a non-targeting ASO (ASO NC) or the targeting ASO. Relative quantitation of R2-S (B) and R3-S (C) eRNA expression levels was performed by qRT-PCR. U6 stands for the control gene RNU6-1. (D–F) Relative quantitation of MYC mRNA expression levels (D) and MYC-controlled mRNA expression levels (E,F). The values are the mean ± SD from triplicate well measurements. *p < 0.05 and **p < 0.01. (G) Cell proliferation was determined using a WST-1 assay and represented by the relative absorbance at 450 nm. Absorbance was measured every 24 h from 0 to 96 h after transfection of ASO. The data represent three biologically independent experiments. **p < 0.01. (H) ASO-transfected Huh7 cells were cultured in a growth medium for the spheroid formation assay. Cells grown for 7 days and 14 days in spheroid-forming conditions are shown in bright-field images taken with a 10X objective. The number of large spheres (over 100 µm) was counted; scale bar = 100 µm. The data represent three biologically independent experiments. **p < 0.01.
Discussion
The enhancer is a DNA region in which the E1A-binding proteins p300 (p300), RNAPII, and TFs are enriched through increased accessibility caused by histone modification. Enhancers help transcriptional activation directly through looping or indirectly affect transcription by expressing eRNAs37,38. Previous studies have revealed the presence of cancer-specific enhancers, the expression of eRNA in cancer cells, and the associated abnormal expression of oncogenes39,40. Since lineage-specific TFs affect the tumorigenesis function of their enhancers, they can also be cancer type-specific41. Therefore, we used ChIP-seq data from HepG2 cells as well as GRO-seq data that included nascent transcripts to identify eRNA expression, an active enhancer marker42,43. Based on these data, we found that the specific MYC enhancers R2 and R3 in HCC cell lines are significantly different from those in other cancers in terms of their location and activity29,39,44. In acute myeloid leukemia (AML), MYC expression is regulated by a SE that consists of five distinct small enhancers located 1.7 Mb downstream of the MYC promoter40. In colorectal cancer (CRC) and prostate cancer, the MYC enhancer is located 335 kb upstream of MYC5. In addition, the Zhang group deleted approximately 1.5 kb of the MYC enhancer located 450 kb downstream of the 3′ end in lung adenocarcinoma cells. MYC expression was reduced by 70%, and clonogenic growth was inhibited by approximately 50%. Note that we epigenetically identified the novel MYC enhancers in HepG2 cells (which are derived from a 15 years-old patient)45 but functionally confirmed these enhancers in Huh7 cells (which are derived from a 57 years-old patient)46. This coincidence strongly suggests the general significance of these newly found enhancers in hepatocyte tumorigenesis. It will be interesting to see whether the same enhancers may also be active in normal proliferating liver cells (such as in embryogenesis or liver regeneration).
eRNAs are generally upregulated in various cancers compared to normal tissues, and they can be used as pan-cancer diagnostic markers47. In addition, tissue-specific highly expressed eRNAs, such as CCAT1 in colorectal cancer and androgen receptor (AR)-induced Kallikrein-related peptidase 3 (KLK3) eRNA (KLK3e) in prostate cancer, are considered new targets for treating various cancers14,48,49. Although the function of eRNAs has not been fully elucidated, eRNA depletion reduces the transcription of target genes by affecting alterations in chromatin structure and contributing to transcriptional initiation of target genes49. eRNA transcription can be regulated by inhibiting enhancer activity or effectively targeting ASO to control target gene expression and cancer cell progression50,51.
Previous studies have found that the inhibition of MYC-related and other eRNAs by using ASOs can effectively inhibit tumor cell progression, suggesting that eRNAs can be helpful as therapeutic targets51,52,53. For example, Epstein-Barr virus (EBV) super-enhancer (ESE) RNAs facilitated the expression of the MYC oncogene in lymphoma, and targeting ESE eRNA showed a therapeutic effect on EBV-related malignancies53. Similarly, in our results, targeting MYC-IEANC RNAs transcribed from MYC enhancers R2 and R3 in Huh7 cells caused an effective decrease in MYC expression along with reduction of proliferation and spheroid formation, suggesting a therapeutic effect on HCC. A previous study demonstrating that downregulation of MYC suppressed spheroid growth of colon CSCs and tumor growth in vivo54, lends support to our suggestion. Of note, the ASO-mediated inhibition of MYC eRNA expression may be superior compared to an alternative strategy, in which BRD4 inhibitors also suppressed tumor cell-associated MYC expression5,55,56,57. Unlike BET inhibitors, the selective targeting of MYC eRNAs probably avoids side effects such as toxicity and resistance.
Conclusion
In this study, we identified the putative MYC enhancers of HCC cells. Enhancer activity and eRNA transcription were analyzed to determine the region involved in MYC expression, and it was found that both the deletion of the MYC enhancers and the ASO-mediated inhibition of the corresponding eRNAs suppressed the proliferation and reduced spheroid formation in HCC cell lines. Thus, our study suggests that for HCC, a strategy for reducing MYC expression through specific targeting with ASO has therapeutic potential without the side effects of gene editing or BRD4 inhibition. Future work will need to evaluate the role of the MYC enhancers identified here in HCC in vivo.
Materials and methods
Cell culture of HCCs
The HepG2 and Huh7 HCC cell lines were purchased from Korean Cell Line Bank (Seoul, Korea) and maintained in minimum essential medium or RPMI-1640 medium supplemented with 10% fetal bovine serum (FBS) and penicillin (100 units/ml)/streptomycin (100 μg/ml) (Thermo Fisher Scientific, Waltham, MA, USA). The medium was replaced every 3–4 days. The cells were maintained in a humidified incubator with 95% air and a 5% CO2 atmosphere at 37 °C. JQ1, OTX015, C646, and 5, 6-dichlorobenzimidazole 1-β-d-ribofuranoside (DRB) were purchased from Tocris Bioscience (Minneapolis, MN, USA). JQ1, OTX015, C646, and DRB were dissolved in dimethyl sulfoxide (DMSO) at a stock concentration of 10 mM. The cells were treated with different concentrations of JQ1 and DRB for different durations.
Cell proliferation assays
Relative cell numbers were assessed using a premixed water-soluble tetrazolium salt (WST-1) cell viability test (Takara, Shiga, Japan) according to the manufacturer’s instructions. The cells were seeded at a density of 1 × 104 cells per well. WST-1 was added to each well, and the absorbance of the microplate at 450 nm was measured after an additional 4 h incubation. The data represent three independent experiments (n = 3). DNA-synthesizing cells were visualized using an Ethynyl deoxyuridine (EdU) kit (Invitrogen, CA, USA) following the manufacturer’s instructions. Then, the cells were washed with phosphate-buffered saline, mounted with a 4’, 6-diamidino-2-phenylindole (DAPI)-containing mounting solution (Vectashield, Vector Laboratories, Burlingame, CA, USA), and imaged by microscopy (Nikon Eclipse 80i, Tokyo, Japan). The percentage of EdU-positive cells was examined in HCC cell lines using ImageJ (Bethesda, MD, USA) software. The data represent three independent experiments (n = 3).
Gene expression analysis using quantitative PCR (qRT-PCR)
Total RNA was extracted from the cells using RNAiso Plus (Takara, Shiga, Japan) according to the manufacturer’s instructions. cDNA was synthesized by PrimeScript reverse transcriptase (Takara, Shiga, Japan) and amplified using gene-specific primers. The primers used for qRT-PCR are listed in Supplementary Table S1. The primers were designed by Primer Bank (https://pga.mgh.harvard.edu/primerbank/). qRT-PCR was performed with TBGreen Premix Ex Taq II (Takara, Shiga, Japan). Glyceraldehyde-3-phosphate dehydrogenase (GAPDH) was used as an internal control. The data represent three independent experiments (n = 3). After performing qRT-PCR, the results were analyzed using the critical threshold (ΔCT) and the comparative critical threshold (ΔΔCT) methods in ABI 7500 (Applied Biosystems, Foster City, CA, USA) software with the NormFinder and geNorm PLUS algorithms.
Genomic data analysis
We re-analyzed two public ChIP-seq data sets in Gene Expression Omnibus (GEO) (GSE29611 and GSM1112809) according to the procedure described previously58, and three GRO-seq data sets in GEO (GSE92375, GSM3271003, and GSM3271012). For our re-analysis, the raw data were trimmed with Trimmomatic (version 0.36)59 and processed using Bowtie2 (version 2.3.5)60 or STAR (version 2.7.8)61 aligner software with a UCSC hg 38 reference. The ChIP-seq and GRO-seq peaks identified were analyzed with Homer (version 4.11)62 and visualized using UCSC Genome Browser (https://www.genome.ucsc.edu).
Luciferase reporter assay
Enhancer regions (R1 ~ R6) were amplified using LongAmp Taq 2X Master Mix (New England Biolabs, Ipswich, MA, USA). Enhancer regions were amplified using forward and reverse primers to generate NheI or SacI and XhoI sites, respectively. These constructs were cloned into the pGL4.26 construct (Promega, Madison, WI, USA). The primers used for cloning are listed in Supplementary Table S2. The cells were seeded into 24-well plates and transfected with Lipofectamine 3000 (Thermo Fisher Scientific, Waltham, MA, USA). According to the manufacturer’s instructions, luciferase activity was measured using the Dual-Glo Luciferase Assay kit (Promega, Madison, WI, USA). PRL-TK (Renilla luciferase expression construct; Promega, Madison, WI, USA) was used as an internal control. Luciferase activity was normalized to Renilla luciferase and the control (empty vector).
Construction of R3 region KO HCCs using the CRISPR/Cas9 system
Forward and reverse oligomers for gRNA synthesis against target sites were designed according to the manufacturer’s instructions. The oligomers were extended into 100-mer insert DNA using Phusion High-Fidelity PCR Master Mix (M0531, New England BioLabs, Ipswich, MA, USA) with the following setup: 2 min at 98 °C, 4 cycles of amplification (10 s at 98 °C, 20 s at 53 °C, 30 s at 72 °C), and 5 min at 72 °C. Then, the insert DNA was purified and combined with gRNA_Cloning Vector using Gibson Assembly Master Mix (E2611, New England Biolabs, Ipswich, MA, USA) at 50 °C for 1 h, followed by transformation and colony PCR. The cloned vectors were then purified and ordered to be sequenced (Macrogen, Seoul, Korea) to confirm the recombination. HCCs were transfected with the recombinant gRNA plasmid vector and hCas9 in a 95:5 ratio using Lipofectamine 3000 (L3000-001, Life Technologies, Carlsbad, CA, USA) according to the manufacturer’s instructions. The gRNA cloning vector and hCas9 was a gift from George Church63. The transfected cells were seeded into 96-well plates at a ratio of less than 1 cell per well. After ~ 2 weeks, cells from wells that showed growth were moved into 24-well plates. After further growth, a portion of each well was used to extract genomic DNA (gDNA) using a Wizard Genomic DNA Purification Kit (A1125, Promega, Madison, WI, USA). Then, gDNA was amplified by PCR with target-specific primers and sequenced to check properly generated deletions. The cells of one confirmed well were expanded in 100 mm dishes and then used for the various gene expression and proliferation assays. These analyses were performed within the first 10 passages.
Knockdown of eRNA using ASO
Locked nucleic acid (LNA)-modified ASOs complementary to eRNA of MYC were designed from Antisense LNA GapmeRs (Qiagen, Hilden, Germany). The ASOs were purchased from Qiagen. The sequences are listed in Supplementary Table S3. For the transfection of Huh7 cells, ASOs were mixed with RNAiMAX in serum-free Opti-MEM (Gibco, Waltham, MA, USA). At varying concentrations of ASOs, dissolved Opti-MEM was added, and the cells were incubated in a growth medium for 4 h at 37 °C and 5% CO2. For total RNA extraction, the cells were harvested 48 h posttransfection.
Colony formation assay
R3-deleted and WT Huh7 cells were seeded on 6-well plates in growth media at a density of 2500 cells/well and incubated in a CO2 incubator for 10 days. Then, the cells were washed with PBS, fixed with 4% paraformaldehyde for 20 min, and washed once with PBS. The cells were stained with 1% Crystal Violet (Sigma, St. Louis, MO, USA) for 30 min. After Crystal Violet was removed, the plates were washed with DW for 5 min and dried. The stained cells were analyzed for colony formation rates using ImageJ (Bethesda, MD, UAS).
Spheroid formation assay
Cells were seeded on 24-well ultralow attachment culture dishes in growth media at a density of 1000 cells/well. DMEM/F12 serum-free medium (Gibco, Waltham, MA, USA) contained 2 mM l-glutamine, 1% sodium pyruvate (Invitrogen, Carlsbad, CA, USA), 100 U/ml penicillin, and 100 µg/ml streptomycin supplemented with 20 ng/ml epithelial growth factor (Invitrogen, Carlsbad, CA, USA), 10 ng/ml fibroblast growth factor-2 (Invitrogen, Carlsbad, CA, USA), N2 (R&D Systems, Minneapolis, MN, USA), and B27 (Invitrogen, Carlsbad, CA, USA). The cells were incubated in a CO2 incubator for one to two weeks, and oncosphere cells over 100 µm were counted with JuLI Br (NanoEnTek, Seoul, Korea).
Statistical analysis
The data are presented as the mean ± standard deviation (SD) of the mean. All statistical analyses were performed using the IBM SPSS Statistics 26.0 program (IBM). We used a one-way analysis of variance followed by Tukey’s honestly significant difference post hoc test. p values < 0.05 were considered significant.
Abbreviations
- HCC:
-
Hepatocellular carcinoma
- eRNA:
-
Enhancer RNA
- ASO:
-
Antisense oligonucleotide
- SE:
-
Super-enhancer
- GRO-seq:
-
Global run-on sequencing
- H2K27ac:
-
Acetylated H3 lysine 27
- Chr:
-
Chromosome
- TAD:
-
Topologically associated domains
- CSC:
-
Cancer stem cell
- qRT-PCR:
-
Quantitative reverse transcription-polymerase chain reaction
References
Pelengaris, S., Khan, M. & Evan, G. c-MYC: More than just a matter of life and death. Nat. Rev. Cancer 2, 764–776. https://doi.org/10.1038/nrc904 (2002).
Adhikary, S. & Eilers, M. Transcriptional regulation and transformation by Myc proteins. Nat. Rev. Mol. Cell Biol. 6, 635–645. https://doi.org/10.1038/nrm1703 (2005).
Gabay, M., Li, Y. & Felsher, D. W. MYC activation is a hallmark of cancer initiation and maintenance. Cold Spring Harb. Perspect. Med. 4, a014241. https://doi.org/10.1101/cshperspect.a014241 (2014).
Lancho, O. & Herranz, D. The MYC Enhancer-ome: Long-Range Transcriptional Regulation of MYC in Cancer. Trends Cancer 4, 810–822. https://doi.org/10.1016/j.trecan.2018.10.003 (2018).
Ahmadiyeh, N. et al. 8q24 prostate, breast, and colon cancer risk loci show tissue-specific long-range interaction with MYC. Proc. Natl. Acad. Sci. U S A 107, 9742–9746. https://doi.org/10.1073/pnas.0910668107 (2010).
Hnisz, D. et al. Super-enhancers in the control of cell identity and disease. Cell 155, 934–947. https://doi.org/10.1016/j.cell.2013.09.053 (2013).
Oktay, Y. et al. IDH-mutant glioma specific association of rs55705857 located at 8q24.21 involves MYC deregulation. Sci. Rep. 6, 27569. https://doi.org/10.1038/srep27569 (2016).
Balogh, J. et al. Hepatocellular carcinoma: A review. J. Hepatocell. Carcinoma 3, 41–53. https://doi.org/10.2147/JHC.S61146 (2016).
Yang, Y. et al. The chromatin remodeling protein BRG1 links ELOVL3 trans-activation to prostate cancer metastasis. Biochim. Biophys. Acta 1862, 834–845. https://doi.org/10.1016/j.bbagrm.2019.05.005 (2019).
Liu, M., Jiang, L. & Guan, X. Y. The genetic and epigenetic alterations in human hepatocellular carcinoma: A recent update. Protein Cell 5, 673–691. https://doi.org/10.1007/s13238-014-0065-9 (2014).
Zender, L. et al. Cancer gene discovery in hepatocellular carcinoma. J. Hepatol. 52, 921–929. https://doi.org/10.1016/j.jhep.2009.12.034 (2010).
Zheng, K., Cubero, F. J. & Nevzorova, Y. A. c-MYC-making liver sick: Role of c-MYC in hepatic cell function, homeostasis disease. Genes https://doi.org/10.3390/genes8040123 (2017).
Kalkat, M. et al. MYC deregulation in primary human cancers. Genes https://doi.org/10.3390/genes8060151 (2017).
Hsieh, C. L. et al. Enhancer RNAs participate in androgen receptor-driven looping that selectively enhances gene activation. Proc. Natl. Acad. Sci. USA 111, 7319–7324. https://doi.org/10.1073/pnas.1324151111 (2014).
Arnold, P. R., Wells, A. D. & Li, X. C. Diversity and emerging roles of enhancer RNA in regulation of gene expression and cell fate. Front. Cell Dev. Biol. 7, 377. https://doi.org/10.3389/fcell.2019.00377 (2019).
Lee, J. H., Xiong, F. & Li, W. Enhancer RNAs in cancer: Regulation, mechanisms and therapeutic potential. RNA Biol. 17, 1550–1559. https://doi.org/10.1080/15476286.2020.1712895 (2020).
Bhagwat, A. S. et al. BET bromodomain inhibition releases the mediator complex from select cis-regulatory elements. Cell Rep. 15, 519–530. https://doi.org/10.1016/j.celrep.2016.03.054 (2016).
Donati, B., Lorenzini, E. & Ciarrocchi, A. BRD4 and cancer: Going beyond transcriptional regulation. Mol. Cancer 17, 164. https://doi.org/10.1186/s12943-018-0915-9 (2018).
Kanno, T. et al. BRD4 assists elongation of both coding and enhancer RNAs by interacting with acetylated histones. Nat. Struct. Mol. Biol. 21, 1047–1057. https://doi.org/10.1038/nsmb.2912 (2014).
Bid, H. K. et al. The Bromodomain BET inhibitor JQ1 suppresses tumor angiogenesis in models of childhood sarcoma. Mol. Cancer Ther. 15, 1018–1028. https://doi.org/10.1158/1535-7163.MCT-15-0567 (2016).
Goel, H. L. & Mercurio, A. M. VEGF targets the tumour cell. Nat. Rev. Cancer 13, 871–882. https://doi.org/10.1038/nrc3627 (2013).
Kaseb, A. O. et al. Vascular endothelial growth factor in the management of hepatocellular carcinoma: A review of literature. Cancer 115, 4895–4906. https://doi.org/10.1002/cncr.24537 (2009).
Benhammou, J. N. et al. Novel lipid long intervening noncoding RNA, oligodendrocyte maturation-associated long intergenic noncoding RNA, regulates the liver steatosis gene stearoyl-coenzyme A desaturase as an enhancer RNA. Hepatol. Commun. 3, 1356–1372. https://doi.org/10.1002/hep4.1413 (2019).
Bouvy-Liivrand, M. et al. Analysis of primary microRNA loci from nascent transcriptomes reveals regulatory domains governed by chromatin architecture. Nucleic Acids Res. 45, 9837–9849. https://doi.org/10.1093/nar/gkx680 (2017).
Consortium, E. P. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74. https://doi.org/10.1038/nature11247 (2012).
Gertz, J. et al. Distinct properties of cell-type-specific and shared transcription factor binding sites. Mol. Cell 52, 25–36. https://doi.org/10.1016/j.molcel.2013.08.037 (2013).
Viiri, L. E. et al. Extensive reprogramming of the nascent transcriptome during iPSC to hepatocyte differentiation. Sci. Rep. 9, 3562. https://doi.org/10.1038/s41598-019-39215-0 (2019).
Bernstein, B. E. et al. The NIH roadmap epigenomics mapping consortium. Nat. Biotechnol. 28, 1045–1048. https://doi.org/10.1038/nbt1010-1045 (2010).
Zhang, X. et al. Identification of focally amplified lineage-specific super-enhancers in human epithelial cancers. Nat. Genet. 48, 176–182. https://doi.org/10.1038/ng.3470 (2016).
Zhu, H. Q. et al. Aberrant expression of CCAT1 regulated by c-Myc predicts the prognosis of hepatocellular carcinoma. Asian Pac. J. Cancer Prev. 16, 5181–5185. https://doi.org/10.7314/apjcp.2015.16.13.5181 (2015).
Wang, Y. et al. The 3D Genome Browser: a web-based browser for visualizing 3D genome organization and long-range chromatin interactions. Genome Biol. 19, 151. https://doi.org/10.1186/s13059-018-1519-9 (2018).
Fernandez, P. C. et al. Genomic targets of the human c-Myc protein. Genes Dev 17, 1115–1129. https://doi.org/10.1101/gad.1067003 (2003).
Khattar, E. & Tergaonkar, V. Transcriptional regulation of telomerase reverse trnascriptase (TERT) by MYC. J. Hepatol. 69, 182–236. https://doi.org/10.1016/j.jhep.2018.03.019 (2018).
Dave, K. et al. Mice deficient of Myc super-enhancer region reveal differential control mechanism between normal and pathological growth. Elife 6, e23382. https://doi.org/10.7554/eLife.23382 (2017).
Tak, Y. G. et al. Effects on the transcriptome upon deletion of a distal element cannot be predicted by the size of the H3K27Ac peak in human cells. Nucleic Acids Res. 44, 4123–4133. https://doi.org/10.1093/nar/gkv1530 (2016).
Florea, V. et al. c-MYC is Essential to prevent endothelial pro-inflammatory senescent phenotype. PLoS ONE 8, e73146. https://doi.org/10.1371/journal.pone.0073146 (2013).
Kim, T. K. et al. Widespread transcription at neuronal activity-regulated enhancers. Nature 465, 182–187. https://doi.org/10.1038/nature09033 (2010).
Sur, I. & Taipale, J. The role of enhancers in cancer. Nat. Rev. Cancer 16, 483–493. https://doi.org/10.1038/nrc.2016.62 (2016).
Loven, J. et al. Selective inhibition of tumor oncogenes by disruption of super-enhancers. Cell 153, 320–334. https://doi.org/10.1016/j.cell.2013.03.036 (2013).
Shi, J. & Vakoc, C. R. The mechanisms behind the therapeutic activity of BET bromodomain inhibition. Mol. Cell 54, 728–736. https://doi.org/10.1016/j.molcel.2014.05.016 (2014).
Pomerantz, M. M. et al. The androgen receptor cistrome is extensively reprogrammed in human prostate tumorigenesis. Nat. Genet. 47, 1346–1351. https://doi.org/10.1038/ng.3419 (2015).
Andersson, R. et al. An atlas of active enhancers across human cell types and tissues. Nature 507, 455–461. https://doi.org/10.1038/nature12787 (2014).
Park, A. et al. Global epigenomic analysis of KSHV-infected primary effusion lymphoma identifies functional MYC superenhancers and enhancer RNAs. Proc. Natl. Acad. Sci. U S A 117, 21618–21627. https://doi.org/10.1073/pnas.1922216117 (2020).
Schuijers, J. et al. Transcriptional dysregulation of MYC reveals common enhancer-docking mechanism. Cell Rep 23, 349–360. https://doi.org/10.1016/j.celrep.2018.03.056 (2018).
Aden, D. et al. Controlled synthesis of HBsAh in a differentiated human liver carcinoma-derived cell line. Nature 282, 615–616. https://doi.org/10.1038/282615a0 (1979).
Nakabayashi, H. et al. Growth of human hepatoma cells lines with differentiated functions in chemically defined medium. Cancer Res. 42, 3858–3863 (1982).
Kaczkowski, B. et al. Transcriptome analysis of recurrently deregulated genes across multiple cancers identifies new pan-cancer biomarkers. Cancer Res. 76, 216–226. https://doi.org/10.1158/0008-5472.CAN-15-0484 (2016).
Oh, S. et al. Enhancer release and retargeting activates disease-susceptibility genes. Nature https://doi.org/10.1038/s41586-021-03577-1 (2021).
Pnueli, L., Rudnizky, S., Yosefzon, Y. & Melamed, P. RNA transcribed from a distal enhancer is required for activating the chromatin at the promoter of the gonadotropin alpha-subunit gene. Proc. Natl. Acad. Sci. USA 112, 4369–4374. https://doi.org/10.1073/pnas.1414841112 (2015).
Pan, C. W. et al. Functional roles of antisense enhancer RNA for promoting prostate cancer progression. Theranostics 11, 1780–1794. https://doi.org/10.7150/thno.51931 (2021).
Zhang, Z. et al. Transcriptional landscape and clinical utility of enhancer RNAs for eRNA-targeted therapy in cancer. Nat. Commun. 10, 4562. https://doi.org/10.1038/s41467-019-12543-5 (2019).
Jiao, W. et al. HPSE enhancer RNA promotes cancer progression through driving chromatin looping and regulating hnRNPU/p300/EGR1/HPSE axis. Oncogene 37, 2728–2745. https://doi.org/10.1038/s41388-018-0128-0 (2018).
Liang, J. et al. Epstein-Barr virus super-enhancer eRNAs are essential for MYC oncogene expression and lymphoblast proliferation. Proc. Natl. Acad. Sci. USA 113, 14121–14126. https://doi.org/10.1073/pnas.1616697113 (2016).
Zhang, H. L., Wang, P., Lu, M. Z., Zhang, S. D. & Zheng, L. c-Myc maintains the self-renewal and chemoresistance properties of colon cancer stem cells. Oncol. Lett. 17, 4487–4493. https://doi.org/10.3892/ol.2019.10081 (2019).
Delmore, J. E. et al. BET bromodomain inhibition as a therapeutic strategy to target c-Myc. Cell 146, 904–917. https://doi.org/10.1016/j.cell.2011.08.017 (2011).
Li, G. Q. et al. Suppression of BRD4 inhibits human hepatocellular carcinoma by repressing MYC and enhancing BIM expression. Oncotarget 7, 2462–2474. https://doi.org/10.18632/oncotarget.6275 (2016).
Sengupta, D. et al. Disruption of BRD4 at H3K27Ac-enriched enhancer region correlates with decreased c-Myc expression in Merkel cell carcinoma. Epigenetics 10, 460–466. https://doi.org/10.1080/15592294.2015.1034416 (2015).
Kang, S. C. et al. Transcriptomic profiling and H3K27me3 distribution reveal both demethylase-dependent and independent regulation of developmental gene transcription in cell differentiation. PLoS ONE 10, e0135276. https://doi.org/10.1371/journal.pone.0135276 (2015).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120. https://doi.org/10.1093/bioinformatics/btu170 (2014).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359. https://doi.org/10.1038/nmeth.1923 (2012).
Dobin, A. et al. STAR: Ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21. https://doi.org/10.1093/bioinformatics/bts635 (2013).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and b cell identities. Mol. Cell 38, 576–589. https://doi.org/10.1016/j.molcel.2010.05.004 (2010).
Mali, P. et al. RNA-guided human genome engineering via Cas9. Science 339, 823–826. https://doi.org/10.1126/science.1232033 (2013).
Funding
This work was supported by the National Research Foundation of Korea (NRF) Grants 2017M3A9G7073033 and 2020R1A2C1014193 (to Y. G. C.), 2016R1D1A1B04934970 (to K. H. J.), and 2014M3C9A3064693 (to Y. S. L.), 2020R1F1A1063217 (to B. B.) from the Korean government.
Author information
Authors and Affiliations
Contributions
H.I.C. designed and performed experiments, analyzed and interpreted the data, and prepared the manuscript; G.Y.A., M.N.B. and E.Y.Y. designed and performed experiments; B.B. analyzed and interpreted the data, edited the manuscript; J.C.C. and Y.S.L. analyzed next-generation sequencing and bioinformatics data; K.H.J. designed experiments, financial support, analyzed and interpreted the data, prepared and edited the manuscript; Y.G.C. designed experiments, financial support, analyzed and interpreted next-generation sequencing and bioinformatics data, final approval of the manuscript, edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Choi, H.I., An, G.Y., Yoo, E. et al. Targeting of non-coding RNAs encoded by novel MYC enhancers inhibits the proliferation of human hepatic carcinoma cells in vitro. Sci Rep 12, 855 (2022). https://doi.org/10.1038/s41598-022-04869-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-04869-w
This article is cited by
-
MYC in liver cancer: mechanisms and targeted therapy opportunities
Oncogene (2023)
-
Function of Long Noncoding RNAs in Glioma Progression and Treatment Based on the Wnt/β-Catenin and PI3K/AKT Signaling Pathways
Cellular and Molecular Neurobiology (2023)
-
The bromodomain inhibitor JQ1 up-regulates the long non-coding RNA MALAT1 in cultured human hepatic carcinoma cells
Scientific Reports (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.