Abstract
Multiple sclerosis (MS) is a chronic inflammatory and autoimmune disorder of the central nervous system characterized by myelin loss and axonal dysfunction. Increased production of inflammatory factors such as cytokines has been implicated in axon destruction. In the present study, we compared the expression level of IL7R, NFATc2, and RNF213 genes in the peripheral blood of 72 MS patients (37 familial MS, 35 sporadic MS) and 74 healthy controls (34 individuals with a family history of the disease, 40 healthy controls without a family history) via Real-time PCR. Our results showed that the expression level of IL7R was decreased in the sporadic patients in comparison with other groups. Additionally, there was an increased NFATc2 expression level in MS patients versus healthy controls. Increased expression of NFATc2 in sporadic and familial groups compared to the controls, and familial group versus FDR was also seen. Our results also represented an increased expression level of RNF213 in familial patients as compared to the control group. The similar RNF213 expression between sporadic and control group, as well as FDR and familial group was also seen. Diagnostic evaluation was performed by receiver operating characteristic (ROC) curve analysis and area under the curve (AUC) calculation. The correlation of clinical parameters including onset age and Expanded Disability Status Scale (EDSS) with our gene expression levels were also assessed. Overall, decreased expression level of IL7R in the sporadic cases and increased expression level of NFATc2 may be associated with the pathogenesis of MS disease. Confirmation of the effects of differential expression of RNF213 gene requires further studies in the wider statistical populations.
Similar content being viewed by others
Introduction
Multiple Sclerosis (MS) is a chronic inflammatory autoimmune disorder of the central nervous system (CNS) affecting about 2–3 million people worldwide1. The main causes of the disease are a breakdown of the blood–brain barrier, gliosis of astrocytes, loss of oligodendrocytes and destruction of axons2. In all of these stages, pro-inflammatory cytokines could cause or exacerbate the disease compromising the blood–brain barrier. Therefore, inflammation plays a key role in the pathogenesis of the disease3. Many of the cells in the innate and adaptive immune system respond to inflammation created within the central nervous system4. Mounting criteria have suggested that the major cause of MS is a combination of both environmental and genetic factors5,6,7. Genetic susceptibility plays an important role in the development of the disease, as more than 200 genetic variants associated with MS risk is identified through genome-wide association studies (GWAS) in the last few years8. The most susceptible locus associated with this disease is the HLA class Π genes9. Mostly, the strongest MS risk factor is human leukocyte antigen (HLA)-DRB*15: 01, a major histocompatibility complex (MHC) class Π allele10,11. The importance of non-HLA genes in MS has also been repeatedly mentioned. One of the most non-HLA genes was recognized as the Interleukin 7 Receptor (IL7R), which is translated to a functional receptor for interleukin 7 (IL7)12. Generally, IL7R can be recognized on the surface of the mature T cells and immature B cells. IL7 has a range of biological activities and is essential for the survival, proliferation, and homeostasis of T-cells. It is also a factor in bone marrow culture that can induce the proliferation of B progenitor cells13. IL7R may also have a primary signaling function through the autoimmunity cascade. This gene may also be involved in the remyelination process. Deletion of IL7R delays the maturation and differentiation of oligodendrocyte cells14. Previous studies have shown the association between IL7R and MS, and also the link between the IL7R haplotypes with the disease has been investigated15.
The Wnt signaling pathway is one of the key cascades in regulating the evolution and stability of the immune and blood cells. Although the disruption of this pathway is associated with many pathological conditions, its exact role is still controversial. This pathway appears to regulate several biological processes, including cell proliferation, migration, polarity, differentiation, and axonal growth. Wnt proteins have been shown to regulate T-cell growth and maturation of dendritic cells and also play a significant function in the expression of inflammatory mediators16. NFATc2 is one of the important genes within the Wnt signaling pathway that plays an important role in immune system inflammation. It is one of the several transcription factors regulated by Calcineurin in the Wnt/Ca2+ signaling pathway. NFATc2 also mediates oligodendrocyte differentiation, T-cell function and coordinates the induction of cytokines and immunomodulatory molecules. NFAT proteins can regulate the expression of genes involved in the differentiation of T cell receptors and T helper cells. It is reported that the extracellular calcium influx through cation channels could lead to the increased expression of NFAT target genes17. RNF213 is an ATPase and E3 ubiquitin ligase protein, which is involved in the angiogenesis, and degradation of FLNA and NFATc2, that leads to the inhibition of Wnt/Ca2+ pathway. Consequently, RNF213 deficiency has been shown to cause an increased expression of NFAT target genes18. Interestingly, these genes play an important role in chronic inflammation, nerve cell destruction, and myelin depletion, and appear to be common factors in the onset of familial MS19. Considering the effect of MS individual's lifestyle, as well as the lack of diagnostic and prognostic biomarkers, it is important to identify therapeutic targets and proper biomarkers in MS condition. Our study focuses on the investigating the expression of IL7R, NFATc2, and RNF213 pro-inflammatory genes. We also investigated their potential application in the diagnosis of the disease in the familial and sporadic MS patients versus healthy individuals with and without a family history of the disease. To investigate the effect of hereditary factors involved in the familial aggregation of the disease, we have divided patients into two different groups: familial and sporadic. Healthy first-degree relatives of the familial patients were also involved in the study to identify a distinct type of gene expression pattern that indicates a predisposition to MS.
Materials and methods
Study subjects
All patients were referred to Kashani hospital, affiliated with Isfahan University of Medical Sciences between June 2018 and September 2019, and blood samples afterwards were taken. The diagnosis of MS was based on the McDonald Criteria20. Blood samples were taken from 72 MS patients (35 sporadic, 37 familial), and 74 healthy individuals (34 healthy first-degree relatives of the familial MS patients (FDR), 40 healthy persons without a family history of MS (control)). All patients received IFN-Beta as a part of their treatment. Patients with coexistence of other neurological disorders were excluded from the study. The control group had no family history of the autoimmune or neurological disorders and were admitted for medical check-up. They were also sex/age-matched to the patient groups. All patients were evaluated for the extended disability status scale (EDSS) and were subjected to magnetic resonance imaging (MRI). The study was performed in accordance with the seventh edition of Helsinki declaration, approved by the ethics committee of the University of Isfahan, and informed consent was obtained from patients and healthy individuals. Table 1 presents more information on the clinical features of patients, and also demographic data of the healthy individuals.
Demographic data of the participants
Our samples were classified into four different groups (35 sporadic patients, 37 familial patients, 34 FDRs, and 40 controls). In brief, 71.4% of the sporadic group (with an average age of 38.0 ± 10.5 years) and 75.6% of the familial group (with an average age of 36.2 ± 9.6 years) were female. Also, healthy participants were largely female (55.88%, 72.5%), with an average age of 38.4 ± 9.2, 37.3 ± 15.9 years in FDR and Control group, respectively. There was no significant difference in age (P = 0.38) and sex (P = 0.16) between separated groups. The Onset age and EDSS results were not altered significantly in sporadic and familial groups. Most of the patients in both groups were classified as relapsing–remitting type of MS (RRMS). The MRI results also were not meaningfully different in both patient groups. Table 1 presents the detailed characteristics of all patients and healthy groups.
RNA isolation and reverse transcription
Total RNA was extracted from the peripheral blood of patients and healthy controls using the Favor Prep blood/cultured cell total RNA mini kit (Favorgen, Taiwan) according to the manufacturer’s protocol. The samples stored at − 70 °C. RNA concentration quantitated by Nano drop ND-100 (Thermo scientific, USA). Only samples containing completely pure RNA were used in the RT-PCR analysis, also the quality of this RNA samples was determined using electrophoresis. Reverse transcription was carried out in 10 μl reaction volume using a prime script RT reagent kit (Takara, Japan), following the standard instruction. The reaction mixture contained 0.5 μl of random hexamer primer, 0.5 μl of oligo dT primers, 2 μl of 5 × prime script buffer, 0.5 μl of prime script RT Enzyme Mix I, and 6.5 μl of extracting RNA solution plus RNase free water. The incubation program was as follows: 15 min at 37 °C, and 5 s at 85 °C. Finally, cDNA samples were stored at −20 °C.
Quantitative real time-PCR
Target gene intron-spanning primers were designed by the NCBI online tool and oligo primer software (Bioneer Company), and primer sequences are presented in Table 2. Real time PCR was carried out on a Bio-Rad (Chromo4 TM, Bio-Rad Laboratories, USA) and also an ABI-7500 Real-Time PCR System (Applied Biosystems) using a Real Q plus 2 × master mix gene (Ampliqon, Odense, Denmark). Each PCR reaction was performed in a total volume of 10 μl, including 5 μl of the master mix, 0.5 μl reverse and 0.5 μl forward primer of target genes, and 4 μl of the reverse-transcribed cDNA. Reactions were run under standard conditions: initial denaturation at 95 °C for 15 min, 40 amplification cycles of 30 s denaturation at 95 °C and 30 s, annealing and elongation at 72 °C. The PCR products were separated on 1.5% agarose gel and produced the expected size amplicons for each specific target (Supplementary Fig. 1). The comparative Ct method (ΔCt) was used to estimate relative expression changes in our target genes. The expression levels were normalized to ACTB as a reference gene.
Statistical analysis
The results were analyzed using the SPSS 26 software package (SPSS Inc. 2019. Chicago) and Graph Pad Prism 8 (Graph Pad Prism Software, Inc. San Diego CA, USA). Independent sample t-test was used to compare the differential expression between patients and control groups. Also the one-way ANOVA followed by LSD post hoc test was used to determine the statistical significance of expression differences among four analyzed groups.
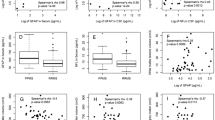
We also investigated the correlation between IL7R, NFATc2, RNF213 relative expressions and clinical variables, such as onset age and Expanded Disability Status Scale (EDSS). Spearman Correlation test were used to examine the association of relative gene expression levels with clinical parameters. The predictive power of the differentially expressed genes was calculated with plotting receiver-operating characteristic (ROC) curve (a graphical presentation of sensitivity versus 1-specificity) and also defining area under the curve (AUC). The AUC can be regarded as an indicator for evaluating the accuracy of a diagnostic test. The AUC value varies from 0 to 1, being close to 1 when the diagnostic assay has a high accuracy21. P ≤ 0.05 were considered statistically significant. The cutoff point is the threshold value of analytical signals below which the samples are negative and above which the samples are evaluated positive22. The optimal cut off point values for relative quantification that separates different groups was determined.
Results
Investigation of the relative gene expression levels of IL7R, NFATc2, and RNF213 among genetically separated groups
We compared the expression level of IL7R in the four study groups. Our results demonstrated a significant down-regulation of IL7R in sporadic MS patients compared with the control group by 80% ± 0.4515 (Pvalue < 0.0001) as well as the FDR group by 59% ± 0.4478 (Pvalue = 0.0005). There was also a significant decrease in the expression of sporadic versus familial patients by 70% ± 0.4697 (Pvalue = 0.0004). We did not observe a significant difference in expression between the familial patients and healthy individuals (both Control and FDR) (Fig. 1A). Comparison of the NFATc2 expression level in the four study groups demonstrated an increased expression in familial compared with control group by 314% ± 0.5975 (Pvalue = 0.0065), and also FDR group by 290% ± 0.6221(Pvalue = 0.031). There was also a significant increase in the expression of sporadic versus control group by 254% ± 0.6230 (Pvalue = 0.0148) (Fig. 1B). In the case of RNF213 gene, the expression comparison among four groups demonstrated a similar gene expression level among familial and FDR groups. A significant increase in the expression of RNF213 in familial by 277% ± 0.5420 (Pvalue = 0.0074), as well as the FDR by 227% ± 0.5621 (Pvalue = 0.0372) versus control group was also seen. The expression comparison demonstrated a significant increase in familial compared with sporadic patients by 225% ± 0.5592 (Pvalue = 0.038). A similar RNF213 gene expression level among sporadic and control group was also seen (Fig. 1C).
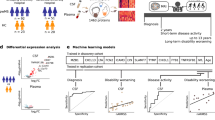
Differential expression levels of (A) IL7R, (B) NFATc2 and (C) RNF213 among genetically separated groups and control group. One-way ANOVA test and in the following, Fisher LSD post hoc test was used for the statistical analysis. Ctrl control group, FDR healthy first-degree relatives of the familial group. Bars represent the standard error of the mean. (*p < 0.05, **p < 0.01, ***p < 0.001).
Investigation of the relative gene expression levels of IL7R, NFATc2, and RNF213 among patients and the control group
In this part of the study all sporadic and familial MS patients were classified as a single group: MS (n = 72). Also all healthy individuals were considered as a definite control group (n = 74). In this type of classification, the expression level of IL7R was lower in MS patients as compared to the control group by 58% ± 0.3586 (Pvalue = 0.0009) (Fig. 2A). Comparison of the NFATc2 expression level in the two study groups showed an up-regulation in NFATc2 mRNA level in the MS versus control groups by 275% ± 0.4586 (Pvalue = 0.0019) (Fig. 2B). No significant change in the RNF213 expression level was observed between MS and control groups (Pvalue = 0.3550) (Fig. 2C).
The relative expression level of (A) IL7R, (B) NFATc2 and (C) RNF213 in all MS patients (n:72) compared to healthy controls (n:74). MS multiple sclerosis, Ctrl control group. Bars represent the standard error of the mean. (*p < 0.05, **p < 0.01, ***p < 0.001) Unpaired t test with Welch’s correction and also Fisher LSD post hoc test was used for statistical analysis.
Diagnostic values of IL7R, NFATc2 and RNF213 among different study groups
Diagnostic test evaluation was performed for genes whose expression had changed significantly by ROC curve and AUC calculation. According to the expression level of IL7R between all patients and healthy controls, AUC was 0.66 (Pvalue = 0.0014) (Fig. 3A). Also, the optimal cut off value was defined as 0.0246 and the sensitivity and specificity values of 55.38% and 80.60% were achived, respectively. Plotting ROC curve for the sporadic patients versus FDR group (AUC = 0.75, Pvalue = 0.0007, cut off point = 0.019, sensitivity = 71.88%, specificity = 89.29%) (Fig. 3B), also sporadic patients versus healthy controls (AUC = 0.79, Pvalue < 0.0001, cut off point = 0.0202, sensitivity = 71.88%, specificity = 82.05%) (Fig. 3C), and sporadic patients versus familial patients (AUC = 0.71, Pvalue = 0.003, cut off point = 0.0201, sensitivity = 71.88%, specificity = 78.79%) (Fig. 3D), showed a significant result. AUC didn’t change significantly in other groups (Supplementary Fig. 2). In the diagnostic performance assessment of the NFATc2 expression in MS versus the control group AUC was 0.66 (Pvalue = 0.0076) (Fig. 3E). Also, the optimal cut off value was defined as 0.0919 and the sensitivity and specificity values of 52.24% and 75.00% were achived, respectively. Plotting ROC curve for the familial versus FDR group (AUC = 0.66, Pvalue = 0.02, cut off point = 0.1181, sensitivity = 52.78%, specificity = 78.79%) (Fig. 3F), and also familial versus control group (AUC = 0.64, Pvalue = 0.028, cut off point = 0.01038, sensitivity = 52.78%, specificity = 87.18%) (Fig. 3G), showed a significant result. AUC didn’t change significantly in other groups (Supplementary Fig. 2). Diagnostic test evaluation was also performed for RNF213 by ROC curve and AUC calculation. According to the expression level of RNF213 in the familial versus control group, AUC was 0.67 (Pvalue = 0.014). Also, the optimal cut off value was defined as 0.0128, and the sensitivity and specificity values of 58.06% and 72.5% were achived, respectively (Fig. 3H). There was no statistical significance in other groups (Supplementary Fig. 2).
ROC curve analysis for determining statistically significant differences between study groups. (A) ROC curve of all patients and healthy controls analyzed for relative expression level of IL7R (AUC: 0.66, Pvalue = 0.0014). (B) ROC curve of sporadic patients and FDR groups analyzed for relative expression level of IL7R (AUC: 0.75, Pvalue = 0.0007). (C) ROC curve of sporadic patients and healthy controls analyzed for relative expression level of IL7R (AUC: 0.79, Pvalue < 0.0001). (D) ROC curve of sporadic patients and familial patients analyzed for relative expression level of IL7R (AUC: 0.71, Pvalue = 0.0030). (E) ROC curve of all patients and healthy controls analyzed for relative expression level of NFATC2 (AUC: 0.63, Pvalue = 0.076). (F) ROC curve of Familial patients and FDR groups analyzed for relative expression level of NFATC2 (AUC: 0.66, Pvalue = 0.021). (G) ROC curve of Familial patients and healthy controls analyzed for relative expression level of NFATC2 (AUC: 0.64, Pvalue = 0.028). (H) ROC curve of Familial patients and healthy controls analyzed for relative expression level of RNF213 (AUC: 0.67, Pvalue = 0.014).
Association of IL7R, NFATc2 and RNF213 gene expressions with clinical parameters
We investigated the association of IL7R, NFATc2 and RNF213 expression levels with EDSS and onset age in all MS patients and control groups, and separated four groups. The results did not show any significant correlation (Table 3).
Discussion
Despite a great deal of research in the past, there is still no proper treatment for patients with MS, but researchers suggest a genetic effect on the etiology of the disease23. Previous Studies have investigated the factors involved in the familial aggregation of MS24. Due to the evidence indicating the association of IL7R, NFATc2, and RNF213 with immunopathology of MS, in the present study, we analyzed their gene expression in familial and sporadic types of the disease and also healthy individuals to identify a distinct gene expression pattern that indicates a predisposition to the disease.
IL7R is a member of the type Ι cytokine receptor family. As mentioned before, the IL7R-IL7Rα ligand-receptor pair play an important role in the survival and proliferation of B and T lymphocytes, and their genetic variations could lead to the immunodeficiency syndromes12. IL7R is located on chromosome 5pl3, and two promoter polymorphisms contribute to the genetic background of MS pathogenesis25. Previous studies have shown that IL7R can interact with Thymic Stromal Lymphopoietin (TSLP). TSLP is thought to trigger dendritic cell–mediated Th2-type inflammatory responses26. Th2 cells are believed to confer an anti-inflammatory response, and have been associated with decreasing inflammation27. Lei et al., have used metronidazole-induced demyelination in a transgenic zebrafish line in which oligodendrocytes expressed green fluorescent protein (GFP). Their results demonstrated a decrease in IL7R expression that induced JAK/STAT signaling pathway leading to oligodendrocytes apoptosis. These findings highlight the role of IL7R in the demyelination that is important in the pathogenesis of MS28, and increase the need to investigate the expression of IL7R in patients with MS. Therefore, interaction of IL7R with TSLP leads to the induction of T-helper 2 (Th2) cells26, which inhibits synthesis of a wide range of proinflammatory cytokines27. Also, according to the studies of the past that were mentioned, the decrease in IL7R expression caused demyelination26,28. Given that the role of this gene in MS is not clearly defined, we can expect a decrease in gene expression in MS patients. Of course, it should also be noted that gene expression can be different in various types of MS and people with different polymorphisms.
Regulatory T cells (Treg) express the transcription factor Foxp3 and show decreased expression of IL7Ra. Therefore the deficiency in suppression of autoimmune mechanisms may have an important role in multiple sclerosis pathogenesis29. In fact, previous studies have shown that single nucleotide polymorphisms (SNPs) in the IL7R and IL2RA may confer a risk in the susceptibility to MS. The identification of IL7R has made great strides in understanding the MS genetic risk factors30. Increasing evidence has shown that this gene also promotes oligodendrocyte survival and myelination26.
Our results showed that the IL7R was significantly decreased in the sporadic patients compared both to familial patients and to healthy controls. Since there is a significant differential expression between the sporadic and familial group, it is expected that familial patients do not show a significant difference in expression compared with FDR and control groups, and the results could directly confirm this .Additionally, the lower IL7R gene expression in all patients versus control groups might be significantly correlated with sporadic MS cases. The differential expression between the sporadic and familial groups can be attributed to SNPs in one of these groups, which in previous studies have shown in IL7R, that have affected the IL7R expression29. Investigation of the IL7R gene expression in MS patients in Australia, in 2005, showed a down-regulation in PPMS, and an up-regulation in SPMS compared to the controls31. In 2017, Bina et al. have evaluate the expression level and controlling role of lnc-IL-7R in the expression of two variants of IL-7Ra in MS patients versus healthy controls. they concluded no significant difference between the expression levels of IL-7RB and IL-7RS isoforms of IL-7R gene and lnc-IL-7R in MS patients (36 MS patient and 30 healthy controls)32. In 2018, a Bioinformatics study has identified miR-199a and miR-142–3-P as crucial biomarkers in MS due to targeting the pivotal susceptibility genes, in particular KRAS and IL7R. They also examined IL7R expression in a small population of MS and control patients (15 MS, 14 controls) and reported an increased expression of this gene, which contradicts our study33.
Wnt signaling pathway plays a key role in activating the myelination process as well as the differentiation and formation of oligodendrocytes34 . Therefore, it can be said that it is one of the key pathways in the MS disease. This pathway is also necessary for the maturation of the brain cells and is effective in reducing the penetration of immune cells. Besides, activation of Wnt/Ca2+ pathway regulates cytokine production in T cells35 .Previous studies have also suggested the role of the Wnt pathway in the development and oligodendrocytes re-myelination36.Therefore, this pathway can be targeted as a potential and effective treatment for MS patients. The Wnt/Ca2+ pathway activates the Nuclear Factor of Activating T-cell (NFAT)37. NFATc2 is a member of the nuclear transcription factor activating a family of T cells, and is one of the Wnt/Ca2+ pathway genes, which translocates to the nucleus via dephosphorylation by calcineurin. As a result, it activates the Wnt/Ca2+ pathway target genes38. In 2019, Vilariño-Güell et al. performed whole-exome sequencing analysis in 132 patients from 34 families. In this study, pathogenic variants for MS in 12 genes of the innate immune system that regulate the transcription and activation of inflammatory mediators were nominated. NFATc2 and RNF213 were among these genes and provide the molecular and biological rationale for the inflammation, demyelination and neurodegeneration observed in MS patients. In this experiment, NFATc2 knockout mice indicated an increase in immune responses, as well as the production of cytokines in T cells19. The induction of EAE in NFATc2 knockout mice causes a markedly reduced clinical score, compared to wild-type animals39. According to these studies, it seems that NFATc2 suppression can be considered as a potential disease-modifying treatment for MS patients. Thus, it would be expected that the expression level of NFATc2 is increased in MS patients compared with controls. In the present study, the relative expression level of NFATc2 is significantly up-regulated in sporadic and also familial groups, compared with healthy controls. Additionally, we observed significant differences in the of NFATc2 expression level among familial patients and FDR. The results of our study indicated that the expression of NFATc2 is not significantly different between sporadic and familial patients, and therefore the NFATc2 can not act as a biomarker in distinguishing familial and sporadic types. However, since NFATc2 expression in familial patients has changed significantly compared with their first-degree relatives, this gene can be considered as a possible prognostic marker in affected families. This hypothesis is acceptable if FDR is no longer supposed to develop the disease. Thus, down regulation of the NFATc2 in FDR versus familial can be introduced as a prognostic biomarker. Since we preferred older healthy first-degree relatives, the possibility of using this gene as a prognostic biomarker could be strengthened. However, more studies are needed to prove this hypothesis. As expected, NFATC2 mRNA level was also up-regulated in all MS patients versus all controls.
RNF213 is located on chromosome 17 and is one of the genes involved in the inflammatory pathways, such as Wnt/Ca2+. This gene is expressed in most tissues of the body, but its physiological function is still much unknown. RNF213 encodes a unique, 591-KDa protein with both a ring finger domain and walker motifs40. Ring-based E3 ligases have been linked to the control of many cellular processes, including proteasome-dependent proteolysis, immunological processes and transcription41. Therefore, RNF213 is an ATPase and E3 ubiquitin ligase protein. RNF213 also targets NFATc2 proteasomal degradation and attenuats the non-canonical Wnt/Ca2+ pathway. As a result, RNF213 deficiency has been shown to trigger an increased expression of NFAT target genes19. Thus, because the expression of the NFATC2 increased in the pathways leading to MS, the expression of the RNF213 gene is expected to decrease. In the present study, a similar RNF213 gene expression level among familial and FDR groups was seen, but gene expression in both groups had significantly increased compared to the control group. Based on the observed results, the differential expression of this gene can be considered as a predisposing factor in FDR individuals. Most of these individuals may remain asymptomatic for the rest of their lives or develop the disease due to different environmental factors. It seems that the existence of a specific variant in all family individuals could lead to an increased gene expression, compared with healthy individuals without a family background. Also different miRNAs have the potential to affect the expression of a gene. Therefore, it is also possible that the RNF213 expression in familial individuals has been affected in this way. Additionally, the expression comparison demonstrated a significant increase in familial compared with sporadic patients and a similar RNF213 gene expression level among sporadic and control group was seen. Therefore, it is concluded that only familial cases show a significant change in expression of this gene, and RNF213 may not be considered as a biomarker in sporadic MS cases. The averaged RNF213 expression is similar for MS and control groups. Here, by averaging all patients real differences are masked, resulting in the wrong conclusion, and misinterpretation of the data. But we prefere to provide these results to show the need to classify patients into sporadic and familial categories in the future studies. However, since no studies have been found to investigate the expression level of RNF213 in MS versus healthy individuals, the results of the study are not comparable. It seems that by examining this gene in different populations and in a larger statistical population, more reliable results will be achieved. Additionally, we could find a study that investigated the association of RNF213 with MS pathogenesis, which was conducted in 2021. In this study, multiple variants in several genes, including RNF213, were introduced as an effective factor in the development of MS42.
This research is the first study to examine the expression of IL7R, NFATc2 and RNF213 in MS patients versus healthy controls. Also, it is the first study that has evaluated the expression level of IL7R, NFATc2 and RNF213 among MS groups with genetic classification, to investigate the effect of hereditary factors involved in the familial aggregation of the disease. However, this research also has some limitations, which can be pointed out by the small statistical population of the participants. Further studies in a larger group are necessary to determine whether IL7R, NFATc2 and RNF213 can be used as biomarkers for the prognosis of MS.
Conclusions
Today, many genetic tests use prognostic biomarkers to predict disease before clinical manifestation, to inform family members at risk. In this study, an attempt was made to help the prognosis of people at risk. Therefore, this research has investigated the potential role of IL7R, NFATc2 and RNF213 as prognostic markers in MS patients. According to the results of our study, the down regulation of IL7R can be considered as a biomarker for sporadic but not familial MS patients. In the case of NFATc2, there is no difference between controls, nor between MS groups. Therefore, its upregulation is associated with the disease in both familial and sporadic MS. Additionally, only familial cases show a significant change in RNF213 expression. But, since there is no difference between the diseased and healthy relatives, it couldn't be used as a prognostic biomarker in MS cases.
The results of our study will improve the development of appropriate treatment plans for sporadic and familial MS patients separately and plays a significant role in the introduction of potential prognostic markers. Also, in the future, similar studies will have interesting implications in personalizing the treatments.
References
Ehtesham, N., Khorvash, F. & Kheirollahi, M. miR-145 and miR20a-5p potentially mediate pleiotropic effects of interferon-beta through mitogen-activated protein kinase signaling pathway in multiple sclerosis patients. J. Mol. Neurosci. 61, 16–24 (2017).
Baecher-Allan, C., Kaskow, B. J. & Weiner, H. L. Multiple sclerosis: Mechanisms and immunotherapy. Neuron 97, 742–768 (2018).
Varatharaj, A. & Galea, I. The blood-brain barrier in systemic inflammation. Brain Behav. Immun. 60, 1–12 (2017).
Kunkl, M., Frascolla, S., Amormino, C., Volpe, E. & Tuosto, L. T helper cells: The modulators of inflammation in multiple sclerosis. Cells 9, 482 (2020).
Al-Eitan, L., Qudah, M. A. & Qawasmeh, M. A. Association of multiple sclerosis phenotypes with single nucleotide polymorphisms of IL7R, LAG3, and CD40 genes in a Jordanian population: A genotype-phenotype study. Biomolecules 10, 356 (2020).
Dendrou, C. A., Fugger, L. & Friese, M. A. Immunopathology of multiple sclerosis. Nat. Rev. Immunol. 15, 545–558 (2015).
Kay, M., Hojati, Z. & Dehghanian, F. The molecular study of IFNβ pleiotropic roles in MS treatment. Iran. J. Neurol. 12, 149 (2013).
Patsopoulos, N. A. Genetics of multiple sclerosis: An overview and new directions. Cold Spring Harbor Perspect. Med. 8, a028951 (2018).
Olerup, O. & Hillert, J. HLA class II-associated genetic susceptibility in multiple sclerosis: A critical evaluation. Tissue Antigens 38, 1–15 (1991).
Consortium, I. M. S. G. A systems biology approach uncovers cell-specific gene regulatory effects of genetic associations in multiple sclerosis. Nat. Commun. 10 (2019).
Patsopoulos, N. A. et al. Fine-mapping the genetic association of the major histocompatibility complex in multiple sclerosis: HLA and non-HLA effects. PLoS Genet. 9, e1003926 (2013).
Galarza-Muñoz, G. et al. Human epistatic interaction controls IL7R splicing and increases multiple sclerosis risk. Cell 169, 72-84e13 (2017).
Sahami-Fard, M. H., Mozhdeh, M., Izadpanah, F., Kashani, H. H. & Nezhadi, A. Interleukin 7 receptor T244I polymorphism and the multiple sclerosis susceptibility: A meta-analysis. J. Neuroimmunol. 341, 577166 (2020).
Lundmark, F. Genetic Analysis of IL7R and Other Immune-Regulatory Genes in Multiple Sclerosis (Institutionen för klinisk neurovetenskap/Department of Clinical Neuroscience, 2007).
Timasheva, Y. et al. Association between allelic variants of IL2, IL2RA, and IL7R genes and multiple sclerosis. Russ. J. Genet. 55, 487–494 (2019).
Meyer, I. S. & Leuschner, F. The role of Wnt signaling in the healing myocardium: A focus on cell specificity. Basic Res. Cardiol. 113, 44 (2018).
Weider, M. et al. Nfat/calcineurin signaling promotes oligodendrocyte differentiation and myelination by transcription factor network tuning. Nat. Commun. 9, 1–16 (2018).
Scholz, B. et al. Endothelial RSPO3 controls vascular stability and pruning through non-canonical WNT/Ca2+/NFAT signaling. Dev. Cell 36, 79–93 (2016).
Vilariño-Güell, C. et al. Exome sequencing in multiple sclerosis families identifies 12 candidate genes and nominates biological pathways for the genesis of disease. PLoS Genet. 15, e1008180 (2019).
Thompson, A. J. et al. Diagnosis of multiple sclerosis: 2017 revisions of the McDonald criteria. Lancet Neurol. 17, 162–173 (2018).
Kamitsuji, S. & Kamatani, N. Estimation of haplotype associated with several quantitative phenotypes based on maximization of area under a receiver operating characteristic (ROC) curve. J. Hum. Genet. 51, 314–325 (2006).
Nutz, S., Döll, K. & Karlovsky, P. Determination of the LOQ in real-time PCR by receiver operating characteristic curve analysis: Application to qPCR assays for Fusarium verticillioides and F. proliferatum. Anal. Bioanal. Chem. 401, 717–726 (2011).
Patsopoulos, N. A. & De Jager, P. L. Genetic and gene expression signatures in multiple sclerosis. Mult. Scler. J. 26, 576–581 (2020).
Khalilian, S. et al. Gene expression profiles of YAP1, TAZ, CRB3, and VDR in familial and sporadic multiple sclerosis among an Iranian population. Sci. Rep. 11, 1–10 (2021).
Benešová, Y., Vašků, A. & Bienertová-Vašků, J. Association of interleukin 6, interleukin 7 receptor alpha, and interleukin 12B gene polymorphisms with multiple sclerosis. Acta Neurol. Belg. 118, 493–501 (2018).
Gregory, S. G. et al. Interleukin 7 receptor α chain (IL7R) shows allelic and functional association with multiple sclerosis. Nat. Genet. 39, 1083–1091 (2007).
Oreja-Guevara, C., Ramos-Cejudo, J., Aroeira, L. S., Chamorro, B. & Diez-Tejedor, E. TH1/TH2 cytokine profile in relapsing-remitting multiple sclerosis patients treated with glatiramer acetate or natalizumab. BMC Neurol. 12, 1–6 (2012).
Lei, X. et al. Down-regulation of interleukin 7 receptor (IL-7R) contributes to central nervous system demyelination. Oncotarget 8, 28395 (2017).
Lundmark, F. et al. Variation in interleukin 7 receptor α chain (IL7R) influences risk of multiple sclerosis. Nat. Genet. 39, 1108–1113 (2007).
Traboulsee, A. L. et al. Genetic variants in IL2RA and IL7R affect multiple sclerosis disease risk and progression. Neurogenetics 15, 165–169 (2014).
Booth, D. et al. Gene expression and genotyping studies implicate the interleukin 7 receptor in the pathogenesis of primary progressive multiple sclerosis. J. Mol. Med. 83, 822–830 (2005).
Bina, P., Kakhki, M. P., Sahraian, M. A. & Behmanesh, M. The expression of lnc-IL-7R long non-coding RNA dramatically correlated with soluble and membrane-bound isoforms of IL-7Ra gene in multiple sclerosis patients. Neurosci. Lett. 642, 174–178 (2017).
Luo, D. & Fu, J. Identifying characteristic miRNAs-genes and risk pathways of multiple sclerosis based on bioinformatics analysis. Oncotarget 9, 5287 (2018).
Soomro, S. H., Jie, J. & Fu, H. Oligodendrocytes development and Wnt signaling pathway. Int. J. Hum. Anat 1, 17–35 (2018).
Staal, F. J., Luis, T. C. & Tiemessen, M. M. WNT signalling in the immune system: WNT is spreading its wings. Nat. Rev. Immunol. 8, 581–593 (2008).
Xie, C., Li, Z., Zhang, G.-X. & Guan, Y. Wnt signaling in remyelination in multiple sclerosis: Friend or foe?. Mol. Neurobiol. 49, 1117–1125 (2014).
Chae, W.-J. & Bothwell, A. L. Canonical and non-canonical Wnt signaling in immune cells. Trends Immunol. 39, 830–847 (2018).
Park, Y.-J., Yoo, S.-A., Kim, M. & Kim, W.-U. The role of calcium–calcineurin–NFAT signaling pathway in health and autoimmune diseases. Front. Immunol. 11, 195 (2020).
Becker, M. et al. Impaired mast cell-driven immune responses in mice lacking the transcription factor NFATc2. J. Immunol. 182, 6136–6142 (2009).
Liu, W. et al. Identification of RNF213 as a susceptibility gene for moyamoya disease and its possible role in vascular development. PLoS ONE 6, e22542 (2011).
Deshaies, R. J. & Joazeiro, C. A. RING domain E3 ubiquitin ligases. Annu. Rev. Biochem. 78, 399 (2009).
Alkhateeb, A. M., Salman, D. S. & Al-Hayk, K. A. Exome sequencing analysis of familial cases of multiple sclerosis and a monozygotic discordant twin. Arab. J. Sci. Eng. 46, 1–7 (2021).
Acknowledgements
The authors appreciate the volunteers for their participation.
Funding
This study was performed at the University of Isfahan (Isfahan, Iran) and was financially supported by the Graduate Studies Offices at this university.
Author information
Authors and Affiliations
Contributions
S.Z.H., S.K., F.D., ME.K. And Z.H. conceived and planned the experiments. S.Z.H., S.K. and F.D. carried out the experiments. S.Z.H., S.K., ME.K., MA.K., O.M. and V.S. contributed to sample preparation and patient's data collection. S.Z.H. analyzed the results. S.Z.H., S.K., F.D. and Z.H. contributed to the interpretation of the results. S.Z.H., S.K., took the lead in writing the manuscript. O.M. carefully revised and Z.H finally approved the manuscript. All authors contributed to and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Imani, S.Z.H., Hojati, Z., Khalilian, S. et al. Expression and clinical significance of IL7R, NFATc2, and RNF213 in familial and sporadic multiple sclerosis. Sci Rep 11, 19260 (2021). https://doi.org/10.1038/s41598-021-98691-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-98691-5
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.