Abstract
Bifidobacteria, which commonly inhabit the primate gut, are beneficial contributors to host wellbeing. Anatomical differences and natural habitat allow an arrangement of primates into two main parvorders; New World monkeys (NWM) and Old World monkeys (OWM). The number of newly described bifidobacterial species is clearly elevated in NWM. This corresponds to our finding that bifidobacteria were the dominant group of cultivated gut anaerobes in NWM, while their numbers halved in OWM and were often replaced by Clostridiaceae with sarcina morphology. We examined an extended MALDI-TOF MS database as a potential identification tool for rapid screening of bifidobacterial distribution in captive primates. Bifidobacterial isolates of NWM were assigned mainly to species of primate origin, while OWM possessed typically multi-host bifidobacteria. Moreover, bifidobacterial counts reflected the feed specialization of captive primates decreasing from frugivore-insectivores, gummivore-insectivores, frugivore-folivores to frugivore-omnivores. Amplicon sequencing analysis supported this trend with regards to the inverse ratio of Actinobacteria and Firmicutes. In addition, a significantly higher diversity of the bacterial population in OWM was found. The evolution specialization of primates seems to be responsible for Bifidobacterium abundance and species occurrence. Balanced microbiota of captive primates could be supported by optimized prebiotic and probiotic stimulation based on the primate host.
Similar content being viewed by others

Introduction
Primates are a remarkably species-rich order of mammals1. Their anatomical differences and natural habitat allow their arrangement into two main parvorders. Platyrrhines, referred as New World monkeys (NWM), naturally occurring in central and southern American tropical and subtropical regions and catarrhines (Cercopithecoidea and Hominoidea), referred as Old World monkeys (OWM), coming from tropical, subtropical, and temperate regions of Asia and Africa2. Many primate species are endangered3 and they must be protected. The conservation of threatened species is a complex and demanding process consisting of elaborated breeding programs and providing of habitat sanctuaries in captive or semi-captive centres, e.g. zoological institutions or forest corridors, which usually aim to reintroduce these species back into their natural habitat4,5. Unfortunately, health of captive animals is compromised by emerging recurring infectious diseases mediated through human contact and habitat modifications, and frequent therapeutic doses of antibiotics6,7. Furthermore, captive breeding modifies primate microbiome8,9 and these microbial shifts can substantially affect the host’s health10,11. Captivity may be also associated with the occurrence of potential pathogens that further increase risk of gut dysbiosis and illnesses12,13.
Besides exposure to antibiotics, dietary changes and lifestyle seem to be significant modifiers of primate gut microbiome14. To provide nutritional needs, primates consume a wide range of plants and animal tissues and possess a variety of dietary specializations based on the proportion of individual dietary components (one type of feed component is dominating only), such as generalist feeders or omnivores, e.g. Cercopithecines15,16,17,18. The generalist feeders are adapted to receive a wide variety of feed components, depending on their availability in the environment, and can be split by extension into groups classified by their majority feeds, with seasonal variation in their ratio. Among the generalist feeders, there are highly frugivorous representatives, namely chimpanzees19,20,21. If there is a lack of fruit, these primates consume various feed reaching from plants, nectar, seeds to insects or small vertebrates. Such a feeding type can be described as frugivore-omnivore. If the preferred fruit is less available during the season, primates start to consume more leaves or other parts of plants. Gibbons, for instance pursue this frugivore-folivore feeding strategy22,23,24,25. Similarly, if the second major component alongside fruit consists of insects, primates are classified as frugivores-insectivores (tamarins)26,27,28,29,30. Exudates are another important nutritious feed apart from fruit and animal prey. Some primate species have specially adapted teeth for gum intake31,32. This type of feeding behaviour is called gummivory. It is typical for marmosets and can either be dominant or it can be supplemented with insect intake33,34,35,36,37. These primates are counted in the gummivore-insectivore feeding category.
Unfortunately, despite all efforts of breeders, composition of diet in captivity does not completely simulate that in the wild, in which primates consume a wider range of natural local plant and animal species9,38. In addition, Amato et al.39 points out the seasonality that is one of the natural phenomena of wild primate diet, which results in a seasonal variation of the gut microbiome.
Deviation from the natural lifestyle in captivity and associated modified diet led to a shift of native gut microbiota and a decrease in diversity and an increased relative abundance of Bacteroidetes8,9,40,41. Furthermore, the microbiome of captive primates displays a reduction in Actinobacteria compared to wild groups14,41. However, members of the Bifidobacteriaceae family (Actinobacteria phylum) are important natural commensals, which possess a large amount of adaptive genes involved in carbohydrate metabolism42,43,44. Moreover, bifidobacteria can utilize a diverse range of dietary carbohydrates that escape degradation in the upper parts of the intestine45.
Although, bifidobacterial abundance in the gut microbiota usually decreases with host aging46, bifidobacteria persist throughout the lifespan of primates42,47. Moreover, their abundance is confirmed by a recent boom of novel bifidobacterial species isolation and characterization connected to primate gut environment48,49,50.
However, data are still scarce about the bifidobacterial microbiota of captive primates and the impact of different diets. We hypothesize that the quantity and species richness of bifidobacteria in captive primates are affected by the host and feed classification. The aim of this study was to compare the quantity and diversity of bifidobacteria in faecal microbiota of captive NWM and OWM by a combination of culture-dependent and culture-independent approaches.
Results
Cultivation analysis
Quantification of cultivable bifidobacteria in primate faecal samples
Non-selective and selective media were used for the quantification of anaerobic bacteria and bifidobacteria in primate faecal samples (FS) (Table 1). Cultivation counts significantly varied between the NWM and the OWM in each monitored group of bacteria (Fig. 1A, Suppl. Tab. 1). NWM harboured significantly more anaerobic bacteria (9.52 ± 0.62 log CFU g-1) compared to OWM (8.62 ± 0.71 log CFU g-1) (t(50) = 4.84, p = 1.30e-05). A similar statistically significant trend was found in colony forming units cultivated on WPS-MUP medium intended for bifidobacteria that reached 8.91 ± 1.38 log CFU g-1 in the NWM compared to 7.02 ± 0.93 log CFU g-1 in the OWM (t(50) = 5.87, p = 3.50e-07). In case of FS with lower numbers of bifidobacteria and the presence of clostridia, this medium was not sufficiently selective also allowing the growth of clostridia51,52. Consequently, a notably greater statistically significant difference was detected between primate parvorders on more selective WSP-NORF medium with bifidobacterial counts of 8.57 ± 2.13 log CFU g-1 for the NWM and 4.32 ± 2.04 log CFU g-1 for the OWM (Z = 5.17, p = 2.38e-07). Cultivation differences between parvorders were also reflected within the primate sub-division based on feed specialization (Fig. 1B). Specifically, gummivore-insectivores (9.63 ± 0.71 log CFU g-1) and frugivore-insectivores (9.46 ± 0.57 log CFU g-1) exhibited significantly higher numbers of anaerobic bacteria including bifidobacteria than frugivore-folivores (8.72 ± 0.49 log CFU g-1) and frugivore-omnivores (8.60 ± 0.78 log CFU g-1). The same statistically significant trend was found on WPS-MUP in gummivore-insectivores (8.99 ± 1.19 log CFU g-1) and frugivore-insectivores (9.19 ± 0.96 log CFU g-1) in comparison with frugivore-folivores (6.58 ± 1.05 log CFU g-1) and frugivore-omnivores (7.07 ± 1.01 log CFU g-1), as well as on WSP-NORF in gummivore-frugivores (8.46 ± 2.34 log CFU g-1) and frugivore-insectivores (9.15 ± 0.76 log CFU g-1) compared to frugivore-folivores (4.29 ± 1.95 log CFU g-1) and frugivore-omnivores (4.22 ± 2.13 log CFU g-1) (Supplementary S5).
Quantification of cultivable anaerobic bacteria (log CFU g-1) in primate faecal samples. (A) Cultivation counts of bacteria per parvorder: New World monkeys (n = 24) and Old World monkeys (n = 28). (B) Cultivation counts of bacteria per feed category: frugivore-folivore (n = 8), frugivore-omnivore (n = 21), frugivore-insectivore (n = 13), gummivore-insectivore (n = 10). Asterisks (*) denote statistically significant differences as determined by t-test and ANOVA (p < 0.05).
Bifidobacterial species detected by MALDI-TOF MS
Bacterial colonies with variable cultivation characteristics from bifidobacterial selective media were isolated for further identifications (Suppl. Tab. 1). From a total of 326 isolates, 210 were F6PPK-positive bifidobacteria and the remaining 116 isolates (isolated mainly from WSP-MUP) were F6PPK-negative gas producing clostridial rods or cells with sarcina morphology. All F6PPK-positive strains were also identified with MALDI-TOF MS using an expanded custom database for bifidobacterial identification. 54% of the strains (n = 112) were assigned to 18 different bifidobacterial species, 36% (n = 76) were assigned only to the Bifidobacterium genus, and 11% (n = 22) were not identified reliably (Fig. 2A, C).
MALDI-TOF MS identification of primate bifidobacterial isolates. (A) MALDI-TOF MS identification of 210 bifidobacterial strains. (B) The closest probable species match of isolates with unambiguous genus MALDI-TOF MS identification (Bifidobacterium spp). (C) Proportion of species assignment, genus assignment and not reliable identification (NRI) of bifidobacterial isolates. Bruker criteria (scores) for assignment: 0.000–1.699 not reliable identification, 1.700–1.999 probable genus identification, 2.000–3.000 genus and species identification.
B. parmae, B. imperatoris/saguini, and B. ramosum were the most frequently identified species in the NWM, whereas B. dentium and B. catenulatum/pseudocatenulatum were most common in the OWM. Interestingly, B. adolescentis was equally represented in both primate parvorders. A more diverse species representation of bifidobacteria was found in the NWM (14 spp.) compared to the OWM (5 spp.). Genus-level assignment and the presence of not reliable identifications (NRI) was mainly detected in the NWM. Related presumed species compliance and the closest match of Bifidobacterium spp. strains was found predominantly with B. parmae and B. stellenboschense in the NWM, and B. angulatum/merycicum in the OWM (Fig. 2B).
Species assignment verification by 16S rRNA gene sequencing
The MALDI-TOF MS identification was verified by 16S rRNA gene Sanger sequencing of 46 strains, whose selection was randomly executed based on determined species frequency and identification scores (Suppl. Tab. 2). Due to similar MALDI-TOF MS spectra, some bifidobacterial species could not be distinguished. However, the results consistently suggest an assignment to either of the two indistinguishable species. These indistinguishable groups were merged to produce consistent MALDI-TOF MS assignment and are presented together in the following groups: B. angulatum/merycicum, B. breve/indicum, B. catenulatum/pseudocatenulatum, and B. imperatoris/saguini.
An agreement between the MALDI-TOF MS species assignment and the sequencing of 16S rRNA gene was confirmed for 38 strains. Only 3 strains were identified differently by the two methods. Namely, strain N127 identified as B. faecale by 16S rRNA gene sequencing was mistaken for B. adolescentis by the MALDI-TOF MS, B. imperatoris for NRI (N40), and PEBJ_s for B. imperatoris/saguini (N50). Interestingly, mentioned strain N50 together with N74, N94, N97, and N115, exhibiting MALDI-TOF MS NRI score (< 1.69), were considered potential novel species of bifidobacteria. In addition, this sample set also contained 5 problematic strains (N16, N70, N81, N119, and N125), whose 16S rRNA gene sequencing failed repeatedly and thus their MALDI-TOF MS identity was not confirmed.
Amplicon sequencing analysis
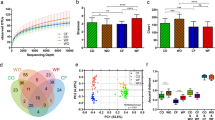
Amplicon sequencing profiles of the FS collected from captive primates were determined by sequencing the V4 region of the 16S rRNA gene. The bacterial α-diversity was expressed as an ASV count, Shannon diversity, and Pielou evenness. Each diversity parameter between the primate parvorders was significantly higher in the OWM (ASV count: F(1,50) = 30.47, p = 1.21 × 10–6, η2 = 0.379, Shannon: F(1,50) = 38.01, p = 1.21e-07, η2 = 0.432, Pielou: F(1,50) = 38.41, p = 1.08e-07, η2 = 0.434) (Fig. 3A). Similarly, there was a significantly higher diversity, evenness, and richness of the bacterial population in the frugivore-folivores and frugivore-omnivores compared to the frugivore-insectivores and gummivore-insectivores (Fig. 3B, Supplementary S1).
Alfa-diversity of primate gut microbiota. (A) Bacterial α-diversity per parvorder: New World monkeys and Old World monkeys. (B) Bacterial α-diversity per feed category: frugivore-folivore, frugivore-omnivore, frugivore-insectivore, gummivore-insectivore. Asterisks (*) denote adjusted statistically significant differences (adj. p < 0.05).
Microbial community shifts were found between the NWM and OWM parvorders. The relative abundance of phylum Actinobacteriota (W = 13) and Campylobacterota (W = 12) was significantly higher in the NWM compared to the OWM as confirmed by the ANCOM statistics. Meanwhile, the phylum Firmicutes showed an opposite trend, which was however not statistically significant (Fig. 4A). The difference in the Actinobacteriota can be attributed specifically to the family Bifidobacteriaceae which was significantly higher in the NWM (16%) compared to the OWM (3%) (W = 139) (Fig. 4B, Supplementary S2). These findings corroborate the cultivation results.
Relative abundance of primate gut microbiota. (A) Relative abundance of bacteria within parvorders on phylum level. (B) Relative abundance of bacteria within parvorders on family level. (C) Relative abundance of bacteria within feed categories on phylum level. (D) Relative abundance of bacteria within feed categories on family level. ANCOM statistically significant differences are denoted with grey links.
The proportion of the phylum Actinobacteriota was statistically significantly different among the primate feed categories (W = 12) and it was the highest in the frugivore-insectivores followed by gummivore-insectivores, frugivore-omnivores, and frugivore-folivores. Furthermore, the phyla Proteobacteria and Campylobacterota were statistically significantly different among the categories (W = 8, W = 7 respectively) with a notable enrichment of both in the frugivore-insectivores followed by gummivore-insectivores compared to frugivore-omnivores and frugivore-folivores. Moreover, although not statistically significant, the opposite ratio of Firmicutes was also detected (Fig. 4C). The relative abundance of Bifidobacteriaceae was significantly different across the categories (W = 140); the most abundant in the frugivore-insectivores (19%), followed by the gummivore-insectivores (12%), the frugivore-omnivores (4%), and the frugivore-folivores (2%) (Fig. 4D).
By comparing 16S rRNA gene sequencing data of cultured bifidobacterial isolates with the results of 16S rRNA gene amplicon sequencing of the FS, we retrospectively confirmed the presence of 18 species within this sample set. B. callitrichos and B. parmae were significantly enriched in the NWM (W = 38, W = 38 respectively), followed by B. saguini (W = 34), B. biavatii (W = 34), B. vansinderenii (W = 34), B. aerophilum (W = 34), unclassified II ASV (W = 33) and sp. I ASV (W = 30). The distribution of bifidobacteria corresponds to the proportion of Bifidobacteriaceae among the total relative bacteria in samples normalized to 42 134 sequences/sample in the primate feed categories as determined by amplicon sequencing.
Discussion
Dynamic microbial communities aid the living and surviving of animals in changing environmental conditions, including habitat degradation, captive breeding, and diet. If microbial balance of the host is disturbed and dysbiosis occurs, there is a presumption of disease development5,53,54. Among others, commensal microorganisms, such as bifidobacteria, play a crucial role in maintaining the gut homeostasis55,56,57. Bifidobacterial diversity and adaptation are connected to their hosts and environments with possession of specific genomic traits58,59,60 which includes primates42.
Two independent approaches, cultivation with subsequent MALDI-TOF MS identification and amplicon sequencing of the V4 region of the 16S rRNA gene, were used to analyse the microbiome composition and the prevalence of bifidobacterial species in primate gut microbiota. NWM are a significant source of cultivable bifidobacteria with average counts of 108 CFU g-1 of faeces compared to the OWM with four orders of magnitude lower counts. Interestingly, although no health complications were evident, FS of primate individuals with reduced or undetectable cultivation counts of bifidobacteria contained Clostridiaceae, mainly displaying sarcina morphology. This was mainly observed in individuals belonging to the OWM parvorder (Suppl. Tab. 1). Spore-forming bacteria identified as Sarcina ventriculi (syn. Clostridium ventriculi) were previously isolated also from primates without apparent health problems61,62,63. Although they are considered pathogens64, this may indicate sarcina as common bacteria of the primate gut microbiota. In the gut of NWM, the abundance of sarcina is probably decreased by the presence of bifidobacteria, which exhibit potential to hamper growth of clostridia65,66,67. The inverse ratio and balancing of the bifidobacteria and clostridia are typically described in the gut microbiome of infants68,69,70.
Timperio et al.71 showed that the screening of bacterial isolates from environmental samples can be performed efficiently, quickly, and inexpensively using MALDI-TOF MS and should be refined by implementation of environmental strains into the database. Within our study, the use of an extended custom database for MALDI-TOF MS allowed reliable species differentiation and identification of wild bifidobacterial isolates. Higher species diversity was observed in NWM. Interestingly, the multi-host species B. adolescentis was present among most screened captive primates. In OWM B. dentium and B. catenulum/pseudocatenulatum, that are common species of the human gut microbiota, as well as B. adolescentis, were found72. Lugli et al.42 detected B. adolescentis and B. dentium in OWM as well, and indicated possible joint development and evolutionary relatedness. In contrast, NWM exhibited the presence of cultivable bifidobacteria mainly with primate origin. Interestingly, Brown et al.73 pointed out that marmoset bifidobacteria are closely related to those in tamarins. Furthermore, we found that bifidobacterial species variability in NWM significantly exceeds that in OWM. Furthermore, we hereby confirmed that we can re-isolate recently described primate Bifidobacterium spp. also from primate species with various captive locations other than those from which bifidobacteria were originally isolated.
Moreover, MALDI-TOF MS screening allowed us to identify 5 potential novel species of bifidobacteria isolated from tamarins that were confirmed by 16S rRNA gene sequencing. That indicates primate gut as a promising environment for the discovery of novel species of bifidobacteria42,48,50. To achieve an accurate identification of potential novel species, a combination with other methods, such as sequencing of phylogenetic markers74,75,76, multi-locus sequence typing77, and genome sequencing78, should be included.
The significantly lower species richness and high relative abundance of bifidobacteria in NWM compared to OWM was confirmed by sequencing of the V4 region of 16S rRNA gene. The relative abundance of Bifidobacteriaceae reached 16% in the NWM and only 3% in the OWM. The same trend was also detected for Prevotellaceae and Veillonellaceae. In particular, marmosets and tamarins exhibited 32% bifidobacterial abundance compared to 0.03% in the OWM42. This high relative bifidobacterial proportion in adult marmosets could be a consequence of their housing as family groups and their constant subjection to the gut microbiota of other individuals73. Conversely, Lachnospiraceae, Oscillospiraceae, Ruminococcaceae, and Spirochaetaceae showed an opposite trend with high abundances in OWM. Interestingly, we showed that the captive NWM have high relative levels of bifidobacteria, which is similar to what they display in the wild47,79,80,81. It indicates that NWM gut is a rich bifidobacterial environment that is also supported by other studies42,82,83. In contrast to our results in captive individuals, some microbiome studies point to a slightly increased bifidobacterial relative proportions in wild OWM as well84,85. Although the captivity was previously described as a factor influencing the presence of Actinobacteria in the primate gut microbiome14,41, our results suggest that it is probably not as strong as the affiliation to the primate parvorder, which seems to be considerably more significant.
Primate gut microbiome seems to be significantly modified by dietary changes of the host species and geography14. Frugivore-insectivores and gummivore-insectivores possessed significantly more abundant Bifidobacteriaceae compared to frugivore-omnivores and frugivore-folivores. Interestingly, if insects constitute an important component of the diet, bifidobacteria are highly abundant. Ecologically beneficial symbionts leading to host evolutionary dependence have been previously described in other animal taxa, such as sap-feeding insects, which generate essential amino acids exclusively for their microbial symbionts86. Bifidobacteria are known as a commensal bacterial group of insects with social life87, whereas the importance of insects in the diet of primates in relation to bifidobacterial occurrence remains unclear.
Although captive feeding inevitably modifies primate gut microbiome to decreased diversity, the feed optimization could improve the animals health condition40. In contrast to Amato et al.88, who state that the host phylogeny is stronger driver in shifts of microbial composition than the diet and geographic location, our results suggest that both diet and the host itself affect the microbiome composition, especially the relative abundance of Bifidobacteriaceae. Moreover, it is important to mention, that the diet of captive animals usually includes fruits, vegetables, and leaves that may not completely match the available components present in the wild. In addition, the natural microbiota reflects diet seasonality and location that may affect trophic interactions in the gastrointestinal tract of the host89,90.
Clayton et al.91 confirms that modified diet in captive primates is related to the alteration of microbiome composition and host health. Captive primate individuals susceptible to health disorders may show clinical signs including chronic diarrhoea, weight loss, lethargy, cardiac disease, and poor reproductive success9,12,92,93. Therefore, it is necessary to further monitor the relationship between the microbiome, diet, and the health of captive primates40. Microbiota modulation is an effective and affordable strategy for host health support of threatened animals5. Therefore, applicable mitigation strategies such as optimized dietary40 and prebiotic interventions94 could be pursued towards supporting balanced microbiota in captive primates. Moreover, probiotic supplementation with focus on bifidobacteria, that naturally colonize primate guts, can be a further promising approach42,43,95. Furthermore, this may provide a potential approach in human probiotic intervention. Due to the ever-decreasing diversity of the human microbiome through diet and antimicrobial intake, the microbiome of originally living evolutionarily close relatives has the potential to design a probiotic that is no longer part of the human microbiota and could have the potential to strengthen health96. Probiotic intervention should be optimized according to the gut microbiota composition and should be supported by appropriately selected prebiotic stimulation in synbiotic mixtures for long-term maintenance of balanced microbiome and host health.
Materials and methods
Sampling and cultivation analysis
Faecal samples of primate hosts (n = 52) belonging to two parvorders, NWM (n = 24) and OWM (n = 28), were preliminary screened for quantitative content of cultivable bifidobacteria. The list of primate hosts and classification into parvorders and feed category is shown in Table 1. Sampling was performed in zoological gardens in Dvur Kralove, Hodonin, Liberec, Olomouc, Pilsen (all Czechia), Bojnice, and Bratislava (both Slovakia) between 2017–2019. FS were collected in tubes containing dilution buffer (5 g L-1 tryptone, 5 g L-1 nutrient broth No. 2, 2.5 g L-1 yeast extract (all Oxoid, Basingstoke, UK), 0.5 g L-1 L-cysteine, 1 mL L-1 Tween 80 (both Sigma-Aldrich, St. Louis, Missouri, USA), 30% glycerol (VWR, Radnor, Pennsylvania, USA), and glass pearls for homogenization. Media were prepared in an oxygen‐free carbon dioxide environment97 and then sterilized. After sampling, the tubes were stored at –20 °C and within the 14 days transported into the laboratory for analysis. Then, decimal serial dilutions of FS were spread on the following media.
Wilkins-Chalgren Anaerobe Agar was supplemented with 5 g L-1 GMO-Free Soya Peptone (both Oxoid), 0.5 g L-1 L-cysteine, and 1 mL L-1 Tween 80 to determine total counts of anaerobic bacteria (WSP medium). Moreover, two selective media were used for bifidobacterial quantification and isolation: WSP-NORF (WSP agar supplemented with 100 mg L-1 of mupirocin, 200 mg L-1 of norfloxacin (both Oxoid), and 1 mL L-1 of acetic acid (Sigma-Aldrich)52) and WSP-MUP (WSP agar supplemented with 100 mg L-1 of mupirocin and 1 mL L-1 of acetic acid98). All plates were incubated anaerobically using GENbag anaer (bioMérieux, Craponne, France) at 37 °C for 2 days.
Isolation and culture identifications
Based on variable cultivation characteristics, the isolation of colonies from selective media and consecutive sub-cultivation was performed in tubes containing WSP broth under anaerobic conditions97 at 37 °C for 1 day. Whether a culture belonged to Bifidobacterium spp. was verified by fructose-6-phosphate phosphoketolase (F6PPK) test with cetrimonium bromide for cell disruption according to Orban and Patterson (2000)99. Subsequently, bifidobacterial isolates were identified to the species level using Matrix-Assisted Laser Desorption/Ionization Mass Spectrometry (MALDI-TOF MS) with ethanol-formic acid extraction procedure with HCCA matrix solution according to the manufacturer’s instructions (Bruker Daltonik GmbH, Bremen, Germany). An extended custom database (based on Bruker Biotyper software tools), which included 50 additional bifidobacterial species in addition to the already available entries, was used for identification. An overview about the database entries is provided in Suppl. Tab. 3. Stock cultures of bifidobacteria were stored at –80 °C in 30% glycerol.
Selected isolates (n = 46) were further identified by 16S rRNA gene amplicon sequencing. DNA was isolated from freshly grown bifidobacterial cultures in WSP broth using PrepMan Ultra™ (Applied Biosystems, Waltham, Massachusetts, USA) according to manufacturer's instructions and stored at –20 °C. Primers 285F (5′-GAGGGTTCGATTCTGGCTCAG-3′) and 261R (5′-AAGGAGGTGATCCAGCCGCA-3′) were used for PCR amplification of nearly the full 16S rRNA gene according to Kim et al.100 enabling longer reads and thus more precise taxonomic identification. PCR products were purified using the E.Z.N.A. Cycle Pure Kit (Omega Bio-Tek, Norcross, Georgia, USA) and sequenced by Eurofins Genomics (Ebersberg, Germany). The obtained sequences were processed in Chromas Lite 2.5.1 (Technelysium Pty Ltd., Tewantin, Australia), BioEdit101 with ClustalW algorithm102, and compared with 16S rRNA gene sequences in BLAST rRNA/ITS (https://blast.ncbi.nlm.nih.gov/) and EZBioCloud databases (https://www.ezbiocloud.net/). The sequences of the 16S rRNA gene are available in the GenBank database under accession numbers MN736337–341, 342, 344–346, 348, 350–355, 357–360, 363–365, 367, 369, 372–378, 381, 387–388, 390–392, and MW678772–74.
Amplicon sequencing analysis
Total genomic DNA was extracted from 200 mg of FS using the Fast DNA SPIN kit for soil (MP Biomedicals, Illkirch-Graffenstaden, France) according to the manufacturer's instructions. The DNA concentration of each sample was determined using the Qubit 1X dsDNA HS Assay Kit (Invitrogen, Paisley, UK) and a Qubit fluorometer. Subsequent library preparation and sequencing were performed by NovoGene (Cambridge, UK). As amplicon sequencing method supports only shorter fragments, the V4 region of the 16S rRNA gene (300 bp fragments) was amplified using primers 515F (5′-GTGCCAGCMGCCGCGGTAA-3′) and 806R (5′-GGACTACHVGGGTWTCTAAT-3′) and a Phusion High-Fidelity PCR Master Mix (New England Biolabs, Ipswich, Massachusetts, USA). The library was prepared using the NEB Next® UltraTM DNA Library Prep Kit for Illumina and paired-end 250 bp sequencing was performed using the NovaSeq machine (Illumina, San Diego, California, USA). The resulting sequences were submitted to the NCBI database with the accession number ERP128111. Amplicon sequence variants (ASV) were obtained using the DADA2 pipeline (bioconductor-dada2 v1.16.0)103 and Silva non redundant database v138104 (Supplementary S3) with custom manual species assignment. The depth of sequencing of the resulting data was normalized by rarefaction to the lowest sequencing depth (42 134 sequences/sample) and a relative abundance on several taxonomic levels in different variable groups were explored (Supplementary S4). Total bacterial diversity was expressed as Shannon entropy105, the population richness was expressed as simple feature or ASV counts and the evenness was expressed as Pielou’s index106.
Statistical analyses
Counts of bacterial colonies in log CFU g-1 within the parvorders and feed categories are shown as boxplots. The normality of data was evaluated by Shapiro–Wilk W test (α = 0.05). Differences in bacterial counts were assessed using a Mann–Whitney U Test (α = 0.05) within the parvorders, and a one-way ANOVA within the feed categories (α = 0.05) using STATISTICA software (StatSoft, Prague, Czechia) and Microsoft Office Professional Plus 2016.
To detect differentially abundant taxa between the sample categories, the ANCOM statistical test107 was used from the package skbio v0.5.2 (scikit-bio.org). The one-way F statistics from the scipy package v1.4.1108 was used to determine that statistical significance with α = 0.05. Several categories of the data were explored on both the Phylum and Family level. Furthermore, the bifidobacterial sub-population was extracted for each sample and the differentially abundant species were calculated. Statistically significant results are presented in form of boxplots (Supplementary S2).
The statistical significance of difference in means of the diversity metrics (Shannon, Pielou, and ASV counts) was assessed using the ordinary least squares method coupled with a pairwise T-test. The data was Box-Cox transformed and the resulting residuals were normally distributed (Jarque-Berra and Omnibus probability > 0.05), however, the groups were highly heteroskedastic. To mitigate this, we have used the ordinary least square method from the package statsmodels v0.11.0109 with MacKinnon and White’s heteroscedasticity robust standard errors110 (Supplementary S1).
Ethical approval
The sampling of primate faeces was performed during routine daily procedures. All procedures involving animals adhered to recommendations of the “Guide for the Care and Use of Animals” by the Czech University of Life Sciences Prague. The research conducted herein was approved by Ethic and Animal Care Committee of the Czech University of Life Sciences Prague (protocol number: CZU/17/19) and was performed in accordance with the relevant guidelines and regulations. All zoological institutions have rigorous standards for animal welfare and are accredited by the European Association of Zoos and Aquaria. The research adhered to the legal requirements of the Czech Republic for the ethical treatment of nonhuman primates as well as in accordance with European Directive 2010/63/EU.
References
Arbour, J. H. & Santana, S. E. A major shift in diversification rate helps explain macroevolutionary patterns in primate species diversity. Evolution 71, 1600–1613 (2017).
Groves, C. Primates (Taxonomy) in The International Encyclopedia of Primatology (ed Augustin Fuentes) (John Wiley & Sons, Inc., 2016).
Cotton, A., Clark, F., Boubli, J. & Schwitzer, C. IUCN red list of threatened primate species in An Introduction to Primate Conservation 31–18 (Oxford University Press, 2016).
Stumpf, R. M. et al. Microbiomes, metagenomics, and primate conservation: New strategies, tools, and applications. Biol. Conserv. 199, 56–66 (2016).
West, A. G. et al. The microbiome in threatened species conservation. Biol. Conserv. 229, 85–98 (2019).
Cunningham, A. A., Daszak, P. & Wood, J. L. N. One Health, emerging infectious diseases and wildlife: two decades of progress?. Philos. Trans. R. Soc. B: Biol. Sci. 372, 20160167 (2017).
Ramey, A. M. & Ahlstrom, C. A. Antibiotic resistant bacteria in wildlife: Perspectives on trends, acquisition and dissemination, data gaps, and future directions. J. Wildl. Dis. 56, 1–15 (2020).
Clayton, J. B. et al. Captivity humanizes the primate microbiome. Proc. Natl. Acad. Sci. 113, 10376–10381 (2016).
Hale, V. L. et al. Gut microbiota in wild and captive Guizhou snub-nosed monkeys. Rhinopithecus brelichi. Am. J. Primatol. 81, e22989 (2019).
Kriss, M., Hazleton, K. Z., Nusbacher, N. M., Martin, C. G. & Lozupone, C. A. Low diversity gut microbiota dysbiosis: drivers, functional implications and recovery. Curr. Opin. Microbiol. 44, 34–40 (2018).
Mahnert, A. et al. Man-made microbial resistances in built environments. Nat. Commun. 10, 1–12 (2019).
Amato, K. R. et al. Using the gut microbiota as a novel tool for examining colobine primate GI health. Glob. Ecol. Conserv. 7, 225–237 (2016).
Zhu, H. et al. Diarrhea-associated intestinal microbiota in captive Sichuan golden snub-nosed monkeys (Rhinopithecus roxellana). Microbes Environ. ME17163 (2018).
Campbell, T. P. et al. The microbiome and resistome of chimpanzees, gorillas, and humans across host lifestyle and geography. ISME J. 14, 1584–1599 (2020).
Buzzard, P. J. Ecological partitioning of Cercopithecus campbelli, C. petaurista, and C. diana in the Taï Forest. Int. J. Primatol. 27, 529–558 (2006).
Chapman, C. A. et al. The guenons: diversity and adaptation in African monkeys. 325–350 (Springer, 2004).
Krishnadas, M., Chandrasekhara, K. & Kumar, A. The response of the frugivorous lion-tailed macaque (Macaca silenus) to a period of fruit scarcity. Am. J. Primatol. 73, 1250–1260 (2011).
Swedell, L., Hailemeskel, G. & Schreier, A. Composition and seasonality of diet in wild hamadryas baboons: preliminary findings from Filoha. Folia Primatol. 79, 476–490 (2008).
Basabose, A. K. Diet composition of chimpanzees inhabiting the montane forest of Kahuzi, Democratic Republic of Congo. Am. J. Primatol. 58, 1–21 (2002).
McLennan, M. R. & Ganzhorn, J. U. Nutritional characteristics of wild and cultivated foods for chimpanzees (Pan troglodytes) in agricultural landscapes. Int. J. Primatol. 38, 122–150 (2017).
Newton-Fisher, N. E. The diet of chimpanzees in the Budongo Forest Reserve Uganda. Afr. J. Ecol. 37, 344–354 (1999).
Bach, T. H., Chen, J., Hoang, M. D., Beng, K. C. & Nguyen, V. T. Feeding behavior and activity budget of the southern yellow-cheeked crested gibbons (Nomascus gabriellae) in a lowland tropical forest. Am. J. Primatol. 79, e22667 (2017).
Fan, P.-F., Fei, H.-L., Scott, M. B., Zhang, W. & Ma, C.-Y. Habitat and food choice of the critically endangered cao vit gibbon (Nomascus nasutus) in China: implications for conservation. Biol. Conserv. 144, 2247–2254 (2011).
Fan, P. F., Fei, H. L. & Ma, C. Y. Behavioral responses of cao vit gibbon (Nomascus nasutus) to variations in food abundance and temperature in Bangliang, Jingxi China. Am. J. Primatol. 74, 632–641 (2012).
McConkey, K. R., Ario, A., Aldy, F. & Chivers, D. J. Influence of forest seasonality on gibbon food choice in the rain forests of Barito Ulu Central Kalimantan. Int. J. Primatol. 24, 19–32 (2003).
Amora, T. D., BeltrÃO-Mendes, R. & Ferrari, S. F. Use of alternative plant resources by common marmosets (Callithrix jacchus) in the semi-arid Caatinga scrub forests of northeastern Brazil. Am. J. Primatol. 75, 333–341 (2013).
Dietz, J. M., Peres, C. A. & Pinder, L. Foraging ecology and use of space in wild golden lion tamarins (Leontopithecus rosalia). Am. J. Primatol. 41, 289–305 (1997).
Garber, P. A. Feeding ecology and behaviour of the genus Saguinus. Marmosets and tamarins: systematics behaviour and ecology (1993).
Heymann, E. W., Knogge, C. & Tirado Herrera, E. R. Vertebrate predation by sympatric tamarins, Saguinus mystax and Saguinus fuscicollis. Am. J. Primatol. 51, 153–158 (2000).
Porter, L. M. Dietary differences among sympatric Callitrichinae in northern Bolivia: Callimico goeldii, Saguinus fuscicollis and S. labiatus. Int. J. Primatol. 22, 961–992 (2001).
Anapol, F. & Lee, S. Morphological adaptation to diet in platyrrhine primates. Am. J. Phys. Anthropol. 94, 239–261 (1994).
Nash, L. T. Dietary, behavioral, and morphological aspects of gummivory in primates. Am. J. Phys. Anthropol. 29, 113–137 (1986).
Abreu, F., De la Fuente, M. F. C., Schiel, N. & Souto, A. Feeding ecology and behavioral adjustments: flexibility of a small neotropical primate (Callithrix jacchus) to survive in a semiarid environment. Mammal Res. 61, 221–229 (2016).
Cunha, A. A., Vieira, M. V. & Grelle, C. E. V. Preliminary observations on habitat, support use and diet in two non-native primates in an urban Atlantic forest fragment: the capuchin monkey (Cebus sp.) and the common marmoset (Callithrix jacchus) in the Tijuca forest Rio de Janeiro. Urban Ecosyst. 9, 351–359 (2006).
Passamani, M. & Rylands, A. B. Feeding behavior of Geoffroy’s marmoset (Callithrix geoffroyi) in an Atlantic forest fragment of south-eastern Brazil. Primates 41, 27–38 (2000).
Veracini, C. Habitat use and ranging behavior of the silvery marmoset (Mico argentatus) at Caxiuanã National Forest (eastern Brazilian Amazonia) in The smallest anthropoids 221–240 (Springer, 2009).
Yépez, P., De La Torre, S. & Snowdon, C. T. Interpopulation differences in exudate feeding of pygmy marmosets in Ecuadorian Amazonia. Am. J. Primatol. 66, 145–158 (2005).
Hale, V. L. et al. Diet versus phylogeny: a comparison of gut microbiota in captive colobine monkey species. Microb. Ecol. 75, 515–527 (2018).
Amato, K. R. et al. The gut microbiota appears to compensate for seasonal diet variation in the wild black howler monkey (Alouatta pigra). Microb. Ecol. 69, 434–443 (2015).
Frankel, J. S., Mallott, E. K., Hopper, L. M., Ross, S. R. & Amato, K. R. The effect of captivity on the primate gut microbiome varies with host dietary niche. Am. J. Primatol. 81, e23061 (2019).
McKenzie, V. J. et al. The effects of captivity on the mammalian gut microbiome. Integr. Comp. Biol. 57, 690–704 (2017).
Lugli, G. A. et al. Evolutionary development and co‐phylogeny of primate‐associated bifidobacteria. Environ. Microbiol. (2020).
Milani, C. et al. Unveiling bifidobacterial biogeography across the mammalian branch of the tree of life. ISME J. 11, 2834–2847 (2017).
Lugli, G. A. et al. Comparative genomic and phylogenomic analyses of the Bifidobacteriaceae family. BMC Genom. 18, 568 (2017).
Pokusaeva, K., Fitzgerald, G. F. & van Sinderen, D. Carbohydrate metabolism in Bifidobacteria. Genes Nutr. 6, 285–306 (2011).
Stewart, C. J. et al. Temporal development of the gut microbiome in early childhood from the TEDDY study. Nature 562, 583–588 (2018).
Orkin, J. D. et al. Seasonality of the gut microbiota of free-ranging white-faced capuchins in a tropical dry forest. ISME J. 13, 183–196 (2019).
Neuzil-Bunesova, V. et al. Five novel bifidobacterial species isolated from faeces of primates in two Czech zoos: Bifidobacterium erythrocebi sp. nov., Bifidobacterium moraviense sp. nov., Bifidobacterium oedipodis sp. nov., Bifidobacterium olomucense sp. nov. and Bifidobacterium panos sp. nov. Int. J. Syst. Evol. Microbiol. (2020).
Duranti, S. et al. Characterization of the phylogenetic diversity of two novel species belonging to the genus Bifidobacterium: Bifidobacterium cebidarum sp. Nov. and Bifidobacterium leontopitheci sp. nov.. Int. J. Syst. Evol. Microbiol. 70, 2288–2297 (2020).
Modesto, M. et al. Bifidobacterium primatium sp. nov., Bifidobacterium scaligerum sp. nov., Bifidobacterium felsineum sp. nov. and Bifidobacterium simiarum sp. nov.: Four novel taxa isolated from the faeces of the cotton top tamarin (Saguinus oedipus) and the emperor tamarin (Saguinus imperator). Syst. Appl. Microbiol. (2018).
Neuzil-Bunesova, V. et al. Bifidobacterium canis sp nov a novel member of the Bifidobacterium pseudolongum phylogenetic group isolated from faeces of a dog (Canis lupus f. familiaris). Int. J. Syst. Evol. Microbiol. 70, 5040–5047 (2020).
Vlková, E. et al. A new medium containing mupirocin, acetic acid, and norfloxacin for the selective cultivation of bifidobacteria. Anaerobe 34, 27–33 (2015).
Carding, S., Verbeke, K., Vipond, D. T., Corfe, B. M. & Owen, L. J. Dysbiosis of the gut microbiota in disease. Microb. Ecol. Health Dis. 26, 26191 (2015).
WagnerMackenzie, B. et al. Bacterial community collapse: a meta-analysis of the sinonasal microbiota in chronic rhinosinusitis. Environ. Microbiol. 19, 381–392 (2017).
Arboleya, S., Watkins, C., Stanton, C. & Ross, R. P. Gut bifidobacteria populations in human health and aging. Front. Microbiol. 7 (2016).
Binda, C. et al. Actinobacteria: a relevant minority for the maintenance of gut homeostasis. Dig. Liver Dis. 50, 421–428 (2018).
Tojo, R. et al. Intestinal microbiota in health and disease: role of bifidobacteria in gut homeostasis. World J. Gastroenterol. 20, 15163 (2014).
Rodriguez, C. I. & Martiny, J. B. H. Evolutionary relationships among bifidobacteria and their hosts and environments. BMC Genom. 21, 1–12 (2020).
Sharma, V., Mobeen, F. & Prakash, T. Exploration of survival traits, probiotic determinants, host interactions, and functional evolution of bifidobacterial genomes using comparative genomics. Genes 9, 477 (2018).
Sun, Z. et al. Comparative genomic analysis of 45 type strains of the genus Bifidobacterium. a snapshot of its genetic diversity and evolution. PLoS One 10, 0117912 (2015).
Frey, J. C. et al. Fecal bacterial diversity in a wild gorilla. Appl. Environ. Microbiol. 72, 3788–3792 (2006).
Makovska, M., Modrackova, N., Bolechova, P., Drnkova, B. & Neuzil-Bunesova, V. Antibiotic susceptibility screening of primate-associated Clostridium ventriculi. Anaerobe, 102347 (2021).
Ushida, K. et al. Draft genome sequences of Sarcina ventriculi strains isolated from wild Japanese macaques in Yakushima Island. Genome announcements 4 (2016).
Owens, L. A. et al. A Sarcina bacterium linked to lethal disease in sanctuary chimpanzees in Sierra Leone. Nat. Commun. 12, 1–16 (2021).
Vlková, E., Rada, V., Šmehilová, M. & Killer, J. Auto-aggregation and co-aggregation ability in bifidobacteria and clostridia. Folia Microbiol. 53, 263–269 (2008).
Wang, L. et al. Adhesive Bifidobacterium induced changes in cecal microbiome alleviated constipation in mice. Front. Microbiol. 10, 1721 (2019).
Wei, Y. et al. Protective effects of bifidobacterial strains against toxigenic Clostridium difficile. Front. Microbiol. 9, 888 (2018).
Guittar, J., Shade, A. & Litchman, E. Trait-based community assembly and succession of the infant gut microbiome. Nature Commun. 10, 1–11 (2019).
Moore, R. E. & Townsend, S. D. Temporal development of the infant gut microbiome. Open Biol. 9, 190128 (2019).
Korpela, K. et al. Probiotic supplementation restores normal microbiota composition and function in antibiotic-treated and in caesarean-born infants. Microbiome 6, 1–11 (2018).
Timperio, A. M., Gorrasi, S., Zolla, L. & Fenice, M. Evaluation of MALDI-TOF mass spectrometry and MALDI BioTyper in comparison to 16S rDNA sequencing for the identification of bacteria isolated from Arctic sea water. PloS One 12, 0181860 (2017).
Bäckhed, F. et al. Dynamics and stabilization of the human gut microbiome during the first year of life. Cell Host Microbe 17, 690–703 (2015).
Brown, C. J. et al. Comparative genomics of Bifidobacterium species isolated from marmosets and humans. Am. J. Primatol. 81, e983 (2019).
Killer, J. et al. Gene encoding the CTP synthetase as an appropriate molecular tool for identification and phylogenetic study of the family Bifidobacteriaceae. MicrobiologyOpen 7, e00579 (2018).
Milani, C. et al. Evaluation of bifidobacterial community composition in the human gut by means of a targeted amplicon sequencing (ITS) protocol. FEMS Microbiol. Ecol. 90, 493–503 (2014).
Srinivasan, R. et al. Use of 16S rRNA gene for identification of a broad range of clinically relevant bacterial pathogens. PloS One 10, e0117617 (2015).
Maiden, M. C. J. et al. MLST revisited: the gene-by-gene approach to bacterial genomics. Nature Rev. Microbiol. 11, 728–736 (2013).
Lugli, G. A. et al. Phylogenetic classification of six novel species belonging to the genus Bifidobacterium comprising Bifidobacterium anseris sp. nov., Bifidobacterium criceti sp. nov., Bifidobacterium imperatoris sp. nov., Bifidobacterium italicum sp. nov., Bifidobacterium margollesii sp. nov. and Bifidobacterium parmae sp. nov. Syst. Appl. Microbiol. 41, 173–183 (2018).
Malukiewicz, J. et al. The effects of host taxon, hybridization, and environment on the gut microbiome of Callithrix marmosets. BioRxiv, 708255 (2019).
Amato, K. R. et al. Phylogenetic and ecological factors impact the gut microbiota of two Neotropical primate species. Oecologia 180, 717–733 (2016).
Hernández‐Rodríguez, D., Vásquez‐Aguilar, A. A., Serio‐Silva, J. C., Rebollar, E. A. & Azaola‐Espinosa, A. Molecular detection of Bifidobacterium spp. in faeces of black howler monkeys (Alouatta pigra). J. Med. Primatol. 48, 99–105 (2019).
Zhu, L. et al. Sex bias in gut microbiome transmission in newly paired marmosets (Callithrix jacchus). Msystems 5, e00910-00919 (2020).
Kap, Y. S. et al. Targeted diet modification reduces multiple sclerosis–like disease in adult marmoset monkeys from an outbred colony. J. Immunol. 201, 3229–3243 (2018).
Ren, T., Grieneisen, L. E., Alberts, S. C., Archie, E. A. & Wu, M. Development, diet and dynamism: longitudinal and cross-sectional predictors of gut microbial communities in wild baboons. Environ. Microbiol. 18, 1312–1325 (2016).
Xu, B. et al. Metagenomic analysis of the Rhinopithecus bieti fecal microbiome reveals a broad diversity of bacterial and glycoside hydrolase profiles related to lignocellulose degradation. BMC Genom. 16, 1–11 (2015).
Baumann, P. Biology of bacteriocyte-associated endosymbionts of plant sap-sucking insects. Annu. Rev. Microbiol. 59, 155–189 (2005).
Killer, J. et al. Bifidobacterium actinocoloniiforme sp. nov. and Bifidobacterium bohemicum sp. nov., from the bumblebee digestive tract. Int. J. Syst. Evol. Microbiol. 61, 1315–1321 (2011).
Amato, K. R. et al. Evolutionary trends in host physiology outweigh dietary niche in structuring primate gut microbiomes. ISME J. 13, 576–587 (2019).
Garber, P. A., Mallott, E. K., Porter, L. M. & Gomez, A. The gut microbiome and metabolome of saddleback tamarins (Leontocebus weddelli): Insights into the foraging ecology of a small‐bodied primate. Am. J. Primatol. 81, e23003 (2019).
Gralka, M., Szabo, R., Stocker, R. & Cordero, O. X. Trophic interactions and the drivers of microbial community assembly. Curr. Biol. 30, R1176–R1188 (2020).
Clayton, J. B. et al. Associations between nutrition, gut microbiome, and health in a novel nonhuman primate model. Sci. Rep. 8, 1–16 (2018).
Koo, B. S. et al. Idiopathic chronic diarrhea associated with dysbiosis in a captive cynomolgus macaque (Macaca fascicularis). J. Med. Primatol. 49, 56–59 (2020).
Krynak, K. L., Burke, D. J., Martin, R. A. & Dennis, P. M. Gut microbiome composition is associated with cardiac disease in zoo-housed western lowland gorillas (Gorilla gorilla gorilla). FEMS Microbiol. Lett. 364 (2017).
Modrackova, N. et al. Prebiotic potential of natural gums and starch for bifidobacteria of variable origins. Bioact. Carbohydr. Diet. Fibre 20, 100199 (2019).
McKenzie, V. J., Kueneman, J. G. & Harris, R. N. Probiotics as a tool for disease mitigation in wildlife: insights from food production and medicine. Ann. N. Y. Acad. Sci. 1429, 18–30 (2018).
Hicks, A. L. et al. Gut microbiomes of wild great apes fluctuate seasonally in response to diet. Nat. Commun. 9, 1–18 (2018).
Hungate, R. E. & Macy, J. The roll-tube method for cultivation of strict anaerobes. Bulletins from the ecological research committee, 123–126 (1973).
Rada, V. & Petr, J. A new selective medium for the isolation of glucose non-fermenting bifidobacteria from hen caeca. J. Microbiol. Methods 43, 127–132 (2000).
Orban, J. I. & Patterson, J. A. Modification of the phosphoketolase assay for rapid identification of bifidobacteria. J. Microbiol. Methods 40, 221–224 (2000).
Kim, B. J., Kim, H.-Y., Yun, Y.-J., Kim, B.-J. & Kook, Y.-H. Differentiation of Bifidobacterium species using partial RNA polymerase β-subunit (rpoB) gene sequences. Int. J. Syst. Evol. Microbiol. 60, 2697–2704 (2010).
Hall, T. A. BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. 41 edn 95–98 ([London]: Information Retrieval Ltd., c1979-c2000.).
Thompson, J. D., Higgins, D. G. & Gibson, T. J. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 22, 4673–4680 (1994).
Callahan, B. J. et al. DADA2: high-resolution sample inference from Illumina amplicon data. Nat. Methods 13, 581–583 (2016).
Quast, C. et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Ress 41, D590–D596 (2012).
Shannon, C. E. & Weaver, W. The mathematical theory of information. Urbana: University of Illinois Press 97 (1949).
Pielou, E. C. The measurement of diversity in different types of biological collections. J. Theor. Biol. 13, 131–144 (1966).
Mandal, S. et al. Analysis of composition of microbiomes: a novel method for studying microbial composition. Microb. Ecol. Health Dis. 26, 27663 (2015).
fundamental algorithms for scientific computing in Python. Virtanen, P. et al. SciPy 1.0. Nat. Methods 17, 261–272 (2020).
Seabold, S. & Perktold, J. Statsmodels: Econometric and statistical modeling with python in Proceedings of the 9th Python in Science Conference 57 (Austin, TX, 2010).
MacKinnon, J. G. & White, H. Some heteroskedasticity-consistent covariance matrix estimators with improved finite sample properties. J. Econom. 29, 305–325 (1985).
Acknowledgements
Research was funded by the European Regional Development Fund‐Project, “Centre for the investigation of synthesis and transformation of nutritional substances in the food chain in interaction with potentially harmful substances of anthropogenic origin: comprehensive assessment of soil contamination risks for the quality of agricultural products” [No: CZ.02.1.01/0.0/0.0/16_019/0000845], by the cooperation project (8J19AT028) between Austria and the Czech Republic – BMBWF and Ministry of Education, Youth and Sports of the Czech Republic, and by METROFOOD‐CZ research infrastructure project [MEYS Grant No: LM2018100] including access to its facilities. The authors thank Tamara Rudavsky and Cristina Ukowitz for their valuable assistance in laboratory experiments. We also thank the zoos in Dvur Kralove, Hodonin, Liberec, Olomouc, Pilsen, Bojnice, and Bratislava for their kind cooperation. The MALDI-TOF-MS (Bruker Biotyper) was kindly provided by the EQ-BOKU VIBT GmbH and the BOKU Core Facility Food & Bio Processing.
Author information
Authors and Affiliations
Contributions
N.-B.V. designed experiments. M.N. and N.-B.V. carried out experiments. M.N. and S.A. participated at data analysis, interpretation, and visualization of results. M.N., N.-B.V., and S.A. drafted the manuscript. B.P. and K.J. were involved in the writing up of the manuscript. B.J. and D.K.J. participated at data evaluation and revised the manuscript critically. The study was supervised by N.-B.V. and D.K.J. All authors reviewed the manuscript and approved the version to be published.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Modrackova, N., Stovicek, A., Burtscher, J. et al. The bifidobacterial distribution in the microbiome of captive primates reflects parvorder and feed specialization of the host. Sci Rep 11, 15273 (2021). https://doi.org/10.1038/s41598-021-94824-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-94824-y
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.