Abstract
DNA-binding with one finger (Dof) are plant-specific transcription factors involved in numerous pathways of plant development, such as abiotic stresses responses. Although genome-wide analysis of Dof genes has been performed in many species, but these genes in spinach have not been analyzed yet. We performed a genome-wide analysis and characterization of Dof gene family in spinach (Spinacia oleracea L.). Twenty-two Dof genes were identified and classified into four groups with nine subgroups, which was further corroborated by gene structure and motif analyses. Ka/Ks analysis revealed that SoDofs were subjected to purifying selection. Using cis-acting elements analysis, SoDofs were involved in plant growth and development, plant hormones, and stress responses. Expression profiling demonstrated that SoDofs expressed in leaf and inflorescence, and responded to cold, heat, and drought stresses. SoDof22 expressed the highest level in male flowers and under cold stress. These results provided a genome-wide analysis of SoDof genes, their gender- and tissue-specific expression, and response to abiotic stresses. The knowledge and resources gained from these analyses will benefit spinach improvement.
Similar content being viewed by others
Introduction
Spinach (Spinacia oleracea L.) is an annual or biennial diploid species, belong to the Amaranthaceae family in the order Caryophyllales1 Its annual worldwide gross production in 2016 was about 26 million tonnes (FAOSTAT; http://faostat3.fao.org). Spinach is a dietary source of Ca, Cu, Fe, K, Mg, Mn, P, Zn, folate, vitamins, and dietary fiber2, providing its great potential for medical economy3,4. However, like many other crops, its development and production is hampered by biotic stresses(diseases, pests and weed infestations,) and abiotic stresses (salinity, drought, and heat)5. Climate change causes elevated temperature and a network of events triggering the response of plants and animals6,7. Although it seems that organisms on earth gradually developed local thermal adaptation to impact their healthy condition8. Spinach is cold tolerant but having heat-sensitive characteristics that influencing its growth and significantly decrease yield and quality under hight temperature9. Winter sweet treatment (WST), termed the cold enrichment technique, has been established for cultivating high-quality leafy spinach during winter10. At that time (early December), the average daily temperature is generally below 5 °C. But staying at a low temperature for a long time would also damage spinach by reactive oxygen species (ROS)11. Although drought stress has no direct effects on the leaf nutrition quality, some physiological indicators could be decreased, such as leaf area, fresh and dry weight, leaf relative water content, and specific leaf area, which might change the shape of plant12.
Dof domain proteins are plant-specific transcription factors that contain a highly conserved 52 amino acid DNA-binding domain at the N-terminalincluding a single Cys2/Cys2 zinc finger structure13. It was projected that Cys2/Cys2 zinc finger specifically binds to a conserved sequence with 5′-(T/A)AAAG-3′ in gene promoters14. At the C-terminal of the Dof proteins, there is a transcription regulation domain with diverse functions involving interaction with a variety of regulatory proteins and activating the gene expression15. Indeed, previous studies corroborated its functional role in plant growth and development, such as in flowering control16,17, maturation18, seed development19, and germination20,21. Specifically, mutant dag1 (encoding a Dof transcription factor in Arabidopsis) seeds are induced to germinate by much lower red light fluence rates22; the COG1 gene (encoding a Dof protein in Arabidopsis) functions as a negative regulator in phytochrome signaling pathways23; CDFs (CYCLING DOF FACTORS, Dof-type transcriptional repressors) that directly suppresses the expression of CONSTANS (CO), which could prevent the expression of photoperiodic gene, the perception of day-length and the floral transition in Arabidopsis24. Moreover, Dof transcription factors also participated in phytohormone and stress responses, such as the TDDF1 (encoding a Dof protein in tomato) which could improve drought, salt, various hormones stress as well as resistance to late blight25; ThZFP1 and ThDof1.4 improve salt and osmotic stress tolerance by increase the proline level and ROS scavenging capability26. Therefore, Dof gene family plays an essential role in the life cycle of plants.
In recent years, with the sequencing of genome, the identification of Dof genes was widely researched in various plant species, such as Arabidopsis, rice27, soybean28, maize29, sorghum30, sugarcane31, and so on. The spinach draft genome was reported in 20171, however, few gene families were analyzed for the genome. The functions of members of Dof genes remain unknown in spinach. As previously reported, plants different sex types show different responses to abiotic stress32. The reproductive potential of male, female, and monoecious spinach differe under water-limited condition33. But the expression of Dof genes in different sex types of spinach under abiotic stresses is still unknown. In this study, we identified 22 Dof genes, showed the structure and motifs, and classified the group of Dof genes in spinach. In addition, duplication events and cis-element on their promoters were predicted. Functional prediction was performed based on gene expression analysis in different tissues and in responses to different abiotic stresses. The results will provide a foundation for gene cloning and functional characterization of Dofs in spinach.
Materials and methods
Identification of SoDof gene family members in the spinach genome
To identify the Dof gene family members in Spinacia oleracea L., all proteins from the spinach genome were scanned by HMMER-3.234 using the Hidden Markov Model (HMM) corresponding to the HMM profile of the Dof domain (PF02701). The spinach genome data was downloaded from SpinachBase (http://www.spinachbase.org/?q=download). The predicted proteins were confirmed for the presence of the conserved Dof domain by NCBI Conserved Domain Database (CDD)35, Pfam36 and SMART37 tools. Similarly, Arabidopsis and sugarbeet (Beta vulgaris L.) Dof genes were identified by scanning Arabidopsis database (ftp://ftp.ensemblgenomes.org/pub/plants/release-42/fasta/arabidopsis_thaliana/) and sugarbeet database (ftp://ftp.ensemblgenomes.org/pub/plants/release-42/fasta/beta_vulgaris/) using HMM and CDD. We performed the ExPASy server38 to detect the theoretical pI and molecular weight of candidate SoDof genes.
Multiple sequences alignment and phylogenetic characterization
For phylogenetic analysis of the Dof gene family, multiple sequence alignments were conducted on the amino acid sequences of Dof protein from spinach, Arabidopsis, and sugarbeet by MUSCLE with default settings. After that, MEGA-X-10.0.4 software was used to construct phylogenetic tree among these three species with the Neighbour-Joining (NJ) method and 1000 bootstraps. Alignment of multiple SoDofs was performed by DNAMAN-6.0.
Chromosomal locations and duplication time
The distribution information for each SoDof gene on chromosome was obtained from their annotation file. MG2C (http://mg2c.iask.in/mg2c_v2.1/) was used to map the chromosomal locations for each SoDof gene with default settings. To estimate the synonymous and non-synonymous substitution, Ka and Ks values were calculated. ClustalW was used to align the nucleotide sequence of SoDof genes. Ka and Ks values were used to estimate by DnaSp-5.10. The time (million years ago, Mya) of segmental duplication events for each SoDof gene was estimated using a formula, T = Ks/2λ which assumed λ of 7.0e−9 synonymous/substitution site/year for spinach1.
Gene structure analysis and conserved motif identification
The exon–intron organizations of the genes with phylogenetic tree and Dof motifs were determined using the Gene Structure Display Server (http://gsds.cbi.pku.edu.cn/). The motifs distribution of the Dof protein in spinach, Arabidopsis, and sugarbeet were statistically identified by the MEME program (http://meme-suite.org/) with the motif length set to 6–100 and the maximum number of motifs was set to 15. Then TBtools-1.08239 was employed to create the motif structure with phylogenetic tree.
Cis-elements identification in promoter regions of SoDofs
To investigate cis-elements in promoter sequences of Dof coding genes in spinach, the upstream sequences (2000 bp) of each SoDof gene were extracted from spinach genome according to the GFF3 (general feature format) file. Then the retrieved sequences were submitted to a search by the PlantCARE (http://bioinformatics.psb.ugent.be/webtools/plantcare/html/)40 for predicting the cis-elements which might be involved in regulation of SoDof genes expression.
Sample collection and preparation
Spinach II9A0073 seeds were obtained from CAAS (China Academy of Agricultural Sciences). Seeds were sown in plots, and seedlings grew in an artificial climate chamber with a photoperiod of 16 h light/8 h dark, temperature at 24 °C and humidity at about 60%. After three weeks, spinach seedlings with consistent growth were selected and prepared for environmental stress treatment. Abiotic stresses were performed by adding 20% (mass fraction) PEG 4000 to simulate the drought condition and adjusting the temperature of the artificial climate box to simulate high-temperature stress (40 °C) and low-temperature stress (4 °C). Under stress conditions, the spinach leaves were sampled at 0, 2, 4, 7, 12, 24 h after treatment. The plants with non-treatment were collected for their roots, leaves, and stems in vegetative growth stage, as well as their male flowers and female flowers. All samples were immediately frozen in liquid nitrogen and stored at − 80 °C.
RNA extraction and quantitative real-time PCR analysis
Total RNA from different samples was extracted using the Trizol reagent. The quality and concentration of RNA were tested on 1.0% agar gel electrophoresis and the NanoDrop 2000 (Thermo Fisher Scientific, USA). The total RNA was reverse transcribed into cDNA with its 200 ng per microliter final work concentration using Evo M-MLV RT Kit with gDNA Clean for qPCR (Accurate Biotechnology, China) according to the manufacturer’s instruction. For qRT-PCR, Actin11 gene was used as a reference gene. The specific primers were designed by IDT (https://sg.idtdna.com/pages) and the sequences of all primers are listed in Supplementary Table S3. The qRT-PCR was conducted with SYBR Green Premix Pro Taq HS qPCR Kit (Accurate Biotechnology, China) following the manufacturer’s protocol. Experiments were repeated three times with technical and biological replications for each sample. The relative gene expression level was calculated by the 2 − ∆∆CT method. Graphpad Prism8 (Graphpad Software Inc., La Jolla, CA) was performed to calculate the p-value. Expression values were calculated as the arithmetic mean and then presented as the heatmap by R package.
Result
Identification and classification of SoDofs genes
To identify the Dof gene family members in spinach, all proteins from the spinach genome were scanned by using HMMER-3.2 and 22 genes were predicted as Dof gene family members in spinach. These Dof candidate genes in spinach were named as SoDof1–SoDof22 (Table 1). The predicted proteins were further confirmed to contain the conserved Dof domain. Similarly, 36 Dof genes had been identified in Arabidopsis and 22 Dof genes were identified in sugarbeet named as BvDof1–BvDof22 (Supplementary Table S1). The full length of the coding sequence (CDS) ranged from 492 (SoDof12) bp to 1485 (SoDof13) bp with an average length of 1060 bp. The quantity of aa (amino acids) for SoDof varied from 163 (SoDof12) to 494 (SoDof13) aa, with an average protein length of ~ 352 aa. The molecular weight (MW) fluctuated between 18.5 kDa (SoDof12) and 54.5 kDa (SoDof13), and the theoretical isoelectric points (pI) ranged from 4.6 (SoDof20) to 8.92 (SoDof9) (Table 1).
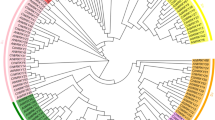
Multiple sequence alignment showed a Dof conserved motif of 52 amino acids located in 22 SoDof genes, with a single Cys2/Cys2 zinc-finger structure at the N-terminal (Fig. 1A). Phylogenetic tree was constructed between 22 SoDof genes, 22 BvDof genes, and 36 Dofs in Arabidopsis (Fig. 2). A total of 22 SoDof TFs from spinach were classified into four main groups (Groups A–D), which could be divided into multiple subgroups, A, B1, B2, C1, C2.1, C2.2, C3, D1, and D2. The number of SoDofs in Group B, C, and D was similar with a total number of 20. Specifically, Group B (contained the most number among all groups) could be divided into subgroup B1 and subgroup B2 with SoDof10, SoDof16, SoDof17, SoDof18 in subgroup B1 and SoDof3, SoDof6, SoDof9, SoDof19 in subgroup B2 (Fig. 2). Subgroup D1 had the largest number of SoDofs (SoDof11, SoDof12, SoDof13, SoDof14, SoDof21) in subgroups. SoDof2 and SoDof4 belonged to Group A (Fig. 2). Over half SoDofs were alkaline which contained all members in Group B, and subgroup D1 (Table 1).
Mapping SoDof genes in spinach chromosomes and Ka/Ks analysis
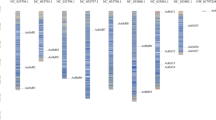
The spinach genome consists of only 6 chromosomes. The 22 putative SoDof genes were found to be distributed in 6 chromosomes, and unplaced contigs (Fig. 3). Only 50% SoDofs genes were anchored in chromosomes. The largest number of SoDof members was located in chromosome 5, which contains SoDof 7, 11, and 4. Compared with the gap of SoDof in other chromosomes, these three genes were closer to each other, especially SoDof11 and SoDof4. There were 2 SoDof genes in chromosomes 1, 4, and 6, respectively. SoDof1 and SoDof3 were located in chromosomes 2 and 3, respectively. Ka and Ks value calculation aims to identify duplication events for each SoDof gene. The duplication of SoDof genes originated from about 5.66 Mya (Ks = 0.793) to 41.27 Mya (Ks = 5.778) with an average of 16.12 Mya (Supplementary Table S2). All values of Ka/Ks were lower than 1 and some SoDof were even lower than 0.1 (Table 2).
Chromsomal location of SoDof genes. The size of a chromosome is indicated by its relative length. Figure was made by MG2C (http://mg2c.iask.in/mg2c_v2.1/).
Gene structure and motif analysis of SoDof genes
Candidate SoDof genes were analyzed using Gene Structure Display Server to investigate the characterization of exon–intron structure. There was no more than two introns in each SoDof (Fig. 4). To further reveal the diversification of SoDof genes, we performed the MEME program to detect motif patterns, and 15 distinct motifs were identified (Fig. 5). It was predicted that motif1 could be considered as the Dof region (Fig. 1B). The schematic distribution of the 15 motifs showed that motif1 (Fig. 1B) and motif2 (Fig. 1C) were highly conserved in all SoDof proteins. Notably, SoDofs shared similar conserved motif compositions in some subgroups. Motif 7 in front of the Dof region were highly conserved in subgroup B1. And members of subgroup C2.2 contained motif13. Interestingly, motif5 was prominently conserved in subgroup D1 (contained the most SoDof members among all subgroups). Specifically, motif5 presented at the N-terminal in all subgroup D1 members, and motif4 appeared at the C-terminal in majority of subgroup D1 members.
The exon–intron structure of Dof genes in Spinach: CDS (yellow), UTR (blue), Intron (black line) and zf-Dof region (pink). SoDof6 contains one intron which is too short to recognize in this figure resolution. Figure was made by the Gene Structure Display Server (http://gsds.cbi.pku.edu.cn/).
The schematic distribution of motifs for Dof genes among spinach, Arabidopsis and sugarbeet. Figure was made by the MEME program (http://meme-suite.org/) and TBtools-1.082.
Cis-regulatory element analysis
PlantCARE was used to analyze the cis-regulatory element for each SoDof gene by retrieving the 2 kb upstream sequence of each candidate, except for SoDof18 because of lack of 2 kb upstream sequence on its scaffold location (Supplementary Data). Dof gene family in spinach had TATA-box and CAAT-box. SoDof genes may also be controlled by many phytohormones, such as methyl jasmonate (MeJA), gibberellins (GA), ethylene, auxin, and salicylic acid (SA). We also detected many other important cis-elements on Dof gene family that involve in plant growth and development. For example, there were a large number of elements associated with physiological processes, such as light responsiveness, circadian control, endosperm expression, meristem and flower meristem expression, root-specific and seed-specific regulation (Supplementary Data). The sum of cis-elements of subgroup D1 was greatest in plant growth and development. The sum of cis-elements of subgroup D1 was also greatest in phytohormones class. The greatest mean of cis-elements in phytohormones class was subgroup C3. The greatest mean of cis-elements in light responsiveness and physiological process were in subgroup C2.2 and C1 respectively (Table 3). In physiological process, some elements, participated in some small molecule pathway, were also found, such as zein metabolism regulation and flavonoid biosynthetic genes regulation (Supplementary Data). Moreover, nine cis-elements (WUN-motif, STRE, TC-rich repeats e.g.) were also predicted, which were related to defense and stress responsiveness. The sum and mean of cis-elements of subgroup A were greatest in stress response.
Tissue-specific expression analysis of SoDof genes
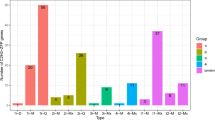
We isolated RNA samples from roots, stems, leaves, male flowers, and female flowers, and detected expression of all SoDof genes in spinach using qRT-PCR. Expression profile of the SoDof genes revealed that nine SoDofsexhibited their highest transcript level in reproductive organs and eight SoDofs in leaves (Fig. 6A). Only two SoDofs (SoDof1 and SoDof5) were expressed in roots and stems, respectively. Notably, SoDof10 and SoDof15 had extremely high expression in leaves; SoDof22 showed high expression in male flowers (Fig. 6B). Comparing with leaves or inflorescences, the transcript level of these three genes in other tissues was neglectable, indicating that their expression was tissue-specific. There were three homologous genes (SoDof16, SoDof17, and SoDof18) with same mRNA sequence, and their expression pattern was not analyzed.
The tissue-specific expression of Dof genes in Spinach by qRT-PCR. (A) Expression level of SoDofs. The color scheme used to present expression level is sky-blue/red: light-yellow boxes indicate low variation in gene expression, sky-blue indicate a fold decrease, and red boxes indicate a fold increase in relation to mean value. The expression value were calculated as the arithmetic mean. (B) The expression level of SoDof10, SoDof15 and SoDof22 in different tissues. The Y-axis indicates relative expression level and the X-axis indicated different tissues: root (gray); stem (light brown); leaf (green); female flower (red); male flower (pink). The error bars were caculated based on three biological repiticates using standard deviation. Figure (A) was made by R package; (B) was made by Graphpad Prism8.
Expression patterns of SoDof genes under abiotic stresses
To investigate the stress responsiveness and expression pattern of SoDof gene between different sex-types, we treated female male plants, and plants at vegetative stage under three types of abiotic stress (low-temperature 4 °C, high-temperature 40 °C, and drought 20%PEG4000). Spinach leaves were collected at 0 h, 2 h, 4 h, 7 h, 12 h, and 24 h after treatment and detected by qRT-PCR.
The majority of SoDof genes in female plants were up regulated under low temperature (Fig. 7A). The greatest increase in expression occurred in SoDof22 (up to the top at 24 h after treatment) in female plants (Supplementary Fig. S2A). SoDof14 experienced the same trend, but the expression level was much lower than that in SoDof22. Compared with other SoDofs, the SoDof22 expressed the most in plants at vegetative stage, and its extreme expression reached the top at 7 h and then went down (Supplementary Fig. S2B). However, in male plants, the expression pattern of SoDof3 and SoDof5 was similar. The expression of SoDof3 reached the highest level at 4 h and the expression of SoDof5 reached the highest level at 7 h (Supplementary Fig. S2C). In vegetative plants, 95% SoDof genes (more than those in male or female plants) were up-regulated and almost all of their highest expression appeared at 7 h (Fig. 7A). Among them, SoDof3, SoDof4. SoDof8 and SoDof9 were down-regulated at 2 h and 4 h. After that, they expressed the highest level at 7 h and then went down. The trends of six SoDofs (SoDof11, SoDof12, SoDof13, SoDof19, SoDof20, and SoDof21) were similar. Their expression went up slightly at 2 h and 4 h and reached the highest at 7 h, and then went down (Supplementary Fig. S2B). But there were difference between female and male plants. In male plants, there were the most number of SoDofs (SoDof6, SoDof8, and SoDof9) down-regulated, indicating that SoDof genes in males showed more negative response under 4 °C (Fig. 7B).
The expression pattern of SoDof genes under stresses. (A) The expression pattern of all SoDof genes under cold stress, heat stress and drought stress. The color scheme used to present expression level is sky-blue/red: light-yellow boxes indicate low variation in gene expression, sky-blue indicate a fold decrease, and red boxes indicate a fold increase in relation to mean value. The Y-axis indicates each SoDof gene and the X-axis indicated the time after treatment. The expression value were calculated as the arithmetic mean. (B) The expression level of down-regulated SoDofs. F-SoDof means the SoDof gene in female plants; V-SoDof means the SoDof gene in vegetative plants; M-SoDof means the SoDof gene in male plants. The Y-axis indicates relative expression level and the X-axis indicated the time after treatment:0 h (gray); 2 h (light brown); 4 h (orange); 7 h (green); 12 h (purple);24 h (pink). Asterisk indicates a significant difference from 0 h (p < 0.05). Error bars indicate standard error of independent technological replicates. Figure (A) and (B) were made by Graphpad Prism8.
Under high temperature, most SoDofs were up-regulated and all SoDof genes were up-regulated in female plants. Compared with other SoDof genes, the expression of SoDof3 (up to the top at 24 h) was the highest in females, males, and vegetative plants (Supplementary Fig. S3). SoDof12, SoDof13, SoDof14, SoDof15, and SoDof22 also exhibited the highest expression at 24 h in female plants. The expression of some genes (SoDof1, SoDof2, SoDof5, SoDof6, SoDof11, SoDof19, and SoDof20) went up to the highest at 4 h which means they responded earlier than others did. In plants at vegetative stage, there was only one down-regulated SoDof gene (SoDof1) (Fig. 7B). Additionally, the expression of SoDof6, SoDof8, and SoDof9 were suppressed in male plants (Fig. 7B). 68% SoDofs showed the highest transcript level at 24 h in plant at vegetative stage, and 84% SoDofs showed the highest transcript level at 7 h or before 7 h in male plants (Supplementary Fig. S3).
To investigate the expression profile for each SoDofs under drought condition.All SoDof genes were up-regulated in female plants. Compared to other SoDof genes, the expression of SoDof15 was highest in females, males, and vegetative plants (Supplementary Fig. S4). But it was up to the top at 24 h in females, at 12 h in vegetative plants, and at 2 h in males. SoDof3 and SoDof7 were down-regulated in plants at vegetative stage (Fig. 7B). In male plants, six SoDof genes (SoDof1, SoDof3, SoDof5, SoDof9 SoDof14, and SoDof20) exhibited suppressed expression, and the expression of all SoDofs was lower than in female and vegetative plants (Supplementary Fig. S4).
Discussion
Identification and characteristics of SoDof genes
The Dof gene family is a plant-specific family of transcription factors. Since the discovery of the first Dof gene in maize41, its members in other species have been uncovered and its function in the growth and development has been characterized. We identified 22 SoDof genes in spinach genome and constructed a phylogenetic tree to divide them into four categories (A, B, C, and D) (Fig. 2). The quantity of SoDofs is lower than that of Arabidopsis (36)27, tomato (34)42, wheat (96)43, rice (30)27, potato (35)44, soybean (78)28, and sugarcane (29)31, but it is same to that of sugarbeet. This is because spinach separated with Arabidopsis just after the ancient whole-genome triplication and there was no whole-genome duplication in spinach genome1. The theoretical isoelectric points (pI) of Dof proteins ranged from 4.6 to 8.92. Only two Dof proteins have an isoelectric point between 6.5 and 7.5, and over half Dof proteins were alkaline. All values of Ka/Ks were lower than 1 (Supplementary Table S2), indicating that SoDof genes were subjected to purifying selection45.
Structural conservation and chromosome location of SoDof genes
From our analysis of the spinach genome1, only half of the Dof genes were assembled in chromosomes. Their distribution was relatively even, but three Dof genes clustered on one end of the chromosome 5 (Fig. 3). Although the spinach genome has no recent whole-genome duplication, partial gene duplications may lead to the formation of specific Dof genes clustered in specific parts of chromosomes. It is the main effect on gene family expansion46. The exon–intron divergence is supporting evidence to determine the evolutionary relationship of plants47. The intron–exon analysis showed that there were no more than two introns in each Dof gene (Fig. 4). The distribution of motifs is indicative of an evolutionary relationship43. The protein sequence analysis of the 80 Dof genes (22 SoDof, 22 BvDof, and 36 Dof in Arabidopsis) revealed that only Dof motifs of these 80 protein sequences are conserved (Fig. 5). The Dof proteins in the same subgroup contain relatively conserved motif structures. Motif 7 is in subgroup B1 and motif13 is in subgroup C2.2. Motif5 were prominently conserved in the subgroup D1. Specifically, motif5,, motif3, and motif14 are only conserved in subgroup D1.
Cis-elements of SoDof genes
Cis-elements play significant roles during the life cycle of plants, such as phytohormone and stress response. In SoDof gene family, most cis-elements we identified were those related to light response, revealing that light signals may influence the regulation of SoDofs expression. Moreover, we identified cis-elements associated with the development of plant tissues in the promoter region of SoDofs, such as AP-148. Cis-elements associated with hormones and stress response were also identified in the promoter region of SoDofs. These results suggested that SoDof genes may participate in plant development and response to hormone and stress.
Potential Role of SoDof genes in different tissues
To figure out the potential roles of SoDofs, we analyzed the expression profiles of 19 SoDof genes in different spinach tissues. The other three genes, SoDof16, SoDof17, and SoDof18, were excluded from the analyses because they shared the mRNA sequences that are not distinguishable from each other. Among the 19 SoDofs expressed in spinach, 42% SoDofs showed a dominant expression in leaves and 47% in reproductive organs (Fig. 6A). In grapevine, eleven of twenty-five Dof gene expressed in inflorescences49 (similar to the number of SoDofs). Over half of Dof genes were expressed in vascular system in spinach, as in Arabidopsis50. Among them, there are six SoDofs (SoDof4, SoDof11, SoDof19, SoDof20, SoDof21, and SoDof22) that expressed at a high level in flowers, indicating that they might be involved in the development of reproductive organs, especially for SoDof22 (Fig. 6B). SoDof22 is orthologous to AT4G21050, which is involved in regenerated shoot numbers51. Comparing with the number of cis-elements of SoDofs, SoDof22 contained the most cis-elements associated with plant hormone. One-third of them were ERE52 which are ethylene-responsive elements. This gene also contained the most auxin-responsive cis-elements, such as AuxRR-core53 and TGA-box54. These Dof genes might involve in the growth and development of spinach reproductive organs.
Potential role of SoDof genes in response to abiotic stress
In the expression profile for abiotic stress, the expression of SoDofs in male plants was lower than that in female plants and the plants at vegetative stage (Supplementary Figs. S2–S4). The trend of expression in each subgroup under each condition is different. SoDof22, SoDof3, and SoDof15 showed the highest level in expression after treatment under cold, heat, and drought stress, respectively (Fig. 7B). As previous studies have shown, Dof genes participate in responding to various stresses. In tomato, SlCDF1-5 genes were induced in response to osmotic, salt, heat, and low-temperature stresses. Over-expressing SlCDF1 or SlCDF3 in Arabidopsis showed an increasing drought and salt tolerance55. In brassica, the BnCDF1 gene was induced in response to low temperatures, and overexpressing BnCDF1 in Arabidopsis could increase freezing tolerance56. In watermelon, nine selected Dof genes showed differential expression under salt stress and ABA treatments57. In Chinese cabbage, most Dof genes were up-regulated quickly under salt, drought, heat and cold stresses58. Higher expression level of SoDof22, SoDof3, and SoDof15 were detected after abiotic stress treatment, indicating that these genes might have an important role in responding to heat, cold and drought stresses. Over-expressing BnCDF1 in Arabidopsis also delayed flowering time by reducing the expression of CO and FT56. SoDof22 showed high expression level both in inflorescence and under cold stress, suggesting that the role of SoDof22 might be similar to BnCDF1 within the interplay between environmental conditions and flowering time.
The promoter of SoDof22 contains an LTR cis-element responding to low-temperature and the promoter of SoDof15 contains an MBS cis-element that participated in drought inducibility59 (Supplementary Data). The response of its cis-element leads to an increased expression under low temperature or PEG4000. According to the expression profile of each stress, there was an expression difference between each sex type in spinach. Under cold stress, SoDof4 was down-regulated in female plants and SoDof7 was down-regulated in female and vegetative plants. While, in male plants, they showed expression increase at 2 h after treatment. Under heat stress, SoDof genes in female plants were all up-regulated, while, vegetative plants and male plants contained down-regulated SoDof genes. Under drought stress, the quantity of down-regulated SoDofs in male plants was much more than that in others. Female plants are more sensitive to drought than male plants, similar to the response in Populus yunnanensis60.
References
Xu, C. et al. Draft genome of spinach and transcriptome diversity of 120;accessions. Nat. Commun. 8, 1–10 (2017).
Qin, J. et al. Genetic diversity and association mapping of mineral element concentrations in spinach leaves. BMC Genom. https://doi.org/10.1186/s12864-017-4297-y (2017).
He, T., Huang, C. Y., Chen, H. & Hou, Y. H. Effects of spinach powder fat-soluble extract on proliferation of human gastric adenocarcinoma cells. Biomed. Environ. Sci. 12, 247–252 (2000).
Gorgi, H. M., Safakhah, H. A. & Haghighi, S. Anxiolytic effects of the aqueous extracts of spinach leaves in mice. Sci. J. Kurdistan Univ. Med. Sci. 15, 43–50 (2010).
Kandel, S. L., Mou, B., Shishkoff, N., Shi, A. & Subbarao, K. V. Spinach downy mildew: Advances in our understanding of the disease cycle and prospects for disease management. Plant Dis. 103, 791–803. https://doi.org/10.1094/pdis-10-18-1720-fe (2019).
Vázquez, D. P., Gianoli, E., Morris, W. F. & Bozinovic, F. Ecological and evolutionary impacts of changing climatic variability. Biol. Rev. Camb. Philos. Soc. 92, 22 (2017).
Trenberth, K. E. Changes in precipitation with climate change. Clim. Res. 47, 123–138 (2011).
Verheyen, J. & Stoks, R. Temperature variation makes an ectotherm more sensitive to global warming unless thermal evolution occurs. J. Anim. Ecol. 88, 624–636 (2019).
Yan, J. et al. De novo transcriptome sequencing and gene expression profiling of spinach (Spinacia oleracea L.) leaves under heat stress. Sci. Rep. 6, 19473 (2016).
Satoh, Y., Katoh, T. & Ozawa, K. Growers’ barriers to a new technique to improve vegetable nutrition using cold Weather. Acta Horticulturae. https://doi.org/10.17660/ActaHortic.2001.559.60 (2001).
Watanabe, M. & Ayugase, J. Effect of low temperature on flavonoids, oxygen radical absorbance capacity values and major components of winter sweet spinach (Spinacia oleracea L.): Winter sweet treatment for spinach cultivation. J. Sci. Food Agric. https://doi.org/10.1002/jsfa.6925 (2014).
Xu, C. & Leskovar, D. Effects of A. nodosum seaweed extracts on spinach growth, physiology and nutrition value under drought stress. Sci. Hortic. https://doi.org/10.1016/j.scienta.2014.12.004 (2015).
Riechmann, J. et al. Arabidopsis transcription factors: Genome-wide comparative analysis among eukaryotes. Science (New York, N.Y.) 290, 2105–2110. https://doi.org/10.1126/science.290.5499.2105 (2001).
Yanagisawa, S. & Schmidt, R. J. Diversity and similarity among recognition sequences of Dof transcription factors. Plant J. 17, 209–214. https://doi.org/10.1046/j.1365-313x.1999.00363.x (1999).
Noguero, M. et al. Role of the DNA-binding One Zinc Finger (DOF) transcription factor family in plants. Plant Sci. 209, 32–45 (2013).
Liu, J., Cheng, Z., Xie, L., Li, X. & Gao, J. Multifaceted role of PheDof12-1 in the regulation of flowering time and abiotic stress responses in Moso Bamboo (Phyllostachys edulis). Int. J. Mol. Sci. 20, 424. https://doi.org/10.3390/ijms20020424 (2019).
Liu, X. et al. Characterization of Dof family in Pyrus bretschneideri and role PbDof9.2 in flowering time regulation. Genomics https://doi.org/10.1016/j.ygeno.2019.05.005 (2019).
Salaria, N. et al. Solanum tuberosum (CYCLING DOF FACTOR) CDF1.2 allele: A candidate gene for developing earliness in potato. S. Afr. J. Bot. 132, 242–248. https://doi.org/10.1016/j.sajb.2020.05.008 (2020).
Dong, G., Ni, Z., Yao, Y., Nie, X. & Sun, Q. Wheat Dof transcription factor WPBF interacts with TaQM and activates transcription of an alpha-gliadin gene during wheat seed development. Plant Mol. Biol. 63, 73–84. https://doi.org/10.1007/s11103-006-9073-3 (2007).
Santopolo, S. et al. DOF AFFECTING GERMINATION 2 is a positive regulator of light-mediated seed germination and is repressed by DOF AFFECTING GERMINATION 1. BMC Plant Biol. 15, 453. https://doi.org/10.1186/s12870-015-0453-1 (2015).
Martinez, M. et al. The barley cystatin gene (Icy) is regulated by DOF transcription factors in aleurone cells upon germination. J. Exp. Bot. 56, 547–556. https://doi.org/10.1093/jxb/eri033 (2005).
Maura, P. et al. Inactivation of the phloem-specific Dof zinc finger gene DAG1 affects response to light and integrity of the testa of Arabidopsis seeds. Plant Physiol. 128, 411–417 (2002).
Park, D. H. et al. The Arabidopsis COG1 gene encodes a Dof domain transcription factor and negatively regulates phytochrome signaling. Plant J. 34, 161–171 (2003).
Ishida, T., Sugiyama, T., Tabei, N. & Yanagisawa, S. Diurnal expression of CONSTANS-like genes is independent of the function of cycling DOF factor (CDF)-like transcriptional repressors in Physcomitrella patens. Plant Biotechnol. 31, 293–299 (2014).
Ewas, M. et al. The Tomato DOF Daily Fluctuations 1, TDDF1 acts as flowering accelerator and protector against various stresses. Sci. Rep. https://doi.org/10.1038/s41598-017-10399-7 (2017).
Zang, D., Wang, L., Zhang, Y., Zhao, H. & Wang, Y. ThDof1.4 and ThZFP1 constitute a transcriptional regulatory cascade involved in salt or osmotic stress in Tamarix hispida. Plant Mol. Biol. https://doi.org/10.1007/s11103-017-0620-x (2017).
Diego, L., Pilar, C. & Jesús, V.-C. Genome-wide comparative phylogenetic analysis of the rice and Arabidopsis Dof gene families. BMC Evol. Biol. 3, 1–11 (2003).
Guo, Y. & Qiu, L. Genome-wide analysis of the Dof transcription factor gene family reveals soybean-specific duplicable and functional characteristics. PLoS One 8, e76809. https://doi.org/10.1371/journal.pone.0076809 (2013).
Chen, Y. & Cao, J. Comparative analysis of Dof transcription factor family in maize. Plant Mol. Biol. Report. 33, 1245–1258 (2015).
Hariom, K., Shubhra, G., Kumar, S. V., Smita, R. & Dinesh, Y. Genome wide identification of Dof transcription factor gene family in sorghum and its comparative phylogenetic analysis with rice and Arabidopsis. Mol. Biol. Rep. 38, 5037–5053 (2011).
Mingxing, C. et al. Allele specific expression of Dof genes responding to hormones and abiotic stresses in sugarcane. PLoS One 15, e0227716 (2020).
Li, C., Ren, J., Luo, J. & Lu, R. Sex-specific physiological and growth responses to water stress in Hippophae rhamnoides L. populations. Acta Physiol. Plant. 26, 123 (2004).
Freeman, D. C. & Vitale, J. J. The influence of environment on the sex ratio and fitness of spinach. Bot. Gaz. 146, 137–142 (1985).
Potter, S. C. et al. HMMER web server: 2018 update. Nucleic Acids Res. 46, W200–W204 (2018).
Shennan, L. et al. CDD/SPARCLE: The conserved domain database in 2020. Nucleic Acids Res. 48, D1 (2019).
Sara, E. G. et al. The Pfam protein families database in 2019. Nucleic Acids Res. 47, D1 (2018).
Ivica, L. & Peer, B. 20 years of the SMART protein domain annotation resource. Nuclc Acids Res. 46, D493–D496 (2018).
Walker, J. M. The Proteomics Protocols Handbook (Humana Press, 2005). https://doi.org/10.1385/1592598900.
Chen, C., Xia, R., Chen, H. & He, Y. TBtools, a Toolkit for Biologists integrating various HTS-data handling tools with a user-friendly interface. bioRxiv. https://doi.org/10.1101/289660 (2018).
Lescot, M., Déhais, P., Thijs, G., Marchal, K. & Rombauts, S. PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res. 30, 325–327 (2002).
Yanagisawa, S. & Izui, K. Molecular cloning of two DNA-binding proteins of maize that are structurally different but interact with the same sequence motif. J. Biol. Chem. 268, 16028–16036 (1993).
Cai, X. et al. Genome-wide analysis of plant-specific Dof transcription factor family in tomato. J. Integr. Plant Biol. https://doi.org/10.1111/jipb.12043 (2013).
Liu, Y.et al. Genome-wide analysis of wheat DNA-binding with one finger (Dof) transcription factor genes: evolutionary characteristics and diverse abiotic stress responses. BMC Genomics. https://doi.org/10.1186/s12864-020-6691-0 (2020).
Venkatesh, J. & Park, S. W. Genome-wide analysis and expression profiling of DNA-binding with one zinc finger (Dof) transcription factor family in potato. Plant Physiol. Biochem. https://doi.org/10.1016/j.plaphy.2015.05.010 (2015).
Hurst, L. The Ka/Ks ratio: Diagnosing the form of sequence evolution. Trends Genet. TIG 18, 486. https://doi.org/10.1016/S0168-9525(02)02722-1 (2002).
Taylor, J. & Raes, J. Duplication and divergence: The evolution of new genes and old ideas. Annu. Rev. Genet. 38, 615–643. https://doi.org/10.1146/annurev.genet.38.072902.092831 (2004).
Koralewski, T. & Krutovsky, K. Evolution of exon–intron structure and alternative splicing. PLoS One 6, e18055. https://doi.org/10.1371/journal.pone.0018055 (2011).
Eckardt, N. Dissecting cis-regulation of FLOWERING LOCUS T. Plant Cell 22, 1422. https://doi.org/10.1105/tpc.110.220511 (2010).
Costenaro-da-Silva, D. et al. Transcriptome analyses of the Dof-like gene family in grapevine reveal its involvement in berry, flower and seed development. Hortic. Res. 3, 16042. https://doi.org/10.1038/hortres.2016.42 (2016).
Le Hir, R. & Bellini, C. The plant-specific Dof transcription factors family: New players involved in vascular system development and functioning in Arabidopsis. Front. Plant Sci. https://doi.org/10.3389/fpls.2013.00164 (2013).
Lardon, R., Wijnker, E., Keurentjes, J. & Geelen, D. The genetic framework of shoot regeneration in Arabidopsis comprises master regulators and conditional fine-tuning factors. Commun. Biol. 3, 549. https://doi.org/10.1038/s42003-020-01274-9 (2020).
Li, X., Li, M. & Bai, X. Upregulation of TLR2 expression is induced by estrogen via an estrogen-response element (ERE). Arch. Biochem. Biophys. 549, 26–31 (2014).
Ballas, N., Wong, L.-M. & Theologis, A. Identification of the Auxin-responsive Element, AuxRE, in the primary indoleacetic acid-inducible gene, PS-IAA4/5, of Pea (Pisum sativum). J. Mol. Biol. 233, 580–596. https://doi.org/10.1006/jmbi.1993.1537 (1993).
Liu, Z.-B., Ulmasov, T., Shi, X., Hagen, G. & Guilfoyle, T. Soybean GH3 promoter contains multiple auxin-inducible elements. Plant Cell 6, 645–657. https://doi.org/10.1105/tpc.6.5.645 (1994).
Corrales, A. et al. Characterization of tomato Cycling Dof Factors reveals conserved and new functions in the control of floweri. J. Exp. Bot. https://doi.org/10.1093/jxb/ert451 (2014).
Xu, J. & Dai, H. Brassica napus Cycling Dof Factor1 (BnCDF1) is involved in flowering time and freezing tolerance. Plant Growth Regul. https://doi.org/10.1007/s10725-016-0168-9 (2016).
Zhou, Y. et al. Genome-wide characterization and expression analysis of the Dof gene family related to abiotic stress in watermelon. PeerJ 8, e8358. https://doi.org/10.7717/peerj.8358 (2020).
Ma, J., Li, M.-Y., Wang, F., Tang, J. & Xiong, A.-S. Genome-wide analysis of Dof family transcription factors and their responses to abiotic stresses in Chinese cabbage. BMC Genom. 16, 33. https://doi.org/10.1186/s12864-015-1242-9 (2015).
Han-Hua Liu, X. T., Li, Y.-J., Chang-Ai, W. & Zheng, C.-C. Microarray-based analysis of stress-regulated microRNAs in Arabidopsis thaliana. RNA-A Publ. RNA Soc. 14, 836–843 (2008).
Chen, L., Zhang, S., Zhao, H., Korpelainen, H. & Li, C. Sex-related adaptive responses to interaction of drought and salinity in Populus yunnanensis. Plant Cell Environ. 33, 1767–1778 (2010).
Acknowledgements
This work was supported by startup fund from Fujian Agriculture and Forestry University and Natural Science Foundation of Fujian Province of China (2019J05055).
Author information
Authors and Affiliations
Contributions
R.M. and H.Y. conceived the project and designed experiments. H.Y., Y.M. and Y.L. performed the qRT-PCR experiments. H.Y. and Y.M. draw the figures. H.Y. and J.Y. discussed the results. H.Y. wrote the manuscript and R.M. revised it.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yu, H., Ma, Y., Lu, Y. et al. Expression profiling of the Dof gene family under abiotic stresses in spinach. Sci Rep 11, 14429 (2021). https://doi.org/10.1038/s41598-021-93383-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-93383-6
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.