Abstract
Cryptococcal meningitis is a life-threatening disease among immune compromised individuals that is caused by the opportunistic fungal pathogen Cryptococcus neoformans. Previous studies have shown that the fungus is phagocytosed by dendritic cells (DCs) and trafficked to the lysosome where it is killed by both oxidative and non-oxidative mechanisms. While certain molecules from the lysosome are known to kill or inhibit the growth of C. neoformans, the lysosome is an organelle containing many different proteins and enzymes that are designed to degrade phagocytosed material. We hypothesized that multiple lysosomal components, including cysteine proteases and antimicrobial peptides, could inhibit the growth of C. neoformans. Our study identified the contents of the DC lysosome and examined the anti-cryptococcal properties of different proteins found within the lysosome. Results showed several DC lysosomal proteins affected the growth of C. neoformans in vitro. The proteins that killed or inhibited the fungus did so in a dose-dependent manner. Furthermore, the concentration of protein needed for cryptococcal inhibition was found to be non-cytotoxic to mammalian cells. These data show that many DC lysosomal proteins have antifungal activity and have potential as immune-based therapeutics.
Similar content being viewed by others
Introduction
Cryptococcus neoformans is an encapsulated fungal pathogen that can cause pneumonia and meningitis in immune compromised individuals1,2. C. neoformans is an environmental yeast that is associated with decayed wood and pigeon droppings, and therefore, humans and animals breathe it in frequently3,4. Fortunately, the pathogen is typically quickly cleared from the lungs by a Th-1 type CD4+ T cell response before it can cause a symptomatic infection2. However, in cases where a person has a compromised immune system, such as reduced CD4+ T cells in patients with HIV/AIDS or in individuals taking immunosuppressive drugs to prevent organ transplant rejection, the fungus can cause an infection5,6. The prevalence between AIDS and cryptococcosis is so high that the CDC has named it an AIDS-defining illness, and the occurrence of cryptococcal meningitis is most prevalent in sub-Saharan Africa7,8. An analysis of the 2014 Joint UN Programme on HIV and AIDS showed that of the 223,100 yearly cases of cryptococcal meningitis, 73% occurred in sub-Saharan Africa (162,500 cases) with estimated yearly death totals of 181,100 and 135,900 worldwide and in sub-Saharan Africa, respectively9. While these numbers have been trending downwards in recent years, due to its high mortality rate there is still a case to be made for the World Health Organization (WHO) to name cryptococcal meningitis as a neglected tropical disease7,10,11. The most popular treatment programs are all based around three drugs: Amphotericin B, flucytosine, and fluconazole12,13. However, each is associated with certain barriers and limitations such as bioavailability, host toxicity, and emergence of resistant cryptococcal strains14,15,16.
While an adaptive T cell response is required for normal host clearance of C. neoformans, initial interactions with the fungus occur with innate immune cells in the lung such as dendritic cells (DCs) which are the cells responsible for antigen presentation to naive T cells17,18. The lungs contain a dense network of DCs, and during a cryptococcal infection, additional monocyte-derived DCs are recruited to the site of infection19,20,21,22. Upon encountering the fungus, DCs will phagocytose the opsonized pathogen, killing the organism in the phagolysosome23,24,25. The lysosome itself contains many different molecules (including proteases and antimicrobial peptides—AMPs) each with their own distinct role in degrading many different pathogens and/or facilitating the presentation of antigens via the MHC II pathway to the adaptive immune system26,27,28,29. For instance, AMPs such as cationic defensins are antifungal and the suggested mechanism of action is insertion into the fungal membrane leading to cell lysis30,31,32. Also, mice deficient in cathepsins L and S, which are cysteine proteases (these make up the majority of proteases in the lysosome), have problems with invariant chain (Ii) cleavage, which is necessary for antigen presentation33. Resolution of C. neoformans requires proper antigen presentation and activation of T cells5,34.
It has been previously shown that DCs (both murine bone marrow-derived DCs and human PMBC-derived DCs) can kill C. neoformans both in vitro and in vivo by oxidative and non-oxidative mechanisms26,35,36. However, it is possible for the fungus to survive and replicate inside other immune cells such as macrophages37,38. Furthermore, the purified contents of DC lysosomes can also kill the fungus in vitro in a dose-dependent manner26,36. These studies illustrate that it is indeed the components of the lysosome that act upon the pathogen. Since the lysosome is a generalized killing organelle designed to destroy many pathogens, it contains many different substances that individually may or may not have an effect on C. neoformans39. Thus far, it has been shown that one component within the DC lysosome can inhibit the growth of this yeast: cathepsin B. This cysteine protease is able to form a hole in the cell wall that results in osmotic lysis of C. neoformans26. In addition, use of enzymatic inhibitors of cathepsin B enhances the antifungal activity, suggesting that this protease may be acting in a non-enzymatic fashion26. Further studies are underway in our laboratory to determine cathepsin B’s mechanism of action against C. neoformans.
Our previous studies showed that in an acidic buffer (pH 5.5) the entire contents of the DC lysosome as well as a specific lysosomal component could kill C. neoformans26,36. The lysosome is rich in enzymes and other antimicrobial mediators27,40, and we hypothesized that additional lysosomal components have antifungal properties. Therefore, in the present study, the contents of the DC lysosome were investigated to identify additional anti-cryptococcal molecules that have activity in similar conditions to the DC lysosome (pH 5.5). First, the lysosome extract was fractionated by molecular weight and individual fractions were tested for antifungal activity. Next, mass spectrometry was performed on the DC lysosome extract fractions to identify all of the components present. A panel of lysosomal components was selected for their antimicrobial properties and screened against C. neoformans in vitro, and those that showed antifungal activity were further studied in dose-dependent assays. Finally, those concentrations that inhibited the growth of the fungus were assessed for cytotoxic effects against mammalian cells near physiological pH (7.5). The purpose of the current study was to identify the anticryptococcal components of the DC lysosome and to assess their potential to be used as therapeutics.
Results
C. neoformans growth is inhibited by DC-derived lysosomal fractions
Previous studies have shown that C. neoformans growth can be inhibited by DCs21,35 as well as by DC-derived lysosomal extract36. Our previous studies showed that cathepsin B, which is found in the DC lysosome, can also significantly inhibit the growth of the fungus26. In order to identify other proteins within the lysosome that can inhibit the growth of C. neoformans, DC-derived lysosomal extract was first fractionated by molecular weight, and fractions were verified by SDS-PAGE (Fig. 1a). Protein concentration was measured in each fraction, and those containing at least 50 μg/ml protein (the protein concentration that was antifungal for DC lysosomal extract36)—fractions 1, 7, 8, and 10—were used in anti-cryptococcal assays. C. neoformans strain H99 was incubated at 1 × 105 cells/ml with either individual fractions in phosphate buffer (pH 5.5), total extract in phosphate buffer, or in phosphate buffer alone for 24 h at 37 °C, 5% CO2. All fractions that were tested (fractions 1, 7, 8, and 10) showed significant killing (p < 0.0001) of the cryptococcal growth when compared to the C. neoformans inoculum (Fig. 1b). These data show that multiple fractions of DC lysosomal extract have anti-cryptococcal activity, and therefore, there are multiple compounds within the lysosomal extract that can kill C. neoformans or inhibit cryptococcal growth. In order to further identify individual proteins within the lysosomal extract with anti-cryptococcal activity, the fractions were analyzed by mass spectrometry to identify the contents of each fraction. Analysis revealed over 3000 different proteins present within the lysosomal fractions (Supplementary Table S1).
Fractions of DC Lysosomal Extract Have Anti-cryptococcal Activity. (a) Crude DC-derived lysosomal extracts were separated by molecular weight using a GELFrEE 8100 system into 12 different fractions and further resolved by standard SDS-PAGE. Proteins in the 12.5% gels were stained using BioSafe Coomassie Stain and visualized using a Gel Doc XR + system. (b) C. neoformans serotype A strain H99 yeast cells (1 × 105 cells/ml) were incubated in phosphate buffer (pH 5.5) with whole lysosomal extract or fractions of lysosomal extract for 24 h at 37 °C, 5% CO2, following which CFUs in the wells were determined. Data shown are means ± standard errors of the means (SEM) of the results of three independent experiments (n = 3), with each condition performed in duplicate. Two-tailed t-tests comparing H99 inoculum to each treatment were performed with asterisks (*) indicating significantly lower CFUs of treatment compared to the H99 inoculum (fraction 8: p = 0.0001, all others: p < 0.0001).
DC lysosomal proteins have anti-cryptococcal activity
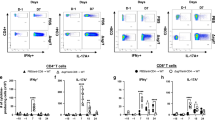
In order to determine the anti-cryptococcal potential of DC lysosomal proteins, ten different lysosomal proteins were tested for anti-cryptococcal activity. From the > 3000 proteins identified by mass spectrometry, ten test proteins were selected based on potential antimicrobial activity, such as those that are known antimicrobial peptides or cysteine proteases41,42,43,44. Concentrations of individual proteins used in our anti-cryptococcal assay were based upon either prior published concentrations45,46,47,48 or were based on our previous studies with cathepsin B26,45,46,47,48, where concentrations of 50 and 10 µg/ml were used. Specific protein concentrations used in this study are provided in the methods section. C. neoformans strain H99 was incubated at 1 × 105 cells/ml in a 50 μl volume separately with each protein in sterile 0.1 mM phosphate buffer, pH 5.5 (phosphate buffer) or in phosphate buffer alone for 24 h at 37 °C, 5% CO2, followed by diluting then plating on YPD agar to quantify CFUs. Percent cryptococcal inhibition was defined as CFUs of the experimental condition (C. neoformans incubated with test protein) divided by the CFUs of the control condition (C. neoformans incubated with phosphate buffer alone). We observed that incubation of C. neoformans with five of the lysosomal proteins (Coronin, HNE, MMP25, MPO, NOSTRIN) led to a significant (p < 0.0001) reduction of CFUs when compared to the C. neoformans grown in phosphate buffer alone which is shown as a high percent inhibition (Fig. 2a), while the remaining five proteins either did not inhibit fungal growth with low percent inhibition (cystatin B, striatin) or significantly enhanced fungal growth (calmodulin, p < 0.0001; calnexin, p < 0.0001; S100A6, p = 0.0034), as shown by negative percent inhibition (Fig. 2b).
Cryptococcal Inhibition by Lysosomal Proteins. C. neoformans strain H99 yeast cells were grown in YPD broth for 18 h in a 30 °C shaking incubator, then washed and adjusted to a concentration of 1 × 105 cells/ml. Fungi were incubated in phosphate buffer alone (pH 5.5) or in phosphate buffer with each protein for 24 h at 37 °C, 5% CO2, and percent inhibition defined as 100%—(test CFUs/control CFUs) × 100%. Data shown are means ± standard error of the mean (SEM) of the percent inhibition of three independent experiments (n = 3), with each condition performed in triplicate. (a) Lysosomal proteins that showed significant cryptococcal inhibition (*) compared to H99 in phosphate buffer alone as indicated by two-tailed t-tests (p < 0.0001 for all). (b) Lysosomal proteins that showed no cryptococcal inhibition or significantly enhanced growth (*) by two-tailed t-tests when compared to H99 in phosphate buffer alone (p = 0.0034 for S100A6, p < 0.0001 for calmodulin and calnexin). (c) Reduced concentrations of select lysosomal proteins with white bars, striped bars, and black bars indicating protein concentration at 100%, 10%, and 1%, respectively. Two-tailed t-test comparing fungal growth with protein against growth in phosphate buffer alone confirmed significant inhibition (*) at 10% for coronin (p < 0.0001), HNE (p < 0.0001), and MMP25 (p = 0.0008). MPO showed significant inhibition at 1% (p = 0.0213) but not 10% while NOSTRIN was not able to reduce growth at any of the lower concentrations. At 1% original concentration, MMP25 significantly (p = 0.0010) increased cryptococcal growth.
Cryptococcal inhibition is dose-dependent
To further elucidate the effective anti-cryptococcal concentration of each lysosomal protein, similar assays with reduced protein concentration were conducted on the five proteins that showed cryptococcal inhibition. Each of the five proteins were diluted to 10% and 1% of original concentration and incubated under the same conditions. Of those tested (coronin, HNE, MMP25, MPO, and NOSTRIN), only coronin (p < 0.0001), HNE (p < 0.0001), and MMP25 (p = 0.0008) significantly inhibited cryptococcal growth at the 10% protein concentration, but did not inhibit growth at the 1% concentration (Fig. 2c). Surprisingly, MMP25 significantly enhanced fungal growth (p = 0.0010) at 1% of the original concentration when compared to growth in phosphate buffer alone. While MPO was not able to reduce the growth of H99 at 10% concentration, when tested at 1%, it did have the capacity to significantly inhibit (p = 0.0213) the growth of the fungus. NOSTRIN was not effective at either of the reduced concentrations.
Minimum inhibitory concentration (MIC) of anti-cryptococcal lysosomal proteins
Next, in order to verify that the concentrations used for the previous experiments were similar to the minimum inhibitory concentrations (MIC), the five anti-cryptococcal lysosomal proteins were evaluated for MICs against C. neoformans strain H99 yeast cells. Highest concentrations of proteins started at the same maximum value as previous assays and ranged from 50 µg/ml to 0.0488 µg/ml (for HNE and MMP25), 25 µg/ml to 0.0224 µg/ml (for coronin), 18 µg/ml to 0.0176 µg/ml (for MPO), and 2 µg/ml to 0.0020 µg/ml (for NOSTRIN). Proteins were assessed in phosphate buffer (pH 5.5) as in our antifungal assays above as well as a more physiological relevant condition in RPMI-MOPS (pH 7.0), which is the media typically used in fungal MIC assays. Assays were completed over 48 h at 35 °C as per Clinical and Laboratory Standards Institute (CLSI)49,50,51,52. Results for these assays are summarized in Table 1. When testing in lysosomal-like conditions (pH 5.5) HNE, MMP25, and NOSTRIN retained similar anti-cryptococcal efficacy. In addition, these three compounds also retained some ability to inhibit yeast growth at pH 7.0. MPO was unable to inhibit cryptococcal growth at the highest tested concentration of 18 µg/ml under both conditions and did not replicate its previous effectiveness (Fig. 2a). MIC values for coronin were not able to be determined in these assays.
Due to the nephrotoxicity associated with high doses of Amphotericin B (AmpB), examining methods to increase efficacy and lower dosage has become an area of intense research53,54,55. One method is the use of a combination of compounds that, by themselves, display anti-cryptococcal properties, such as current guidelines that advise treating cryptococcal meningitis using combination therapy with AmpB and flucytosine56. Therefore, we tested our anti-cryptococcal lysosomal proteins in combination with AmpB against the fully virulent C. neoformans strain H99 using a checkerboard method and categorized data based on the fractional inhibitory concentration indices (FICI)57. While still retaining their previously stated anti-cryptococcal capacity, none of the lysosomal proteins showed any synergistic effects when used in combination with AmpB (Supplemental Table S2). Each protein/AmpB combination scored between 1.0 and 1.8 placing them comfortably within the indifference category (0.5–4.0) and provided neither antagonistic nor synergistic results.
Time-kill kinetics of lysosomal proteins against C. neoformans
MIC assays are useful starting points for understanding antimicrobial properties of unknown compounds, since they have been standardized by various laboratories and agencies. The time-kill assay studies the kinetic rates of inhibition over a period of time rather than examining only the end result. While they give great quantitative results and the ability to explore fungicidal rates, their methods are not as widely uniform and they can become time consuming the more time points and concentration one examines58. For this, we used a modified version of the method proposed in a paper by Klepser et. al which has since been cited over 200 times since its publication59. Cryptococcal cells were incubated with 2X MIC values (listed in Table 1 or 2X Table 1 or 2X highest concentration tested if none could be determined) in an acidic phosphate buffer (pH 5.5) for 48 h. As conditions change within the test chamber, properties of the proteins may also be altered, therefore, pH was monitored throughout the study, but we did not observe any changes in pH over the course of the study. We measured the percent inhibition of cryptococcal growth over time using the five anti-cryptococcal lysosomal proteins (Fig. 3). Three of the five (NOSTRIN, HNE, and MPO) displayed sustained inhibition starting from as early as three hours post inoculation. Meanwhile, at the 3 h time point, coronin and MMP25 exhibited intermediate and no inhibition, respectively. However, by 24 h, the latter two proteins showed inhibition and caught up to the other three by 48 h.
Time-kill Plots Demonstrating Cryptococcal Inhibition by Lysosomal Proteins. C. neoformans strain H99 yeast cells were grown in YPD broth for 18 h in a 30 °C shaking incubator, then washed and adjusted to a concentration of 1 × 105 cells/ml. Fungi were incubated in phosphate buffer alone (pH 5.5) or in phosphate buffer with each protein at 2X MIC for 48 h at 35 °C. Aliquots were taken and plated for CFUs for each hour 0–8, and at 12, 24, 36, and 48 h. Percent inhibition is defined as 100%—(protein CFUs/control CFUs) × 100%. Data shown are means of two independent experiments (n = 2). Lysosomal proteins NOSTRIN, HNE, and MPO showed sustained cryptococcal inhibition by 3 h post-inoculation while coronin displayed intermediate inhibition and MMP25 exhibited no cryptococcal inhibition at the same time point. From 24–48 h all proteins inhibited cryptococcal growth.
Cytotoxicity of DC lysosomal proteins
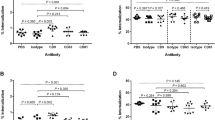
To test the relative cytotoxicity of these anti-cryptococcal lysosomal proteins on mammalian cells, a murine macrophage cell line (J774A.1) was used in conjunction with the Vybrant Cytotoxicity Assay Kit (G6PD assay) (Molecular Probes, Eugene, OR). Mammalian cells were incubated with the highest concentration of anti-cryptococcal protein (stated in methods section) in cell culture media or in cell culture media alone for 24 h at 37 °C, 5% CO2, then cytotoxicity assay was performed at near physiological pH (7.5). After fluorescence was normalized to control wells (J774A.1 macrophages in cell culture media alone), percent cytotoxicity was defined as fluorescence of experimental wells (macrophages and proteins) divided by positive control wells (macrophages and cell-lysis buffer). A compound is defined as “cytotoxic” if there is a reduction of more than 30% of viable cells when compared to control wells60. All proteins showed low cytotoxicity (< 20%) at the highest concentration tested for cryptococcal inhibition assays (Fig. 4).
Cytotoxicity of Lysosomal Proteins. J774A.1 macrophages (5 × 105 cells/ml) were incubated in cell culture media alone or cell culture media with each antifungal protein at 100% concentration for 24 h at 37 °C, 5% CO2, and percent cytotoxicity was calculated as per manufacturer’s instructions (pH 7.5). Data shown are means ± standard error of the means (SEM) of the cumulative results of three independent experiments (n = 3), with each condition performed in triplicate. All proteins tested had low cytotoxicity (< 20%).
Discussion
The primary function of the DC lysosome is to break down many different biomolecules including pathogens that may cause harm to a host in order to generate peptide fragments for antigen presentation39,61. This rapid degradation by the lysosome favors presentation through the MHC II pathway rather than cross presentation via MHC I62. Studies have shown that it can kill a wide variety of pathogens regardless of whether they are fungal, bacterial, or viral63,64,65,66,67. Previous studies have shown that C. neoformans traffics to the lysosomal compartment of DCs and is killed by its components35,36. It was also shown that contents of the DC lysosomal extract have activity against the fungus in vitro26,36. Furthermore, the DC lysosomal component cathepsin B can inhibit or kill the pathogen26. Due to the problems associated with current anti-cryptococcal drug therapies, finding new sources of treatment has become a major area of research.
To combat these ever-changing problems, most current research focuses on either discovering novel anti-fungal drugs or repurposing older drugs68,69,70,71. Our research focuses on identifying DC lysosomal components that are anti-cryptococcal and determining how these can be used to develop immune-based therapies to combat cryptococcosis. In accordance with our previous study on DC lysosomal proteins and their anti-cryptococcal effects, our current study has shown there are multiple DC lysosomal compounds that can inhibit and/or kill C. neoformans in an acidic buffer (pH 5.5) in a dose dependent manner. Three (HNE, MMP25, MPO) of the five compounds that showed anti-cryptococcal activity are known proteases that have been shown to have antimicrobial activity72,73,74. The other two are both associated with antimicrobial activity by other means: coronin is an actin binding protein that is involved with phagocytosis in leukocytes75 and NOSTRIN modulates the release of the oxidative stressor nitric oxide76. Several of these proteins still displayed inhibition at reduced concentrations which shows dose-dependent activity. Contrary to these results, however, reduced concentrations of MMP25 showed significant increase in fungal growth and may be a cause for concern if used as a therapeutic.
In order to verify that the concentrations used in these studies were in the range of minimum inhibitory concentration (MIC) concentrations, we tested the five antifungal proteins (Fig. 2) (coronin, HNE, MPO, MMP25, and NOSTRIN) in MIC assays in accordance with CLSI guidelines49,50,51,52. Three proteins (HNE, MMP25, and NOSTRIN) showed promising results, as not only did they recapitulate their effects in low pH, but also displayed some anti-cryptococcal activity at the higher pH, albeit at a lower efficiency. This was expected as pH changes can affect the activity of many lysosomal proteins, notably cathepsins77. Despite the lower activity, these results point to a future of promising therapeutics for systemic cryptococcosis, since some of these proteins can also be antifungal at neutral pH. Two proteins had different MIC results compared to our original lysosomal assays. However, differences in methodology between standard MIC assays (conducted in non-CO2 conditions and at 35 °C, as is standard MIC procedures in the field) and our antifungal assays (conducted at 37 °C, 5% CO2), may have contributed to these discrepancies.
Currently, there are only a few classes of antifungal compounds, all of which can become toxic to the host at high concentrations53,54. Despite being the drug of choice when treating cryptococcal meningitis, AmpB does have these same concerns, especially targeting the kidneys, causing nephrotoxicity. We sought ways to reduce standard treatment concentrations by investigating synergistic activity that may exist between it and our anti-cryptococcal lysosomal proteins. However, none of the compounds showed any synergy when combined with AmpB at any of concentrations tested. These results are not too surprising owing to the fact that the H99 strain, while fully virulent, is still susceptible to AmpB at low doses and other compounds may not show synergistic effects. These results may differ and show synergistic effects when testing against a cryptococcal strain that is resistant to AmpB. Furthermore, synergistic combinations tend to target both pathogenic processes and cell growth to completely inhibit a fungal infection78. More pre-clinical research in this area is needed as these studies were conducted in vitro. We also wanted to further understand the kinetics of these anti-cryptococcal lysosomal proteins, so a time-kill assay was conducted to evaluate growth over 48 h, the same as a standard MIC assay. Three (HNE, MPO, NOSTRIN) of the five proteins showed sustained inhibition quickly, within 3 h post-incubation. However, by the time point of our typical lysosomal assays (24 h), the remaining lysosomal proteins all showed inhibition of cryptococcal growth, which was sustained through the 48 h experiment.
Despite previous reports of antimicrobial activity against bacterial pathogens79,80,81,82 and/or interference with virulence pathways (such as the calcium-binding pathway44,83), several proteins tested in this study (calmodulin, calnexin, cystatin B, S100A6, and striatin) showed either no cryptococcal inhibition or enhanced fungal growth. We hypothesized that this enhanced growth may be attributed to secretion of nutrients or trace metals that may enhance cryptococcal growth and we are currently analyzing the contents of media in these conditions. Additional studies are also underway in our laboratory to examine the mechanism(s) by which these components enhance cryptococcal growth. Cystatin B is a cysteine protease that inhibits cathepsin B by tightly binding to it84. While cystatin B did not show any efficacy in our studies, previous work has shown that when cathepsin B and other inhibitors (calpain and CA-074) are used in conjunction with cathepsin B, they lead to increased inhibition C. neoformans growth when compared to cathepsin B treatment alone26. Striatin proteins have a diverse range of functions in both filamentous fungi and mammals including the signaling of motor functions and cell division85,86,87. This family and the closely related pyristriatins that have select antimicrobial activity against Gram positive bacteria and some filamentous fungi as well as yeasts80. When tested against our yeast, C. neoformans, striatin showed no change in the cryptococcal growth. The S100 series of proteins are known antimicrobial peptides and are also known as alarmins, and these proteins have a range of antimicrobial activity88,89. S100A8 and S100A9 form a heterodimer named calprotectin, which has anti-cryptococcal activity by the depletion of zinc from the fungus90,91,92. However, the related protein S100A6, which has similar zinc-binding potential, significantly enhanced the cryptococcal growth93. The final two proteins tested, calmodulin and calnexin, also significantly enhanced cryptococcal growth. Calmodulin is a calcium binding protein that activates calcineurin and is implicated in increased pathogen defense44,94. The membrane associated protein calnexin is also a signaling protein that functions as a chaperone but has been also associated with increased adaptive immunity95. DCs that express higher than normal levels of calnexin were able to induce T cell function greater than unmodified DCs96. It is currently unknown why these proteins had either limited or opposite effects than previously stated literature would suggest or by what means they increased cryptococcal growth.
To our knowledge, the cytotoxicity of DC lysosomal proteins has not been tested in mammalian cells. Our findings indicated that of the proteins tested, none were cytotoxic at the concentrations needed for anti-cryptococcal activity. This shows promise that these compounds may be used further in the treatment for cryptococcosis. However, the physiological relevance and bioavailability of these compounds in a live model remains to be seen. Our laboratory is currently in the process of testing these lysosomal components in a murine model of cryptococcosis. Furthermore, the complex interactions of these proteins with other proteins residing within the lysosome has not been investigated. None of these proteins are usually present alone or act upon pathogens individually, and we recognize that there are many more proteins and protein combinations that may add to the complexity of these interactions. To help us understand these problems, we are actively working on elucidating the mechanism(s) of action for these anti-cryptococcal proteins, studying interaction of multiple lysosomal proteins, and identifying the mechanism(s) of enhanced cryptococcal growth in our lab.
The present study has investigated the anti-cryptococcal activity of specific proteins of the DC lysosome. Five different fractions showed similar anti-cryptococcal activity to the entire extract. Mass spectrometry revealed many different compounds present within this extract that could potentially inhibit cryptococcal growth. Our results show that not only are there several that indeed inhibit cryptococcal growth, but these are also not cytotoxic to mammalian cells at these concentrations. From these findings, novel immune-based anti-cryptococcal treatments may be developed from immune-based proteins that could ease the burden brought on by this deadly disease.
Methods
Strains and media
Cryptococcus neoformans strain H99 (serotype A, mating type α) was recovered from 15% glycerol stocks stored at − 80 °C and were cultured for 18 h at 30 °C with shaking in yeast extract-peptone-dextrose (YPD) broth (BD Difco; Franklin Lakes, NJ) and collected by centrifugation. Organisms were washed three times with sterile phosphate-buffered saline (PBS), and viable yeast cells were quantified using trypan blue dye exclusion in a hemocytometer. Cryptococcal cells were resuspended in appropriate medium at the concentration needed for each experiment.
Mice
Female and male BALB/c (H-2d) mice were purchased from the National Cancer Institute/Charles River Laboratories and were housed either at Oklahoma State University Animal Resources or at The University of Texas at San Antonio Small Animal Laboratory Vivarium. Mice were handled according to approved guidelines, authors complied with the ARRIVE guidelines, and the study and experimental design was approved by the Oklahoma State University Institutional Animal Care and Use Committee (IACUC) or the University of Texas at San Antonio IACUC.
Generation of bone marrow-derived dendritic cells (BMDCs)
BMDC culture was performed as previously described26,36. Briefly, bone marrow was flushed from the femurs and tibiae of mice using sterile HBSS. Bone marrow cells were washed, counted, and plated in complete medium (RPMI-1640 supplemented with 10% FBS, 2 mM l-glutamine, 100 U penicillin/ml, 100 μg streptomycin/ml, and 50 mM 2-mercaptoethanol) supplemented with 20 ng/ml recombinant murine GM-CSF (Peprotech, Rocky Hill, NJ) at a concentration of 2 × 105 cells/ml in 10 ml (2 × 106 cells/plate). Cells were incubated at 37 °C, 5% CO2 for a total of eight days. At day 3 of incubation, 10 ml complete medium + GM-CSF was added to each plate. At day 6 of incubation, half of the medium was removed, and 10 ml fresh complete medium + GM-CSF was added. BMDCs were harvested at day 8, and DCs were purified by negative selection using anti-F4/80 microbeads (Miltenyi Biotec, Auburn, CA) (to remove contaminating macrophages) followed by positive selection using magnetically labeled anti-CD11c microbeads according to the manufacturer’s protocol (Miltenyi Biotec). Dendritic cell purity following this procedure was > 90% by flow cytometry.
Generation of lysosomal extract from BMDCs
DC lysosomal extracts were obtained as previously described using the lysosome isolation kit (Sigma-Aldrich, St. Louis, MO)26,36. Briefly, DCs were lysed using 1X extraction buffer (Sigma-Aldrich) followed by homogenization with a PowerGen 700 homogenizer (Fisher Scientific, Pittsburgh, PA) using the 7 × 110 mm homogenizer tip passed through the cells 15–20 times to disrupt 75–80% of the cells. The homogenized cells were then centrifuged at 1000 × g for 10 min to remove intact cells and cellular debris. The first supernatant was removed and centrifuged at 20,000×g for 20 min to pellet lysosomes. Lysosome purity was verified by flow cytometry for lysosomal marker LAMP1 (BD Biosciences, San Jose, CA), which showed > 90% pure lysosomes from this procedure. The pellet containing lysosomes was resuspended and sonicated for 20 s at 40% power on a Model 300 VT Ultrasonic Homogenizer (BioLogics, Inc., Manassas, VA), resulting in lysosome extract (600 mg/ml).
Fractionation and SDS-PAGE of lysosomal extract
Twelve molecular weight protein fractions of crude lysosomal extract were separated from crude lysosomal extract using GELFREE 8100 System (Expedeon, San Diego, CA). The system uses HEPES running buffer and Tris Acetate sample buffer (Expedeon), and the system was run according to manufacturer’s instructions as previously described97. Briefly, lysosomal extract was mixed with sample buffer and loaded into the 8% Tris–Acetate cartridge (Expedeon). Following loading of the sample, the instrument was paused for sample collection at predefined intervals. This process was repeated until all 12 fractions were collected. After the collection of fractions, standard SDS-PAGE was performed. For this, a 12.5% Precast Gel (Bio-Rad, Hercules, CA) was loaded with a Precision Protein Plus ladder (Bio-Rad) and each of the 12 samples. Gels were run for 55 min at 200 V in Tris/glycine/SDS running buffer (Bio-Rad). Protein bands in the gel were stained using BioSafe Coomassie Stain (Bio-Rad), and a Gel Doc XR + system (Bio-Rad) was used for visualization.
Anti-cryptococcal assays with lysosomal fractions
Anti-cryptococcal assays were performed as previously described26,36. Briefly, C. neoformans was resuspended in phosphate buffer (0.1 mM potassium phosphate monobasic supplemented with 2% RPMI-1640, pH 5.5) at concentration of 1 × 105 cells/ml and added to triplicate wells of a 96-well plate in a 50 μl volume. Lysosomal fractions were added to individual triplicate well sets at 50 μl volume for a final concentration of 50 μg/ml. Negative control wells included 50 μl C. neoformans with 50 μl of phosphate buffer to equal the volume of experimental wells. Positive control wells contained 50 μl C. neoformans and 50 μl DC lysosomal extract (at 50 ug/ml). Plates were incubated for 24 h at 37 °C, 5% CO2. Following incubation, cryptococcal cells in the plates were diluted in PBS and plated onto yeast extract-peptone-dextrose plates supplemented with 1 microgram/ml (ug) chloramphenicol (YPD agar). The plates were incubated at 30 °C for 2 days, and then CFUs were enumerated. Killing was defined as significantly reduced CFUs compared to the inoculum, and inhibition was defined as significantly reduced CFUs compared to Cryptococcus grown alone in phosphate buffer. Each assay was conducted in three independent experiments (n = 3), with each condition performed in duplicate.
Identification of lysosomal proteins by HPLC–ESI–MS/MS
Peptide identifications were performed at the Institutional Mass Spectrometry Laboratory at The University of Texas Health Science Center at San Antonio as previously described97. Briefly, individual fractions were digested overnight at 37 °C with trypsin (Promega, sequencing grade, Madison, WI) in 40 mM NH4CO3/10% ACN. The tryptic peptides were extracted with 0.1% TFA followed by 0.1% TFA/50% ACN. The extracts were dried by vacuum centrifugation and resuspended in 0.5% TFA. Digests were analyzed by capillary HPLC–ESI–MS/MS using a Thermo Fisher LTQ linear ion trap mass spectrometer fitted with a New Objective PicoView 550 nanospray interface. On-line HPLC separation of the digests was accomplished with an Eksigent NanoLC micro HPLC: column, PicoFrit (New Objective; 75 μm id) packed to 10 cm with C18 adsorbent (Vydac; 218MS 5 μm, 300 Å); mobile phase A, 0.5% acetic acid/0.005% TFA; mobile phase B, 90% ACN/0.5% acetic acid/0.005% TFA; gradient 2 to 42% B in 30 min; flow rate, 0.4 μl/min. MS conditions were: ESI voltage, 2.9 kV; isolation window for MS/MS, 3; relative collision energy, 35%; scan strategy, survey scan followed by acquisition of data-dependent CID spectra of the seven most intense ions in the survey scan above a set threshold. Methionine oxidation and cysteine carbamidomethylation were considered as variable modifications for all searches. Scaffold 4.0 (Proteome Software, Portland, OR, USA) was used to conduct an X! Tandem subset search of the Mascot data followed by cross-correlation of the results of both searches. The Scaffold confidence levels for acceptance of peptide assignments and protein identifications were 95 and 99%, respectively. (Supplementary Table S1).
Anti-cryptococcal assays using DC lysosomal proteins
Anti-cryptococcal assays were performed as described above. Proteins were chosen based on previously-described antimicrobial activity, cysteine protease activity (similar to cathepsin B)26 or calcium interference, which affects calcineurin, a known virulence factor for C. neoformans44. All proteins were derived from commercial sources and were > 95% pure. Each protein was prepared at a concentration determined by either published studies or from our previous studies with lysosomal proteins26,45,46,47,48. Lysosomal proteins tested included calmodulin (1 mM) (Enzo Life Sciences, Farmingdale, NY), calnexin (50 μg/ml) (Novus Biologicals, Centennial, CO), coronin 1a (CORO1A) (25 μg/ml) (LSBio, Seattle, WA), cystatin B (5 μg/ml) (R&D Systems), human neutrophil elastase (HNE) (50 μg/ml) (LSBio), matrix metalloproteinase-25 (MMP25) (50 μg/ml) (OriGene, Rockville, MD), myeloperoxidase (MPO) (18 μg/ml) (Novus Biologicals), recombinant nitric oxide synthase traffic inducer (NOSTRIN) (2 μg/ml) (Novus Biologicals), S100A6 (25 μg/ml) (Novus Biologicals), and striatin (100 μg/ml) (Novus Biologicals). Lysosomal proteins were diluted to a 2X concentration in 0.1 mM phosphate buffer, pH 5.5 (phosphate buffer) and added to wells (50 μl) with 1 × 105 cells/ml C. neoformans strain H99 yeast cells diluted in phosphate buffer (50 μl) to equal 100 μl in each well, and plates were incubated at 37 °C, 5% CO2 for 24 h. After incubation, contents of wells were diluted and plated on YPD agar to quantify CFUs. Each protein was tested in three independent experiments (n = 3), with each condition performed in triplicate. Additional anti-cryptococcal assays were performed to test dose-dependent activity of the DC lysosomal proteins. For these, proteins used were diluted from working concentrations to 1:10 and 1:100 with phosphate buffer for 10% and 1% concentrations, respectively, and the anti-cryptococcal assay was conducted as described above. Each anti-fungal assay was conducted in three independent experiments (n = 3), with each condition performed in triplicate. Percent inhibition was defined as 100% (positive inhibition control) minus the experimental condition CFUs divided by the negative inhibition CFUs × 100%. Negative percent inhibition is the result of higher CFUs in the experimental compared to negative inhibition control.
Minimum inhibitory concentration (MIC) assays of anti-cryptococcal DC lysosomal proteins
Fungal growth inhibition of anti-cryptococcal DC lysosomal proteins were performed according to Clinical Laboratory Standards Institute (CLSI) guidelines M27: Reference Method for Broth Dilution Antifungal Susceptibility Testing of Yeasts, 4th edition49,50,51,52. Proteins were evaluated in twofold dilutions in concentrations ranging from 50 µg/ml to 0.0488 µg/ml (for HNE and MMP25), 25 µg/ml to 0.0224 µg/ml (for coronin), 18 µg/ml to 0.0176 µg/ml (for MPO), and 2 µg/ml to 0.0020 µg/ml (for NOSTRIN). Dilutions occurred in either phosphate buffer (0.1 mM potassium phosphate monobasic supplemented with 2% RPMI-1640, pH 5.5) or RPMI-MOPS (RPMI-1640 supplemented with 34.5 g/L MOPS, pH 7.0) in a 96-well microtiter plate. MICs assay were conducted with a non-visible concentration (0.5 × 103 cells/ml) of C. neoformans strain H99 yeast cells. Plates were incubated for 48 h at 35 °C and optical densities of 490 nm were taken on a Synergy HTX multi-mode plate reader (BioTek). MIC was defined as the lowest concentration of protein with no visible fungal growth. Each MIC assay was conducted in two independent experiments (n = 2), with each condition performed in at least duplicate.
Checkerboard assay using AmpB in combination with lysosomal proteins against C. neoformans
Anti-cryptococcal activity of AmpB in combination with anti-cryptococcal lysosomal proteins was measured in vitro using a checkerboard method as previously described57. Assays were conducted in either phosphate buffer (pH 5.5) or RPMI-MOPS (pH 7.0) (both described above) in 100 µl total. A 96-well plate was used with rows A-G and columns 1–9 used to create the checkerboard titrations between AmpB and a lysosomal protein. Controls used were protein only (row H), AmpB only (column 10), cryptococcal growth control (column 11), and media control (column 12). Concentrations were evaluated in twofold dilutions ranging from 4 μg/ml to 0.062 μg/ml (for AmpB), 25 μg/ml to 0.098 μg/ml (for coronin), 50 μg/ml to 0.195 μg/ml (for HNE and MMP25), 72 μg/ml to 0.281 μg/ml (for MPO), and 2 μg/ml to 0.0078 μg/ml (for NOSTRIN). C. neoformans strain H99 (0.5 × 103 cells/ml) was used to evaluate efficacy of combinations. Plates were incubated for 48 h at 35 °C and optical densities of 490 nm were taken on a Synergy HTX multi-mode plate reader (BioTek). Combinatory results were analyzed using the Fractional Inhibitory Concentration Index (FICI, a non-parametric model based on the Loewe additivity theory) whereas FICI ≤ 0.5 is synergistic, FICI 0.5–4 is indifference, and FICI ≥ 4 is antagonistic. FICIs were defined as the summation of individual FICs (FICI = FICAmp B + FICProtein) with FICs being defined as the MIC in combination divided by the MIC alone (FIC = MICCombination/MICAlone). Off-scale MICs were considered to be the highest or lowest concentration tested in the assay. Each checkerboard assay was conducted in two independent experiments (n = 2) for each lysosomal protein and for each buffer.
Time-kill assay
Rate of cryptococcal killing by lysosomal protein was measured using a time-kill assay as previously described59. C. neoformans strain H99 was tested at a concentration of 1 × 105 cells/ml with lysosomal proteins at 2X their respective MICs (coronin: 25 μg/ml, HNE: 3.125 μg/ml, MPO: 18 μg/ml, MMP25: 0.78 μg/ml, NOSTRIN: 0.25 μg/ml) in 1000 μl of phosphate buffer (pH 5.5). Solutions were incubated for 48 h at 35 °C with 50 μl aliquots being taken at hours 0–8, 12, 24, 36, and 48 post inoculation. Aliquots were diluted and plated on YPD agar for 48 h at 30 °C to enumerate CFUs. Each time-kill assay was conducted in two independent experiments (n = 2) for each protein along with fungus only growth control. Percent inhibition was calculated as described above with any negative inhibition values displayed as 0% inhibition.
Cytotoxicity assay
The macrophage cell line J774A.1 (TIB-67; ATCC, Manassas, VA) was used for cytotoxicity assays of the present study. J774A.1 cells were grown in cell culture medium (DMEM supplemented with 10% heat-inactivated fetal bovine serum (FBS), 10% NCTC-109, 1% non-essential amino acids, 100 U penicillin/ml, and 100 μg streptomycin/ml) in CytoOne T75 tissue culture flasks (USA Scientific, Ocala, FL) at 37 °C, 5% CO2. J774A.1 cells were passaged according to manufacturer’s directions. For detection of cytotoxicity of DC lysosomal proteins, the Vybrant Cytotoxicity Assay Kit (Molecular Probes, Eugene, OR) was used according to manufacturer’s instruction at near physiological pH (7.5). Briefly, J774A.1 cells were added to a 96-well plate in triplicate at a concentration of 10 × 105 cells/ml in 25 μl. Lysosomal proteins were prepared similarly to the anti-cryptococcal assay except cell culture media was used for dilutions instead of phosphate buffer. Each protein was used at 25 μl per well. Controls included no-cell, untreated cells, and fully lysed cells. All wells were prepared in triplicate. Plates were incubated for 24 h at 37 °C, 5% CO2. After incubation, 50 μl of reaction mixture was added to all wells. Fluorescent readings were taken on a Synergy HTX multi-mode plate reader (BioTek, Winooski, VT) with filters for 530/25 (excitation) and 590/20 (emission). Each cytotoxicity assay was conducted in three independent experiments (n = 3), with each condition performed in triplicate.
Statistical analysis
Unpaired two-tailed t-tests were used to compare CFUs using GraphPad Prism version 5.00 for Windows (GraphPad Software, San Diego, CA). Significant differences were defined as p < 0.05 and α = 0.05 unless noted otherwise.
Data availability
The mass spectrometry datasets generated during and/or analyzed during the current study are included in this published article (Supplementary Table S1).
References
Neal, L. M. et al. CD4(+) T cells orchestrate lethal immune pathology despite fungal clearance during Cryptococcus neoformans meningoencephalitis. MBio https://doi.org/10.1128/mBio.01415-17 (2017).
Wozniak, K. L., Olszewski, M. A. & Wormley, F. L. Jr. Molecules at the interface of Cryptococcus and the host that determine disease susceptibility. Fungal Genet. Biol. 78, 87–92. https://doi.org/10.1016/j.fgb.2014.10.013 (2015).
Wiesner, D. L. et al. Chitin recognition via chitotriosidase promotes pathologic type-2 helper T cell responses to cryptococcal infection. PLoS Pathog 11, e1004701. https://doi.org/10.1371/journal.ppat.1004701 (2015).
Levitz, S. M. The ecology of Cryptococcus neoformans and the epidemiology of cryptococcosis. Rev. Infect. Dis. 13, 1163–1169 (1991).
Gibson, J. F. & Johnston, S. A. Immunity to Cryptococcus neoformans and C. gattii during cryptococcosis. Fungal Genet. Biol. 78, 76–86. https://doi.org/10.1016/j.fgb.2014.11.006 (2015).
Pappas, P. G. et al. Invasive fungal infections among organ transplant recipients: Results of the Transplant-Associated Infection Surveillance Network (TRANSNET). Clin. Infect. Dis. 50, 1101–1111. https://doi.org/10.1086/651262 (2010).
Nyazika, T. K. et al. Epidemiology and aetiologies of cryptococcal meningitis in Africa, 1950–2017: Protocol for a systematic review. BMJ Open 8, e020654. https://doi.org/10.1136/bmjopen-2017-020654 (2018).
Warkentien, T. & Crum-Cianflone, N. F. An update on Cryptococcosis among HIV-infected patients. Int. J. STD AIDS 21, 679–684. https://doi.org/10.1258/ijsa.2010.010182 (2010).
Rajasingham, R. et al. Global burden of disease of HIV-associated cryptococcal meningitis: an updated analysis. Lancet Infect. Dis. 17, 873–881. https://doi.org/10.1016/s1473-3099(17)30243-8 (2017).
Molloy, S. F. et al. Cryptococcal meningitis: A neglected NTD?. PLoS Negl. Trop. Dis. 11, e0005575. https://doi.org/10.1371/journal.pntd.0005575 (2017).
Armstrong-James, D., Meintjes, G. & Brown, G. D. A neglected epidemic: Fungal infections in HIV/AIDS. Trends Microbiol. 22, 120–127. https://doi.org/10.1016/j.tim.2014.01.001 (2014).
Denning, D. W. & Hope, W. W. Therapy for fungal diseases: Opportunities and priorities. Trends Microbiol. 18, 195–204. https://doi.org/10.1016/j.tim.2010.02.004 (2010).
Molloy, S. F. et al. Antifungal combinations for treatment of cryptococcal meningitis in Africa. N. Engl. J. Med. 378, 1004–1017. https://doi.org/10.1056/NEJMoa1710922 (2018).
Bicanic, T. et al. Toxicity of amphotericin B deoxycholate-based induction therapy in patients with HIV-associated cryptococcal meningitis. Antimicrob. Agents Chemother. 59, 7224–7231. https://doi.org/10.1128/AAC.01698-15 (2015).
Sudan, A. et al. Pharmacokinetics and pharmacodynamics of fluconazole for cryptococcal meningoencephalitis: Implications for antifungal therapy and in vitro susceptibility breakpoints. Antimicrob. Agents Chemother. 57, 2793–2800. https://doi.org/10.1128/AAC.00216-13 (2013).
Bicanic, T., Harrison, T., Niepieklo, A., Dyakopu, N. & Meintjes, G. Symptomatic relapse of HIV-associated cryptococcal meningitis after initial fluconazole monotherapy: The role of fluconazole resistance and immune reconstitution. Clin. Infect. Dis. 43, 1069–1073. https://doi.org/10.1086/507895 (2006).
Wozniak, K. L. Interactions of Cryptococcus with dendritic cells. J. Fungi (Basel) https://doi.org/10.3390/jof4010036 (2018).
Mukaremera, L. & Nielsen, K. Adaptive immunity to Cryptococcus neoformans infections. J. Fungi https://doi.org/10.3390/jof3040064 (2017).
Holt, P. G., Schonhegrad, M. A. & Oliver, J. Mhc Class-Ii antigen-bearing dendritic cells in pulmonary tissues of the rat: Regulation of antigen presentation activity by endogenous macrophage populations. J. Exp. Med. 167, 262–274. https://doi.org/10.1084/jem.167.2.262 (1988).
Schonhegrad, M. A., Oliver, J., Mcmenamin, P. G. & Holt, P. G. Studies on the density, distribution, and surface phenotype of intraepithelial class-Ii major histocompatability complex antigen (Ia)-bearing dendritic cells (Dc) in the conducting airways. J. Exp. Med. 173, 1345–1356. https://doi.org/10.1084/jem.173.6.1345 (1991).
Wozniak, K. L., Vyas, J. M. & Levitz, S. M. In vivo role of dendritic cells in a murine model of pulmonary cryptococcosis. Infect. Immun. 74, 3817–3824. https://doi.org/10.1128/IAI.00317-06 (2006).
Wozniak, K. L. et al. Insights into the mechanisms of protective immunity against Cryptococcus neoformans infection using a mouse model of pulmonary cryptococcosis. PLoS ONE 4, e6854. https://doi.org/10.1371/journal.pone.0006854 (2009).
Bright, N. A., Gratian, M. J. & Luzio, J. P. Endocytic delivery to lysosomes mediated by concurrent fusion and kissing events in living cells. Curr. Biol. 15, 360–365. https://doi.org/10.1016/j.cub.2005.01.049 (2005).
de Duve, C. The lysosome turns fifty. Nat. Cell. Biol. 7, 847–849. https://doi.org/10.1038/ncb0905-847 (2005).
Luzio, J. P. et al. Lysosome-endosome fusion and lysosome biogenesis. J. Cell. Sci. 113, 1515–1524 (2000).
Hole, C. R., Bui, H., Wormley, F. L. Jr. & Wozniak, K. L. Mechanisms of dendritic cell lysosomal killing of Cryptococcus. Sci. Rep. 2, 739. https://doi.org/10.1038/srep00739 (2012).
Lennon-Dumenil, A. M. et al. Analysis of protease activity in live antigen-presenting cells shows regulation of the phagosomal proteolytic contents during dendritic cell activation. J. Exp. Med. 196, 529–540. https://doi.org/10.1084/jem.20020327 (2002).
Guermonprez, P., Valladeau, J., Zitvogel, L., Thery, C. & Amigorena, S. Antigen presentation and T cell stimulation by dendritic cells. Annu. Rev. Immunol. 20, 621–667. https://doi.org/10.1146/annurev.immunol.20.100301.064828 (2002).
Wozniak, K. L. Dendritic cells at the interface of fungal immunity. Curr. Fungal Infect. Rep. 1, 89–95 (2007).
de Medeiros, L. N. et al. Backbone dynamics of the antifungal Psd1 pea defensin and its correlation with membrane interaction by NMR spectroscopy. Biochim. Biophys. Acta 1798, 105–113. https://doi.org/10.1016/j.bbamem.2009.07.013 (2010).
Silva, P. M., Goncalves, S. & Santos, N. C. Defensins: antifungal lessons from eukaryotes. Front. Microbiol. 5, 97. https://doi.org/10.3389/fmicb.2014.00097 (2014).
Thevissen, K. et al. Defensins from insects and plants interact with fungal glucosylceramides. J. Biol. Chem. 279, 3900–3905. https://doi.org/10.1074/jbc.M311165200 (2004).
Honey, K. & Rudensky, A. Y. Lysosomal cysteine proteases regulate antigen presentation. Nat. Rev. Immunol. 3, 472–482. https://doi.org/10.1038/nri1110 (2003).
Nelson, B. N., Hawkins, A. N. & Wozniak, K. L. Pulmonary macrophage and dendritic cell responses to Cryptococcus neoformans. Front. Cell. Infect. Microbiol. https://doi.org/10.3389/fcimb.2020.00037 (2020).
Kelly, R. M., Chen, J., Yauch, L. E. & Levitz, S. M. Opsonic requirements for dendritic cell-mediated responses to Cryptococcus neoformans. Infect. Immun. 73, 592–598. https://doi.org/10.1128/IAI.73.1.592-598.2005 (2005).
Wozniak, K. L. & Levitz, S. M. Cryptococcus neoformans enters the endolysosomal pathway of dendritic cells and is killed by lysosomal components. Infect. Immun. 76, 4764–4771. https://doi.org/10.1128/IAI.00660-08 (2008).
Davis, M. J. et al. Cryptococcus neoformans-induced macrophage lysosome damage crucially contributes to fungal virulence. J. Immunol. 194, 2219–2231. https://doi.org/10.4049/jimmunol.1402376 (2015).
Diamond, R. D. & Bennett, J. E. Growth of Cryptococcus neoformans within human macrophages in vitro. Infect. Immun. 7, 231–236 (1973).
Luzio, J. P., Pryor, P. R. & Bright, N. A. Lysosomes: Fusion and function. Nat. Rev. Mol. Cell Biol. 8, 622–632. https://doi.org/10.1038/nrm2217 (2007).
Savina, A. & Amigorena, S. Phagocytosis and antigen presentation in dendritic cells. Immunol. Rev. 219, 143–156. https://doi.org/10.1111/j.1600-065X.2007.00552.x (2007).
Zhang, L. J. & Gallo, R. L. Antimicrobial peptides. Curr. Biol. 26, R14-19. https://doi.org/10.1016/j.cub.2015.11.017 (2016).
Bahar, A. A. & Ren, D. Antimicrobial peptides. Pharmaceuticals (Basel) 6, 1543–1575. https://doi.org/10.3390/ph6121543 (2013).
Verma, S., Dixit, R. & Pandey, K. C. Cysteine proteases: Modes of activation and future prospects as pharmacological targets. Front. Pharmacol. 7, 107. https://doi.org/10.3389/fphar.2016.00107 (2016).
Steinbach, W. J., Reedy, J. L., Cramer, R. A. Jr., Perfect, J. R. & Heitman, J. Harnessing calcineurin as a novel anti-infective agent against invasive fungal infections. Nat. Rev. Microbiol. 5, 418–430. https://doi.org/10.1038/nrmicro1680 (2007).
Walkup, W. G. et al. A model for regulation by SynGAP-alpha1 of binding of synaptic proteins to PDZ-domain “Slots” in the postsynaptic density. Elife https://doi.org/10.7554/eLife.16813 (2016).
Rahayu, P., Marcelline, F., Sulistyaningrum, E., Suhartono, M. T. & Tjandrawinata, R. R. Potential effect of striatin (DLBS0333), a bioactive protein fraction isolated from Channa striata for wound treatment. Asian Pac. J. Trop. Biomed. 6, 1001–1007. https://doi.org/10.1016/j.apjtb.2016.10.008 (2016).
Liachko, N. F. et al. The phosphatase calcineurin regulates pathological TDP-43 phosphorylation. Acta Neuropathol. 132, 545–561. https://doi.org/10.1007/s00401-016-1600-y (2016).
de Jong, N. W. M. et al. Immune evasion by a staphylococcal inhibitor of myeloperoxidase. Proc. Natl. Acad. Sci. U S A 114, 9439–9444. https://doi.org/10.1073/pnas.1707032114 (2017).
CLSI. Reference Method for Broth Dilution Antifungal Susceptibility Testing of Yeasts. CLSI Standard M27 4th edn. (Clinical and Laborartory Standards Institutue, 2017).
Cordeiro, R. A. et al. Minimum inhibitory concentrations of amphotericin B, azoles and caspofungin against Candida species are reduced by farnesol. Med. Mycol. 51, 53–59. https://doi.org/10.3109/13693786.2012.692489 (2013).
Singh-Babak, S. D. et al. Global analysis of the evolution and mechanism of echinocandin resistance in Candida glabrata. PLoS Pathog 8, e1002718. https://doi.org/10.1371/journal.ppat.1002718 (2012).
Xie, J. L., Singh-Babak, S. D. & Cowen, L. E. Minimum inhibitory concentration (MIC) assay for antifungal drugs. Bio-Protoc. 2, e252. https://doi.org/10.21769/BioProtoc.252 (2012).
Abassi, M., Boulware, D. R. & Rhein, J. Cryptococcal meningitis: Diagnosis and management update. Curr. Trop. Med. Rep. 2, 90–99. https://doi.org/10.1007/s40475-015-0046-y (2015).
Houst, J., Spizek, J. & Havlicek, V. Antifungal drugs. Metabolites https://doi.org/10.3390/metabo10030106 (2020).
Perfect, J. R. The antifungal pipeline: A reality check. Nat. Rev. Drug Discov. 16, 603–616. https://doi.org/10.1038/nrd.2017.46 (2017).
Perfect, J. R. et al. Clinical practice guidelines for the management of cryptococcal disease: 2010 update by the infectious diseases society of america. Clin. Infect. Dis. 50, 291–322. https://doi.org/10.1086/649858 (2010).
Bonifacio, B. V. et al. Antifungal activity of a hydroethanolic extract from Astronium urundeuva leaves against Candida albicans and Candida glabrata. Front. Microbiol. 10, 2642. https://doi.org/10.3389/fmicb.2019.02642 (2019).
Bidaud, A. L., Schwarz, P., Herbreteau, G. & Dannaoui, E. Techniques for the assessment of in vitro and in vivo antifungal combinations. J. Fungi https://doi.org/10.3390/jof7020113 (2021).
Klepser, M. E., Ernst, E. J., Lewis, R. E., Ernst, M. E. & Pfaller, M. A. Influence of test conditions on antifungal time-kill curve results: Proposal for standardized methods. Antimicrob. Agents Chemother. 42, 1207–1212. https://doi.org/10.1128/AAC.42.5.1207 (1998).
ISO. Tests for In Vitro Cytotoxicity. Biological Evaluation of Medical Devices 3rd edn. (International Organization for Standardization, Switzerland, 2009).
Thakur, A., Mikkelsen, H. & Jungersen, G. Intracellular pathogens: Host immunity and microbial persistence strategies. J. Immunol. Res. 2019, 1356540. https://doi.org/10.1155/2019/1356540 (2019).
Embgenbroich, M. & Burgdorf, S. Current concepts of antigen cross-presentation. Front Immunol. 9, 1643. https://doi.org/10.3389/fimmu.2018.01643 (2018).
Newman, S. L. & Holly, A. Candida albicans is phagocytosed, killed, and processed for antigen presentation by human dendritic cells. Infect. Immun. 69, 6813–6822. https://doi.org/10.1128/IAI.69.11.6813-6822.2001 (2001).
Gildea, L. A., Ciraolo, G. M., Morris, R. E. & Newman, S. L. Human dendritic cell activity against Histoplasma capsulatum is mediated via phagolysosomal fusion. Infect. Immun. 73, 6803–6811. https://doi.org/10.1128/IAI.73.10.6803-6811.2005 (2005).
Clemens, D. L., Lee, B. Y. & Horwitz, M. A. Virulent and avirulent strains of Francisella tularensis prevent acidification and maturation of their phagosomes and escape into the cytoplasm in human macrophages. Infect. Immun. 72, 3204–3217. https://doi.org/10.1128/IAI.72.6.3204-3217.2004 (2004).
Wong, C. O. et al. Lysosomal degradation is required for sustained phagocytosis of bacteria by macrophages. Cell Host Microbe 21, 719–730. https://doi.org/10.1016/j.chom.2017.05.002 (2017).
Lin, Y. et al. Hepatitis B virus is degraded by autophagosome-lysosome fusion mediated by Rab7 and related components. Protein Cell 10, 60–66. https://doi.org/10.1007/s13238-018-0555-2 (2019).
Krysan, D. J. Toward improved anti-cryptococcal drugs: Novel molecules and repurposed drugs. Fungal Genet. Biol. 78, 93–98. https://doi.org/10.1016/j.fgb.2014.12.001 (2015).
Wall, G., Herrera, N. & Lopez-Ribot, J. L. Repositionable compounds with antifungal activity against multidrug resistant Candida auris identified in the medicines for malaria venture’s pathogen box. J. Fungi 5, 15. https://doi.org/10.3390/jof5040092 (2019).
Van Dijck, P. et al. Methodologies for in vitro and in vivo evaluation of efficacy of antifungal and antibiofilm agents and surface coatings against fungal biofilms. Microb. Cell 5, 300–326. https://doi.org/10.15698/mic2018.07.638 (2018).
Castro-Lopez, N. & Hung, C. Y. Immune response to coccidioidomycosis and the development of a vaccine. Microorganisms https://doi.org/10.3390/microorganisms5010013 (2017).
Belaaouaj, A., Kim, K. S. & Shapiro, S. D. Degradation of outer membrane protein A in Escherichia coli killing by neutrophil elastase. Science 289, 1185–1188. https://doi.org/10.1126/science.289.5482.1185 (2000).
Pei, D. Leukolysin/MMP25/MT6-MMP: A novel matrix metalloproteinase specifically expressed in the leukocyte lineage. Cell Res. 9, 291–303. https://doi.org/10.1038/sj.cr.7290028 (1999).
Klebanoff, S. J. Myeloperoxidase: Friend and foe. J. Leukoc. Biol. 77, 598–625. https://doi.org/10.1189/jlb.1204697 (2005).
Pieters, J. In The Coronin Family of Proteins. Subcellular Biochemistry Vol. 48 (eds Clemen, C. S. et al.) 116–123 (Landes Bioscience, 2008).
Zimmermann, K. et al. NOSTRIN: A protein modulating nitric oxide release and subcellular distribution of endothelial nitric oxide synthase. Proc. Natl. Acad. Sci. USA 99, 17167–17172. https://doi.org/10.1073/pnas.252345399 (2002).
Bonam, S. R., Wang, F. & Muller, S. Lysosomes as a therapeutic target. Nat. Rev. Drug Discov. 18, 923–948. https://doi.org/10.1038/s41573-019-0036-1 (2019).
Cui, J., Ren, B., Tong, Y., Dai, H. & Zhang, L. Synergistic combinations of antifungals and anti-virulence agents to fight against Candida albicans. Virulence 6, 362–371. https://doi.org/10.1080/21505594.2015.1039885 (2015).
Fajardo, M. et al. Calnexin, calreticulin and cytoskeleton-associated proteins modulate uptake and growth of Legionella pneumophila in Dictyostelium discoideum. Microbiology (Reading) 150, 2825–2835. https://doi.org/10.1099/mic.0.27111-0 (2004).
Richter, C., Helaly, S. E., Thongbai, B., Hyde, K. D. & Stadler, M. Pyristriatins A and B: Pyridino-Cyathane Antibiotics from the Basidiomycete Cyathus cf. striatus. J. Nat. Prod. 79, 1684–1688. https://doi.org/10.1021/acs.jnatprod.6b00194 (2016).
Yano, J. et al. The acute neutrophil response mediated by S100 alarmins during vaginal Candida infections is independent of the Th17-pathway. PLoS ONE 7, e46311. https://doi.org/10.1371/journal.pone.0046311 (2012).
Xiao, P. P., Hu, Y. H. & Sun, L. Scophthalmus maximus cystatin B enhances head kidney macrophage-mediated bacterial killing. Dev. Comp. Immunol. 34, 1237–1241. https://doi.org/10.1016/j.dci.2010.07.008 (2010).
Donato, R., Sorci, G. & Giambanco, I. S100A6 protein: functional roles. Cell Mol. Life Sci. 74, 2749–2760. https://doi.org/10.1007/s00018-017-2526-9 (2017).
Pol, E. & Bjork, I. Role of the single cysteine residue, Cys 3, of human and bovine cystatin B (stefin B) in the inhibition of cysteine proteinases. Protein Sci. 10, 1729–1738. https://doi.org/10.1110/ps.11901 (2001).
Castets, F. et al. A novel calmodulin-binding protein, belonging to the WD-repeat family, is localized in dendrites of a subset of CNS neurons. J. Cell Biol. 134, 1051–1062. https://doi.org/10.1083/jcb.134.4.1051 (1996).
Benoist, M., Gaillard, S. & Castets, F. The striatin family: A new signaling platform in dendritic spines. J. Physiol. Paris 99, 146–153. https://doi.org/10.1016/j.jphysparis.2005.12.006 (2006).
Poggeler, S. & Kuck, U. A WD40 repeat protein regulates fungal cell differentiation and can be replaced functionally by the mammalian homologue striatin. Eukaryot. Cell 3, 232–240. https://doi.org/10.1128/ec.3.1.232-240.2004 (2004).
Cole, A. M. et al. Calcitermin, a novel antimicrobial peptide isolated from human airway secretions. FEBS Lett. 504, 5–10. https://doi.org/10.1016/s0014-5793(01)02731-4 (2001).
Yano, J., Noverr, M. C. & Fidel, P. L. Jr. Cytokines in the host response to Candida vaginitis: Identifying a role for non-classical immune mediators, S100 alarmins. Cytokine 58, 118–128. https://doi.org/10.1016/j.cyto.2011.11.021 (2012).
Mambula, S. S., Simons, E. R., Hastey, R., Selsted, M. E. & Levitz, S. M. Human neutrophil-mediated nonoxidative antifungal activity against Cryptococcus neoformans. Infect. Immun. 68, 6257–6264. https://doi.org/10.1128/Iai.68.11.6257-6264.2000 (2000).
Wozniak, K. L., Hole, C. R., Yano, J., Fidel, P. L. Jr. & Wormley, F. L. Jr. Characterization of IL-22 and antimicrobial peptide production in mice protected against pulmonary Cryptococcus neoformans infection. Microbiology 160, 1440–1452. https://doi.org/10.1099/mic.0.073445-0 (2014).
Donato, R. et al. Functions of S100 proteins. Curr. Mol. Med. 13, 24–57 (2013).
Deloulme, J. C. et al. S100A6 and S100A11 are specific targets of the calcium- and zinc-binding S100B protein in vivo. J. Biol. Chem. 275, 35302–35310. https://doi.org/10.1074/jbc.M003943200 (2000).
Chiasson, D., Ekengren, S. K., Martin, G. B., Dobney, S. L. & Snedden, W. A. Calmodulin-like proteins from Arabidopsis and tomato are involved in host defense against Pseudomonas syringae pv. tomato. Plant Mol Biol 58, 887–897. https://doi.org/10.1007/s11103-005-8395-x (2005).
Fregno, I. et al. ER-to-lysosome-associated degradation of proteasome-resistant ATZ polymers occurs via receptor-mediated vesicular transport. Embo J. https://doi.org/10.15252/embj.201899259 (2018).
Wang, B., Han, S., Lien, L. & Chang, L. J. Lentiviral calnexin-modified dendritic cells promote expansion of high-avidity effector T cells with central memory phenotype. Immunology 128, 43–57. https://doi.org/10.1111/j.1365-2567.2009.03067.x (2009).
Chaturvedi, A. K., Weintraub, S. T., Lopez-Ribot, J. L. & Wormley, F. L. Jr. Identification and characterization of Cryptococcus neoformans protein fractions that induce protective immune responses. Proteomics 13, 3429–3441. https://doi.org/10.1002/pmic.201300213 (2013).
Acknowledgements
This work was supported in part by Oklahoma State University start-up funds (KLW), the National Institutes of Health (NIH) National Institute of General Medical Sciences (GMS) pilot award under Award Number P20GM103648 (KLW) and NIH GMS under Award Number P20GM134973 (KLW), and NIH National Institute of Allergy and Infectious Diseases Award Number AI123837 (JS).
Author information
Authors and Affiliations
Contributions
B.N.N., K.L.W. and J.S. conceived the experiments. B.N.N. and K.L.W. wrote the main manuscript text, B.N.N. and K.L.W. prepared all figures, B.N.N., S.B., S.P., B.C., E.M., and K.L.W. conducted the experiments. B.N.N., S.B., S.P., B.C., E.M., J.S., and K.L.W. analyzed the data. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Nelson, B.N., Beakley, S.G., Posey, S. et al. Antifungal activity of dendritic cell lysosomal proteins against Cryptococcus neoformans. Sci Rep 11, 13619 (2021). https://doi.org/10.1038/s41598-021-92991-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-92991-6
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.