Abstract
Invasive species contribute to deteriorate the health of ecosystems due to their direct effects on native fauna and the local parasite-host dynamics. We studied the potential impact of the invasive hornet Vespa velutina on the European parasite-host system by comparing the patterns of diversity and abundance of pathogens (i.e. Microsporidia: Nosematidae; Euglenozoa: Trypanosomatidae and Apicomplexa: Lipotrophidae) in European V. velutina specimens with those in the native European hornet Vespa crabro, as well as other common Hymenoptera (genera Vespula, Polistes and Bombus). We show that (i) V. velutina harbours most common hymenopteran enteropathogens as well as several new parasitic taxa. (ii) Parasite diversity in V. velutina is most similar to that of V. crabro. (iii) No unambiguous evidence of pathogen release by V. velutina was detected. This evidence together with the extraordinary population densities that V. velutina reaches in Europe (around of 100,000 individuals per km2 per year), mean that this invasive species could severely alter the native pathogen-host dynamics either by actively contributing to the dispersal of the parasites and/or by directly interacting with them, which could have unexpected long-term harmful consequences on the native entomofauna.
Similar content being viewed by others
Introduction
Insect diversity and abundance are declining worldwide1,2. This phenomenon has been associated with parallel effects on wild and managed pollinators, and interpreted as evidence for the deteriorating health of ecosystems and an ongoing major extinction event3,4,5,6. Pollinator loss is known to have serious consequences on a wide variety of species that rely on them either to feed or to reproduce4,7,8, thus threatening biodiversity, ecosystem services and crop-production9,10. Anthropogenic drivers of insect loss include climate change, habitat loss, agricultural intensification, exposure to pesticides and pathogens, and the introduction of invasive alien species6,11,12,13,14.
Over two-thousand invasive alien species have stablished in the terrestrial European Union territory, sixty-six of which have been labelled as of Union concern owing to their potential harm to ecosystems (https://ec.europa.eu/environment/nature/invasivealien/). One such species is Vespa velutina Lepeletier, 1836 (Hymenoptera: Vespidae), a hornet of East Asian origin which was first detected in France in 2004 and since then it has spread to all neighbouring countries15,16. V. velutina can reach very high population densities17,18 and is a serious threat to the ecosystem16,19. Its most obvious impact derives from its intense predatory activity: in Europe Apis mellifera (Hymenoptera: Apidae) represents 30–60% of its catches and it has been estimated that hornet hunting pressure can drive the collapse of around 30% of the colonies20,21,22 and these figures can be as high as 80% for small apiaries (X. Maside unpublished). Their protein diet is completed with other important pollinators, mainly flies, social wasps as well as other arthropods20,23. In addition, the sudden irruption of a large hornet population in Europe might also interfere with the native host–pathogen dynamics, further threatening the pollinator system. There is mounting evidence for between-species transmission of protozoan pathogens across a wide variety of pollinator species24,25,26, likely mediated through shared fomites27,28. Epidemiological and population genetics data suggest that Nosema ceranae (Fungi; Microsporidia; Nosematidae), first described in the Asian honey bee Apis cerana, has spread across the A. mellifera worldwide population25,29,30,31, from where it might have jumped to bumblebees24. Also, Crithidia bombi (Euglenozoa: Kinetoplastida: Trypanosomatidae) has been horizontally transferred from commercial to wild bumblebees32,33. This and other Trypanosomatidae (i.e. Crithidia mellificae and Lotmaria passim) are increasingly often found in many pollinators34,35,36. These spillover processes have been related to severe losses in the target host populations and are considered a major threat to ecosystem functioning24,37,38. Apart from hunting other insects, the foraging behaviour of V. velutina also involves frequent visiting of flowers and other parts of the plants in search of nectar and sap as a source of carbohydrates. Thus, the hornets are frequently exposed to most native pollinator pathogens, some of which might benefit from the hornet’s active contribution to their spread during the hornets’ visits to apiaries and plants, and from the presence of a novel and highly abundant host. These factors could prompt significant variations of the population sizes of the pollinators’ pathogens in Europe39,40, but also of their pathogenic properties, such as virulence and contagiousness41. Furthermore, the possibility that the founder V. velutina individuals that initiated the invasion process might have carried over alien pathogens from its original distribution range in SE Asia, cannot be discarded.
To evaluate the extent to which V. velutina interacts with the European native pathogens, a detailed knowledge of the parasites hosted by the invading population and the native entomofauna are needed. But so far, to the best of our knowledge, only three reports on V. velutina parasites have been published, and refer to isolated cases of infection by fungi (Beauveria and Metarhizium), a mermithid nematode (Pheromermis sp.) and a parasitoid wasp (Conops vesicularis) in France42,43,44. Also, several viruses commonly found in other hymenoptera, including the Israeli acute paralysis virus, the aphid lethal paralysis virus, the black queen cell virus and the Sacbrood virus have been detected in V. velutina45,46,47,48. Here, we hypothesize that in Europe V. velutina occupies an ecological niche that largely overlaps with that of native hornets (V. crabro) and wasps, such that they interact with a similar profile of parasite species. To falsify this hypothesis, we characterized and compared the patterns of diversity and prevalence of common pollinator enteroparasite species of the groups Nosematidae, Trypanosomatidae and Lipotrophidae, in a collection of samples from Galiza (NW- Iberian Peninsula) representing an European population of V. velutina, V. crabro, wasps (gen. Vespula and Polistes) and bumblebees (gen. Bombus).
Results
Nosematidae
Optical microscopy analysis revealed the presence of spores of Nosema-like parasites in the guts of specimens from 19 samples: seven V. velutina, four Bombus spp., three Polistes spp., three Vespula spp. and two V. crabro (Supplementary Figure S1; Table 1). PCR amplification of the SSU-rRNA locus yielded products of the expected size in 24 samples: 12 of them were new positives, but it failed to amplify from seven samples where spores had been detected by microscopy (Supplementary Table S1). This indicates that some samples might harbour Nosema-like organisms that were not amplified by the primers used in this study. Only PCR-positive samples were considered in what follows. The most common species was detected in twelve samples (nine of V. velutina and three of V. crabro) and it was nearly identical to other Nosema thomsoni sequences available in GenBank, albeit a single nucleotide substitution (Table 1 and Supplementary Table S1). The sequence reads of two of these V. velutina samples (labelled 70 and 163; Supplementary Table S1) displayed double peaks precisely at the six nucleotides of divergence between N. thomsoni and Nosema ceranae. Cloning of these amplicons confirmed the presence of haplotypes of these two species in both samples. The three positive bumblebee samples produced haplotypes that were nearly identical to Nosema bombi. Finally, eight samples—four V. velutina, two Vespula spp, one Polistes spp. and one V. crabro—produced a distinct haplotype that displayed 7% nucleotide divergence with N. bombi and 13% with Nosema apis reference sequences.
A phylogenetic reconstruction of the evolutionary relationships of the SSU-rRNA sequences revealed that most Microsporidia from wasps and bumblebees clustered into two main groups (Fig. 1): the largest one included haplotypes closely related to Nosema thomsoni (11), Nosema ceranae (3) and Nosema bombi (3). The second group, which branched off earlier in the tree, included highly differentiated Nosema-like sequences from V. velutina, V. crabro, Vespula spp. and Polistes spp. samples, as well as two other sequences that were retrieved from GenBank, one isolated from a bumblebee in China and the other from the European corn borer (Ostrinia nubilalis¸ Lepidoptera: Crambidae) in France49,50. The low diversity, high differentiation and strong bootstrap support for this branch suggests that this clade might correspond to a yet undescribed new microsporidian species present in a wide variety of hymenopteran hosts.
Phylogenetic relationships of the SSU-rRNA sequences from Nosematidae. Sequence names indicate the sample name and are color-coded by host groups: V. velutina (grey), V. crabro (yellow), Bombus spp. (orange), Vespula spp. (blue), Polistes spp. (green). Some sequences that were slightly shorter or with double peaks were excluded. The evolutionary history was inferred using the NJ method. Bootstrap values higher than 70% are shown next to branches. The evolutionary distances were computed using the Tamura 3-parameter method and are in the units of the number of base substitutions per site.
Trypanosomatidae
PCR amplification of rpb1 and topoII allowed the identification of 56 Trypanosomatidae-positive samples (Table 1 and Supplementary Table S1). The two loci were amplified in twenty-nine of them and just one locus in the remainder: rpb1 in 13 and topoII in 14. In 20 of the former samples the two loci allowed the detection of the same parasite species (Supplementary Table S1), whereas there were discrepancies in the other nine either because one locus allowed the detection of more species than the other (N = 5) or because they identified different species (N = 4). These differences could not be attributed to a systematic bias of the detection technique—i.e. a differential sensibility of the PCR-primer pairs designed for each locus across parasite species, as both loci allowed the detection of a similar number of positives samples (N = 42 and 43, for rpb1 and topoII, respectively) and there were no statistically significant differences between the frequency spectra of the parasites identified by either of them (X2 = 7.38; d.f. = 4; P = 0.11; in a Chi-squared test of homogeneity).
Most positive samples presented topoII and rpb1 haplotypes that were nearly identical to Trypanosomatidae species previously found in honeybees and bumblebees (Fig. 2). The average pairwise neutral divergence between those closely related to Crithidia acanthocephali, C. bombi, C. mellificae or to L. passim was 1.1% ± 0.72 and 0.5% ± 0.24 for topoII and rpb1, respectively (± SE; Supplementary Table S2).
Phylogenetic relationships of the topoII (a) and rpb1 (b) sequences from Trypanosomatidae. Sequence names include the sample name and the clone number when applicable (C-followed by clone numbers separated by dashes if more than one). Host groups are color-coded: V. velutina (grey), V. crabro (yellow), Bombus spp. (orange), Vespula spp. (blue), Polistes spp. (green). Red arrows indicate new clades. Some sequences that were slightly shorter or with double peaks were excluded. The evolutionary history was inferred using the NJ method. Bootstrap values higher than 70% are shown next to branches. The evolutionary distances were computed using the modified Nei-Gojobori method (assumed transition/transversion bias = 2) and are in the units of the number of synonymous differences per synonymous site.
Contrastingly, other topoII and rpb1 haplotypes displayed high divergence from any previously reported sequences. At topoII these distinct haplotypes formed three clusters: T1, which included four haplotypes from samples 8 and 14; T2, one from sample 82; and T3, one from 41B (Fig. 2a). Similarly, at rpb1 there were up to four novel clusters (R1-R4). Three of them were detected in the same samples as those of topoII: R1, three haplotypes from sample 8, R3 in samples 41B, 42 and 82, and R4, which included sequences from samples 82 and 42 closer to C. mellificae (Fig. 2b). Most of these novel clades displayed levels of genetic differentiation of similar magnitude to those observed between the regular species. For instance, the closest relative of any of the new topoII lineages was Blastocrithidia culicis, which presented a level of synonymous divergence from clade T3 of 34.7% ± 4.84 (Supplementary Tables S3and S4), a value slightly larger than that observed between B. culicis, L. passim and C. acanthocephali (average KS = 33.7% ± 0.04). This is also true for most rpb1 new clades, except R4 which displayed lower divergence from C. mellificae (KS = 7.0 ± 2.46 Supplementary Table S4). At any rate, this value was nearly ten times larger than the average diversity observed within species (see above).
These results are consistent with the presence of new Trypanosomatidae taxa in the samples. However, to make further inferences, such as to estimate the total number of new taxa by combining the information from the two loci, is not straight forward because: (i) the evolutionary relationships among taxa might vary significantly across loci due to both deterministic (differential patterns of natural selection) and stochastic effects (i.e. mutation and drift). (ii) The resolution of the PCR technique is expected to vary across samples and loci due to differences in template DNA quality across samples and in the ability of primers to bind new sequences. (iii) There is no minimum threshold value of neutral divergence to unambiguously determine whether variation occurs between or within species.
C. bombi was the most common Trypanosomatidae in the sample (22.2%). It was the most prevalent parasite in all hosts but in V. crabro, and it reached up to 48.0% prevalence in bumblebees (Table 1). C. mellificae and L. passim were also detected in all hosts, although at lower prevalence (8.6% and 10.4%, respectively). C. acanthocephali was found in four V. velutina and one Vespula spp. samples (5.1% and 4.8%, respectively).
Forty of the positive samples (71.4%) harboured a single Trypanosomatidae species, thirteen (23.2%) had two and three had three (5.4%) (Supplementary Table S1). Co-occurrence of parasites usually involved the three most prevalent species (C. bombi, C. mellificae and L. passim). A significant excess of mixed samples relative to random expectations was detected between these three species (P < 0.05 in pairwise comparisons assuming a hypergeometric distribution; Supplementary Table S5).
Lipotrophidae
Apicystis parasites were detected in 43 samples (25.7%; Table 1 and Supplementary Table S1). Sequences were identical to other Apicystis bombi haplotypes previously found in honeybees (e.g. AB738024), except for two sequences that displayed a single nucleotide substitution. A. bombi prevalence was largest in Polistes spp. (45.5%) and Vespula spp. (42.9%) and lowest in V. velutina (17.7%).
Overall between-hosts comparisons
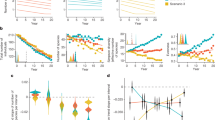
Eighty-eight out of the 167 samples tested positive for any of the parasites (52.7%; from data in Supplementary Table S1). The Vespula spp. and Polistes spp. groups presented the highest fractions of positive samples (81.0% and 72.7%, respectively; Table 1), followed by Bombus spp. (68.0%). V. velutina and V. crabro presented lower frequencies of positive samples (42.0%; Table S1). The two hornet species also presented similar prevalence of parasites of the three groups (X2 = 1.22; d.f. = 2; P = 0.54, in a Chi-square test of homogeneity; data from Table S1). But these figures only represent overall parasite prevalence, and do not consider species variation. For a more comprehensive comparison we used the Shannon diversity and equitability indexes across the five hosts (Fig. 3). V. velutina displayed the highest parameter values (H = 2.0 and EH = 0.91) although they were only slightly larger than those for the other three Vespidae hosts. Indeed, there were no significant differences between H estimates for V. velutina and any of the other Vespidae groups, (P > 0.05 in pairwise Student’s t tests). Contrastingly, the bumblebees produced the lowest diversity scores, as expected given that they hosted fewer parasite species than any other group (Supplementary Table S1) and that just two of them (C. bombi and A. bombi) accounted for 75.0% of the positive samples. The differences between H for Bombus spp. and the Vespidae (pooled data) were statistically significant (t = 5.47; d.f. = 31; P < 0.001 in a Student’s t-test).
The number of parasites species per sample varied significantly across hosts (Table 2). The two hornet species presented very similar distributions (X2 = 0.05; d.f. = 2; P = 0.97, in a chi-square test of homogeneity; samples with 2, 3 or 4 parasites were grouped to avoid zero values), which differed significantly from those of the other three hosts (X2 = 19.5; d.f. = 2; P < 0.001; data from Table 2). Nearly 60% of the hornet samples did not harbour any parasite, about 20% had just one species and a similar fraction had two or more parasites. In contrast, most wasps and bumblebees’ samples were positive (an average of 73.7% across the three hosts), most of them with one parasite (47.4%) and 26.3% with two or more. No significant associations between the three main parasite groups were detected (P > 0.05 in all pairwise comparisons, Supplementary Table S5).
Discussion
Microscopy and molecular methods were used to determine the prevalence and diversity of parasites of the groups Nosematidae, Trypanosomatidae and Lipotrophidae in a collection of 167 samples representing the populations of V. velutina (79), V. crabro (31), Vespula spp. (21), Polistes spp. (11), and Bombus spp. (25), in Galiza (NW-Iberian Peninsula). Most pathogens commonly found in native European Hymenoptera were detected in the samples. Trypanosomatidae were the most prevalent group of parasites. C. bombi was present in up to 48.0% of the bumblebees, a figure slightly larger than previous reports on bumblebees from Scotland51, England27,33, Argentina52 and the Iberian Peninsula53. This parasite was also detected in all Vespidae hosts along with C. mellificae and L. passim; the two latter species at an average frequencies below 10%, in agreement with previous records from honeybees from Galiza54 or North America55. There was a significant excess of co-occurrence of Trypanosomatidae species in the whole sample. This is consistent with data from honeybees35,56. No other associations were detected.
Apicystis bombi, a Neogregarine, was the most prevalent parasite of the series (25.7%), reaching frequencies of 40% and greater in Vespula spp. and Polistes spp. These figures are even greater than those previously reported for this parasite in bumblebees, which are considered as their primary host, elsewhere in Europe and in South America27,33,51,52. In the Iberian Peninsula A. bombi had been exclusively detected in bumblebees from southern Spain, where Bombus terrestris are extensively used in greenhouse tomato production53, and its dispersal has been linked to the use of managed B. terrestris, as in other parts of the world33,52. The present data suggest that this pathogen is widespread across the whole of the Iberian Peninsula and can be found in a wide variety of hosts, including bumblebees, wasps and hornets, as suggested by recent reports of A. bombi in honeybees and in wild bumblebees form various locations across Iberia36,56.
Nosematidae were the less prevalent of the three parasite groups (14.4%). N. apis was not detected, and N. ceranae and N. bombi were restricted to five samples: two hornet and three bumblebee samples, respectively. This is at odds with recent reports from honeybees and bumblebees, where Nosematids (particularly the latter two species) are usually found at high frequencies in Galiza54 and elsewhere51,57,58,59,60,61, and with previous evidence that N. bombi was circumscribed to bumblebees from the southern Iberian Peninsula53. On the other hand, N. thomsoni, a species that was first described in the moth Choristoneura conflictana (Lepidoptera: Trotricidae) and has been found only sporadically in Apidae, including honeybees, bumblebees and solitary bees49,56,62,63, was the most prevalent Nosematidae, although it was detected only in both hornet species.
In addition to the species commonly found in honeybees and bumblebees, the molecular analysis allowed the detection of several new taxa: one microsporidium found in the four Vespidae hosts and five Trypanosomatidae (three in V. velutina and two in Vespula spp. and Polistes spp). Nosema sp. had been previously found in a honeybee in China and in the European corn borer (Ostrinia nubilalis) in France49,50 and one of the Trypanosomatidae in honeybees in the Iberian Peninsula56. All these reports postdate the arrival of V. velutina in Europe. It could thus be speculated that some of these organisms might have been brought into Europe by the invading V. velutina. But the scarcity of available data on pathogen diversity in hymenopterans other than bees and bumblebees from Europe and SE-Asia64 hampers the possibility of reliably testing this hypothesis. Provided that the V. velutina population in Europe probably derived from a single female65, the possibility that it brought along novel parasitic taxa into Europe and that they survived over sixteen generations seems somewhat remote. This goes in line with the lack of evidence for pathogen release by invasive populations of V. vulgaris in South America and New Zealand66,67. At any rate, the finding of these new parasite taxa highlights that pollinators other than the commonly studied honeybees and bumblebees might harbour a plethora of parasites and that their spread across other pollinator species might have undesirable health consequences for the entire ecosystem.
Three main patterns of parasite presence across the five hosts groups were observed: (i) hornets and wasps presented similar levels of parasite species diversity, somewhat greater than the bumblebees, whose parasite repertoire was dominated by just two species (C. bombi and A. bombi; Table 1). (ii) Among the Vespidae, the parasite profile of V. velutina was most similar to that of V. crabro; both displayed a varied representation of the Nosematidae and Trypanosomatidae, whereas the presence of Nosematidae in Vespula spp. and Polistes spp. was only testimonial and limited to three samples with rare new taxa (Table 1). Also, (iii) most hornet samples were parasite-free, whereas wasps and bumblebee samples often presented one or more parasite species.
The above discordances contrast with the facts that all of them visit flowers in search of food24,27,28,68 and that the Vespidae diet includes honeybees and bumblebees, amongst other insects20,69. However, subtle variation in their feeding habits might help explain the differences. For instance, (i) Vespidae and Apidae might not have the same flower preferences, such that the spectra of parasite species that they encounter might be different. (ii) The Vespidae feed on nectar, but do not seem to actively harvest pollen while flower visiting, thus they may not frequently ingest the parasites in the flower surface brought about by other pollinators69,70. (iii) Wasps often discard the abdomens of their prey, which might reduce their exposure to gut pathogens such as the ones studied here. And (iv) hornets, besides feeding on carrions, use only the thoraxes of their preys and discard all other body parts20 (see also: http://www.vespa-crabro.com/) whereas Vespula spp. rely more often on dead preys and sporadically include the heads and the abdomens in their diets69. Further knowledge of their feeding habits should help ascertain the extent to which they can explain the different patterns of parasite distribution across hosts.
The arrival of V. velutina into Europe together with its enormous reproductive success mean that this invasive hornet is a serious threat to the health of the native insect pollinator community. Theoretical approaches identify vast areas of Europe where climatic conditions are now suitable for V. velutina reproduction. They comprise most of the Mediterranean and Atlantic coastal areas, including the British Isles71, but global warming could facilitate its expansion throughout most continental Europe72. In addition, V. velutina can reach very high population densities: in Galiza (NW-Iberian Peninsula) the public services removed nearly 70,000 nests in three years (2018 to 2020; data released by Xunta de Galiza). Most of them were in the areas of highest incidence of V. velutina, which comprise about 1/5 of the Galician territory (approx. 6000 km2; as estimated from Rodríguez-Lado73). This means that about four nests were destroyed per km2 in these areas. Provided that between 60–70% of the extant nests remain undetected74, nest density in Galiza can be approximated as 14–17 nests per km2, which fits well with observations of up to 12 nests per km2 in Andernos-les-Bains (France)18. Considering that each nest on average produce 6150 hornets per reproductive season (June–October)17, highly infested areas harbour on average 100,000 V. velutina specimens per km2 per year. The presence of this huge population is likely to have at least two main effects on the native entomofauna. One is directly derived from the intense predatory activity they exert over a wide variety of insects, including common wasps, paper wasps, bumblebees, honeybees, dipterans (syrphids and brachycera), hymenopterans (halictids) among others20,23,75. Indeed, this pressure is so strong that they can destroy nests of Vespula germanica (X. Maside unpublished) and drive to extinction up to 30% of the honeybee colonies, on average21. These effects are inversely associated with the size of the apiaries due to a difussion effect of the predatory activity on individual colonies. Indeed, smaller-scale professional beekeepers have reported the death of all the colonies in small apiaries (up to 50 colonies) due to V. velutina predation76.
The second effect has been so far unexplored and refers to the indirect impact of this large novel population on the local host-parasite dynamics. Our data revealed that the invading V. velutina harbours most if not all parasites commonly found in native European pollinators. Therefore, its presence is causing a significant change in the host community whose effect will be determined by the specific relationships between V. velutina and the parasites, and how its presence modifies the abundance of parasite spreaders, the rate of parasite-host encounters, and the probability of disease transmission77. For instance, the presence of a new host might have a dilution effect, particularly for those parasites that find that V. velutina is not a competent host, as they will experience a reduction in their chance to find a suitable one77,78,79. Alternatively, it might produce an amplification effect on those pathogens that successfully reproduce in them80. Furthermore, a surplus of a competent hosts might prompt selection for virulence41,81, threatening the overall health of native pollinators. V. velutina might further contribute to pathogen evolution by favouring the encounter and sexual reproduction between strains of pathogens usually found in different hosts on which they prey. The potential role of recombination and natural selection on pathogen evolution is well illustrated by the recent spread of a novel hypervirulent strain of the fungus Batrachochytrium dendrobatidis that threatens amphibian diversity worldwide82.
Understanding the impact of invasive species in biodiversity is critical to properly protect native ecosystems. Here we showed that specimens of the invasive V. velutina in Europe harbour a wide variety of parasites commonly found in local pollinators, but have not investigated the extent to which this interaction alters local host–pathogen dynamics. Further studies are needed to quantify and comprehend the potential consequences of this disturbance and to help put forward strategies to minimize its impact and to protect the already badly threatened native insect populations.
Methods
Sampling
Between August and November 2016 a total of 564 specimens of V velutina, V. crabro and other Hymenoptera belonging to genera Bombus, Vespula and Polistes were collected throughout the distribution area of the invasive species in Galiza (NW Iberian Peninsula) (Fig. 4). Specimens were captured using homemade traps or directly from nests (mainly V. velutina, V. crabro). Traps were filled with 80 ml of liquid bait: 95% alcohol free dark beer (Super Bock; Super Bock Group SGPS, S.A.), 4% white wine (various brands were used) and 1% grenadine syrup (Rives). Traps were fitted with a fine mesh to avoid insects from drowning in the bait and producing cross-contamination (Supplementary Figure S2). Captured insects were collected daily. Whole nests were removed by the fire brigade of Santiago de Compostela at night, when most of the hornets were inside the nest. This means that the specimens collected are a random sample of the population of the nest.
Geographic distribution of the samples. Circles identify the council of origin of the samples. Circles are drawn to scale to represent the number of samples and are color coded: V. velutina (grey), V. crabro (yellow), Vespula spp. (blue), Polistes spp. (green), Bombus spp. (orange). Map: South-West Europe and Galiza (enlarged).
Sample processing and DNA extraction
All captured specimens were classified into five groups: V. velutina, V. crabro, Vespula spp. (including mostly V. germanica and/or V. vulgaris), Polistes spp. (P. austroccidentalis, P. dominula, P. nimpha) and Bombus spp. (B. terrestris, B. pascuorum and other species) and preserved in 80% EtOH at 4 °C. Before dissection, each specimen was thoroughly washed in fresh 80% EtOH and rinsed three times in sterile distilled water. Abdomens were dissected and abdominal contents (including midgut, Malpighian tubules and fat body) of each specimen were extracted and homogenised separately in a 1.5 μL tube using disposable plastic pestles (VWR). Dissection tools and materials were cleaned with fresh 5% bleach for 20 min, rinsed two times in sterile distilled water, once in 80% EtOH and air-dried prior to manipulation of each specimen. A fraction of the homogenates of one to five specimens of the same species from each sampling location was combined in a pooled sample (depending on availability). In total there were 167 samples representing the populations of V. velutina (79), V. crabro (31), Vespula spp. (21), Polistes spp. (11), and Bombus spp. (25). The number of individuals pooled in each sample was found to be unrelated to the number of parasite species or to the fraction of positive samples (Supplementary Figure S3). This probably reflects that most pooled samples were made up with individuals collected from the same nest, so there was little variation between specimens. Whole genomic DNA was extracted using the phenol–chloroform method: 100 μL of homogenate were digested overnight at 56 °C in 300 μL of lysis buffer (50 mM Tris–HCl pH 8, 100 mM EDTA, 100 mM NaCl, 1% SDS) with 5 μL of proteinase K (~ 20 mg/mL, Thermo Scientific) and DNA was extracted with phenol:chloroform:isoamyl alcohol (Sigma-Aldrich), precipitated with isopropanol, washed in 80% EtOH and resuspended in 100 μL of PCR grade water.
Microscopic detection of Nosematids
Air-dried smears of homogenates from the pooled samples were stained with Calcofluor White (CFW) (0.1% w/v CFW stock solution diluted 3:7 in a 10% w/v KOH, 10% v/v Glycerine solution). CFW binds to the cell wall of Nosema spores and after being excited in UV emits in blue. Microscopic slides were observed at 400 × on a Nikon Eclipse C1000M fluorescent microscope using a UV 2-A fluorescent filter (Nikon; excitation filter: 330–380 nm, dichroic mirror: 400 nm, barrier filter: 420 nm).
Molecular detection
The presence of parasites was determined by PCR amplification of selected marker loci and Sanger sequencing of PCR amplicons. The small subunit rRNA (SSU rRNA) loci were used for the detection of Nosematidae and Lipotrophidae species, respectively, and the DNA-directed RNA polymerase II subunit (rpb1) and Type II topoisomerase II (topo II) for Trypanosomatidae. Primer details are given in Table S6.
PCR amplifications were performed using Phusion High-Fidelity PCR Kit (Thermo Scientific) in a total volume of 15 μL under manufacturer’s conditions. DNA templates were 1:20 dilutions (in H2O) of the DNA pools. PCR cycling was as follows: initial denaturation at 98 °C for 30′; 35 cycles of melting at 98 °C for 10″, annealing for 30″ and extension at 72 °C for 30″; and final extension at 72 °C for 5′. Annealing temperatures are shown in Table S6.
Five microliters of PCR products were tested by electrophoresis in 2% agarose gels. When PCR yields were too low for direct sequencing, additional reamplifications for 10–15 cycles were performed under the same conditions, using 1 μL of a 1/20 dilution of the initial PCR product as templates.
Sequencing and cloning of PCR products
PCR products were purified using NZYGelpure kit (NZYTech, Lda.) following manufacturer’s instructions and sent to GATC-Biotech (Eurofins Scientific) for Sanger sequencing of both strands. Sequences were corrected for accurate base calling by comparison of both strands using CodonCode Aligner (CodonCode Corporation). Sequence alignments were performed with ClustalW83 and manually edited with BioEdit84. Sequences have been deposited in GenBank under accession numbers: MW288771–MW288823, MW284939–MW284941 and MW284942–MW284950.
The presence of double peaks throughout some of the sequence reads were taken as evidence for genetic variation in the sample. In these cases, the PCR products were cloned using CloneJET PCR Cloning Kit (ThermoFisher Scientific Inc.) and DH5α Escherichia coli competent cells (Invitrogen) under manufacturer’s conditions using half of the indicated volumes. Plasmids were isolated using the NZYMiniprep Kit (NZYtech) and 5 clones per locus were sent to GATC-Biotech for Sanger sequencing.
Species identification and phylogenetic analysis
Identification of parasite species was performed by comparing the resulting haplotypes with relevant reference sequences from each marker locus available in public databases (GenBank and ENA). For Trypanosomatidae the data from the two loci (rpb1 and topoII) were taken independently: parasites detected by one marker locus were taken as true positives. For the samples with more than one parasite (mixed samples), only the species whose presence was further confirmed by sequencing the clones were considered. In a few cases the haplotypes were a chimera of fragments from more than one species. Given that the chance that they corresponded to true between-species recombinants is negligible, they were taken as experimental artefacts during PCR amplification and were excluded from the analyses.
Phylogenetic analyses were conducted using MEGA version 685. The evolutionary relationships were inferred using the Neighbor-Joining (NJ) method, and evolutionary distances were computed at all sites using the Tamura 3-parameter method for SSU-rRNA sequences and at synonymous sites using the Nei-Gojobori method for rpb1 and topoII. Bootstrap values were calculated using 500 replicates.
Parasite diversity
To assess how parasite diversity varied across hosts we used the Shannon’s diversity index (H), which provides a quantitative measure of the species diversity by combining abundance and evenness of the species present in a dataset, and the Shannon’s equitability index (EH), which is the ratio of the H relative to its maximum value and measures the evenness in the distribution of species (takes values between 0 and 1). 95% CIs were estimated assuming that H follows a normal distribution86,
where S and pi are the number of species and the proportion of individuals of the ith species in each dataset, respectively.
References
Habel, J. C. et al. Butterfly community shifts over two centuries. Conserv. Biol. 30, 754–762 (2016).
Hallmann, C. A. et al. More than 75 percent decline over 27 years in total flying insect biomass in protected areas. PLoS One 12, e0185809 (2017).
Dirzo, R. et al. Defaunation in the Anthropocene. Science 345, 401–406 (2014).
Inger, R. et al. Common European birds are declining rapidly while less abundant species’ numbers are rising. Ecol. Lett. 18, 28–36 (2015).
Thomas, J. A. et al. Comparative losses of British butterflies, birds, and plants and the global extinction crisis. Science 303, 1879–1881 (2004).
Vanbergen, A. J. & The Insect Pollinators Initiative. Threats to an ecosystem service: Pressures on pollinators. Front. Ecol. Environ. 11, 251–259 (2013).
Biesmeijer, J. C. et al. Parallel declines in pollinators and insect-pollinated plants in Britain and the Netherlands. Science 313, 351–354 (2006).
Stanton, R. L., Morrissey, C. A. & Clark, R. G. Analysis of trends and agricultural drivers of farmland bird declines in North America: A review. Agric. Ecosyst. Environ. 254, 244–254 (2018).
Gallai, N., Salles, J.-M., Settele, J. & Vaissière, B. E. Economic valuation of the vulnerability of world agriculture confronted with pollinator decline. Ecol. Econ. 68, 810–821 (2009).
Potts, S. G. et al. Safeguarding pollinators and their values to human well-being. Nature 540, 220 (2016).
Gonzalez-Varo, J. P. et al. Combined effects of global change pressures on animal-mediated pollination. Trends Ecol. Evol. 28, 524–530 (2013).
Banks, N. C., Paini, D. R., Bayliss, K. L. & Hodda, M. The role of global trade and transport network topology in the human-mediated dispersal of alien species. Ecol. Lett. 18, 188–199 (2015).
Sánchez-Bayo, F. & Wyckhuys, K. A. G. Worldwide decline of the entomofauna: A review of its drivers. Biol. Conserv. 232, 8–27 (2019).
Essl, F. et al. Drivers of future alien species impacts: An expert-based assessment. Glob. Change Biol. 26, 4880–4893 (2020).
Laurino, D., Lioy, S., Carisio, L., Manino, A. & Porporato, M. Vespa velutina: An alien driver of honey bee colony losses. Diversity 12, 5 (2020).
Monceau, K., Bonnard, O. & Thiéry, D. Vespa velutina: A new invasive predator of honeybees in Europe. J. Pest Sci. 87, 1–16 (2013).
Rome, Q. et al. Caste differentiation and seasonal changes in Vespa velutina (Hym.: Vespidae) colonies in its introduced range. J. Appl. Entomol. 139, 771–782 (2015).
Monceau, K. & Thiery, D. Vespa velutina nest distribution at a local scale: An 8-year survey of the invasive honeybee predator. Insect Sci. 24, 663–674 (2017).
Snyder, W. E. & Evans, E. W. Ecological effects of invasive arthropod generalist predators. Annu. Rev. Ecol. Evol. Syst. 37, 95–122 (2006).
Villemant, C. et al. Bilan des travaux (MNHN et IRBI) sur l’invasion en France de Vespa velutina, le frelon asiatique prédateur d’abeilles. In Proceedings of the Journée Scientifique Apicole (eds Barbançon, J.-M. & L’Hostis, M.) 3–12 (ONIRIS-FNOSAD, 2011).
Monceau, K. & Thiery, D. Vespa velutina: Current situation and perspectives (Atti Accademia Nazionale Italiana di Entomologia, 2017).
Requier, F. et al. Predation of the invasive Asian hornet affects foraging activity and survival probability of honey bees in Western Europe. J. Pest Sci. 92, 567–578 (2019).
Rome, Q. et al. Not just honeybees: Predatory habits of Vespa velutina (Hymenoptera: Vespidae) in France. Ann. Soc. Entomol. Fr. (N.S.). 57, 1–11. https://doi.org/10.1080/00379271.2020.1867005 (2021).
Furst, M. A., McMahon, D. P., Osborne, J. L., Paxton, R. J. & Brown, M. J. F. Disease associations between honeybees and bumblebees as a threat to wild pollinators. Nature 506, 364–366 (2014).
Maside, X. et al. Population genetics of Nosema apis and Nosema ceranae: One host (Apis mellifera) and two different histories. PLoS One 10, e0145609 (2015).
Plischuk, S. et al. South American native bumblebees (Hymenoptera: Apidae) infected by Nosema ceranae (Microsporidia), an emerging pathogen of honeybees (Apis mellifera). Environ. Microbiol. Rep. 1, 131–135 (2009).
Graystock, P., Goulson, D. & Hughes, W. O. Parasites in bloom: Flowers aid dispersal and transmission of pollinator parasites within and between bee species. Proc. Biol. Sci. 282, 20151371 (2015).
Graystock, P. et al. Dominant bee species and floral abundance drive parasite temporal dynamics in plant-pollinator communities. Nat. Ecol. Evol. 4, 1358–1367 (2020).
Gómez-Moracho, T. et al. Recent worldwide expansion of Nosema ceranae (Microsporidia) in Apis mellifera populations inferred from multilocus patterns of genetic variation. Infect. Genet. Evol. 31, 87–94 (2015).
Higes, M., Martin, R. & Meana, A. Nosema ceranae, a new microsporidian parasite in honeybees in Europe. J. Invertebr. Pathol. 92, 93–95 (2006).
Klee, J. et al. Widespread dispersal of the microsporidian Nosema ceranae, an emergent pathogen of the western honey bee, Apis mellifera. J. Invertebr. Pathol. 96, 1–10 (2007).
Otterstatter, M. C. & Thomson, J. D. Does pathogen spillover from commercially reared bumble bees threaten wild pollinators?. PLoS One 3, e2771 (2008).
Graystock, P., Goulson, D. & Hughes, W. O. The relationship between managed bees and the prevalence of parasites in bumblebees. PeerJ 2, e522 (2014).
Ravoet, J. et al. Differential diagnosis of the honey bee trypanosomatids Crithidia mellificae and Lotmaria passim. J. Invertebr. Pathol. 130, 21–27 (2015).
Bartolomé, C. et al. A new multiplex PCR protocol to detect mixed trypanosomatid infections in species of Apis and Bombus. J. Invertebr. Pathol. 154, 37–41 (2018).
Bartolomé, C. et al. Wide diversity of parasites in Bombus terrestris (Linnaeus, 1758) revealed by a high-throughput sequencing approach. Environ. Microbiol. 23, 478–483 (2020).
Meeus, I., Brown, M. J. F., De Graaf, D. C. & Smagghe, G. Effects of invasive parasites on bumble bee declines. Conserv. Biol. 25, 662–671 (2011).
Goulson, D. & Hughes, W. O. H. Mitigating the anthropogenic spread of bee parasites to protect wild pollinators. Biol. Conserv. 191, 10–19 (2015).
Burdon, J. & Chilvers, G. Host density as a factor in plant disease ecology. Annu. Rev. Phytopathol. 20, 143–166 (1982).
Parker, I. M. et al. Phylogenetic structure and host abundance drive disease pressure in communities. Nature 520, 542 (2015).
Borovkov, K., Day, R. & Rice, T. High host density favors greater virulence: A model of parasite–host dynamics based on multi-type branching processes. J. Math. Biol. 66, 1123–1153 (2013).
Darrouzet, E., Gévar, J. & Dupont, S. A scientific note about a parasitoid that can parasitize the yellow-legged hornet, Vespa velutina nigrithorax, Europe. Apidologie 46, 130–132 (2014).
Villemant, C. et al. Can parasites halt the invader? Mermithid nematodes parasitizing the yellow-legged Asian hornet in France. PeerJ 3, e947 (2015).
Poidatz, J., López Plantey, R. & Thiéry, D. Indigenous strains of Beauveria and Metharizium as potential biological control agents against the invasive hornet Vespa velutina. J. Invertebr. Pathol. 153, 180–185 (2018).
Dalmon, A. et al. Viruses in the Invasive Hornet Vespa velutina. Viruses 11, 1041 (2019).
Marie-Pierre Chauzat et al. First detections of honey bee pathogens in nest of the Asian hornet (Vespa velutina) collected in France. CIHEAM Watch Letter 33 (2015).
Mazzei, M. et al. Detection of replicative Kashmir Bee Virus and Black Queen Cell Virus in Asian hornet Vespa velutina (Lepelieter 1836) in Italy. Sci. Rep. 9, 10091 (2019).
Yañez, O., Zheng, H.-Q., Hu, F. L., Neuman, P. & Dietemann, V. A scientific note on Israeli acute paralysis virus infection of Eastern honeybee Apis cerana and vespine predator Vespa velutina. Apidologie 43, 587–589 (2012).
Li, J. et al. Diversity of Nosema associated with bumblebees (Bombus spp.) from China. Int. J. Parasitol. 42, 49–61 (2012).
Tokarev, Y. S. et al. Redefinition of Nosema pyrausta (Perezia pyraustae Paillot 1927) basing upon ultrastructural and molecular phylogenetic studies. Parasitol. Res. 114, 759–761 (2015).
Goulson, D., Whitehorn, P. & Fowley, M. Influence of urbanisation on the prevalence of protozoan parasites of bumblebees. Ecol. Entomol. 37, 83–89 (2012).
Plischuk, S., Antúnez, K., Haramboure, M., Minardi, G. M. & Lange, C. E. Long-term prevalence of the protists Crithidia bombi and Apicystis bombi and detection of the microsporidium Nosema bombi in invasive bumble bees. Environ. Microbiol. Rep. 9, 169–173 (2017).
Jabal-Uriel, C. et al. Short communication: First data on the prevalence and distribution of pathogens in bumblebees (Bombus terrestris and Bombus pascuorum) from Spain. Span. J. Agric. Res. 15, 3 (2017).
Meana, A. et al. Risk factors associated with honey bee colony loss in apiaries in Galicia, NW Spain. Span. J. Agric. Res. 15, e0501 (2017).
Cavigli, I. et al. Pathogen prevalence and abundance in honey bee colonies involved in almond pollination. Apidologie 47, 251–266 (2016).
Bartolomé, C. et al. Longitudinal analysis on parasite diversity in honeybee colonies: New taxa, high frequency of mixed infections and seasonal patterns of variation. Sci. Rep. 10, 10454 (2020).
Ravoet, J. et al. Comprehensive bee pathogen screening in Belgium reveals Crithidia mellificae as a new contributory factor to winter mortality. PLoS One 8, e72443 (2013).
Arbulo, N. et al. High prevalence and infection levels of Nosema ceranae in bumblebees Bombus atratus and Bombus bellicosus from Uruguay. J. Invertebr. Pathol. 130, 165–168 (2015).
Plischuk, S. et al. Pathogens, parasites, and parasitoids associated with bumble bees (Bombus spp.) from Uruguay. Apidologie 48, 298–310 (2017).
Sinpoo, C., Disayathanoowat, T., Williams, P. H. & Chantawannakul, P. Prevalence of infection by the microsporidian Nosema spp. in native bumblebees (Bombus spp.) in northern Thailand. PLoS One 14, e0213171 (2019).
Michalczyk, M., Bancerz-Kisiel, A. & Sokół, R. Lotmaria passim as third parasite gastrointestinal tract of honey bees living in tree trunk. J. Apic. Sci. 64, 143–151 (2020).
Ravoet, J. et al. Widespread occurrence of honey bee pathogens in solitary bees. J. Invertebr. Pathol. 122, 55–58 (2014).
Schoonvaere, K., Smagghe, G., Francis, F. & de Graaf, D. C. Study of the metatranscriptome of eight social and solitary wild bee species reveals novel viruses and bee parasites. Front. Microbiol. 9, 177 (2018).
Rose, E. A. F., Harris, R. J. & Glare, T. R. Possible pathogens of social wasps (Hymenoptera: Vespidae) and their potential as biological control agents. N. Z. J. Zool. 26, 179–190 (1999).
Arca, M. et al. Reconstructing the invasion and the demographic history of the yellow-legged hornet, Vespa velutina, Europe. Biol. Invasions 17, 2357–2371 (2015).
Torchin, M. E., Lafferty, K. D., Dobson, A. P., McKenzie, V. J. & Kuris, A. M. Introduced species and their missing parasites. Nature 421, 628–630 (2003).
Lester, P. J. et al. No evidence of enemy release in pathogen and microbial communities of common wasps (Vespula vulgaris) in their native and introduced range. PLoS One 10, e0121358 (2015).
Adler, L. S. et al. Disease where you dine: Plant species and floral traits associated with pathogen transmission in bumble bees. Ecology 99, 2535–2545 (2018).
Pusceddu, M., Mura, A., Floris, I. & Satta, A. Feeding strategies and intraspecific competition in German yellowjacket (Vespula germanica). PLoS One 13, e0206301 (2018).
Ueno, T. Flower-visiting by the invasive hornet Vespa velutina nigrithorax (Hymenoptera: Vespidae). Int. J. Chem. Environ. Biol. Sci. 3, 444–448 (2015).
Barbet-Massin, M. et al. Climate change increases the risk of invasion by the Yellow-legged hornet. Biol. Conserv. 157, 4–10 (2013).
Barbet-Massin, M., Salles, J.-M. & Courchamp, F. The economic cost of control of the invasive yellow-legged Asian hornet. NeoBiota 55, 11–25 (2020).
Rodríguez-Lado, L. velutina | Visor. Monitorización da distribución de Vespa velutina. http://webs-gis.cesga.es/velutina/ (2017).
Robinet, C., Suppo, C. & Darrouzet, E. Rapid spread of the invasive yellow-legged hornet in France: The role of human-mediated dispersal and the effects of control measures. J. Appl. Ecol. 54, 205–215 (2017).
Rojas-Nossa, S. V. & Calviño-Cancela, M. The invasive hornet Vespa velutina affects pollination of a wild plant through changes in abundance and behaviour of floral visitors. Biol. Invasions 22, 2609–2618 (2020).
Ferreira-Golpe, M., Garcia-Arias, A. I. & Pérez-Fra, M. M. Costes de la lucha contra la especie invasora Vespa velutina soportados por los apicultores en la provincia de A Coruña. in XII Congreso Iberoamericano de Estudios Rurales. Territorios Globales, Ruralidades Diversas 294–297 (2019).
Keesing, F. et al. Impacts of biodiversity on the emergence and transmission of infectious diseases. Nature 468, 647–652 (2010).
Civitello, D. J. et al. Biodiversity inhibits parasites: Broad evidence for the dilution effect. Proc. Natl. Acad. Sci. 112, 8667–8671 (2015).
Ostfeld, R. & Keesing, F. Biodiversity and disease risk: The case of Lyme disease. Conserv. Biol. 14, 722-728 (2000).
Luis, A. D., Kuenzi, A. J. & Mills, J. N. Species diversity concurrently dilutes and amplifies transmission in a zoonotic host–pathogen system through competing mechanisms. Proc. Natl. Acad. Sci. U.S.A. 115, 7979–7984 (2018).
Berngruber, T. W., Froissart, R., Choisy, M. & Gandon, S. Evolution of virulence in emerging epidemics. PLoS Pathog. 9, e1003209 (2013).
Farrer, R. A. et al. Multiple emergences of genetically diverse amphibian-infecting chytrids include a globalized hypervirulent recombinant lineage. Proc. Natl. Acad. Sci. 108, 18732–18736 (2011).
Thompson, J. D., Higgins, D. G. & Gibson, T. J. CLUSTAL W: Improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 22, 4673–4680 (1994).
Hall, T. A. BioEdit: A user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucl. Acids Symp. Ser. 41, 95–98 (1999).
Tamura, K., Stecher, G., Peterson, D., Filipski, A. & Kumar, S. MEGA6: Molecular Evolutionary Genetics Analysis Version 6.0. Mol. Biol. Evol. 30, 2725–2729 (2013).
Hutcheson, K. A test for comparing diversities based on the Shannon formula. J. Theor. Biol. 29, 151–154 (1970).
Acknowledgements
We are indebted to X. Asorey, J.M. Bello Santos, R. Díaz Nieto, M. Rodríguez Gómez and J.R. Suárez Estévez of the Asociación Galega de Apicutores (AGA) and the Servizo de Bombeiros de Santiago de Compostela for providing samples. We thank Damian Dasilva for critical reading of the manuscript. This work was partially financed by a joint partnership between the USC and Real Deputación de A Coruña, and the Interreg Atlantic Area Program (European Regional Development Fund—ERDF, European Union): EAPA_800/2018—Atlantic-Positive, to XM. Financial support from Xunta de Galicia (Centro singular de investigación de Galicia accreditation 2019–2022) and the European Union (ERDF) is also acknowledged.
Author information
Authors and Affiliations
Contributions
L.G.G., C.B. and X.M. conceived and designed the study. L.G.G., J.Ll., C.B. and C.G.T. performed the experiments. S.R. contributed samples. L.G.G. and X.M. wrote the paper. All authors reviewed and approved the submitted manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Gabín-García, L.B., Bartolomé, C., Guerra-Tort, C. et al. Identification of pathogens in the invasive hornet Vespa velutina and in native Hymenoptera (Apidae, Vespidae) from SW-Europe. Sci Rep 11, 11233 (2021). https://doi.org/10.1038/s41598-021-90615-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-90615-7
This article is cited by
-
Development of a TaqMan qPCR assay for trypanosomatid multi-species detection and quantification in insects
Parasites & Vectors (2023)
-
Bee Trypanosomatids: First Steps in the Analysis of the Genetic Variation and Population Structure of Lotmaria passim, Crithidia bombi and Crithidia mellificae
Microbial Ecology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.