Abstract
CLEC16A has been shown to play a role in autophagy/mitophagy processes. Additionally, genetic variants in CLEC16A have been implicated in multiple autoimmune diseases. We generated an inducible whole-body knockout, Clec16aΔUBC mice, to investigate the loss of function of CLEC16A. The mice exhibited a neuronal phenotype including tremors and impaired gait that rapidly progressed to dystonic postures. Nerve conduction studies and pathological analysis revealed loss of sensory axons that are associated with this phenotype. Activated microglia and astrocytes were found in regions of the CNS. Several mitochondrial-related proteins were up- or down-regulated. Upregulation of interferon stimulated gene 15 (IGS15) were observed in neuronal tissues. CLEC16A expression inversely related to IGS15 expression. ISG15 may be the link between CLEC16A and downstream autoimmune, inflammatory processes. Our results demonstrate that a whole-body, inducible knockout of Clec16a in mice results in an inflammatory neurodegenerative phenotype resembling spinocerebellar ataxia.
Similar content being viewed by others
Introduction
Several studies have described a role of CLEC16A in autophagic processes, particularly an inhibitory role on autophagy induction1,2,3,4 and mitophagy regulation5,6,7. CLEC16A is a membrane-associated endosomal protein, E3 ubiquitin ligase, that forms a ubiquitin-dependent complex with NRDP1 and USP85. Maintenance of the CLEC16A–NRDP1–USP8 mitophagy complex is necessary to sustain mitochondrial functions important for optimal cellular functions. Mitophagy, clearing of damaged or dysfunctional mitochondria, is crucial for mitochondrial quality control. Previously, we demonstrated the connection between CLEC16A and pancreatic ß cell function through mitophagy7. Loss of CLEC16A leads to an increase in the NRDP1 target PARKIN (PARK2), a master regulator of mitophagy7. We have found the loss of CLEC16A in immune cells results in dysregulated mitophagy and upregulated inflammatory cytokine response that increases the risk of autoimmunity8.
CLEC16A also is genetically linked with many distinct autoimmune disorders, including but not limited to type-1 diabetes, multiple sclerosis, systemic sclerosis, systemic lupus erythematosus, and rheumatoid arthritis9,10,11,12,13. Despite strong genetic evidence implicating CLEC16A in autoimmunity, this association remains unexplained mechanistically. CLEC16A may be interacting with other proteins to connect mitophagy/autophagy with autoinflammation/autoimmune processes. In preliminary studies, Tripartite motif 25 [TRIM25, aka estrogen-responsive finger protein (EFP)], an E3 ubiquitin, interferon stimulated gene 15 (ISG15) ligase, was identified as interacting with CLEC16A (Supplemental Table 1). ISG15, a ubiquitin-like modifier that inhibits polyubiquitination and is involved in a post-translational modification process called ISGylation, has a role in autophagy by conjugating with other proteins to control clearance of protein aggregates14. Activation of ISIG15 occurs after cell stress15, when neuronal damage is present16,17,18, and is hypothesized to be a potential cause of defective mitophagy in neurodegenerative diseases19. ISG15 and ISGylation are implicated in the control of mitochondrial OXPHOS and recycling in bone marrow-derived macrophages20. ISG15 conjugation to Parkin enhances its E3 ubiquitin ligase activity; this is an example of how ISGylation affects mitochondrial processes in neurodegeneration21. Free ISG15 stimulates the production of IFN-γ22. Because of its many roles in ubiquitination and inflammatory processes, we hypothesized ISG15 could be the link between CLEC16A and downstream autoimmune and inflammatory processes.
To better understand the role of CLEC16A in mitophagy and other autoimmune-inflammatory disorders besides diabetes, we created a Clec16a whole-body, inducible-knockout mouse model (Clec16aΔUBC). Clec16aΔUBC/ΔUBC homozygote mice (constitutive knockout) are embryonic lethal based on unequal genotype distribution in heterozygous breeding crosses, therefore, an inducible model was utilized. In 2016, Redmann et al. described two constitutive Clec16a mutant mouse models (Clec16aGT/GT and Clec16aCURT/CURT) with a neurodegenerative phenotype including Purkinje cell loss and locomotor deficits1. Because of the embryonic lethality of the constitutive Clec16aΔUBC/ΔUBC mice, we anticipated the severity of the phenotype may be greater in our inducible model than in the other constitutive mutants. In addition, the previous study only examined CNS areas where Clec16a is highly expressed, namely the cerebellar Purkinje cells and neurons of the deep cerebellar nuclei23.
In the present study, we describe a neurological phenotype in which Clec16aΔUBC mice develop severe neurologic dysfunction, including impaired gait and dystonic postures, in association with the degeneration of primary sensory axons. Astrogliosis and activated microglia are present in regions with axonopathy, and evidence of dysregulated autophagy and mitophagy in neural regions with and without axonopathy is observed. Alterations in mitochondrial-related proteins, ER stress markers, and IGS15 were observed in neuronal tissues. Hence, our Clec16aΔUBC mice are a suitable animal model of sensory neuron degeneration, and further illuminate a role for CLEC16A in inflammatory neurodegeneration.
Results
Rapidly progressive neurological disease in Clec16a ΔUBC mice
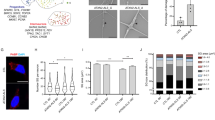
Young adult UBC-Cre-ERT2-Clec16aloxP/loxP mice were treated with tamoxifen for five successive days to generate adult Clec16a knockout (Clec16aΔUBC) mice as described previously8. Prior to treatment, Clec16aloxP/loxP and UBC-Cre-ERT2-Clec16aloxP/loxP were indistinguishable. Approximately 8 days after tamoxifen induction, UBC-Cre-ERT2-Clec16aloxP/loxP (Clec16aΔUBC) mice displayed weight loss that advanced quickly over days (Pandey et al., submitted). Tremors and other mild neurological behaviors (disability score = 1; Fig. 1B) started ~ 10 days and progressed rapidly, with about 60% of mice exhibiting the highest level of disability with dystonic postures after 25 days (disability score = 4; Fig. 1A; see also Supplemental Figure 1). A subset of the mice did not progress beyond a mild or moderate disability even at 40 days, when the experiment was terminated. These mice had higher Clec16a mRNA levels likely due to inefficient LoxP recombination. As observed in a previous study on Clec16a mouse models (Redman et al.), Clec16aΔUBC mice exhibited aberrant clasping and/or extension of the hindlimbs instead of the normal splay when picked up by their tails (Supplemental Figure 2). Some Clec16aΔUBC mice exhibited abnormal hind limb posturing by day 11, and all treated mice exhibited this behavior after 30 days. CLEC16A expression was decreased in multiple regions of the nervous system, including those with and without axonopathy (Supplemental Figure 3).
Clec16aΔUBC (whole-body, inducible knockout) mice exhibit neurological disability. (A) Image illustrating dystonic posturing in a representative ubiquitous, inducible KO (Clec16aΔUBC) mouse around 15 days post induction. (B) Timeline to onset of disability (scores 1–4) in a cohort of control/Clec16aloxP/loxP and KO/Clec16aΔUBC mice after treating P73–79 mice with tamoxifen for 4 successive days. Number of mice/group indicated on each graph; both sexes were used. *P < 0.05, **P < 0.01, ***P < 0.001 (Control vs. KO) for each disability score as analyzed by the Mantel–Cox test for survival curves.
Diminished amplitude of the caudal nerve action potential in Clec16a ΔUBC mice
The movement disorders we observed in Clec16aΔUBC mice were reminiscent of dystonic mutant mice, which have a recessive disorder that result in the progressive degeneration of primary sensory axons24. To investigate this issue, we examined Clec16aΔUBC mice that had reached a disability score of 4, 24 days after initiating tamoxifen treatment, along with age-matched control/Clec16aloxP/loxP mice. We measured the compound action potentials (CAPs) of the tail, which is a mixed nerve response from the bilateral dorsal and ventral caudal nerves, mainly comprised of myelinated and unmyelinated sensory axons25,26,27. Compared to control/Clec16aloxP/loxP mice, Clec16aΔUBC mice had a profoundly smaller CAP amplitude and a mildly decreased conduction velocity (Fig. 2).
Reduced caudal nerve amplitude and conduction velocity in Clec16aΔUBC mice. The graphs represent mean ± SEM of (A) compound action potential (CAP) and (B) conduction velocity in the caudal/tail nerves of a cohort of KO/Clec16aΔUBC mice that reached a disability scale of 4, 25 days after initiating tamoxifen treatment, and a cohort of control/Clec16aloxP/loxP mice. Number of mice/group indicated on each graph; both sexes were used. *P < 0.05, Student’s t test.
Pathological changes in Clec16a ΔUBC mice
To determine the anatomical location and substrates of the above abnormalities, we first examined various tissues, including the brains and spinal cords, of Clec16aΔUBC mice with the most severe disability vs. control mice, with hematoxylin and eosin (H&E) staining of paraffin sections. These experiments did not reveal any gross morphological changes (data not shown). We next stained frozen sections of the brain and spinal cord with fluoromyelin which labels myelin lipids and ASPA which labels oligodendrocyte cell bodies28,29,30,31, Calbindin to identify cerebellar Purkinje cells32 and GFAP-immunostaining which labels activated astrocytes33. This analysis revealed significant overall cerebellar Purkinje cell loss (22%, P = 0.01) and reactive astrocytosis (Supplemental Figure 4), in keeping with a previous report1. However, when the number of Calbindin + Purkinje cells and astrocytes were quantified as cells per unit area of cerebellum, a significant difference from controls was not detected (P = 0.17). The average area of the cerebellum was smaller in the Clec16aΔUBC mice, though not significantly (18% smaller, P = 0.12). We found significant increases in CD68-positive cells, IBA1 positive cells, and GFAP positive cells in the dorsal columns (Fig. 3) but not in other brain regions (data not shown). No differences were found between the Clec16aΔUBC mice and control mice in the amount of myelin lipids or oligodendrocyte cell bodies in spinal cord (Supplemental Figure 5).
Activated microglia and astrogliosis in the spinal cords, particularly dorsal columns, of Clec16aΔUBC mice. (A–C) Immunofluorescence images of the spinal cords from control/Clec16aloxP/loxP mice and KO/Clec16aΔUBC mice with severe disability and quantification of positive cell labelling. (A) Spinal cord sections were labeled with antibody to IBA1 (red), which labels microglia. (B) Spinal cord sections were labeled with antibody to CD68 (red) which labels activated microglia. (C) Spinal cord sections were labeled with antibody to GFAP (red), which labels astrocytes. For all, the number of positive labeled cells per spinal cord section was counted and divided by the area of the section in three sections from three mice per category and normalized to control. DAPI (blue) labels nuclei. Scale bars = 200 μm. *P < 0.05, ***P < 0.001, Student’s t test.
We also processed the spinal cord and various portions of the peripheral nervous system for transmission electron microscopy. In Clec16aΔUBC mice, we found prominent degenerating myelinated axons in the dorsal columns of the lumbar spinal cord, both in the fasciculis gracilis and the fasciculis cuneatus (Fig. 4), which contain the central axons of mechanoreceptors and proprioceptors, respectively34, but not in the lateral and ventral funiculi of the spinal cord, which contain ascending and descending axons from diverse types of neurons. In accord, we found degenerating myelinated axons in the dorsal roots, which are comprised only of sensory axons, but not in the ventral roots, which are comprised only of motor axons (Fig. 5). Similarly, we found degenerating myelinated axons in the femoral sensory and motor branches (Supplemental Figure 6) and in the tibial, peroneal, and sural branches of the sciatic nerve (data not shown). Axon size distributions were examined and the frequency of smaller axons in Clec16aΔUBC mice was higher than in control mice (median 5.4 vs 7.4 μm2, mean ± SEM. 5.4 ± 0.10 vs 9.7 ± 0.10 μm2, respectively; Supplemental Figure 7). This pattern of axonal degeneration, involving dorsal roots in addition to their more distal aspects, is distinct from what one sees in other animal models of peripheral neuropathy, in which one finds distal axonal degeneration, sparing the more proximal axons in the dorsal roots35. Because we did not find degenerating sensory neurons in semi-thin sections of the dorsal root ganglia (DRG), it appears that Clec16aΔUBC mice develop a sensory neuropathy that affects proximal and distal axons of both the roots and the peripheral nerves.
Degenerating sensory axons in the dorsal columns of Clec16aΔUBC mice. These are representative images of semi-thin sections of the lumbar spinal cord from control/Clec16aloxP/loxP and KO/Clec16aΔUBC mice with severe disability (score 4), as indicated. The upper panels show the locations of the corticospinal tract (CST), fasciculus gracilis (FG), and fasciculus cuneatus (FC). The lower panels correspond to the rectangular regions depicted in the upper panels, and show many degenerating myelin sheaths in the FG and FG (the boundaries of which are shown by the dotted curves) in the KO/Clec16aΔUBC sample. Scale bar = 100 μm upper panel, 10 μm lower panel.
Selective degeneration of sensory and not motor axons in Clec16aΔUBC mice. These are images of semi-thin sections from the dorsal and ventral roots from control/Clec16aloxP/loxP and KO/Clec16aΔUBC with severe disability (score 4) mice, as indicated. Degenerating myelinated axons in nerves are only found in the dorsal roots from KO/Clec16aΔUBC mice. Scale bar = 10 microns.
To compare the myelin thickness between control and Clec16aΔUBC mice, the ratio between the inner axon radius and the outer myelinated axon radius (g ratio) of femoral sensory nerves from 6 mice (each group) was measured (Supplemental Figure 8). Though a statistically significant decrease was found, the difference between the groups was very small and unlikely to be of biological relevance (control 0.73 ± 0.0008 SEM, n = 5359 neurons, KO group 0.70 ± 0.0011, n = 4326 neurons).
Clusters of vacuoles were the most conspicuous finding in the DRG neurons of Clec16aΔUBC mice, as previously noted in Clec16a-null (Clec16aGT/GT) cerebellar Purkinje cells1 (Fig. 6). In a single, semi-thin section, 11% (range 4–22%) of DRG neurons had clusters of vacuoles (N = 9 animals) in Clec16aΔUBC mice; none were found in Clec16aloxP/loxP control mice (N = 5 animals). By electron microscopy (EM; Fig. 6A,B,G), these vacuoles had a single membrane structure, and adjacent vacuoles were seen to protrude into each other, like a finger poking into a balloon, such that some vacuoles appeared to contain a double-membrane structure. We did find clusters of vesicles in a few of the stem processes of DRG neurons (data not shown), but not in the axons of the dorsal and ventral roots or in femoral and sciatic nerves. The vacuoles were postulated to be derived from the Golgi apparatus1 but we did not recognize structures that looked like transitional structures between Golgi apparatus and vacuoles, and axons are not known to contain Golgi apparatus.
Abnormal vacuoles and mitochondria in Clec16aΔUBC neurons. These are electron micrographs of sensory (A-F) and motor (G&H) neurons from KO/Clec16aΔUBC with severe disability (score 4; A–D, G) and control/Clec16aloxP/loxP (E,F,H) mice. In panel (A), there are five sensory neurons, three of which (1, 2, and 4) contain cytoplasmic clusters of clear vesicles; the area enclosed by the rectangle is enlarged in panel (B). The arrows in panel (B) mark two examples of a vesicle that protrudes into an adjacent vesicle; the arrowheads mark three examples in which the origin of the protrusion is not evident in the plane of section. Panel (C) shows another sensory neuron that contains abnormal mitochondria labeled in panel (D) (asterisks), which is an enlargement of the area shown in the rectangle. Panel (E) shows a sensory neuron from a control/Clec16aloxP/loxP mouse; the area enclosed by the rectangle is enlarged in panel (F), and shows a Golgi apparatus (g) and normal appearing mitochondria (asterisks). Panels (G) and (H) show portions of cervical motor neurons from a KO/Clec16aΔUBC and a control/Clec16aloxP/loxP mouse, respectively. Note the clusters of clear vesicles in the KO/Clec16aΔUBC motor neuron, and the normal appearing Golgi apparatus (g) in the control/Clec16aloxP/loxP neuron. Scale bars, 2 μm (A,C,E) and 600 nm (B,D,H).
We also noted vacuoles in a few (> 1%) of the larger neurons in transverse, semi-thin sections of the cervical and lumbar spinal cords of Clec16aΔUBC mice, between 3–13 such neurons were found per section (N = 7 animals), in both the dorsal and ventral horns. None were found in Clec16aloxP/loxP control mice (N = 7 animals). By EM (Supplemental Figure 9), a minority of affected neurons had vacuoles that were similar in appearance to those seen in the DRG neurons. Most of the affected neurons, however, had vacuoles that were much larger (Supplemental Figure 9B) than those seen in the DRG neurons, and these affected neurons also had the more typical, smaller vacuoles, too. The relationship between the smaller and larger vacuoles warrants further investigations.
We also noted some DRG neurons (less than 10%) had large numbers of abnormal mitochondria, with an electron-lucent matrix and reduced cristae (Fig. 6C,D). We do not think that these abnormal mitochondria were merely the result of poor fixation, as cells with abnormal mitochondria were found adjacent to cells with normal mitochondria in Clec16aΔUBC mice, and cells with abnormal mitochondria were not found in control mice. In Clec16aΔUBC mice, the cells with abnormal mitochondria usually did not contain clusters of vacuoles.
DRG sensory neurons from Clec16aΔUBC mice also exhibited myelin debris and altered myelin-like structures (Fig. 4, lower right panel). The myelin debris found in the peripheral nerves is likely a consequence of Wallerian degeneration, which occurs when the part of the axon distal to the injury degenerates, followed by degeneration of the myelin sheath and infiltration of macrophages which then clear the debris, as shown previously36.
Alterations in mitophagy-related proteins, endoplasmic reticulum (ER) stress, and oxidative phosphorylation (OXPHOS) related proteins in Clec16a ΔUBC mice
Previous reports from our laboratory and others have shown an important role of CLEC16A in mitophagy functions in immune cells, islet β-cells, and cerebellum1,5,6,7,8. Similarly, in this study we found increased expression of the mitophagy proteins, PINK1, Parkin, and p62, as well as the autophagic protein, lipidated form of LC3 (Supplemental Figure 10). The global effects of a whole-body Clec16a deletion on these mitophagy and autophagy proteins contrasts to its specific effects on sensory neurons.
We next examined the effect of Clec16a knockout on ER stress and OXPHOS signaling in mitochondrial in light of previous observations7,37. Unfolded-protein response and OXPHOS signaling related genes and proteins were examined, focusing on the sensory neuron tissues. The ER stress marker genes were evaluated at mRNA level in sensory neuron tissues, the trigeminal ganglia and DRG, with mild disability (Day = 10) and severe disability (Day = 22). As found previously in other tissues, ER stress markers (GRP78, ATF4, and CHOP) showed significant upregulation at mRNA levels in the Clec16aΔUBC mice with mild and severe disability in both tissues (Supplemental Figure 11A–D). XBP1 was increased in DRG and trigeminal ganglia (TG) of Clec16aΔUBC mice with severe disability. Immunoblot analysis revealed significant increase in expression of CHOP in both DRG and TG’s of mice with severe disability (Supplemental Figure 11C).
We also measured OXPHOS respiratory complex protein levels in sensory neuron tissues (Supplemental Figure 11E–F). The Complex 1 protein, NADH dehydrogenase (ubiquinone) 1 beta subcomplex 8 (NDUFB8) and Complex IV protein, cytochrome c oxidase subunit 2 (COXII), were increased in both DRG and TG of the Clec16aΔUBC mice. The TG of Clec16aΔUBC mice showed an increase of Complex III ubiquinol-cytochrome c reductase core protein II (UQCR2). Complex V protein ATP synthase 5A (ATP 5A) was decreased in both DRG and TG of Clec16aΔUBC mice.
CLEC16A expression is inversely related to IGS15 protein expression
In search of potential candidates interacting with CLEC16A, MSMS analysis was performed on lysates of YTS and YTS-Clec16a overexpressing cells immunoprecipitated with CLEC16A. Tripartite motif 25 (TRIM25, aka estrogen-responsive finger protein (EFP)), an E3 ubiquitin, interferon stimulated gene 15 (ISG15) ligase, was identified as one of the top ten hits among the potential candidate proteins interacting with CLEC16A (Supplemental Table 1). A recent study shows TRIM25 is significantly induced upon ER stress, promoting ER-degradation in an attempt to restore ER homeostasis38. Because TRIM25 regulates ISG15, ISG15 expression is low under normal conditions, the ubiquitin pathway regulates mitophagy, ISG15 inhibits the ubiquitin pathway, and activation of ISG15 occurs when neuronal damage is present, we hypothesized that a constitutively elevated ISG15 pathway inhibits the ubiquitin pathway and consequently ubiquitin-dependent-mitophagy, which may contribute to neurodegeneration in Clec16aΔUBC mice and CLEC16A’s actions in sensory neurons. We measured ISG15 protein levels in several neuronal tissues in Clec16aΔUBC mice with mild disability (score = 1) and severe disability (score = 4) compared to Clec16aloxP/loxP/control mice. ISG15 levels in control mice were low as predicted. Significantly increased levels of ISG15 were observed in all neuronal tissues tested from Clec16aΔUBC mice with mild disability compared to controls (Fig. 7). The increase in ISG15 levels was more pronounced in the spinal cord, TG and DRG in Clec16aΔUBC mice with severe disability, while levels were unaltered in cerebellum, cortex or striatum.
ISG15 expression in neuronal tissues of Clec16aloxP/loxP and Clec16aΔUBC mice. (A) Representative western blot from cerebellum, cortex, spinal cord, striatum, TG and DRG lysates of control/Clec16aloxP/loxP and KO/Clec16aΔUBC mice depicting expression levels of ISG15. Membranes were striped and re-probed for β-actin as a loading control. Image is cropped from the full membrane image; the full membrane image can be found in the Supplemental Figure 12. (B) Quantitation graph depicting expression levels of ISG15. Data expressed as mean ± SE of three independent repeats of two mice each. *P < 0.05, ***P < 0.001 (Control vs. KO-Score 1), ##P < 0.01 (Control vs. KO-Score 4), Student’s t test.
Discussion
We show for the first time that whole body deletion of Clec16a in adult mice results in an overt neurological phenotype that is associated with loss of primary sensory axons but not the loss of Purkinje cells seen in other Clec16a mutant models. These results are similar to the recent finding that homozygous mice carrying a gene-trap insertion in Clec16a (Clec16aGT/GT) develop an overt neurological phenotype at 7–8 weeks of age, ascribed to the loss of cerebellar Purkinje cells. Clec16acurt/curt mice, which have 4-nucleotide deletion allele in exon 21, resulting in a frameshift mutation, also lose cerebellar Purkinje cells1,39. It remains to be determined whether primary sensory neurons are affected in Clec16aGT/GT and Clec16acurt/curt mice. Further studies will be completed to understand why a greater loss of Purkinje cells was not observed in the Clec16aΔUBC/ΔUBC mice.
Clec16aΔUBC/ΔUBC homozygote mice have a severe phenotype and are embryonic lethal based on unequal genotype distribution in heterozygous breeding crosses. In contrast, Clec16aGT/GT mice have a delayed and mild behavioral phenotype as well as longer survival1; the Clec16acurt/curt mice, have decreased body weight, skeletal abnormalities, and prenatal/perinatal death1,39,40. In view of the above, we infer that the Clec16aGT allele likely produces a partial loss of function.
We observed weight loss and lipodystrophy in the Clec16aΔUBC mice prior to the manifestation of the neuronal phenotype (Pandey et al., submitted). Lipid molecules are important for key brain structure and functions and comprise about 50% of the brain composition; most of this is contained in the myelin sheath around neurons41. The neuronal phenotype was not the result of demyelination as myelin-related proteins were not altered in the Clec16aΔUBC mice with severe disability as compared to Clec16aloxP/loxP/control mice. Likewise, only the sensory neurons were affected and the motor neurons and other CNS structures were intact indicating a global demyelination did not occur. However, ER stress mRNAs were increased in trigeminal and dorsal root ganglia. The brain generates its own phospholipids and, therefore, may be protected from the lipolytic cascade present in adipose tissue. Previous studies show that high stress in neurons stimulates the formation of lipid droplets in nearby glial cells preventing fatty acid toxicity in the neurons themselves serving as neuronal protection42.
Sensory neurons are responsible for conveying external environmental stimuli to the central nervous system. When there is a loss of these neurons, the transmission of information is incomplete and consequently the brain cannot send motor impulses to respond correctly to its surroundings. The selective involvement of cerebellar and/or primary sensory neurons in Clec16a KO mice models a human syndrome known as spinocerebellar ataxia, which has diverse systemic and genetic causes43,44,45. Though CLEC16A has not been implicated in human spinocerebellar ataxia, other mitophagic and autophagic proteins have been identified. Some mouse models of other autophagocytic-related proteins exhibit gait abnormalities, ataxia, cerebellar/Purkinje cell deficits, fewer dendritic spines, axonal swelling, and other CNS abnormalities. There are few mouse models of a sensory neuronopathy—Atg7 conditional KO using Advillin-Cre mice, and Dstdt transgenic mice46,47. Our Clec16aΔUBC mice model represents a sensory ataxia with dystonia, immune and inflammatory components seen in some human cerebellar ataxias and sensory neuronopathies45,48.
Sensory neuropathy and/or cerebellar degeneration have been observed in autoimmune disorders that are associated with CLEC16A such as type I diabetes, celiac disease, multiple sclerosis, systemic lupus erythematosus, autoimmune thyroid disease, and systemic sclerosis (scleroderma)45,49,50,51,52,53,54,55. The connection between CLEC16A variants and sensory neuropathy or cerebellar degeneration have not been considered clinically in any of these disorders. Cerebellar Purkinje cells express high levels of CLEC16A23 which may contribute to some loss of cells in the Clec16aΔUBC mice. Others have noted that mechanisms underlying spinocerebellar ataxia, as well as dystonia, involves specific pathways likely crucial for Purkinje cell function43,56, indicating CLEC16A is connected to one of these pathways.
The role of TRIM25 has been further developed recently to not only include its role in innate immunity57,58 and metabolism59 but also to oncology38,60 and mediating clearance of misfolded proteins61. It is unclear if TRIM25 is interacting with CLEC16A and affecting neuropathy in our Clec16aΔUBC mice. Future studies will address the direct CLEC16A interaction with TRIM25 and its function or impact in neuronal loss. The contribution of IGS15 to neurodegeneration by inhibiting the ubiquitin pathway is intriguing, but not yet confirmed19. One role of free ISG15 is stimulating the production of IFN-γ from CD3+ T cells to enhance the proliferation and cytotoxicity of natural killer (NK) cells22. A recent study found that stimulated NK cells degenerate partially injured sensory axons62. Since increased NK cell cytotoxicity is observed in the Clec16aΔUBC mice37, future studies could address whether this is a potential mechanism for selective sensory neuron loss.
Autophagy and apoptosis are important and interconnected stress response mechanisms. Dead or damaged cells constitute a source of novel antigens and proinflammatory molecules that can provoke autoimmune inflammatory immune response. ISG15 is in the type-I-interferon (IFN) signaling cascade and is increased after cell stress, in neuronal injury and motor neuron disease (ALS) in humans and mice15,16,63,64. A role for ISG15 in the mitochondria is established as 17% of free ISG15 and about 5% of total ISG15 target proteins are localized in the mitochondria. ISG15 and ISGylation are implicated in the control of mitochondrial OXPHOS and recycling in bone marrow-derived macrophages20. We found increases in ISG15 in neuronal tissues during the early phases of neuronal disability with a concomitant decrease in the brain regions at the later, more severe disability timepoint. Upregulation of ISG15 has been noted in several neuronal injury models involving inflammation and in a mouse model of ALS motor neuropathy16,64. However, in the models described, ISG15 was not increased in unaffected CNS areas. These data suggest that CLEC16A directly or indirectly more broadly impacts IGS15 activation and protein ISGylation, which are associated with enhanced and prolonged JAK-STAT signaling, initiating SRC family tyrosine kinases, and ultimately increased cytokine and chemokine production in the Clec16aΔUBC mice.
Targeting any of these pathways often induces harmful side effects on top of positive effects thereby negating potential benefits. While our current exploratory intervention experiments in the Clec16aΔUBC mice show only partial rescue of the phenotype (Pandey et al., submitted), newer, more selective inhibitors targeting IFN-STAT signaling (Ruxolitinib) alone or in combination with autophagy/mitophagy modulators, represent potential opportunities for autoimmune and inflammatory disorders without the harmful side effects65.
The complexity of each condition, and the presence of different types of neuropathies across different autoimmune diseases, complicates the elucidation of how CLEC16A is related to sensory neuron loss. Whether the above altered pathways contribute to the selective loss of primary sensory neurons in Clec16aΔUBC mice remains to be determined. Autophagy is necessary for neuronal homeostasis and survival66,67, but this alone would not account for the selective vulnerability of affected neurons. Nor would astrogliosis and activated microglia be the logical explanation, as both are found in a wide variety of CNS insults. The Clec16aloxP/loxP mice provide a valuable murine model to elucidate these issues in sensory ataxia as well as mitochondrial function in neurodegeneration.
Materials and methods
The study is reported in accordance with ARRIVE guidelines.
Mice
The Institutional Animal Care and Use Committee of the Children’s Hospital of Philadelphia approved all animal studies. All methods were performed in accordance with the IACUC guidelines and regulations. Mice were group-housed on an individually-ventilated cage rack system on a 12:12 light:dark cycle. Mice were fed standard rodent chow and water ad libitum.
Mice were generated as described previously8 Clec16aloxP mice were generated by flanking exon 3 (Ozgene). Mice with targeted insertion in the Clec16a gene were crossed to the Flpo Deleter line (mouse Strain: 129S4/SvJae-Gt (ROSA) 26Sortm2(FLP*) Sor/J; The Jackson Laboratory) to achieve deletion of the FRT-flanked Neomycin cassette. Clec16aloxP mice were mated to UBC-Cre-ERT2 mice (inducible cre recombinase driven by the human ubiquitin C promoter) obtained from The Jackson Laboratory to generate UBC-Cre-Clec16aloxP mice. Mice were kept on a mixed background C57BL/6-129S1.
Ten-week old UBC-Cre-ERT2-Clec16aloxP/loxP male and female mice were treated orally by gavage with tamoxifen (100 mg/kg/day) at 24-h intervals for four consecutive days to create experimental Clec16aΔUBC groups. Clec16aloxP/loxP littermates dosed with either tamoxifen or vehicle (10% ethanol:90% corn oil) were used as controls. There were no differences in behavior between the two control groups, so they were combined for phenotypic analyses. All images depicting mouse behavior were taken by one of the authors of the manuscript.
Behavioral observations
Mice were an average of 10 weeks old at beginning of study. Mice were weighed 3 times a week. Mice were scored for gross abnormalities and behavior, including disability and hindlimb splay.
Disability was scored 0–4 as follows:
-
0 = Normal behavior, appearance and movement
-
1 = No impairment; abnormal behavior or appearance (ex: tremor, not grooming)
-
2 = Mild impairment; uneven gait (ex: waddling, abdomen pressed to floor)
-
3 = Moderate impairment; body constrictions, unsteady gait upon initiation of movement
-
4 = Severe impairment; abdominal twisting and limb contractions upon initiation of movement, little to no forward movement, ambulation impaired
Hindlimb splay was scored 0–2 as follows:
-
0 = normal, hind limbs are (splayed) out when lifted
-
1 = One hind limb is partially grasping or contracted out towards tail; abnormal, one or both hind limbs are drawn partially into the body or partially grasping more than 50% of observation time
-
2 = Both hind limbs are grasping together immediately; impaired, both hind limbs are grasping together or contracted out towards tail more than 50% of observation time
Observations were completed prior to treatment and then 3 times per week until clinical signs were noticed at which point mice were observed and scored daily. Once a mouse reached the highest disability score of ‘4’ or greater than 20% body weight loss from baseline, it was euthanized and tissues were collected. Some mice were euthanized at different disability scores (without the opportunity to progress a more severe score) to understand the molecular mechanism of CLEC16A. Mice remaining at the end of the study (40 days) were euthanized and organs were harvested for future analyses.
Nerve conduction study
A portable EMG/NCS machine (Nicolet) was used to record the CAPs from caudal nerves. Mice were anesthetized with ketamine: xylazine (10:100 mg/kg, IP). After full anesthesia was verified by toe pinch, mice were placed supine with the tail fully extended. Electrodes were placed on the tail following the method of Maia et al.25 Wire loop electrodes were used for recording and placed with the reference loop 0.2 cm from the tail base and the active electrode 0.5 cm more distal along the tail. One-pin electrodes were used as ground electrodes (separated by 0.6 cm). More distally along the tail, a pair of loop electrodes was used for stimulation (separated by 0.5 cm). The distance between the recording electrode and stimulating cathode electrode is three cm. The intensity of electrical stimulation for mice is equivalent to the intensity used in clinical nerve conduction studies in awake human. A series of 5 responses were averaged to reduce noise and variability. Tail surface temperature was maintained at 30–32 °C using a heating pad and monitored using infrared non-contact thermometer during recording. Upon completion of recording, mice were returned to the cage and closely observed until they fully recovered.
Western blot
As described in Pandey et al.8,37, sample lysis was performed with NP40 lysis buffer (Invitrogen). The lysates were electrophoresed on 4–12% NuPAGE Bis–Tris gels in MOPS SDS running buffer and transferred onto nitrocellulose membranes (Invitrogen). The membranes were blocked in 3% BSA and incubated with indicated primary antibodies where specified: CLEC16A, PINK1 (Abgent), Parkin, p62/SQSTM1, LC3 I/II for mitophagy/autophagy (Santa Cruz), CHOP for ER stress (Novus Biologicals) and ISG15 (Cell Signaling Technology). For OXPHOS signaling, a commercially available cocktail antibody (ab110413, Abcam) comprising the following subunits of respiratory complex proteins were used: NADH dehydrogenase (ubiquinone) 1 beta subcomplex 8 (NDUFB8; complex I), succinate dehydrogenase complex, subunit B, iron sulfur (SDHB/Ip; complex II), ubiquinol-cytochrome c reductase core protein II (UQCR2; complex III), cytochrome c oxidase subunit 2 (COXII; complex IV) and ATP synthase 5A (ATP 5A, Complex V).
The membranes were washed and incubated with a respective secondary antibody as recommended by the manufacturer. Bound antibody was detected with WesternBright ECL chemiluminescence detection system (Advansta, San Jose, CA). Membranes were stripped and re-probed with mouse anti-β-Actin mAb (Abcam) as a loading control. Membranes were cut in parts and probed for protein of interest where specified. Band intensities were measured using Image J software (v1.53, NIH Shareware), scanned in grey scale mode at 300 DPI and saved in TIFF format, measuring the area under each peak for each band.
Quantitative real-time PCR
As described in our previous publication37, total RNA was isolated with Trizol reagent (Invitrogen) following RNA purification using the RNeasy Mini Kit (Qiagen) and converted to cDNA by High Capacity RNA-to-cDNA Kit (Applied Biosystems), according to the manufacturer’s protocols. Assay comprising known human and murine CLEC16A RNA transcripts as well as control genes (β-actin and HRPT1) were measured by real time PCR on a ViiA 7 Real Time PCR System using predesigned 20X FAM-MGB TaqMan gene expression assays available from Applied Biosystems. All assays had primers covering exon-exon boarders to avoid DNA contamination. Triplicates were used for all samples included in the experiment. All PCR runs were performed on ViiA 7 Real Time PCR System using ViiA7 RUO software v1.2.2 (Life Technologies).
Immunohistochemistry
Mice were euthanized, perfused with 4% paraformaldehyde, brains were embedded in OTC, and frozen sections were cut on a Leica cryostat at 12-micron thicknesses, all according to established protocols68,69. Hematoxylin and eosin (H&E) staining was performed by the histology core at Children’s Hospital of Philadelphia. Oligodendrocytes were labeled with fluoromyelin which labels myelin lipids and ASPA which labels oligodendrocyte cell bodies28,29,30,31. Microglia/macrophages were labeled with anti-IBA-1 (Wako Pharmaceuticals, 1:1000) and anti-CD68 (Abcam, 1:100)70,71. Astrocytes were labeled with used anti-GFAP antibody (rat monoclonal, undiluted, gift of Dr. Virginia Lee, University of Pennsylvania)33. Purkinje cells were labeled with an anti-calbindin antibody (Swant, 1:2000)32. Sections were incubated with primary antibodies overnight at 4 °C. Secondary antibodies of appropriate species and isotype used for external and internal antigens were purchased from Jackson Immunoresearch, West Grove, PA and used at 1:200 for 30 min. Coverslips were mounted onto glass slides in 4′,6-diamidino-2-phenylindole (DAPI)-containing Vectashield mounting medium (Vector Laboratories, Burlingame, CA, USA). For all, the number of positive labeled cells per spinal cord or cerebellar section was counted and divided by the area of the section in three sections from three mice per category and normalized to control.
Transmission electron microscopy
Anesthetized mice were transcardially perfused with 2% paraformaldehyde and 2% glutaraldehyde in 0.1 M phosphate buffer (pH 7.4). DRG and cervical spinal cords were dissected, and fixed for at least 4 more hours at 4 °C. The tissues were osmicated, dehydrated, infiltrated, and then embedded in Embed 812 mixture (Electron Microscopy Sciences) as previously described72. For light microscopy, cross-sections were cut at a thickness of 1 μm on an American Optical Reichert Ultracut Ultra microtome, and stained with alkaline toluidine blue. For electron microscopy, cross-sections were cut at a thickness of 90 nm, and stained with lead citrate and uranyl acetate. The ultra-thin sections were imaged using a JEOL 1010 electron microscope.
Axon size distribution and measurement of g ratio of femoral sensory nerves
Femoral sensory nerves from six mice in each group (KO and control) were analyzed. All the myelinated axons of one LM images (× 100) from each mouse were analyzed. The g-ratio was calculated by the square root of the ratio of inner to outer axonal area since most axons were not perfectly circular. Inner and outer axonal area were measured using Image J software (v1.53, NIH Shareware).
Mass spectrometry analysis of Clec16a
Methods for the mass spectrometry are as described in Greco et al.73 As described in Pandey et al.37, the YTS cell line were grown in 5% CO2 at 37 °C in complete RPMI medium (Gibco). The “complete” medium indicates it was supplemented with 10% FBS (Thermo Scientific), non-essential amino acids, HEPES, l-glutamine, sodium pyruvate, and penicillin–streptomycin (all from Gibco). The YTS-CLEC16A (Clec16a overexpressing) cell line37 was grown in 5% CO2 at 37 °C in complete RPMI medium as described above with G418 (1.6 g/L, Mediatech). Briefly, 5 × 106 YTS-CLEC16A and YTS cells were lysed in ice cold IP-lysis buffer. After centrifugation at 13,000 rpm for 10 min at 4 °C, supernatants were pre-cleared with 50 μl of agarose G beads (Invitrogen) and IgG for 45 min at 4 °C. The pre-cleared lysates were incubated with CLEC16A antibody for 1 h at 4 °C and then with 50 μl of agarose G beads for an additional 1 h at 4 °C. Immune complexes were washed three times, dissolved in SDS sample buffer, and ~ 50 μg was resolved by SDS-PAGE. Gel regions were excised from the Coomassie stained gels and cut into 1 mm3 cubes (2, 3)(2, 3)(2, 3)(2, 3)(2, 3)(2). The latter served as controls for gel loading and internal standards for MS analysis. The excised gel pieces were then washed with 100 μl of 50 mM ammonium bicarbonate, followed by 50 mM ammonium bicarbonate in 50% acetonitrile and then solvent removal in a vacuum centrifuge. Dehydrated gel pieces were then reswollen in 100 μl of a digestion buffer containing 50 mM ammonium bicarbonate, and 12.5 ng/ml of trypsin and enzymatic cleavage was allowed to continue overnight at 37 °C. Peptides were extracted into 20 mM ammonium bicarbonate (100 μl) followed by two separate extractions into 100 μl of water/acetonitrile/formic acid (10:10:1; v/v/v). After evaporation to dryness, peptides were re-dissolved in 10 μl of 5% acetonitrile and 0.1% FA for LC/MS/MS analysis. For peptide mass fingerprinting of samples, 0.5 μl of each Tryptic digest was mixed with 1 μl of matrix (saturated aqueous solution of 2,5-dihydroxybenzoic acid) and spotted onto a sample target plate and dried. Matrix-assisted laser desorption/ionization time-of-flight (MALDI-TOF) mass spectra were acquired using an externally calibrated Ultraflex TOF/TOF mass spectrometer (Bruker Daltonics) in the reflector mode. After internal calibration using trypsin autolysis peaks, prominent peaks in the mass range m/z 700–4000 were used to generate a peptide mass fingerprint that was searched against the updated human IPI database using Mascot version 2.1.3 (Matrix Sciences, London, UK). Identifications were accepted when a minimum of six peptide masses matched to a particular protein (mass error of ± 50 ppm allowing 1 missed cleavage), sequence coverage was > 25%, MOWSE scores were higher than the threshold value (P = 0.05) and the predicted protein mass agreed with the gel-based mass.
Nano-HPLC electrospray ionization (ESI) collision induced dissociation (CID) MS/MS was performed on a Q-TOF mass spectrometer (Waters, Manchester, UK), coupled to an Ultimate system LC (Dionex) with a PepMap C18 75 μm inner diameter column at a flow rate of 300 nl/min. Spectra were processed using MassLynx software (v4.0, Waters) and submitted to Mascot database search routines. Positive identifications were made when at least two peptide sequences matched an entry and MOWSE scores were above the significance threshold value (P = 0.05). The data analysis and sample report were created using Proteome Software Scaffold (version: Scaffold_3.3.3).
Statistical analysis
All graphs denote mean values, except where noted, and error bars represent the SEM. Comparisons of groups of data were performed using Log-rank (Mantel-Cox) test for survival curves or two-tailed Student’s t tests for parametric data (Prism ver7, GraphPad Software, Inc.). Statistical significance is shown in figures (*P < 0.05, **P < 0.01, ***P < 0.001). No phenotypic difference was observed between the Clec16aloxP/loxP treated with vehicle or Clec16aloxP/loxP mice treated with tamoxifen, so data were combined for comparison to Clec16aΔUBC mice.
References
Redmann, V. et al. Clec16a is critical for autolysosome function and Purkinje cell survival. Sci. Rep. 6, 23326. https://doi.org/10.1038/srep23326 (2016).
Schuster, C. et al. The autoimmunity-associated gene CLEC16A modulates thymic epithelial cell autophagy and alters T cell selection. Immunity 42, 942–952. https://doi.org/10.1016/j.immuni.2015.04.011 (2015).
Tam, R. C. et al. Human CLEC16A regulates autophagy through modulating mTOR activity. Exp. Cell Res. 352, 304–312. https://doi.org/10.1016/j.yexcr.2017.02.017 (2017).
Kim, S., Naylor, S. A. & DiAntonio, A. Drosophila Golgi membrane protein Ema promotes autophagosomal growth and function. Proc. Natl. Acad. Sci. U.S.A. 109, E1072–E1081. https://doi.org/10.1073/pnas.1120320109 (2012).
Pearson, G. et al. Clec16a, Nrdp1, and USP8 form a ubiquitin-dependent tripartite complex that regulates beta-cell mitophagy. Diabetes 67, 265–277. https://doi.org/10.2337/db17-0321 (2018).
Soleimanpour, S. A. et al. Diabetes susceptibility genes Pdx1 and Clec16a function in a pathway regulating mitophagy in beta-cells. Diabetes 64, 3475–3484. https://doi.org/10.2337/db15-0376 (2015).
Soleimanpour, S. A. et al. The diabetes susceptibility gene Clec16a regulates mitophagy. Cell 157, 1577–1590. https://doi.org/10.1016/j.cell.2014.05.016 (2014).
Pandey, R. et al. CLEC16A regulates splenocyte and NK cell function in part through MEK signaling. PLoS ONE 13, e0203952. https://doi.org/10.1371/journal.pone.0203952 (2018).
Hakonarson, H. et al. A genome-wide association study identifies KIAA0350 as a type 1 diabetes gene. Nature 448, 591–594. https://doi.org/10.1038/nature06010 (2007).
Berge, T., Leikfoss, I. S. & Harbo, H. F. From identification to characterization of the multiple sclerosis susceptibility gene CLEC16A. Int. J. Mol. Sci. 14, 4476–4497. https://doi.org/10.3390/ijms14034476 (2013).
Gorlova, O. Y. et al. Gene-level association analysis of systemic sclerosis: A comparison of African-Americans and White populations. PLoS ONE 13, e0189498. https://doi.org/10.1371/journal.pone.0189498 (2018).
Skinningsrud, B. et al. A CLEC16A variant confers risk for juvenile idiopathic arthritis and anti-cyclic citrullinated peptide antibody negative rheumatoid arthritis. Ann. Rheum. Dis. 69, 1471–1474. https://doi.org/10.1136/ard.2009.114934 (2010).
Tam, R. C., Lee, A. L., Yang, W., Lau, C. S. & Chan, V. S. Systemic lupus erythematosus patients exhibit reduced expression of CLEC16A isoforms in peripheral leukocytes. Int. J. Mol. Sci. 16, 14428–14440. https://doi.org/10.3390/ijms160714428 (2015).
Villarroya-Beltri, C., Guerra, S. & Sanchez-Madrid, F. ISGylation—A key to lock the cell gates for preventing the spread of threats. J. Cell Sci. 130, 2961–2969. https://doi.org/10.1242/jcs.205468 (2017).
Zhang, D. & Zhang, D. E. Interferon-stimulated gene 15 and the protein ISGylation system. J. Interferon Cytokine Res. 31, 119–130. https://doi.org/10.1089/jir.2010.0110 (2011).
Wang, R. G., Kaul, M. & Zhang, D. X. Interferon-stimulated gene 15 as a general marker for acute and chronic neuronal injuries. Sheng Li Xue Bao 64, 577–583 (2012).
Kim, C. D., Reed, R. E., Juncker, M. A., Fang, Z. & Desai, S. D. Evidence for the deregulation of protein turnover pathways in Atm-deficient mouse cerebellum: An organotypic study. J. Neuropathol. Exp. Neurol. 76, 578–584. https://doi.org/10.1093/jnen/nlx038 (2017).
Liu, C. et al. Identification of imidazo[1,2-b]pyridazine derivatives as potent, selective, and orally active Tyk2 JH2 inhibitors. ACS Med. Chem. Lett. 10, 383–388. https://doi.org/10.1021/acsmedchemlett.9b00035 (2019).
Desai, S., Juncker, M. & Kim, C. Regulation of mitophagy by the ubiquitin pathway in neurodegenerative diseases. Exp. Biol. Med. (Maywood) 243, 554–562. https://doi.org/10.1177/1535370217752351 (2018).
Albert, M., Becares, M., Falqui, M., Fernandez-Lozano, C. & Guerra, S. ISG15, a small molecule with huge implications: Regulation of mitochondrial homeostasis. Viruses https://doi.org/10.3390/v10110629 (2018).
Im, E., Yoo, L., Hyun, M., Shin, W. H. & Chung, K. C. Covalent ISG15 conjugation positively regulates the ubiquitin E3 ligase activity of parkin. Open Biol. https://doi.org/10.1098/rsob.160193 (2016).
D’Cunha, J., Knight, E. Jr., Haas, A. L., Truitt, R. L. & Borden, E. C. Immunoregulatory properties of ISG15, an interferon-induced cytokine. Proc. Natl. Acad. Sci. U.S.A. 93, 211–215. https://doi.org/10.1073/pnas.93.1.211 (1996).
Sunkin, S. M. et al. Allen Brain Atlas: An integrated spatio-temporal portal for exploring the central nervous system. Nucleic Acids Res. 41, D996–D1008. https://doi.org/10.1093/nar/gks1042 (2013).
Duchen, L. W. Dystonia musculorum—An inherited disease of the nervous system in the mouse. Adv. Neurol. 14, 353–365 (1976).
Maia, J. N. et al. Electrophysiological study of the caudal nerve on developing rats. Acta Cir. Bras. 25, 144–147 (2010).
Miyoshi, T. & Goto, I. Serial in vivo determinations of nerve conduction velocity in rat tails. Physiological and patholgoical changes. Electroencephalogr. Clin. Neurophysiol. 35, 125–131 (1973).
Schmelzer, J. D. & Low, P. A. Electrophysiological studies on the effect of age on caudal nerve of the rat. Exp. Neurol. 96, 612–620 (1987).
Kirmani, B. F., Jacobowitz, D. M. & Namboodiri, M. A. Developmental increase of aspartoacylase in oligodendrocytes parallels CNS myelination. Brain Res. Dev. Brain Res. 140, 105–115. https://doi.org/10.1016/s0165-3806(02)00592-8 (2003).
Hershfield, J. R. et al. Aspartoacylase is a regulated nuclear-cytoplasmic enzyme. FASEB J. 20, 2139–2141. https://doi.org/10.1096/fj.05-5358fje (2006).
Monsma, P. C. & Brown, A. FluoroMyelin Red is a bright, photostable and non-toxic fluorescent stain for live imaging of myelin. J. Neurosci. Methods 209, 344–350. https://doi.org/10.1016/j.jneumeth.2012.06.015 (2012).
Mohan, S., Coto Hernandez, I., Selig, M. K., Shibata, S. & Jowett, N. Stain-free resolution of unmyelinated axons in transgenic mice using fluorescence microscopy. J. Neuropathol. Exp. Neurol. 78, 1178–1180. https://doi.org/10.1093/jnen/nlz099 (2019).
Garcia-Segura, L. M., Baetens, D., Roth, J., Norman, A. W. & Orci, L. Immunohistochemical mapping of calcium-binding protein immunoreactivity in the rat central nervous system. Brain Res. 296, 75–86. https://doi.org/10.1016/0006-8993(84)90512-2 (1984).
Middeldorp, J. & Hol, E. M. GFAP in health and disease. Prog. Neurobiol. 93, 421–443. https://doi.org/10.1016/j.pneurobio.2011.01.005 (2011).
Niu, J. et al. Modality-based organization of ascending somatosensory axons in the direct dorsal column pathway. J. Neurosci. 33, 17691–17709. https://doi.org/10.1523/JNEUROSCI.3429-13.2013 (2013).
Lancaster, E. et al. Myelinated axons fail to develop properly in a genetically authentic mouse model of Charcot-Marie-Tooth disease type 2E. Exp. Neurol. 308, 13–25. https://doi.org/10.1016/j.expneurol.2018.06.010 (2018).
Coleman, M. Axon degeneration mechanisms: Commonality amid diversity. Nat. Rev. Neurosci. 6, 889–898. https://doi.org/10.1038/nrn1788 (2005).
Pandey, R. et al. The autoimmune disorder susceptibility gene CLEC16A restrains NK cell function in YTS NK cell line and Clec16a knockout mice. Front. Immunol. 10, 68. https://doi.org/10.3389/fimmu.2019.00068 (2019).
Liu, Y. et al. TRIM25 promotes the cell survival and growth of hepatocellular carcinoma through targeting Keap1-Nrf2 pathway. Nat. Commun. 11, 348. https://doi.org/10.1038/s41467-019-14190-2 (2020).
Harris, B. S., Ward-Bailey, P. F., Bergstrom, D. E., Bronson, R. T. & Donahue, L. R. Curvy tail: a new skeletal mutation that maps to Chromosome 16. http://www.informatics.jax.org/reference/J:172931 (2011).
Fairfield, H. et al. Exome sequencing reveals pathogenic mutations in 91 strains of mice with Mendelian disorders. Genome Res. 25, 948–957. https://doi.org/10.1101/gr.186882.114 (2015).
Bruce, K. D., Zsombok, A. & Eckel, R. H. Lipid processing in the brain: A key regulator of systemic metabolism. Front. Endocrinol. (Lausanne) 8, 60. https://doi.org/10.3389/fendo.2017.00060 (2017).
Liu, L. et al. Glial lipid droplets and ROS induced by mitochondrial defects promote neurodegeneration. Cell 160, 177–190. https://doi.org/10.1016/j.cell.2014.12.019 (2015).
Huang, M. & Verbeek, D. S. Why do so many genetic insults lead to Purkinje cell degeneration and spinocerebellar ataxia?. Neurosci. Lett. https://doi.org/10.1016/j.neulet.2018.02.004 (2018).
Synofzik, M., Puccio, H., Mochel, F. & Schols, L. Autosomal recessive cerebellar ataxias: Paving the way toward targeted molecular therapies. Neuron 101, 560–583. https://doi.org/10.1016/j.neuron.2019.01.049 (2019).
Mitoma, H. et al. Consensus paper: Neuroimmune mechanisms of cerebellar ataxias. Cerebellum 15, 213–232. https://doi.org/10.1007/s12311-015-0664-x (2016).
Zhou, X. et al. Deletion of PIK3C3/Vps34 in sensory neurons causes rapid neurodegeneration by disrupting the endosomal but not the autophagic pathway. Proc. Natl. Acad. Sci. U.S.A. 107, 9424–9429. https://doi.org/10.1073/pnas.0914725107 (2010).
Ferrier, A. et al. Disruption in the autophagic process underlies the sensory neuropathy in dystonia musculorum mice. Autophagy 11, 1025–1036. https://doi.org/10.1080/15548627.2015.1052207 (2015).
Sullivan, R., Yau, W. Y., O’Connor, E. & Houlden, H. Spinocerebellar ataxia: An update. J. Neurol. 266, 533–544. https://doi.org/10.1007/s00415-018-9076-4 (2019).
Amaral, T. N., Peres, F. A., Lapa, A. T., Marques-Neto, J. F. & Appenzeller, S. Neurologic involvement in scleroderma: A systematic review. Semin. Arthritis Rheum. 43, 335–347. https://doi.org/10.1016/j.semarthrit.2013.05.002 (2013).
Hadjivassiliou, M. et al. Neurological dysfunction in coeliac disease and non-coeliac gluten sensitivity. Am. J. Gastroenterol. 111, 561–567. https://doi.org/10.1038/ajg.2015.434 (2016).
Kutzelnigg, A. et al. Widespread demyelination in the cerebellar cortex in multiple sclerosis. Brain Pathol. 17, 38–44. https://doi.org/10.1111/j.1750-3639.2006.00041.x (2007).
Luostarinen, L., Himanen, S. L., Luostarinen, M., Collin, P. & Pirttila, T. Neuromuscular and sensory disturbances in patients with well treated coeliac disease. J. Neurol. Neurosurg. Psychiatry 74, 490–494. https://doi.org/10.1136/jnnp.74.4.490 (2003).
Oomatia, A., Fang, H., Petri, M. & Birnbaum, J. Peripheral neuropathies in systemic lupus erythematosus: Clinical features, disease associations, and immunologic characteristics evaluated over a twenty-five-year study period. Arthritis Rheumatol. 66, 1000–1009. https://doi.org/10.1002/art.38302 (2014).
Redondo, J. et al. Purkinje cell pathology and loss in multiple sclerosis cerebellum. Brain Pathol. 25, 692–700. https://doi.org/10.1111/bpa.12230 (2015).
Selim, M. & Drachman, D. A. Ataxia associated with Hashimoto’s disease: Progressive non-familial adult onset cerebellar degeneration with autoimmune thyroiditis. J. Neurol. Neurosurg. Psychiatry 71, 81–87. https://doi.org/10.1136/jnnp.71.1.81 (2001).
Shakkottai, V. G. et al. Current opinions and areas of consensus on the role of the cerebellum in dystonia. Cerebellum 16, 577–594. https://doi.org/10.1007/s12311-016-0825-6 (2017).
Ozato, K., Shin, D. M., Chang, T. H. & Morse, H. C. 3rd. TRIM family proteins and their emerging roles in innate immunity. Nat. Rev. Immunol. 8, 849–860. https://doi.org/10.1038/nri2413 (2008).
Kimura, T. et al. TRIM-mediated precision autophagy targets cytoplasmic regulators of innate immunity. J. Cell Biol. 210, 973–989. https://doi.org/10.1083/jcb.201503023 (2015).
Lee, J. M. et al. The E3 ubiquitin ligase TRIM25 regulates adipocyte differentiation via proteasome-mediated degradation of PPARgamma. Exp. Mol. Med. 50, 1–11. https://doi.org/10.1038/s12276-018-0162-6 (2018).
Zhang, P. et al. TRIM25 has a dual function in the p53/Mdm2 circuit. Oncogene 34, 5729–5738. https://doi.org/10.1038/onc.2015.21 (2015).
Zhang, L., Afolabi, L. O., Wan, X., Li, Y. & Chen, L. Emerging roles of tripartite motif-containing family proteins (TRIMs) in eliminating misfolded proteins. Front. Cell Dev. Biol. 8, 802. https://doi.org/10.3389/fcell.2020.00802 (2020).
Davies, A. J. et al. Natural killer cells degenerate intact sensory afferents following nerve injury. Cell 176, 716-728 e718. https://doi.org/10.1016/j.cell.2018.12.022 (2019).
Schwartzenburg, J., Juncker, M., Reed, R. & Desai, S. Increased ISGylation in cases of TBI-exposed ALS veterans. J. Neuropathol. Exp. Neurol. 78, 209–218. https://doi.org/10.1093/jnen/nly129 (2019).
Wang, R., Yang, B. & Zhang, D. Activation of interferon signaling pathways in spinal cord astrocytes from an ALS mouse model. Glia 59, 946–958. https://doi.org/10.1002/glia.21167 (2011).
Burke, J. R. et al. Autoimmune pathways in mice and humans are blocked by pharmacological stabilization of the TYK2 pseudokinase domain. Sci. Transl. Med. https://doi.org/10.1126/scitranslmed.aaw1736 (2019).
Maday, S. Mechanisms of neuronal homeostasis: Autophagy in the axon. Brain Res. 1649, 143–150. https://doi.org/10.1016/j.brainres.2016.03.047 (2016).
Hara, T. et al. Suppression of basal autophagy in neural cells causes neurodegenerative disease in mice. Nature 441, 885–889. https://doi.org/10.1038/nature04724 (2006).
Ara, J. et al. Bone morphogenetic proteins 4, 6, and 7 are up-regulated in mouse spinal cord during experimental autoimmune encephalomyelitis. J. Neurosci. Res. 86, 125–135 (2008).
Feigenson, K., Reid, M., See, J., Crenshaw, E. B. II. & Grinspan, J. B. Wnt signaling is sufficient to perturb oligodendrocyte maturation. Mol. Cell. Neurosci. 42, 255–265 (2009).
Hendrickx, D. A. E., van Eden, C. G., Schuurman, K. G., Hamann, J. & Huitinga, I. Staining of HLA-DR, Iba1 and CD68 in human microglia reveals partially overlapping expression depending on cellular morphology and pathology. J. Neuroimmunol. 309, 12–22. https://doi.org/10.1016/j.jneuroim.2017.04.007 (2017).
Hoogland, I. C., Houbolt, C., van Westerloo, D. J., van Gool, W. A. & van de Beek, D. Systemic inflammation and microglial activation: Systematic review of animal experiments. J. Neuroinflammation 12, 114. https://doi.org/10.1186/s12974-015-0332-6 (2015).
Potter, K. A. et al. Central nervous system dysfunction in a mouse model of FA2H deficiency. Glia 59, 1009–1021. https://doi.org/10.1002/glia.21172 (2011).
Greco, T. M., Seeholzer, S. H., Mak, A., Spruce, L. & Ischiropoulos, H. Quantitative mass spectrometry-based proteomics reveals the dynamic range of primary mouse astrocyte protein secretion. J. Proteome Res. 9, 2764–2774. https://doi.org/10.1021/pr100134n (2010).
Acknowledgements
This work was supported by an Institute Development Award to the Center for Applied Genomics from the Children’s Hospital of Philadelphia and by the endowed chair in genomic research to Dr. Hakonarson.
Author information
Authors and Affiliations
Contributions
H.S.H., M.B., R.P., S.S.S., J.G., and H.H. conceptualized the experiments; Methods were decided by H.S.H., R.P., S.S.S., J.G., and M.B.; Experimental investigations were completed by B.P.S., M.B., M.R., E.L., J.L., D.H., L.R., J.G., H.S.H. and W.W.M.; H.S.H. and S.S.S. wrote the original draft of the manuscript. Writing, review and editing of the manuscript was completed by H.S.H., S.S.S., J.G., H.H., E.L., R.P., and M.B.; Funding acquisition and resources were supported by H.H., S.S.S., and J.G.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hain, H.S., Pandey, R., Bakay, M. et al. Inducible knockout of Clec16a in mice results in sensory neurodegeneration. Sci Rep 11, 9319 (2021). https://doi.org/10.1038/s41598-021-88895-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-88895-0
This article is cited by
-
CLEC16A interacts with retromer and TRIM27, and its loss impairs endosomal trafficking and neurodevelopment
Human Genetics (2023)
-
The genetics of autoimmune Addison disease: past, present and future
Nature Reviews Endocrinology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.