Abstract
Korla pear (Pyrus sinkiangensis Yü) is a landrace selected from a hybrid pear species in the Xinjiang Autonomous Region in China. In recent years, pericarp roughening has been one of the major factors that adversely affects fruit quality. Compared with regular fruits, rough-skin fruits have a greater stone cell content. Stone cells compose sclerenchyma tissue that is formed by secondary thickening of parenchyma cell walls. In this work, we determined the main components of stone cells by isolating them from the pulp of rough-skin fruits at the ripening stage. Stone cell staining and apoptosis detection were then performed on fruit samples that were collected at three different developmental stages (20, 50 and 80 days after flowering (DAF)) representing the prime, late and stationary stages of stone cell differentiation, respectively. The same batches of samples were used for parallel transcriptomic and proteomic analysis to identify candidate genes and proteins that are related to SCW biogenesis in Korla pear fruits. The results showed that stone cells are mainly composed of cellulose (52%), hemicellulose (23%), lignin (20%) and a small amount of polysaccharides (3%). The periods of stone cell differentiation and cell apoptosis were synchronous and primarily occurred from 0 to 50 DAF. The stone cell components increased abundantly at 20 DAF but then decreased gradually. A total of 24,268 differentially expressed genes (DEGs) and 1011 differentially accumulated proteins (DAPs) were identified from the transcriptomic and proteomic data, respectively. We screened the DEGs and DAPs that were enriched in SCW-related pathways, including those associated with lignin biosynthesis (94 DEGs and 31 DAPs), cellulose and xylan biosynthesis (46 DEGs and 18 DAPs), S-adenosylmethionine (SAM) metabolic processes (10 DEGs and 3 DAPs), apoplastic ROS production (16 DEGs and 2 DAPs), and cell death (14 DEGs and 6 DAPs). Among the identified DEGs and DAPs, 63 significantly changed at both the transcript and protein levels during the experimental periods. In addition, the majority of these identified genes and proteins were expressed the most at the prime stage of stone cell differentiation, but their levels gradually decreased at the later stages.
Similar content being viewed by others
Introduction
Korla pear (Pyrus sinkiangensis Yü) is a native species of the Xinjiang Autonomous Region in China. It is known for its aroma, juicy flesh, and crisp texture. Increased stone cell content, which leads to the formation of rough-skin fruits, is one of the most critical factors that result in decreased fruit quality. In regular fruits, stone cells are nearly invisible, but in rough-skin fruits, the high stone cell content imparts a very gritty texture to the fruit pulp. The rough skins are invisible phenotypically on the early fruits and gradually become visible in the later stage (from late July to mid-August). Hence, it is difficult to determine the time when the pericarp roughening is actually initiated. This makes it challenging to study the growth mechanism of rough-skin formation on pear fruits. Therefore, in order to find out the stress factors that induce the formation of rough-skinned fruits, we must first understand the underlying physiological and molecular mechanisms of stone cell differentiation.
Previous studies have shown that the formation of stone cells is closely related to the synthesis, transfer, and deposition of lignin1,2. Apart from tracheary elements (TEs) and fibers, sclereids (stone cells) are also formed by the secondary thickening of cell walls3. Cellulose, hemicellulose, and lignin constitute the main components of secondary walls, and their proportions may vary among different plant species. While we understand a great deal about the genes involved in lignin synthesis and deposition during stone cell differentiation in pear fruits4,5,6,7,8,9,10, we know surprisingly little about the underlying physiological and molecular mechanisms of stone cell differentiation because cellulose and hemicellulose account for the largest percentage of secondary cell wall (SCW) biomass. Therefore, the formation of stone cells cannot be fully explained merely by the process of lignin deposition. After their biosynthesis, lignin monomers must be transported to the cell wall where they are oxidized for polymerization. Reactive oxygen species (ROS) play a vital role in this process11,12,13. In our previous work, we found that apoptosis occurs along with the differentiation of stone cells during fruit development, and the apoptosis period overlaps with the period of stone cell formation and ROS accumulation14. ROS is the main stress factor leading to apoptosis. Therefore, we hypothesized that the formation of stone cells might essentially be linked to the process of apoptosis triggered by ROS. H2O2 induces cell wall thickening by polymer cross-linking, leading to growth inhibition of some parenchyma cells. As a result, some parenchyma cells are forced to differentiate. However, few studies have linked the formation of stone cells to the synthesis of cell wall polymers and cross-linkages among them. To resolve this scientific problem and theoretical inference, in this study, parallel analyses of the transcriptome and proteome of Korla pear fruits at three developmental stages were carried out to identify essential regulators and pathways involved in stone cell formation. The results gained through this study will help us understand the mechanisms underlying stone cell formation in pear fruits.
Results
Morphological and physiological analysis
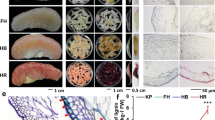
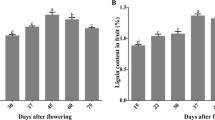
Stone cells were isolated from rough-skin fruits at the ripening stage, and the main components were analyzed. The results revealed that the stone cells are mainly composed of cellulose (52%), hemicellulose (23%), lignin (20%), and a small amount of polysaccharides (3%) (Fig. 1D). The variation trends of these polymers, stone cell staining and cell apoptosis were then observed at three important time points of stone cell differentiation (20 DAF, 50 DAF and 80 DAF, representing the prime, late and stationary stages of stone cell differentiation, respectively). It is evident from the data that stone cell formation primarily occurred from 0 to 50 DAF, and the apoptotic period overlapped with this period (Fig. 1A, B). All the stone cell components were abundant at 20 DAF, after which the levels decreased gradually (Fig. 1C), and the trends of these components were consistent with trends of stone cell contents during fruit development. These results also indicated that the formation of stone cells could not be fully explained by lignin deposition in the secondary walls.
Morphological and physiological differences in Korla pear fruit pulp during the experimental time course. (A) Safranin staining of stone cells. Magnification: 10 × 4, Scale: 100 μm. (B) TUNEL detection of pear fruit cell apoptosis during the experimental time course. Magnification: 10 × 4, Scale: 100 μm. (C) Variation in cell wall polymers during fruit development. (D) Composition of stone cells. The bars represent the means ± SDs (n = 3).
Overview of transcriptomic and proteomic data
To identify essential regulators and pathways involved in stone cell formation in Korla pear fruit, transcriptomic and proteomic analyses were performed via RNA sequencing (RNA-seq) and tandem mass tag (TMT) techniques. A total of 42,893 transcripts and 7904 proteins were identified, of which 24,268 genes and 1011 proteins showed differential expression during the development time course. Correlation analysis showed that approximately 13% (1010/7904) of the proteins correlated with DEGs, including 700 DAPs. Of the 1011 DAPs, 311 DAPs (30.8%) were not correlated with DEGs (Fig. 2A). Stone cells are formed by secondary thickening of parenchyma cell walls, and SCWs mainly consist of cellulose, hemicellulose, and lignin. Therefore, in this work, we closely focused on the DEGs and DAPs that were enriched in SCW-related pathways, including those associated with lignin biosynthesis (94 genes and 31 proteins), cellulose and xylan biosynthesis (46 genes and 18 proteins), S-adenosylmethionine (SAM) metabolic processes (10 genes and 3 proteins), apoplastic ROS production (16 genes and 2 proteins), and cell death (14 genes and 6 proteins) (Fig. 2B). Here, we list only the genes and proteins with greater differences in expression during the critical stage of stone cell differentiation (Figs. 3, 4, 5); information on all DEGs and DAPs in these pathways can be found in Supplementary Data S1-S3. Fifteen genes were randomly selected from among the DEGs for qPCR to validate the RNA-seq results. As shown in Fig. S1, the validity of the RNA-seq data was demonstrated by the high agreement between the RNA-seq data and qPCR data.
Other genes and proteins involved in apoplastic ROS production, SAM metabolic processes and PCD. The heat maps were produced using standardized figures that were transformed to a value between 0.0 and 1.0 by the min–max normalization method. In the proteomic data, the value 0 for all three periods indicates that no corresponding protein was detected.
DEGs and DAPs involved in lignin biosynthesis
The omics data revealed 94 DEGs and 31 correlated DAPs enriched in the lignin synthesis pathway in Korla pear pulp (Data S1). These genes and proteins encode upstream enzymes of the phenylpropanoid pathway, including phenylalanine ammonialyase (PAL), cinnamate-4-hydroxylase (C4H), p-coumarate 3-hydroxylase (C3′H), 4-coumarate: CoA ligase (4CL), p-hydroxycinnamoyl-CoA: quinate shikimate p-hydroxycinnamoyl transferase (HCT), caffeoyl shikimate esterase (CSE), and caffeoyl-CoA O-methyltransferase (CCoAOMT); enzymes in the monolignol-specific pathway, including cinnamoyl-CoA reductase (CCR), caffeic acid-5-O-methyltransferase (COMT), ferulate 5-hydroxylase (F5H), and cinnamoyl alcohol dehydrogenase (CAD); and enzymes involved in lignin polymerization (peroxidase (PRX) and laccase (LAC)). The majority of these genes showed peak expression at 20 DAF, after which the expression gradually decreased from 20 to 80 DAF, showing trends similar to the trend of lignin content during fruit development. Some transcripts, however, showed no differential expression from 50 to 80 DAF, indicating that lignin accumulation mainly occurred at the early stage of fruit development. Interestingly, the expression patterns of some transcripts encoding PAL (one transcript), C4H (two transcripts), 4CL (two transcripts), HCT (one transcript), CCR (five transcripts), CAD (two transcripts), CSE (two transcripts) and PRX (five transcripts) showed the opposite trend, indicating that they may be involved in the biosynthesis of other secondary metabolites involved in the phenylpropanoid pathway. However, the expression levels of these genes were almost negligible compared with those of the genes positively related to lignin content.
DEGs and DAPs associated with cellulose biosynthesis
Cellulose is the most abundant component of SCWs. Twenty-nine DEGs in the transcriptome profile were assigned to cellulose biosynthesis, while 7 DAPs were annotated to the same process in the proteomic data (Fig. 4). UDP-glucose is the direct precursor of cellulose. Therefore, pathways that lead to UDP-glucose production can potentially have an impact on cellulose biosynthesis. UDP-glucose pyrophosphorylase (UGP) and sucrose synthase (SuSy) are the main enzymes that have been shown to produce UDP-glucose in plants15. Although we did not find any differentially expressed genes encoding UGPs, four DEGs and one DAP were found to encode sucrose synthase (SuSy), and their expression pattern was consistent with the trend of cellulose content, indicating that SuSy is tightly associated with cellulose biosynthesis. The hexose phosphate pool can also affect the UDP-glucose pool size. The biosynthesis of sucrose is inhibited by the feedback regulation of fructose, and fructokinase (FRK) phosphorylates fructose to produce fructose-6-phosphate. According to our omics data, 4 DEGs and 2 DAPs that encode FRK were selected. The expression of all of these genes and proteins was similar to that of SuSy during the experimental period, indicating that the synthesis of cellulose may be affected by FRK through reducing intracellular fructose pools, which can modulate SuSy activity and UDP-glucose production.
Cellulose synthase (CesA) uses UDP-glucose as a substrate for the biosynthesis of cellulose chains. In our study, we obtained a total of 21 functional genes and 4 proteins related to CesA enzymes, among which the expression patterns of the homologous genes of Arabidopsis CesA4 (2 transcripts), CesA7 (2 transcripts), and CesA8 (2 transcripts) were the same as those of the lignin synthesis-related genes, following trends of lignin and stone cell contents during the experimental time course (Fig. 1). In addition, we screened 2 DEGs and 1 DAP annotated as COBRA-like 4 (COBL4) proteins. In Arabidopsis, COBL4 is required for cellulose synthesis in secondary cell walls. The expression trends of these genes were similar to those of CesAs associated with secondary wall biosynthesis, implying that this protein is most likely involved in the assembly of cellulose synthase complexes. However, homologous genes of Arabidopsis CesA1 (2 transcripts), CesA2 (2 transcripts), CesA3 (2 transcripts) and most cellulose synthase-like proteins (CSLs) (5 transcripts) showed the opposite expression trend.
DEGs and DAPs associated with xylan biosynthesis
UDP-xylose is a nucleotide sugar required for xylan backbone elongation. According to our data, 25 DEGs and 13 DAPs were identified as xylan biosynthesis-related genes and proteins. All of these genes and proteins showed the same expression trends as the genes related to cellulose and lignin synthesis (Fig. 4). Among them, one gene with no associated protein identified encoded UDP-glucose dehydrogenase (UGD), which converts UDP-glucose to UDP-glucuronic acid; 6 genes and 5 associated proteins encoded UDP-xylose synthase (UXS), which is responsible for the conversion of UDP-glucuronic acid to UDP-xylose; 4 genes and 4 associated proteins encode xylosyltransferases (XylTs) needed for elongation of the xylan backbone; 4 genes with no associated proteins encoded glucuronosyltransferase (GUX), which is essential for the side chain addition; 6 genes and 3 associated proteins encoded glucuronoxylan 4-O-methyltransferase (GXMT) required for the glucuronic acid (GlcA) side chain methylation; and 3 genes (IRX7) with one associated protein required for the formation of the xylan reducing end sequence (XRES), which is speculated to be closely related to the initiation or termination of glucuronoxylan biosynthesis, were found16 (Fig. 4 and Data S2). As shown in Fig. 4, the expression levels of genes related to UDP-xylose formation (103,928,376, 103,928,424, 103,940,426, 103,948,402, 103,930,040 and 103,944,949)), elongation of the xylan backbone (103,947,711, 103,950,395, 103,942,677, etc.), and side chain methylation (GXM) were significantly higher than those of other xylan-related genes, implying their essential roles in xylan or glucuronoxylan synthesis. Most of the SCW- genes and xylan-related genes were preferentially expressed at 20 DAF, while primary wall-related genes were relatively abundant in the day-80 samples (Data S2). Taken together, these results suggested that the secondary thickening of parenchymal cell walls mainly occurred at the early stage of fruit development.
Other secondary cell wall-related genes and proteins
Genes related to apoplastic ROS production (16 genes and 2 proteins), SAM metabolic processes (10 genes and 3 proteins), and cell death (12 genes and 6 proteins) were well represented among the candidate genes (Fig. 5 and Data S3). Apoplastic ROS can be used as oxidants in cell wall cross-linking and are required for lignin polymerization. ROS, which constitute a major stress factor, also play an essential role in apoptosis induction. Genes and proteins related to apoplastic ROS preferentially accumulated at 20 DAF, with markedly lower expression at later stages. For example, 3 genes and 1 associated protein annotated as respiratory burst oxidase homolog (RBOH) showed the highest expression at the early stage, after which the expression rapidly decreased during fruit development. Increased accumulation of transcripts encoding superoxide dismutase (SOD), polyamine oxidase (PAO), PRXs and germin-like proteins (GLPs) was also observed in early fruits.
The expression patterns of genes encoding enzymes related to the SAM metabolic process were similar to those of ROS-related genes (Fig. 5 and Data S3). Polyamines are important substrates for apoplastic H2O2 production, and polyamine generation also requires SAM to provide carboxyl propyl groups. Moreover, methyl groups required for the synthesis of cell wall polymers are also provided by SAM. Numerous genes related to programmed cell death (PCD) were also expressed in pear pulp and showed their highest expression at 20 DAF, which is consistent with the trends of genes that lead to ROS generation during fruit development (Fig. 5 and Data S3).
Discussion
In pear fruits, some parenchyma cells undergo secondary thickening during fruit development, and lignin is deposited in the thickened cell walls to form stone cells17,18,19. Our results showed that the stone cell formation primarily occurred in the early fruits. Similar results have been reported for other pear varieties1,2,9. Cellulose, hemicellulose and lignin are the main components of secondary walls, and their proportions can vary among different plant species. The secondary walls of some special tissues consist mainly of one or two polymers. For example, cotton fibers are abundant in cellulose (> 90%), and phloem fibers, on the other hand, contain no lignin4. We found that pear stone cells were mainly composed of cellulose (52%), hemicellulose (23%), lignin (20%) and a small amount of polysaccharides (3%). Our results are also supported by several previous reports 3,20, indicating that stone cell formation is more likely to result from the differentiation of distinct cell types. In support of this, we found that many secondary cell wall-related genes and proteins (53 genes and 20 proteins) showed peak expression in early fruits, which was consistent with the change trends of stone cells and lignin.
The lignification process varies among different cell types; for instance, in TEs, cell apoptosis is a prerequisite for cell wall lignification, in which enzymes and/or substrates needed for cell wall lignification are provided by neighboring cells12,21,22. In contrast to that of TEs, lignification of fiber cells is carried out by cell-autonomous processes, in which cell apoptosis and cell wall lignification occur simultaneously21,22. The lignin composition varies between these cell types. In TEs, lignin is primarily composed of G-units, while S-lignin is abundant in the secondary walls of fibers23,24. It is a part of the folklore of plant biology that, like fibers, stone cells comprise sclerenchyma cells. Supporting this theory, in our previous work, we found that sinapyl alcohol, which gives rise to S-units, is the dominant lignin monomer in Korla pear fruit pulp during the whole period of fruit development17. These observations suggest that the differentiation of stone cells, which are types of sclerenchyma cells, must begin with the formation of a framework of secondary walls that are primarily composed of cellulose and hemicellulose.
Cellulose is the most abundant component of secondary cell walls. We found that the cellulose content accounted for more than half of the stone cell biomass, indicating that cellulose composes the skeleton of secondary walls of stone cells. Cellulose synthase (CesA) uses UDP-glucose as a substrate for the biosynthesis of cellulose chains25,26. Therefore, pathways that lead to UDP-glucose production can potentially have an impact on cellulose biosynthesis. SuSy and UGPs are the main enzymes that produce UDP-glucose in plants15. In hybrid aspen, inhibition of SuSy was shown to reduce carbon allocation to all secondary cell wall polymers; however, its overexpression increased only cellulose deposition in hybrid poplar26,27. In accordance with these findings, in our work, several genes (103950289, 103940505, 103940043 and 103964096) encoding SuSy showed peak expression at 20 DAF, after which their expression decreased more than 20-fold at 80 DAF. The change trends were consistent with those of the cellulose content, indicating that SuSy is tightly associated with the processes o cellulose biosynthesis. In our previous research, we found that SuSy showed higher enzymatic activity in early fruits, and SuSy activity decreased first but then increased gradually 60 DAF. Accordingly, the sucrose abundance increased slowly at the later stage and began to increase at 60 DAF28, suggesting that, in early fruits, sucrose is mainly used for cellulose synthesis. The hexose-phosphate pool can also affect the UDP-glucose pool size29. The biosynthesis of sucrose is inhibited by feedback regulation of fructose, and FRK phosphorylates fructose to produce fructose-6-phosphate29. We identified 4 DEGs and 2 DAPs that encode FRK and whose expression was similar to that of the SuSy gene during the experimental period, indicating that the synthesis of cellulose may be affected by FRK by reducing intracellular fructose pools, which can modulate SuSy activity and UDP-glucose production15,29,30.
In our study, CESA genes showed differential expression throughout the experimental time course. As expected, homologs of AtCESAs that encode CESAs in Arabidopsis and that are essential for secondary cell wall synthesis (AtCESA4, AtCESA7 and AtCESA8)31,32 showed peak expression in early fruits, and the expression levels gradually declined at the later stage. In contrast, homologous genes of AtCESAs required for primary wall cellulose synthesis (AtCESA1, AtCESA2, and AtCESA3) in Arabidopsis31,32 showed the opposite expression trends. In addition, associated proteins of AtCESA7 and AtCESA8 were also found to be differentially expressed during the experimental time course and showed the same expression patterns as their corresponding genes did. These genes were reported to constitute secondary wall CesA complexes33,34,35. In addition, studies have shown that cesa4, cesa7, and cesa8 mutants have severe cellulose defects and reduced secondary wall thickness31,32. The same results were reported in rice, Brachypodium and Populus34,35,36, implying that CESA genes are required for secondary cell wall cellulose synthesis during the stone cell differentiation process.
Hemicellulose is another main component of pear stone cells, accounting for 23% of stone cell biomass. Xylan, a major component of hemicellulose, is essential for secondary wall strength 37. UDP-xylose is a nucleotide sugar required for xylan backbone elongation. Serial actions of UGD and UXS form UDP-xylose. Genes encoding these two enzymes showed relatively high expression in early fruits. The expression levels of UXSs were more than 100-fold higher than those of UGDs, indicating that UXS is the primary enzyme responsible for UDP-xylose. This is supported by the evidence that antisense downregulation of UXS resulted in fewer xylose-containing polymers and showed altered organization and reduced xylan phenotypes38,39.
Several enzymes have been identified to be involved in the elongation of the xylan backbone and its side chain addition. Genes (IRX10, IRX15, IRX15-L) encoding these enzymes and their associated proteins responsible for the synthesis of secondary cell wall xylan40,41 showed peak expression in early fruits, after which their expression decreased at the later stage. In Arabidopsis, all irx mutants have severely decreased xylan contents, chain length, and secondary wall thickness40,41,42,43,44. Interestingly, there is no significant reduction in xylan: XylT activity in the irx15 and irx15-L mutants, and the irx15 irx15-L double mutant replaced GlcA with the meGlcA xylan side chain. Therefore, IRX15 and IRX15-L define a new class of genes involved in xylan biosynthesis, but their function remains unknown45,46,47.
Another group of genes involved in the addition of GlcA side chains and side chain methylation, such as GUXs and GXMTs, showed peak expression in early fruits but were expressed at low levels at the later stages. These results are supported by evidence that GUX and GXMT are preferentially expressed in secondary thickening cell types, and gux and gxmt mutants are also found with a lack of GlcA or methylated GlcA in xylan side chains and reduced secondary wall thickness48,49,50. IRX7 is required for the biosynthesis of XRES51,52,53. In accordance with this, we screened three DEGs (and one DAP) that were annotated as probable IRX7s and whose expression was substantially upregulated at the early stage and downregulated at the later stage, indicating that these genes are essential for xylan synthesis. Notably, mutations in XRES-related genes does affect either XylT or GlcAT enzyme activity, despite the loss of XRES45,46.
Lignin composes approximately 20% of stone cell biomass (Fig. 1). In previous studies, lignin-related genes were associated with the development of xylem and fibers in stems54,55. Similarly, in the present study, more than 90 genes and 31 proteins related to lignin biosynthesis were differentially expressed during pear fruit development (Data S1). The majority of these genes showed peak expression in early fruits and were positively correlated with lignin content, but some transcripts showed the opposite expression trends. However, the expression levels of the genes that were positively regulated with lignin content were overall much higher than those of negatively regulated genes, suggesting that these genes are essential for lignin biosynthesis during stone cell differentiation in pear fruits. Lignin-related genes have been studied extensively in different pear varieties1,2,9,10,17,56. Therefore, the role of these genes during lignin accumulation in pear pulp is not discussed in detail here. Instead, we mainly focused on the genes involved in lignin polymerization. After their biosynthesis, lignin monomers must be transported to the cell walls, where they are oxidized for polymerization57,58,59,60. Laccase and peroxidase are responsible for the oxidation of monolignols. In this work, the expression profiles of all LAC genes and the majority of PRX genes were similar to those of monolignol biosynthesis-related genes. Although some PRX genes showed the opposite trend, their expression levels were nearly negligible during the entire period compared to those of upregulated PRXs at the early stage. LACs are reported to be coregulated together with monolignol synthesis-related genes and secondary cell wall formation-related genes46,61. We found 6 of 17 LAC genes, three transcripts of LAC4-L, two transcripts of LAC17-L and one transcript of LAC2-L whose expression was 10- to 150-fold higher than that of the other transcripts, indicating that these are the core LACs for the formation of monolignol radicals. Studies have shown that the lac4 lac17 double mutant presents a phenotype involving reduced lignin content, and the lac4 lac11 lac17 triple mutant presents a phenotype involving severe growth defects and a loss of lignification of root vessels61,62. In model plant species, downregulating or silencing PRXs (PRX2, PRX3, PRX25, PRX60, PRX71, PRX72, etc.) resulted in reduced lignin accumulation and altered lignin composition60,61,62,63. Consistent with these findings, the results of our study showed that six transcripts of PRXs (two PRX12-L, two PRX42-L, one PRX25 and one PRX72-L) were expressed markedly higher (up to several hundred times) than other transcripts were throughout the experiment. The abundance of the associated proteins PRX12-L and PRX72-L was also 10- to 100-fold higher than that of other associated proteins, suggesting their importance for lignin polymerization in Korla pear.
In addition to lignin monomers, peroxidases and laccases require H2O2 and O2, respectively, to form monolignol radicals. Our study showed that ROS-related genes and their encoded proteins were more abundantly expressed at the early stage and downregulated at the later stage. Specific ROS, such as H2O2, can be used as oxidants in cell wall cross-linking or as signaling molecules for controlling various biological processes64,65. It has been reported that scavenging H2O2 results in a reduction in lignification in the Casparian strip and TEs21,66,67. Together, these findings indicate that apoplastic ROS play an important role in the lignification process. H2O2 is also a key modulator of PCD, which is essential for the differentiation of TEs, fibers, etc.68 The present study also showed several highly expressed cell death-related genes and proteins (AED3-like, MC1-like, ACD11-like), and their expression patterns were similar to those of ROS-associated genes.
In conclusion, the differentiation of stone cells primarily occurred during cell division period of pear fruits, and no longer differentiated in the later stage of fruit development under the normal growth condition. Our results indicate that cellulose, hemicellulose, and lignin were the main components of stone cells. In addition to lignin synthetic genes, many other genes related to secondary wall development, cell apoptosis and ROS production also highly expressed during the critical period of stone cell differentiation, suggesting that stone cell formation in pear fruits required synergistic regulation of lignin synthetic genes and other genes involved in other SCW-related pathways.
Methods
Plant materials and sampling period
Fruits of Korla pear were collected from 20-year-old pear trees grown in an orchard in Korla (Xinjiang, China). The fruits were sampled at 20, 50 and 80 days after flowering (DAF), respectively. Stone cells are commonly present in all pear varieties. The differentiation of pear stone cells primarily occurs during cell division period of pear fruit development which usually takes place within 50 DAF, and after then, the stone cells are no longer differentiated in the later stage of fruit development under the normal growth condition. Therefore, the selected experimental periods in this study can represent the prime, late and stationary stages of stone cell differentiation in Korla pear fruits, respectively. At the early stages (20 DAF and 50 DAF), each biological replicate consisted of fifteen individual fruit samples collected from five independent trees, and ten fruits were collected for each biological replicate at the later stage. The fruit skins were peeled with a peeler, and fruit pulps were sampled, frozen in liquid nitrogen, ground into a powder, and stored at -80 °C until further analysis. The same batch of samples was used for transcriptomic, proteomic, physiological and biological analyses.
Stone cell separation and structural carbohydrate detection
The rough-skin phenotype is not apparent for early fruits and gradually appears at the later stage of fruit development (late July to early August). Thus, stone cells were separated from 5 kg of rough-skin fruits at 100 DAF (August 15) using the method described by Brahem et al. (2017)3. The stone cells were collected, oven dried, and ground into a uniform powder. The ground powder was used for analysis of secondary cell wall components. The lignin content was determined by the Klason method69, the hemicellulose content was determined via 2% hydrochloric acid hydrolysis with the DNS method70, and the cellulose content was determined by the anthrone-sulfuric acid colorimetry method70. The polysaccharide content was tested by the phenol–sulfuric acid method71.
Stone cell staining and apoptosis detection
Fruit pulps were cut into appropriate sizes and fixed with FAA fixative solution (formalin:glacial acetic acid:ethanol (90%) = 5:5:90, v/v). The preparation and tissue staining of paraffin sections were performed using the phloroglucinol method described by Tao et al. (2009)2. The same paraffin sections were used for apoptosis detection. Apoptosis was detected by TUNEL (terminal deoxynucleotidyl transferase-mediated dUTP nick-end labeling) assay kits from Shanghai Beyotime (www.beyotime.com). Briefly, paraffin sections were heated for 2 h at 65 °C in an oven and then dewaxed in xylene for 10 min. The tissues were then hydrated sequentially in a 100%, 100%, 95%, and 80% ethanol series and purified water for 5 min. The slices were then transferred to a wet box, 50 µg/ml proteinase K working solution was added to each sample, and the reaction was carried out at 37 °C for 30 min. The splices were then washed thoroughly with phosphate buffer solution (PBS; pH 7.2–7.4) 3 times, each for 5 min. During this step, protease K was washed away; otherwise, it strongly interfered with the subsequent marking reaction. The PBS around the tissue was absorbed with blotting paper, and a sufficient amount of TUNEL test solution was added to each glass slide. The slides were then incubated at 45 °C in the dark for 2 h, after which they were washed thoroughly with PBS 3 times for 5 min each. The PBS around the tissue was absorbed with blotting paper. Apoptosis was observed under a fluorescence microscope after sealing the slices with antifluorescent quenching agent. The regulated dead cells showed green fluorescence.
Transcriptomic sequencing
Transcriptomic sequencing was performed at Novogene Bioinformatics Technology Co., Ltd. (Beijing, China). In brief, total RNA was isolated by the use of TRIzol reagent, and the purity, concentration and integrity of the RNA were determined. Once the RNA samples were qualified, oligo-dT beads were used for the isolation of mRNA. The isolated mRNA was fragmented into short sections and used for the synthesis of first-strand cDNA. The synthesis of second-strand cDNA was performed by use of dNTPs, DNA polymerase I and RNase H. The purification of double-stranded cDNA was carried out by using AMPure XP beads, and then the purified double-stranded cDNA was used for end reparation, polyadenylation and ligation of adapters. After size selection with AMPure XP beads, PCR amplification was performed, and the product was purified using AMPure XP beads for library construction. Finally, the library consisting of 2 × 150 bp paired-end reads was sequenced on an Illumina HiSeq P150 sequencer (Illumina, San Diego, CA, USA). The raw data were cleaned by removing reads with adapters, reads of low quality (> 50%) or reads with a high proportion of unknown bases (> 10%) and then assembled with the Trinity software package. Annotation of transcriptome sequences was performed by using the following public databases: the Nr (NCBI nonredundant protein sequences), Nt (NCBI nucleotide sequences), Pfam (Protein family), KOG/COG (eukaryotic Ortholog Groups and Clusters of Orthologous Groups of proteins), Swiss-Prot (a manually annotated and reviewed protein sequence database), KEGG (Kyoto Encyclopedia of Genes and Genomes)72 and GO (Gene Ontology) databases. The mapped read numbers were transformed into FPKM (fragments per kilobase of transcript sequence per million base pairs sequenced) values for gene expression quantification and differential expression analysis. KEGG pathway analysis was conducted, and the corrected P-value cutoff was set at 0.05. The sequencing data have been deposited in the NCBI Sequence Read Archive (SRA) database under the accession number PRJNA588520.
Proteomic sequencing
Proteomic sequencing was performed at Novogene Bioinformatics Technology Co., Ltd. (Beijing, China). In brief, the concentrations of proteins were determined using the Bradford assay, and 100 µg of protein from each replicate was digested with Trypsin Gold (Promega, Madison, WI) at 37 ℃ for 16 h. The peptides were then dried by vacuum centrifugation, and the desalted peptides were labeled with TMT6/10-plex reagents (TMT6/10plex Isobaric Label Reagent Set, Thermo Fisher) according to the manufacturer’s instructions. After labeling, the samples were mixed and lyophilized. The TMT-labeled peptide mix was fractionated using a C18 column (Waters BEH C18 4.6 × 250 mm, 5 µm) on a Rigol L3000 HPLC instrument. Eluent was collected every minute and then combined to generate 15 fractions. Shotgun proteomic analyses were performed using an EASY-nLC 1200 UHPLC system (Thermo Fisher) coupled to an Orbitrap Q Exactive HF-X mass spectrometer (Thermo Fisher) operating in the data-dependent acquisition (DDA) mode. Peptides were separated on a Reprosil-Pur 120 C18-AQ analytical column (15 cm × 150 μm, 1.9 μm). Full MS scans from 350 to 1500 m/z were obtained at a resolution of 60,000 (at 200 m/z), with an AGC target value of 3 × 106 and a maximum ion injection time of 20 ms. From the full MS scan, a maximum number of 40 of the most abundant precursor ions were selected for higher energy collisional dissociation (HCD) fragment analysis at a resolution of 15,000 for TMT6-plex (at 200 m/z) or 45,000 for TMT10-plex (at 200 m/z), with an automatic gain control (AGC) target value of 1 × 105, a maximum ion injection time of 45 ms, a normalized collision energy of 32%, an intensity threshold of 8.3 × 103, and a dynamic exclusion parameter of 60 s. Differentially accumulated proteins (DAPs) were selected based on p < 0.05 and |log2FC|> 0.585. To identify the most important biochemical metabolic pathways and signal transduction pathways involved in the DAPs, KEGG analysis was performed. GO enrichment analysis of DAPs was also performed to identify the major biological functions.
Quantitative real-time PCR validation
Fifteen genes that were mapped to the lignin, cellulose and xylan biosynthesis pathways were randomly selected for quantification via real-time PCR. The sequences of the primers used were designed using the Oligo 7.0 software package. The specificity of the primer sets was examined by running the PCR products on agarose gels to ensure single-band amplification. Approximately 1 μg of total RNA was used for the synthesis of cDNA with RR037A reverse transcriptase (Takara, USA). qRT-PCR was performed in a 20 μl system consisting of 1 μl each of both forward and reverse primers, SYBR Green Real-Time PCR Master Mix (Invitrogen, USA) and 1 μl of cDNA template on a Step-One Plus Real-Time PCR System (Applied Biosystems, USA). The cycling conditions were as follows: 95 °C for 15 min followed by 40 cycles of 95 °C for 10 s, 58 °C for 20 s, and 72 °C for 20 s. Each sample comprised three individuals as replicates. The expression level of each gene was normalized to that of tubulin, and the relative gene expression level was calculated with the 2-ΔΔCt method73.
Statistical analysis
The statistical significance of differences in this study was analyzed using one-way ANOVA followed by Tukey’s post hoc test with a significance level of 0.05 (p < 0.05) (GraphPad Prism 7.0 software for Windows).
Ethics approval and consent to participate
The fruit samples were collected in an orchard at Korla (Xingjiang, China) with the permission of the local grower. The experimental research on plants, including collection of plant material, was complied with institutional, national, or international guidelines. And field studies were conducted in accordance with local legislation. And also comply with the Convention on the Trade in Endangered Species of Wild Fauna and Flora.
Data availability
① The raw data of transcription sequencing generated during the current study are available in the NCBI Bioproject repository (https://www.ncbi.nlm.nih.gov/bioproject/PRJNA588520). ② The raw data of proteome sequencing generated during the current study are available in the iProX repository (https://www.iprox.org/page/project.html?id=IPX0002167000). It also can be found in the repository of ProteomeXchange (http://proteomecentral.proteomexchange.org/cgi/GetDataset?ID=PXD018829). ③ All data analyzed during this study are included in this published article (and its supplementary data files).
References
Cai, Y. et al. Study of the structure and biosynthetic pathway of lignin in stone cells of pear. Sci. Hortic. https://doi.org/10.1016/j.scienta.2010.04.029 (2010).
Tao, S., Khanizadeh, S., Zhang, H. & Zhang, S. Anatomy, ultrastructure and lignin distribution of stone cells in two Pyrus species. Plant Sci. 176, 413–419. https://doi.org/10.1016/j.plantsci.2008.12.011 (2009).
Brahem, M., Renard, C. M. G. C., Gouble, B., Bureau, S. & Le Bourvellec, C. Characterization of tissue specific differences in cell wall polysaccharides of ripe and overripe pear fruit. Carbohyd. Polym. 156, 152–164. https://doi.org/10.1016/j.carbpol.2016.09.019 (2017).
Haigler, C. H., Betancur, L., Stiff, M. R. & Tuttle, J. R. Cotton fiber: a powerful single-cell model for cell wall and cellulose research. Front. Plant Sci. 3, 104. https://doi.org/10.3389/fpls.2012.00104 (2012).
Timell, T. E. Recent progress in the chemistry of wood hemicelluloses. Wood Sci. Technol. 1, 45–70. https://doi.org/10.1007/bf00592255 (1967).
Cao, Y., Han, Y., Li, D., Lin, Y. & Cai, Y. Systematic analysis of the 4-coumarate: coenzyme a ligase (4CL) related genes and expression profiling during fruit development in the Chinese pear. Genes (Basel) 7, 89, https://doi.org/https://doi.org/10.3390/genes7100089 (2016).
Jin, Q. et al. Structural characterization and deposition of stone cell lignin in Dangshan Su pear. Sci. Hortic. 155, 123–130. https://doi.org/10.1016/j.scienta.2013.03.020 (2013).
Lu, G., Li, Z., Zhang, X., Wang, R. & Yang, S. Expression analysis of lignin-associated genes in hard end pear (Pyrus pyrifolia Whangkeumbae) and its response to calcium chloride treatment conditions. J. Plant. Growth Regul. 34, 251–262. https://doi.org/10.1007/s00344-014-9461-x (2014).
Wang, Y. et al. Heterogenous expression of Pyrus pyrifolia PpCAD2 and PpEXP2 in tobacco impacts lignin accumulation in transgenic plants. Gene 637, 181–189. https://doi.org/10.1016/j.gene.2017.09.056 (2017).
Yan, C. et al. Stone cell distribution and lignin structure in various pear varieties. Sci. Hortic. 174, 142–150. https://doi.org/10.1016/j.scienta.2014.05.018 (2014).
Barros, J., Serk, H., Granlund, I. & Pesquet, E. The cell biology of lignification in higher plants. Ann. Bot. 115, 1053–1074. https://doi.org/10.1093/aob/mcv046 (2015).
Suzuki, N. et al. Respiratory burst oxidases: the engines of ROS signaling. Curr. Opin. Plant Biol. 14, 691–699. https://doi.org/10.1016/j.pbi.2011.07.014 (2011).
Voxeur, A., Wang, Y. & Sibout, R. Lignification: different mechanisms for a versatile polymer. Curr. Opin. Plant Biol. 23, 83–90. https://doi.org/10.1016/j.pbi.2014.11.006 (2015).
Mamat, A. et al. Study on the relationship between the stone cell formation and apoptosis during the fruit development of Korla Xiangli pear. J. Fruit. Sci. 37, 59–67, https://doi.org/https://doi.org/10.13925/j.cnki.gsxb.20180494 (2020).
Mahboubi, M. A. H. New Insights into Carbon Transport and Incorporation to Wood Ph.D. thesis, Swedish University of Agricultural Sciences, (2014).
York, W. & Oneill, M. Biochemical control of xylan biosynthesis — which end is up?. Curr. Opin. Plant Biol. 11, 258–265. https://doi.org/10.1016/j.pbi.2008.02.007 (2008).
Mamat, A. et al. Pulp lignification in Korla fragrant pear. Eur. J. Hortic. Sci. 263–273, https://doi.org/https://doi.org/10.17660/eJHS.2019/84.5.2 (2019).
Nie, J. et al. The anatomic study on relationship of stone cells and parenchyma cells during fruit development of Pyrus bretschneideri. Acta Hortic. Sin. 36, 1209–1214 (2009).
Qiao, Y., Zhang, S., Tao, S., Zhang, Z. & Liu, Z. Advances in research on developing mechanism of stone cells in pear fruit. J. Fruit. Sci. 22, 367–371 (2005).
Ben-Arie, R., Kislev, N. & Frenkel, C. Ultrastructural changes in the cell walls of ripening apple and pear fruit. Plant Physiol. 64, 197–202. https://doi.org/10.1104/pp.64.2.197 (1979).
Pesquet, E. et al. Non-cell-autonomous postmortem lignification of tracheary elements in Zinnia elegans. Plant Cell 25, 1314–1328. https://doi.org/10.1105/tpc.113.110593 (2013).
Smith, R. A. et al. Neighboring parenchyma cells contribute to Arabidopsis xylem lignification, while lignification of interfascicular fibers is cell autonomous. Plant Cell 25, 3988–3999. https://doi.org/10.1105/tpc.113.117176 (2013).
Higuchi, T. Lignin biochemistry: biosynthesis and biodegradation. Wood Sci. Technol. 24, 23–63. https://doi.org/10.1007/bf00225306 (1990).
Terashima, N. & Fukushima, K. Biogenesis and structure of macromolecular lignin in the cell wall of tree xylem as studied by microautoradiography. Plant Cell Wall Polym. ACS Symp. Ser. 399, 160–168, https://doi.org/https://doi.org/10.1021/bk-1989-0399.ch011 (1989).
Arioli, T. et al. Molecular analysis of cellulose biosynthesis in Arabidopsis. Science 279, 717–720. https://doi.org/10.1126/science.279.5351.717 (1998).
Coleman, H. D., Yan, J. & Mansfield, S. D. Sucrose synthase affects carbon partitioning to increase cellulose production and altered cell wall ultrastructure. Proc. Natl. Acad Sci. USA 106, 13118–13123. https://doi.org/10.1073/pnas.0900188106 (2009).
Gerber, L. et al. Deficient sucrose synthase activity in developing wood does not specifically affect cellulose biosynthesis, but causes an overall decrease in cell wall polymers. New Phytol. 203, 1220–1230. https://doi.org/10.1111/nph.12888 (2014).
Maimaiti, A. et al. Soluble sugar accumulation and related enzyme activity in the fruits of Korla fragrant pear. Xinjiang Agric. Sci. 55, 82–91 (2018).
Roach, M. et al. Fructokinase is required for carbon partitioning to cellulose in aspen wood. Plant J. 70, 967–977. https://doi.org/10.1111/j.1365-313X.2012.04929.x (2012).
Verbančič, J., Lunn, J. E., Stitt, M. & Persson, S. Carbon supply and the regulation of cell wall synthesis. Mol. Plant 11, 75–94. https://doi.org/10.1016/j.molp.2017.10.004 (2018).
Taylor, N. G., Howells, R. M., Huttly, A. K., Vickers, K. & Turner, S. R. Interactions among three distinct CesA proteins essential for cellulose synthesis. Proc. Natl. Acad Sci. USA 100, 1450–1455. https://doi.org/10.1073/pnas.0337628100 (2003).
Timmers, J. et al. Interactions between membrane-bound cellulose synthases involved in the synthesis of the secondary cell wall. FEBS Lett. 583, 978–982. https://doi.org/10.1016/j.febslet.2009.02.035 (2009).
Gardiner, J. C., Taylor, N. G. & Turner, S. R. Control of cellulose synthase complex localization in developing xylem. Plant Cell 15, 1740–1748. https://doi.org/10.1105/tpc.012815 (2003).
Handakumbura, P. P. et al. Perturbation of Brachypodium distachyon CELLULOSE SYNTHASE A4 or 7 results in abnormal cell walls. BMC Plant Biol. 13, 131. https://doi.org/10.1186/1471-2229-13-131 (2013).
Tanaka, K. et al. Three distinct rice cellulose synthase catalytic subunit genes required for cellulose synthesis in the secondary wall. Plant Physiol. 133, 73–83. https://doi.org/10.1104/pp.103.022442 (2003).
Joshi, C. P. et al. Perturbation of wood cellulose synthesis causes pleiotropic effects in transgenic aspen. Mol. Plant. 4, 331–345. https://doi.org/10.1093/mp/ssq081 (2011).
Simmons, T. J. et al. Folding of xylan onto cellulose fibrils in plant cell walls revealed by solid-state NMR. Nat. Commun. 7, 13902. https://doi.org/10.1038/ncomms13902 (2016).
Du, Q., Pan, W., Tian, J., Li, B. & Zhang, D. The UDP-glucuronate decarboxylase gene family in Populus: structure, expression, and association genetics. PLoS ONE 8, e60880. https://doi.org/10.1371/journal.pone.0060880 (2013).
Pan, Y., Wang, X., Liu, H., Zhang, G. & Ma, Z. Molecular cloning of three UDP-glucuronate decarboxylase genes that are preferentially expressed in Gossypium fibers from elongation to secondary cell wall synthesis. J. Plant Biol. 53, 367–373. https://doi.org/10.1007/s12374-010-9124-9 (2010).
Lee, C., O’Neill, M. A., Tsumuraya, Y., Darvill, A. G. & Ye, Z. H. The irregular xylem9 mutant is deficient in xylan xylosyltransferase activity. Plant Cell Physiol. 48, 1624–1634. https://doi.org/10.1093/pcp/pcm135 (2007).
Peña, M. J. et al. Arabidopsis irregular xylem8 and irregular xylem9: implications for the complexity of glucuronoxylan biosynthesis. Plant Cell 19, 549–563. https://doi.org/10.1105/tpc.106.049320 (2007).
Brown, D. M. et al. Comparison of five xylan synthesis mutants reveals new insight into the mechanisms of xylan synthesis. Plant J. 52, 1154–1168. https://doi.org/10.1111/j.1365-313X.2007.03307.x (2007).
Brown, D. M., Zhang, Z., Stephens, E., Dupree, P. & Turner, S. R. Characterization of IRX10 and IRX10-like reveals an essential role in glucuronoxylan biosynthesis in Arabidopsis. Plant J. 57, 732–746. https://doi.org/10.1111/j.1365-313X.2008.03729.x (2009).
Wu, A.-M. et al. The Arabidopsis IRX10 and IRX10-LIKE glycosyltransferases are critical for glucuronoxylan biosynthesis during secondary cell wall formation. Plant J. 57, 718–731. https://doi.org/10.1111/j.1365-313X.2008.03724.x (2009).
Brown, D. et al. Arabidopsis genes IRREGULAR XYLEM (IRX15) and IRX15L encode DUF579-containing proteins that are essential for normal xylan deposition in the secondary cell wall. Plant J. 66, 401–413. https://doi.org/10.1111/j.1365-313X.2011.04501.x (2011).
Brown, D. M., Zeef, L. A. H., Ellis, J., Goodacre, R. & Turner, S. R. Identification of novel genes in Arabidopsis involved in secondary cell wall formation using expression profiling and reverse genetics. Plant Cell 17, 2281–2295. https://doi.org/10.1105/tpc.105.031542 (2005).
Jensen, J. K. et al. The DUF579 domain containing proteins IRX15 and IRX15-L affect xylan synthesis in Arabidopsis. Plant J. 66, 387–400. https://doi.org/10.1111/j.1365-313X.2010.04475.x (2011).
Lee, C., Teng, Q., Zhong, R. & Ye, Z. H. Arabidopsis GUX proteins are glucuronyltransferases responsible for the addition of glucuronic acid side chains onto xylan. Plant Cell Physiol. 53, 1204–1216. https://doi.org/10.1093/pcp/pcs064 (2012).
Mortimer, J. C. et al. Absence of branches from xylan in Arabidopsis gux mutants reveals potential for simplification of lignocellulosic biomass. Proc. Natl. Acad Sci. USA 107, 17409–17414. https://doi.org/10.1073/pnas.1005456107 (2010).
Rennie, E. A. et al. Three members of the Arabidopsis glycosyltransferase family 8 are xylan glucuronosyltransferases. Plant Physiol. 159, 1408–1417. https://doi.org/10.1104/pp.112.200964 (2012).
Lee, C., Teng, Q., Huang, W., Zhong, R. & Ye, Z. H. The F8H glycosyltransferase is a functional paralog of FRA8 involved in glucuronoxylan biosynthesis in Arabidopsis. Plant Cell Physiol. 50, 812–827. https://doi.org/10.1093/pcp/pcp025 (2009).
Lee, C. et al. The PARVUS gene is expressed in cells undergoing secondary wall thickening and is essential for glucuronoxylan biosynthesis. Plant Cell Physiol. 48, 1659–1672. https://doi.org/10.1093/pcp/pcm155 (2007).
Persson, S. et al. The Arabidopsis irregular xylem8 mutant is deficient in glucuronoxylan and homogalacturonan, which are essential for secondary cell wall integrity. Plant Cell 19, 237–255. https://doi.org/10.1105/tpc.106.047720 (2007).
Raes, J., Rohde, A., Christensen, J. H., Van de Peer, Y. & Boerjan, W. Genome-wide characterization of the lignification toolbox in Arabidopsis. Plant Physiol. 133, 1051–1071. https://doi.org/10.1104/pp.103.026484 (2003).
Vanholme, R. et al. Metabolic engineering of novel lignin in biomass crops. New Phytol. 196, 978–1000. https://doi.org/10.1111/j.1469-8137.2012.04337.x (2012).
Li, H. et al. The effect of 1-methylcyclopropene, methyl jasmonate and methyl salicylate on lignin accumulation and gene expression in postharvest ‘Xuxiang’ kiwifruit during cold storage. Postharvest Biol. Technol. 124, 107–118. https://doi.org/10.1016/j.postharvbio.2016.10.003 (2017).
Bonawitz, N. D. & Chapple, C. The genetics of lignin biosynthesis: connecting genotype to phenotype. Annu. Rev. Genet. 44, 337–363. https://doi.org/10.1146/annurev-genet-102209-163508 (2010).
Herrero, J. et al. Bioinformatic and functional characterization of the basic peroxidase 72 from Arabidopsis thaliana involved in lignin biosynthesis. Planta 237, 1599–1612. https://doi.org/10.1007/s00425-013-1865-5 (2013).
Lu, S. et al. Ptr-miR397a is a negative regulator of laccase genes affecting lignin content in Populus trichocarpa. Proc. Natl. Acad Sci. USA 110, 10848–10853. https://doi.org/10.1073/pnas.1308936110 (2013).
Shigeto, J., Kiyonaga, Y., Fujita, K., Kondo, R. & Tsutsumi, Y. Putative cationic cell-wall-bound peroxidase homologues in Arabidopsis, AtPrx2, AtPrx25, and AtPrx71, are involved in lignification. J. Agric. Food Chem. 61, 3781–3788. https://doi.org/10.1021/jf400426g (2013).
Berthet, S. et al. Disruption of LACCASE4 and 17 results in tissue-specific alterations to lignification of Arabidopsis thaliana stems. Plant Cell 23, 1124–1137. https://doi.org/10.1105/tpc.110.082792 (2011).
Zhao, Q. et al. Loss of function of cinnamyl alcohol dehydrogenase 1 leads to unconventional lignin and a temperature-sensitive growth defect in Medicago truncatula. Proc. Natl. Acad Sci. USA 110, 13660–13665. https://doi.org/10.1073/pnas.1312234110 (2013).
Shigeto, J. et al. Simultaneously disrupting AtPrx2, AtPrx25 and AtPrx71 alters lignin content and structure in Arabidopsis stem. J. Integr. Plant Biol. 57, 349–356. https://doi.org/10.1111/jipb.12334 (2015).
Fry, S. C. Oxidative coupling of tyrosine and ferulic acid residues: Intra- and extra-protoplasmic occurrence, predominance of trimers and larger products, and possible role in inter-polymeric cross-linking. Phytochem Rev. 3, 97–111. https://doi.org/10.1023/b:phyt.0000047808.74647.43 (2004).
Novo-Uzal, E. et al. From Zinnia to Arabidopsis: approaching the involvement of peroxidases in lignification. J. Exp. Bot. 64, 3499–3518. https://doi.org/10.1093/jxb/ert221 (2013).
Blokhina, O. B., Chirkova, T. V. & Fagerstedt, K. V. Anoxic stress leads to hydrogen peroxide formation in plant cells. J. Exp. Bot. 52, 1179–1190. https://doi.org/10.1093/jexbot/52.359.1179 (2001).
Kotula, L., Ranathunge, K., Schreiber, L. & Steudle, E. Functional and chemical comparison of apoplastic barriers to radial oxygen loss in roots of rice (Oryza sativa L.) grown in aerated or deoxygenated solution. J. Exp. Bot. 60, 2155–2167, https://doi.org/https://doi.org/10.1093/jxb/erp089 (2009).
Gechev, T. S., Van Breusegem, F., Stone, J. M., Denev, I. & Laloi, C. Reactive oxygen species as signals that modulate plant stress responses and programmed cell death. BioEssays 28, 1091–1101. https://doi.org/10.1002/bies.20493 (2006).
Raiskila, S. et al. FTIR spectroscopic prediction of klason and acid soluble lignin variation in Norway spruce cutting clones. Silva Fenn https://doi.org/https://doi.org/10.14214/sf.301 (2007).
Xiong, S., Zuo, X. & Zhu, Y. Determination of cellulose, hemi-cellulose and ligin in rice hull. Cereal Feed Ind 8, 40–41 (2005).
Mao, S., Xu, J., Zhou, S., Zhang, Y. & Liu, Y. Determination of polysaccharide from Lonicera japonica at different flowering phases by phenol-sulfuric acid method. Cent S. Pharm. 13, 65–67 (2015).
Kanehisa, M. et al. KEGG for linking genomes to life and the environment. Nucl. Acids Res. 36, D480–D484. https://doi.org/10.1093/nar/gkm882 (2008).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25, 402–408. https://doi.org/10.1006/meth.2001.1262 (2001).
Acknowledgements
This research was funded by the National Natural Science Foundation of China (NO. 31760565) and Natural Science Foundation of Xinjiang Province (CN) (NO. 2019D01A63). Funders have no role in the design of the study and collection, analysis, and interpretation of data and in writing the manuscript. We would like to thank the members of Novogene Bioinformatics Technology Co., Ltd for technical sport for during the proteome and transcriptome sequencing.
Author information
Authors and Affiliations
Contributions
A.M. designed the research. A.M., K.T., and J.X. performed most of the experiments and analyzed the data. A.M. wrote the manuscript. P.Y., C.M. and J.X.W. participated to the result analyses and interpretation of data. A.M. and K.T. revised the manuscript critically. All authors have read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Mamat, A., Tusong, K., Xu, J. et al. Integrated transcriptomic and proteomic analysis reveals the complex molecular mechanisms underlying stone cell formation in Korla pear. Sci Rep 11, 7688 (2021). https://doi.org/10.1038/s41598-021-87262-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-87262-3
This article is cited by
-
Genome-Wide Investigation of NAC Family Genes Potentially Related to SCW Biogenesis in Korla Pear Fruit
Plant Molecular Biology Reporter (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.