Abstract
Dogs are known to be the oldest animals domesticated by humans. Although many studies have examined wolf domestication, the geographic and temporal origin of this process is still being debated. To address this issue, our study sheds new light on the early stages of wolf domestication during the Magdalenian period (16–14 ka cal BP) in the Hegau Jura region (Southwestern Germany and Switzerland). By combining morphology, genetics, and isotopes, our multidisciplinary approach helps to evaluate alternate processes driving the early phases of domestication. The isotope analysis uncovered a restricted, low δ15N protein diet for all analyzed Gnirshöhle specimens, while morphological examinations and phylogenetic relationships did not unequivocally assign them to one or the other canid lineage. Intriguingly, the newly generated mitochondrial canid genomes span the entire genetic diversity of modern dogs and wolves. Such high mitochondrial diversity could imply that Magdalenian people tamed and reared animals originating from different wolf lineages. We discuss our results in light of three ecological hypotheses and conclude that both domestication and the existence of a specialized wolf ecomorph are highly probable. However, due to their proximity to humans and a restricted diet, we propose domestication as the most likely scenario explaining the patterns observed herein.
Similar content being viewed by others
Introduction
In line with several theories detailing biological diversity and evolution, the survival of species is often predicated on their ability to adapt and thrive within a changing environment. To do so, plant and animal species have developed several strategies, including the adaptation to existing, or the development of new ecological niches1. This is exemplified by various studies that have investigated faunal adaptation to the environment before and after the Last Glacial Maximum (LGM), a period of climatic fluctuation and dramatic landscape changes2. One possibility to survive unfavorable conditions is to retreat into refugia, i.e. restricted and often isolated areas still harboring ecological and environmental features beneficial for the species in question3,4. Specifically, during the LGM, various species retreated to warmer areas such as the Iberian Peninsula5, subsequently preserving and shaping patterns of genetic variation prevalent during this epoch6.
An alternative pathway for survival is to form so-called ecomorphs and thus adapt to environmental conditions and habitats7. Ecomorphs are characterized by a genetic variety within a species or a variety of several species exposing the same phenotypic features, due to the adaptation to a local ecology. One prominent example of a species represented by many ecomorphs since prehistoric periods is the grey wolf (Canis lupus). The wolf is a mobile carnivore with an opportunistic diet8, and can occupy different trophic niches, best exemplified by Late Pleistocene wolves, which were characterized by two different dietary habits9,10.
During the Late Pleistocene, as humans became more numerous and intrusive11,12,13,14, they started to re-shape their local environment and thereby became a driving factor within the landscape providing artificial ecological and dietary (= trophic) niches15,16,17. The concept of humans as niche constructors is vividly debated and recently contributed to our understanding of domestication18,19. This debate includes the origin of modern dogs, which may have occurred intentionally by pet-keeping, a side-effect of goal-orientated domestication, or unintentionally (self-domestication), when humans created a niche for commensal scavengers with their food waste. The uniting element of these theories is the subsequent selection for tameness and reduced fear, resulting in decreased wariness and aggression, high tolerance of penning, and sexual precocity20,21.
Although the when and where of wolf domestication are still disputed, it is now accepted that dogs are indeed the oldest domesticates10,21,22,23,24. Once the process of domestication began, humans quickly gained control over the diet, reproduction, and health of their new companions and thus set the stage for a lasting human–dog bond21,24,25. For example, stable isotope analyses of canids found in Předmostí, a Gravettian site in the Czech Republic dated to 31,500 years ago, showed evidence of adaptation to two different dietary niches21,26,27; however, it remains unclear whether or not one of these two different canid groups are indeed early domesticated wolves24,27,28. While such older dates for potential onsets of wolf domestication have been challenged, it is widely accepted that 16,000 years ago is the approximate time for the first emergence of dogs10. From this time onwards, dog remains have been discovered at several archaeological sites throughout Europe, such as the famous dog from Bonn-Oberkassel excavated alongside two human burials25,29,30. Another important site with possible early dog remains is Kesslerloch31,32 (Canton Schaffhausen) located in the Hegau Jura and dating to the late Magdalenian period (17.5–14.3 ka cal BP33). Morphological examinations of these remains revealed a distinction between wolves and dogs32,33 and dietary reconstruction analyses16 demonstrated parallel occurrences of two distinct canid trophic niches at this site. However, it remains unclear whether Kesslerloch is a unique case or if wolf domestication was practiced in the entire region over a longer period of time. The Hegau Jura, including the Kesslerloch site and two other cave sites, Petersfels and Drexlerloch, represents a Magdalenian hotspot for human activity in the pre-Alpine region. In the heart of this setting lies the cave site Gnirshöhle, which also shows evidence of human occupation, with butchered animal bones, worked antlers, and bone needles34 (Supplementary Note 1), as well as provides a cohort of large canid remains, making this cave particularly valuable for investigating pre-Alpine canids.
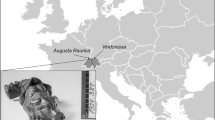
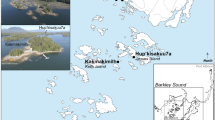
In the present study, we applied a multidisciplinary approach to study canid remains from the Gnirshöhle and assessed canid population dynamics potentially shaped by environmental conditions and anthropogenic pressure during the Magdalenian in the Hegau Jura (Fig. 1). By combining isotopic dietary reconstruction, metric assessment, and paleogenetics, we were able to test various hypotheses to better elucidate the onset of wolf to dog transition in the Hegau Jura region and derived a refined proposal of wolf domestication.
Map of the archaeological sites from which the canid remains were retrieved and investigated. Dots represent the sites from which new data are included in this study, the triangle marks a site with important comparative data. Additional information about sample sizes and the archaeological dates are provided in Table S1. Dates are given in ka (kilo annos) cal BP, for samples younger than 2000 years dates are given in century AD.
Results
Archaeozoology: morphological and metric results
All samples except one (GN-999) were assigned to be large canid specimens (Canis sp.) by archaeozoological classification. The mandible GN-999 (Figure S2) was described earlier as a wolf-like specimen with morphological traits of domestication34. The mandible is relatively short and exhibits tooth crowding between P4 and M1. GN-999 contains a tooth row with P2, P3, P4, M1, and M2, while P1 is missing and M3 was lost premortem; the alveolus of these two missing teeth is still visible.
The length of the tooth row of the mandible GN-999 (ALP1M3 = 94.4 mm) plotted against the length of the first molar (CLM1 = 27.3 mm) falls into the observed absolute ranges of recent Northern wolves and ‘Palaeolithic dog’ group35. Although, GN-999 is considerably shorter by one standard deviation than the lower bound of the range for modern and Pleistocene wolves, as well as the ‘Paleolithic dog’ group35. This specimen is also morphologically similar to a small wolf from Kesslerloch32 (Fig. 2, Table S4). Therefore, we cannot exclude GN-999 assigned as a wolf using metric data (other standardized measurements after von den Driesch36 concerning GN-999 and the maxilla GN-192 are given in Supplementary Note 2 and Tables S2 and S3). Tooth crowding of the premolars is a typical trait for dogs and is related to shortening of the snout during the process of domestication37. However, crowding between P4 and M1 is also observed in wolves38. Additionally, a diastema is present between the P2 and the P3, which is rather typical for recent and Pleistocene wolves35. Thus, the morphological and metric assignments for GN-999 remain unresolved.
Metric data of GN-999 (black star) in comparison to metric data of seven canid groups (see Table S2): Modern dogs (n = 2, reference collection Uni Tübingen); Modern wolves (n = 4, reference collection Uni Tübingen and Bonn); Pleistocene wolves (n = 7, Brillenhöhle, Kesslerloch, Geißenklösterle. The four groups ‘Archaic dogs’ (n = 27), ‘Palaeolithic dogs’ (n = 31), ‘Northern wolves’ (n = 35) and ‘Pleistocene wolves’ (n = 36) are defined by Germonpré and colleagues35 and defined by mean value and standard deviation.
Stable isotope analysis: niche modeling and dietary reconstruction
The percentage of nitrogen in bone was measured for a total of ten canids (Table S1 and Supplementary Note 4); only six of the Gnirshöhle canids, the red fox from Bockstein, and the pre-LGM wolf from Hohle Fels had sufficient collagen content for analysis (1.1–3.1% Nbone)39. Specimen HF-912 was removed from the isospace published by Baumann and colleagues16, as it was dated to the pre-LGM (Supplementary Note 3). The newly analyzed HF-1712 sample, contextually dated to pre-LGM, was also excluded. The slightly younger specimen, HF-1250.2 (Supplementary Note 3), was dated to 11,400 ± 30 BP (13,308–13,150 cal BP), and therefore belonged to the Late Palaeolithic period. However, since both archaeological periods are temporarily close and the environmental conditions did not change dramatically, we decided to keep the isotopic values of this specimen in the reconstructed Magdalenian isospace. We calculated three canid niches: niche A, niche B, and niche C (Fig. 3A,B), in accordance to the δ13C and δ15N values of each specimen. For niche A and C, we observed no changes compared to the previously published isospace16. Niche B, however, is further extended with respect to the δ13C values. Briefly, five wolves16 from Kesslerloch and Hohle Fels, as well as one red fox16 from Vogelherd fell into niche A (Fig. 3A), with δ13C values ranging from − 20.1 to − 19.4‰ and δ15N values ranging from + 7.1 to + 9.1‰. Niche B included three dogs16,40, one red fox40 from Kesslerloch, and all newly analyzed canids from Gnirshöhle (Fig. 3A). The isotopic values of this niche ranged from − 20.0 to − 19.0‰ and from + 4.7 to + 6.4‰ for δ13C and δ15N, respectively. Finally, niche C included three red foxes16 from Geißenklösterle, Gnirshöhle, and Bockstein, one arctic fox16 from Hohle Fels, and one wolf40 from Schussenquelle (Fig. 3A). This niche is characterized by δ13C values ranging from − 20.6 to − 20.3‰ and δ15N values ranging from + 4.5 to + 5.8‰.
(A) Cluster analysis of stable isotopic values of the canids. (B) Reconstructed trophic niches and (C) dietary reconstruction of the ancient Magdalenian canids based on stable isotope data of carbon and nitrogen preserved in bone collagen. Labels of the new samples from this study are in bold and marked with an orange rectangular frame. Numbers in (B) correspond to these in (A).
We reconstructed the percentages of three different dietary sources for each niche (i.e., megafauna, small game, ungulates; see methods section, and Table S5), using the Bayesian statistic model MixSIAR. The dietary preferences strongly varied among the niches (Fig. 3C, Table 1). Specimens of niche A had a preference for megaherbivores, such as mammoths. Members of niche B fed on small mammals, such as hares, and in addition, on ungulates, such as reindeer and horse, and megaherbivores. Lastly, individuals in niche C had a high preference for small mammals.
Palaeogenetics: phylogeny, genetic diversity, and evolutionary trajectory
Although our genetic investigations were focused on analyzing the mitochondrial DNA sequences of the canid samples from Gnirshöhle (n = 8), additional mitochondrial genomes from canid remains of diverse temporal and geographic origin (Fig. 1, Supplementary Note 1, Table S1) were also generated and included to increase the genetic diversity. Out of 28 extracted samples, we were able to reconstruct a total of 23 (77%) complete or nearly complete mitochondrial genomes, including five Magdalenian mitochondrial genomes from Gnirshöhle, and one from Hohle Fels (Table S1). The mean coverage ranged from 7.0 to 341.2 (Table 2), and all reconstructed mitochondrial genomes had a threefold coverage between 93.5 and 100% of the ~ 16,000 mitochondrial nucleotides (Table 2).
The phylogenetic arrangement, estimated in a Maximum Likelihood framework (Fig. 4), did not reveal any clear chronological or spatial differentiation of our six Magdalenian samples compared to the assemblage of ancient and modern canids. Our novel Magdalenian mitochondrial genomes clustered with ancient mitochondrial genomes from Belgium, the Czech Republic, and Russia, ranging in age from the pre-LGM (ca. 50—28 ka cal BP) to the post-LGM (19.5—16 ka cal BP), in addition to modern canids of global origins (Fig. 4). Interestingly, one mitochondrial genome, from the Magdalenian specimen GN-192 (ca. 15.6—15.2 ka cal BP, Supplementary Note 3, Figure S1), fell within a very basal cluster that was previously assigned exclusively to Belgian pre-LGM canids. Taking the age of this specimen into account (Supplementary Note 3), this finding implies a genetic continuity of one maternal canid lineage from the pre-LGM to the Magdalenian. Furthermore, the specimens GN-14, GN-106, and GN-133 are placed closely to the two ancient wolves KSL-5825 and KSL-6141; additionally, the specimen HF-530 is placed with a third Swiss mitochondrial genome, KSL-62 (Fig. 4).
Maximum likelihood (ML) consensus tree of mitochondrial canid genomes. Newly generated samples are highlighted in orange; with the GN canids further bolded. Statistical support was assessed by generating 10,000 bootstrap replicates and nodes with bootstrap support higher than 95% are shown with an asterisk. Important nodes are labeled with respective node ages (given in ka cal BP) estimated using time-aware Bayesian phylogenetic inference (Figure S7). Ancient dogs are labeled in dark blue, modern dogs in light blue. Ancient wolves in dark green, modern wolves in light green. The ages of samples older 1,000 years are given in ka (cal BP).
Altogether, these results demonstrate a close maternal relationship of temporally spaced specimens from the same region in southwestern Germany. Despite the close relationship of these specimens to others within the region, we also observed a close genetic affinity of temporally and geographically distantly spaced maternal haplotypes, e.g., GN-999 was placed close to mitochondrial DNA sequences of ancient and modern canids originating from northeastern Europe, Siberia, and eastern Beringia. In general, we observed that our reconstructed mitochondrial genomes for specimens older than the Mesolithic, an archaeological period of the early Holocene, are maternally more closely related to modern and ancient wolves than to modern and ancient dogs (Figure S5). Nevertheless, an assignment of the canids as dogs or wolves cannot be unambiguously performed based solely on the genetic affinity.
As evidenced by the broad distribution of our samples throughout the phylogeny, the Magdalenian canids from southwestern Germany harbor a vast amount of genetic diversity. To assess the genetic diversity of the Magdalenian Gnirshöhle canids, we compared the nucleotide diversity (π) of those specimens with the observed nucleotide diversity of the Kesslerloch canids, in addition to modern dogs and wolves sampled worldwide (Tables S7, S8, Figure S7). Nucleotide diversity is defined as the average number of differences per site between any two DNA sequences in the population42, and is less biased by sample size and potential sequencing errors than, for instance, haplotype diversity. The estimated nucleotide diversity of our five novel sequences from the Gnirshöhle canids (n = 5; π = 0.00306 ± 0.00106) was almost as high as that observed in a global assemblage of modern dogs (n = 79; π = 0.00339 ± 0.00021) and slightly lower than the diversity determined for the Kesslerloch canids (n = 5; π = 0.003379 ± 0.00098). These estimates of genetic diversity illustrate the high genetic variation in Magdalenian canids originated from caves located in the Hegau Jura region.
To evaluate the evolutionary timescale of the canid mitochondrial phylogeny, we performed BEAST analyses (Figure S6). The results produced a very similar topology compared to the Maximum Likelihood tree (Figure S5), except for minor differences whenever the statistical support was low, as for example the dogs within clade A, or amongst North American wolves. As part of the phylogenetic timescale estimation, the ages of the samples not directly dated by 14C analyses were estimated. While the median ages for most samples do not differ substantially from their respective archaeological age (Table S6), the age of the three canid samples from Frankfurt did; for F-1986.2 approximately 4 ka cal BP, for F-1986.1 and F-α19496 about 6 ka cal BP. However, the archaeological ages of these samples fall within the 95% credibility intervals of the age estimates and the departure of median estimated age is likely caused by the young age of the samples being close to the minimum age constraint (0 ya) of the wide priors (uniform 0–100 ka) for the age parameter. Interestingly, for HF-912, the estimated age (~ 42 ka BP) was older than expected considering the sample was excavated from a layer contextually dated to the Magdalenian. This finding agrees with the taphonomic observations concerning the bone color that suggested the sample may be older than the layer in which it was found and has been additionally confirmed by 14C dating (31.4 ka cal BP, Supplementary Note 3).
To further elucidate the dynamics of wolf domestication, we estimated the timescale of the inferred canid mitogenomic evolutionary tree. As outlined by Loog and colleagues41, the coyote outgroup and the modern wolf samples from the Himalayas were excluded from this analysis. The TMRCA of all dogs and wolves was estimated to approximately 152 ka cal BP (111–231 cal ka BP 95% HPD; Fig. 4). The TMRCA for clade A was estimated to around 34 ka cal BP, clade B to circa 6.5 ka cal BP, clade C to approximately 22.5 ka cal BP, and clade D to about 3.5 ka cal BP. Our six novel Magdalenian mitogenomes fall outside these clades, but four are genetically close to either clade A or D, showing a presence of close relatives to modern dogs in the Magdalenian Hegau Jura. On the other hand, the same small region and period also hosted very genetically distant canids that diverged with modern dog lineages as early as 135 ka cal BP.
Discussion
Most studies focusing on ancient canids and the proposal of various scenarios for potential wolf domestication have previously used one line of evidence, such as morphological examinations, isotopic analyses, or genetic investigations21,22,25,26,27,35,41. Hence, studies analyzing the same archaeological material often lacked a consensus and often resulted in divergent outcomes25,26,35,41. Our multidisciplinary approach combined various archaeological disciplines to analyze pre-Alpine Magdalenian canids and revealed high mitochondrial variation prevalent in a geographically restricted area—the Gnirshöhle (Fig. 4). While Thalmann and colleagues25 had observed high variation among Pleistocene canids in general, our findings now highlight the Hegau Jura region, encompassing Magdalenian caves such as Gnirshöhle and Kesslerloch, as a hot-spot for genetic variation in Pleistocene canids. Interestingly, the variation within the Gnirshöhle mirrors the collective variation of almost all canids analyzed herein.
Utilizing one of the most comprehensive assemblages of canid mitochondrial genomes varying in time and space, we were able to associate dog and wolf haplotypes of modern and ancient origin and infer the time to their common ancestor. Intriguingly, the large data assemblage enabled us to genetically assign the Kesslerloch specimen KSL-189 as dog-like in contrast to the study of Loog and colleagues41, in which the canid was included in genetic examination of ancient wolves. Due to the close genetic relatedness to the Bonn-Oberkassel canid, a widely accepted early dog, combined with dietary analysis16 (Fig. 2A,B) and morphological study32, we revised the status of KSL-189 as dog-like.
In general, our TMRCA estimates are in line with previous findings placing a common ancestor of dogs and wolves in the late Pleistocene (oldest clade dog A—wolf: ~ 46 ka BP24,25,43). It should be emphasized that the TMRCA is not equivalent to a population split44, nor does it represent the actual onset of domestication; it can, however, be used to assess an upper time limit of such events. We attribute slight discrepancies between the previously published TRMCAs and our new estimates to varying sampling regimes and parameter settings used in the different studies24,25,41,43. The most recent common ancestor of all closely contemporaneous samples from Gnirshöhle dates back to 135 ka BP (including GN-192). This is an intriguing finding for several reasons. First, Magdalenian mitochondrial genomes reconstructed from specimens originating from a single cave in southwestern Germany captured almost the entire breadth of genetic diversity of all contemporary and ancient dogs and most wolves. Second, the newly generated mitochondrial genomes from the Gnirshöhle canids introduce a yet unrecognized, ancient mitochondrial canid lineage that had survived into the Magdalenian. Several authors have now proposed that mitochondrial dog lineages in prehistoric Europe were replaced by expanding lineages arriving from the East24,45,46. This fate was possibly shared by the Gnirshöhle canids as an example of diversity that was replaced by the incoming lineages represented by today's dog clade A43. The age of the Gnirshöhle canids exceeds those of the samples analyzed by Frantz and colleagues24, and implies that with more samples, we may still discover divergent lineages representative of early dogs47, a prospect that helps to finally derive a more nuanced picture of modern wolf’s domestication history (see also Bergström and colleagues46).
To better explain the observed genetic diversity, we expanded our focus to include niche and dietary reconstructions to assess the trophic behavior, as well as a comparison with two other Magdalenian cave sites, Hohle Fels and Kesslerloch, located less than 100 km away. With respect to their isospace, all specimens from the Gnirshöhle were assigned to the trophic niche B that also includes the Kesslerloch dogs16, a finding that is in contrast to the detected closer genetic affinity of the Gnirshöhle canids to modern and ancient wolves (Fig. 4) and warrants further explanation. Carbon and nitrogen isotopes in bone collagen accumulate over several years before tissue turnover occurs48,49,50, therefore, tissue samples rather than indicating seasonal or single dietary events reflect a food resource that would have been regularly available over a long period of time. The newly generated data allowed us to test three hypotheses explaining the diversity patterns in Magdalenian canids from the Hegau Jura: (1) the refugium hypothesis, (2) the ecomorph hypothesis, and 3) the domestication hypothesis (Fig. 5).
Previous studies of genetic diversity in refugia have shown that a direct correlation exists between the size of the refugium and the observed diversity (e.g.,51). The larger an area, the more genetic variation could have accumulated consequently resulting in higher estimates of the genetic diversity of the species under consideration. If we accept the Hegau Jura region as a potential canid refugium, we should observe lower diversity in the mitochondrial genomes, as it is rather small compared to, for instance, the Iberian Peninsula refugium5,51,52. Furthermore, a retreat into glacial refugia would not necessarily require a shift in the trophic niche of the focal species, and hence under the refugium hypothesis, we would predict a diet composition similar to that of pre-LGM canids from southwestern Germany. Results of dietary reconstructions from the pre-LGM periods of the Swabian Jura17 have shown that all studied canids belonged exclusively to one group with high δ15N values that primarily fed on megafauna. While we observed the same pattern in the Magdalenian ‘wolf’ niche (niche A), canids from Gnirshöhle behaved differently, which led us to reject the refugium hypothesis.
With regard to the ecomorph and domestication hypotheses, we are not able to unequivocally differentiate between either solely based on our mitochondrial DNA data. Both hypotheses are equally supported by the high genetic diversity within the Magdalenian canids in addition to its dissociation from space and time. This demonstrates the limitation of genetic analysis of the maternally inherited mitochondrial genome to appropriately examine such complex hypotheses. From the archaeozoological perspective, neither the metrics nor tooth crowding38,53 was sufficient to differentiate whether the canids of Gnirshöhle were dogs or wolves.
Considering the trophic niche observed among the Gnirshöhle canids being defined as a Late Pleistocene ecomorph and bearing in mind the properties of a surviving wolf ecomorph from the pre-LGM, we predict a restriction to a similar diet, as was observed for an Eastern Beringia Pleistocene ecomorph9. All Gnirshöhle canids showed signals indicating they consumed a low δ15N protein diet (niche B), while those from the two nearby cave sites, Kesslerloch and Hohle Fels, showed a high δ15N protein dietary source like those consumed by other canids16. Thus it is unlikely that the Gnirshöhle canids represent a surviving wolf ecomorph with a similar high δ15N protein diet as the pre-LGM wolves17. However, it is possible that canids consumed reindeer and small game and may be defined by a new type of ecomorph, similar to the hypothesis proposed by Perri10 concerning canids from the Gravettian open-air site Předmostí (Czech Republic). Members of the Magdalenian niches A and B show similar reconstructed diets compared to those canid groups analyzed from Předmostí26. As Bocherens and colleagues26 showed, the ‘Pleistocene wolf’ group had free access to all dietary resources and consumed mostly megafauna, while the ‘Palaeolithic dog’ group consumed primarily reindeer26,27. We observed a dietary distribution in the Magdalenian isospace within niches A and B, similar to the Předmostí canid groups16. Ultimately, the ecomorph hypothesis becomes a reasonable explanation for the observed genetic and isotopic patterns, but it fails to answer the question of why the Gnirshöhle canids would form a separate ecomorph while a high δ15N protein diet source was likely available nearby.
Besides environmental changes, humans can be a major driving factor to create ecological niches, and thus, domestication can lead to an ecomorph shaped by them54,55. Direct dating of the Gnirshöhle samples implied that canids could have lived in close vicinity of Magdalenian people, occupying the Hegau Jura, and subsequently adapted to a restricted diet, possibly under human influence. Thus, we consider the Gnirshöhle canids to likely represent an early phase in wolf domestication—facilitated by humans actively providing a food resource for those early domesticates (niche B16). Moreover, the high mitochondrial genome diversity could be explained by the fact that Magdalenian people would have arbitrarily drawn individuals from a large pool of canid genetic variation in the region some 15,000 years ago. We suggest that the proximity Magdalenian hunter-gatherers is the most parsimonious explanation for the genetic, isotopic, and archaeozoological patterns observed in the Gnirshöhle canids.
In conclusion, future investigations should include the analyses performed in our study, in addition to others for more detailed diet reconstructions (i.e. microwear analysis, amino acid isotope analysis), identification of the specimens’ geographic origin or range (i.e., sulfur, lead and strontium isotope analysis), as well as including nuclear genomes to further decipher details of the wolf’s domestication history. Intriguingly, a recent study focusing on the analysis of nuclear genomes of various ancient dogs suggested a single origin of modern dogs, but it failed to provide a geographic location for such an event46. While we cannot address the question of the domestication event’s singularity, our results support the hypothesis that the Hegau Jura was a potential center of early European wolf domestication. Such a scenario becomes plausible considering a close proximity of canids and humans thereby introducing a controlled, or at least a restrictive diet. This would foster their differentiation from wild conspecifics and may thus constitute a driving factor in the process of domesticating wolves. Lastly, we reiterated the importance of multidisciplinary approaches to investigate the origin of modern dogs, serving as a model for similar studies in the future.
Material and methods
Archaeological context and sample information
The Gnirshöhle (GN) is a small cave with two chambers (GN I and GN II) situated in the Bruder Valley close to Engen in the Hegau Jura of southwestern Germany. The new dates of the Gnirshöhle canid remains range from 15.5 to 15 ka cal BP (Figure S1, Table S1, Supplementary Note 3). Nearby are two other Magdalenian sites, Petersfels56,57,58, and Drexlerloch59. All three sites contain faunal remains dominated by reindeer and horses; in addition, Gnirshöhle provides a considerable number of canids34 (Supplementary Note 1). These remains are well-suited to explore early interactions between humans and canids. To shed light on these scenarios, we combined diverse datasets generated from various disciplines of archaeological sciences (archaeozoology, stable isotope analyses, and paleogenetics).
The archaeozoological investigations included 65 canid remains from Gnirshöhle (GN I: n = 60, GN II: n = 5) of which only two specimens were measurable, namely a mandible (GN-999, Figure S2, Tables S2 and S3), and a maxilla (GN-192, Figure S3, Tables S2 and S3). Well-preserved samples (larger than 0.5 cm, Figure S4) were selected for isotopic and genetic analysis. New isotopic data (δ13C, δ15N) were obtained for six out of eight canid remains of Gnirshöhle, one red fox from Bockstein Cave (BS-21), and one pre-LGM wolf from Hohle Fels (HF-1712) in the Swabian Jura (Supplementary Note 1, Table S1). Additionally, for the isotopic analysis, we included previously published isotope values from the specimens from Hohle Fels and Kesslerloch, which were genetically analyzed in this study. The palaeogenetic analyses focused on the mitochondrial genomes of canid samples from Gnirshöhle (n = 8, Fig. 3). We further generated mitochondrial genomes from additional canid remains (Supplementary Note 1), namely Hohle Fels (Swabian Jura, n = 11) including samples from the Aurignacian to the Late Paleolithic (Fig. 1): Umingmak (n = 5), a Palaeoeskimo site in northwestern Canada, Le Parc du Château (Auneau, n = 2), a Mesolithic site in France, and Frankfurt a. M. (Germany, n = 3), assigned either to Roman (n = 1) or Medieval periods (n = 2) and combined those with publicly available canid mitochondrial genomes (Table S9), including the specimens from Bonn-Oberkassel and Kesslerloch25,41. Although some of the analyzed samples come from outside of the region and period central to this manuscript, we included them in this paper to get as high as available to us diversity covered in our phylogenetic inference. Since canid mitochondrial lineages are widely geographically and temporally spread, the inclusion of North American wolves allowed us additionally to confirm the genetic continuity within the region.
Archaeozoology: morphological and metrical methods
Archaeozoological identification of species and morphological classification of canid remains were done by taking measurements after von den Driesch36 and comparing them with the archaeozoological reference collection housed at the University of Tübingen, Germany. If fragmented specimens did not allow measurement, the size of the canid remains were compared to wolf or dog specimens from the reference collection and assigned as being either wolf-like, dog-like or Canis sp.
For metrical analysis of the GN-999 mandible, we focused on two measurements: the length of the tooth row (ALP1M3) and the maximal length of the first molar (M1) (CLM1 from mesial to distal). To enable a two-dimensional classification, we plotted the two measurements against each other and compared them with published metrical data from Germonpré and colleagues35 (‘Paleolithic dogs’, n = 31, ALP1M3 = 94.9 ± 3.5 mm, CLM1 = 28.9 ± 1.5 mm; ‘Pleistocene wolves’, n = 36, ALP1M3 = 101.2 ± 2.9 mm, CLM1 = 29.9 ± 1.3 mm; ‘Archaic dogs’, n = 27, ALP1M3 = 75.9 ± 3.6 mm, CLM1 = 22.5 ± 1.2 mm; modern ‘Northern wolves’, n = 35, ALP1M3 = 99.3 ± 3.8 mm, CLM1 = 29.5 ± 1.6 mm), and Pleistocene wolves (Nobis60, n = 1; Boessneck and von den Driesch61, n = 1; Napierala33, n = 9; Napierala and Uerpmann32, n = 1; Münzel62, n = 1, Modern wolves60, n = 4), post-LGM dogs60, n = 4) in addition to metric data from modern wolves (n = 3) from the archaeozoological reference collection mentioned above (Table S4). Other standardized measurements for GN-999 and the maxilla GN-192 after Von den Driesch36 are given in Supplementary Note 2.
Stable isotopes: elemental and isotopic analysis
We performed new isotopic analyses from bone collagen for seven samples of canids from Gnirshöhle (Canis sp., n = 6), Hohle Fels (Canis lupus, n = 1) and Bockstein (Vulpes vulpes, n = 1). Collagen extraction followed the protocol outlined by Bocherens and colleagues48 and is further detailed explained in Supplementary Note 4. The process was performed in the Biogeology working group laboratory at the University of Tübingen (Germany). Isotopic measurements of collagen were undertaken in duplicate at the Institute of Environmental Science and Technology of the Universitat Autònoma de Barcelona (ICTA-UAB) using a Thermo Flash 1112 (Thermo ScientificVC) elemental analyzer coupled to a Thermo Delta V Advantage mass spectrometer with a Conflo III interface. All details for elemental and isotopic analysis are provided in Supplementary Note 4.
The isotopic ratios are expressed using the δ (delta) value as follows:
Stable isotopes: niche modeling and dietary reconstruction
To reconstruct the niches of the sampled canids, we included newly analyzed δ13C and δ15N values of six canids from Gnirshöhle, one newly analyzed red fox from Bockstein, and published isotopic values of 14 canids16,40, and applied a multivariate cluster analysis (using JMP 14) to the δ13C and δ15N isotopic values of all canids. We then used R63 and the R package SIBER to calibrate the core niches (standard ellipse area = SEA) out of the clusters64. The core niche depicts the center of a niche that is calculated by using a Bayesian most-likelihood estimation and explains roughly 40% of the expected variability64. This method is more reliable for analyzing small sample sizes and recommended for niche interpretations by Jackson and colleagues64. For dietary reconstructions, we utilized the same prey groups that were defined by Baumann and colleagues16: ‘Megaherbivores’, ‘Ungulates’, and ‘Small mammals’ that were constructed from a database of 91 carbon and nitrogen stable isotopic values from herbivores (Table S5). To reconstruct the proportions of different prey groups in the protein fraction of the canids’ diet, we used MixSIAR65. More detailed information is presented in the Supplementary Note 4.
Palaeogenetics: ancient DNA laboratory workflow and sequence analysis
All pre-amplification steps, i.e. the DNA extraction and library preparation, were performed in clean room facilities at the University of Tübingen fulfilling all requirements for ancient DNA work66,67. Amplification and mitochondrial canid DNA enrichment steps were performed in a separate laboratory also located at the University of Tübingen. Lastly, DNA libraries were either sequenced at the Max-Planck-Institute for Science of Human History in Jena, Germany, or at the Functional Genomic Center Zürich, Zürich, Switzerland. Detailed information can be found in Supplementary Note 5.
We used the Efficient Ancient Genome Reconstruction (EAGER) pipeline68, version 1.92.37 for bioinformatic analysis of the sequencing data (Supplementary Note 5). The quality of the sequencing reads was estimated by FastQC tool69 and the adapters were trimmed by AdapterRemoval70 version 2.2.1a, both tools are integrated in the EAGER pipeline. MarkDuplicates v2.15.0 (Picard Tools) was applied to remove duplicates for genome reconstruction68. To demonstrate the authenticity of ancient DNA reads, the program MapDamage71 is utilized by the EAGER pipeline to estimate the distribution and frequency of any misincorporations at the 3′- and 5′-ends of the DNA reads. Potential contaminated DNA reads in sequencing data for the individuals GN-14 and GN-106 were identified by a low frequency of misincorporations estimated by MapDamage71. To overcome potentially false reconstruction of mitochondrial genomes due to contamination, we applied PMD-tools72 to separate endogenous ancient DNA reads from modern contaminant reads with a threshold PMD score of three. For these two specimens, filtered reads were then used for mitochondrial genome reconstruction.
Paleogenetics: phylogenetic analysis and timescale estimation
We used canid mitochondrial genome data published in previous studies24,25,41,43,73 to reconstruct a comprehensive phylogeny. In total, a multiple sequence alignment database of 244 genomes was constructed (MAFFT74), including 221 mitochondrial genomes recruited for our comparative dataset24,25,41,43,73 and the reconstructed genomes of our study. Before the phylogenetic tree reconstruction, we defined the best-fit model for our data75, using IQ-TREE76, to obtain the highest statistically supported phylogenetic tree. Subsequently, a phylogenetic tree was generated employing IQ-TREE76 using a maximum likelihood approach with an estimation of 10,000 bootstrap replications.
In order to estimate time-aware phylogeny in coalescent framework, we excluded the coyote mitogenomes, as well as three modern Tibetan/Mongolian wolves from China (NC011218, EU442884 and FJ032363) and one ancient Siberian wolf (MK936996) that in preliminary runs revealed an unexpectedly old divergence time, which could be attributed either to genetic isolation or comparatively stronger selective forces. The alignment of 236 mitogenomes was partitioned using PartitionFinder77 2.1.1 into four partitions: (1) protein coding, gene position one (3782 nt length, TRN + I substitution model), (2) position two (3780 nt, TRN + G), (3) position three with rRNA and tRNA (7832 nt, TRN + I), and (4) non-coding (1050 nt, HKY + G). Bayesian phylogenetic and timescale estimation was performed using BEAST78 1.10.4. Sample ages were used as tipdates for molecular clock calibration. Undated samples were assigned uniform age prior (0-100ky). Uncorrelated lognormal relaxed clock for each partition and Bayesian SkyGrid population model79 were used, as supported by the data. The MCMC chain was run for 200 M steps with sampling every 20,000th step. The first 20 M steps were removed as burn-in. Convergence and mixing were inspected (all ESS exceeded 100) and the SkyGrid plot was generated in Tracer80 v 1.7.1. Maximum Clade Credibility trees were generated in TreeAnnotator (part of the BEAST package) and visualized in FigTree81 1.4.2.
Palaeogenetics: estimation of genetic diversity
To investigate the genetic diversity in more detail, the mitochondrial genomes from Kesslerloch and Gnirshöhle were compared to modern dog and wolf mitochondrial DNA sequences24,25,41,43,73 assigned into four canid population groups: Kesslerloch (KSL), Gnirshöhle (GN), modern dogs, and modern wolves (Table S7 and S8). We calculated the nucleotide diversity for each predefined group via DnaSP82 v5 after excluding all sites with gaps for each group individually. The program estimated the nucleotide diversity Pi (π), sampling variance, and the standard deviation, defined by the square root of the variance82, according to well-established statistical algorithms42,83.
Data Availability
The data will become available upon publication. The genetic data (raw sequencing data) can be found on NCBI (BioProject ID: PRJNA703747).
References
Darwin, C. On the Origin of Species by Means of Natural Selection Vol. 167 (John Murray, London, 1859).
Kahlke, R.-D. The origin of Eurasian mammoth faunas (Mammuthus–Coelodonta faunal complex). Quatern. Sci. Rev. 96, 32–49 (2014).
Ashcroft, M. B. Identifying refugia from climate change. J. Biogeogr. 37, 1407–1413 (2010).
Tarkhnishvili, D., Gavashelishvili, A. & Mumladze, L. Palaeoclimatic models help to understand current distribution of Caucasian forest species. Biol. J. Lin. Soc. 105, 231–248 (2012).
Hewitt, G. The genetic legacy of the Quaternary ice ages. Nature 405, 907–913 (2000).
Lumibao, C. Y., Hoban, S. M. & McLachlan, J. Ice ages leave genetic diversity ‘hotspots’ in Europe but not in Eastern North America. Ecol. Lett. 20, 1459–1468 (2017).
Williams, E. E. in Evolutionary Biology 47–89 (Springer, 1972).
Peters, G. in Handbuch der Säugetiere Europas: Raubsäuger (Teil 1) (eds M. Stubbe & F. Krapp) 47–106 (AULA-Verlag, 1993).
Leonard, J. A. et al. Megafaunal extinctions and the disappearance of a specialized wolf ecomorph. Curr. Biol. 17, 1146–1150 (2007).
Perri, A. A wolf in dog’s clothing: Initial dog domestication and Pleistocene wolf variation. J. Archaeol. Sci. 68, 1–4. https://doi.org/10.1016/j.jas.2016.02.003 (2016).
Weniger, G.-C. in The Pleistocene Old World: Regional Perspectives (ed Olga Soffer) 201–215 (Springer, 1987).
Weniger, G.-C. Magdalenian settlement and subsistence in South-west Germany. Proc. Prehist. Soc. 53, 293–307. https://doi.org/10.1017/S0079497X0000623X (1987).
Taller, A., Bolus, M. & Conard, N. The Magdalenian of Hohle Fels Cave and the resettlement of the Swabian Jura after the LGM. in Modes de Contacts et de Déplacements au Paléolithique Eurasiatique/Modes of Contact and Mobility During the Eurasian Palaeolithic. ERAUL Vol. 140, 383–399 (2014).
Maier, A. Population and settlement dynamics from the Gravettian to the Magdalenian. Mitteilungen Gesellschaft Urgeschichte 26, 83–101 (2017).
Hulme-Beaman, A., Dobney, K., Cucchi, T. & Searle, J. B. An ecological and evolutionary framework for commensalism in anthropogenic environments. Trends Ecol. Evol. 31, 633–645. https://doi.org/10.1016/j.tree.2016.05.001 (2016).
Baumann, C. et al. Dietary niche partitioning among Magdalenian canids in southwestern Germany and Switzerland. Quatern. Sci. Rev. 227, 106032 (2020).
Baumann, C., Bocherens, H., Drucker, D. G. & Conard, N. J. Fox dietary ecology as a tracer of human impact on Pleistocene ecosystems. PLoS ONE 15, e0235692. https://doi.org/10.1371/journal.pone.0235692 (2020).
Smith, B. D. A cultural niche construction theory of initial domestication. Biol. Theory 6, 260–271 (2011).
Zeder, M. A. Domestication as a model system for niche construction theory. Evol. Ecol. 30, 325–348 (2016).
Belyaev, D. K., Plyusnina, I. Z. & Trut, L. N. Domestication in the silver fox (Vulpes fulvus Desm): Changes in physiological boundaries of the sensitive period of primary socialization. Appl. Anim. Behav. Sci. 13, 359–370. https://doi.org/10.1016/0168-1591(85)90015-2 (1985).
Germonpré, M., Láznickova-Galetova, M., Sablin, M. V. & Bocherens, H. in Hybrid Communities, Biosocial Approaches to Domestication and Other Trans-Species Relationships Routledge Studies in Anthropology (eds C. Stépanoff & J.-D. Vigne) (Routledge, 2018).
Thalmann, O. & Perri, A. Population Genetics 1–34 (Springer, Cham, 2018).
MacHugh, D. E., Larson, G. & Orlando, L. Taming the past: Ancient DNA and the study of animal domestication. Annu. Rev. Anim. Biosci. 5, 329–351. https://doi.org/10.1146/annurev-animal-022516-022747 (2017).
Frantz, L. A. et al. Genomic and archaeological evidence suggest a dual origin of domestic dogs. Science 352, 1228–1231 (2016).
Thalmann, O. et al. Complete mitochondrial genomes of ancient canids suggest a European origin of domestic dogs. Science 342, 871–874 (2013).
Bocherens, H. et al. Reconstruction of the Gravettian food-web at Předmostí I using multi-isotopic tracking (13C, 15N, 34S) of bone collagen. Quatern. Int. 359–360, 211–228. https://doi.org/10.1016/j.quaint.2014.09.044 (2015).
Prassack, K. A., DuBois, J., Lázničková-Galetová, M., Germonpré, M. & Ungar, P. S. Dental microwear as a behavioral proxy for distinguishing between canids at the Upper Paleolithic (Gravettian) site of Předmostí, Czech Republic. J. Archaeol. Sci. 115, 105092 (2020).
Wilczyński, J. et al. Friend or foe? Large canid remains from Pavlovian sites and their archaeozoological context. J. Anthropol. Archaeol. 59, 101197. https://doi.org/10.1016/j.jaa.2020.101197 (2020).
Street, M., Napierala, H. & Janssens, L. The late Paleolithic dog from Bonn-Oberkassel in context. in The Late Glacial Burial from Oberkassel Revisited 253–274. (Verlag Phillip von Zabern, Darmstadt, 2015).
Janssens, L. et al. A new look at an old dog: Bonn-Oberkassel reconsidered. J. Archaeol. Sci. 92, 126–138. https://doi.org/10.1016/j.jas.2018.01.004 (2018).
Rütimeyer, L. Die Knochenhöhle von Thayingen bei Schaffhausen. (F. Vieweg & Sohn, 1875).
Napierala, H. & Uerpmann, H.-P. A ‘new’ palaeolithic dog from central Europe. Int. J. Osteoarchaeol. 22, 127–137. https://doi.org/10.1002/oa.1182 (2012).
Napierala, H. Die Tierknochen aus dem Kesslerloch: Neubearbeitung der paläolithischen Fauna (Baudepartement des Kantons Schaffhausen, Kantonsarchäologie Schaffhausen, 2008).
Albrecht, G., Drautz, D. & Kind, J. Eine Station des Magdalénien in der Gnirshöhle bei Engen- Bittelbrunn im Hegau. Archäol. Korrespondenzblatt 7, 161–179 (1977).
Germonpré, M., Lázničková-Galetová, M., Losey, R. J., Räikkönen, J. & Sablin, M. V. Large canids at the Gravettian Předmostí site, the Czech Republic: The mandible. Quatern. Int. 359–360, 261–279. https://doi.org/10.1016/j.quaint.2014.07.012 (2015).
Von den Driesch, A. A Guide to the Measurement of Animal Bones from Archaeological Sites: as Developed by the Institut für Palaeoanatomie, Domestikationsforschung und Geschichte der Tiermedizin of the University of Munich. Vol. 1 (Peabody Museum Press, 1976).
Benecke, N. Archäozoologische Studien zur Entwicklung der Haustierhaltung. (Akademie Verlag, 1994).
Ameen, C. et al. A landmark-based approach for assessing the reliability of mandibular tooth crowding as a marker of dog domestication. J. Archaeol. Sci. 85, 41–50. https://doi.org/10.1016/j.jas.2017.06.014 (2017).
Bocherens, H., Drucker, D., Billiou, D. & Moussa, I. Une nouvelle approche pour évaluer l'état de conservation de l'os et du collagène pour les mesures isotopiques (datation au radiocarbone, isotopes stables du carbone et de l'azote). l'Anthropologie 109, 557–567 (2005).
Bocherens, H. et al. Isotopic evidence for dietary ecology of cave lion (Panthera spelaea) in North-Western Europe: Prey choice, competition and implications for extinction. Quatern. Int. 245, 249–261. https://doi.org/10.1016/j.quaint.2011.02.023 (2011).
Loog, L. et al. Ancient DNA suggests modern wolves trace their origin to a late Pleistocene expansion from Beringia. Mol. Ecol. 1–15 (2019).
Nei, M. & Miller, J. C. A simple method for estimating average number of nucleotide substitutions within and between populations from restriction data. Genetics 125, 873–879 (1990).
Botigué, L. R. et al. Ancient European dog genomes reveal continuity since the Early Neolithic. Nat. Commun. 8, 1–11 (2017).
Nichols, R. Gene trees and species trees are not the same. Trends Ecol. Evol. 16, 358–364 (2001).
Ollivier, M. et al. Dogs accompanied humans during the Neolithic expansion into Europe. Biol. Lett. 14, 20180286 (2018).
Bergström, A. et al. Origins and genetic legacy of prehistoric dogs. Science 370, 557–564 (2020).
Larson, G. et al. Rethinking dog domestication by integrating genetics, archeology, and biogeography. Proc. Natl. Acad. Sci. 109, 8878 (2012).
Bocherens, H. et al. Paleobiological implications of the isotopic signatures (13C, 15N) of fossil mammal collagen in Scladina Cave (Sclayn, Belgium). Quatern. Res. 48, 370–380 (1997).
Bocherens, H. & Drucker, D. Trophic level isotopic enrichment of carbon and nitrogen in bone collagen: Case studies from recent and ancient terrestrial ecosystems. Int. J. Osteoarchaeol. 13, 46–53. https://doi.org/10.1002/oa.662 (2003).
Krajcarz, M. T., Krajcarz, M. & Bocherens, H. Collagen-to-collagen prey-predator isotopic enrichment ( Δ 13C, Δ 15N) in terrestrial mammals - A case study of a subfossil red fox den. Palaeogeogr. Palaeoclimatol. Palaeoecol. 490, 563–570. https://doi.org/10.1016/j.palaeo.2017.11.044 (2018).
von Seth, J., Niemann, J. & Dalén, L. in Paleogenomics 393–418 (Springer, 2018).
Orlando, L. in Paleogenomics 325–351 (Springer, 2018).
Janssens, L. A. A. From Wolf to Dog, Uitgever Niet Vastgesteld (2019).
Zeder, M. A. The domestication of animals. J. Anthropol. Res. 68, 161–190 (2012).
Zeder, M. A. Pathways to animal domestication. in Biodiversity in Agriculture: Domestication, Evolution, and Sustainability, 227–259 (2012).
Albrecht, G. Magdalénien-Inventare vom Petersfels: Siedlungsarchäologische Ergebnisse der Ausgrabungen 1974–1976. Vol. 6 (Verlag Archaeologica Venatoria, 1979).
Albrecht, G. in Urgeschichte in Baden-Württemberg (ed Hansjürgen Müller-Beck) 331–353 (Theiss Verlag, 1983).
Albrecht, G. & Berke, H. in De la Loire à l'Oder BAR International Series (ed Marcel Otte) 465–473 (1988).
Albrecht, G., Wong, G. L. & Münzel, S. C. in „All der holden Hügel ist keiner mir fremd…“ Festschrift zum 65. Geburtstag von Claus-Joachim Kind. (eds M. Baales & C. Pasda) 301–310 (Archaeologica Venatoria, 2019).
Nobis, G. Die Wildsäugetiere in der Umwelt des Menschen von Oberkassel bei Bonn und das Domestikationsproblem von Wölfen im Jungpaläolithikum. in Bonner Jahrbücher, 367–376 (1986).
Boessneck, J., von den Driesch, A., Lepiksaar, J., Riek, G. & Storch, G. Das Paläolithikum der Brillenhöhle bei Blaubeuren (Schwäbische Alb) II: Die Jungpleistozänen Tierknochenfunde aus der Brillenhöhle. (Verlag Müller & Gräff, 1973).
Münzel, S. C. in Geißenklösterle: Chronostratigraphie, Paläoumwelt und Subsistenz im Mittel- und Jungpaläolithikum der Schwäbischen Alb (eds Nicholas J. Conard, M. Bolus, & Susanne C. Münzel) 147–327 (Kerns Verlag, 2019).
Team, R. C. R: A Language and Environment for Statistical Computing, https://www.R-project.org.
Jackson, A. L., Inger, R., Parnell, A. C. & Bearhop, S. Comparing isotopic niche widths among and within communities: SIBER—stable isotope Bayesian ellipses in R. J. Anim. Ecol. 80, 595–602. https://doi.org/10.1111/j.1365-2656.2011.01806.x (2011).
Stock, B. C. & Semmens, B. X. MixSIAR GUI User Manual v3.1. (2016).
Cooper, A. & Poinar, H. N. Ancient DNA: Do it right or not at all. Science 289, 1139–1139 (2000).
Knapp, M. & Hofreiter, M. Next generation sequencing of ancient DNA: Requirements, strategies and perspectives. Genes 1, 227–243 (2010).
Peltzer, A. et al. EAGER: Efficient ancient genome reconstruction. Genome Biol. 17, 60 (2016).
Andrews, S. Babraham Bioinformatics (Babraham Institute, Cambridge, 2010).
Schubert, M., Lindgreen, S. & Orlando, L. AdapterRemoval v2: Rapid adapter trimming, identification, and read merging. BMC Res. Notes 9, 1–7 (2016).
Ginolhac, A., Rasmussen, M., Gilbert, M. T. P., Willerslev, E. & Orlando, L. mapDamage: Testing for damage patterns in ancient DNA sequences. Bioinformatics 27, 2153–2155 (2011).
Skoglund, P. et al. Separating endogenous ancient DNA from modern day contamination in a Siberian Neandertal. Proc. Natl. Acad. Sci. 111, 2229–2234 (2014).
Skoglund, P., Ersmark, E., Palkopoulou, E. & Dalén, L. Ancient wolf genome reveals an early divergence of domestic dog ancestors and admixture into high-latitude breeds. Curr. Biol. 25, 1515–1519 (2015).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: Improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Kalyaanamoorthy, S., Minh, B. Q., Wong, T. K., von Haeseler, A. & Jermiin, L. S. ModelFinder: Fast model selection for accurate phylogenetic estimates. Nat. Methods 14, 587–589 (2017).
Nguyen, L.-T., Schmidt, H. A., Von Haeseler, A. & Minh, B. Q. IQ-TREE: A fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 32, 268–274 (2015).
Lanfear, R., Frandsen, P. B., Wright, A. M., Senfeld, T. & Calcott, B. PartitionFinder 2: New methods for selecting partitioned models of evolution for molecular and morphological phylogenetic analyses. Mol. Biol. Evol. 34, 772–773 (2017).
Suchard, M. A. et al. Bayesian phylogenetic and phylodynamic data integration using BEAST 1.10. Virus Evolut. 4, vey016 (2018).
Gill, M. S. et al. Improving Bayesian population dynamics inference: A coalescent-based model for multiple loci. Mol. Biol. Evol. 30, 713–724 (2013).
Rambaut, A., Drummond, A. J., Xie, D., Baele, G. & Suchard, M. A. Posterior summarization in Bayesian phylogenetics using Tracer 1.7. Syst. Biol. 67, 901 (2018).
Rambaut, A. FigTree 1.4. 2 Software. (Institute of Evolutionary Biology, Univ. Edinburgh, 2014).
Librado, P. & Rozas, J. DnaSP v5: A software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25, 1451–1452 (2009).
Nei, M. Molecular Evolutionary Genetics. (Columbia University Press, 1987).
Acknowledgements
We are grateful to Prof. Dr. Hans-Peter Uerpmann, who helped with morphological issues. We are also thankful for the financial and technical support to Prof. Dr. Johannes Krause, head of the Max Planck Institute for the Science of Human History (Jena, Germany), who supported the laboratory work and DNA sequencing of canid samples included in this study. Furthermore, we want to thank Dr. Christoph Wißing and Peter Tung for technical support in the isotope laboratory as well as Angel Blanco-Lapaz for photographing and optical editing of the finds (University of Tübingen). Next, we appreciate the map design performed by Corina Steiner (IEM, Zurich, Switzerland). We would like to thank the collaborators of Zooarchaeology and Palaeogenetic working groups for their helpful support at the department of archaeological sciences (University of Tübingen), the Biogeology working group of the department of geology (University of Tübingen), and the Paleogenetic working group at the Institute of Evolutionary Medicine (University of Zurich). For financial support, we are thankful to the UNESCO (Stipend, CB), Senckenberg (CB, HB, VJS), to the Regierungspräsidium Stuttgart, Landesamt für Denkmalpflege Baden-Württemberg (SCM, CB), to the University of Tübingen (CB, SP, HB), and to the University of Zurich’s University Research Priority Program “Evolution in Action: From Genomes to Ecosystems” (VJS). Additionally, we thank the National Science Center in Poland (OT, 2017/26/E/NZ5/00851; MM, 2018/31/B/HS3/01464) for their financial support. Lastly, we appreciate the reviewers for their critical and helpful suggestions leading to an improvement of the paper as well as the university library of Tübingen as well as the University of Zurich for funding the open access publication.
Author information
Authors and Affiliations
Contributions
C.B., S.P., S.C.M., O.T., and V.J.S. conceived and designed the study. V.J.S., H.B., and O.T. supervised the study. S.C.M., T.R.F., G.A., C.J.K., C.V., C.L., N.J.C., D.G.D., and L.G. provided samples and archaeological context. S.C.M. and C.B. performed metrical and morphological analyses. C.B. and H.B. provided isotopic sampling, analysis, dietary reconstruction, and niche modeling. S.P., M.M., E.R., O.T., T.R.F., and V.J.S. performed genetic sampling, DNA extraction, library preparation of sequencing libraries, bioinformatics, and population genetic analysis. Selection for direct dating of the finds was done by C.B. and S.C.M. and financially supported by V.J.S., C.J.K. and H.B. C.B., S.P., S.C.M., A.B., O.T., V.J.S., and M.M. wrote the manuscript with input from all authors. S.C.M., S.P., C.B., C.V., C.L., L.G., O.T., and V.J.S. wrote the supplementary notes with input from all authors. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Baumann, C., Pfrengle, S., Münzel, S.C. et al. A refined proposal for the origin of dogs: the case study of Gnirshöhle, a Magdalenian cave site. Sci Rep 11, 5137 (2021). https://doi.org/10.1038/s41598-021-83719-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-83719-7
This article is cited by
-
The paleo-synanthropic niche: a first attempt to define animal’s adaptation to a human-made micro-environment in the Late Pleistocene
Archaeological and Anthropological Sciences (2023)
-
Grey wolf genomic history reveals a dual ancestry of dogs
Nature (2022)
-
Domestication Through Dingo Eyes: An Australian Perspective on Human-Canid Interactions Leading to the Earliest Dogs
Human Ecology (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.