Abstract
Milk small extracellular vesicles (sEV) contain proteins that provide potential information of host physiology and immunology. Bovine leukemia virus (BLV) is an oncogenic virus that causes progressive B-cell lymphosarcoma in cattle. In this study, we aimed to explore the proteomic profile of milk sEV from BLV-infected cattle compared with those from uninfected cattle. Milk sEV were isolated from three BLV-infected and three uninfected cattle. Proteomic analysis was performed by using a comprehensive nanoLC-MS/MS method. Furthermore, gene ontology (GO) annotation and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways were used to evaluate the candidates for uniquely or differentially expressed proteins in milk sEV from BLV-infected cattle. Proteomic analysis revealed a total of 1330 common proteins in milk sEV among BLV-infected cattle, whereas 118 proteins were uniquely expressed compared with those from uninfected cattle. Twenty-six proteins in milk sEV were differentially expressed proteins more than two-fold significant difference (p < 0.05) in BLV-infected cattle. GO and KEGG analyses indicated that the candidates for uniquely or differentially expressed proteins in milk sEV had been involved in diverse biological activities including metabolic processes, cellular processes, respond to stimulus, binding, catalytic activities, cancer pathways, focal adhesion, and so on. Taken together, the present findings provided a novel insight into the proteomes of milk sEV from BLV-infected cattle.
Similar content being viewed by others
Introduction
Extracellular vesicles (EV) are membranous particles, secreted by a wide variety of cells found in all biological fluids in humans and animals such as blood, amniotic fluid, ascitic fluid, urine, saliva, tears, and milk1. There is still some variability in the different classes of EV that included exosomes, ectosomes or shedding microvesicles, apoptotic bodies, and other EV subsets according to their size, biogenesis, and releasing pathway2. Among all classes of EV, one of the EV is classified as small EV (sEV), so called exosomes (30–150 nm in diameter)1. The International Society for Extracellular Vesicles (ISEV) has been established to accelerate research activities including all classes of EV. Recently, ISEV suggested that the term “sEV” should be used instead of “exosomes” in the Minimal Information for Studies of Extracellular Vesicles guidelines 2018 (MISEV2018)3.
Milk is a diverse source of sEV that contains proteins, microRNAs (miRNAs), mRNAs, DNA, and lipids that play an important role in many biological activities including cell growth, development, immune modulation and regulation4,5. For example, sEV derived from beneficial bacteria in human breast milk has been involved in the transfer of gut microbiota from mother to infant6. More recent study described that bovine milk sEV also contained proteins, miRNAs, mRNAs, DNA, and lipids that were considered to transport biologically active cargos from donor to recipient cells for exchanging genetic information7. Over the last two decades, proteomic analysis has been widely used to detect changes of proteins in milk sEV in relation with many physiological information8. A recent study reported that milk sEV contained distinctive proteins that provided the potential information of mammary physiology of cattle9. Moreover, the proteomic analysis of milk sEV provided pathological information of disease of cattle. For example, a previous study reported that miRNAs such as bta-miR-142-5p and bta-miR-223 had been up-regulated in milk sEV from Staphylococcus aureus-infection, considered as a potential biomarker for monitoring of physiological and pathological status in cattle10.
Bovine leukemia virus (BLV) is one of the tumorigenic virus that causes enzootic bovine leukosis (EBL) characterised by B-cell lymphosarcoma and is present worldwide, including Japan11. A nationwide survey in Japan reported that approximately 40.9% of dairy cattle and 28.7% of beef cattle had antibodies against BLV12. Among cattle infected with BLV, 2–5% of cattle develop clinical signs of B-cell lymphosarcoma and 20–30% of cattle progress to persistent lymphocytosis (PL). Approximately 70% of BLV-infected cattle do not show any clinical signs and remain sub-clinically infected for life13. To date, evidence has indicated that host genetic factors may play a vital role in the stages of BLV infection: from early infection to developing of PL to lymphoma14. Very recently, our study revealed that BLV infection caused profound effects on host cellular activity resulting in changes of encapsulated mRNA in milk sEV obtained from BLV-infected cattle15. We hypothesised that there would be a probable change in encapsulating proteins in milk sEV also during BLV infection in cattle. However, to date, no study has reported on the proteomic analysis of milk sEV from BLV-infected cattle.
In this study, a comprehensive proteomic analysis in milk sEV was performed to identify the changes in protein levels in BLV-infected cattle. The proteomic analysis revealed a large number of proteins along with candidates for uniquely or differentially expressed proteins in milk sEV from BLV-infected cattle. Further, gene ontology (GO) annotation and Kyoto Encyclopaedia of Genes and Genomes (KEGG) pathways provided a new perspective to understand and unveil the pathological roles of milk sEV during BLV infection in cattle. Our study suggested that the candidates for uniquely or differentially expressed proteins in milk sEV could be the land mark for future investigations of clinical stages of BLV infection and it’s pathogenesis in cattle by using of milk sEV.
Results
Clinical status of cattle
The clinical status including BLV infection, hematology, and serum chemistry parameters of cattle were assessed and shown in Table 1. BLV provirus and BLV antibody was checked either by nested polymerase chain reaction (nested PCR) and enzyme-linked immunosorbent assay (ELISA) for the confirmation of BLV infection in cattle. Three BLV-infected cattle had high proviral load (HPL) > 30,000/105 white blood cells (WBCs) DNA along with high lactate dehydrogenase isozymes 2 and 3 (LDH2 + 3) percentage > 30%. The high WBCs and lymphocyte counts along with age were indicating either ‘Suspect’ or ‘Lymphocytic’ according to the European Community's (EC) key parameter16.
Protein concentration of milk sEV
Protein concentration of milk sEV from BLV-infected and uninfected cattle were 9.95 mg/mL and 9.92 mg/mL, respectively (p > 0.05) (Supplementary Fig. 1).
Milk sEV characterization. The peak (mode) of particle size distribution of the milk sEV from BLV-infected and uninfected cattle were showed by NTA. Representative data from B1 and U1 cattle were shown (A). Similar spherical morphology of milk sEV between BLV-infected and uninfected cattle were observed by TEM analysis. Scale bar shows 200 nm (B). Milk sEV were successfully detected by WB analysis using antibodies against sEV-surface-marker protein CD63 and internal protein HSP70 (C).
Detection of BLV genomic RNA and BLV proteins in milk sEV
To select BLV genomic RNA and BLV protein free milk sEV from BLV-infected cattle, reverse transcription-nested PCR and western blot (WB) analyses were performed. The results indicated that both BLV genomic RNA and BLV protein were not detected in milk sEV from BLV-infected cattle (Supplementary Fig. 2A and 1B).
Characterization of milk sEV
For the characterization of isolated milk sEV from BLV-infected and uninfected cattle biophysically, nanoparticle tracking analysis (NTA) and transmission electron microscopy (TEM) analysis were performed. NTA showed that the peak (mode) for particle size distribution of milk sEV from BLV-infected and uninfected cattle were 145.6 nm and 145.7 nm, respectively (p > 0.05) (Fig. 1A). TEM indicated that a similar spherical bilayer shape of milk sEV from BLV-infected and uninfected cattle was observed (Fig. 1B). WB analysis successfully detected milk sEV-surface-marker CD63 and internal protein HSP70 from BLV-infected and uninfected cattle (Fig. 1C and Supplementary Fig. 3).
Volcano plot demonstrated the expression of milk sEV proteins from BLV-infected and uninfected cattle. Up/down-regulated sEV proteins were indicated in orange and blue colours, respectively. Proteins not classified as up/down-regulated were plotted in grey colour. X-axis and Y-axis indicated a more than two-fold change (in log2 scale) and − log10 with significance (p < 0.05) (A). Heat map of hierarchical clustering indicated the candidates for differentially expressed proteins milk sEV from BLV-infected cattle. Up/down-regulated proteins were indicated by red/green colour, respectively (B). GraphPad Prism software, version 8 (https://www.graphpad.com) was used to generate the volcano plot and heat map.
Proteomic profiling of milk sEV
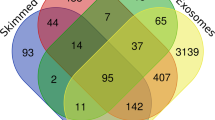
Isolated sEV were analysed using nano LC–MS/MS method to compare the milk sEV proteins of BLV-infected cattle with that of uninfected cattle. The proteomic analysis revealed a large number of proteins in milk sEV from both BLV-infected and uninfected cattle (Table 2). A total of 1330 proteins in milk sEV were found as common proteins among three BLV-infected cattle. Whereas, a total of 1512 proteins in milk sEV were found as common proteins among three uninfected cattle. From the result, a total of 1212 proteins in milk sEV were overlapped after comparing BLV-infected cattle with those of uninfected cattle (Fig. 2). Most significantly, our study identified a total of 118 proteins that were the candidates for uniquely expressed proteins in milk sEV from BLV-infected cattle (Supplement Table 1). The volcano plot indicated the differential encapsulation of milk sEV proteins between BLV-infected and uninfected cattle (Fig. 3A). The heat map showed that a total of 26 proteins in milk sEV showed more than two-fold up/down-regulation with a significant difference (p < 0.05) in BLV-infected cattle (Fig. 3B and Table 3).
Functional and protein–protein interaction (PPI) network analysis
The GO analysis was performed to evaluate the candidates for uniquely (Supplementary Fig. 4A-4D) or differentially expressed proteins (Fig. 4A–4D) to obtain a comprehensive image of the changes in proteins in milk sEV from BLV-infected cattle. The candidates for uniquely or differentially expressed proteins in milk sEV were engaged in a broad range of biological processes such as cellular process, metabolic processes, response to stimulus, signalling, biogenesis, and so on. As for the molecular function, the majority of the proteins appeared to participate in binding and catalytic activity. In the cellular component category, proteins were mainly localised in the cell, membrane, and organelles. Furthermore, the majority of the proteins were classified as cytoskeletal proteins, protein modifying enzymes, translational proteins, and metabolite interconversion enzyme.
GO analysis of the candidates for differentially expressed proteins in milk sEV from BLV-infected cattle. The candidates for differentially expressed proteins in milk sEV from BLV-infected cattle were analysed using Panther software and categorised according to the biological process (A), molecular function (B), cellular component (C), and protein class (D).
The candidates for uniquely or differentially expressed proteins in milk sEV were illustrated in the protein–protein interaction (PPI) network by STRING analysis. Importantly, the unique proteins in milk sEV have been illustrated a connectivity PPI network (Supplementary Fig. 5A) relating to several KEGG pathways including ribosome, pathways in cancer, regulation of actin cytoskeleton, chemokine signalling pathway, focal adhesion, and so on (top 10 KEGG pathways were shown in Supplementary Fig. 5B). The candidates for differentially expressed proteins in milk sEV made a strong cluster of networking in PPI (Fig. 5A) relating to several KEGG pathways including pathways in cancer, cytokine-cytokine receptor interaction, JAK-STAT signalling pathways, proteoglycans in cancer, focal adhesion, and so on (top 10 KEGG pathways were shown in Fig. 5B).
PPI network and KEGG pathway analysis. The PPI network was analysed by STRING software to evaluate the candidates for differentially expressed proteins in milk sEV from BLV-infected cattle (A). In the network, nodes and edges indicated milk sEV proteins and physical and/or functional interactions of the milk sEV proteins, respectively. Empty nodes represent the proteins of unknown three-dimensional structure, and filled nodes represent the proteins with some known or predicted three-dimensional structure. Different coloured lines between the proteins represent the various types of interactions in STRING (databases, experiments, neighbourhood, gene fusion, co-occurrence, text mining, co-expression, and homology). The candidates for differentially expressed proteins in milk sEV from BLV-infected cattle were analysed using KEGG software, and the top 10 KEGG pathways were demonstrated in the bar diagram (B).
Discussion
In this study, milk sEV were characterized by NTA, TEM, and WB analyses. The results by NTA, TEM and WB analyses were indicating the typical definition and enrichment of sEV according to the MISEV2018 guidelines3. Proteomic analysis revealed a total of 1330 proteins in milk sEV were common among three BLV-infected cattle, of which 118 proteins were the candidates for uniquely expressed proteins compared to those from uninfected cattle. The identified protein numbers in milk sEV from BLV-infected cattle were distinctive and relatively larger to those reported in previous studies9,17,18,19. These results indicated that our current study was efficiently isolated milk sEV from BLV-infected cattle which allowed the protein count in milk sEV to increase. Moreover, nanoLC-MS/MS method had detected of low-abundance compounds in small amount of proteins resulting a high-throughput protein count in milk sEV from BLV-infected cattle20.
Although the mortality rate is low, the most noticeable negative outcome of BLV infection is that cattle develop to lymphosarcoma. However, it is difficult to suspect and detect which BLV-infected cattle could be developed with lymphosarcoma soon or later. Collection and analysis of milk sEV proteins hold to promise as a novel monitoring platform for BLV infection of cattle. Though sEV play a vital role in cell to cell communication7 and influence the physiological processes8, their function in infectious diseases is still under progress. Huang et al.21 reported that Staphylococcus aureus modified the protein expression in mammary tissue of cattle indicating that infectious diseases are capable of changing the normal proteomic profile of infected tissues. Recently, we demonstrated that mRNAs profile in milk sEV from BLV-infected cattle were altered compared with those of uninfected cattle15. However, to date, no research has been performed on identifying the changes of proteins in milk sEV from BLV-infected cattle. Therefore, the current study was conducted to evaluate the proteins encapsulation in milk sEV from BLV-infected cattle.
The present study detected a total of 26 proteins which were the candidates for differentially expressed proteins in milk sEV from BLV-infected cattle compared with those of uninfected cattle. Previous studies have been identified different types of proteins from blood or tumour tissues that were feasible in understanding the tumour progression in human. Some of the up/down-regulated proteins in our present study such as CNTFR, DDAH1, TUBB1, MARCH8, KRT1, KRT18, APOE, PDCD6, MMP2, ITGA6, MYOF, and TGFB2 were previously reported to be associated with certain tumorigenesis and progression in malignancies22,23,24,25,26,27,28,29,30,31,32,33. Taken together, these results indicated that as an oncogenic viral disease in cattle, BLV infection could alter the encapsulated sEV proteins; therefore, the new proteins are appearing within milk sEV. The results also suggested that these proteins could be crucial in obtaining information regarding BLV infection and its pathogenesis in cattle.
GO and KEGG analyses identified several functional terms that were enriched by the candidates for uniquely or differentially expressed proteins in milk sEV. Attention was paid to the aforementioned milk sEV proteins to unveil diverse GO terms and numerous KEGG pathways for BLV infection. In the biological process, where proteins were predominantly associated with the cellular process, metabolic process, response to stimulus, and developmental process suggesting cellular proliferation, alteration, and attachment, extensively participated during BLV infection. Binding and catalytic activity were the most prevalent molecular functions, which indicated that direct regulation of protein–protein interactions might be the key process during BLV infection and its pathogenicity in the host. The cellular component indicated that the identified proteins were enriched in the cell and membrane, thus suggesting that the contents of sEV shared close ties with their host cells. The KEGG analysis revealed that the candidates for uniquely or differentially expressed proteins involved in many pathways including pathways in cancers, cytokine-cytokine interaction, PI3K-Akt signalling pathway, Jak-STAT signalling pathway, and focal adhesion. Several of these KEGG pathways played an important role in tumour cell proliferation, metastasis, leukemogenesis as well as formation of solid tumours reported previously in human34. The results suggested that the biologically important pathways are probably involved in development and/or progression of BLV-induced leukemogenesis and tumour formation in cattle. Previous studies reported that mRNAs in milk and blood from BLV-infected cattle involved in diverse biological functions and many other pathways that are consistent with our current study15,35.
In conclusion, proteomic analysis identified a total of 1330 common proteins in milk sEV from BLV-infected cattle, from which 118 proteins were the candidates for uniquely expressed proteins compared with those of uninfected cattle. Besides, GO and KEGG analyses showed a new biological phenomenon and distinct pathways that may have a great contribution into the BLV pathogenesis. The results of this study could be solid ground to facilitate future development of milk sEV-based disease monitoring. This is the first study to present the proteomic analysis of milk sEV from BLV-infected cattle that could have wide applicability in molecular biology.
In this study, since verification of the data by ELISA or other, equivalent means has not been performed, further studies are required.
Materials and methods
All experiments were performed in accordance with relevant guidelines and regulations of the Gifu University Animal Care and Use Committee (approval number 17046 and approved on 4 September, 2017). Additionally, experiments using cows were carried out in compliance with the standards of animal rights, welfare, and with minimum distress by following relevant guidelines and regulations of the Gifu University Animal Care and Use Committee.
Blood samples
Blood samples of 10 ml of each of the 16 Holstein cows were collected from two different farms and directly allocated to vacuum blood collection tubes with or without an anti-coagulant (VP-AS076K, VP-NA050K, and VP-H070K, Terumo, Tokyo, Japan). Total WBCs and lymphocyte counts were measured by an automatic cell counter Celltac α (Nihon Kohden, Tokyo, Japan). The increased lymphocyte count was checked based on the European Community’s leukosis key16. After WBCs and lymphocyte counts, 1.3 ml of each of the anticoagulated blood samples were centrifuged at 2500 × g for 15 min at 25 °C for plasma separation by a centrifuge, MAX-307 (Tomy Seiko, Tokyo, Japan). Plasma samples were collected from the top portion of the tube and used for lactate dehydrogenase (LDH) isozymes measurement later.
Detection of antibodies
Serum was separated from blood by centrifugation at 3000 × g for 15 min at 25 °C by using a centrifuge, MAX-307. Anti-BLV antibodies in serum were measured by using an anti-BLV antibody ELISA kit (JNC, Tokyo, Japan) according to the manufacturer’s instructions.
Detection of BLV
WBC was isolated from blood by hemolysis of red blood cells with 0.83% ammonium chloride followed by washing twice with phosphate buffer saline (PBS). Total amount of DNA was extracted from WBCs by using QIAamp DNA Mini Kit (51304, Qiagen, Hilden, Germany) according to the manufacturer’s instructions. After measurement of DNA concentration of WBC DNA by a spectrophotometer NanoDropLite (Thermo Fisher Scientific, Waltham, MA, USA). Primers to amplify the envelope or pX region of BLV were used for nested PCR according to the protocols by Fechner et al.36 and Murakami et al.37. PCR was carried out in a total reaction volume of 20 µl containing 0.5 U of polymerase from GoTaq Hot Start Green Master Mix (M5122, Promega, Madison, WI, USA) or SapphireAmp Fast PCR Master Mix (RR350A, Takara Bio, Kusatsu, Japan), 0.5 µM of forward and reverse primers, and 1 µl of extracted WBC DNA (100–400 ng). Thermal cycling condition was as follows: 95 °C for 2 min, followed by 35 cycles of 94 °C for 45 s, 62 °C for 30 s, 72 °C for 30 s, and finally 72 °C for 4 min.
Measurement of proviral load
BLV-infected cattle with HPL in blood were selected for this study. It was reported that BLV-infected cattle with HPL in blood were considered as cattle at high risk to be BLV spreaders and might be one of the factors of disease progression38. BLV proviral load was measured by using 100 ng of WBC DNA by a quantitative real-time PCR (qRT-PCR). The amplification was carried out in a reaction mixture containing 10 µl of THUNDERBIRD Probe qPCR Mix (A4250K, Toyobo, Osaka, Japan), 0.3 µl of CoCoMo-BLV Primer/Probe (A803, Riken Genesis, Tokyo, Japan), 5 µl of a template DNA sample, and PCR grade water to increase the volume up to 20 µl. For the proviral quantification, BLV BoLA-DRA gene Plasmid DNA was used from the kit (A804, Riken Genesis) and BLV proviral DNA was measured by a Thermal Cycler Dice Real Time System III (TP970, Takara Bio) according to the manufacturer’s instructions. After the measurement, BLV proviral copies of > 30,000 in 105 WBCs DNA was considered as HPL in BLV-infected cattle (Table 1). Hematological test, detection of serum antibodies against BLV, detection of BLV provirus, and measurement of BLV proviral load were conducted by the Gifu Central Livestock Hygiene Service Center (Gifu, Japan).
Measurement of lactate dehydrogenase (LDH) isozymes
A previous study reported that LDH activities in the serum, mainly, increased LDH 2 and 3 isozyme percentages, reflected progression of EBL, thereby making it a key parameter for the diagnosis of lymphosarcoma39. Therefore, we also focused on serum LDH isozyme activity. BLV-infected cattle with LDH 2 + 3 > 30% was selected. LDH isozymes were measured by a Hydrasys 2 Scan (Sebia, Paris, France) using HYDRAGEL 7 ISO-LDH (Sebia), which was conducted by a clinical laboratory testing company, Fujifilm Vet Systems (Tokyo, Japan).
Collection of milk samples
Milk samples were collected from three BLV-infected and three uninfected healthy cattle. After collection, both milk samples were transported quickly to the laboratory in a cool box to maintain the temperature and were stored at 4 °C for further use.
Isolation of sEV
For the isolation of milk sEV from BLV-infected and uninfected cattle, we followed the procedure previously described by Yamauchi et al.40 and Rahman et al.41 with slight modifications. Importantly, after defatting of milk, milk sEV were purified using acetic acid followed by sequential filtration through 1.0, 0.45, and 0.2 μm filters (GA-100, C045A047A, and C020A047A, Advantec, Tokyo, Japan). Subsequently, milk sEV were concentrated by ultracentrifugation (UC) at 100,000 × g at 4 °C for 1 h using a P42A angle rotor (Hitachi Koki, Tokyo, Japan) in a Himac CP80WX ultracentrifuge (Hitachi Koki). After the first UC, the supernatant was discarded and the pellet was resuspended with PBS up to 10 ml into a 13PET tube (Hitachi Koki) followed by another UC at 100,000 × g at 4 °C for 1 h using a P40ST swing rotor (Hitachi Koki). sEV pellet was collected and resuspended with 100 µl of PBS for further use.
Protein concentration of milk sEV
Protein concentration of recovered sEV were estimated by Lowry’s method42 using a DC protein assay kit (500-0113, 500-0114, 500-0115, and 5,000,007; Bio-Rad Laboratories, Hercules, CA, USA) with a spectrophotometer, GeneQuant100 (GE Healthcare, Chicago, IL, USA).
Isolation of RNA and detection of BLV genomic RNA and BLV proteins in milk sEV
Total RNA was extracted from milk sEV from BLV-infected cattle by using QIAamp viral RNA Mini Kit (52906, Qiagen, Hilden, Germany) according to the manufacturer’s instructions. The RNA concentration was measured by a spectrophotometer NanoDropLite (Thermo Fisher Scientific). cDNA was synthesis by reverse transcription reaction using 5 × PrimeScript RT Master Mix (RR036A-1, Takara Bio) followed by nested PCR was performed to amplify the envelope or pX region of BLV36,37. WB analysis was performed as described in a previous study41. After gel electrophoresis and trans-blotting, membranes were blocked with 5% non-fat skim milk in Tris-buffered saline [0.1 M Tris–HCl (pH 8.0) and 0.03 M NaCl] containing 0.1% Tween-20 (TBST) at room temperature (RT) for 30 min. For the detection of BLV proteins gp51 in milk sEV from BLV-infected cattle, monoclonal antibody specific to a D-D' epitope on BLV gp51 (Env) (1:400, VMRD, Pullman, WA, USA) was used. The membrane was incubated for 1 h at RT, diluted in 1% non-fat skim milk in TBST, followed by washing thrice with TBST. The secondary antibody, anti-mouse IgG sheep antibody (1:1000, NA9310, GE Healthcare, Little Chalfont, UK) conjugated with horseradish peroxidase, were diluted with TBST incubated for 1 h at RT followed by washing thrice with TBST. Peroxidase activity was detected using a Pierce ECL Plus substrate (Thermo Fisher Scientific) and visualised by using a chemiluminesence apparatus (ChemiDoc XRS + , Bio-Rad Laboratories).
Characterization of milk sEV
NTA analysis of milk sEV from BLV-infected and uninfected cattle was performed using a NanoSight LM10V-HS, NTA 3.4 instrument by an assigning company (Quantum Design Japan, Tokyo, Japan). Morphological examination of isolated milk sEV from BLV-infected and uninfected cattle was carried out by TEM as described in a previous study41 with slight modifications. The sEV pellet was diluted 10 times from its original concentration with distilled water, applied to glow-discharged carbon support films on copper grids followed by stained with 2% uranyl acetate. The samples were then visualized under an electron microscope, JEM-2100F (JEOL, Tokyo, Japan) at 200 kV. WB analysis was carried out as described in a previous study41 with slight modifications. The primary antibodies, anti-CD63 (1:400, M-13, SC-31214, Santa Cruz Biotechnology, Santa Cruz, CA, USA) or anti-HSP70 (1:100, N27F3-4, Enzo Life Science, Farmingdale, NY, USA) following the secondary antibodies, anti-goat IgG donkey antibody (1:2000, SC-3851, Santa Cruz Biotechnology) or anti-mouse IgG sheep antibody (GE Healthcare) conjugated with horseradish peroxidase, were used as described above.
Proteomic and Scaffold Data Independent Acquisition (Scaffold DIA) analysis
For the proteomic profiling, all milk sEV samples were analyzed by the nanoLC-MS/MS method (UltiMate 3000 RSLCnano System, Thermo Fisher Scientific) as described previously by Nguyen et al.43 with slight modifications. All procedures of proteomic analysis were performed by an entrusted company, Hakarel (Osaka, Japan). Accession number of each sEV protein was obtained from the UniPort database for Bos taurus (https://www.uniprot.org/ proteomes/UP000009136). The results were further analyzed by the Scaffold DIA software (Proteome Software, Portland, OR, USA) to compare the peptide counts of the identified proteins, considering the false discovery rate > 1%. A moderated t-test with Benjamini–Hochberg test44 were performed to assess the statistical significance of the data. Corrected p-value cut off of 0.05 was applied. From our data sets, common proteins present in all three samples from BLV-infected and those from three uninfected cattle were considered for listing up and the proteins without a gene name or those with a dual gene name were excluded.
Functional and STRING interaction network analysis
GO and KEGG pathway45 analyses were performed using the Database for Annotation, Visualization and Integrated Discovery (DAVID version 6.8) (https://david.ncifcrf.gov/)46. The biological process, molecular function, cellular component, and protein class were determined using Panther software, online version (http://www.pantherdb.org/). An adjusted p-value of less than 0.05 was considered statistically significant. In this study, we used the Search Tool for the Retrieval of Interacting Genes (STRING) database, version 10.5 (https://string-db.org) to construct the interaction network of the candidates for uniquely or differentially expressed proteins in milk EVs from BLV-infected cattle.
Statistical analysis
Statistical analysis was performed using the Student's t-test to compare the milk sEV proteins from BLV-infected and uninfected cattle. The p value 0.05 was considered as statistically significant.
Data availability
Raw data of proteomic analysis of milk sEV from BLV-infected and uninfected cattle was deposited in the Mendeley data repository (Direct URL: Mendeley Data, V1, https://doi.org/10.17632/zxb5vhjrf 5.1 and https://doi.org/10.17632/7c2ddgwcgt.1).
References
van Niel, G. & DAngelo, G. & Raposo, G. Shedding light on the cell biology of extracellular vesicles. Nat. Rev. Mol. Cell Biol. 19, 212–228 (2018).
Raposo, G. & Stoorvogel, W. Extracellular vesicles: exosomes, microvesicles, and friends. J. Cell. Biol. 200, 373–383 (2013).
Théry, C. et al. Minimal information for studies of extracellular vesicles 2018 (MISEV2018): a position statement of the International society for extracellular vesicles and update of the MISEV2014 guidelines. J. Extracell. Vesicles. 7, 1 (2018).
Hata, T. et al. Isolation of bovine milk-derived micro vesicles carrying mRNAs and microRNAs. Biochem. Biophys. Res. Commun. 396, 528–533 (2010).
Sanwlani, R. et al. Milk-derived extracellular vesicles in inter-organism, cross-species communication and drug delivery. Proteomes. 8, 11 (2020).
Kim, S. Y. & Yi, D. Y. Analysis of the human breast milk microbiome and bacterial extracellular vesicles in healthy mothers. Exp. Mol. Med. 52, 1288–1297 (2020).
Izumi, H. et al. Bovine milk exosomes contain microRNA and mRNA and are taken up by human macrophages. J. Dairy Sci. 98, 2920–2933 (2015).
Hata, T. et al. Isolation of bovine milk-derived microvesicles carrying mRNAs and microRNAs. Biochem. Biophys. Res. Commun. 396, 528–533 (2010).
Reinhardt, T. A. et al. Bovine milk exosome proteome. J. Proteom.. 75, 1486–1492 (2012).
Sun, J. et al. MicroRNA expression profiles of bovine milk exosomes in response to Staphylococcus aureus infection. BMC Genom. 16, 806. https://doi.org/10.1186/s12864-015-2044-9 (2015).
Gillet, N. et al. Mechanisms of leukemogenesis induced by bovine leukemia virus: prospects for novel anti-retroviral therapies in human. BMC Retrovirol. 4, 18. https://doi.org/10.1186/1742-4690-4-18 (2007).
Murakami, K. et al. Nationwide survey of bovine leukemia virus infection among dairy and beef breeding cattle in Japan from 2009–2011. J. Vet. Med. Sci. 75, 1123–1126 (2013).
Burny, A. et al. Bovine leukemia: facts and hypotheses derived from the study of an infectious cancer. Vet. Microbiol. 17, 197–218 (1988).
Glass, E. J. et al. (2010) Bovine viral diseases: the role of host genetics, Chap 6. In Breeding for Disease Resistance in Farm Animals 3rd edn (eds Bishop, S. C. et al.) 88–140 (Oxfordshire, CAB International Oxfordshire, 2010).
Ishikawa, H. et al. mRNA profile in milk extracellular vesicles from bovine leukemia virus-infected cattle. Viruses 12, 669. https://doi.org/10.3390/v12060669 (2020).
Mekata, H. et al. New hematological key for bovine leukemia virus-infected Japanese Black cattle. J. Vet. Med. Sci. 80, 316–319 (2018).
Somiya, M., Yoshioka, Y. & Ochiya, T. Biocompatibility of highly purified bovine milk-derived extracellular vesicles. J. Extracell. Vesicles. 7, 1440132. https://doi.org/10.1080/20013078.2018.1440132 (2018).
Samuel, M. et al. Bovine milk-derived exosomes from colostrum are enriched with proteins implicated in immune response and growth. Sci. Rep. 7, 5933. https://doi.org/10.1038/s41598-017-06288-8 (2017).
Yang, M. et al. Comparative proteomic analysis of milk-derived exosomes in human and bovine colostrum and mature milk samples by iTRAQ-coupled LC-MS/MS. Food Res. Int. 92, 17–25 (2017).
Wilson, S. R. et al. Nano-LC in proteomics: recent advances and approaches. Bioanalysis. 7, 1799–1815 (2015).
Huang, J. et al. iTRAQ proteomics and bioinformatics analyses of mammary tissue from cows with clinical mastitis due to natural infection with Staphylococci aureus. BMC Genom.. 15, 839 (2014).
Fan, J. et al. MARCH8 is associated with poor prognosis in non-small cell lung cancers patients. Oncotarget. 8, 108238–108248 (2017).
Iyer, S. V. et al. Understanding the role of keratins 8 and 18 in neoplastic potential of breast cancer derived cell lines. PLoS ONE 8, e53532 (2013).
Nami, B. & Wang, Z. Genetics and expression profile of the tubulin gene superfamily in breast cancer subtypes and its relation to taxane resistance. Cancers. 10, 274. https://doi.org/10.3390/cancers10080274 (2018).
Singh, S. et al. Increased expression of MARCH8, an E3 ubiquitin ligase, is associated with growth of esophageal tumor. Cancer Cell Int. 17, 116. https://doi.org/10.1186/s12935-017-0490-y (2017).
Sorlie, T. et al. Gene expression patterns of breast carcinomas distinguish tumor subclasses with clinical implications. Proc. Natl. Acad. Sci. 98, 10869–10874 (2001).
Ye, J. et al. DDAH1 mediates gastric cancer cell invasion and metastasis via Wnt/b-catenin signaling pathway. Mol. Oncol. 11, 1208–1224 (2017).
Massagué, J. TGFβ in cancer. Cell 134, 215–230 (2008).
Pellikainen, J. M. et al. Expression of matrix metalloproteinase (MMP)-2 and MMP-9 in breast cancer with a special reference to activator protein-2, HER-2, and prognosis. Clin. Cancer Res. 10, 7621–7628 (2004).
Wu, Y. et al. ITGA6 and RPSA synergistically promote pancreatic cancer invasion and metastasis via PI3K and MAPK signaling pathways. Exp. Cell Res. 379, 30–47 (2019).
Yoon, J. H. et al. Programmed cell death 6 (PDCD6) as a prognostic marker for gastric Cancers. Tumor Biol. 33, 485–494 (2012).
Zhao, Z. et al. Apolipoprotein E overexpression is associated with tumor progression and poor survival in colorectal cancer. Front. Genet. https://doi.org/10.3389/fgene.2018.00650 (2018).
Zhu, W. et al. Myoferlin, a multifunctional protein in normal cells, has novel and key roles in various cancers. J. Cell. Mol. Med. 23, 7180–7189 (2019).
Harvey, A. J. Overview of cell signaling pathways in cancer. In Predictive Biomarkers in Oncology (eds Badve, S. & Kumar, G.) 167–182 (Switzerland, Springer, Cham, 2019).
Brym, P. & Kamiński, S. Microarray analysis of differential gene expression profiles in blood cells of naturally BLV-infected and uninfected Holstein-Friesian cows. Mol. Biol. Rep. 44, 109–127 (2017).
Fechner, H. et al. Provirus variants of the bovine leukemia virus and their relation to the serological status of naturally infected cattle. Virology 237, 261–269 (1997).
Murakami, K. et al. Bovine Leukemia virus induces CD5-B cell lymphoma in sheep despite temporarily increasing CD5+B cells in asymptomatic stage. Virology 202, 458–465 (1994).
Takeshima, S. N. et al. BLV-CoCoMo-qPCR-2: improvements to the BLV-CoCoMoqPCR assay for bovine leukemia virus by reducing primer degeneracy and constructing an optimal standard curve. Arch. Virol. 160, 1325–1332 (2015).
Sobiech, P., Kuleta, Z. & Jałyński, M. Serum LDH isoenzyme activity in dairy and beef cows. Acta Sci. Pol. 1, 39–43 (2002).
Yamauchi, M. et al. Efficient method for isolation of exosomes from raw bovine milk. Drug Dev. Ind. Pharm. 45, 359–364 (2019).
Rahman, M. M. et al. Acidification effects on isolation of extracellular vesicles from bovine milk. PLoS ONE 14, e0222613 (2019).
Schacterle, G. R. & Pollack, R. L. A simplified method for the quantitative assay of small amounts of protein in biologic material. Anal. Biochem. 51, 654–655 (1973).
Nguyen, E. V. et al. Identification of novel response and predictive biomarkers to HSP90 inhibitors through proteomic profiling of patient-derived prostate tumor explants. Mol. Cell. Proteom. https://doi.org/10.1074/mcp.RA118.000633 (2018).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B. 57, 289–300 (1995).
Kanehisa, M. et al. KEGG: new perspectives on genomes, pathways, diseases and drugs. Nucleic Acids Res. 45, 353–361 (2017).
Huang, D. W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Acknowledgements
We are grateful to Yanagido Farm, Gifu University, and Yamagoe Farm for providing bovine milk. We are also thankful to the Gifu Prefectural Chuo Livestock Hygiene Service Center for providing information of these cattle.
Funding
This study was partly supported by a regulatory research project for food safety, animal health and plant protection (JPJ008617.17935709) of the Ministry of Agriculture, Forestry, and Fisheries of Japan. This study was also supported in part by the OGAWA Science and Technology Foundation, the Morinaga Foundation for Health and Nutrition, and The Sasakawa Scientific Research Grant from The Japan Science Society.
Author information
Authors and Affiliations
Contributions
M.M.R. collected samples, performed laboratory experiments, interpreted data, and wrote the original manuscript. M.M.R. and S.T. performed statistical and bioinformatic analysis. M.M.R. and Y.O.K. performed transmission electron microscopy. M.M.R. and Y.K. performed additional experiments. M.M.R., Y.B., K.S., A.O., and Y.I. revised, edited, and approved the final version of the manuscript. Y.I. conceptualized, designed, supervised, and acquired fund for the entire experiments.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Rahman, M.M., Takashima, S., Kamatari, Y.O. et al. Proteomic profiling of milk small extracellular vesicles from bovine leukemia virus-infected cattle. Sci Rep 11, 2951 (2021). https://doi.org/10.1038/s41598-021-82598-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-82598-2
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.