Abstract
After 25 years of genetically modified cotton cultivation in Mexico, gene flow between transgenic individuals and their wild relatives represents an opportunity for analysing the impacts of the presence of novel genes in ecological and evolutionary processes in natural conditions. We show comprehensive empirical evidence on the physiological, metabolic, and ecological effects of transgene introgression in wild cotton, Gossypium hirsutum. We report that the expression of both the cry and cp4-epsps genes in wild cotton under natural conditions altered extrafloral nectar inducibility and thus, its association with different ant species: the dominance of the defensive species Camponotus planatus in Bt plants, the presence of cp4-epsps without defence role of Monomorium ebeninum ants, and of the invasive species Paratrechina longicornis in wild plants without transgenes. Moreover, we found an increase in herbivore damage to cp4-epsps plants. Our results reveal the influence of transgene expression on native ecological interactions. These findings can be useful in the design of risk assessment methodologies for genetically modified organisms and the in situ conservation of G. hirsutum metapopulations.
Similar content being viewed by others
Introduction
The introgression of transgenes from economically important GM crops to their wild relatives1 (e.g. cotton2,3, sunflower4, rice5, and native papaya breeds6) has been frequently reported in the last decade within the natural distribution of the latter. However, the consequences of transgene expression in natural ecosystems remain unexplored, leading to uncertainty about its impact on local evolutionary and ecological processes. Research on this topic has been challenging largely because, while agro-biotechnological innovations move rapidly, the information required for adequate risk assessment is still limited7. However, knowledge on transgene introgression effects in natural conditions and in particularly in the species centre of origin can offer valuable information for in situ conservation of crop wild relatives (CWR).
The different disciplines interested in assessing the expected functions of genetically engineered genes have produced a fair amount of research in controlled environments, with the only purpose of applying the generated information to improving technological packages8,9,10. However, there is great uncertainty and limited evidence regarding transgene behaviour in wild genotypes that survive in natural conditions under a set of complex ecological challenges7. Consequently, studying transgenes in the wild represents an excellent opportunity to integrate a new stage of scientific knowledge, combining genetic, ecological, evolutionary, and environmental information.
Recently, several studies have looked for differences in genetic expression, fitness, and diversity in non-target organisms, between transgenic organisms and their isogenic lines, as well as between experimental parameters and settings (e.g. laboratory, crops fields, and greenhouse conditions)11,12,13,14. From these studies, we can infer that differences found in seed production and attraction of herbivores’ natural enemies are linked to the interaction of transgenes (i.e. cry and cp4-epsps). Such differences have been associated with defence mechanisms and/or main metabolic pathways involved in plant growth, development, and reproduction, such as the shikimate or the octadecanoid pathways15,16. As a consequence, the quantity and quality of metabolic products are modified, compromising not only plant performance but also their interactions with insects, bacteria, and soil fungi17.
To assess the consequences of transgenic expression in the absence of target pests and glyphosate as selection agents, we studied wild upland cotton (Gossypium hirsutum L.) and its associated ant communities in Mexico where the wild to domesticated complex exists. Upland cotton is a Mesoamerican native species in which introgression of transgenes conferring resistance to specific lepidopteran larvae (Helicoverpa armigera, H. punctigera, Heliothis virescens, Pectinophora gossypiella) (Bt or cry genes) and tolerance to glyphosate (cp4-epsps) have been detected, at its center of origin, in seven out of eight metapopulations (Baja California Sur, BCSM; North Pacific, NPM; Central Pacific, CPM; South Pacific, SPM; Gulf North, GNM; Gulf South, GSM; and Yucatán Peninsula, YPM)2,3. We examined the potential effects of introgressed transgenes on G. hirsutum production of extrafloral nectar, which constitutes a reward for defensive ants who patrol the plant to repel herbivores18,19. Assessing the ant-cotton interaction is relevant because ants are not-target insects of transgenic technology and their role is protecting cotton plants against herbivores20. Furthermore, in the coastal sand dunes where the studied populations are established, ants play important ecological functions acting as soil engineers and seed dispersers, influencing plant germination, reproductive success, and distribution patterns. To this end, we analyzed three wild cotton genotypes, two of which are introgressed plants with transgenes. All of them are growing in the species centre of origin: (1) wild genotypes without transgenes; (2) wild genotypes with cry; and (3) wild genotypes with cp4-epsps (hereafter W, Wcry, and Wcp4-epsps, respectively). We evaluated the inducibility of extrafloral nectar (EFN) after exogenous application of methyl jasmonate (MeJA), which has been widely used as a tool when studying plant reactions to stress and understanding the interactions between the elicitation of plant responses and their consequences for herbivores and beneficial insects21. We also measure the response of the ant community and herbivore damage associated with the three cotton genotypes.
Material and methods
Study site
This study was conducted in the costal dunes of the Ría Lagartos Biosphere Reserve (hereafter RL), Yucatan, Mexico. We located it in the stub zone, where wild cotton, G. hirsutum, is distributed22 (Table S1). We chose this site for the purpose of keeping track of the introgressive hybridisation between GM domesticated and wild plants3 within the RL, a place that allows long-term research without direct evidence of the anthropisation effect (i.e. land-use change).
Experimental plants
To determine the presence of the herbicide-tolerant transgenes and those with insecticide effects within the studied population, in March 2018 we collected the foliar tissue of 61 cotton plants (six leaves per plant) of comparable age, georeferencing each plant to recognize them for further experiments into a natural patch of cotton plants of 2 ha into a the RL to 6.1 km from the nearest Town, the most distance between plants was 280 m. Genomic DNA was isolated from foliar tissue using the Miniprep CTAB method2. DNA concentrations were analyzed with a Quibit 3.0 Fluorometer (ThermoFisher Scientific, Massachusetts, USA). DNA sequences were amplified through a PCR endpoint, DNA polymerase GoTaq Flexi (Promega, USA), and the primers: Cry1Ab/Ac, Cry2Ab, and CP4EPSPS from Eurofins Scientific (Brussels, Belgium). The thermocycler was programmed for a denaturation period of 8 min at 95 °C, followed by 30 cycles of 20 s at 95 °C, 1 min at 60 °C, 1 min 72 °C, and then 8 min at 72 °C for the final cycle. Reactions were carried out in a volume of 25 µL containing 5 µL × buffer, 25 mM of magnesium chloride, 100 mM of dTTP, 10 µM primer, 70 nM of genomic DNA, and one unit of Taq polymerase (GoTaq Flexi by Promega). PCR products were separated in 2% denaturing agarose gel. We recorded the presence of three transgenes, cry1ab/ac, cry2ab, and cp4epsps, in individual and stacked form (Table 1). Based on the above, we combined in a single category all the plants that expressed the cry gene because they have the same insecticide effect. The initial sample size for our experiments of 61 plants was reduced to 21 plants, because the maximum amount of the Wcry category was seven (Table 1). These plants were divided into three groups, with 7 individuals for each genotype (wild, introgressed with cry, and introgressed with cp4-epsps): (1) wild without transgenes, W (control); (2) wild with cry, Wcry; and (3) wild with cp4-epsps, Wcp4-epsps. For the purpose of this investigation we decided to exclude the plants expressing both transgenes, to avoid non additive effects. As described below, even with this limited amount of plants we were able to detect interesting results due to the genetic expression of these genes in a natural population, where plants coexist with their antagonists and mutualists. These results, of course, must be taken as a first line of evidence that requires further investigation with larger sample sizes.
Experiment 1: effect of MeJA induction on extrafloral nectar production
Each experimental plant of the three genotypes was divided into two zones, in which we applied the control and induction treatments in the most distal branches23. Previously, we had evaluated whether induction through methyl jasmonate (MeJA) was active in cotton plants and found that indeed, cotton plants can respond to this elicitor by inducing extrafloral nectar in the branch where it is applied, hence the response was local and not systemic21. Considering this, control and induction treatments were applied in the same individual on different branches, using ten leaves that were sprayed with 2 mL of distilled water or MeJA solution (450 mM), respectively. To exclude insects from extrafloral nectaries, we coated plant branches with Tanglefoot in its original concentration (25% of natural gum resins) (Tangletrap, The Tanglefoot Corporation)24. Induction was carried out at 6:00 a.m. during three days of the summer of 2018 and concentration measurements of soluble solids were taken six hours later, at noon, based on previous observations on the peak of extrafloral nectar production. Nectar secretions from 10 leaves in each treatment(one nectary per leaf) per plant, in total 84 branches and 420 leaves were quantified; the amount of soluble solids was measured with 3 µL graduated micropipettes, and nectar concentration was obtained with a temperature-compensated portable refractometer (ATAGO hand refractometer, L Kübler, Karlsruhe, Germany). To remove the nectar, we applied 3 µL of distilled water to each nectary and repeated the procedure until we reached concentrations lower than 1%. Then, we added values from all collections in one leaf, to quantify leaf overall production of solid EFN compounds23,25.
Experiment 2: effect of MeJA induction on abundance, richness, and composition of ants
After nectar measurements concluded, MeJA induction was carried out for two more days following the same experimental design as described above but allowing ant foraging. We conducted two censuses per day during a two-day period, at 10:00 a.m. and 4:00 p.m., when ant activity was at its highest (according to previous observations by our lab group). We registered the number of ants feeding and patrolling extrafloral nectaries and collected a sample of each ant species for taxonomic identification.
Ants were identified to species level using specialized publications or by comparison with specimens in the ant collection of the Laboratorio de Invertebrados del Suelo (INECOL, Xalapa, Mexico). Voucher specimens for all collected species were deposited in the latter collection (Table S2).
Herbivore damage
We had to estimate the percentage of herbivore damage on the leaves, that is to say, the part of the leaves eaten by herbivores. Pictures of fresh leaves were captured (n = 10) for each branch of the genotypes described above. We used a Canon EOS Rebel T6i 18 MP camera without flash, positioned at 15 cm from the leaves placed above a scaled portable background. Pictures were taken in the field, preventing the leaves from being removed from the branches. Images were saved as JPG format and automatic quantification of leaf damage was performed with BioLeaf Analyze Foliar (version 1.0)26.
Data analysis
Given that our data did not fit normal assumptions, our simples were analyzed using Generalized Linear Models with Quasipoisson distribution (GLM Ime4 package in R version 3.5.0), based on the dispersion of the data.The Poisson distribution and zero inflated models were also considered27. Consequently, we individually evaluated the effect of the induction treatment on EFN for each genotype using the Generalized Lineal Model with Quasipoisson distribution with the package GLM Ime4 in R version 3.5.0. Likewise, ant abundance and the percentage of herbivore damage were evaluated with Generalized Linear Models with Quasipoisson distribution (GLM Ime4 package in R version 3.5.0, where theinduction treatment (MeJA or control) and the genotype of the wild plants (Wcry, Wcp4-esps, and W) were considered as a fixed factor. We compared the effect of the induction treatment, genotype, date, and their interactions. Finally, ant richness and composition were expressed as the total of morphospecies per genotype. The scripts utilized for the analyses are available online at https://github.com/conservationgenetics/Ongoing-ecological-and-evolutionary-consequences-by-the-presence-of-transgenes-in-a-wild-cotton-popu.
Results
Effect of MeJA induction treatment on extrafloral nectar production in wild cotton with and without transgenes
Exogenous application of MeJA significantly increased EFN secretion in wild plants without transgenes (W) (P(χ2) = 0.048, df = 1). In contrast, transgenic genotypes (Wcry and Wcp4-epsps) did not respond to this stimulus (Wcry: P(χ2) = 0.375, df = 1; Wcp4-epsps: P(χ2) = 0.388, df = 1), because of different reasons regarding the genotypes with introgressed genes. Whereas Wcry control plants had a high EFN production equivalent to the induced state of W plants, EFN production in Wcp4-epsps plants was inhibited, yielding less than 8 µg/mL in both treatments (Fig. 1). When comparing the control response between genotypes, the extrafloral production in the Wcry genotype was significantly different to the one found in W and Wcp4-epsps (P(χ2) < 0.001, df = 2) (Fig. 1). And, when comparing the induction response between induced and control treatment of the W and Wcry genotypes, respectively, we observed no differences (P(χ2) = 0.637, df = 1).
Effect of the induction treatment with MeJA on secretion of extrafloral nectar by wild Gossypium hirsutum plants without transgenes (W), wild with cry (Wcry) and wild with cp4-epsps (Wcp4-epsps) and within each plant. Grey bars represent the control treatment and yellow bars are the MeJA induction treatment. Values are means ± SE (N = 21, n = 84). The asterisk (*) indicates significant differences according to a post-hoc test.
Abundance, richness and composition of ants associated with cotton’s EFN
We collected 109 ants associated with the EFN of G. hirsutum, which belonged to eight species. The highest abundance and richness was registered in the W genotype (42 ants and 7 species), meanwhile in Wcry and Wcp4-epsps we registered 33 and 34 ants, belonging to four and six species, respectively (Table 2). Four species of ants displayed defensive behaviour: Camponotus planatus, Camponotus rectangularis aulicus, Dorymyrmex bicolor, and Pseudomyrmex gracilis. Likewise, we registered two ant species without defensive behaviour associated with the wild cotton' EFN, and one of these is invasive (Paratrechina longicornis).
The ant' community composition associated with cotton' EFN was different between genotypes. W plants had two exclusive species, Wcp4-epsps had an exclusive ant species, while Wcry didn't have exclusives ants, and four of the ants collected were presented in all genotypes (Table 2).
Effect of MeJA induction treatment on the abundance of ants in wild cotton with and without transgenes
Exogenous application of MeJA significantly affected the abundance of ants in the Wcry genotype (P(χ2) < 0.001, df = 1), with C. planatus being the most abundant ant species. In contrast, in the W and Wcp4-epsps genotypes the induction treatment did not have any effect on ant abundance (W: P(χ2) = 0.436, df = 1; Wcp4-epsps: (P(χ2) = 0.210, df = 1) (Fig. 2).
Effect of induction treatment on ant abundance in three genotypes of wild Gossypium hirsutum L.: wild cotton without transgenes (W), wild with cry (Wcry), and wild with cp4-epsps (Wcp4-epsps). Grey bars are for the untreated control and yellow bars for MeJA (induction treatment). Bars show means ± SE (N = 21 n = 42). n.s. indicating non-significant differences, and *** indicates significant differences (P < 0.001) according to the Tukey post-hoc test between constitutive and induced conditions.
Herbivore damage in wild cotton with and without transgenes
Wcp4-epsps was the genotype with the highest herbivore damage value that was significantly different (P(χ2) < 0.001, df = 2) when compared to W and Wcry genotypes. The lowest percentage of herbivore damage value was registered for Wcry plants (Fig. 3).
Discussion and conclusions
In this study, we showed that the expression of cry and cp4-epsps genes in wild cotton altered the secretion of EFN, the associations with different ant species, and the levels of herbivore damage on target plants. Wcry constantly maintained a high production of EFN, regardless of the MeJA treatment, but nectar production was minimal in Wcp4-epsps. These changes in nectar inducibility seem to modify the composition of ant communities, foster the dominance of the generalist and defensive species C. planatus in Bt plants and the presence of ants without defensive role, M. ebeninum, in the herbicide tolerant genotype, while W plants had both defending species (C. planatus, C. rectangularis aulicus and P. gracilis) and invasive ant species (P. longicornis) in the same proportion. Furthermore, herbivore damage and its associated ant community were different according to the introgressed transgene.
Wild and introgressed cotton do not display phenotypic equivalence in natural conditions
In general, it has been assumed that introgressed and wild genotypes should display similar phenotypes in the absence of the selection agents targeted by transgenes. However, when we compared the control group and the three genotypes, we registered different nectar secretion patterns among them (Fig. 1). Similar results have been registered in populations of bt rice and glyphosate-tolerant sunflowers living in natural conditions where introgressed plants are different from their wild relatives5.
Transgene expression modified indirect induced defences in wild cotton
Most plants are able to induce responses after herbivore damage and/or phytohormone exogenous application (i.e. jasmonic acid, JA; methyl jasmonate, MeJA; and salicylic acid, SA)11,28,29. However, unlike wild plants without transgenes, individuals with transgenes were not sensitive to the induction treatment with MeJA for increasing their EFN production (Fig. 1). These results contrast with previous reports on cultivated varieties, such as Bt and glyphosate-resistant (cp4-epsps), in which direct defences such as gossypol terpenoids (160%), hemigossypolone (160%), helicoids 1|4 (213%) and indirect defenses, such as volatile compounds (VOCs) (171.2%) and extrafloral nectar (EFN) (133%), were reported to increase in plants sprinkled with JA and MeJA21,28,29,30.
The inability of plants with transgenes to have the production of extrafloral nectar induced in them was related to different processes dependent on the identity of the transgenes in question. Whereas Wcry control plants had a high EFN production equivalent to the induced state of W plants, EFN production in Wcp4-epsps plants was inhibited. Contrasting these findings with results obtained under controlled conditions (i.e. greenhouse and crop conditions)3,21, we suggest that EFN production is linked to genotypes with transgenes and abiotic stress in the coastal dunes, because transgenes are connected to main metabolic pathways that respond to stressful conditions21.
Wild cotton with cp4-epsps
In the absence of herbicides acting as a selection agent, wild plants with cp4-epsps exhibited large differences compared to wild plants without them. Their low nectar production (> 8 µg/mL) (Fig. 1) could be linked to the crosstalk between the jasmonate and the salicylate (SA) pathways (Fig. 4, orange and purple section). In G. hirsutum and other species, SA signalling has been proven to negatively affect JA signalling (e.g. Zea mays, Solanum lycopersicum, Nicotiana tabacum and Arabidopsis thaliana)31,32,33: therefore, we suggest an interference between the SA and JA pathways given previous reports that an over-expression of the cp4-epsps gene modifies the second part of the shikimate pathway (post-chorismate), which leads to the synthesis of essential amino acids as phenylalanine, tryptophan, or tyrosine, the latter being a precursor of benzoic acid BE, and SA34,35 (Fig. 4, purple section). This evidence highlights that hidden crosstalk effects among different metabolic pathways can scale up and modify plant phenotypes (e.g. extrafloral nectar production).
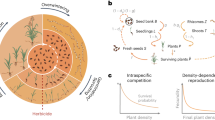
A diagram illustrating how the expression of cry (A) and cp4-epsps (B) in absence of their selection agent (pests and glyphosate) can affect the extrafloral nectar production. The extrafloral nectar (EFN) production is an induced defence that can be triggered by foliar herbivory, mechanical damage, and exogenous application of phytohormones (i.e. jasmonic acid, methyl jasmonate, and salicylic acid). These factors activate the octadecanoid pathway, and therefore, the production of extrafloral nectar, (A) aqua rectangle. The (C) section is an example of this reaction in a wild cotton plant (without transgenes). After damage, the key genes (yellow mesh) of the octadecanoid pathway are activated and produce extrafloral nectar. Another scenario is when the wild cotton expresses cry genes (A section), in this case, the key genes of the octadecanoid pathway interact synergistically with the cry transgene (green mesh). This triggers an over-expression of the production of EFN (aqua thick arrow), switching from inducible to constitutive responses. When the plants express cp4-epsps (B section), the production of extrafloral nectar is reduced or inhibited. A possible answer is an over-expression of the epsps gene (gold curve arrow), that increased production of salicylic acid which creates a crosstalk between shikimate and octadecanoid pathways (black cross-talk arrow). When the shikimate pathway is activated, the principal inducible defence is the production of volatile organic compounds (VOCs) (pink rectangle).
Wild cotton with cry
Wild cotton plants with cry genes continuously produced EFN as a constitutive defence (Fig. 1), in equivalent quantities as the induced state of W plants. EFN production is regulated by the octadecanoid signalling pathway, which can be activated by herbivore damage, mechanical damage, and phytohormones, such as JA and MeJA21,28 (Fig. 4, green section). However, for cotton, a specific elicitor is not necessary36. Four key genes for the synthesis of JA and MeJA have been described: AOS, AOC, HPL, and COI137. In Bt maize, studies comparing GM corn and its isogenic lines report an increase of 24% in phenols and 63% of DIMBOA (2,4-dihidroxi-7-metoxi-1,4-benzoxazin-3-ona; natural defences against lepidopteran herbivores)11. This is consistent with observations of a synergy between maize direct defences and Bt genes, after exogenous applications of JA (Fig. 4, orange section). Considering the latter, we suggest that Wcry cotton may present a similar response, as the genes activating the JA pathway are GhAOS and GhCOI1 (homologs to maize JA biosynthesis genes: ZmAOS and ZmCOI1), in addition to Ghppo1, which confers natural resistance to lepidopteran pest, such as H. armigera38. The interaction of cry with other genes could modify the production of EFN in Wcry plants.
Effect of the transgenes’ expression on ants associated to wild cotton
We identified eight species of ants harvesting EFN (Table 2), but with distinctive communities as a function of the plant genotype. This result suggests that the change in quantity, and possibly the composition and quality of EFN, can influence the ant community associated with G. hirsutum39,40,41.
Changes in plant reward production could potentially compromise the attraction of natural enemies of herbivores42. In our study, the availability of EFN was modified. Although species richness was the same as in W plants (Table 2), the most abundant ant species associated with Wcp4-epsps plants, M. ebeninum, is considered a generalist species. Moreover, due to the lack of aggressive behaviour, this species does not represent an effective biotic defence43. The high abundance of this non-defensive species could be associated with the greater herbivore damage observed in Wcp4-epsps plants (Fig. 2). In contrast, W or Wcry plants showed a greater abundance of more aggressive ant species such as C. planatus, C. rectangulatus, and P. brunneus and significantly less herbivore damage.
In Wcry cotton, the community of patrolling ants was mainly dominated by C. planatus, in both treatments (control and induction). Interestingly, although the amount of nectar did not vary between treatments, the abundance of ants was significantly different. The dominance of a single ant species could have benefited the plants with increased indirect defence, reducing herbivore damage and promoting a greater seed production per plant, as described in Turnera ulmifolia44, Schomburgkia tibicinis45, and Opuntia stricta42. However, considering the aggressive and dominant behaviour of C. planatus, there may be ecological costs through antagonistic relationships with pollinators. Ants can interrupt pollination and affect plant fitness25,46,47. The outcome of these mutualistic and antagonistic interactions requires further study.
Effects of transgenes on herbivore damage
Considering that the type of mutualism that cotton sustains with ants is defensive, we suggest that the change we observed in the composition of ants is likely to have influenced herbivore damage in the different genotypes, which in turn has the potential to reduce fitness as shown by other studies of cotton48,49,50. However, a study carried out on wild upland cotton reported that plants tolerate intermediate levels of leaf damage inflicted by leaf-chewing insects (< 50%)51. Thus, the increased herbivore damage observed in Wcp4-epsps plants (< 10%; 4.402 ± 0.863) does not necessarily jeopardize plant growth or reproductive success. For example, in the absence of glyphosate, transgenic hybrids of rice and soybean show changes in their phenology (i.e. early flowering and shorter germination times) and an increase in their fitness (i.e. larger fruit and seed sets) compared to their wild relatives4,5. It has been suggested that the over-expression of the epsps gene is responsible for these compensation effects on plant fitness, in view of the biotic and abiotic stressors that activate the shikimate pathway5,15. Further investigation to find out if this is the case for Wcp4-epsps cotton is currently being developed under natural conditions.
In contrast, Wcry plants exhibited the lowest herbivore damage, although not significantly different from wild plants without transgenes (Fig. 3). The expression of cry genes, conferring a new defensive trait against lepidopterans, may represent an advantage for plants under selection when target herbivores are present. For example, within the natural distribution of wild sunflowers, genotypes with cry genes showed less herbivore damage than their wild relatives (WR), increasing their seed production by 55%4. Nonetheless, in G. hirsutum we did not find differences in herbivore damage when compared to the W and Wcry genotypes, probably because lepidopteran species targeted by Bt cotton were not present in our study site, hence, cry expression might not have a role in increasing fitness. However, indirect effects could be present through its interaction with native herbivores. For example, experiments under laboratory and experimental conditions conducted on maize with cry genes, have shown non-lethal effects in the physical condition of non-target caterpillars (e.g. smaller size, lower weight, lower survival, and more larval instars)52. Likewise, if the nutritious quality of herbivore tissues changes after consuming Bt toxins, these effects could cascade to higher trophic levels53,54, increasing mortality and decreasing longevity or development of predators. This has been shown for chewing predators (e.g. Chrysoperla carnea and lady beetles, Coccinellidae) because they ingest the gut of the prey, where most of the toxins are concentrated54. Parasitoids have also been found to be affected after the consumption of Bt toxins contained in their preys55. When the wasp Microplitis mediator (parasitoid of H. armigera) was fed with larvae containing cry toxins, they extended its egg and larval development time by 1–2 days, significantly decreasing its pupal weight by 35%, and its overall longevity when the toxin concentrations were high (4–8 μg g−1). Hence, a question that warrants further investigation is how Wcry cotton genotypes in the wild affect herbivore and predator communities and their interactions.
By means of an integrative methodology, we evaluated the effect of the cry and cp4-epsps gene expression in wild cotton plants. As a result, we obtained the following noteworthy results: (1) differential response in the induced defence mechanism (extrafloral nectar production) concordat to plant genotype; and (2) modification of biotic interactions between introgressed cotton and relevant organisms, under natural conditions.
Although several hypotheses have been raised regarding the consequences of GMO release into the environment (e.g. gene flow, hybridization, and introgression), these have only been tested under controlled conditions. We found that some theory-based concerns can be confirmed when performing functional experimental designs in the wild. First, it is possible to investigate ecological and evolutionary impacts of new genes under natural conditions by studying community processes, such as changes in tri-trophic interactions. Second, we detected physiological and possible metabolic alterations generated by the expression of transgenes in wild cotton plants without the pressure of selection agents (pests and herbicides) targeted by those genes. Third, until 2008, 4 wild metapopulations of upland cotton showed evidence of introgression with GM cotton2, and recently, introgression in the Baja California Sur, BCSM, Central Pacific, CPM, and Yucatan Peninsula YPM metapopulations has been reported3. In this study, we reaffirm the presence of transgenes in the YPM, but it's important to consider that the establishment of these has been fast. Whereas during the first monitoring in 2008, YPM metapopulation didn't register the presence of transgenes, in 2018, 60.64% of 61 plants had them (Table 1).
In natural ecosystems, changes at different scales (i.e. genetic, individual, and community), following the introduction of novel genes lead to endless research possibilities. Through them we can integrate broad information regarding biological control, agro-biotechnologies, and conservation biology, with promising further applications. Although we do not know the routes of transgene dispersion, we provide evidence of some of the mechanisms that could favour the establishment and persistence of these new genes, mainly due to their interaction with key defence metabolic pathways. However, transgene frequency in wild populations and the associated ecological consequences must continue to be monitored and evaluated so as to contribute information that allows us to make decisions for the conservation of the primary genetic pool and the ecological and evolutionary processes that have shaped its diversity.
Alterations in the defence mechanisms of wild relatives in one of the most important crops for humanity represents critical evidence on the threats of introgressed genes to biological and cultural heritage for the following generations. Such negative consequences would be enough to envision GMO liberation into the environment, short and long distance from its origin centre, from a different perspective. At this point, we are in a watershed moment to: (1) develop the necessary research to mitigate the ecological, evolutionary caused by the introgression of GMOs into the wild-to-domesticated complex of the species used, as well as reformulating risk assessments for protection goals; (2) fill gaps in our knowledge of the complex dispersion routes of transgenes from crop to wild relatives and native varieties; and finally, (3) prioritize the in-situ conservation of the primary gene pool without transgenes. All these efforts would contribute to the integration of the available disperse information so we may understand the consequences of transgenic plants coexisting with their relatives. As we demonstrated in the current study, the presence of these genes can cause intrinsic changes in wild populations of cotton (allelic frequency), and changes in their ecological interactions. If we want to conserve in-situ the primary gene pool of wild relatives, we must work to identify the ecological and evolutionary processes affected by the existence and permanence of these transgenes within their populations. Upon the detection of these genes, mitigation strategies to reduce the magnitude of the damage can be promptly designed.
References
Ellstrand, N. C., Prentice, H. C. & Hancock, J. F. Gene flow and introgression from domesticated plants into their wild relatives. Annu. Rev. Ecol. Syst. 30, 539–563 (1999).
Wegier, A. et al. Recent long-distance transgene flow into wild populations conforms to historical patterns of gene flow in cotton (Gossypiumhirsutum) at its centre of origin. Mol. Ecol. 20, 4182–4194 (2011).
Hernández-Terán, A. et al. In vitro performance in cotton plants with different genetic backgrounds: The case of Gossypiumhirsutum in Mexico, and its implications for germplasm conservation. PeerJ 7, 1–18 (2019).
Snow, A. A. et al. A Bt transgene reduces herbivory and enhances fecundity in wild sunflowers. Ecol. Appl. 13, 279–286 (2003).
Yang, X. et al. Genetically engineered rice endogenous 5-enolpyruvoylshikimate-3-phosphate synthase (epsps) transgene alters phenology and fitness of crop-wild hybrid offspring. Sci. Rep. 7, 1–12 (2017).
Manshardt, R., Bishaw, D., Pitz, K. & Stewart, C. N. Gene flow from commercial transgenic papaya fields into feral populations in Hawaii. Acta Hortic. 1124, 33–40 (2016).
Ellstrand, N. C. “Born to run”? Not necessarily: Species and trait bias in persistent free-living transgenic plants. Front. Bioeng. Biotechnol. 6, 1–10 (2018).
Elmore, R. W. et al. Glyphosate-resistant soybean cultivar yields compared with sister lines. Agron. J. 93, 408 (2001).
Zhao, J. Z. et al. Transgenic plants expressing two Bacillusthuringiensis toxins delay insect resistance evolution. Nat. Biotechnol. 21, 1493–1497 (2003).
Halpin, C. Gene stacking in transgenic plants—The challenge for 21st century plant biotechnology. Plant Biotechnol. J. 3, 141–155 (2005).
Feng, Y. J., Wang, J. W. & Luo, S. M. Effects of exogenous jasmonic acid on concentrations of direct-defense chemicals and expression of related genes in Bt (Bacillusthuringiensis) corn (Zeamays). Agric. Sci. China 6, 1456–1462 (2007).
Torres, J. B., Ruberson, J. R. & Whitehouse, M. Transgenic Cotton for Sustainable Pest Management: A Review 15–53 (Springer, Dordrecht, 2009). https://doi.org/10.1007/978-1-4020-9654-9_4.
Rauschen, S., Schultheis, E., Pagel-Wieder, S., Schuphan, I. & Eber, S. Impact of Bt-corn MON88017 in comparison to three conventional lines on Trigonotyluscaelestialium (Kirkaldy) (Heteroptera: Miridae) field densities. Transgenic Res. 18, 203–214 (2009).
Chen, Y. H., Shapiro, L. R., Benrey, B. & Cibrián-Jaramillo, A. Back to the origin: In situ studies are needed to understand selection during crop diversification. Front. Ecol. Evol. 5, 1–8 (2017).
Strapasson, P., Pinto-Zevallos, D. M. & Zarbin, P. H. G. Soybean (Glycine max) plants genetically modified to express resistance to glyphosate: Can they modify airborne signals in tritrophic interactions?. Chemoecology 26, 7–14 (2016).
Erb, M. Volatiles as inducers and suppressors of plant defense and immunity—Origins, specificity, perception and signaling. Curr. Opin. Plant Biol. 44, 117–121 (2018).
Kolseth, A. K. et al. Influence of genetically modified organisms on agro-ecosystem processes. Agric. Ecosyst. Environ. 214, 96–106 (2015).
Ckers, F. L. W. & Bonifay, C. How to be sweet? Extrafloral nectar allocation by Gossypiumhirsutum fits optimal defense theory predictions. Spec. Featur. Ecol. 85, 1512–1518 (2004).
Rudgers, J. A., Strauss, S. Y. & Wendel, J. F. Trade-offs among anti-herbivore resistance traits: insights from Gossypieae (Malvaceae). Am. J. Bot. 91, 871–880 (2004).
Rudgers, J. A., Hodgen, J. G. & White, J. W. Behavioral mechanisms underlie an ant–plant mutualism. Oecologia 135, 51–59 (2003).
Williams, L., Rodriguez-Saona, C. & del Conte, S. C. C. Methyl jasmonate induction of cotton: A field test of the ‘attract and reward’ strategy of conservation biological control. AoB Plants 9, 1–15 (2017).
Ramos-Zapata, J., Parra-Tabla, V., Leirana-Alcocer, J., González-Moreno, A. & Chiappa-Carrara, X. Ecología Funcional de la Reserva de la Biósfera de Ría Lagartos (2017).
Heil, M. Induction of two indirect defences benefits Lima bean (Phaseoluslunatus, Fabaceae) in nature. J. Ecol. 92, 527–536 (2004).
Mooney, K. A. & Agrawal, A. A. Plant genotype shapes ant–aphid interactions: Implications for community structure and indirect plant defense. Am. Nat. 171, E195–E205 (2008).
Hernandez-Cumplido, J., Forter, B., Moreira, X., Heil, M. & Benrey, B. Induced floral and extrafloral nectar production affect ant–pollinator interactions and plant fitness. Biotropica 48, 342–348 (2016).
Machado, B. B. et al. BioLeaf: A professional mobile application to measure foliar damage caused by insect herbivory. Comput. Electron. Agric. 129, 44–55 (2016).
Lindén, A. & Mäntyniemi, S. Using the negative binomial distribution to model overdispersion in ecological count data. Ecology 92, 1414–1421 (2011).
Wang, L. & Wu, J. The essential role of jasmonic acid in plant-herbivore interactions—Using the wild tobacco Nicotianaattenuata as a model. J. Genet. Genomics 40, 597–606 (2013).
Park, S. H., Scheffler, J., Scheffler, B., Cantrell, C. L. & Pauli, C. S. Chemical defense responses of upland cotton, Gossypiumhirsutum L. to physical wounding. Plant Direct 3, 1–15 (2019).
Hagenbucher, S., Eisenring, M., Meissle, M. & Romeis, J. Interaction of transgenic and natural insect resistance mechanisms against Spodopteralittoralis in cotton. Pest Manag. Sci. 73, 1670–1678 (2017).
Zhang, P. et al. Suppression of jasmonic acid-dependent defense in cotton plant by the mealybug Phenacoccussolenopsis. PLoS ONE 6, e22378 (2011).
Kunkel, B. N. & Brooks, D. M. Cross talk between signaling pathways in pathogen defense. Curr. Opin. Plant Biol. 5, 325–331 (2002).
Thaler, J. S., Humphrey, P. T. & Whiteman, N. K. Evolution of jasmonate and salicylate signal crosstalk. Trends Plant Sci. 17, 260–270 (2012).
Verberne, M. C., Verpoorte, R., Bol, J. F., Mercado-Blanco, J. & Linthorst, H. J. M. Overproduction of salicylic acid in plants by bacterial transgenes enhances pathogen resistance. Nat. Biotechnol. 18, 779–783 (2000).
Dempsey, D. A., Vlot, A. C., Wildermuth, M. C. & Klessig, D. F. Salicylic acid biosynthesis and metabolism. Arab. B. 9, e0156 (2011).
Wäckers, F. L. & Wunderlin, R. Induction of cotton extrafloral nectar production in response to herbivory does not require a herbivore-specific elicitor. in Proceedings of the 10th International Symposium on Insect-Plant Relationships 149–154 (Springer Netherlands, 1999). https://doi.org/10.1007/978-94-017-1890-5_18.
Yang, F. et al. Analysis of key genes of jasmonic acid mediated signal pathway for defense against insect damages by comparative transcriptome sequencing. Sci. Rep. 5, 16500 (2015).
Chu, B. et al. Genetic regulation of defence responses in cotton to insect herbivores. AoB Plants 9, plx048 (2017).
Heil, M. Nectar: Generation, regulation and ecological functions. Trends Plant Sci. 16, 191–200 (2011).
Rico-Gray, V. & Oliveira, P. S. The Ecology and Evolution of Ant–Plant Interactions (University of Chicago Press, Chicago, 2013). https://doi.org/10.7208/chicago/9780226713540.001.0001.
Pacelhe, F. T., Costa, F. V., Neves, F. S., Bronstein, J. & Mello, M. A. R. Nectar quality affects ant aggressiveness and biotic defense provided to plants. Biotropica 51, 196–204 (2019).
Oliveira, P. S., Rico-Gray, V., Díaz-Castelazo, C. & Castillo-Guevara, C. Interaction between ants, extrafloral nectaries and insect herbivores in Neotropical coastal sand dunes: Herbivore deterrence by visiting ants increases fruit set in Opuntiastricta (Cactaceae). Funct. Ecol. 13, 623–631 (1999).
Koptur, S., William, P. & Olive, Z. Ants and plants with extrafloral nectaries in fire successional habitats on Andros (Bahamas). Fla. Entomol. 93, 89–99 (2010).
Cuautle, M., Rico-Gray, V. & Diaz-Castelazo, C. Effects of ant behaviour and presence of extrafloral nectaries on seed dispersal of the Neotropical myrmecochore Turneraulmifolia L. (Turneraceae). Biol. J. Linn. Soc. 86, 67–77 (2005).
Rico-Gray, V. & Thien, T. B. Ant-mealybug interaction decreases reproductive fitness of Schomburgkiatibicinis (Orchidaceae) in Mexico. J. Trop. Ecol. 5, 109–112 (1989).
Raine, N. E., Gammans, N., Macfadyen, I. J., Scrivner, G. K. & Stone, G. N. Guards and thieves: Antagonistic interactions between two ant species coexisting on the same ant–plant. Ecol. Entomol. 29, 345–352 (2004).
Villamil, N., Boege, K. & Stone, G. N. Testing the distraction hypothesis: Do extrafloral nectaries reduce ant–pollinator conflict?. J. Ecol. 107, 1377–1391 (2019).
Rico-Gray, V. & Thien, L. B. Effect of different ant species on reproductive fitness of Schomburgkiatibicinis (Orchidaceae). Oecologia 81, 487–489 (1989).
Diaz-Castelazo, C., Rico-Gray, V., Oliveira, P. S. & Cuautle, M. Extrafloral nectary-mediated ant–plant interactions in the coastal vegetation of Veracruz, Mexico: Richness, occurrence, seasonality, and ant foraging patterns. Ecoscience 11, 472–481 (2004).
Author, M., Rudgers, J. A. & Rudgers, J. A. Enemies of herbivores can shape plant traits: Selection in a facultative ant–plant. Source Ecol. Ecol. 85, 192–205 (2004).
Quijano-Medina, T., Covelo, F., Moreira, X. & Abdala-Roberts, L. Compensation to simulated insect leaf herbivory in wild cotton (Gossypiumhirsutum): Responses to multiple levels of damage and associated traits. Plant Biol. 21, 805–812 (2019).
Lang, A. & Otto, M. A synthesis of laboratory and field studies on the effects of transgenic Bacillusthuringiensis (Bt) maize on non-target Lepidoptera. Entomol. Exp. Appl. 135, 121–134 (2010).
Schmidt, J. E. U., Braun, C. U., Whitehouse, L. P. & Hilbeck, A. Effects of activated Bt transgene products (Cry1Ab, Cry3Bb) on immature stages of the ladybird Adaliabipunctata in laboratory ecotoxicity testing. Arch. Environ. Contam. Toxicol. 56, 221–228 (2009).
Romeis, J., Meissle, M. & Bigler, F. Transgenic crops expressing Bacillusthuringiensis toxins and biological control. Nat. Biotechnol. 24, 63–71 (2006).
Yu, H., Romeis, J., Li, Y., Li, X. & Wu, K. Acquisition of Cry1Ac protein by non-target arthropods in Bt soybean fields. PLoS ONE 9, e103973 (2014).
Acknowledgements
We are grateful for the support of all people in the communities where our work was developed. The research about wild cotton in Mexico is possible thanks to the support and mentorship of the Daniel Piñero, Francisca Acevedo, and José Sarukhan. Also, we thank Martin Heil and Rosa Adame for their advice and consultation throughout the research process. In addition, we would like to acknowledge Eunice Rojas, Pamela Rodríguez, Sergio Soria and Javier Pérez for their help and support in the fieldwork. We wish to thank Valeria Alavez, Ivalú Cacho, Ek del Val, Alejandra Hernández-Terán, and Deborah Gordon for their comments on our work. Furthermore, VV-B thanks Horacio Sepulveda, Denise Arroyo, Greta Ramirez, Edgar Reyna, and Monica Pérez Monterrubio for their help in reviewing the grammar of this work. KB acknowledges logistic support by Ruben Pérez-Ishiwara while writing this article. This research is dedicated to Alejandra Barrios Pérez, who through her work in SEMARNAT has defended the genetic resources of Mexico. This work constitutes the master’s degree research of VV-B, who received a scholarship from the Consejo Nacional de Ciencia y Tecnología (CONACyT, scholarship no. 477713), and extends thanks to the Graduate Program Posgrado en Ciencias Biológicas at Universidad Nacional Autónoma de México (UNAM).
Funding
This work was financially supported by the project “Program for the conservation of wild populations of Gossypium hirsutum in Mexico”, DGAP003/WN003/18 funded by CONABIO, and the project UNAM-PAPIIT No. IN214719; complemented with the support of CONACYT scholarships (477713). The funders had no role in the study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
V.V.-B. and A.W. conceived the study and performed the field and analytical work. V.V.-B., K.B., and A.W. undertook and discussed the data analysis. V.V.-B. and T.S.-F. performed the lab experiments, and A.W. supervised the laboratory tests. P.R.-F. and V.V.-B. identified the species of ants. V.V.-B., K.B., and A.W. wrote the manuscript. A.W. coordinated the study. All authors contributed to and approved the final draft.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Vázquez-Barrios, V., Boege, K., Sosa-Fuentes, T.G. et al. Ongoing ecological and evolutionary consequences by the presence of transgenes in a wild cotton population. Sci Rep 11, 1959 (2021). https://doi.org/10.1038/s41598-021-81567-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-81567-z
This article is cited by
-
Agronomic and phenotypic plant traits as indicators for environmental risks of genetically modified plants
Environmental Sciences Europe (2024)
-
Hybrid Bt cotton is failing in India: cautions for Africa
Environmental Sciences Europe (2023)
-
The need for assessment of risks arising from interactions between NGT organisms from an EU perspective
Environmental Sciences Europe (2023)
-
Wild papaya shows evidence of gene flow from domesticated Maradol papaya in Mexico
Genetic Resources and Crop Evolution (2023)
-
An experimental test of ant effects on herbivory and pathogen infection on wild cotton (Gossypium hirsutum L.)
Arthropod-Plant Interactions (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.