Abstract
Viruses in the Luteoviridae family, such as Potato leafroll virus (PLRV), are transmitted by aphids in a circulative and nonpropagative mode. This means the virions enter the aphid body through the gut when they feed from infected plants and then the virions circulate through the hemolymph to enter the salivary glands before being released into the saliva. Although these viruses do not replicate in their insect vectors, previous studies have demonstrated viruliferous aphid behavior is altered and the obligate symbiont of aphids, Buchnera aphidocola, may be involved in transmission. Here we provide the transcriptome of green peach aphids (Myzus persicae) carrying PLRV and virus-free control aphids using Illumina sequencing. Over 150 million paired-end reads were obtained through Illumina sequencing, with an average of 19 million reads per library. The comparative analysis identified 134 differentially expressed genes (DEGs) between the M. persicae transcriptomes, including 64 and 70 genes that were up- and down-regulated in aphids carrying PLRV, respectively. Using functional classification in the GO databases, 80 of the DEGs were assigned to 391 functional subcategories at category level 2. The most highly up-regulated genes in aphids carrying PLRV were cytochrome p450s, genes related to cuticle production, and genes related to development, while genes related to heat shock proteins, histones, and histone modification were the most down-regulated. PLRV aphids had reduced Buchnera titer and lower abundance of several Buchnera transcripts related to stress responses and metabolism. These results suggest carrying PLRV may reduce both aphid and Buchnera genes in response to stress. This work provides valuable basis for further investigation into the complicated mechanisms of circulative and nonpropagative transmission.
Similar content being viewed by others
Introduction
Aphids are a member of the superfamily Aphidoidea, are distributed world-wide, and cause major damage to global agricultural1. Despite there being over 4000 species, only about 400 are known as significant pests2. Aphids are effective pests partially because they do not require sexual reproduction and can use parthenogenesis to quickly increase their numbers1,2. Another aspect of aphid biology that makes them an effective pest is their host range. Although many aphids are very specialized herbivores, only feeding on a few related species, some species feed on many taxa of plants. Myzus persicae is one of these polyphagous pests, feeding on over 40 different families, including Solanaceae1,2.
Along with causing direct feeding damage, aphids are important plant virus vectors, representing over 50% of all known insect vectors for plant viruses3,4,5. Increasing evidence has shown that plant viruses alter vector host finding, dispersal, and inoculation through changes in host physiology, however the underlying mechanisms are largely unknown6,7,8. Recent evidence suggests that viruses may also directly affect aphid biology9,10,11,12. For example, Rhopalosiphum padi that had been fed an artificial diet with purified Barley yellow dwarf virus (BYDV) virions prefer non-infected host plants, while virus-free R. padi prefer infected hosts9. By separating the virus from the host plant the authors demonstrate that the virus alone can impact insect behavior.
Although their life history or host may change, all aphids depend on Buchnera aphidicola as their primary obligate endosymbiont13. Buchnera provide the aphid with essential amino acids and nutrients that are limited in the aphid’s diet14,15,16,17, and because of this aphids can no longer survive without Buchnera. For example, when Buchnera is reduced by using antibiotics, studies have shown lower body mass, lower fecundity, and changes to feeding behavior18,19. As Buchnera co-diversified with aphids over time13,20, essential components of the Buchnera genome have also been lost21,22. Because of this Buchnera also depends on aphids for survival, living inside special aphid cells, known as “bacteriocytes”. Previous studies have speculated Buchnera may have a role in aphid transmission of plant viruses23,24,25,26,27. Specifically, the Buchnera chaperone protein GroEL, a homologue from Escherichia coli28, has been implicated in transmission for a number of viruses23,24,27,29,30,31,32. Direct interactions are thought to be unlikely due to the spatial separation of bacteriocytes and circulating virions23, however, GroEL from Buchnera is found in aphid saliva and has been shown to trigger plant defenses and reduce aphid fecundity using transgenic plants33,34.
Potato leafroll virus (PLRV) is a positive sense ssRNA virus and the type member of the genus Polerovirus (family Luteroviridae). PLRV is phloem limited and transmitted in a circulative nonpropagative manner. This means the virus particles will travel across the gut membrane on specific receptors into the insect hemolymph. From here it will traverse to the salivary gland and duct so that it may be injected back into the phloem tissue6,35. Previous studies have shown that aphid vectors prefer to settle on plants infected with PLRV and that insect vectors have higher fecundity when feeding on these plants compared to controls36,37. Recently we demonstrated PLRV induces changes in plant nutrients and defenses in infected host plant38, however, the impacts of these changes on symbiont-aphid interactions are unknown. To address this lack of knowledge we examined changes in the transcriptome of M. persicae with and without PLRV, Buchnera titer, and changes in aphid and Buchnera transcripts from aphids feeding on PLRV-infected plants. By providing evidence that nonpropagative circulative plant viruses can affect insect vectors through changes in the transcriptome and alter Buchnera titer, our study will contribute to growing knowledge of the insect microbiome at a plant–insect interface.
Methods
Plant and insect growth conditions
Solanum tuberosum were propagated using leaf-bud cutting from cv. Désirée39 in laboratory experiments. Plants were grown in growth chambers under controlled conditions (25/23 °C day/night with a photoperiod of 16/8 h day/night). Non-viruliferous and viruliferous aphid clones of a potato-adapted red strain of Myzus persicae were reared under controlled conditions (25/23 °C day/night with a photoperiod of 16/8 h day/night) on healthy potato. We confirmed our colony was free of secondary symbionts using universal primers for the bacteria 16S–23S ribosomal RNA intergenic spacer region (Supplementary Table S1). All experiments were conducted in the same environmental chambers and conditions, so there were no environmental differences in treatments (25/23 °C day/night with a photoperiod of 16/8 h day/night).
Pathogen infection
Agrobacterium tumefaciens (LBA4404) containing the infectious clone of PLRV40 was grown at 28 °C in LB broth (+ 10 mM MgSO4), with kanamycin (50 µg/mL), carbenicillin (100 µg/mL) and rifampicin (50 µg/mL) for selection. After 24 h, bacteria were centrifuged to concentrate and resuspended in 10 mM MgCl2. One-week-old S. tuberosum were inoculated at an optical density (OD) of 0.70. Three weeks post infection, tissue was collected from all plants, RNA was extracted using the SV Total Isolation Kit as per manufacturer’s instructions (Promega, Madison, WI, USA), and cDNA was synthesized using 1500 ng of total RNA and random hexamers (20 ng/µL) with the SMART® MMLV as per manufacturer’s instructions (Takara Bio USA, Mountain view, CA, USA). cDNAs were used in PCRs with PLRV specific primers (F-5’ATGAGTACGGTCGTGGTT-3’ and R-5 ‘CTATTTGGGGTTTTGCAAAGC-3’). A set of uninfected potato cuttings were grown at the same time as the plants above to serve as controls. After systemic plant infection was verified plants were immediately used in experiments.
RNAseq, qRT-PCR, and qPCR aphid experiments
One week after infection was verified, five adult aphids were placed on the first fully expanded leaflet of three infected and three healthy plants. After 24 h, all adults were removed and 20 larvae were left to develop. Seven days later all aphids were at the same developmental stage. Ten adult aphids were collected into a tube from each plant (N = 3 plants with 10 aphids per plant) and immediately frozen in liquid nitrogen until use in RNAseq experiments. The entire experiment was repeated a second time for confirmation of RNAseq results using qRT-PCR and to examine the titer of the bacterial symbiont, Buchnera, in aphids. For this experiment 5 aphids were collected for RNA extraction and 5 aphids were collected for DNA extractions from each plant. We also prepared 6–7 plants for each treatment instead of 3 (N = 6–7 with 5 aphids per plant), however all other methods were the same.
RNA and DNA isolation from aphids
RNA was extracted from aphid tissue collected in the first two experiments as described above. The RNA concentration and purity were measured using a NanoDrop. The integrity of RNA was confirmed using the Bioanalyzer 2100 system (Agilent, Santa Clara, CA, USA). DNA was extracted from aphid tissue collected in the second experiment using cetyl trimethylammonium bromide. The integrity of DNA was confirmed using an agarose gel. The DNA concentration and purity were measured using a NanoDrop (Thermo Fisher Scientific, Waltham, MA, USA).
Library preparation, and sequencing
Sequencing libraries were prepared using a multiplexing library protocol41. Briefly, oligo(dT) 25 Dynabeads were used to purify mRNA, which was then fragmented, and the first-strand cDNA was synthesized using random primers, dNTP, and reverse transcriptase. The second-strand was synthesized using a dUTP mix, DNA Polymerase I, and RNase H, ends repaired, and adenylated. The cDNA fragments were ligated to adapters, selectively enriched by PCR, and purified using the AMPure XP beads. The library quality was assessed using the Agilent Bioanalyzer 2100 system and sequenced using an Illumina HiSeq 2000 instrument.
Read mapping, differential expressed gene (DEG) analysis, and Gene Ontology (GO) classification
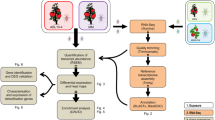
RNA-Seq data were analyzed using RStudio (Version 1.1.383) and Bioconductor according to Anders et al. (2013) with some modifications39 (See Supplementary Fig. S1). Sequence quality was determined, trimmed, and poor-quality reads removed using ShortRead42,43 and FastQC44. Reads were mapped to the Myzus persicae clone G006 genome v2.0 from AphidBase45 using TopHat246. Mapped reads were assigned to genes and counted with HTSeq47, and normalized by size factors obtained from the negative binomial-based DESeq2 package48. Gene annotation files were downloaded from NCBI. After normalization, clusterization profiles of the samples were assessed by hierarchical clustering (with Euclidean distance metric and Ward´s clustering method) and principal component analysis (PCA). Differentially expressed genes (DEGs) between infected and control treatments were identified using DESeq248. Genes with False Discovery Rate (FDR)-corrected p-values ≤ 0.1 were classified as differentially expressed. For Gene ontology (GO) analysis, Blast2GO software49 was utilized for annotation as previously described50.
qRT-PCR
cDNA was synthesized using random hexamer (20 ng/µL) and quantitative RT-PCR (qRT-PCR) was performed. Transcript abundance was quantified for the M. persicae genes, Hsp68-like (MYZPE13164_G006_v1.0_000070430.1) and M. persicae cuticle protein5-like (Mpcp5-like) (MYZPE13164_G006_v1.0_000133030.2), and for the Buchnera genes, argE (BUMPUSDA_CDS00542), dnaK (BUMPUSDA_CDS00441), and groEL (BUMPUSDA_CDS00567), using gene specific primers (Supplementary Table S1). qRT-PCR reactions were carried out using SYBR green PCR master mix (Applied Biosystems, Carlsbad, CA, USA), in an CFX384 instrument (Bio-Rad, Hercules, CA, USA). Three technical replicates were performed for each individual sample, and a digital pipette was used for all pipetting. Relative transcript abundance was calculated utilizing a standard curve produced from tenfold series dilution of cDNA synthesized from 1000 ng/μL of total RNA according to the standard curve method (Applied Biosystems, Carlsbad, CA, USA). Technical replicates of raw CT values were averaged and transcripts of interest were normalized to the house-keeping transcript ribosomal protein L7 rpl7 for aphids or the 50S ribosomal subunit gene rpIN for Buchnera, as previously described51,52,53.
Buchnera titer
Buchnera titer here is defined as the ratio of Buchnera single copy genes to aphid single copy genes. To determine Buchnera titer in whole aphid bodies we used qPCR and measured the ratio of a single copy Buchnera gene (rplN) to a single copy aphid gene (RPL7). qPCR reactions were carried out using SYBR green PCR master mix (Applied Biosystems, Carlsbad, CA, USA), in the CFX384 instrument (Bio-Rad, Hercules, CA, USA). Reactions were performed in triplicate for each sample, and the average was used for analysis. Relative abundance was calculated utilizing a standard curve produced from tenfold serial dilution of DNA.
Statistical analyses
RNAseq data analyses were performed as described above. All statistical analyses for qRT-PCR and qPCR were determined using a P < 0.05. Analysis of variance (ANOVA) was used to determine significant difference in transcript abundance. For Buchnera titer single factor ANOVA was used to determine difference in relative abundance. The statistical analyses were performed using JMP 8 software (SAS Institute, Cary, NC, USA).
Compliance statement
All research in this paper complied with relevant institutional, national, and international guidelines and legislation.
Results
Differential gene expression in the presence of PLRV
Over 150 million paired-end reads were obtained through Illumina sequencing, with an average of 125,815, 247 100-bp reads per library (Table 1, Fig. 1A). A higher number of average reads were sequenced in control 1 compared to other samples. Nevertheless, reads mapped to the target genome in the same relative proportions across treatments (Fig. 1B) revealing that the sample was not contaminated with reads other than for the target genome. Further, read counts were normalized before differential gene analyses (see methods) accounting for variation in library sizes among treatments. About 73% of the reads mapped to the M. persicae reference genome, with around only 27% being uniquely mapped (Table 1, Fig. 1B). In order to examine biological variability, a principal component analysis (PCA) of the normalized count data was performed (Fig. 1C). The first component of variance separated samples by treatments and accounted for 54% of the variance. Hierarchical clustering confirmed PCA results in visual representation of DEG expression (Fig. 1D). The transcriptome of viruliferous M. persicae was compared to the transcriptome of virus-free M. persicae using the negative binomial-based DESeq248. Overall, 96 differentially expressed genes (DEGs) were detected using an FDR adjusted p-value ≤ 0.05 and log2 fold change (FC) ≥ 0.5, however, by relaxing our FDR adjusted p-value to ≤ 0.1 we were able to include 38 additional DEGs (134 DEGs total included; Fig. 1E; Supplementary Table S2). The presence of PLRV in the aphid vector caused a down-regulation of 70 genes and an up-regulation of 64 genes (Fig. 1F; FDR adjusted p-value ≤ 0.1 and log2 fold change (FC) ≥ 0.5). Overall, 0.8% of the aphid genome was significantly impacted by the presence of PLRV.
Overview of Myzus persicae transcriptome after Potato leafroll virus (PLRV) acquisition. (A) Number of paired-end reads generated for each library by Illumina HiSeq sequencing. The dashed line represents the average of paired-end reads from all 6 libraries. (B) Proportion of uniquely mapped, multimapped, and unmapped reads obtained for each library. Reads were mapped in the Myzus persicae clone G006 genome (AphidBase). (C) Principal component analysis of normalized count data from all samples. (D) Hierarchical clustering analysis of normalized count data z-scores exhibited by differentially expressed genes (DEGs) within each sample. (E) Volcano-plots of − log10p and log2FC exhibited by each gene in viruliferous aphids compared to controls. Up- and down-regulated genes are presented in red and green, respectively. (F) Numbers of up- and down-regulated DEGs in viruliferous aphids in comparison to control aphids. DEGs were identified using DESeq2 and defined by |log2FC|≥ 0.5 and false discovery rate (FDR)-corrected p-value ≤ 0.1. C = control aphids without virus; V = viruliferous aphids carrying PLRV, p = FDR-corrected p-value.
Functional roles of differentially expressed genes
Gene ontology (GO) enrichment analyses were performed with the DEGs from each treatment to identify functions and pathways disturbed in aphids carrying PLRV. One or more gene ontology terms were assigned to each transcript from biological processes, molecular functions, and cellular compartments term using Blast2GO functional gene annotation49. The 134 DEGs were assigned to functional GO terms within the three categories, including 125 biological processes, 118 molecular functions, and 148 cellular compartments. Of the 134 DEGs, 53 (39.55% of total DEGs) were classified as “uncharacterized proteins.” The majority of DEGs assigned to biological processes were categorized as metabolic processes (41%), cellular-protein processes (11%), and oxidation–reduction processes (11%) (Fig. 2A). As for DEGs assigned to molecular functions, almost half were associated with catalytic activity (33%) or nucleic acid binding (21%) (Fig. 2B). Within the cellular component category, 24% were related to the membrane and 24% were related to intracellular locations (Fig. 2C).
Blast2GO Gene Ontology of DEGs arranged by functional categories. (A) Biological processes (BP), (B) molecular function (MF), and (C) cellular component (CC). The predicted gene functions of differentially expressed genes as assigned by Blast2GO at level 2–3 in each aforementioned category. Each DEG may be assigned to one or more GOterm, with a total of 391 GOterms from the three functional groups assigned to the 134 DEGs.
Next each DEGs was annotated using a single Blast2GO consensus description. Many of the genes up-regulated in PLRV aphids were related to cuticle formation and development (16%), and catalytic activity (16%), however the majority of up-regulated transcripts were uncharacterized (31%; Fig. 3A). The largest groups of down-regulated genes in PLRV aphids were related to histones (10%), catalytic activity (10%), transmembrane transport (9%), proteolysis or protein ubiquitination (7%), and nucleic acid binding and metabolic processes (7%; Fig. 3B). A significant proportion of the down-regulated transcripts in PLRV aphids were also uncategorized (47%). The most highly expressed DEGs included transcripts related to cuticle formation and development, and 4C1-like cytochrome P450s (Table 2, Supplemental Table 2). The most down-regulated transcripts in PLRV aphids were related to histones and histone modifying proteins (Table 3).
Validation of select aphid transcripts via RT-qPCR
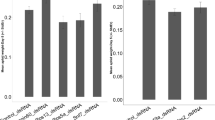
To validate the RNAseq we conducted qRT-PCR analysis of select DEGs in a separate experiment using the same experimental design. We selected one of the most up-regulated genes (MYZPE13164_G006_v1.0_ 000133030.2 (Mpcp5-like)) and one of the most down-regulated gene (MYZPE13164_G006_v1.0_ 000070430.1 (Hsp68-like)) and measured transcript abundance using qRT-PCR (bolded genes in Tables 2 and 3). Mpcp5-like is related to cuticle formation and development and Hsp68-like encodes a heat shock protein, which are related to immunity and stress responses. Consistent with the RNA-seq data, abundance of the Mpcp5-like transcript was significantly higher in viruliferous M. persicae compared to virus-free controls (10.045, 2.017, relative expression respectively; p = 0.019; Fig. 4A). Abundance of the Hsp68-like transcript was significantly lower in viruliferous M. persicae when compared to virus-free controls (1.44, 5.51, relative expression respectively; p < 0.01) (Fig. 4A,B).
Relative transcript abundance of two genes in Myzus persicae with and without Potato leafroll virus (PLRV). (A) A cuticle related protein (Mpcp5-like) transcript was significantly up-regulated in expression in individuals with PLRV compared to virus free controls. (B) A predicted heat shock protein (Hsp68-like) was significantly down-regulated in expression in individuals with PLRV compared to controls. Transcripts were measured relative to a housekeeping gene RPL7. Significant differences were calculated using an ANOVA (*P < 0.05; Error bars represent ± SEM).
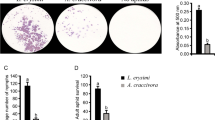
The impact of PLRV on Buchnera aphidicola titer
Buchnera has been previously implicated in transmission of PLRV and other luteoviruses23,25,27,31,54, however Buchnera titer and coding sequence transcripts have not been examined in aphids carrying PLRV. From our experiments, Buchnera titer was ~ 1.5 times higher for virus-free aphids compared to aphids carrying PLRV (ratios 6.42, 4.20 respectively; p = 0.037; Fig. 5A). To investigate the potential mechanisms mediating decreases in Buchnera titer we measured abundance of two transcripts related to stress, dnaK55,56 and groEL56,57, and one transcript related to metabolism, argE58. Abundance of all three transcripts were reduced in aphids carrying PLRV compared to controls. Viruliferous aphids had 63.32% less argE transcripts (p = 0.026), 83.33% less groEL transcripts (p = 0.024), and 81.23% less dnaK transcripts (p = 0.046) compared to that of the virus free aphids (Fig. 5B–D).
Buchnera aphidicola titer and transcript changes in Myzus persicae with Potato leafroll virus (PLRV). (A) Ratio of a single copy Buchnera gene rpIN relative to a single copy aphid gene RPL7, demonstrates M. persicae with PLRV have a decreased Buchnera aphidicola titer relative to control aphids. (B–D) Buchnera transcripts groEL, dnaK, and argE relative to the Buchnera gene rpIN housekeeping gene. All three transcripts were down-regulated in viruliferous M. persicae compared to virus free controls. Significant differences were calculated using an ANOVA (*P < 0.05; Error bars represent ± SEM).
Discussion
The main focus of this paper was to examine the effect that Potato leafroll virus has on the transcriptome of M. persicae, and their primary endosymbiont Buchnera aphidicola. The largest category of known up-regulated transcripts in viruliferous aphids compared to controls were related to the cuticle and cuticle development. Insect cuticles are largely composed of a protein matrix embedded with chitin filaments59. Cuticle proteins (CPs) have been shown to be involved in general development, molting, transmission of non-persistent viruses, and insecticide resistance through changes in cuticle permeability30,60,61,62,63. In Acyrthosiphon pisum, 19 CPs were found to be regulated by photoperiodism and suspected to be involved in the transition from asexual to sexual production64. Further, cuticle proteins have been implicated as potentially facilitating transmission of Barley yellow dwarf virus (BYDV-GPV), Cereal yellow dwarf virus (CYDV-RPV), and Turnip yellows virus (TuYV), three related Luteoviridae viruses65,66,67. Whilst we cannot know the function of changes in CP transcripts in PLRV-aphid interactions from these experiments, these genes represent promising targets for further investigation.
In addition to many cuticle related proteins, five cytochrome P450s genes were significantly up-regulated in viruliferous aphids compared to controls. Cytochrome P450s play important roles in hormone and pheromone metabolism but are more famous for their roles in the metabolism of insecticides and host plant chemicals. Polyphagous insects, like M. persicae, encounter many different hosts and tend to have a higher number of P450s related to the metabolism of allelochemicals compared to more specialized aphids68. Previous work has shown that a cytochrome 450 gene (CYP6CY3) was found to increase nicotine tolerance and aphid host adaptation69,70. It has been previously hypothesized that up-regulation of p450s could help insect vectors tolerate less desirable hosts which could be beneficial to the virus71.
Transcripts encoding a heat shock protein (HSP68-like) was among the most down-regulated in viruliferous aphids compared to controls. HSP68 is a member of the HSP70 family, which are important chaperone proteins that are known to be up-regulated in response to stress. One study found that the Hsp70 from Bemisia tabaci is up-regulated after acquisition of Tomato yellow leaf curl virus (TYLCV)72. They went on to show that HSP70 protein can directly interact with TYLCV using in vitro studies and that they co-localize together in insect midgut cells using in situ hybridization. The authors suggest HSP70 may play an inhibitory role in virus transmission, as transmission was increased when whiteflies were fed HSP70 antibodies. Given Hsp68-like transcripts were down-regulated in aphids carrying PLRV in our study, it would be interesting to investigate if this has any impact on PLRV transmission. Porras et al.73 demonstrated that BYDV-PAV, a strain that is only transmitted by Rhopalosiphum padi (bird-cherry oat aphid), up-regulated the abundance of three Hsp70 transcripts in the aphid vector. The authors found BYDV infection increases plant surface temperature and aphid heat tolerance, suggesting a protective role of HSP70 proteins in virus-aphid-plant interactions73. Although it is not known if PLRV increases plant surface temperature and vector heat tolerance, it has been shown that potato plants kept at higher temperatures are more susceptible to PLRV than compared to lower temperatures74. Also aphid acquisition and transmission at higher temperatures resulted in higher transmission rates compared to lower temperatures75, however at very high temperatures differences were reduced76. It is not known how decreases in Hsp68-like transcripts in aphids carrying PLRV may alter aphid heat tolerance.
In this study there was a significant reduction of Buchnera titer and Buchnera gene expression of three genes (dnaK, groEL, and argE) in aphids carrying PLRV compared to control aphids. In general, gene regulation at the mRNA level in Buchnera is thought to be minimal because Buchnera transcription factors are reduced58 and very few transcriptional responses had been observed previously77. Only two transcription initiation factors (\(\sigma\) 32 and \(\sigma\) 70), the heat shock and housekeeping transcription factors, respectively, remain in Buchnera Myzus’s genomes78 similar to other Buchnera taxa79,80. The housekeeping sigma factor (\(\sigma\) 70) initiates transcription of argE which is regulated by the repressor ArgR when bound to arginine in Escherichia coli81. Similar to other Buchnera taxa, Buchnera Myzus’s genome78 has lost the repressor ArgR so it is unclear how this gene is down-regulated in virus-infected aphids compared to un-infected aphids. The other two Buchnera genes (dnaK and groEL) that were down-regulated in this study in aphids carrying PLRV compared to control aphids are associated with the heat shock regulon80. Moreover, these Buchnera genes still retain recognizable \(\sigma\) 32 promoter sites up-stream of dnaK and groEL in the Myzus Buchnera G006 genome (NCBI Reference Sequence: NZ_MJNC01000001; Supplemental Table 3) similar to other Buchnera taxa80. The \(\sigma\) 32 heat shock response is highly conserved in bacteria and is initiated in response to stress, such as heat shock or other environmental stressors that destabilize proteins78,79,80,81. In this study it is unclear how PLRV is either directly or indirectly dampening Buchnera’s expression of dnaK and groEL and if it is through a similar mechanism that is also down-regulating the aphid’s stress response genes including Hsp70.
A decrease in Buchnera titer has previously been associated with different aphid clones83, plant diets13, increasing aphid nymphal age82,84, and heat shock81,84,86. Most obligate pathogens and symbionts, including Buchnera, overexpress the protein GroEL during non-heat shock conditions to rescue misfolded proteins87. We hypothesize that PLRV is reducing Buchnera’s ability to up-regulate genes that are associated with the heat shock regulon (Fig. 5C,D) and this may lead to increased stress, lysing of Buchnera cells, and ultimately a reduction of Buchnera titer (Fig. 5A). Other insect-plant pathogen systems are known to modulate obligate symbiont titer. For example, in whiteflies Portiera titer is modulated by the co‐occurrence of its facultative symbiont Rickettsia and TYLCV88. Alternatively, as PLRV-infected plants have higher concentrations of free amino acids38, the change in host plant diet may have influenced Buchnera titer similar to Zhang et al.13. Because aphids carrying PLRV may obtain higher levels of essential amino acids from virus-infected plants, Buchnera genes that are involved in arginine biosynthesis, such as argE, may be down-regulated compared to aphids feeding on un-infected plants with lower amounts of essential amino acids.
Parasites of plants and animals can modify host behavior to improve their own transmission and survival6,7,89,90. This work explores the complex relationships that exist between hosts, viruses, vectors, and endosymbionts, and opens up more questions regarding the complexity and depth of these relationships. Aphids and bacterial endosymbionts may benefit from relationships with plant-infecting viruses indirectly or directly but additional studies are needed. Although it is known that Buchnera titer and gene expression responses vary with aphid linages91, it is not known how this is impacted by long term associations with plant-infecting viruses. In regions where virus pressure is high or where poor hosts dominate, aphids may more often be associated with plant infecting viruses. This study also expands on previous work that given the mounting evidence of virus manipulation of insect vectors, this could have lasting impacts on the population structures of these insect vectors and their obligate endosymbiont.
References
Remaudiere, G., & Remaudiere, M. Catalogue of the World’s Aphididae: Homoptera Aphidoidea. 473–1275. (Institut National de la Recherche Agronomique (INRA), 1997).
Fereres, A., Irwin, M.E., & Kamppeier, G.E. Aphid movement: Process and consequences. in (van Emden H.F.R.H. ed.) Aphids as Crop Pests. 2nd edn. 196–200. (CABI, 2017).
Ng, J. C. K. & Perry, K. L. Transmission of plant viruses by aphid vectors. Mol. Plant Pathol. 5(5), 505–511. https://doi.org/10.1111/j.1364-3703.2004.00240.x (2004).
Whitfield, A. E., Falk, B. W. & Rotenberg, D. Insect vector-mediated transmission of plant viruses. Virology 479–480, 278–289. https://doi.org/10.1016/j.virol.2015.03.026 (2015).
Elena, S. F., Bernet, G. P. & Carrasco, J. L. The games plant viruses play. Curr. Opin. Virol. 8, 62–67. https://doi.org/10.1016/j.coviro.2014.07.003 (2014).
Casteel, C.L., & Falk, B.W. Plant virus-vector interactions: More than just for virus transmission. in (Wang, A., & Zhou, X. eds.) Current Research Topics in Plant Virology. 2016. 217–240. https://doi.org/10.1007/978-3-319-32919-2_9 (2016).
Eigenbrode, S. D., Bosque-Pérez, N. A. & Davis, T. S. Insect-borne plant pathogens and their vectors: Ecology, evolution, and complex interactions. Annu. Rev. Entomol. 63, 169–191. https://doi.org/10.1146/annurev-ento-020117-043119 (2018).
Blanc, S. & Michalakis, Y. Manipulation of hosts and vectors by plant viruses and impact of the environment. Curr. Opin. Insect Sci. 16, 36–43. https://doi.org/10.1016/j.cois.2016.05.007 (2016).
Ingwell, L. L., Eigenbrode, S. D. & Bosque-Pérez, N. A. Plant viruses alter insect behavior to enhance their spread. Sci. Rep. 2(1), 578. https://doi.org/10.1038/srep00578 (2012).
Stafford, C. A., Yang, L. H., Mcmunn, M. S. & Ullman, D. E. Virus infection alters the predatory behavior of an omnivorous vector. Oikos 123, 1384–1390. https://doi.org/10.1111/oik.01148 (2014).
Wang, Q. et al. Rice dwarf virus infection alters green rice leafhopper host preference and feeding behavior. PLoS ONE 13(9), 1–16. https://doi.org/10.1371/journal.pone.0203364 (2018).
Stafford, C. A., Walker, G. P. & Ullman, D. E. Infection with a plant virus modifies vector feeding behavior. Proc. Natl. Acad. Sci. 108(23), 9350–9355. https://doi.org/10.1073/pnas.1100773108 (2011).
Zhang, Y. C., Cao, W. J., Zhong, L. R., Godfray, H. C. J. & Liu, X. D. Host plant determines the population size of an obligate symbiont (Buchnera aphidicola) in aphids. Appl. Environ. Microbiol. 82(8), 2336–2346. https://doi.org/10.1128/AEM.04131-15 (2016).
Hansen, A. K. & Moran, N. A. Aphid genome expression reveals host-symbiont cooperation in the production of amino acids. Proc. Natl. Acad. Sci. 108(7), 2849–2854. https://doi.org/10.1073/pnas.1013465108 (2011).
Nakabachi, A. et al. Transcriptome analysis of the aphid bacteriocyte, the symbiotic host cell that harbors an endocellular mutualistic bacterium, Buchnera. Proc. Natl. Acad. Sci. 102(15), 5477–5482. https://doi.org/10.1073/pnas.1013465108 (2005).
Wernegreen, J. J. Strategies of genomic integration within insect-bacterial mutualisms. Biol Bull. 223(1), 112–122. https://doi.org/10.1086/BBLv223n1p112 (2012).
Zhang, Y. et al. Genetic structure of the bacterial endosymbiont, Buchnera aphidicola, from its host aphid, Schlechtendalia chinensis, and evolutionary implications. Curr. Microbiol. 75(3), 309–315. https://doi.org/10.1007/s00284-017-1381-0 (2018).
Zhang, F. et al. Bacterial symbionts, Buchnera, and starvation on wing dimorphism in English grain aphid, Sitobion avenae (F) (Homoptera: Aphididae). Front. Physiol. 6, 155. https://doi.org/10.3389/fphys.2015.00155 (2015).
Machado-Assefh, C. R., Lopez-Isasmendi, G., Tjallingii, W. F., Jander, G. & Alvarez, A. E. Disrupting Buchnera aphidicola, the endosymbiotic bacteria of Myzus persicae, delays host plant acceptance. Arthropod. Plant Interact. 9(5), 529–541. https://doi.org/10.1007/s11829-015-9394-8 (2015).
Douglas, A. E. Nutritional interactions in insect-microbial symbioses: Aphids and their symbiotic bacteria Buchnera. Annu. Rev. Entomol. 43(1), 17–37. https://doi.org/10.1146/annurev.ento.43.1.17 (1998).
Tamas, I. et al. 50 million years of genomic stasis in endosymbiotic bacteria. Science 296(5577), 2376–2379. https://doi.org/10.1126/science.1071278 (2002).
Van Ham, R. C. H. J. et al. Reductive genome evolution in Buchnera aphidicola. Proc. Natl. Acad. Sci. 100(2), 581–586. https://doi.org/10.1073/pnas.0235981100 (2003).
Bouvaine, S., Boonham, N. & Douglas, A. E. Interactions between a Luteovirus and the GroEL chaperonin protein of the symbiotic bacterium Buchnera aphidicola of aphids. J. Gen. Virol. 92(6), 1467–1474. https://doi.org/10.1099/vir.0.029355-0 (2011).
Rana, V. S., Singh, S. T., Priya, N. G., Kumar, J. & Rajagopal, R. Arsenophonus GroEL interacts with CLCuV and is localized in midgut and salivary gland of whitefly B. tabaci. PLoS ONE 7(8), e42168. https://doi.org/10.1371/journal.pone.0042168 (2012).
Kliot, A. & Ghanim, M. The role of bacterial chaperones in the circulative transmission of plant viruses by insect vectors. Viruses 5(6), 1516–1535. https://doi.org/10.3390/v5061516 (2013).
Filichkin, S. A., Brumfield, S., Filichkin, T. P. & Young, M. J. In vitro interactions of the aphid endosymbiotic SymL chaperonin with Barley yellow dwarf virus. J. Virol. 71(1), 569–577. https://doi.org/10.1128/JVI.71.1.569-577.1997 (1997).
van den Heuvel, J. F., Verbeek, M. & van der Wilk, F. Endosymbiotic bacteria associated with circulative transmission of Potato leafroll virus by Myzus persicae. J. Gen. Virol. 75(Pt 10), 2559–2565. https://doi.org/10.1099/0022-1317-75-10-2559 (1994).
Gray, S. M. & Gildow, F. E. Luteovirus-aphid interactions. Annu. Rev. Phytopathol. 41(1), 539–566. https://doi.org/10.1146/annurev.phyto.41.012203.105815 (2003).
Li, C., Cox-Foster, D., Gray, S. M. & Gildow, F. Vector specificity of Barley yellow dwarf virus (BYDV) transmission: Identification of potential cellular receptors binding BYDV-MAV in the aphid, Sitobion avenae. Virology 286(1), 125–133. https://doi.org/10.1006/viro.2001.0929 (2001).
Dombrovsky, A., Gollop, N., Chen, S., Chejanovsky, N. & Raccah, B. In vitro association between the helper component-proteinase of Zucchini yellow mosaic virus and cuticle proteins of Myzus persicae. J. Gen. Virol. 88(5), 1602–1610. https://doi.org/10.1099/vir.0.82769-0 (2007).
van den Heuvel, J. F. et al. The N-terminal region of the luteovirus readthrough domain determines virus binding to Buchnera GroEL and is essential for virus persistence in the aphid. J. Virol. 71(10), 7258–7265. https://doi.org/10.1128/JVI.71.10.7258-7265.1997 (1997).
Morin, S. et al. A GroEL homologue from endosymbiotic bacteria of the whitefly Bemisia tabaci is implicated in the circulative transmission of Tomato yellow leaf curl virus. Virology 256(1), 75–84. https://doi.org/10.1006/viro.1999.9631 (1999).
Chaudhary, R., Atamian, H. S., Shen, Z., Briggs, S. P. & Kaloshian, I. GroEL from the endosymbiont Buchnera aphidicola betrays the aphid by triggering plant defense. Proc. Natl. Acad. Sci. 111(24), 8919–8924. https://doi.org/10.1073/pnas.1407687111 (2014).
Vandermoten, S. et al. Comparative analyses of salivary proteins from three aphid species. Insect Mol. Biol. 23(1), 67–77. https://doi.org/10.1111/imb.12061 (2014).
Gray, S. M., Cilia, M. & Ghanim, M. Circulative, “nonpropagative” virus transmission: An orchestra of virus-, insect-, and plant-derived instruments. Adv. Virus Res. 2014, 89. https://doi.org/10.1016/B978-0-12-800172-1.00004-5 (2014).
Eigenbrode, S. D., Ding, H., Shiel, P. & Berger, P. H. Volatiles from potato plants infected with Potato leafroll virus attract and arrest the virus vector, Myzus persicae (Homoptera: Aphididae). Proc. Biol. Sci. 269(1490), 455–460. https://doi.org/10.1098/rspb.2001.1909 (2002).
Rajabaskar, D., Wu, Y., Bosque-Pérez, N. A. & Eigenbrode, S. D. Dynamics of Myzus persicae arrestment by volatiles from Potato leafroll virus-infected potato plants during disease progression. Entomol. Exp. Appl. 148(2), 2. https://doi.org/10.1111/eea.12087 (2013).
Patton, M. F., Bak, A., Sayre, J. M., Heck, M. L. & Casteel, C. L. A polerovirus, Potato leafroll virus, alters plant–vector interactions using three viral proteins. Plant Cell Environ. 43(2), 387–399. https://doi.org/10.1111/pce.13684 (2020).
Sadowy, E., Juszczuk, M., David, C., Gronenborn, B. & Danuta Hulanicka, M. D. Mutational analysis of the proteinase function of Potato leafroll virus. J. Gen. Virol. 82(Pt 6), 1517–1527. https://doi.org/10.1099/0022-1317-82-6-1517 (2001).
DeBlasio, S. L. et al. Insights into the polerovirus– plant interactome revealed by coimmunoprecipitation and mass spectrometry. Mol. Plant-Microbe Interact. 28(4), 467–481. https://doi.org/10.1094/MPMI-11-14-0363-R (2015).
Zhong, S. et al. High-throughput illumina strand-specific RNA sequencing library preparation. Cold Spring Harb. Protoc. 2011(8), 940–949. https://doi.org/10.1101/pdb.prot5652 (2011).
Anders, S. et al. Count-based differential expression analysis of RNA sequencing data using R and bioconductor. Nat. Protoc. 8(9), 1765–1786. https://doi.org/10.1038/nprot.2013.099 (2013).
Morgan, M. et al. ShortRead: A bioconductor package for input, quality assessment and exploration of high-throughput sequence data. Bioinformatics 25(19), 2607–2608. https://doi.org/10.1093/bioinformatics/btp450 (2009).
Andrews, S. FastQC: A Quality Control Tool for High Throughput Sequence Data. http://www.bioinformatics.babraham.ac.uk/projects/fastqc/. (2010).
Gauthier, J. P., Legeai, F., Zasadzinski, A., Rispe, C. & Tagu, D. AphidBase: A database for aphid genomic resources. Bioinformatics 23(6), 783–784. https://doi.org/10.1093/bioinformatics/btl682 (2007).
Kim, D. et al. TopHat2: Accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36. https://doi.org/10.1186/gb-2013-14-4-r36 (2013).
Anders, S., Pyl, P. T. & Huber, W. HTSeq-A Python framework to work with high-throughput sequencing data. Bioinformatics 31(2), 166–169. https://doi.org/10.1093/bioinformatics/btu638 (2015).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15(12), 550. https://doi.org/10.1186/s13059-014-0550-8 (2014).
Conesa, A. et al. Blast2GO: A universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21(18), 3674–3676. https://doi.org/10.1093/bioinformatics/bti610 (2005).
Patton, M. F., Arena, G. D., Salminen, J. P., Steinbauer, M. J. & Casteel, C. L. Transcriptome and defence response in Eucalyptus camaldulensis leaves to feeding by Glycaspis brimblecombei Moore (Hemiptera: Aphalaridae): A stealthy psyllid does not go unnoticed. Austral. Entomol. 57(2), 247–254. https://doi.org/10.1111/aen.12319 (2017).
Casteel, C. L. et al. Disruption of ethylene responses by Turnip mosaic virus mediates suppression of plant defense against the green peach aphid vector. Plant Physiol. 169(1), 209–218. https://doi.org/10.1104/pp.15.00332 (2015).
Nikoh, N. et al. Bacterial genes in the aphid genome: Absence of functional gene transfer from Buchnera to its host. PLoS Genet. 6(2), e1000827. https://doi.org/10.1371/journal.pgen.1000827 (2010).
Hansen, A. K. & Degnan, P. H. Widespread expression of conserved small RNAs in small symbiont genomes. ISME J. 8(12), 2490–2502. https://doi.org/10.1038/ismej.2014.121 (2014).
Hogenhout, S. A., van der Wilk, F., Verbeek, M., Goldbach, R. W. & van den Heuvel, J. F. Potato leafroll virus binds to the equatorial domain of the aphid endosymbiotic GroEL homolog. J. Virol. 72(1), 358–365. https://doi.org/10.1128/JVI.72.1.358-365.1998 (1998).
Camberg, J.L., Doyle, S.M., Johnston, D.M., & Wickner, S. Molecular Chaperones. in Brenner’s Encyclopedia of Genetics. 2nd Edn. 456–60. (Elsevier, 2013). https://doi.org/10.1016/B978-0-12-809633-8.06723-6.
Segal, G. & Ron, E. Z. Regulation and organization of the groE and dnaK operons in Eubacteria. FEMS Microbiol. Lett. 138(1), 1–10. https://doi.org/10.1111/j.1574-6968.1996.tb08126.x (1996).
Zhang, L., Pelech, S. & Uitto, V. J. Bacterial GroEL-like heat shock protein 60 protects epithelial cells from stress-induced death through activation of ERK and inhibition of caspase 3. Exp. Cell Res. 292(1), 231–240. https://doi.org/10.1016/j.yexcr.2003.08.012 (2004).
Shigenobu, S., Watanabe, H., Hattori, M., Sakaki, Y. & Ishikawa, H. Genome sequence of the endocellular bacterial symbiont of aphids Buchnera sp. APS Nat. 407(6800), 81–86. https://doi.org/10.1038/35024074 (2000).
Dombrovsky, A., Sobolev, I., Chejanovsky, N. & Raccah, B. Characterization of RR-1 and RR-2 cuticular proteins from Myzus persicae. Comp. Biochem. Physiol. B Biochem. Mol. Biol. 146(2), 256–264. https://doi.org/10.1016/j.cbpb.2006.11.013 (2007).
Dombrovsky, A., Huet, H., Zhang, H., Chejanovsky, N. & Raccah, B. Comparison of newly isolated cuticular protein genes from six aphid species. Insect Biochem. Mol. Biol. 33(7), 709–715. https://doi.org/10.1016/s0965-1748(03)00065-1 (2003).
Liang, Y. & Gao, X. W. The cuticle protein gene MPCP4 of Myzus persicae (Homoptera: Aphididae) plays a critical role in cucumber mosaic virus acquisition. J. Econ. Entomol. 110(3), 848–853. https://doi.org/10.1093/jee/tox025 (2017).
Silva, A. X., Jander, G., Samaniego, H., Ramsey, J. S. & Figueroa, C. C. Insecticide resistance mechanisms in the green peach aphid Myzus persicae (Hemiptera: Aphididae) I: A transcriptomic survey. PLoS ONE 7(6), e36366. https://doi.org/10.1371/journal.pone.0036366 (2012).
Deshoux, M., Monsion, B. & Uzest, M. Insect cuticular proteins and their role in transmission of phytoviruses. Curr. Opin. Virol. 33, 137–143. https://doi.org/10.1016/j.coviro.2018.07.015 (2018).
Gallot, A. et al. Cuticular proteins and seasonal photoperiodism in aphids. Insect Biochem. Mol. Biol. 40(3), 235–240. https://doi.org/10.1016/j.ibmb.2009.12.001 (2010).
Cilia, M. et al. Genetics coupled to quantitative intact proteomics links heritable aphid and endosymbiont protein expression to circulative polerovirus transmission. J. Virol. 85(5), 2148–2166. https://doi.org/10.1128/JVI.01504-10 (2011).
Wang, H., Wu, K., Liu, Y., Wu, Y. & Wang, X. Integrative proteomics to understand the transmission mechanism of Barley yellow dwarf virus-GPV by its insect vector Rhopalosiphum padi. Sci. Rep. 5, 10971. https://doi.org/10.1038/srep10971 (2015).
Seddas, P. et al. Rack-1, GAPDH3, and actin: proteins of Myzus persicae potentially involved in the transcytosis of Beet western yellows virus particles in the aphid. Virology 325(2), 399–412. https://doi.org/10.1016/j.virol.2004.05.014 (2004).
Yang, Z., Zhang, F., Zhu, L. & He, G. Identification of differentially expressed genes in brown planthopper Nilaparvata lugens (Hemiptera: Delphacidae) responding to host plant resistance. Bull. Entomol. Res. 96(1), 53–59. https://doi.org/10.1079/ber2005400 (2006).
Bass, C. et al. Gene amplification and microsatellite polymorphism underlie a recent insect host shift. Proc. Natl. Acad. Sci. 110(48), 19460–19465. https://doi.org/10.1073/pnas.1314122110 (2013).
Ramsey, J. S. et al. Adaptation to nicotine feeding in Myzus persicae. J. Chem. Ecol. 40(8), 869–877. https://doi.org/10.1007/s10886-014-0482-5 (2014).
Casteel, C. L. & Jander, G. New synthesis: Investigating mutualisms in virus-vector interactions. J. Chem. Ecol. 39(7), 809. https://doi.org/10.1007/s10886-013-0305-0 (2013).
Götz, M. et al. Implication of Bemisia tabaci HEAT SHOCK PROTEIN 70 in Begomovirus-whitefly interactions. J. Virol. 86(24), 13241–13252. https://doi.org/10.1128/JVI.00880-12 (2012).
Porras, M. F. et al. Enhanced heat tolerance of viral-infected aphids leads to niche expansion and reduced interspecific competition. Nat. Commun. 11(1), 1184. https://doi.org/10.1038/s41467-020-14953-2 (2020).
Syller, J. The influence of temperature on transmission of potato leaf roll virus by Myzus persicae Sulz. Potato Res. 30(1), 47–58. https://doi.org/10.1007/BF02357683 (1987).
Syller, J. The effects of temperature on the susceptibility of potato plants to infection and accumulation of Potato Leafroll Virus. J. Phytopathol. 133(3), 216–224. https://doi.org/10.1111/j.1439-0434.1991.tb00156.x (1991).
Chung, B. N. et al. The effects of high temperature on infection by Potato virus Y, Potato virus A, and Potato leafroll virus. Plant Pathol. J. 32(4), 321–328. https://doi.org/10.5423/PPJ.OA.12.2015.0259 (2016).
Hansen, A. K. & Moran, N. A. The impact of microbial symbionts on host plant utilization by herbivorous insects. Mol. Ecol. 23(6), 1473–96 (2014).
Jiang, Z. et al. Comparative analysis of genome sequences from four strains of the Buchnera aphidicola Mp endosymbion of the green peach aphid, Myzus persicae. BMC Genom. 14(1), 917. https://doi.org/10.1186/1471-2164-14-917 (2013).
Enders, L. S. et al. Abiotic and biotic stressors causing equivalent mortality induce highly variable transcriptional responses in the soybean aphid. G3 (Bethesda) 5(2), 261–270. https://doi.org/10.1534/g3.114.015149 (2014).
Wilcox, J. L., Dunbar, H. E., Wolfinger, R. D. & Moran, N. A. Consequences of reductive evolution for gene expression in an obligate endosymbiont. Mol. Microbiol. 48(6), 1491–1500. https://doi.org/10.1046/j.1365-2958.2003.03522.x (2003).
Karp, P. D. et al. The BioCyc collection of microbial genomes and metabolic pathways. Brief Bioinform. 20(4), 1085–1093. https://doi.org/10.1093/bib/bbx085 (2019).
Zhang, B., Leonard, S. P., Li, Y. & Moran, N. A. Obligate bacterial endosymbionts limit thermal tolerance of insect host species. Proc. Natl. Acad. Sci. 116(49), 24712–24718. https://doi.org/10.1073/pnas.1915307116 (2019).
Chong, R. A. & Moran, N. A. Intraspecific genetic variation in hosts affects regulation of obligate heritable symbionts. PNAS 113(46),13114–13119. https://doi.org/10.1073/pnas.1610749113 (2016).
Pers, D. & Hansen, A. K. The boom and bust of the aphid's essential amino acid metabolism across nymphal development. G3 (Bethesda). 11(9), jkab115. https://doi.org/10.1093/g3journal/jkab115 (2021).
Dunbar, H. E., Wilson, A. C. C., Ferguson, N. R. & Moran, N. A. Aphid thermal tolerance is governed by a point mutation in bacterial symbionts. PLoS Biol. 5(5), e96. https://doi.org/10.1371/journal.pbio.0050096 (2007).
Moran, N. A. & Yun, Y. Experimental replacement of an obligate insect symbiont. Proc. Natl. Acad. Sci. 112(7), 2093–2096. https://doi.org/10.1073/pnas.1420037112 (2015).
Fares, M. A., Barrio, E., Sabater-Muñoz, B. & Moya, A. The evolution of the heat-shock protein GroEL from Buchnera, the primary endosymbiont of aphids, is governed by positive selection. Mol. Biol. Evol. 19(7), 1162–1170. https://doi.org/10.1093/oxfordjournals.molbev.a004174 (2002).
Kliot, A., Cilia, M., Czosnek, H., & Ghanim, M. Implication of the bacterial endosymbiont Rickettsia spp. in interactions of the whitefly Bemisia tabaci with Tomato yellow leaf curl virus. J. Virol. 88(10), 5652–5660. https://doi.org/10.1128/JVI.00071-14 (2014).
Dheilly, N. M. et al. Who is the puppet master? Replication of a parasitic wasp-associated virus correlates with host behaviour manipulation. Proc. R. Soc. B Biol. Sci. 2015(282), 20142773 (1803).
Mohan, P. & Sinu, P. A. Does the solitary parasitoid Microplitis pennatulae use a combinatorial approach to manipulate its host?. Entomol. Exp. Appl. 168(4), 295–303 (2020).
Smith, T. E. & Moran, N. A. Coordination of host and symbiont gene expression reveals a metabolic tug-of-war between aphids and Buchnera. Proc. Natl. Acad. Sci. 117(4), 2113–2121. https://doi.org/10.1073/pnas.1916748117 (2020).
Acknowledgements
We thank Laura Baldwin, Sarah Kate Muehleck, and the many undergraduates that helped with experiments. We thank Gabriella Arena for advice on RNAseq analysis. This work was supported by USDA-NIFA award 2017-67013-26537 and award 2013-2013-03265 to CLC.
Author information
Authors and Affiliations
Contributions
C.L.C. conceived the project. M.F.P. and C.L.C. designed the research. M.F.P. and C.L.C. performed research and analyzed the data. A.K.H., M.F.P., and C.L.C. interpreted the data. A.K.H., M.F.P., and C.L.C. wrote the article.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Patton, M.F., Hansen, A.K. & Casteel, C.L. Potato leafroll virus reduces Buchnera aphidocola titer and alters vector transcriptome responses. Sci Rep 11, 23931 (2021). https://doi.org/10.1038/s41598-021-02673-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-02673-6
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.