Abstract
Evolutionary trends may underlie some aspects of the risk for common, non-communicable disorders, including psychiatric disease. We analyzed whole exome sequencing data from 80 unique individuals from India coming from families with two or more individuals with severe mental illness. We used Population Branch Statistics (PBS) to identify variants and genes under positive selection and identified 74 genes as candidates for positive selection. Of these, 20 were previously associated with Schizophrenia, Alzheimer’s disease and cognitive abilities in genome wide association studies. We then checked whether any of these 74 genes were involved in common biological pathways or related to specific cellular or molecular functions. We found that immune related pathways and functions related to innate immunity such as antigen binding were over-represented. We also evaluated for the presence of Neanderthal introgressed segments in these genes and found Neanderthal introgression in a single gene out of the 74 candidate genes. However, the introgression pattern indicates the region is unlikely to be the source for selection. Our findings hint at how selection pressures in individuals from families with a history of severe mental illness may diverge from the general population. Further, it also provides insights into the genetic architecture of severe mental illness, such as schizophrenia and its link to immune factors.
Similar content being viewed by others
Introduction
Severe mental illnesses (SMI) such as schizophrenia and bipolar disorder (BD) have a lifetime prevalence of 1%; and this seems to have remained relatively stable across geography and time1,2. Psychiatric disease syndromes are common, usually begin in young adulthood, are a source of considerable personal and social distress, associated with premature mortality and the treatments have limited efficacy. Hence, detecting underlying mechanisms that may contribute to risk and recovery will be useful.
These syndromes are known to be heritable and their genetic architecture is quite likely to be polygenic, with a combination of common variants of small effect and rare variants of relatively larger effect being implicated3. The apparently uniform genetic epidemiology of these syndromes in different parts of the world seems to suggest that there is no specific selection for, or against, these conditions. This has been attributed to theories of balancing selection, ancestral neutrality or polygenic mutation selection balance4.
From a population genetics standpoint, admixture, migrations and selection all have an impact on our understanding of the genetic contributions to risk of psychiatric illness5. Summary statistics generated from genome wide association study (GWAS) data have been commonly used to investigate the contribution of natural selection on the genetic architecture of complex traits, such as psychiatric syndromes6. Findings from studies investigating the role of natural selection in mental illness have been ambiguous with a few implicating the role of positive selection7,8,9, while others have shown either no evidence for selection or negative selection4,10,11.
Whole exome sequencing, which documents the variation in protein-coding sequences, has also been used as a tool to investigate natural selection. A number of studies in isolated populations have identified genetic variation that confers protection against environmental conditions such as adaptation to hypoxia at high altitudes among Tibetans12 or arctic climate among Nunavik Inuit13 and Siberians14. Signatures of natural selection are detected even in the context of more recent population divergence and this influences many aspects of physiology, underscored by variations in genes that impact on height, blood coagulation, pigmentation, diet availability and resistance to infections15,16.
People with psychiatric illness such as schizophrenia and BD are known to have protective alleles in their genomes17, and these may be associated with resilience18. A study which investigated the evolutionary pattern in the SLC39A8 gene, found that a schizophrenia risk variant in the European population had experienced recent positive selection in Europeans, and that it may have offered protection from the risk of hypertension, and also helped them adapt to the cold environment19. These patterns have been suggested, and detected, for many diseases, especially those that have an onset in adult life20.
In addition to the role of natural selection, there has also been growing interest in understanding the contribution of archaic sequences of DNA to liability for disease21. We know that there has been more than one instance of admixture between early human populations along with Neanderthals and Denisovans22,23. This has resulted in the persistence of a number of introgressed sequences of archaic (Neanderthal and Denisovan) DNA that account for around 2–4% of the genome in modern (Homo sapiens) human populations. Studies have demonstrated the influence of such sequences on immune functioning and susceptibility to infections including COVID-1924. These sequences have also been found to be depleted in genes related to specific brain regions25 and influence functional connectivity in the brain as well26. Consequently, the impact of archaic introgression and admixtures on psychiatric disorders merits further exploration. South Asia has been inhabited by modern humans for the last several thousand years, and the population displays admixture with both extinct hominins, as well as significant migrations and bottlenecks in the recent past27,28. These admixture events may thus have a noteworthy influence on the susceptibility and prevalence of disease.
Hence, in this study, we investigated signatures of positive selection in unrelated individuals from families with multiple affected individuals with severe mental illness from southern India. We also used allele frequency differences between the cases and controls from the same population to prune out potential regions directly associated with caseness, and concentrated on regions with strong positive selection. Additionally, we specifically explored whether genes which showed evidence of positive selection had any evidence of Neanderthal introgression.
Results
We used whole exome sequence (WES) data of 80 unrelated individuals each of whom was diagnosed with psychiatric illness, as cases. These individuals were drawn from 80 separate and distinct families in which multiple members (at least 2 first-degree relatives in a nuclear family) were diagnosed to have a major psychiatric disorder [schizophrenia, BD, obsessive compulsive disorder (OCD), dementia and substance use disorders (SUD)]29 (A description of the sample is provided in the “Methods” section).
Since WES data is highly dispersed, we decided to use Population Branch Statistics (PBS) for our selection analysis30. PBS is based on allele frequency differentiation between populations using three populations. Unlike FST31, PBS is directional and gives us a clear idea as to which population is under selection for the given allele. A high PBS value corresponds to a highly deviated allele frequency of the target population compared to the reference population caused by positive selection.
Here, we used our data set consisting of 80 cases as the target population, the South Asian and African ancestry genomes from the gnomAD dataset as reference and outgroup populations respectively32. We also used WES data from 10 unrelated individuals from the same population as controls in the analysis to exclude PBS differences that may be attributable to case status rather than selection (A description of the same is provided in the “Methods” section).
Further, since the gnomAD dataset only reports South Asian ancestry Samples (SAS), which is a super population, we also tried to test if using a super population may bias our results when using the same as a reference population for the PBS analysis. We used 1000 genome 3rd phase data, which provides labels of South Asian subpopulations [such as Indian Telugus from the United Kingdom (ITU) and Gujarati Indian from Houston (GHU)], for this purpose. We found that PBS values coming from a subpopulation and superpopulation were highly correlated (R2 = 0.8534), especially SNPs with top values were common between both the results (Fig. S1). This reiterates our previous results33, and supports the conclusion that the use of the SAS super population in our study did not influence the findings of the PBS analysis.
Identification of SNPs and genes under influence of selection
We followed an approach that defined the SNPs that fell in the top 0.1% (99.9th percentile) of the PBS value distribution as the most likely candidates for selection. Further, to increase the confidence that the SNPs that fell under extreme PBS values were caused by selection rather than sampling of cases, we calculated the frequency difference of these SNPs between cases and controls. We then excluded all SNPs (Table S9) that fell within the top 0.1% (99.9th percentile) of the frequency difference distribution between cases and controls, also supported by the Fisher’s exact test P value and Odds ratio (OR) for frequency difference in cases and controls. This provided a list of candidate genes which were a plausible target of adaptive evolution specific to our test population. Further, we filtered genes with at least two SNPs with high PBS value to reduce the chance of false positives.
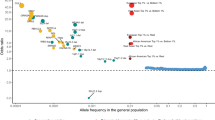
A total of 398 SNPs located in 190 genes were found to lie in the top 0.1% of the PBS value distribution. 115 genes had one SNP per gene (Table S1), while 75 genes had more than one SNP per gene (Table S2). A total of 10 SNPs from 10 genes were excluded due to case - control differences (Table S9). After this, we had 110 genes that had only one SNP per gene (Table S3) and 74 genes that had more than one SNP per gene (Table S3). For these 74 genes, we calculated an average PBS value per gene (Fig. 1; Table S3). We then used this list of 74 genes to look for any indications of an underlying shared biology.
Functions of putatively selected genes
Many of the 74 genes are involved with immunological and defense responses including activation and regulation of interferon-gamma, cytokine and immune system, and different signaling pathways. We manually curated these genes using the GWAS Catalogue (https://www.ebi.ac.uk/gwas/) and ENSEMBL (https://www.ensembl.org/Homo_sapiens/Info/Index) and found that several of the listed genes were reported for having potential roles in cancer, liver disease and diabetes.
We found 20 genes (MAML3, SNX8, WRN, ATXN3, PAK6, TXNDC2, MICA, PK1LD2, AHNAK2, PI4KA, C17orf97, FCER2, SNTG2, GGT1, FLG, IGFN1, PCDHA4, ANTXRL SULT1A1), in this list of 74 genes, that have been previously associated with elevated risk for schizophrenia, Parkinson disease, Alzheimer’s Disease and cognitive abilities or intelligence (Table S4).
Functional enrichment and pathway enrichment analysis
Furthermore, to evaluate whether genes with extremely high PBS values (the 99.9th percentile) were enriched in any functional category or metabolic pathways, we evaluated our list of 74 genes for the three Gene Ontology (GO) categories: biological processes (Table S5), cellular components (Table S6), and molecular function (Table S7). In addition, we analyzed the gene list for pathway over-representation using the IMPaLa tool (Table S8).
In GO analysis, we observed that several of selected genes of the target population were functionally enriched (P < 0.05) for different signaling and regulatory mechanisms related to immune system, and viral defense such as negative regulation of natural killer cell mediated cytotoxicity, interferon-gamma-mediated signaling pathway and antigen processing and presentation of exogenous peptide or polysaccharide antigen via MHC class II.
Using the IMPaLa pathway analysis tool, we again observed an over-representation for the enrichment of genes (below Q-value 0.05) involved in different immune related pathways including antigen processing and presentation, Graft-versus-host disease, Type I diabetes mellitus and autoimmune thyroid disease.
We repeated the functional enrichment analysis and pathway analysis with our list of 74 genes after exclusion of the HLA region (which is known to be extremely polymorphic). It was seen that while the GO analysis did not show any evidence of functional enrichment, the pathway analysis using IMPaLA suggested pathways involved with immune function (Table 1).
Archaic introgression in putatively selected genes
Further, we looked for the archaic introgression in these 74 genes that were chosen as candidates of selection. We applied Haplostrips34 to the exome data of our 80 unrelated case samples. However, we could not detect any definitive traces of archaic introgression in any of the genes in the gene sets except AHNAK2 gene (Fig. 2). Though we observed a few Neanderthal derived SNPs in the AHNAK2 gene region showing a pattern of unique haplotype sharing among the continental populations, none of these SNPs were found in high PBS value during selection scan.
Haplostrips plot of AHNAK2 gene: (A) Clustered and sorted by increasing distance with Neanderthals. A few Neanderthal derived SNPs show a pattern of unique haplotype sharing among the continental populations. However, none of these SNPs were found in high PBS value during selection scan and therefore have no significance in the adaptive role of AHNAK2 in this study. Population label abbreviations are as follows: NEAN Neanderthal, OIS Our Indian Samples, ITU Indian Telugus, CEU Central Europeans from Utah, CHB Chinese Han from Beijing, YRI Yoruba. (B) This plot visualizes the extent of closeness (based on SNP difference) between the haplotypes shared by continental populations and Neanderthal.
The fact that Neanderthal introgression sequence in the AHNAK2 gene region does not contain any of the variants identified as putative candidates of selection, indicates that the introgressed haplotype of the gene was probably not selected, thus, rules out its significance in the adaptive role of AHNAK2 in this study.
Discussion
Our results identify a total of 74 genes that show definitive evidence of selection, in individuals coming from families with multiple individuals with severe mental illness from southern India, based on the PBS analysis. Many of the identified genes have complex biology with numerous being linked to immune processes, cancer and neuropsychiatric illnesses.
There is some prior suggestion that many genes implicated in brain function, and disease, may be under selection35. This was supported by our findings where genes that we identified to be under selection have been previously implicated in a number of neuropsychiatric disease phenotypes. These include; SNX8 and PAK6, which were seen to harbour common variants that were associated with schizophrenia in GWAS10,36,37,38, SNTG2 and PKD1L2 that were associated with neurodevelopmental disorders such as autism39 and attention deficit hyperactivity disorder40, and ATXN3 and C17orf97 that have variants associated with neurodegenerative disorders like Amyotrophic Lateral Sclerosis41 and Parkinson's disease42. We also found genes related to disorders of neuronal migration and brain malformations such as polymicrogyria, like AHNAK2 and PI4KA43.
Aside from associations with disease phenotypes, a number of the genes we identified have been implicated in brain biology, including neurodevelopment, maintenance of neuronal and axonal integrity and apoptosis. The SARM1 protein is a member of MyD88/TIR-domain containing superfamily of proteins, which are involved in innate immune responses. Deficiency of the protein is seen to influence the apoptotic cascade of a variety of neural cells, including microglia; and cytokine expression in the brain44. The HYDIN protein, associated with cilia in the brain, is critical for development of ventricles and brain; and it interacts with other genes like FOXP2 which are well known to be related to neurodevelopmental disorders such as ASD and ADHD45. The neurodevelopmental disorders of autism and schizophrenia have also been hypothesised to be linked to UV exposure possibly via vitamin D levels46. Thus, it was interesting to identify the HORNERIN / HRNR gene which is very sensitive to UV light and is believed to also be involved in the species-specific differentiation of the outer layer of skin47. Further, genes identified by us such as MAML3, WRN and SULT1A1 also harbour variants linked to intelligence and cognitive abilities48,49,50. The contributions of these genes to neurodevelopment, intelligence and cognitive abilities suggests why they may be plausible candidates undergoing positive selection.
We also identified a number of genes linked to immune functions that were under selection in our sample. This was reflected in our functional enrichment and pathway enrichment analyses, where biological processes and pathways linked to the immune system were strongly implicated. This is expected, since immune function related genes are known to undergo significant selection pressure by virtue of factors such as the need to adapt to ecological diversity, and biological factors such as the threat posed by new and ever-changing infectious pathogens51,52.
The concurrent selection of genes that influence distinct, and apparently disparate biological processes, raises a number of extremely interesting questions about the interplay between the immune system and neuropsychiatric phenotypes. A well-known example is the role of the ApoE ε4 allele, which confers protection against viral illness on the one hand, but increases the risk of cognitive impairment later in life53. The role of the HLA region on Chromosome 6 has also been a consistently reported finding from GWAS and WGS of schizophrenia54. A study which investigated the role of balancing selection using exome data from modern and archaic humans also found an excess of SNPs across species in a gene set associated with the immune system, of which six were located within genes previously associated with schizophrenia55. Another study also suggests shared genetic pathways linking white blood cell indices and complex diseases such as schizophrenia, autoimmune, and coronary heart diseases56.
A number of genes that we identified in our study, such as the KIR family of genes have been implicated in a number of immune related functions, and also influence neurodevelopment and immune reactions in the brain. The KIR3DL1 has been linked to several aspects of natural killer cell responses, which in turn have been linked to susceptibility to multiple sclerosis, especially in African-Americans57. The KIR2DL4 immunoglobulin-like receptor has also been implicated in the development and maintenance of oligodendrocytes58; and also thought to have been positively selected for enabling uterine tolerance for embryonic implantation in humans59. Similarly, the KIR3DL2 have also been implicated in the interface between immunity genes and brain development, inflammation and responses to damage. They are the receptors for the major histocompatibility complex class I (MHC-I) like HLA-F, and protect neurons from astrocyte induced toxicity, as is seen in ALS60. The IGSF3 gene forms a complex with Tetraspanin (TSPAN7), which is involved in neurodevelopment (many mutations in this gene are linked to X-linked syndromes), and thus mediates a cross-talk between immune mechanisms and development61.
Although we observed archaic introgression in a single gene (AHNAK2) out of the identified set of putative selected genes, we did not find any evidence of archaic introgression under positive selection in any of the genes that were identified using the PBS analysis. These could be related to the fact that exome sequences have a restricted utility when it comes to finding introgressed regions, as the exonic regions are short and spaced, in the context of the whole genome. Additionally, from an evolutionary standpoint, selection tends to happen either upstream or downstream of the genes in areas such as transcription binding sites, rather than in the exonic regions which are well conserved62. Furthermore, exonic regions are generally under negative selection and therefore may not exhibit differences between modern and archaic hominins. Therefore, even if some highly differentiated exonic segment introgressed, it is most likely to have been weeded out by negative selection from the population, as Neanderthals have many more deleterious SNPs due to their low effective population size63. Thus, while it is not impossible to find an introgressed exonic region being selected for, these are rare in the literature64.
A few previous studies looking for selection signatures in south Asian populations using different methodologies have found evidence for positive selection in genes related to lipid metabolism and glucose uptake and have posited a link between the same and the predilection towards development of type 2 diabetes and obesity65 and height in Andaman Island populations66. Hence, our study also provides fresh insights into selection in a population from southern India.
In conclusion, our findings show that families with multiple members affected with severe mental illness can be used to detect signatures of selection. Immune related genes showed the greatest evidence of selection in these families. This underscores the contribution of immune mechanisms and infection susceptibility, to the genetic architecture of severe mental illness.
Methods
Study population
The study population consisted of 80 unrelated individuals (Females, N = 34); each of whom was diagnosed with psychiatric illness. The diagnoses were made by two trained psychiatrists based on DSM-IV TR criteria, and included BD (N = 26), schizophrenia (N = 25), dementia (N = 23), OCD (N = 3), SUD (N = 2) and Major Depressive Disorder (N = 1). These individuals were drawn from 80 separate and distinct families who were recruited as a part of the Accelerator Program for Discovery in Brain Disorders using Stem Cells (ADBS) study, which has been approved by the ethics committee of the National Institute of Mental Health and Neurosciences, Bengaluru, India. The study was carried out in accordance with the Declaration of Helsinki for research involving human participants. Written informed consent was obtained from all recruited individuals and their family members, wherever required.
Sequencing and quality control
As described in our previous study67, whole exome sequencing was carried out on the Illumina Hiseq NGS platform with libraries prepared using Illumina exome kits. Reads were aligned with the reference human genome hg19 using the Burrows-Wheeler algorithm tool (https://academic.oup.com/bioinformatics/article/25/14/1754/225615).
Variant calling
We used bcftools-1.968 to do the variant calling for all our samples from the bam files. First, we used bcftools mpileup to create genotype likelihoods. We then used a minimum base quality of 20 and a minimum mapping quality of 20 to accept it as a true variant. We also used an adjusted mapping quality of 50 to downgrade reads containing excessive mismatches (as recommended in bcftools for BWA). Additionally, we annotated the file using FORMAT/DP so we had depth information in the vcf file. The output was then piped to bcftools call. We used -m for multi allelic caller. We only used SNPs which were present in the gnomAD vcf file using -T command. An example of the code is presented here:
bcftools mpileup–ignore-RG -q 20 -Q 20 -C 50 -r <chr> -a FORMAT/DP-f <ucsc.hg19.fasta> <*.bam>| bcftools call-m-T <gnomead.vcf.gz> -O z-o <out.vcf.gz>.
A similar approach was also used for the 1000 genome phase 3 release data69, where we merged the 1000 genome vcf file using the bcftools merge command. We only kept our target population, Yoruba in Ibadan, Nigeria (YRI), Gujarati Indians in Houston, TX (GIH) and Indian Telugu in the UK (ITU) for further analysis.
Filtering
After variant calling, we first did a liftover of the vcf file from hg19 to GRCh37 using picard tools. We kept only SNPs (using –remove-indels flag). We removed any genotype where the coverage was less than 10 × using (–minDP 10) and removed any SNPs where we had more than 50% missing genotype data (using –max-missing 0.5). All these commands were done by using vcftools70. We kept only unrelated individuals from the affected multiplex families and disease free (control group) individuals for further analysis.
PBS calculation
We took the vcf file generated from the previous step and used vcftools to create a frequency file for both unrelated individuals and the control group. We used bcftools query to extract information about the frequencies of gnomAD vcf files. We only extracted AF_afr (alternate allele frequency of African-American/African ancestry individuals), AF_sas (alternate allele frequency of South Asian ancestry individuals), AN_afr [(total number of alleles in samples of African-American/African ancestry individuals) and AN_sas (total number of alleles in samples of South Asian ancestry individuals] from the info columns from gnomAD vcf files. These frequency data were used to calculate PBS (X, SAS, AFR) [where X is our data consisting of diseased unrelated individuals] using in house code with scikit-allele71. We then extracted the top SNPs by their PBS values and tried to find their impact on phenotype.
We also calculated PBS (X, ITU, YRI) and PBS (X, ITU + GIH, YRI) using scikit-allele to estimate the impact of using a super population instead of using a subpopulation. The R2 was calculated using scipy.stat.linregress function from scipy-1.5.3.
Frequency differentiation between cases and controls
As our target population consists of cases, some of the top PBS values can come from regions which might simply be associated with caseness due to sampling bias. To circumvent this problem, we also calculated the allele frequency differences between case and control data set (|Freq case − Freq control|). Subsequently, SNPs (> 99.9th percentile distribution of PBS) were only considered as potential targets of selection if they had allele frequency difference between cases and controls < 99.9th percentile and SNPs with allele frequency difference of > 99.9th percentile of the frequency difference distribution between case and controls were dropped. The rationale being that if the PBS value of a SNP is high due to selection instead of the caseness then the allele frequency difference between cases and controls should not vary as much.
We also implemented an alternative approach (Fisher’s exact test) to find out the top differentiated frequencies in case and control studies. We used plink-1.9.072 –assoc fisher and –allow-no-sex to calculate the p-value and the odds ratio (OR).
Identification of top candidate genes
We used python to select genes at the top 0.1% (> 99.9th percentile) of the overall PBS distribution and calculated the number of SNPs per gene. We chose 99.9 percentile as significant based on previously published analysis15. We thus identified 74 genes as the putative candidates for selection in the target population.
Analyses of functional enrichment
To perform the enrichment analysis, we used the set of genes obtained via the above method. For GO enrichment, we used the online tool in http://www.geneontology.org/page/go-enrichment-analysis; (GO Ontology database https://doi.org/10.5281/zenodo.5080993 Released 2021-07-02; last accessed August 9, 2021). We analyzed each of the four gene lists with the three GO categories (biological processes, cellular components, and molecular function) using FDR correction with a significance based on P value < 0.05 (ran on 9th August 2021).
The pathway over-representation analysis on the same gene sets was run using the IMPaLA online tool73 available at: http://impala.molgen.mpg.de (ran on 8th Dec, 2020) and we considered only pathways with a Q-value less than 0.05 to minimize the false positives because Q-value < 0.05 implies only 5% of results can be false positives.
Detection of archaic introgression in selected gene regions
We applied haplostrips tool to detect archaic introgression74 in putative positively selected genes. Haplostrips uses phased genetic data and visualizes polymorphisms of a particular genomic region by clustering and sorting haplotypes independently. The data was phased using shapeit75 with 1000 genome third phase reference69.
References
McGrath, J., Saha, S., Chant, D. & Welham, J. Schizophrenia: A concise overview of incidence, prevalence, and mortality. Epidemiol. Rev. 30, 67–76 (2008).
Rowland, T. A. & Marwaha, S. Epidemiology and risk factors for bipolar disorder. Ther. Adv. Psychopharmacol. 8, 251–269 (2018).
Wray, N. R. & Visscher, P. M. Narrowing the boundaries of the genetic architecture of schizophrenia. Schizophr. Bull. 36, 14–23 (2010).
Liu, C., Everall, I., Pantelis, C. & Bousman, C. Interrogating the evolutionary paradox of schizophrenia: A novel framework and evidence supporting recent negative selection of schizophrenia risk alleles. Front. Genet. 10, 389 (2019).
Pattabiraman, K., Muchnik, S. K. & Sestan, N. The evolution of the human brain and disease susceptibility. Curr. Opin. Genet. Dev. 65, 91–97 (2020).
Guo, J., Yang, J. & Visscher, P. M. Leveraging GWAS for complex traits to detect signatures of natural selection in humans. Curr. Opin. Genet. Dev. 53, 9–14 (2018).
Polimanti, R. & Gelernter, J. Widespread signatures of positive selection in common risk alleles associated to autism spectrum disorder. PLoS Genet. 13, e1006618 (2017).
Srinivasan, S. et al. Genetic markers of human evolution are enriched in schizophrenia. Biol. Psychiatry 80, 284–292 (2016).
Xiang, B. et al. The role of genes affected by human evolution marker GNA13 in schizophrenia. Prog. Neuropsychopharmacol. Biol. Psychiatry 98, 109764 (2020).
Pardiñas, A. F. et al. Common schizophrenia alleles are enriched in mutation-intolerant genes and in regions under strong background selection. Nat. Genet. 50, 381–389 (2018).
Yao, Y. et al. No evidence for widespread positive selection signatures in common risk alleles associated with schizophrenia. Schizophr. Bull. 46, 603–611 (2020).
Peng, Y. et al. Down-regulation of EPAS1 transcription and genetic adaptation of tibetans to high-altitude hypoxia. Mol. Biol. Evol. 34, 818–830 (2017).
Zhou, S. et al. Genetic architecture and adaptations of Nunavik Inuit. Proc. Natl. Acad. Sci. USA 116, 16012–16017 (2019).
Hallmark, B. et al. Genomic evidence of local adaptation to climate and diet in indigenous Siberians. Mol. Biol. Evol. 36, 315–327 (2019).
Ávila-Arcos, M. C. et al. Population history and gene divergence in native Mexicans inferred from 76 human exomes. Mol. Biol. Evol. 37, 994–1006 (2020).
Reynolds, A. W. et al. Comparing signals of natural selection between three Indigenous North American populations. PNAS 116, 9312–9317 (2019).
Nishino, J. et al. Empirical Bayes estimation of semi-parametric hierarchical mixture models for unbiased characterization of polygenic disease architectures. Front. Genet. 9, 115 (2018).
Hess, J. L. et al. A polygenic resilience score moderates the genetic risk for schizophrenia. Mol. Psychiatry 26, 800–815 (2021).
Li, M. et al. Recent positive selection drives the expansion of a schizophrenia risk nonsynonymous variant at SLC39A8 in Europeans. Schizophr. Bull. 42, 178–190 (2016).
Benton, M. L. et al. The influence of evolutionary history on human health and disease. Nat. Rev. Genet. 22, 269–283 (2021).
Dannemann, M. & Racimo, F. Something old, something borrowed: Admixture and adaptation in human evolution. Curr. Opin. Genet. Dev. 53, 1–8 (2018).
Browning, S. R., Browning, B. L., Zhou, Y., Tucci, S. & Akey, J. M. Analysis of human sequence data reveals two pulses of archaic denisovan admixture. Cell 173, 53-61.e9 (2018).
Jacobs, G. S. et al. Multiple deeply divergent Denisovan ancestries in papuans. Cell 177, 1010-1021.e32 (2019).
Zeberg, H. & Pääbo, S. The major genetic risk factor for severe COVID-19 is inherited from Neanderthals. Nature 587, 610–612 (2020).
Dolgova, O. & Lao, O. Evolutionary and medical consequences of archaic introgression into modern human genomes. Genes 9, 1–10 (2018).
Gregory, M. D. et al. Neanderthal-derived genetic variation is associated with functional connectivity in the brains of living humans. Brain Connect 11, 38–44 (2021).
Narasimhan, V. M. et al. The formation of human populations in South and Central Asia. Science 365, 10 (2019).
Pathak, A. K. et al. The genetic ancestry of modern indus valley populations from Northwest India. Am. J. Hum. Genet. 103, 918–929 (2018).
Viswanath, B. et al. Discovery biology of neuropsychiatric syndromes (DBNS): A center for integrating clinical medicine and basic science. BMC Psychiatry 18, 106 (2018).
Yi, X. et al. Sequencing of fifty human exomes reveals adaptation to high altitude. Science 329, 75–78 (2010).
Wright, S. The genetical structure of populations. Ann. Eugen. 15, 323–354 (1951).
Karczewski, K. J. et al. The mutational constraint spectrum quantified from variation in 141,456 humans. Nature 581, 434–443 (2020).
Juyal, G. et al. Population and genomic lessons from genetic analysis of two Indian populations. Hum. Genet. 133, 1273–1287 (2014).
Marnetto, D. & Huerta-Sánchez, E. Haplostrips: Revealing population structure through haplotype visualization. Methods Ecol. Evol. 8, 1389–1392 (2017).
Huang, Y. et al. Recent adaptive events in human brain revealed by meta-analysis of positively selected genes. PLoS ONE 8, e61280 (2013).
Bergen, S. E. et al. Genome-wide association study in a Swedish population yields support for greater CNV and MHC involvement in schizophrenia compared with bipolar disorder. Mol. Psychiatry 17, 880–886 (2012).
Ripke, S. et al. Genome-wide association analysis identifies 13 new risk loci for schizophrenia. Nat. Genet. 45, 1150–1159 (2013).
Ripke, S. et al. Biological insights from 108 schizophrenia-associated genetic loci. Nature 511, 421–427 (2014).
Bacchelli, E. et al. An integrated analysis of rare CNV and exome variation in Autism Spectrum Disorder using the Infinium PsychArray. Sci. Rep. 10, 3198 (2020).
Anney, R. J. L. et al. Conduct disorder and ADHD: Evaluation of conduct problems as a categorical and quantitative trait in the international multicentre ADHD genetics study. Am. J. Med. Genet. B Neuropsychiatr. Genet. 147B, 1369–1378 (2008).
Nicolas, A. et al. Genome-wide analyses identify KIF5A as a Novel ALS gene. Neuron 97, 1268-1283.e6 (2018).
Simón-Sánchez, J. et al. Genome-wide association study reveals genetic risk underlying Parkinson’s disease. Nat. Genet. 41, 1308–1312 (2009).
Pagnamenta, A. T. et al. Germline recessive mutations in PI4KA are associated with perisylvian polymicrogyria, cerebellar hypoplasia and arthrogryposis. Hum. Mol. Genet. 24, 3732–3741 (2015).
Figley, M. D. & DiAntonio, A. The SARM1 axon degeneration pathway: Control of the NAD+ metabolome regulates axon survival in health and disease. Curr. Opin. Neurobiol. 63, 59–66 (2020).
Fang, L. et al. Comparative analyses of sperm DNA methylomes among human, mouse and cattle provide insights into epigenomic evolution and complex traits. Epigenetics 14, 260–276 (2019).
Hart, P. H., Norval, M., Byrne, S. N. & Rhodes, L. E. Exposure to ultraviolet radiation in the modulation of human diseases. Annu. Rev. Pathol. 14, 55–81 (2019).
Shen, Y., Ha, W., Zeng, W., Queen, D. & Liu, L. Exome sequencing identifies novel mutation signatures of UV radiation and trichostatin A in primary human keratinocytes. Sci. Rep. 10, 4943 (2020).
Hill, W. D. et al. A combined analysis of genetically correlated traits identifies 187 loci and a role for neurogenesis and myelination in intelligence. Mol. Psychiatry 24, 169–181 (2019).
Lam, M. et al. pleiotropic meta-analysis of cognition, education, and schizophrenia differentiates roles of early neurodevelopmental and adult synaptic pathways. Am. J. Hum. Genet. 105, 334–350 (2019).
Rietveld, C. A. et al. Common genetic variants associated with cognitive performance identified using the proxy-phenotype method. PNAS 111, 13790–13794 (2014).
Shultz, A. J. & Sackton, T. B. Immune genes are hotspots of shared positive selection across birds and mammals. Elife 8, e41815 (2019).
Quintana-Murci, L. Human immunology through the lens of evolutionary genetics. Cell 177, 184–199 (2019).
Zhao, C. et al. APOE ϵ4 modifies the relationship between infectious burden and poor cognition. Neurol. Genet. 6, e462 (2020).
Birnbaum, R. & Weinberger, D. R. A genetics perspective on the role of the (neuro)immune system in schizophrenia. Schizophr. Res. 217, 105–113 (2020).
Viscardi, L. H. et al. Searching for ancient balanced polymorphisms shared between Neanderthals and Modern Humans. Genet. Mol. Biol. 41, 67–81 (2018).
Astle, W. J. et al. The allelic landscape of human blood cell trait variation and links to common complex disease. Cell 167, 1415-1429.e19 (2016).
Hollenbach, J. A., Pando, M. J., Caillier, S. J., Gourraud, P.-A. & Oksenberg, J. R. The killer immunoglobulin-like receptor KIR3DL1 in combination with HLA-Bw4 is protective against multiple sclerosis in African Americans. Genes Immun. 17, 199–202 (2016).
Banerjee, P. P. et al. KIR2DL4-HLAG interaction at human NK cell-oligodendrocyte interfaces regulates IFN-γ-mediated effects. Mol. Immunol. 115, 39–55 (2019).
Vallender, E. J. Chapter 1: Genetics of human brain evolution. in Progress in Brain Research (ed. Hofman, M. A.) vol. 250, 3–39 (Elsevier, 2019).
Song, S. et al. Major histocompatibility complex class I molecules protect motor neurons from astrocyte-induced toxicity in amyotrophic lateral sclerosis. Nat. Med. 22, 397–403 (2016).
Perot, B. P. & Ménager, M. M. Tetraspanin 7 and its closest paralog tetraspanin 6: Membrane organizers with key functions in brain development, viral infection, innate immunity, diabetes and cancer. Med. Microbiol. Immunol. 209, 427–436 (2020).
Dannemann, M., Prüfer, K. & Kelso, J. Functional implications of Neandertal introgression in modern humans. Genome Biol. 18, 61 (2017).
Juric, I., Aeschbacher, S. & Coop, G. The strength of selection against Neanderthal introgression. PLOS Genet. 12, e1006340 (2016).
Racimo, F., Sankararaman, S., Nielsen, R. & Huerta-Sánchez, E. Evidence for archaic adaptive introgression in humans. Nat. Rev. Genet. 16, 359–371 (2015).
Metspalu, M. et al. Shared and unique components of human population structure and genome-wide signals of positive selection in South Asia. Am. J. Hum. Genet. 89, 731–744 (2011).
Mondal, M. et al. Genomic analysis of Andamanese provides insights into ancient human migration into Asia and adaptation. Nat. Genet. 48, 1066–1070 (2016).
Ganesh, S. et al. Exome sequencing in families with severe mental illness identifies novel and rare variants in genes implicated in Mendelian neuropsychiatric syndromes. Psychiatry Clin. Neurosci. 73, 11–19 (2019).
Li, H. A statistical framework for SNP calling, mutation discovery, association mapping and population genetical parameter estimation from sequencing data. Bioinformatics 27, 2987–2993 (2011).
Auton, A. et al. A global reference for human genetic variation. Nature 526, 68–74 (2015).
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158 (2011).
Miles, A. & Harding, N. cggh/scikit-allel: v1. 1.8 (Version v1. 1.8). Zenodo. (2017).
Purcell, S. et al. PLINK: A tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Kamburov, A., Cavill, R., Ebbels, T. M. D., Herwig, R. & Keun, H. C. Integrated pathway-level analysis of transcriptomics and metabolomics data with IMPaLA. Bioinformatics 27, 2917–2918 (2011).
Green, R. E. et al. A draft sequence of the Neandertal genome. Science 328, 710–722 (2010).
Delaneau, O. & Marchini, J. Integrating sequence and array data to create an improved 1000 Genomes Project haplotype reference panel. Nat. Commun. 5, 3934 (2014).
Acknowledgements
This research is funded by the Accelerator program for discovery in brain disorders using stem cells (ADBS) (jointly funded by the Department of Biotechnology, Government of India, and the Pratiksha trust; Grant BT/PR17316/MED/31/326/2015). AKP was supported by the European Union through the European Regional Development Fund (Project No. 2014-2020.4.01.15-0012). MM was supported by the European Union through Horizon 2020 Research and Innovation Programme under Grant No. 810645 and through the European Regional Development Fund Project No. MOBEC008. The authors are grateful to all the patients, and their family members who participated in the study.
Author information
Authors and Affiliations
Consortia
Contributions
Study concept and design—S.J., M.P. and M.M., Acquisition of data—R.K.N., B.V., S.J., Bio-informatics and Data Analysis—A.K.P., A.V., M.M., Interpretation of data—J.M., A.K.P., S.J., M.P., M.M., Manuscript drafting—J.M., Critical revision of manuscript—All.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Mahadevan, J., Pathak, A.K., Vemula, A. et al. Analysis of whole exome sequencing in severe mental illness hints at selection of brain development and immune related genes. Sci Rep 11, 21088 (2021). https://doi.org/10.1038/s41598-021-00123-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-00123-x
This article is cited by
-
Significance of an altered lncRNA landscape in schizophrenia and cognition: clues from a case–control association study
European Archives of Psychiatry and Clinical Neuroscience (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.