Abstract
Inferences from genetic association studies rely largely on the definition and description of the underlying populations that highlight their genetic similarities and differences. The clustering of human populations into subgroups (population structure) can significantly confound disease associations. This study investigated the fine-scale genetic structure within Cameroon that may underlie disparities observed with Cameroonian ethnicities in malaria genome-wide association studies in sub-Saharan Africa. Genotype data of 1073 individuals from three regions and three ethnic groups in Cameroon were analyzed using measures of genetic proximity to ascertain fine-scale genetic structure. Model-based clustering revealed distinct ancestral proportions among the Bantu, Semi-Bantu and Foulbe ethnic groups, while haplotype-based coancestry estimation revealed possible longstanding and ongoing sympatric differentiation among individuals of the Foulbe ethnic group, and their Bantu and Semi-Bantu counterparts. A genome scan found strong selection signatures in the HLA gene region, confirming longstanding knowledge of natural selection on this genomic region in African populations following immense disease pressure. Signatures of selection were also observed in the HBB gene cluster, a genomic region known to be under strong balancing selection in sub-Saharan Africa due to its co-evolution with malaria. This study further supports the role of evolution in shaping genomes of Cameroonian populations and reveals fine-scale hierarchical structure among and within Cameroonian ethnicities that may impact genetic association studies in the country.
Similar content being viewed by others
Introduction
Advances in the genome-wide analysis of human genomic variation in disease states have led to the discovery of thousands of disease-associated loci1. Although genome-wide association studies (GWASs) have been successful in uncovering the contribution of single nucleotide polymorphisms (SNPs) to common diseases in populations with European and Asian ancestries, studies on African populations have been less successful. Population structure/substructure, characterized by extensive genetic variation and low linkage disequilibrium (LD) among and within African populations is a major cause of the poor performance of GWAS and other statistical genetics tools in Africa2,3.
Population structure analysis is crucial to the design, analysis, and interpretation of genetic association studies. While human genetic structure is well understood at the continental scale4,5,6,7, the fine-scale genetic structure of specific African populations at fine geographic ranges remains largely understudied. Previous genetic association analyses in Africa have relied on principal component analysis (PCA)8, with the top principal components (PCs), included as covariates in the association analyses to control for confounding by population structure9,10,11,12,13,14,15. More recently, approaches like mixed models (MM)16,17,18 and Bayesian statistics19 have proven particularly effective in accounting for genetic structure among and within populations in GWASs. However, given the extensive genetic heterogeneity in Africa that usually accompanies ethno-linguistic, cultural, and religious disparities, scars of genetic substructure remain ubiquitous in GWAS on the continent leading to the high false discovery rates (FDRs) and deficiency of significant SNPs from such studies10,15.
The human genetic diversity in Africa remains complex and multi-factorial, accrued from ancient and recent migration events. The outcome of this has been significant genetic admixture despite cultural, ethno-linguistic, and eco-geographic barriers to gene flow7,20. The genomes of African populations have also been shaped by evolutionary and selection pressures from infectious diseases and the environment. The relatively high frequency of the Sickle cell trait and the glucose-6-phosphate dehydrogenase (G6PD) deficiency in sub-Saharan Africa (sSA) perpetrated by malaria21,22 are classic examples of such disease pressure. Perhaps, it is the clustering of African populations into thousands of demes of local ethnicities20 that mirrors the extent of the genetic diversity within the continent.
In Cameroon for instance, there are over 250 tribes distributed within 3 broad ethnic groups; Bantu (BA), Semi-Bantu (SB), and Sudanese (which includes the Foulbe). A recent population genetic study revealed that chunks of the genome of individuals of the BA and SB ethnicities are shared with their African counterparts20. While this is expected, the ancestral relationship and extent of genetic differentiation between individuals of different ethnicities in Cameroon have not been explored. Considered the World’s most culturally diverse nation, with a complex colonial history23,24, Cameroon’s populations may have been subject to both genetic admixture and numerous barriers to gene flow, leading to differences in allele frequency and haplotype structure between ethnic populations. A recent severe malaria GWAS in Africa highlighted some heterogeneity in the effect and genotype frequency of some key malaria-associated loci in Cameroonian individuals relative to their sSA counterparts15. Hence, detailed analysis of the fine-scale structure of the ethnic groups may shed some light on specificities of the sub-populations.
This study aimed at characterizing the genetic structure of the Bantu, Semi-Bantu, and Foulbe (FO) ethnic groups from Cameroon. SNP data from 1073 individuals living in the South West, Littoral, and Centre regions of Cameroon were analyzed. The Wright’s F-statistic (FST), model-based clustering, and PCA were used to determine the relationship between structure and ethnicity while information derived from patterns of haplotype sharing among individuals (coancestry) was used to depict fine-scale structure and ancestral admixture. Additionally, the genetic architecture of individuals in the study population was compared to those from the 1000 Genomes Phase 3 populations and genome scan was performed to map genomic regions with significantly differentiated alleles and extended haplotype homozygosity that may be due to selective forces.
Results
Genetic distance (FST) and PCA show correlation with geography
A total of 1073 individuals from Cameroon (BA = 492, FO = 25, SB = 556) and 2504 individuals from the 1,000 Genomes reference panel phase 3 (1KGP3) populations were studied25,26. FST, PCA and other measures of genetic proximity were computed from 763,806 autosomal SNPs ascertained to be polymorphic in the Mende population from Sierra Leon (MSL) (see “Discussions” and “Methods” section) and shared (polymorphic) across all the populations. When Cameroonian populations were analyzed separately, 57,374 of these SNPs, polymorphic across all the ethnic groups, were used. We note that several estimators of FST have been developed and a majority of them are not adequately robust against varying population sizes. The Hudson estimator implemented in smartpca27 is, to the best of our knowledge, the only documented estimator that is immune to sample size differences and was (smartpca version 16000) thus employed in this analysis. Estimates for within- and among-continent population comparisons similar to those previously reported were observed (supplementary Data S1). Here, Cameroonian populations generally clustered with other African populations (Fig. 1a). The SB appeared genetically closer to the Yoruba of Nigeria (YRI) (FST SBvsYRI = 0.002) than did the BA (FST BAvsYRI = 0.003) contrary to previous estimates20. We further found that the FO ethnicity is relatively less genetically related to the YRI (FST = 0.004) compared to Cameroonian SB and BA populations. In addition, contrary to prior estimates that the Cameroonian BA population was genetically closer to the Luhya population of Kenya (LWK), we observed no difference in FST among the LWK population and all Cameroonian populations herein studied (all pairwise FST = 0.005)20. Interestingly, the FO, like the LWK population appeared to be more genetically close to populations of European and Asian ancestries as compared to the BA and SB, supporting possible influence of Eurasian migration back into Africa28. PCA generally showed positive concordance with FST results (Fig. 1b).
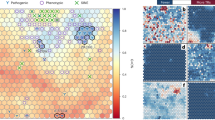
Pairwise Fst and PCA analysis of Cameroonian and world populations. (a) Clustered heatmap showing genetic distance by pairwise population FST (Hudson) estimation. AFR = African, EAS = East Asian, EUR = European, AMR = American and SAS = South Asian ancestry. The red color denotes closely related population, hence low FST while the decrease in redness to yellow represents increasing genetic distance (high FST). Five clusters are apparently corresponding to the five continental proxy ancestry (distinguished broadly by five colors) in the 1000 Genomes project. (b) PCA of Cameroonian populations with the 1000 Genomes populations. Cameroonian populations clustered within African populations. All plots were produced using R 3.6.129.
In an Africa-only analysis, Cameroonian populations clustered between Nigerian and Kenyan populations based on principal components 1 (PC1) and 2 (PC2) (Fig. 2a). There was a general West–East cline (adjusted axes) consistent with the concept of isolation by distance (geographic location). Cameroonian populations clustered West-Centrally reflecting their location on the continent. Initial PCA with smartpca’s default 5 outlier removal iterations (numoutlieriter) resulted in ancestry outliers (Supplementary Fig. S1) which were eventually removed by increasing the numoutlieriter to 10. Cameroonian ethnicities could already be distinguished at this point, although a high degree of spread remained apparent. In addition, FO individuals differentiated from the BA and SB, with SB clustering closer to Nigerian populations than the other ethnicities.
PCA of Cameroonian populations and other Africa populations. (a) Cameroonian populations with other African populations produce a clustering pattern correlated with geography where West African populations (GWD, MSL, ESN, YRI) clustered to the West, the lone east African population (LWK) clustered to the East, while Cameroonian populations clustered West-Centrally. (b) PCA for Cameroonian ethnicities only. PC1 and PC2 separate the three ethnicities, (c) PC1 and PC3 separate the Bantu and Semi-Bantu. All plots were produced using R 3.6.129.
Further PCA of only Cameroonian ethnicities revealed three clusters (Supplementary Fig. S2). To increase the resolution of the clusters, we ran PCA with SNPs from smartpca’s pairwise FST ≥ 0.003 classified here as “ancestry informative markers” (AIMs). In principle, AIMs are loci with substantial allele frequency difference across populations30. They can be used to assign individuals into clusters based on biological (ancestral) or geographical (continental) boundaries31. Although pre-calculated lists of AIMs exist, they mostly apply to cross-continental populations32. Furthermore, there are several measures of marker information content as listed in Table 1 of Rosenberg et al.31. We therefore used the FST measure in this study to generate AIMs by selecting all SNPs with FST greater than or equal to the highest FST (genetic distance) between Cameroonian ethnic groups. FST between the BA and SB was 0.001 whereas FST between the FO and both BA and SB was 0.003. Hence, AIMs were considered as SNPs with pairwise population FST (Hudson) ≥ 0.003. This strategy saw an increased resolution of the clusters, clearly separating the FO from the BA and SB based on PC2 (Fig. 2b). Furthermore, a substructure within the FO population was resolved, showing two separate clusters. At fine scale, using AIMs, a substructure within the BA population based on PC1 against PC3 was resolved (Fig. 2c). We note that increasing the FST at which AIMs were selected only further separated the FO from the BA and SB without achieving further resolution of the BA and SB clusters.
Distinct ancestral proportions and fine structure among Cameroonian ethnicities
Using a total of 81,415 high quality independent AIMs (MAF > 0.05, LD < 0.2), we computed ancestral proportions (Q estimates) for our three ethnic groups with five k-parameters using model-based clustering implemented in Admixture33. At K = 2, model-based clustering differentiated the three ethnicities albeit with low resolution. However, at K = 3 where the lowest cross-validation error was recorded (Fig. 3a), the three ethnicities were clearly differentiated (Fig. 3b). Ancestral proportions (Q) estimated [green predominant in the BA (~ 45%), red in the FO (~ 75%) and blue in the SB (~ 45%)] (Fig. 3c) show that the different ethnic groups differ by allele frequencies (Supplementary material) and/or haplotype structure (see below). When we applied Structure v2.3.434 in the admixture mode using 3 pre-defined clusters on 50 BA, 50 SB, and 25 FO individuals setting 20,000 burnin and 100,000 main iterations, the ancestral proportions were resolved even better (FO = 76.6%, BA = 73.3%, and SB = 68.4%) (Supplementary Fig. S4). Here, the FO appeared to harbor a large chunk of ancestry from a non-Cameroonian source.
Model-based clustering and Coancestry estimation. (a) Model-based clustering cross-validation (CV) error. Lowest CV error recorded at k = 3 indicating three clusters. (b) and (c) show ancestral proportions Q, for each ethnicity colored using the RGB color scheme. (d) Coancestry estimation by FineStructure. All plots were produced using R 3.6.129.
While single marker approaches that measure genetic distance usually require markers in approximate linkage equilibrium (unlinked), haplotype-based approaches have the potential to exploit the linkage information in genotype datasets to estimate genetic distances with finer resolutions. Therefore, we used a haplotype-based approach of coancestry estimation by “chromosome painting” implemented in fineStructure35 for fine scale resolution of Cameroonian ethnicities. In this analysis, we thinned the samples using Plink1.9′s—thin-indiv-count (which removes samples at random until only the required number is left) to obtain 50 BA and 50 SB individuals together with the initial 25 FO individuals to make a final analysis set of 125 individuals. We observed isolation of a subgroup of FO individuals from the BA and SB by PC1 (Fig. 3d). PC2 separated the Bantu and Semi-Bantu, with significant numbers of BA individuals clustering with SB. Generally, all the ethnicities showed a cline into a central cluster that appeared to be a set of admixed individuals, consistent with their longstanding cohabitation while their separation suggests some evidence of ancient genetic isolation and/or gene flow from other populations. The three extreme clusters may represent individuals with the basal ancestry for each respective ethnicity.
Selective sweeps target genes associated with response to disease
Genomic regions under positive selective sweeps are usually characterized by extended haplotype homozygosity (EHH) and low genetic diversity. A genome-wide scan for such regions harboring signatures of selection by the standardized integrated haplotype score (iHS) which measures the EHH identified strong signatures on multiple chromosomes. This included missense and regulatory region variants in genes overwhelmingly associated with response to infections. The scan identified a total of 133 SNPs within 57 overlapped genes and 173 overlapped transcripts across chromosomes 1 to 12, 14, 16, 17, 19, and 20 with significant signatures of selection at iHS threshold of ± 4 (Fig. 4a). Chromosome 6 and 11 harbored the highest proportion of variants involved in selection. Table 1 shows the variants with strong signatures of selection that occurred in coding regions. The entire list of 133 selected variants can be found as Supplementary Data S2 online along with the corresponding variant effect predictor (VEP)36 and BioMart-annotated gene list. Although the strongest signature occurred on chromosome 1 around the REG4 gene (iHS = − 7.23, p-value = 4.67 × 10−13), the most consistent signatures were recorded on chromosome 6 spanning the HLA region which has been reported in several previous studies of selection37,38,39,40. The SNP rs10947368, a missense variant on HLA-DOA emerged with the strongest signal within the HLA region (iHS = − 6.42, p-value = 1.38 × 10−10) (Table 1). In addition, suggestive signatures of selection were recorded in the hemoglobin-beta (HBB) gene cluster of chromosome 11, a region with longstanding knowledge of balancing selection under the influence of malaria41,42. However, the strongest signal on chromosome 11 was a relatively uncommon missense variant (rs7943508) in the APLNR gene, implicated in hypertension and some cancers43,44. Generally, selection signatures were not isolated, but occurred in clusters of consecutive SNPs positions, consistent with “genetic hitchhiking” around a positively selected variant, typical of selective sweeps. Our genome-wide significance threshold was estimated at 6.033 × 10−08 (see “Methods” section). The iHS values obtained were generally normally distributed as expected under neutral evolution with a slight deviation from the expected distribution reflecting the regions showing evidence of selection (Fig. 4b).
iHS and corresponding –log10(p-values) Manhattan plots. (a) iHS plot for the autosomes. Negative values signify selection on derived alleles while positive values are associated with selection on ancestral alleles. (b) Distribution of iHS values as observed in the populations (blue) and as expected under neutral evolution (red). Lower plot represents quantile–quantile (Q–Q) plot of iHS p-values. The plot shows that the test statistics are not inflated. The plots were generated using the R package qqman45.
Allele frequency differences due to selection pressures among Cameroonian ethnicities
Allele frequency differences among populations can be caused by several factors including random genetic drift following isolation, whether by distance, or more commonly in Africa, by cultural and religious disparities (sympatric differentiation). Also, allele frequency differences may arise due to changes in population dynamics like rapid population growths or contractions, and selection pressures among others. Therefore, to isolate only selection pressure effect on allele frequency differences among Cameroonian ethnicities, we computed the extended Lewontin–Krakauer FST outlier statistic (FLK) which accounts for unequal effective population sizes (Ne) and hierarchical population structure46. We used the MSL population as an out-group as earlier mentioned. Several genomic regions with subtle allele frequency differences between the ethnicities were observed although none of these regions remained significant after correction for multiple testing by the Benjamin–Hockerbeg method47,48. However, evidence of positive selection remained apparent in the HLA region on chromosome 6. Specifically, positions on chromosomes 2, 6, 8, 10, 17, 18, and 22 were subtly differentiated among the ethnicities (Supplementary Fig.S5 & Supplementary Table S1). We note that the FLK test is not robust to low levels of heterozygosity and it was shown to perform poorly (with a high false positive rate) at SNPs with ancestral allele frequency (AAF) < 0.2 and AAF > 0.8 (that is less common SNPs and SNPs nearing fixation)46. Therefore our analysis was based on SNPs within the AAF range of 0.2–0.8 (see “Methods” section). A cluster-stratified analysis of the derived allele frequencies revealed a great heterogeneity in the frequency spectra of the three ethnic groups at selected loci (Supplementary Fig. S6).
Genes involved in food/drug metabolism targeted by selection
Given that genomic regions under strong positive selection usually pull along nearby SNPs (genetic hitchhiking), we screened for selection using clusters of loci (haplotypes) by the haplotype variant of the FLK test (hapFLK). This resulted in a strong signature on chromosome 6 as was recorded by iHS, while several other regions on multiple chromosomes showed suggestive signals (Fig. 5). Of note were signatures on chromosomes 10, 16, 17, and 22 occurring in genes associated with food/drug metabolism. We observed signals on chromosome 10 associated with missense variants on the ACSM6 gene associated with acetyl coenzyme-A production (p = 1.62 × 10−06), and on the CYP2C8 gene, a cytochrome P450 superfamily enzyme member associated with drug metabolism (p = 6.09 × 10−06). Multiple missense variant signals were also observed in the ABCC11/12 gene on chromosome 16 (p = 5.39 × 10−07), an ATP binding cassette subfamily member involved in multi-drug resistance. In addition, signals were observed on the MTTP gene on chromosome 4 (p = 3.73 × 10−06) involved in triglyceride transfer and lipoprotein assembly, the TMEM199 gene on chromosome 17 (p = 5.35 × 10−06) whose deficiency is associated with abnormal glycosylation49, and the TCN2 gene on chromosome 22 (p = 6.76 × 10−06) involved in the absorption of vitamin B12 (cobalamin). Our threshold for genome-wide significance was estimated at 6.02 × 10−08 (see “Methods” section).
Manahattan plot of hapFLK p-values. Genome-wide significance threshold (red line), suggestive threshold (blue line). The plot was generated using qqman45.
Extant Cameroonian ethnic populations may be under differential selective pressure
To further investigate which genomic positions are differentially selected among the ethnic groups, we performed a cross-population selection scan using the Rsb statistic implemented in the REHHv3.0.1 R package50,51. We found chromosome 6 to be strongly selected in the BA and SB and only subtly selected in the FO. This could be due to the low sample size of the FO population. The BA population showed additional signatures on chromosome 6 involving the missense variant rs9276 on the HLA-DPB1 consistent with previous reports of selection in this gene in Bantu-speaking African populations (BSP)52 and the variant rs1419638 on the OR5V1 gene, as well as on chromosome 7 unique to the BA ethnic group. Likewise, the SB showed specific signatures on chromosomes 16 and 20, while strong signatures specific to the FO population were recorded on chromosomes 1, 7, 9, 10, 16, and 19 (Supplementary Fig.S8a-d). Again, these selection signatures primarily implicated genes involved in disease response.
Discussion
An in-depth understanding of population structure is critical in genetic studies, particularly in highly diverse and admixed African and American populations because of its potential to impact genetic association studies that usually have small effect sizes and rare variant analyses53,54. Characterizing the genetic architecture of specific populations is thus a “higher level of priority” under the “Basic Genomics and Genomic Technologies” focus area of the NHGRI (https://www.genome.gov/about-nhgri/strategic-plan). This study sought to ascertain the fine-scale genetic structure in Cameroon by analyzing the genotype of 1073 individuals from the South West, Littoral, and Centre regions.
The subtle differences in pairwise FST observed among Cameroonian ethnic populations sampled in this study indicates substantial mixing and common recent coancestries. However, the fact that the estimates were non-zero values supports the existence of distinct ancestral proportions. The FO (belonging to the Sudanese ethnic division) appear to have North African roots related to Fulani migration from north eastern Nigeria and Chad. The BA, on the other hand, are thought to have been the earliest inhabitants of Cameroon, with traces of their ancient civilization still prominent in the pigmies of the South and East. In fact, some studies have associated the spread of the BA ancestral proportions found in Central, South and East Africa to a Bantu expansion that originated somewhere around South Western Cameroon20,55,56,57. The SB mainly inhabit the Western highlands and grass fields of the West and North West of Cameroon. Together with the BA of the South West, the SB of the North West also appear to have endured a four-decade complex cohabitation with Eastern Nigerian populations during colonial era23. Therefore, the ancient interactions, and interactions of the recent past of Cameroonian populations with other populations may have paved the way for substantial genetic admixture and drift.
The close similarity in ancestral proportions among the BA and SB is expected (Fig. 3a). The dissimilarities may be attributable to many factors including varying degrees of contact with different external populations with subsequent genetic drift as could be have been possible during their pre-colonial and colonial era as well as following different selective pressures. According to historical records, Portuguese slave-trading on the South-Western coast of Cameroon began as far back as the fifteenth century and was followed by British commercial hegemony in palm oil and ivory in the nineteenth century, and an eventual German sovereignty in 1884 that was succeeded by French and British rule after World War I58. Following these events, parts of Nigeria, Chad, Central African Republic, Congo, Gabon, and Equatorial Guinea today were parts of Cameroon under the German and subsequently French and British rule23. These were therefore good recipes for genetic admixture and drift.
The considerable dissimilarity in the ancestral proportion of the FO ethnicity from the others is not surprising as was apparent in their allele frequency distribution. However, their splitting into two distinct clusters may provide evidence of genetic heterogeneity within the ethnic group. They are believed to have northern African roots associated with the ancient Sao civilization that flourished around the shores of Lake Chad around ninth–fifteenth century AD, and a Hausa-Fulani land invasion from Nigeria by the 1800s that led to the establishment of a large Islamic empire involving much of the northern regions of Cameroon. Given the barriers imposed by religious differences, as much of the South of Cameroon practiced Christianity23, it is possible that some isolation of the populations was automatically imposed. Large disparities in ancestral proportions among the FO and the BA and SB ethnicities may, thus be due to a longstanding barrier to gene flow due to a sympatric differentiation among the populations (That is they live together but do not interbreed to a great extent). Such a relationship has been documented in the Malian and other African populations59,60,61,62,63,64. The separation of the FO ethnicity into two distinct clusters by chromosome painting, one close to the BA and SB and the other quite apart, further indicate that the FO is not homogeneous due to some limited but detectable coancestry with the other ethnic groups. Indeed, it is known that non-FO individuals integrated in to the Fulani–Hausa populations as herdsmen and these may have contributed to the admixed FO sub-group.
We, however, note that such a retrospective stochastic assessment of the genetic differences among Cameroonian ethnicities and other populations with respect to their demographic histories has several limitations; first, our analysis relied on self-reported ethnicity of the father and mother of each participant which may not have been accurate. Second, not only the SB of the Western Highlands cohabited with Nigerian populations during the British and French sovereignty, but the BA population of the South West as well. Perhaps, other factors may have been in play to account for the close genetic relatedness between the SB and the YRI of Nigeria. Nevertheless, these results highlight key differences in the genetic architecture of Cameroonian ethnicities that may have significant bearing on genetic association studies for this population. For instance, the fine-scale clustering of the three ethnicities based on information provided by patterns of ancestral haplotype sharing35 indicates that different haplotype structures may underlie the various ethnicities and consequently different linkage disequilibrium and recombination patterns. Given that genotype imputation, which is constantly employed to boost the power of association studies, relies upon linkage disequilibrium between SNPs9,19,65,66, it may be inferred from this analysis that the different populations would perform differently on imputation3. Therefore, to optimize the imputation on Cameroonian datasets, an imputation panel including individuals across Cameroon’s ethnic groups, or a panel—as that recently provided by the NHLBI’s Trans-Omics for Precision Medicine (TOPMed) program67—including individuals with close genetic ancestry to Cameroon’s ethnic populations would be required. In addition, considering that these genetic differences would characterize different axes of genetic variation in an association study, one would expect significant false positive results when all the ethnicities are analyzed together. Therefore, larger sample sizes would be required for association studies in such a highly structured population to be sufficiently powered to identify markers associated with specific phenotypes. Hence, association analysis performed on each ethnicity separately would be more profitable given that ethnic information is accurately captured.
Genetic association studies involving Cameroonian populations have reported disparities in the association pattern of SNPs in Cameroonians when compared to other sub-Saharan African (sSA) countries. In their recent study15, the sickle cell trait (HbAS), encoded by the rs334 locus, known to contribute the strongest protective effect against severe malaria in sSA, was found to have its weakest effect in Cameroon. One would have expected a high protective effect at this locus in the Cameroonian populations, owing to the country’s high sickle cell disease (SCD) burden68. Although the complex polygenic nature of malaria and environmental factors may be at play, it is possible that the prevalence of the HbAS trait differs significantly among Cameroonian ethnicities such that a joint analysis could have masked the real effect. More so, MalariaGEN’s analysis was based on the BA and SB ethnicities of Cameroon only. Interestingly, the FO ethnicity is more genetically close to Gambian populations (supplementary Data S1) reported to have the highest protective effect against severe malaria15. Hence this may be a classic example of the potential for population structure to bias allele frequency distribution and affect the outcome of GWA analysis.
Population genetic approaches that measure genetic distance and quantify shared ancestry are more robust when SNPs are ascertained to be polymorphic in an out-group7,8,27. However, out-group ascertainment in African populations remains a challenge as the “most recent common ancestor” of African populations remains to be established. Although the roots of anatomically modern humans have recently been traced to Botswana69, the Mende population from Sierra Leone (MSL), shown to harbor the largest proportion of ancestry from a basal African lineage7 fitted well as an out-group in our analysis. A couple of test analyses supported this observation; (1) pairwise FST estimates with the 1000 Genomes populations without SNP ascertainment required either over a million SNPs or > 50,000 SNPs with MAF > 0.35 to observe estimates similar to those previous reported, (2) SNP ascertainment with all African populations except the MSL did not result in FST estimates as have been previously reported (3) Finally, SNP ascertainment in the MSL population resulted in similar FST estimates as have been previously reported using less than a million SNPs with MAF as low as 0.05. Moreover, the MSL population was estimated to have differentiated ~ 300 thousand years ago (ka)—200 ka7, about the same time modern humans are thought to have originated from Botswana (Southern Africa)69. Hence, in the absence of a publicly available and well-established out-group for African populations, the usage of the MSL population may serve such a purpose.
Finally, we observed signatures of selection in this study, suggesting that Cameroonian populations have come under strong disease pressure. The strong signatures targeted primarily immune response and food/drug metabolism genes, suggestive of polygenic adaptation of the population to diseases and changes in diet. Selection, therefore, acts on multiple loci across multiple genes to simultaneously drive phenotypic adaptation7, although one would expect, in principle, the core locus affecting a particular trait to be under a selective sweep. Indeed, African populations have had to endure immense pressure from infectious diseases being the oldest populations of anatomically modern humans (formerly hunter-gatherers)6,7,20. The case of malaria and SCD is well established, and both conditions are thought to have emerged around the same time (4000–5000 years ago) coinciding with the adoption of agriculture in Central Africa70. Recent reports however suggest that the human Plasmodium falciparum malaria and SCD arose much earlier, and at different times; ~ 60,000 and ~ 22,000 years ago respectively42,71. We also observe weak signatures of selection in the hemoglobin-beta (HBB) gene cluster. It is expected to pick weak signatures of selection in this region by iHS (which mostly targets positive selection) in particular since the method looks at the homozygosity of clusters of SNPs72. Signatures of balancing selection would appear weak because, although alleles occur at intermediate frequencies, an extended homozygosity of alleles of the same type as the core/focal allele would signify selection.
Meanwhile, as SNP-based approaches for selection scan continue to yield useful information regarding the adaptive evolution of specific genomic loci, valuable information may still be missed at loci with more complex genetic architectures. This is particularly true for regions with structural variants and weak linkage disequilibrium such as the Dantu [GYPA-B] rearrangement in the glycophorin gene cluster, a region of ancient balancing selection73 and the common alpha-thalassemia deletions74, both of which are prevalent in malarious areas75,76,77. Therefore, novel tools that incorporate such variants in selection scans would be highly beneficial.
Conclusion
Population structure remains a major confounding factor in genetic association studies, diminishing the power of GWAS in Africa. Smarter sampling strategies and analysis designs are therefore needed to effectively detect and significantly minimize/correct population structure effects in African populations. We observe considerable substructure within our Cameroonian populations to warrant that GWAS is performed on an ethnicity basis. In addition, the analysis of a larger number of Fulani individuals in Cameroon with their Bantu and Semi-Bantu counterparts may shed further light on the contribution of their possibly non-Cameroonian ancestral proportions and the broader demographic history of this West-Central African region. The characterization of the genetic structure within other African populations will elucidate the real extent of their effects so as to inform genetic association studies on the continent.
Methods
Samples and genotyping
Our analysis was conducted with samples from unrelated individuals belonging to the Bantu, Semi-Bantu and Foulbe ethnicities from the South West, Littoral, and Centre regions of Cameroon. The individuals were recruited (1471) as part of a malaria cross-sectional study in Cameroon between 2003/05 and 2007/08 and DNA extraction was performed as previously described78. The samples were then contributed to the Malaria Genomic Epidemiology Network (MalariaGEN) Consortial Project 1 (CP1) where genotyping, genotype calling, and alignment to the human reference genome build 37 (hg19) were performed as previously described10. Data access, retrieval and analysis were performed according MalariaGEN policies79.
Quality control
Sample QC was performed on the autosomes and the X chromosome (of the 1471 samples) using Plink1.980. We explicitly removed individuals whose reported nationality was “Non-Cameroonian” or “Missing”. Individuals with discordant sex information were identified and excluded using Plink1.9′s—heck-sex filter. Furthermore, individuals with more than 10% missing data rate (missingness), and individuals with outlying heterozygosity were excluded from further analysis. Related samples were excluded by computing an identity-by-descent (IBD) matrix with 88252 high quality independent SNPs (MAF > 0.35, SNP missingness < 5%), LD-pruned such that all pairs with correlation, r2 > 0.2 within any 5 kb region, using a step size of 50b, were excluded along with all those that failed the Hardy–Weinberg equilibrium (HWE) test at p-value 1 × 10−8. One individual from each pair with Pi_HAT (mean IBD) > 0.1875 (halfway between expected IBD for third- and second-degree relatives) were removed, removing only the individual with a higher missingness (Fig.S1) and resulting in 1185 individuals. To further exclude individuals of outlying ancestry, we projected Cameroonian populations against African populations of the 1000 Genomes reference panel phase 3 (1KGP3)81,82 using smartpca version 16000 (Cambridge, MA, USA) from the EIGENSOFTv7.2 package8. This saw the removal 111 individuals, resulting in a final set of 1073 individuals. SNP quality control and subsequent analyses were performed on autosomal SNPs (2,261,351). SNPs with minor allele frequency (MAF) < 1%, missingness > 4%, and SNPs that showed significant deviation from the HWE at p-value < 1 × 10−8 were excluded. A total of 1,588,393 autosomal SNPs passed all QC procedures and were retained for further analysis.
Phasing
Prior to phasing, we excluded palindromic A/T, C/G SNPs which are usually problematic to the procedure. We then used the WRayner perl script v4.2.9 (https://www.well.ox.ac.uk/~wrayner/tools/) to check and validate the dataset against the 1KGP3 for ref/alt allele assignment, ID names, positions and alleles. VcfCooker v1.1.1 (https://github.com/statgen/gotcloud/blob/gotcloud.1.17.5/bin/vcfCooker) was used to convert the dataset from plink binary format to VCF format, while checking the validity of the dataset against the human reference genomic build 37. To further ascertain for ancestral/derived allele assignment, we used Plink280 to scrap ancestral allele assignments from ENSEMBL genome’s 1000 Genomes Phase 3 release 98 variation file downloaded from ftp://ftp.ensembl.org/pub/release-98/variation/vcf/homo_sapiens/ (Accessed 2019-09-28)83. We sorted the dataset according to base-pair position with bcftools v1.9 from the SAMtools package84 and phased each chromosome separately with EAGLE v2.4.185 against the 1KGP3 reference panel, using the combined HapMap recombination map in build 37 coordinates included in the software. We used a k parameter (–Kpbwt) of 50,000 (default 10,000) conditioning haplotypes and 10 iterations (–pbwtIters) to increase the phasing accuracy. Phasing accuracy was assessed from the log file produced by the software.
FST estimation
Prior to FST and PCA analyses, we LD-pruned the SNPs such that pairs with r2 > 0.1 within a window of 5 kb were pruned out using a step size of 50 bp. In addition, we excluded SNPs with MAF < 0.05. Pairwise FST estimates among the three ethnic groups within our study populations and among our study populations and the 1KGP3 reference populations were computed using smartpca. The tool computes the Hudson estimator of Wright’s FST which is immune to sample size differences, and uses a jackknife approach for ‘bootstrapping’, estimating a standard error and a Z score for the FST measurements. Regions of the genome with long-range LD listed in Table 1 of Price et al.,86 were excluded from FST analysis and we further utilized smartpca’s LD regression function to correct for remnants of LD prior to FST estimation.
PCA, model-based clustering, and chromosome painting
Axes of genetic variation (principal components—PCs) were computed using smartpca. Ten PCs were computed with 10 outlier removal iterations (default = 5), while maintaining all other default parameters. PC plots and FST heatmaps showing the clustering of the populations into subgroups were generated using R 3.6.129. To further investigate the population structure in our data set, we performed model-based clustering as implemented in Admixture33. Admixture utilizes a maximum likelihood approach to estimate the underlying ancestral coefficients and allele frequencies and then a moving block bootstrap approach for estimating standard errors. This analysis was performed on the set of “ancestry informative markers” described above, which were LD-pruned for MAF ≥ 0.05. The analysis was done with 5 cross-validation runs (K = 1–5) and 300 bootstrap runs. Also, the samples were sorted using Plink1.9′s –indiv-sort flag in the order BA-FO-SB so as to allow for proper grouping in the ancestral coefficients (Q) bar plot. We further resolved the clusters by running Structure v2.3.434 in the admixture mode using 3 pre-defined clusters with 20,000 burnin and 100,000 main iterations. To explore the fine-scale genetic relatedness captured by haplotype structure, we ran ChromoPainter in the linked (LD) mode on a subset of 125 Cameroonian individuals (BA = 50, SB = 50, FO = 25) (as this haplotype-based method is powerful enough to capture similar information as it would in a large sample size, thereby, effectively reducing computational cost) obtained by thinning the full data-set using Plink1.9′s –thin-indiv-count command which does a random sampling of the number of samples required. ChromoPainter summarizes the genomic proportions shared among each donor and recipient individual as a “Coancestry matrix” (Xij) of the expected number of genetic elements donated to individual i from individual j35. We then used FineStructure to assign individuals into clusters setting 100,000 burnin and sampling iterations each. Finally, we generated a tree file with FineStructure for a better assignment of the individuals into different clusters. Clusters were visualized and cluster plots produced using the FineStructure graphical user interphase (fineStructureGUI) and R.
Genome scan for signatures of selection
Signatures of selection were investigated by computing the integrated extended haplotype homozygosity (EHH) score (iHS) and cross-population locus-specific integrated (EHH) score (Rsb) using the R package REHHv3.01 (rehh)51. rehh estimates the probability that a locus, randomly shuffled from any chromosome is identical by descent to a core (focal) locus (ancestral/derived). As this probability breaks down with increasing distance upstream and downstream from the focal locus, the integral of the area under the EHH plot when the focal allele is an ancestral allele to the integral when the focal allele is a derived allele yields the iHS score. Therefore, large negative iHS values would signify selection at the derived allele and vice versa87. In the absence of a focal locus, Rsb is computed using EHH scores for all loci. Replacing the focal and derived alleles with the two populations being compared as popA and popB respectively, Rsb yields selection signatures for pairwise population comparisons. Thus, as above, large negative Rsb values would signify selection in popB and vice versa. iHS and Rsb statistics were computed in our data set using phased haplotypes with MAF ≥ 0.05. First, iHS was computed on the pooled data set (with all the ethnicities), then Rsb was computed with separate pairs of the different ethnicities (SBvsBA, SBvsFO, and BAvsFO). To assess the significance of selection signatures, rehh computes a two-sided p-value from the Gaussian cumulative distribution function of iHS estimates. The p-values were adjusted by the Benjamin–Hochberg (BH) and Bonferroni (BF) methods. Furthermore, rehh generates manhattan plots for visualization of iHS and Rsb results with their corresponding p-values. Genome-wide significance threshold for the manhattan plots was estimated by dividing 0.05 by the effective number of SNPs (those that passed all filters and were used to compute iHS) (Bonferroni method).
Extended Lewontin–Krakauer Fst outlier test (FLK)
We further computed the extended Lewontin–Krakauer Fst outlier test (FLK)46 on a dataset of unlinked loci, LD-pruned to exclude pairs of SNPs with LD (r2) > 0.1 within 5 kb genomic regions with a step size of 50 bp using hapFLKv1.4088. FLK accounts for unequal effective population sizes (Ne) and hierarchical population structure by estimating a kinship matrix which is used to model the covariance matrix of the population allele frequencies. Since the FLK test is not robust to very low levels of heterozygosity, we excluded SNPs with ancestral allele frequency (AAF) < 0.2 as recommended46. FLK test for haplotypes (hapFLK)88 was performed per chromosome using linked loci with MAF ≥ 0.05. hapFLK screens for positive selection signatures using the Sheet and Stephens model89 to summarize local haplotype diversity in a sample by clustering similar haplotypes together. The clusters are then considered as alleles to compute the FLK statistic. A K-parameter (number of clusters to model) of 30 was used while all other options were set to default. Although the genome-wide distribution of hapFLK appears to be bimodal with a large proportion of the values showing good fit to a normal distribution, Fareillo et al., implemented a means of estimating p-values via robust estimators of mean and variance from a standard normal distribution, and reducing the influence of outliers, using a rlm function in R. The p-values were adjusted following the procedure previously described. Genome-wide significance threshold was computed as previously described. hapFLK manhattan plots were generated using the qqman package in R45.
Ethical approval
All experiments in this study were conducted in adherence to the set of ethical principles of the Declaration of Helsinki. Ethical clearance for the study was obtained from the Institutional Review Board of the Faculty of Health Sciences, University of Buea (proposal number: ID D7.1.A/MPH/SWP/PDPH/PS.CH/2340/811) while administrative authorization was sought from the South West Regional Delegation of Public Health. Authorization to conduct the surveys in primary schools was obtained from the Regional Delegation of Basic Education or the Catholic Education Secretariat. Informed consent was obtained from each participant or their caregiver following a clear explanation of the content of the information sheet for the cases and blood bank donors used as controls. Authorization to enroll participants from health facilities or schools was obtained from the Director or Head teacher and only subjects/caregivers who volunteered to participate by signing a written informed consent were enrolled.
Data availability
Data used in this study are available at MalariaGEN’s Oxford Resource Center and data access and retrieval are granted according MalariaGEN’s policies via https://www.malariagen.net/. All scripts and software information including additional resources used in the analysis are present in the GitHub project (https://github.com/esohkevin/CamPopStruct).
References
Buniello, A. et al. The NHGRI-EBI GWAS Catalog of published genome-wide association studies, targeted arrays and summary statistics 2019. Nucl. Acids Res. 47, D1005–D1012 (2019).
Need, A. C. & Goldstein, D. B. Next generation disparities in human genomics: concerns and remedies. Trends Genet. 25, 489–494 (2009).
Teo, Y., Small, K. S. & Kwiatkowski, D. P. Methodological challenges of genome-wide association analysis in Africa. Nat. Rev. Genet. 11, 149–160 (2010).
Bush, W. S., Oetjens, M. T. & Crawford, D. C. Unravelling the human genome-phenome relationship using phenome-wide association studies. Nat. Rev. Genet. 17, 129–145 (2016).
Hammer, M. F., Woerner, A. E., Mendez, F. L., Watkins, J. C. & Wall, J. D. Genetic evidence for archaic admixture in Africa. Proc. Natl. Acad. Sci. 108, 15123–15128 (2011).
Uren, C. et al. Fine-scale human population structure in Southern Africa reflects ecogeographic boundaries. Genetics 204, 303–314 (2016).
Skoglund, P. et al. Reconstructing prehistoric African population structure. Cell 171, 59-71.e21 (2017).
Patterson, N., Price, A. L. & Reich, D. Population structure and eigenanalysis. PLoS Genet. 2, 2074–2093 (2006).
Band, G. et al. Imputation-based meta-analysis of severe malaria in three African populations. PLoS Genet. 9, e1003509 (2013).
Band, G., Rockett, K. A., Spencer, C. C. A. & Kwiatkowski, D. P. A novel locus of resistance to severe malaria in a region of ancient balancing selection. Nature 526, 253–257 (2015).
Timmann, C. et al. Genome-wide association study indicates two novel resistance loci for severe malaria. Nature 489, 443–446 (2012).
Ravenhall, M. et al. Novel genetic polymorphisms associated with severe malaria and under selective pressure in North-eastern Tanzania. PLoS Genet. 14, e1007172 (2018).
Adeyemo, A. A. et al. ZRANB3 is an African-specific type 2 diabetes locus associated with beta-cell mass and insulin response. Nat. Commun. 10, 3195 (2019).
Chen, J. et al. Genome-wide association study of type 2 diabetes in Africa. Diabetologia 62, 1204–1211 (2019).
Network, M. G. E. Insights into malaria susceptibility using genome-wide data on 17,000 individuals from Africa Asia and Oceania. Nat. Commun. 10, 5732 (2019).
Kang, H. M. et al. Variance component model to account for sample structure in genome-wide association studies. Nat. Genet. 42, 348–354 (2010).
Loh, P. R. et al. Efficient Bayesian mixed-model analysis increases association power in large cohorts. Nat. Genet. 47, 284–290 (2015).
Yang, J., Lee, S. H., Goddard, M. E. & Visscher, P. M. REPORT GCTA : a tool for genome-wide complex trait analysis. Am. J. Hum. Genet. 88, 76–82 (2011).
Marchini, J., Howie, B., Myers, S., McVean, G. & Donnelly, P. A new multipoint method for genome-wide association studies by imputation of genotypes. Nat. Genet. 39, 906–913 (2007).
Busby, G. B. et al. Admixture into and within sub-Saharan Africa. Elife 5, 1–44 (2016).
Uyoga, S. et al. Glucose-6-phosphate dehydrogenase deficiency and the risk of malaria and other diseases in children in Kenya: a case-control and a cohort study. Lancet Haematol. 2, e437–e444 (2015).
Clarke, G. M. et al. Characterisation of the opposing effects of G6PD deficiency on cerebral malaria and severe malarial anaemia. Elife 6, e15085 (2017).
Gardinier, D. E., DeLancey, M. W. & DeLancey, M. D. Historical dictionary of the Republic of Cameroon. Int. J. Afr. Hist. Stud. 34, 248 (2001).
Fearon, J. D. Ethnic and cultural diversity by country. J. Econ. Growth 8, 195–222 (2003).
Auton, A. et al. A global reference for human genetic variation. Nature 526, 68–74 (2015).
Sudmant, P. H. et al. An integrated map of structural variation in 2,504 human genomes. Nature 526, 75–81 (2015).
Bhatia, G., Patterson, N., Sankararaman, S. & Price, A. L. Estimating and interpreting F. Genome Res. https://doi.org/10.1101/gr.154831.113.23 (2013).
Callaway, E. First ancient African genome reveals vast Eurasian migration. Nature https://doi.org/10.1038/nature.2015.18531 (2015).
R Core Team. R: A Language and Environment for Statistical Computing. R Foundation for. Statistical. Computing. Vienna, Austria (2019). https://www.r-project.org. Accessed 25 Sept 2020.
Shriver, M. D. et al. Skin pigmentation, biogeographical ancestry and admixture mapping. Hum. Genet. 112, 387–399 (2003).
Rosenberg, N. A., Li, L. M., Ward, R. & Pritchard, J. K. Informativeness of genetic markers for inference of ancestry. Am. J. Hum. Genet. 73, 1402–1422 (2003).
Huckins, L. M. et al. Using ancestry-informative markers to identify fine structure across 15 populations of European origin. Eur. J. Hum. Genet. 22, 1190–1200 (2014).
Alexander, D. H., Novembre, J. & Lange, K. Fast model-based estimation of ancestry in unrelated individuals. Genome Res. 19, 1655–1664 (2009).
Falush, D., Stephens, M. & Pritchard, J. K. Inference of population structure using multilocus genotype data: linked loci and correlated allele frequencies. Genetics https://doi.org/10.1080/13235818.1996.10673675 (2003).
Lawson, D. J., Hellenthal, G., Myers, S. & Falush, D. Inference of population structure using dense haplotype data. PLoS Genet. https://doi.org/10.1371/journal.pgen.1002453 (2012).
McLaren, W. et al. The ensembl variant effect predictor. Genome Biol. 17, 122 (2016).
Bhatia, G. et al. Genome-wide comparison of African-ancestry populations from CARe and other cohorts reveals signals of natural selection. Am. J. Hum. Genet. 89, 368–381 (2011).
dos Santos, F. R. et al. HLA supertype variation across populations: new insights into the role of natural selection in the evolution of HLA-A and HLA-B polymorphisms. Immunogenetics 67, 651–663 (2015).
Gineau, L. et al. Balancing immunity and tolerance: genetic footprint of natural selection in the transcriptional regulatory region of HLA-G. Genes Immun. 16, 57–70 (2015).
Nielsen, R. Molecular signatures of natural selection. Annu. Rev. Genet. 39, 197–218 (2005).
Shriner, D. & Rotimi, C. N. Whole-genome-sequence-based haplotypes reveal single origin of the sickle allele during the holocene wet phase. Am. J. Hum. Genet. 102, 547–556 (2018).
Laval, G. et al. Recent adaptive acquisition by African rainforest hunter-gatherers of the late pleistocene sickle-cell mutation suggests past differences in malaria exposure. Am. J. Hum. Genet. 104, 553–561 (2019).
Lee, T., Park, C.-K. & Ha, S. Y. Prognostic role of apelin receptor expression in hepatocellular carcinoma treated with curative surgical resection. Anticancer Res. 39, 3025–3031 (2019).
Wu, X.-D. et al. Gender-specific association between Apelin/APJ gene polymorphisms and hypertension risk in Southeast China. Gene 669, 63–68 (2018).
Turner, D. S. qqman: an R package for visualizing GWAS results using Q–Q and manhattan plots. J. Open Source Softw. 3, 731 (2018).
Bonhomme, M. et al. Detecting selection in population trees: the Lewontin and Krakauer test extended. Genetics 186, 241–262 (2010).
Chen, S.-Y., Feng, Z. & Yi, X. A general introduction to adjustment for multiple comparisons. J. Thorac. Dis. 9, 1725–1729 (2017).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B 57, 289–300 (1995).
Jansen, J. C. et al. TMEM199 deficiency is a disorder of golgi homeostasis characterized by elevated aminotransferases, alkaline phosphatase, and cholesterol and abnormal glycosylation. Am. J. Hum. Genet. 98, 322–330 (2016).
Gautier, M. & Vitalis, R. rehh: an R package to detect footprints of selection in genome-wide SNP data from haplotype structure. Bioinformatics 28, 1176–1177 (2012).
Gautier, M., Klassmann, A. & Vitalis, R. rehh 2.0: a reimplementation of the R package rehh to detect positive selection from haplotype structure. Mol. Ecol. Resour. 17, 78–90 (2017).
Patin, E. et al. Dispersals and genetic adaptation of Bantu-speaking populations in Africa and North America. Science 356, 543–546 (2017).
Karakachoff, M. et al. Fine-scale human genetic structure in Western France. Eur. J. Hum. Genet. 23, 831–836 (2015).
Retshabile, G. et al. Whole-exome sequencing reveals uncaptured variation and distinct ancestry in the Southern African population of Botswana. Am. J. Hum. Genet. 102, 731–743 (2018).
Grollemund, R. et al. Bantu expansion shows that habitat alters the route and pace of human dispersals. Proc. Natl. Acad. Sci. USA 112, 13296–13301 (2015).
Lipson, M. et al. Ancient West African foragers in the context of African population history. Nature 577, 665–670 (2020).
Patin, E. & Quintana-Murci, L. The demographic and adaptive history of central African hunter-gatherers and farmers. Curr. Opin. Genet. Dev. 53, 90–97 (2018).
Fearon, J. & Laitin, D. Cameroon. (2005).
Israelsson, E. et al. Differences in Fcgamma receptor IIa genotypes and IgG subclass pattern of anti-malarial antibodies between sympatric ethnic groups in Mali. Malar. J. 7, 175 (2008).
Cherif, M. et al. Distribution of FcgammaR gene polymorphisms among two sympatric populations in Mali: differing allele frequencies, associations with malariometric indices and implications for genetic susceptibility to malaria. Malar. J. 15, 29 (2016).
Maiga, B. et al. Fc gamma Receptor IIa-H131R polymorphism and malaria susceptibility in sympatric ethnic groups, Fulani and Dogon of Mali. Scand. J. Immunol. 79, 43–50 (2014).
Nasr, A. et al. FcγRIIa (CD32) polymorphism and anti-malarial IgG subclass pattern among Fulani and sympatric ethnic groups living in eastern Sudan. Malar. J. 8, 43 (2009).
Israelsson, E. et al. Marked differences in CRP genotype frequencies between the Fulani and sympatric ethnic groups in Africa. Malar. J. 8, 136 (2009).
Maiga, B. et al. Glucose-6-phosphate dehydrogenase polymorphisms and susceptibility to mild malaria in Dogon and Fulani Mali. Malar. J. 13, 270 (2014).
Marchini, J. & Howie, B. Genotype imputation for genome-wide association studies. Nat. Rev. Genet. 11, 499–511 (2010).
Howie, B. N., Donnelly, P. & Marchini, J. A flexible and accurate genotype imputation method for the next generation of genome-wide association studies. PLoS Genet. 5, e1000529 (2009).
Taliun, D. et al. Sequencing of 53,831 diverse genomes from the NHLBI TOPMed program. bioRxiv 2, 563866 (2019).
Mulumba, L. L. & Wilson, L. Sickle cell disease among children in Africa: an integrative literature review and global recommendations. Int. J. Afr. Nurs. Sci. 3, 56–64 (2015).
Chan, E. K. F. et al. Human origins in a southern African palaeo-wetland and first migrations. Nature 575, 185–189 (2019).
Phillipson, D. W. African Archaeology. 17 (2006).
Otto, T. D. et al. Genomes of all known members of a Plasmodium subgenus reveal paths to virulent human malaria. Nat. Microbiol. 3, 687–697 (2018).
Ghansah, A. et al. Haplotype analyses of haemoglobin C and haemoglobin S and the dynamics of the evolutionary response to malaria in Kassena-Nankana district of Ghana. PLoS ONE 7, e34565 (2012).
Kariuki, S. N. et al. Red blood cell tension protects against severe malaria in the Dantu blood group. Nature 585, 579 (2020).
Piel, F. B. & Weatherall, D. J. The α-thalassemias. N. Engl. J. Med. 371, 1908–1916 (2014).
Locke, D. P. et al. Linkage disequilibrium and heritability of copy-number polymorphisms within duplicated regions of the human genome. Am. J. Hum. Genet. 79, 275–290 (2006).
Teshima, K. M. & Innan, H. The coalescent with selection on copy number variants. Genetics 190, 1077–1086 (2012).
Chakraborty, M., Emerson, J. J., Macdonald, S. J. & Long, A. D. Structural variants exhibit widespread allelic heterogeneity and shape variation in complex traits. Nat. Commun. 10, 1–11 (2019).
Achidi, E. A. et al. Severe and uncomplicated falciparum malaria in children from three regions and three ethnic groups in Cameroon: prospective study. Malar. J. 11, 1 (2012).
Malaria Genomic Epidemiology Network. MalariaGEN Consortium Internal Data Management and Access Policy. 317, 2007–2010 (2009).
Chang, C. C. et al. Second-generation PLINK: rising to the challenge of larger and richer datasets. Gigascience 4, 7 (2015).
Gibbs, R. A. et al. A global reference for human genetic variation. Nature 526, 68–74 (2015).
Altshuler, D. L. et al. A map of human genome variation from population-scale sequencing. Nature 467, 1061–1073 (2010).
Zerbino, D. R. et al. Ensembl 2018. Nucl. Acids Res. 46, D754–D761 (2018).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Loh, P. R., Palamara, P. F. & Price, A. L. Fast and accurate long-range phasing in a UK Biobank cohort. Nat. Genet. 48, 811–816 (2016).
Price, A. L. et al. Long-range LD can confound genome scans in admixed populations. Am. J. Hum. Genet. 83, 132–135 (2008).
Voight, B. F., Kudaravalli, S., Wen, X. & Pritchard, J. K. A map of recent positive selection in the human genome. PLoS Biol. 4, e72 (2006).
Fariello, M. I., Boitard, S., Naya, H., SanCristobal, M. & Servin, B. Detecting signatures of selection through haplotype differentiation among hierarchically structured populations. Genetics 193, 929–941 (2013).
Scheet, P. & Stephens, M. A fast and flexible statistical model for large-scale population genotype data: applications to inferring missing genotypes and haplotypic phase. Am. J. Hum. Genet. 78, 629–644 (2006).
Acknowledgements
We are indebted to the participants and healthcare workers from the communities who made this study possible. E.A.A. received funding for research leading to these results from MalariaGEN programme of the Wellcome Trust Sanger Institute, Hinxton, UK. We thank Victoria Nembaware, Gaston K. Mazandu, and Oluwafemi Gabriel Oluwole for their valuable comments. We also thank the MalariaGEN Oxford Resource Center for the genotyping work. This work was supported through the DELTAS Africa Initiative [DELGEME Grant 107740/Z/15/Z]. The DELTAS Africa Initiative is an independent funding scheme of the African Academy of Sciences (AAS)’s Alliance for Accelerating Excellence in Science in Africa (AESA) and supported by the New Partnership for Africa’s Development Planning and Coordinating Agency (NEPAD Agency) with funding from the Wellcome Trust [DELGEME Grant 107740/Z/15/Z] and the UK government. Funding for bioinformatics skill development and data analysis was provided by the H3ABioNet project. The H3ABioNet is supported by the National Institutes of Health (NIH) Common Fund [Grant Number U41HG006941]. The views expressed in this publication are those of the author(s) and not necessarily those of AAS, NEPAD Agency, Wellcome Trust, NIH or the UK government.
Author information
Authors and Affiliations
Contributions
K.K.E., A.A.N., T.O.A. and S.G.N. designed the study, analyzed data and wrote the manuscript. K.K.E. and E.R.C. carried out bioinformatics analysis of genotype data. E.A.A. and T.O.A. coordinated field sampling and laboratory work. A.A.N., L.A.E., A.W., and E.R.C. advised on population genetics. Supervision; T.O.A., S.G.N., A.W., and E.R.C. All authors read and suggested revision on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Esoh, K.K., Apinjoh, T.O., Nyanjom, S.G. et al. Fine scale human genetic structure in three regions of Cameroon reveals episodic diversifying selection. Sci Rep 11, 1039 (2021). https://doi.org/10.1038/s41598-020-79124-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-79124-1
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.