Abstract
Allogeneic stem cell transplantation is currently the only curative treatment option for myelodysplastic syndromes (MDS). Pre-transplant debulking treatment have been employed for advanced MDS and we previously reported that marrow response (blast ≤ 5%) following the bridging therapy with hypomethylating agent was an independent favorable factor for survival; however, it is still not clear which patients will respond to hypomethylating agent and which genomic features can predict the response. In this study, we performed RNAseq for 23 MDS patients among which 14 (61%) and 9 (39%) patients showed marrow complete remission and primary resistance to azacitidine, respectively. Differential expression-based analyses of treatment-naive, baseline gene expression profiles revealed that molecular functions representing mitochondria and apoptosis were up-regulated in responders. In contrast, we identified genes involved in the Wnt pathway were relatively up-regulated in non-responders. In independent validation cohorts of MDS patients, the expression of gene sets specific to non-responders and responders distinguished the patients with favorable prognosis and those responded to azacitidine highlighting the prognostic and predictive implication. In addition, a systems biology approach identified genes involved in ubiquitination, such as UBC and PFDN2, which may be key players in the regulation of differential gene expression in treatment responders and non-responders. Taken together, identifying the gene expression signature may advance our understanding of the molecular mechanisms of azacitidine and may also serve to predict patient responses to drug treatment.
Similar content being viewed by others
Introduction
Myelodysplastic syndromes (MDS) are clonal stem cell disorders characterized by peripheral blood cytopenia due to ineffective hematopoiesis and by their potential to evolve into acute myeloid leukemia (AML)1. The ineffective hematopoiesis is associated with elevated apoptosis of hematopoietic clones, a phenomenon that may be related to signaling mediated by TRAIL ligands/receptors2, FAS molecules3 and/or myelosuppressive TGFβ4. Excessive apoptosis occurs in the early stages of MDS and declines as MDS progresses toward the leukemic stage. About one third of MDS patients progress to AML, which is probably associated with expansion and evolution of subclones with certain mutations at the stem cell level5,6. The genetic profiles of individual patients are heterogeneous and an individual patient’s unique genetic makeup may impact their clinical phenotype, prognosis, and response to therapy. An MDS patient’s genome frequently harbors somatic mutations which may contribute to disease progression; however, there is no single gene that appears to be sufficient to elicit disease and the majority of genes mutated in MDS are present in fewer than 5% of cases, highlighting the complexity of the disease7,8. These heterogenity of MDS have led to the development of risk-based stratification systems, and treatment options such as hypomethylating agent (HMA) or allogeneic stem cell transplantation (SCT) are selected systematically according to risk groups9,10,11.
The use of HMA, such as azacitidine (AZA) and decitabine, has become a standard treatment for higher-risk MDS patients, as HMA confer a survival benefit over conventional care12,13, but many issues, including determining the optimum dose or duration of treatment, remain to be addressed. Importantly, it is still not clear which patients will respond to HMA or not. A number of predictive genomic markers of the response to HMA treatment have been reported among the frequent genomic aberrations in MDS genomes. For example, mutations in genes involved in DNA methylation (DNMT3A, TET2, IDH1, and IDH2) and genes encoding epigenetic regulations (ASXL1, EZH2, and TET2) have been proposed to predict AZA sensitivity14,15,16, but the clinical relevance of these associations remains controversial17,18,19. A DNA methylation-based scoring system using a suite of 10 genes has been proposed, but while the score was correlated with patient survival, it could not predict clinical responses to demethylating agents20. The expression of a number of other genes, such as BCL2L1021, FAS22, and PI-PLCbeta123, may also be indicative of HMA activity. Moreover, associations between repressed expression of anti- DNMT1 miRNAs and HMA resistance have been noted24. All of these findings have contributed substantially to a better understanding of individual variability in HMA responses among MDS patients; However, mutation, methylation, or expression information involving limited genes are not full enough to understand disease heterogeneity with respect to different response to HMA treatment. In this regard, genome-wide profiling of mRNA expression may facilitate comprehensive prediction of drug sensitivity.
One important usage of HMA is to serve as a bridge to SCT by diminishing marrow blasts to an acceptable level prior to transplantation25. Several retrospective studies have shown that pre-SCT HMA could be a feasible alternative to induction chemotherapy26,27,28 even for patients with excessive blasts or AML29. As the influence of HMA response on transplant outcomes is still controversial, we previously completed a retrospective analysis of 98 patients who received HMA for higher-risk MDS with > 5% marrow blasts30. Our study showed that continued marrow response at the time of SCT was an independent positive predictor of overall and disease-free survival after transplantation. In light of our findings and those of another study in which a positive correlation between marrow clearance and genomic mutations was reported31, we aimed to identify the molecular markers that could predict a patient’s marrow response to HMA.
In this study, we performed RNAseq-based gene expression profiling of advanced MDS patients with excess blasts (EB) prior to their receiving AZA treatment. Comparison of gene expression profiles between 14 responders and 9 non-responders revealed a number of molecular functions (e.g., apoptosis and cellular respiration) were relatively activated in responders. Functional scores of these molecular pathways were also correlated with patient survival in an independent MDS cohort, and the results suggested their potential prognostic value.
Results
Sequencing data from enrolled patients
A total of 23 patients who received AZA for MDS with excess blasts 1 (MDS-EB-1; n = 9) and MDS-EB-2 (n = 14) before SCT were enrolled in the study (Table 1). The non-responder group consisted of 9 cases showing primary resistance, which included 8 cases with disease progression to AML (n = 6) or MDS-EB-2 (n = 2) and 1 case with stable disease without any hematological improvement (SD-HI) after 5 cycles of AZA treatment. The responder group consisted of cases with complete remission (CR) (n = 7) or marrow CR (mCR) with or without any hematological improvement (n = 7). The patients’ bone marrow derived mononuclear cells were subjected to transcriptome sequencing by RNAseq. The sequencing-related information is presented in Supplementary Table 1.
Differential-expression-based identification of marker genes and molecular functions
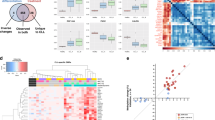
We first selected differentially expressed genes (DEGs) between responders and non-responders (n = 300; P < 0.01, t-test, unadjusted, FPKM and Combat adjusted data). The hierarchical clustering of 300 DEGs was able to segregate responders and non-responders, with the exception of one outlier (Fig. 1a). These findings suggest the baseline mRNA expression of MDS patients may have the potential to predict sensitivity to AZA. The list of DEGs is presented in Supplementary Table 2. To identify the molecular functions associated with the DEGs, we performed Fisher's exact test with Gene Ontology (GO) categories (MSigDB C5 category) and found significant associations in 22 GO categories (Bonferroni corrected P < 0.05). To summarize the identified GO categories by taking into account the redundancy in GO categories, we measured the extent of overlap for all possible pairs of the 22 GO categories identified above. Of note, all possible pairs of the 22 GO categories showed significant overlap between DEGs (P values of Fisher's exact test in the range of 1.2E−25 to 5.4E−55), which suggested the gene–gene co-expression networks were highly interconnected with respect to GO functional categories. The 22 GO categories are illustrated in a network diagram (Fig. 1b). The examination of 22 GO functional annotations in MDS patients indicated that the DEGs between AZA responders and non-responders largely represented 'cellular respiration' and 'mitochondria'. The most significant pairing between GO categories was for the 'electron transport chain' and 'respiratory chain' (P = 5.4E−55; Fisher's exact test) with overlapping DEGs for NDUFS7, COX8A, UBA52, UQCR11, NDUFA2, NDUFB7, NDUFB8, NDUFA7, NDUFB2, COX4I1, NDUFA13, ETFB, NDUFA1, NDUFB9, NDUFA11, and UBC. Oxidative stress and mitochondrial dysfunction have been postulated to play a role in the development of MDS32. Although the association between sensitivity to AZA and mitochondrial dysfunction has not been clearly defined, a number of reports suggest that increased oxidative phosphorylation relative to glycolysis may indicate metabolic plasticity of tumor cells and is associated with the resistance to chemotherapy in colorectal cancers33 and targeted therapeutics in melanomas34.
Hierarchical clustering and functional enrichment analysis of DEGs. (a) The 300 DEGs (P < 0.01; t-test, unadjusted) are shown in a heatmap with gene- and sample-wise dendrograms by hierarchical clustering. Non-responders (red; n = 9) are largely segregated from responders (green; n = 14) except for an outlier. (b) Twenty-two GO categories with significant enrichment (Bonferroni corrected P < 0.05; Fisher's exact test) with DEGs are shown as nodes in a network. All the pairs of nodes showed significant overlap of DEGs and were connected by edges in the network. The GO annotations of 22 nodes converge on the function of ‘cellular respiration' and 'mitochondria'. The node size corresponds to the gene number of the corresponding GO categories.
Enrichment analyses
To further identify subtle but functionally coordinated mRNA expression changes that cannot be captured by DEG analyses, we performed additional gene set enrichment analyses (GSEA) to identify molecular functions for which the gene members are differentially expressed between AZA responders and non-responders. Table 2 lists the 10 most significant GO categories relatively up-regulated in responders compared to non-responders, as well as 10 GO categories up-regulated in non-responders compared to responders. To summarize the GO categories according to the overlap of gene members, we also calculated the significance of the overlap of gene members and a total of 20 GO categories are shown in a network diagram (Fig. 2a). In the network, a main network composed of eight GO categories largely representing oxidative phosphorylation and the electron transport chain were found to be relatively up-regulated in responders compared to non-responders (green nodes in Fig. 2a), which is consistent with the results of the DEG analyses. In addition, GO categories of 'leukocyte apoptotic process' and 'regulation of transcription elongation' were evident, suggesting that these functions are relatively activated in responders. With respect to GO categories relatively up-regulated in non-responders, 'dendrite development/morphogenesis' (3 GO categories) and 'Wnt protein binding/brain morphogenesis' (3 GO categories) were found to be major functional categories (red nodes in Fig. 2a).
GSEA analysis. (a) The top 20 significant gene sets relatively up- and down-regulated (green and red, n = 10 and 10, respectively) in responders compared to non-responders are shown as nodes in a network. The node size is proportional to the number of genes in the gene set. Edges in the network represent the significant overlap of the leading edge genes between two nodes. (b) An enrichment plot of the leukocyte apoptotic process is shown as a snapshot of the GSEA analysis. A heatmap showing the expression of genes belonging to the set is shown below with the annotation of genes ordered. Yellow indicates the leading-edge gene subsets. (c) A similar plot of the gene set of Wnt protein binding.
Two examples of GO categories ('leukocyte apoptosis process' and ‘Wnt protein binding’, which are up- and down-regulated, respectively, in responders) are selected to show their enrichment plots as well as their expression heat maps and leading-edge genes (Fig. 2b,c), respectively. Ineffective hematopoiesis is one of cardinal features of MDS, and is thought to be due to increased apoptosis of myeloid progenitors; the acquisition of proliferative capacity by clonal progenitors has been considered the key event in the progression of MDS to acute myeloid leukemia8. Thus, it is reasonable to assume that elevated expression of apoptosis-related genes at baseline may predict a favorable prognosis and responsiveness to AZA. In our study, we found no significant differences in FAS expression between responders and non-responders (Fig. 2b; P = 0.48; t-test, unadjusted), or between BCL2L10 and PLCB1 expression (P = 0.69 and 0.42; t-test, unadjusted), both of which have previously been proposed as expression markers of AZA sensitivity21,23. In our study, only two genes, BAX and IFNG, had significantly different expressions between the responders and non-responders (P = 0.03 and 0.04, respectively; t-test, unadjusted). Wnt/β-catenin signaling has been presumed to be involved in the ineffective hematopoiesis of patients with myeloid neoplasms with 5q deletions, and has also been presumed to have therapeutic implications in MDS through inhibition of this signaling35. Our results further suggest that the mRNA expression levels in Wnt/β-catenin pathway genes including a number of frizzled receptors may indicate the responsiveness to AZA with similar implications in MDS-derived stromal cells35. It is postulated that up-regulated expression of Wnt pathway genes at baseline in non-responders encodes cellular resistance to AZA treatment. Other GO categories that showed substantial differences in expression with respect to HMA responses are shown in Supplementary Fig. 1, in which enrichment plots and heatmaps of two metabolic functions (‘Nucleoside monophosphate metabolic process’ and ‘Oxidoreductase activity acting on NADPH’) are demonstrated.
In addition to assessing DEG and GSEA, we further employed a DeMAND network-based system biological approach36 to identify the genes related to the mechanism of action (MoA) of AZA. In this process, the level of perturbation was measured for individual genes and their interacting partners in regulatory networks by comparing the gene expression profiles of responders and non-responders following drug perturbation. The 20 most significant candidate MoA genes are listed in Table 3. The modeling revealed UBC and PFDN2 as candidate MoA genes whose protein interactions with interacting partners were the most perturbed in the context of the regulatory network.
Validation of azacitidine sensitivity signatures in independent cohorts
To confirm the impact of the identified gene markers on prognosis and HMA response, we applied the gene set-level scores (i.e., the mean expression of leading edge genes of 20 GO terms in Table 2) in independent data sets. We obtained publicly available gene expression profiles of 123 MDS patients with overall survival (GSE58831)37 and 32 patients with AZA treatment history (13 responders and 19 non-responders; GSE77750)38,39. For survival (GSE58831), MDS patients whose gene expression profiles resembled those of AZA responders in our study (‘responder-like’) showed better survival (green in Fig. 3a,b) compared to those whose gene expression resembled non-responders (‘non-responder-like’, red in Fig. 3a,b; log-rank test P = 0.017). This result suggests that the baseline expression profile from AZA responders may be associated with a favorable prognosis in MDS patients highlighting the prognostic implication of the identified genes in our study. For AZA responsiveness (GSE77750), the clustering using identified markers segregated the 32 MDS patients into ‘responder-like’ and ‘non-responder-like’ cases with the modest level of prediction accuracy 0.687 (Fig. 3c). Thus, our gene markers may have both prognostic and predictive implication with the AZA treatment for MDS patients.
Survival analysis of independent MDS cohorts. (a) Clustering of marker gene expression by gene-set level scores segregated the 123 MDS patients (GSE58831) is shown with a heatmap. Column-wise green/red bars below indicate the patients whose expression profiles resemble those of azacitidine responders/non-responders, respectively (‘responder-like’ and ‘non-responder-like’) (b) MDS patients segregated according to expression profiles showed significant survival differences (P = 0.017, log-rank test). The green and red lines indicate the patients whose scores resemble those of azatidicine responders and non-responders (green and red bars below in Fig. 3a), respectively. (c) Clustering of marker gene expression segregated the 32 MDS patients (GSE77750) into responder-like and non-responder-like groups. Responder-like patients were enriched with AZA responders (green, top) while non-responder-like patients were enriched with non-responders (here, progressed/red and stable/orange were considered as AZA non-responders).
We also tested MoA genes in an independent cohort (GSE58831) to test the prognostic values of the markers. Although not significant, the expression of MoA genes was able to segregate the MDS patients with a substantial difference in overall survival suggesting that the MoA genes, in spite of their small size, may be utilized for prognostic markers (Supplementary Fig. 2a,b).
Discussion
Although HMA response in MDS is known to be associated with better survival both in transplant and non-transplant settings12,30,40, it has not been possible to clearly identify which patients will respond to drug therapy. In this study, we performed next-generation transcriptome sequencing to identify gene-expression-based predictive markers that can discriminate responders from non-responders. For prediction, we first identified a set of DEGs that can segregate the AZA responders and non-responders. The DEGs identified in this study were also able to group the patients from an independent MDS cohort into subgroups of distinguished drug response and overall survival, indicating the selected genes may potentially identify AZA responders and MDS patients with favorable or unfavorable prognosis. Given our study was based on baseline gene expression data prior to therapy, our findings may potentially support a clinical decision to select AZA as a bridging treatment for MDS patients with excess blasts.
The expression of previously proposed single genetic markers of favorable treatment outcomes, such as FAS, BCL2L10, and PI-PLCbeta1 were not able to segregate AZA responders and non-responders in our study. One possible explanation for this discrepancy is that our study only included MDS-EB patients and the definition of a responder in this study was confined to bone marrow response and did not include either partial response or stable disease with hematological improvement. The identification of MDS at the level of single-gene markers is complicated by tumor heterogeneity: in this regard, it has been proposed that multi-gene signatures or gene-set-level classifiers may be more clinically relevant than single-gene markers. Although the repressed expression of the apoptosis-related molecule FAS by aberrant DNA methylation may be indicative of AZA sensitivity, the baseline expression of FAS was not significantly different between responders and non-responders in this study. However, we note that ‘leukocyte apoptosis process’ was identified as one of the significant GO terms showing differential expression in a set of genes in our study. Among the genes belonging to the GSEA top 20 gene set list, we identified two genes, BAX and IFNG belonging to the leukocyte apoptosis process has only significant differential expression (P < 0.05, t-test, unadjusted, Fig. 2b) and are up-regulated in responders compare with non-responders. The pro-apoptotic protein encoded by BAX is transcriptionally activated in MDS patients with more favorable survival outcomes41 and IFNG is known to trigger apoptosis in undifferentiated progenitor cells such as hematopoietic stem cells42. Although the potential roles of these genes in the context of marrow response to AZA require further validation in an independent cohort, the molecular pathways and marker genes identified in our functional-gene-set-level differential analyses may have clinical utility. Supplementary Fig. 3 shows the level of differential expression of BAX and IFNG were modest, e.g., ranked 429th and 442nd in the differential expression-ordered list of total genes, respectively. In addition, 'Wnt protein binding' was also identified as one of the molecular functions relatively up-regulated in non-responders at baseline. The Wnt pathway represents one of the key molecular pathways in carcinogenic processes43 and we assume that this pathway may provide survival signals to the tumor clones. In the survival analyses, we found that the gene set level of 20 GO terms can segregate the patients according to survival outcomes. Notably, gene expression profiles characteristic of AZA marrow responders identified in this study were significantly linked to better survival in an independent MDS cohort, which suggests that the genes identified in our study may have potential clinical utility as general prognostic markers. However, we acknowledge that the results must be interpreted with a caution given the substantial differences between our cohort and validation cohort regarding examined cells, patients’ characteristics and clinical outcomes of interests. We examined mononuclear cells in this study whereas CD34 + cells were used in two validation cohorts (GSE58831, GSE77750), although recent study by Shiozawa et al. showed the concordance of expression levels for selected genes between bone marrow CD34 + cells and mononuclear cells from MDS patients44. In addition, advanced MDS patients with excess blasts who were treated with HMA were only included in our study, while all subtypes of MDS patients with no information on treatment (GSE58831) or AZA treated AML patients, not only MDS patients (GSE77750) were analyzed in validation cohorts. Actually, when we selected EB patients only in validation cohort (GSE58831), the prognostic impact according to the expression of marker genes was not significant (Supplementary Fig. 2c,d), probably due to the small sample size. Based on these discrepancies between the studies along with the small sample size, our results should be explored in a further investigation.
The DeMAND algorithm was employed as a systems biology approach to identify network-level perturbation and to reveal key MoA genes. Through this process, UBC and PFDN2 were designated as potential MoA proteins whose interactions were substantially perturbed by AZA treatment. Of interest, two genes have roles in protein metabolism such as ubiquitination (UBC) and as protein chaperone (PFDN2). It has been previously reported that ubiquitination is implicated in histone modification45 and the ubiquitin–proteasome pathway is associated with the stability of the DNMT1 protein, findings which indicate a potential molecular link between ubiquitination and the demethylating agent AZA46. Therefore, we may assume that UBC may have a role in the regulation of gene expression associated with AZA treatment.
Taken together, the markers for the AZA sensitivity identified in this study may predict drug responsiveness prior to AZA treatment and may have general prognostic implications.
Methods
Patient selection
To identify molecular markers in MDS genomes associated with marrow responses following pre-transplant bridging treatment with AZA patients who received the standard schedule of AZA (75 mg/m2/day for 7 consecutive days) for MDS with > 5% marrow blasts and cases showing extremes of treatment response (marrow complete remission versus primary treatment failure) were screened. Response to treatment was assessed using the modified International Working Group response criteria47. Cases achieving CR or mCR with or without hematological improvement were categorized as responders, while non-responders consisted of those who experienced primary treatment failure defined either by primary disease progression or stable disease without hematological improvement (SD-HI)48. Minimum 4 cycles of AZA was administered before response assessment of SD-HI, whereas assessment of CR, mCR or disease progression was allowed to assess even before the 4th cycles. The patients did not receive previous treatment before AZA for their MDS with excess blasts. Patients with bone marrow samples available for research purposes were enrolled in the final study population. This study was approved by the Institutional Review Board of the Seoul St. Mary’s Hematology Hospital at the Catholic University of Korea, and complied with the tenets of the Declaration of Helsinki. Informed consent was obtained from all subjects.
mRNA sequencing
Bone marrow samples (10 mL) were mixed with 0.3 mL of heparin to prevent coagulation, then diluted with 20 mL of phosphate‐buffered saline (PBS, Welgene, Daegu, Korea). The cells were then fractionated on a Lymphoprep density gradient (Axis‐Shield, Oslo, Norway) through centrifugation at 600 g for 10 min. Interface mononuclear cells were isolated and washed with PBS. An erythrocyte (RBC) lysis buffer (0.154 M NH4Cl, 10 mM KHCO3, 0.1 mM EDTA; Sigma Aldrich, St. Louis, MO) was then added to destroy contaminating RBCs. The RNA was isolated using the standard Tri-Reagent1(Sigma Chemicals) protocol. Preparation of mRNA libraries was performed using an illumina (REF. RS-122-2101 ~ 2) TruSeq stranded kit as manufacturer’s recommendation. The sequencing was performed using the NextSeq500 platform (illumina, REF. SY-415-1001). Paired-end 75 bp sequencing reads were generated and the sequencing information is presented in Supplementary Table 1. Raw sequencing reads in FASTQ files were mapped and aligned by TopHat (version.2.1.1)49. Transcript-level alignment was completed using Cufflinks50 using annotated transcripts of hg19 GTF (UCSC, TCGA.hg19.June2011.gaf). The expression of each gene was represented in terms of fragments per kilobase million (FPKM) and used for the subsequent analyses. RNA sequencing data have been submitted to the Sequence Read Archive (SRA) under accession number of PRJNA650236.
GSEA
Gene set enrichment analysis51 (GSEA, version 2.0, https://software.broadinstitute.org/gsea) was used to identify differential expression of genes associated with specific molecular functions between drug responders and non-responders52. For molecular functions, we used Gene Ontology gene sets as available in MSigDB (MSigDB, C5:GO terms, version 6.0).
References
Ades, L., Itzykson, R. & Fenaux, P. Myelodysplastic syndromes. Lancet (London, England) 383, 2239–2252. https://doi.org/10.1016/s0140-6736(13)61901-7 (2014).
Zang, D. Y., Goodwin, R. G., Loken, M. R., Bryant, E. & Deeg, H. J. Expression of tumor necrosis factor-related apoptosis-inducing ligand, Apo2L, and its receptors in myelodysplastic syndrome: effects on in vitro hemopoiesis. Blood 98, 3058–3065. https://doi.org/10.1182/blood.v98.10.3058 (2001).
Claessens, Y. E. et al. In vitro proliferation and differentiation of erythroid progenitors from patients with myelodysplastic syndromes: evidence for Fas-dependent apoptosis. Blood 99, 1594–1601. https://doi.org/10.1182/blood.v99.5.1594 (2002).
Bhagat, T. D. et al. miR-21 mediates hematopoietic suppression in MDS by activating TGF-beta signaling. Blood 121, 2875–2881. https://doi.org/10.1182/blood-2011-12-397067 (2013).
Chen, J. et al. Myelodysplastic syndrome progression to acute myeloid leukemia at the stem cell level. Nat. Med. 25, 103–110. https://doi.org/10.1038/s41591-018-0267-4 (2019).
da Silva-Coelho, P. et al. Clonal evolution in myelodysplastic syndromes. Nat. Commun. 8, 15099. https://doi.org/10.1038/ncomms15099 (2017).
Bejar, R. Myelodysplastic syndromes diagnosis: what is the role of molecular testing?. Curr. Hematol. Malig. Rep. 10, 282–291. https://doi.org/10.1007/s11899-015-0270-5 (2015).
Raza, A. & Galili, N. The genetic basis of phenotypic heterogeneity in myelodysplastic syndromes. Nat. Rev. Cancer 12, 849–859. https://doi.org/10.1038/nrc3321 (2012).
Greenberg, P. et al. International scoring system for evaluating prognosis in myelodysplastic syndromes. Blood 89, 2079–2088 (1997).
Greenberg, P. L. et al. Revised international prognostic scoring system for myelodysplastic syndromes. Blood 120, 2454–2465. https://doi.org/10.1182/blood-2012-03-420489 (2012).
Malcovati, L. et al. Impact of the degree of anemia on the outcome of patients with myelodysplastic syndrome and its integration into the WHO classification-based Prognostic Scoring System (WPSS). Haematologica 96, 1433–1440. https://doi.org/10.3324/haematol.2011.044602 (2011).
Fenaux, P. et al. Efficacy of azacitidine compared with that of conventional care regimens in the treatment of higher-risk myelodysplastic syndromes: a randomised, open-label, phase III study. Lancet Oncol. 10, 223–232. https://doi.org/10.1016/S1470-2045(09)70003-8 (2009).
Griffiths, E. A. & Gore, S. D. DNA methyltransferase and histone deacetylase inhibitors in the treatment of myelodysplastic syndromes. Semin. Hematol. 45, 23–30. https://doi.org/10.1053/j.seminhematol.2007.11.007 (2008).
Bejar, R. et al. TET2 mutations predict response to hypomethylating agents in myelodysplastic syndrome patients. Blood 124, 2705–2712. https://doi.org/10.1182/blood-2014-06-582809 (2014).
Itzykson, R. et al. Impact of TET2 mutations on response rate to azacitidine in myelodysplastic syndromes and low blast count acute myeloid leukemias. Leukemia 25, 1147–1152. https://doi.org/10.1038/leu.2011.71 (2011).
Traina, F. et al. Impact of molecular mutations on treatment response to DNMT inhibitors in myelodysplasia and related neoplasms. Leukemia 28, 78–87. https://doi.org/10.1038/leu.2013.269 (2014).
Meldi, K. et al. Specific molecular signatures predict decitabine response in chronic myelomonocytic leukemia. J. Clin. Investig. 125, 1857–1872. https://doi.org/10.1172/jci78752 (2015).
Braun, T. et al. Molecular predictors of response to decitabine in advanced chronic myelomonocytic leukemia: a phase 2 trial. Blood 118, 3824–3831. https://doi.org/10.1182/blood-2011-05-352039 (2011).
Jung, S. H. et al. Somatic mutations predict outcomes of hypomethylating therapy in patients with myelodysplastic syndrome. Oncotarget 7, 55264–55275. https://doi.org/10.18632/oncotarget.10526 (2016).
Shen, L. et al. DNA methylation predicts survival and response to therapy in patients with myelodysplastic syndromes. J. Clin. Oncol. 28, 605–613. https://doi.org/10.1200/jco.2009.23.4781 (2010).
Cluzeau, T. et al. BCL2L10 is a predictive factor for resistance to azacitidine in MDS and AML patients. Oncotarget 3, 490–501. https://doi.org/10.18632/oncotarget.481 (2012).
Ettou, S. et al. Fas expression at diagnosis as a biomarker of azacitidine activity in high-risk MDS and secondary AML. Leukemia 26, 2297–2299. https://doi.org/10.1038/leu.2012.152 (2012).
Follo, M. Y. et al. Reduction of phosphoinositide-phospholipase C beta1 methylation predicts the responsiveness to azacitidine in high-risk MDS. Proc. Natl. Acad. Sci. USA 106, 16811–16816. https://doi.org/10.1073/pnas.0907109106 (2009).
Solly, F. et al. An miRNA-DNMT1 axis is involved in azacitidine resistance and predicts survival in higher-risk myelodysplastic syndrome and low blast count acute myeloid Leukemia. Clin. Cancer Res. 23, 3025–3034. https://doi.org/10.1158/1078-0432.Ccr-16-2304 (2017).
NCCN.org. Myelodysplastic syndromes. NCCN guidelines Version 2.2019 (2018).
Damaj, G. et al. Impact of azacitidine before allogeneic stem-cell transplantation for myelodysplastic syndromes: a study by the Societe Francaise de Greffe de Moelle et de Therapie-Cellulaire and the Groupe-Francophone des Myelodysplasies. J. Clin. Oncol. 30, 4533–4540. https://doi.org/10.1200/jco.2012.44.3499 (2012).
Field, T. et al. 5-Azacitidine for myelodysplasia before allogeneic hematopoietic cell transplantation. Bone Marrow Transplant. 45, 255–260. https://doi.org/10.1038/bmt.2009.134 (2010).
Gerds, A. T. et al. Pretransplantation therapy with azacitidine vs induction chemotherapy and posttransplantation outcome in patients with MDS. Biol. Blood Marrow Transpl. J. Am. Soc. Blood Marrow Transpl. 18, 1211–1218. https://doi.org/10.1016/j.bbmt.2012.01.009 (2012).
Lubbert, M. et al. Non-intensive treatment with low-dose 5-aza-2’-deoxycytidine (DAC) prior to allogeneic blood SCT of older MDS/AML patients. Bone Marrow Transpl. 44, 585–588. https://doi.org/10.1038/bmt.2009.64 (2009).
Yahng, S. A. et al. Better transplant outcome with pre-transplant marrow response after hypomethylating treatment in higher-risk MDS with excess blasts. Oncotarget 8, 12342–12354. https://doi.org/10.18632/oncotarget.12511 (2017).
Welch, J. S. et al. TP53 and decitabine in acute Myeloid Leukemia and myelodysplastic syndromes. N. Engl. J. Med. 375, 2023–2036. https://doi.org/10.1056/NEJMoa1605949 (2016).
Goncalves, A. C. et al. Oxidative stress and mitochondrial dysfunction play a role in myelodysplastic syndrome development, diagnosis, and prognosis: a pilot study. Free Radical Res. 49, 1081–1094. https://doi.org/10.3109/10715762.2015.1035268 (2015).
Vellinga, T. T. et al. SIRT1/PGC1alpha-dependent increase in oxidative phosphorylation supports chemotherapy resistance of colon cancer. Clin. Cancer Res. 21, 2870–2879. https://doi.org/10.1158/1078-0432.Ccr-14-2290 (2015).
Haq, R. et al. Oncogenic BRAF regulates oxidative metabolism via PGC1alpha and MITF. Cancer Cell 23, 302–315. https://doi.org/10.1016/j.ccr.2013.02.003 (2013).
Bhagat, T. D. et al. Epigenetically Aberrant Stroma in MDS propagates disease via Wnt/beta-catenin activation. Can. Res. 77, 4846–4857. https://doi.org/10.1158/0008-5472.Can-17-0282 (2017).
Walter, R. B. et al. Comparison of minimal residual disease as outcome predictor for AML patients in first complete remission undergoing myeloablative or nonmyeloablative allogeneic hematopoietic cell transplantation. Leukemia 29, 137–144. https://doi.org/10.1038/leu.2014.173 (2015).
Gerstung, M. et al. Combining gene mutation with gene expression data improves outcome prediction in myelodysplastic syndromes. Nat. Commun. 6, 5901. https://doi.org/10.1038/ncomms6901 (2015).
Merkerova, M. D. et al. Relationship between altered miRNA expression and DNA methylation of the DLK1-DIO3 region in azacitidine-treated patients with myelodysplastic syndromes and acute Myeloid Leukemia with Myelodysplasia-related changes. Cells 7, 138. https://doi.org/10.3390/cells7090138 (2018).
Monika Belickova, M. et al. Up-regulation of ribosomal genes is associated with a poor response to azacitidine in myelodysplasia and related neoplasms. Int. J. Hematol. 104, 566–573. https://doi.org/10.1007/s12185-016-2058-3 (2016).
Potter, V. T. et al. Comparison of intensive chemotherapy and hypomethylating agents before allogeneic stem cell transplantation for advanced myelodysplastic syndromes: a study of the myelodysplastic syndrome subcommittee of the chronic malignancies working party of the European society for blood and marrow transplant research. Biol. Blood Marrow Transpl. J. Am. Soc. Blood Marrow Transpl. 22, 1615–1620. https://doi.org/10.1016/j.bbmt.2016.05.026 (2016).
Diamantidis, M. et al. The role of apoptosis and current therapeutic challenges in myelodysplastic syndromes. Hippokratia 11, 178–182 (2007).
Morales-Mantilla, D. E. & King, K. Y. The role of interferon-gamma in hematopoietic stem cell development, homeostasis, and disease. Curr. Stem Cell Rep. 4, 264–271. https://doi.org/10.1007/s40778-018-0139-3 (2018).
Bhavanasi, D. & Klein, P. S. Wnt signaling in normal and malignant stem cells. Curr. Stem Cell Rep. 2, 379–387. https://doi.org/10.1007/s40778-016-0068-y (2016).
Shiozawa, Y. et al. Gene expression and risk of leukemic transformation in myelodysplasia. Blood 130, 2642–2653. https://doi.org/10.1182/blood-2017-05-783050 (2017).
Moschos, M. M. et al. The role of histone deacetylase inhibitors in uveal melanoma: current evidence. Anticancer Res. 38, 3817–3824. https://doi.org/10.21873/anticanres.12665 (2018).
Ghoshal, K. et al. 5-Aza-deoxycytidine induces selective degradation of DNA methyltransferase 1 by a proteasomal pathway that requires the KEN box, bromo-adjacent homology domain, and nuclear localization signal. Mol. Cell. Biol. 25, 4727–4741. https://doi.org/10.1128/mcb.25.11.4727-4741.2005 (2005).
Cheson, B. D. et al. Clinical application and proposal for modification of the International Working Group (IWG) response criteria in myelodysplasia. Blood 108, 419–425. https://doi.org/10.1182/blood-2005-10-4149 (2006).
Prebet, T. et al. Outcome of high-risk myelodysplastic syndrome after azacitidine treatment failure. J. Clin. Oncol. 29, 3322–3327. https://doi.org/10.1200/jco.2011.35.8135 (2011).
Trapnell, C., Pachter, L. & Salzberg, S. L. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics (Oxford, England) 25, 1105–1111. https://doi.org/10.1093/bioinformatics/btp120 (2009).
Trapnell, C. et al. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 28, 511–515. https://doi.org/10.1038/nbt.1621 (2010).
Mootha, V. K. et al. PGC-1α-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat. Genet. 34, 267–273. https://doi.org/10.1038/ng1180 (2003).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 102, 15545–15550. https://doi.org/10.1073/pnas.0506580102 (2005).
Acknowledgements
This research was supported by the Basic Science Research Program of the National Research Foundation of Korea funded by the Ministry of Science, ICT and Future Planning (NRF-2015R1A2A2A04002756 and NRF-2018R1A2B2006820). This study was also supported by the National Research Foundation funded by the Korean government (2014R1A1A3054664) a Grant (HI15C3224) from the Korea Health Technology R&D Project through the Korea Health Industry Development Institute (KHIDI), which is funded by the Ministry of Health & Welfare, Republic of Korea, and a Grant from the Bio & Medical Technology Development Program of the National Research Foundation (NRF) funded by the Ministry of Science & Information and Communication Technology, Korea (2019M3E5D3073090).
Author information
Authors and Affiliations
Contributions
Y.K. and T.K. conceived the idea and designed the study. K.K. and S.P collected and analyzed data, and K.K and S.P. wrote the main manuscript text. K.K., S.P and D.R. prepared Figures and Tables included supplementary materials. H.C., H.K., Y.K., and M.K. contributed by providing study material. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kim, K., Park, S., Choi, H. et al. Gene expression signatures associated with sensitivity to azacitidine in myelodysplastic syndromes. Sci Rep 10, 19555 (2020). https://doi.org/10.1038/s41598-020-76510-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-76510-7
This article is cited by
-
ABAT gene expression associated with the sensitivity of hypomethylating agents in myelodysplastic syndrome through CXCR4/mTOR signaling
Cell Death Discovery (2022)
-
Dynamic change in peripheral blood WT1 mRNA levels within three cycles of azacitidine predict treatment response in patients with high-risk myelodysplastic syndromes
Annals of Hematology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.