Abstract
An incomplete understanding of the molecular mechanisms behind impairment of lung pathobiology by COVID-19 complicates its clinical management. In this study, we analyzed the gene expression pattern of cells obtained from biopsies of COVID-19-affected patient and compared to the effects observed in typical SARS-CoV-2 and SARS-CoV-infected cell-lines. We then compared gene expression patterns of COVID-19-affected lung tissues and SARS-CoV-2-infected cell-lines and mapped those to known lung-related molecular networks, including hypoxia induced responses, lung development, respiratory processes, cholesterol biosynthesis and surfactant metabolism; all of which are suspected to be downregulated following SARS-CoV-2 infection based on the observed symptomatic impairments. Network analyses suggest that SARS-CoV-2 infection might lead to acute lung injury in COVID-19 by affecting surfactant proteins and their regulators SPD, SPC, and TTF1 through NSP5 and NSP12; thrombosis regulators PLAT, and EGR1 by ORF8 and NSP12; and mitochondrial NDUFA10, NDUFAF5, and SAMM50 through NSP12. Furthermore, hypoxia response through HIF-1 signaling might also be targeted by SARS-CoV-2 proteins. Drug enrichment analysis of dysregulated genes has allowed us to propose novel therapies, including lung surfactants, respiratory stimulants, sargramostim, and oseltamivir. Our study presents a distinct mechanism of probable virus induced lung damage apart from cytokine storm.
Similar content being viewed by others
Introduction
The recent Coronavirus Disease (COVID-19) pandemic has affected approximately 30 million people across 212 countries, and territories and the number of active cases is still on the rise (on 18 September, 2020)1. At the time of this article, approximately 4% of the infected population has suffered death1, and the fatality rate is continuously increasing due to the lack of detailed knowledge of the molecular mechanism of Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) infection and proper-targeted therapeutic approaches against it.
SARS-CoV-2 is an enveloped RNA virus, which contains single-stranded positive-sense RNA and belongs to the betacoronavirus genus of coronavirus2. It has 11 protein coding genes encompassing its ~ 29.9 Kb genome3. About 90% genomic identity was observed between SARS-CoV-2 and bat-derived SARS-like coronavirus, while SARS-CoV-2 genome is ~ 79% and ~ 50% identical with that of Severe Acute Respiratory Syndrome Coronavirus (SARS-CoV) and Middle East Respiratory Syndrome related Coronavirus (MERS-CoV), respectively2,4,5. Lu et al.2 showed the considerable differences between SARS-CoV-2 and SARS-CoV genomes, including 380 amino acids substitution, ORF8a deletion, ORF8b elongation, and ORF3b truncation2. Despite their identical genomic features, SARS-CoV-2 presents unique clinical and pathophysiological features, such as prolonged incubation period6, and latency inside the host7; which complicate its clinical management.
Based on the clinical exhibitions of COVID-19, most of the mild to critically affected patients show respiratory complications including moderate to severe pneumonia, which can further progress into acute respiratory distress syndrome (ARDS), sepsis, and multiple organ dysfunction (MOD) in severely ill patients8. Most of these clinical symptoms are associated with respiratory system, specifically the lungs9 resulting in the depleted lung functionality. Complications in other systems such as the cardiovascular and nervous systems were also reported10,11. Recently, cases of pulmonary embolism in the lungs of COVID-19 patients have been reported12.
In SARS-CoV and MERS-CoV infections, increased level of pro-inflammatory cytokines were evident13, which in turn increased the activation and recruitment of inflammatory cells into the lung tissues, facilitating acute lung injury14. Similarly, increased levels of many pro-inflammatory cytokines were also detected in moderate-to-critically affected COVID-19 patients15, leading to respiratory failure from ARDS. However, the complex interplays between pro-inflammatory and anti-inflammatory cytokines have not been completely illustrated. Apart from the cytokine storm, other factors such as host innate immunity, autoimmunity against the pulmonary epithelial and endothelial cells, and host genetic and epigenetic factors also play important roles in the pathogenesis of SARS-CoV infection16,17. Moreover, the multifaceted host-virus interactions are also found to be a key player in the pathogenesis reported for other coronavirus infections18.
Previously, transcriptional responses in COVID-19 were experimentally recorded using various in vitro and in vivo approaches such as cell lines, animal models, and SARS-CoV-2-infected lungs19, Nasopharyngeal (NP) swabs20, and bronchoalveolar lavage fluid of COVID-19 patients21. However, SARS-CoV-2-mediated deregulation of the lung transcriptome and its potential implications in pathogenesis of acute lung failure remained elusive. Therefore, designing of prospective therapeutics for the clinical management of the COVID-19 is still in its infancy. To this end, we analyzed publicly available transcriptome data from the lung biopsy of a COVID-19 patient and summarized the probably altered pathways in COVID-19. Furthermore, the probable roles of SARS-CoV-2 in these dysregulations, and resulting acute lung damage, were also investigated.
Results
Antiviral immune responses and organ specific functions are dysregulated in lungs
During respiratory virus infections, many host pathways are fine-tuned to battle against the invading pathogen; on the other side, the infecting viruses also try to hijack and modulate host pathways for immune evasion and survival inside the host22. These complex interactions lead to the disease complexity, causing several critical pathophysiological conditions in host’s respiratory system22. To explore which particular biological processes/pathways are dysregulated in SARS-CoV-2 infection, we first identified the dysregulated-genes in both SARS-CoV and SARS-CoV-2 infections and performed comparative functional enrichment analyses23.
We detected 3031 (2408 upregulated and 623 downregulated), 142 (91 upregulated and 51 downregulated, and 6714 (2476 upregulated and 4238 downregulated) dysregulated genes from SARS-CoV infected 2B4 cells, SARS-CoV-2-infected NHBE cells, and lung biopsy of COVID-19 patient, respectively (Supplementary Fig. 1). We observed a wide array of differentially expressed genes in SARS-CoV-2-infected lung whose expression profiles differed from those recorded in SARS-CoV-2-infected NHBE cells and SARS-CoV-infected 2B4 cells (Supplementary Fig. 1).
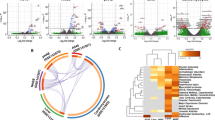
As expected, enrichment analyses revealed biological processes related to antiviral inflammatory responses, and viral processes were overrepresented in all three infection models (Fig. 1A). Several biological processes, such as negative regulation of viral replication, immune system process, response to hypoxia, and heart development were only enriched in the SARS-CoV-2 infection models (Fig. 1A). However, few pivotal processes were uniquely enriched for the dysregulated genes from COVID-19-affected lung, namely viral transcription, adaptive immune response, brain development, lung development, and respiratory gaseous exchange by respiratory system (Fig. 1A).
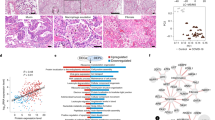
Enrichment analysis and comparison between dysregulated genes in SARS-CoV, SARS-CoV-2 (NHBE cells) and SARS-CoV-2 (Lung biopsy) infections using (A) GOBP41 module, (B) Bioplanet pathway24 module, (C) Reactome pathway28 module, (D) DisGeNet25 module, (E) HumanCyc108 module. Selected significant terms are represented in heatmap. Significance of enrichment in terms of adjusted p-value (< 0.05) is represented in color-coded P-value scale for all heatmaps. Color towards red indicates higher significance and color towards yellow indicates less significance, while grey means non-significant. The column headers are indicating the dysregulation status of the genes used for the enrichment analysis (DE: differentially expressed/Upregulated/Downregulated), while the row labels are pointing the enriched terms.
Similarly, enrichment using the ‘Bioplanet’24 module suggests host antiviral immune responses through various inflammatory cytokine signaling pathways, apoptosis, and interferon-I signaling in all of these infection models (Fig. 1B,C). While, HIF-1 signaling, heart development, asthma, and type-II interferon signaling pathways were found in both SARS-CoV-2 infection data (Fig. 1B, Supplementary Fig. 2). Intriguingly, some pathways, such as disease associated with surfactant metabolism, ROS/RNS production, fatty acyl-CoA biosynthesis, ER-phagosome pathway, and inflammasomes, were only found in the SARS-CoV-2-infected lung (Fig. 1C, Supplementary Fig. 2). Functional enrichment using the ‘DisGeNet’25 module revealed that the dysregulated genes of SARS-CoV and SARS-CoV-2 infections are also involved in other diseases, namely viral disease, lung diseases, asthma, pneumonitis, and hypoxia (Fig. 1D, Supplementary Fig. 2). Interestingly, several cholesterol biosynthesis pathways were found to be dysregulated only in the lung biopsy of COVID-19 patient (Fig. 1E, Supplementary Fig. 2). As cholesterol in lung plays important roles in maintaining normal lung physiology and protection against many diseases26, downregulation of these indicates possible association with lung-related comorbidities of COVID-19 patients.
These enrichment analyses suggest that several pathways related to lung’s function are likely dysregulated directly or indirectly by the infecting SARS-CoV-2 virus. While hunting for more definitive clues on which particular processes are being modulated during SARS-CoV-2 infection, we again performed enrichment analysis with our in-house combined module (Supplementary file 4). This enrichment analysis revealed several important lung-function-related processes only for the dysregulated genes from the COVID-19 patient’s lung. These includued, lung development, pulmonary surfactant metabolism disease or dysfunction, respiratory processes, regulation of respiratory gaseous exchange, and some antiviral responses (Fig. 2, Supplementary Fig. 3). We also exported a list of significantly enriched terms in color-coded heatmaps. Intriguingly, while equating the expression of these enriched genes, we discovered that genes of these key lung-related processes are significantly altered in SARS-CoV-2-infected lungs compared to the SARS-CoV-2-infected NHBE cells and SARS-CoV infection model (Supplementary Fig. 4).
Enrichment analysis and comparison between dysregulated genes and the genes of some selected processes in SARS-CoV, SARS-CoV-2 (NHBE cells) and SARS-CoV-2 (Lung biopsy) infections using combined module. Selected significant terms are represented in heatmap in upper panel. Color schemes are similar as Fig. 1. Lower panel heatmaps presents enriched genes for some selected terms from upper panel enrichment analysis. For individual processes, blue means presence (significantly differentially expressed gene) while grey means absence (not significantly differentially expressed genes for this module for this experimental condition). The column headers are indicating the dysregulation status of the genes used for the enrichment analysis (DE: differentially expressed, Upregulated and Downregulated), while the row labels are pointing the enriched terms.
Genes in lung surfactant metabolism pathway are dysregulated in COVID-19 patient’s lung
Pulmonary surfactant proteins play an important role in maintaining the surface tension necessary for efficient gaseous exchange at the air–liquid interface in the alveoli, and can also modulate functions of lung’s innate immune cells to eliminate pathogens27. Additionally, they have been shown to impede inflammatory responses and the clearance of apoptotic cells in lungs following viral infection27. Taking cue from the results of previous section, it was apparent that surfactant metabolism could be a potential target of viral modulation. Therefore, we sought to decipher the probable routes of SARS-CoV-2-induced dysregulations of this pathway and its downstream signaling. To attain this, we mapped the differentially expressed genes into this pathway using the Reactome pathway browser28. Next, we scrutinized the mechanisms of this pathway to elucidate the probable alterations happening in COVID-19-affected lung.
In normal lung, TTF1-CCDC59 complex can transactivate SFTPB and SFTPC gene expression, which play an important role by regulating the alveolar surface tension29. But in COVID-19-affected lung, TTF1 and SFTPC genes were found to be downregulated, whereas SFTPB was upregulated (Fig. 3). GATA6 transcription factor promotes the transcription of SFTPA gene30 which is involved in immune and inflammatory responses, and lowers the surface tension in the alveoli31. Both GATA6 and SFTPA genes were upregulated in SARS-CoV-2-infected lung, while the GATA6 antagonist LMCD1 was downregulated (Fig. 3). The CSF2RA-CSF2RB complex can bind GM-CSF to induce activation of macrophages32 and helps in the degradation of STFPs in the alveolar macrophages33. We found CFSF2RA downregulated and CSF2RB upregulated in the lung of the COVID-19-affected patient (Fig. 3). Pro-SFTPB and Pro-SFTPC are cleaved by NAPSA, CTSH, and PGA3-5 to produce active SFTPB and SFTPC34,35. Surprisingly, NAPSA was dysregulated in the COVID-19-affected lung (Fig. 3). Overall, the transcription of surfactant genes, production of active surfactant proteins, and their turnover may be dysregulated in the COVID-19 patient’s lung, which could have resulted in the severely lethal disease complications. As these mechanisms are found to be altered upon SARS-CoV-2 infections, the virus may be positively facilitating these anomalies.
Schematic representation of lung surfactant metabolism pathway from Reactome pathway database28. Color towards yellow indicates upregulation and while blue indicates downregulation.
Host proteins which interact with the virus are involved in different respiratory-function-related pathways and diseases
In previously reported human coronavirus infections, SARS and MERS coronaviruses are often found to commandeer host machineries, suppressing host immune responses and other important biological processes for their continued existence within the infected cells36. We performed functional enrichment analyses using previously reported SARS-CoV and SARS-CoV-2 host factor proteins37,38,39 to illuminate the pathways which may be targeted by viral proteins, causing the lung injuries in COVID-19 patients.
As anticipated, the enrichment analyses showed association of both SARS-CoV and SARS-CoV-2 infections with several immune signaling pathways, such as interleukin signaling, interferon signaling, apoptosis, and inflammasomes (Fig. 4A,C). However, this approach also revealed several vital pathways for respiratory function such as HIF-1 signaling, and hypoxic and oxygen homeostasis regulation of HIF-1 alpha (Fig. 4A,C). Using the DisGenNet module25, lung disease, asthma, hyperoxia, respiratory failure, and pulmonary hypertension were found to be enriched (Fig. 4B). These results enlightened that SARS-CoV-2 is likely utilizing its proteins to modulate normal lung’s physiological and immune responses, which we further explored by linking these viral-host protein–protein interactions (PPI) to our previously identified essential lung processes.
Enrichment analysis and comparison between host proteins interacting SARS-CoV proteins (Pfefferle et al.38), SARS-CoV-2 proteins (Srinivasan et al.39) and SARS-CoV-2 proteins (Gordon et al.37) and the genes of some selected processes using (A) Bioplanet pathway24 module, (B) DisGeNet25 module, (C) KEGG pathway109 module. Selected significant terms are represented in heatmap. Color schemes are similar as Figs. 1 and 2.
SARS-CoV-2 proteins and host epigenetic regulators can modulate the functions of lung and other respiratory processes
Results from the previous sections rationalized that both the dysregulated genes in COVID-19-affected lung and SARS-CoV-2 interacting proteins are involved in several respiratory functions. Hence, we produced several functional networks of dysregulated genes, viral protein-host protein interactions, and host epigenetic regulators involved in those processes to gain insight on virus-mediated deregulations and the resulting pathophysiological effects of COVID-19. We have mainly addressed four broad biological processes that can significantly affect COVID-19 patients: response to hypoxia, lung development, respiratory processes, and surfactant metabolism.
Numerous genes in hypoxia response and HIF-1 alpha signaling were abruptly dysregulated in the SARS-CoV-2-infected lung (Supplementary Fig. 5). This included PLAT, a tissue plasminogen activator with profound roles in lung homeostasis, whose aberrant regulation can lead to many lung injuries40. In the PPI map of differentially-expressed genes of the GOBP41 module ‘response to hypoxia’ and SARS-CoV-2 target genes37, this PLAT protein was found to be directly targeted by viral protein ORF8 (Fig. 5). Moreover, several indirect responses from the viral protein interactions were also revealed (Fig. 5). SARS-CoV-2 M protein can target STOM which interacts with SLC2A1. SLC2A1 can also be targeted by host miRNA miR-320a (Fig. 5). SLC2A1 encoding GLUT1 protein is upregulated by hypoxic responses in alveolar epithelial cells42.
Network representing the interactions between proteins in response to hypoxia process (combined module) along with SARS-CoV-2 proteins (Gordon et al.37), and host miRNAs. Hexagon, ellipse, rounded rectangle, octagon represents viral proteins, process related genes, proteins that interacts viral proteins and host miRNAs, respectively. Blunted arrow indicates suppression by miRNAs, dotted arrow pointed with open half-circle indicates downregulated miRNAs failing to modulate host gene, arrowed line pointed with open half-circle indicates targets of viral proteins, and arrowed line pointed with open diamond indicates transcription factors of a gene.
Viral ORF8 interaction with OS9 can modulate EGLN1 and EGLN2, which might disrupt the functions of the EGLN-HIF oxygen sensing system43. KCNMA1 was found to interact with host proteins ATP1B1 and PRKACA that interact with viral M and NSP13 proteins, respectively (Fig. 5). ORF9c may modulate EDNRA indirectly through F2RL1 (Fig. 5) and can alter the vasoconstrictor effect resulting from EDNRA44. PML functions might be altered through the viral N and MOV10 proteins’ interaction. Functions of SLCBA1 may be modulated by NSP13 through PRKACA (Fig. 5). NSP7 can regulate ALDH3A1 which can affect CYP1A1 and NR4A2 (Fig. 5). NSP5 interacting HDAC2 can curb PML and REST which have hypoxia responsive functions45,46 (Fig. 5). NSP12 can modulate a wide range of hypoxia functions related proteins, namely TGFBR2, MECP2, MTHFR, CBFA2T3, EGR1, and ANGPTL4 by interacting with the transcription factor TCF12 (Fig. 5). NSP12 through RIPK1 interactions can dampen the apoptosis regulating function of CFLAR47 (Fig. 5). Apart from this, several host miRNAs could possibly downregulate the expression of some genes, namely- miR-320a, miR-3188, miR-3661, miR-217, miR-421 and miR-429 (Fig. 5). These viral mediated deviations found in the hypoxia responses might be a decisive factor in the lung injury found in COVID-19 patients48.
In the lung development network, viral protein NSP13 is found to indirectly target RC3H2, SOX9, GLI3 through CEP350, PRKACA, PRKAR2A proteins (Fig. 6). While GLI3 can also be modulated through several viral-host interactions, for instance ORF10-RBX1, NSP5-HDAC2, and the M-PSMD8 interactions (Fig. 6). ORF8 can indirectly modulate ADAMTS2, CHI3L1, NOTCH1, TCF12, and FLT4 (Fig. 6). Transcription factor TCF12 is targeted by viral NSP12 protein which can in turn affect transcriptions of PDGFRA, PDGFRB, TGFBR2, ID1, HES1, and LTBP3 genes (Fig. 6). NSP5 can modulate NOTCH1 through HDAC2 (Fig. 6). Moreover, host miRNAs such as miR-630 can downregulate TGFBR2; while miR-206, miR-320a and miR-375 are downregulated and so SOX9, and WYHAZ are overexpressed (Fig. 6). Virus could hamper all these components of different growth factor signaling pathways which are crucial for various lung injury repair mechanisms49,50.
From the respiratory process network, we can delineate that several transcription factors are dysregulated. Many of these are associated with ECSIT that can directly interact with viral ORF9c and indirectly by viral ORF8, and NSP7 (Fig. 7A). Moreover, ECSIT itself is directly targeted by ORF9c (Fig. 7A). NSP12 can modulate SFTPB, SFTPC, SLC5A3, DUSP10, and SAMM50 by targeting TCF12 (Fig. 7A). Also, in this network we have observed suppressive actions of miRNAs- miR-206, miR-217, miR-375 on NR4A2 and NDST1; as well as upregulation of YWHAZ due to the probable downregulation of miR-320a (Fig. 7A). Virus might be dampening the host immune response in the lung by targeting ECSIT51 and by diminishing, respiratory gaseous exchange by negatively modulating the surfactant proteins52.
Surfactant metabolism is found to be modulated not only by viral proteins, but also through irregular host responses (Fig. 7B). Viral proteins NSP12 and NSP5 can target transcription factor TCF12 and epigenetic regulator HDAC2, respectively, which in turn modulate important members of surfactant metabolism process, namely- TTF1, CCDC59, SFTPB, SFTPC, CSF2RA, CSF2RB, NAPSA, SFTPD, and DMBT1 (Fig. 7B). These can also be modulated through CKAP4 which is observed to interact with viral M, NSP2, NSP9, E, and ORF8 proteins (Fig. 7B). Furthermore, we have observed that viral M and S proteins can interact with proteins of this process both directly and indirectly (Supplementary Fig. 6). Host miRNA miR-421 was found to downregulate LMCD1; while miR-137, miR-375, and miR-429 fail to modulate CCDC59 and GATA6 because of their probable inactivation/suppression by differential expression of the regulating TFs (Fig. 7B). As SFTPD and SFTPC are downregulated along with several regulatory partners, their primary function of immunomodulation and efficient air exchange in lung53,54 might be seriously hindered by the viral proteins; which could further lead to pathogenic lung injury55.
SARS-CoV-2 proteins can target several epigenetic factors, such as HDAC2, DNMT1, CUL2, MOV10, RBX1 and TLE1, to alter the above-mentioned processes. Epigenetic factors play a key role in balancing normal lung pathobiology, and anomalies in their regulation can lead to many lung diseases56. For instance, HDAC2 and DNMT1 have significant roles in chronic obstructive pulmonary disease (COPD) progression57,58.
Drug enrichment suggests lung surfactant replacement therapy as a potential treatment option
Our results pointed towards the probable SARS-CoV-2 directed dysregulation in several important respiratory processes and surfactant metabolism pathway in COVID-19-affected lung. We then sought to dig out possible therapies/drugs for the improvement of the lung and respiratory conditions of COVID-19-affected patients. In this regard, we performed drug enrichment analysis for differentially expressed genes present in the combined module terms ‘response to hypoxia’, ‘lung development’, ‘respiratory process’ and ‘surfactant metabolism pathway’ using the WebGestalt functional enrichment tool59. We have found significant enrichment scores for lung surfactants, respiratory stimulants, sargramostim, and oseltamivir (Fig. 8, Supplementary Fig. 7).
Bubble plot of drug enrichment results using WebGestalt tool59. Color towards red indicates higher significance and color towards yellow indicates less significance. Bubble size indicates the enrichment ratio.
Discussion
Though there are a wide variety of symptoms and clinical features seen in COVID-19 patients, almost every mild-to-critically affected patient showed respiratory and breathing complications, ranging from pneumonia to acute lung injury60. Severely ill patients are greatly supported by artificial ventilation, as no therapeutic drugs for mitigating these complications have been discovered. This is due to the lack of understanding on the molecular aspects of lung-related abnormalities in COVID-19. We have identified dysregulated genes related to respiratory and lung-related processes upon the SARS-CoV-2-infection. We have prioritized these pathways and searched for potential therapies/drugs for the treatment which could lessen the resultant effects of gene dysregulation.
Several SARS-CoV-2 entry receptor (ACE2)/entry associated proteins (TMPRSS2, BSG, CTSL, and DPP4) have been previously described61. All of these viral entry associated factors are readily expressed in the lung, except ACE2 (Supplementary Fig. 8A). While analyzing the expression profiling in SARS-CoV-2-infected lung, we discovered a quite unexpected scenario. Astoundingly, in COVID-19-affected lung, ACE2 was upregulated while the genes of the entry associated proteins were downregulated (Supplementary Fig. 8B). More COVID-19 patient data must be analyzed to validate this striking finding.
“Cytokine storm” is a much discussed phenomenon in previously reported pathogenic human coronavirus infections that can be lethal for the host due to the destruction of its own respiratory systems62. Similar responses can also occur following SARS-CoV-2 infection63. We observed that several inflammatory and antiviral responses are readily dysregulated in the SARS-CoV-2-infected lung that could lead to the abnormalities in overall respiratory functions.
Apart from this, our study identified dysregulation in several lung function related vital processes, namely- response to hypoxia, lung development, respiratory process, and surfactant metabolism; alterations in these pathways lead to abnormal lung pathobiology in COVID-19. Networks generated from combining the viral-host protein interactions suggest that viral proteins might be actively involved in these dysregulations along with several host factors, which might also be decontrolled due to the viral infections.
Hypoxic conditions are common in respiratory infections due to reduced oxygenation of the blood64. Several genes involved in hypoxia-induced responses were found to be dysregulated (Fig. 5) in the COVID-19-affected lung. ALDH3A1 can protect airway epithelial cells from destruction65. Though it is upregulated, its functions can be impeded through viral proteins (Fig. 5). CFLAR functions in shutting down apoptotic responses by interacting with RIPK147, but viral interactions with RIPK1 might prevent this (Fig. 5). SARS-CoV-2 can induce the transcription of ANGPTL4 by utilizing TCF12 transcription factor (Fig. 5) which in turn could cause pulmonary tissue damage66. Viral proteins could promote the activity of EGLN1 and EGLN2 (Fig. 5) to suppress the transcription of HIF-induced genes in hypoxia67; on the other hand the regulation through these proteins might be hampered due to viral interactions. Moreover, the constant overexpression of HIF-mediated inflammatory genes could also occur, which could lead to inflammation-induced lung damage. Severe hypoxia-induced responses can occur through the downregulation of MECP268 in SARS-CoV-2 patients (Fig. 5). In hypoxia, REST is induced and can act as a negative regulator of gene expression to maintain a balance between different processes45, that were found to be downregulated and can be targeted through miR-421 in COVID-19 (Fig. 5). GLUT1 (SLC2A1 gene) promotes increased glucose transport into hypoxic cells for its prolonged adaptation during this condition69, but it was found to be downregulated in lung of COVID-19 patient and this can occur through miR-320a (Fig. 5). During hypoxia, TGFβ signaling regulates inflammation and vascular responses70; but attenuated TGFβ expression might lead to increased disease severity (Fig. 5). EGR1 transcription factor can be modulated through viral proteins (Fig. 5) that might prevent the hypoxia induced EGR1 activation of HIF-1 alpha71. This probable route of viral-mediated HIF-1 signaling inhibition could curb the whole hypoxia induced survival responses. Moreover, defective hypoxia response can occur due to the overactivity of EGR1 that can result in anomalous thrombosis72. Also, PLAT is a crucial factor in splitting down clots73; this function might be directly targeted by SARS-CoV-2 ORF8 (Fig. 5). Many COVID-19 patients were reported to have pulmonary embolism and thrombosis74, which could be the effect of altered hypoxic responses.
Similarly, several lung development and respiratory process genes/proteins were found to be dysregulated in the COVID-19 patient’s lung (Fig. 6, 7). SOX9, an important regulator for the recovery from acute lung injury75, can be down-modulated by the virus (Fig. 6). Though the host miRNAs cannot target the YWHAZ pro-survival protein76, they can induce expression surfactant protein A277, which might be indirectly modulated through viral protein NSP7 (Fig. 6). Likewise, viral protein ORF8 can indirectly modulate CHI3L1 (Fig. 6) that normally suppresses lung epithelium injury78. Virus can block TCF12 and inhibit the expression of PDGFRA and PDGFRB (Fig. 6) which could play a role in lung maturation and injury response79. GLI3, which exerts essential role in developing the lung and regulating the innate immune cells80, is downregulated in COVID-19 lungs (Fig. 6). LTBP3 can promote lung alveolarization81, but can be modulated by SARS-CoV-2 protein NSP12 (Fig. 6). Abnormal NOTCH signaling contributes significantly in various lung diseases82, and in SARS-CoV-2 infection NOTCH1 and HES1 are found downregulated (Fig. 6). CD55, a member of the complement system which plays crucial role in host defense in airway epithelium83, can be modulated by viral protein NSP12 (Fig. 7A). CYSLTR1 is upregulated in SARS-CoV-2 infection (Fig. 7A), which is correlated to COPD84. This protein can be modulated by viral ORF9c and ORF3a (Fig. 7A). ECSIT was found to directly interact with viral ORF9c (Fig. 7A) which might stop ECSIT-mediated antiviral innate immune response51. Several mitochondrial genes, for instance NDUFA10, NDUFAF5, and SAMM50 were downregulated in COVID-19-affected lung (Fig. 7A). As mitochondria plays important role in cellular respiration and lung diseases85, aberrant expression of these genes might also lead to lung-related complications. DUSP10 can regulate unusual inflammatory responses upon viral infections86, but in SARS-CoV-2-infected lung this gene was found downregulated, possibly through NSP12-TCF12 interactions (Fig. 7A).
Pulmonary surfactant proteins are lipoproteins that mainly function to lower the alveolar surface tension87 and can elicit immune stimulatory roles against some respiratory pathogens88. Among the surfactant proteins, SP-A and SP-D mainly evoke immune responses, while SP-B and SP-D play roles in maintaining efficient respiratory gas exchange54. Several lung diseases including asthma, acute respiratory distress syndrome (ARDS), COPD are reportedly associated with aberration in the function of surfactant proteins55,89. In SARS-CoV-2-affected lung, production of the surfactant proteins is found to be dysregulated (Fig. 7B). Viral protein NSP5 can recruit HDAC2 and downregulate the expression of TTF1 which is needed for the expression of SP-B and SP-C (Fig. 7B). SP-A and SP-D can be targeted indirectly by several viral proteins (Fig. 7B) which might dampen the production of surfactant proteins in lungs, thus complicating the disease condition. The CSF2RA-CSF2RB complex modulate the surfactant recycling that maintains the overall balance of surfactant content33. We found that CSF2RA is dysregulated in SARS-CoV-2-infected lung, which could lead to abnormal surfactant recycling (Fig. 7B). While performing enrichment analysis, we witnessed a significant downregulation of several cholesterol biosynthesis pathways in COVID-19 patient’s lung cells (Fig. 1E) that could lead to low accumulation of phospholipids in lungs. As phosphatidylcholine (PC) and phosphatidylglycerol (PG) are the principal phospholipids of surfactant proteins90, disrupted production of lipids might make surfactant proteins non-functional.
Considering all these, lung surfactants might be useful in the treatment of COVID-19 patients, as lung surfactant therapies were previously reported to be successful in other respiratory infections and acute lung injury to reduce the lung damage of the patients87, 91,92. Also, other drugs including respiratory stimulants for COPD93, sargramostim for treating pulmonary alveolar proteinosis94, and oseltamivir in curing influenza-related lower respiratory tract complications95 showed potential for improving the lung’s and respiratory system’s overall condition.
From our results, we can suggest that surfactant protein production along with other respiratory responses in lung could be dysregulated in COVID-19. However, further experimental proteomic analyses of this dysregulation are required for the functional implications of this study. Along with the antiviral drugs to mitigate the viral responses, drugs that can improve lung conditions in COVID-19 could also be considered as a treatment option for the patients. Our generated results can be useful for obtaining greater insight on the probable dampened surfactant production upon SARS-CoV-2 infection, and the potential implications of surfactant therapy as a therapeutic agent for COVID-19 treatment.
Methods
Analysis of microarray expression data
Microarray expression data from both SARS-CoV-infected 2B4 cells and uninfected controls (both maintained for 24 h) obtained from Gene Expression Omnibus (GEO) (https://www.ncbi.nlm.nih.gov/geo)96, accession: GSE17400. Raw Affymatrix CEL files were background corrected, normalized using Bioconductor package “affy v1.28.1” using 'rma' algorithm. Quality of microarray experiment (data not shown) was verified by Bioconductor package “arrayQualityMetrics v3.44.0”97. Differentially expressed (DE) between two experimental conditions were called using Bioconductor package Limma98. Probe annotations were converted to genes using in-house python script basing the Ensembl gene model (Biomart 99)99. The highest absolute expression value was considered for the probes that were annotated to the same gene. We have considered the genes to be differentially expressed, which have FDR100 p-value ≤ 0.05 and Log2 fold change value ≥ 0.25 (Supplementary file 1).
Analysis of RNA-seq expression data
Illumina sequenced RNA-seq raw FastQ reads were extracted from GEO database96, accession: GSE147507. This data includes independent biological triplicates of primary human lung epithelium (NHBE) cell lines which were mock treated or infected with SARS-CoV-2 for 24hrs. This data also contains two technical replicate of post-mortem lung biopsy sample of a deceased COVID-19 patient, along with lung biopsy samples of two different healthy persons as control. We have checked the raw sequence quality using FastQC program (v0.11.9)101, and found that the "Per base sequence quality" and "Per sequence quality scores" were high over the threshold for all sequences (data not shown). Mapping of reads was done with TopHat (tophat v2.1.1 with Bowtie v2.4.1)102. Short reads were uniquely aligned allowing at best two mismatches to the human reference genome (GRCh38) as downloaded from UCSC database103. Sequences matched exactly more than once with equal quality were discarded to avoid bias104. The reads that were not mapped to the genome were utilized to map against the transcriptome (junctions mapping). Ensembl gene model105 (version 99, as extracted from UCSC) was used for this mapping. After mapping, we used SubRead package featureCount (v2.21)106 to calculate absolute read abundance (read count, rc) for each transcript/gene associated to the Ensembl genes. For differential expression (DE) analysis we used DESeq2 (v1.26.0) with R (v3.6.2; 2019-07-05)107 that uses a model based on the negative binomial distribution. To avoid false positives, we considered only those transcripts where at least 10 reads are annotated in at least one of the samples used in this study and also applied a minimum Log2 fold change of 0.5 for to be differentially expressed transcripts apart from adjusted p-value cut-off of ≤ 0.05 by FDR. Raw read counts of this experiment are provided in supplementary file 2. To assess the fidelity of the RNA-seq data used in this study and normalization method applied here, we checked the normalized Log2 expression data quality using R/Bioconductor package “arrayQualityMetrics (v3.44.0)”97. From this analysis, in our data no outlier was detected by “Distance between arrays”, “Boxplots”, and “MA plots” methods and replicate samples were clustered together (Supplementary file 3).
Retrieval of the host proteins that interact with SARS-CoV and SARS-CoV-2 proteins
We have obtained the list of human proteins that form high confidence interactions with SARS-CoV and SARS-CoV-2 proteins from previously conducted studies37,38,39 and processed their provided protein names into the associated HGNC official gene symbol.
Functional enrichment analysis
We utilized Gitools (v1.8.4) for enrichment analysis and heatmap generation23. We have utilized the Gene Ontology Biological Processes (GOBP)41, Reactome pathway28, Bioplanet pathways24, HumanCyc database108, DisGeNet25, KEGG pathway109 modules, and a custom in-house built combined module (Supplementary file 4) for the overrepresentation analysis. This combined module was generated from related modules with few genes to a parent term/process which otherwise would have been left out from analysis due to statistical stringency cutoff (module with at least 10 genes are selected) during enrichment analysis. Resulting p-values were adjusted for multiple testing using the Benjamin and Hochberg's method of False Discovery Rate (FDR)100.
Mapping of the differentially expressed genes from SARS-CoV-2-infected lung onto biological pathways
We utilized the Reactome pathway browser28 for the mapping of dysregulated genes of SARS-CoV-2 infection onto different biological pathways. We then focused on the pathways which were found to be enriched for lung-related functions.
Obtaining the transcription factors which can modulate the differential gene expression
We obtained the transcription factors (TFs) which bind to a given differentially expressed gene using a custom TFs module created using ENCODE110, TRRUST111, and ChEA112 databases.
Obtaining human miRNAs target genes
We extracted the experimentally validated target genes of human miRNAs from miRTarBase database113.
Extraction of transcription factors modulate human miRNA expression
We downloaded experimentally validated TFs which bind to miRNA promoters and modules from TransmiR (v2.0) database which provides regulatory relations between TFs and miRNAs114. Because miRNAs play roles in transcriptional regulation, we considered TFs that are expressed (upregulated) and can ‘activate’ or ‘regulate’ miRNAs, or in the absence of TFs (downregulation), those could otherwise ‘suppress’ miRNAs.
Identification of the host epigenetic factors genes
We used the EpiFactors database115 to find human genes related to epigenetic activity.
Construction of biological networks
Construction, visualization and analysis of biological networks with differentially expressed genes, their associated transcription factors, associated human miRNAs, and interacting viral proteins were executed in the Cytoscape software (v3.8.0)116. We used the STRING117 database to extract only the highest confidences (0.9) edges for protein–protein interactions to reduce any false positive connection.
Drug enrichment analysis
We used the WebGestalt tool59 for predicting potential drugs targeting the given differentially expressed genes. We selected the drugs based on FDR (BH) ≤ 0.05100, using both DrugBank118 and GLAD4U119 drug database combined.
Data availability
Publicly available data were utilized. Analyses generated data are deposited as supplementary files.
References
Worldometer. 1–22 (2020).
Lu, R. et al. Genomic characterisation and epidemiology of 2019 novel coronavirus: implications for virus origins and receptor binding. Lancet 395, 565–574 (2020).
NCBI-Gene. (2020)
Jiang, S., Du, L. & Shi, Z. An emerging coronavirus causing pneumonia outbreak in Wuhan, China: calling for developing therapeutic and prophylactic strategies. Emerg. Microb. Infect. 9, 275–277 (2020).
Ren, L.-L. et al. Identification of a novel coronavirus causing severe pneumonia in human: a descriptive study. Chin. Med. J. 133(9), 1015–1024. https://doi.org/10.1097/CM9.0000000000000722 (2020).
Lauer, S. A. et al. The incubation period of coronavirus disease 2019 (COVID-19) from publicly reported confirmed cases: estimation and application. Intern. Med. Ann. https://doi.org/10.7326/m20-0504 (2020)
Lan, L. et al. Positive RT-pcr test results in patients recovered from COVID-19. JAMA 323, 1502–1503. https://doi.org/10.1001/jama.2020.2783 (2020).
Huang, C. et al. Clinical features of patients infected with 2019 novel coronavirus in Wuhan China. Lancet 395, 497–506 (2020).
Galiatsatos, P. What Coronavirus Does to the Lungs, https://www.hopkinsmedicine.org/health/conditions-and-diseases/coronavirus/what-coronavirus-does-to-the-lungs (2020).
Mao, L. N. et al. Neurologic manifestations of hospitalized patients with coronavirus disease 2019 in Wuhan. JAMA Neurol China. https://doi.org/10.1001/jamaneurol.2020.1127 (2020).
Zheng, Y.-Y., Ma, Y.-T., Zhang, J.-Y. & Xie, X. COVID-19 and the cardiovascular system. Nat. Rev. Cardiol. 17, 259–260. https://doi.org/10.1038/s41569-020-0360-5 (2020).
Rotzinger, D. C., Beigelman-Aubry, C., von Garnier, C. & Qanadli, S. D. Pulmonary embolism in patients with COVID-19: time to change the paradigm of computed tomography. Thromb. Res. 190, 58–59. https://doi.org/10.1016/j.thromres.2020.04.011 (2020).
Yoshikawa, T. et al. Dynamic innate immune responses of human bronchial epithelial cells to severe acute respiratory syndrome-associated coronavirus infection. PLoS ONE 5, e8729–e8729. https://doi.org/10.1371/journal.pone.0008729 (2010).
Ye, Q., Wang, B. & Mao, J. The pathogenesis and treatment of the `Cytokine Storm’ in COVID-19. J. Infect. https://doi.org/10.1016/j.jinf.2020.03.037 (2020).
Mehta, P. et al. COVID-19: consider cytokine storm syndromes and immunosuppression. Lancet 395, 1033–1034 (2020).
Gu, J. & Korteweg, C. Pathology and pathogenesis of severe acute respiratory syndrome. Am. J. Pathol. 170, 1136–1147. https://doi.org/10.2353/ajpath.2007.061088 (2007).
Schäfer, A. & Baric, R. S. Epigenetic landscape during coronavirus infection. Pathogens 6, 8. https://doi.org/10.3390/pathogens6010008 (2017).
Fung, S.-Y., Yuen, K.-S., Ye, Z.-W., Chan, C.-P. & Jin, D.-Y. A tug-of-war between severe acute respiratory syndrome coronavirus 2 and host antiviral defence: lessons from other pathogenic viruses. Emerg. Microb. Infect. 9, 558–570. https://doi.org/10.1080/22221751.2020.1736644 (2020).
Blanco-Melo, D. et al. Imbalanced host response to SARS-CoV-2 drives development of COVID-19. Cell https://doi.org/10.1016/j.cell.2020.04.026 (2020).
Butler, D. J. et al. Host, Viral, and Environmental Transcriptome Profiles of the Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2). bioRxiv. https://doi.org/10.1101/2020.04.20.048066 (2020).
Xiong, Y. et al. Transcriptomic characteristics of bronchoalveolar lavage fluid and peripheral blood mononuclear cells in COVID-19 patients. Emerg. Microb. Infect. 9, 761–770. https://doi.org/10.1080/22221751.2020.1747363 (2020).
Manjarrez-Zavala, M. E., Rosete-Olvera, D. P., Gutiérrez-González, L. H., Ocadiz-Delgado, R. & Cabello-Gutiérrez, C. Pathogenesis of viral respiratory infection. In Respiratory Disease and Infection - A New Insight (ed Mahboub, B. H.) (IntechOpen, 2013). https://doi.org/10.5772/54287.
Perez-Llamas, C. & Lopez-Bigas, N. Gitools: analysis and visualisation of genomic data using interactive heat-maps. PLoS ONE 6, e19541 (2011).
Huang, R. et al. The NCATS bioplanet—an integrated platform for exploring the universe of cellular signaling pathways for toxicology, systems biology, and chemical genomics. Front. Pharmacol. https://doi.org/10.3389/fphar.2019.00445 (2019).
Pinero, J. et al. The DisGeNET knowledge platform for disease genomics: 2019 update. Nucleic Acids Res. 48, D845-d855. https://doi.org/10.1093/nar/gkz1021 (2020).
Fessler, M. B. A new frontier in immunometabolism. Cholesterol in lung health and disease. Ann. Am. Thorac. Soc. 14, 399–405. https://doi.org/10.1513/AnnalsATS.201702-136AW (2017).
Glasser, J. R. & Mallampalli, R. K. Surfactant and its role in the pathobiology of pulmonary infection. Microbes Infect. 14, 17–25. https://doi.org/10.1016/j.micinf.2011.08.019 (2012).
Jassal, B. et al. The reactome pathway knowledgebase. Nucleic Acids Res. 48, D498-d503. https://doi.org/10.1093/nar/gkz1031 (2020).
Yang, M. C., Guo, Y., Liu, C. C., Weissler, J. C. & Yang, Y. S. The TTF-1/TAP26 complex differentially modulates surfactant protein-B (SP-B) and -C (SP-C) promoters in lung cells. Biochem. Biophys. Res. Commun. 344, 484–490. https://doi.org/10.1016/j.bbrc.2006.03.158 (2006).
Bruno, M. D., Korfhagen, T. R., Liu, C., Morrisey, E. E. & Whitsett, J. A. GATA-6 activates transcription of surfactant protein A. J. Biol. Chem. 275, 1043–1049. https://doi.org/10.1074/jbc.275.2.1043 (2000).
Silveyra, P. & Floros, J. Genetic complexity of the human surfactant-associated proteins SP-A1 and SP-A2. Gene 531, 126–132. https://doi.org/10.1016/j.gene.2012.09.111 (2013).
Martinez, F. O. & Gordon, S. The M1 and M2 paradigm of macrophage activation: time for reassessment. F1000Prime Rep. 6, 13–13. https://doi.org/10.12703/P6-13 (2014).
Ikegami, M. Surfactant catabolism. Respirology (Carlton, Vic.) 11, 24–27. https://doi.org/10.1111/j.1440-1843.2006.00803.x (2006).
Johansson, J., Jornvall, H. & Curstedt, T. Human surfactant polypeptide SP-B. Disulfide bridges, C-terminal end, and peptide analysis of the airway form. FEBS Lett. 301, 165–167. https://doi.org/10.1016/0014-5793(92)81239-i (1992).
Johansson, J. et al. Hydrophobic 3.7 kDa surfactant polypeptide: structural characterization of the human and bovine forms. FEBS Lett. 232, 61–64. https://doi.org/10.1016/0014-5793(88)80386-7 (1988).
Fung, T. S. & Liu, D. X. Human coronavirus: host-pathogen interaction. Annu. Rev. Microbiol. 73, 529–557. https://doi.org/10.1146/annurev-micro-020518-115759 (2019).
Gordon, D. E. et al. A SARS-CoV-2 protein interaction map reveals targets for drug repurposing. Nature https://doi.org/10.1038/s41586-020-2286-9 (2020).
Pfefferle, S. et al. The SARS-coronavirus-host interactome: identification of cyclophilins as target for pan-coronavirus inhibitors. PLoS Pathog. 7, e1002331–e1002331. https://doi.org/10.1371/journal.ppat.1002331 (2011).
Srinivasan, S. et al. Structural genomics of SARS-CoV-2 indicates evolutionary conserved functional regions of viral proteins. Viruses 12, 360 (2020).
Sisson, T. H. & Simon, R. H. The plasminogen activation system in lung disease. Curr. Drug Targets 8, 1016–1029. https://doi.org/10.2174/138945007781662319 (2007).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. Nat. Genet. 25, 25–29 (2000).
Ouiddir, A., Planès, C., Fernandes, I., VanHesse, A. & Clerici, C. Hypoxia upregulates activity and expression of the glucose transporter GLUT1 in alveolar epithelial cells. Am. J. Respir. Cell Mol. Biol. 21, 710–718. https://doi.org/10.1165/ajrcmb.21.6.3751 (1999).
Ivan, M. & Kaelin, W. G. Jr. The EGLN-HIF O(2)-sensing system: multiple inputs and feedbacks. Mol. Cell 66, 772–779. https://doi.org/10.1016/j.molcel.2017.06.002 (2017).
Calabrò, P. et al. Analysis of endothelin-1 and endothelin-1 receptor A gene polymorphisms in patients with pulmonary arterial hypertension. Intern. Emerg. Med. 7, 425–430. https://doi.org/10.1007/s11739-011-0643-2 (2012).
Cavadas, M. A. S. et al. REST is a hypoxia-responsive transcriptional repressor. Sci. Rep. 6, 31355. https://doi.org/10.1038/srep31355 (2016).
Salsman, J. et al. PML nuclear bodies contribute to the basal expression of the mTOR inhibitor DDIT4. Sci. Rep. 7, 45038. https://doi.org/10.1038/srep45038 (2017).
Faiz, A. et al. Cigarette smoke exposure decreases CFLAR expression in the bronchial epithelium, augmenting susceptibility for lung epithelial cell death and DAMP release. Sci. Rep. 8, 12426. https://doi.org/10.1038/s41598-018-30602-7 (2018).
Shimoda, L. A. & Semenza, G. L. HIF and the lung: role of hypoxia-inducible factors in pulmonary development and disease. Am. J. Respir. Crit. Care Med. 183, 152–156. https://doi.org/10.1164/rccm.201009-1393PP (2011).
Gouveia, L., Betsholtz, C. & Andrae, J. Exploring the effect of PDGF-A deletion in the adult lung: implications in homeostasis and injury. Retrieved from http://urn.kb.se/resolve?urn=urn:nbn:se:uu:diva-347031 (n.d.).
Olajuyin, A. M., Zhang, X. & Ji, H.-L. Alveolar type 2 progenitor cells for lung injury repair. Cell Death Discov. 5, 63. https://doi.org/10.1038/s41420-019-0147-9 (2019).
Lei, C. Q. et al. ECSIT bridges RIG-I-like receptors to VISA in signaling events of innate antiviral responses. J. Innate Immun. 7, 153–164. https://doi.org/10.1159/000365971 (2015).
Veldhuizen, E. J. A. & Haagsman, H. P. Role of pulmonary surfactant components in surface film formation and dynamics. Biochim. Biophys. Acta 1467, 255–270. https://doi.org/10.1016/S0005-2736(00)00256-X (2000).
Mulugeta, S. & Beers, M. F. Surfactant protein C: its unique properties and emerging immunomodulatory role in the lung. Microbes Infect. 8, 2317–2323. https://doi.org/10.1016/j.micinf.2006.04.009 (2006).
Nayak, A., Dodagatta-Marri, E., Tsolaki, A. & Kishore, U. An insight into the diverse roles of surfactant proteins, SP-A and SP-D in innate and adaptive immunity. Front. Immunol. https://doi.org/10.3389/fimmu.2012.00131 (2012).
Sorensen, G. L. Surfactant protein D in respiratory and non-respiratory diseases. Front. Med. (Lausanne) 5, 18–18. https://doi.org/10.3389/fmed.2018.00018 (2018).
Mortaz, E., Masjedi, M. R., Barnes, P. J. & Adcock, I. M. Epigenetics and chromatin remodeling play a role in lung disease. Tanaffos 10, 7–16 (2011).
Barnes, P. J. Role of HDAC2 in the pathophysiology of COPD. Annu. Rev. Physiol. 71, 451–464. https://doi.org/10.1146/annurev.physiol.010908.163257 (2009).
He, X., Chen, L., Chen, Y. & Zeng, H. In A28. Advances in Copd and Asthma A1200-A1200.
Liao, Y., Wang, J., Jaehnig, E. J., Shi, Z. & Zhang, B. WebGestalt 2019: gene set analysis toolkit with revamped UIs and APIs. Nucleic Acids Res. 47, W199–W205 (2019).
Cascella, M., Rajnik, M., Cuomo, A., Dulebohn, S. C. & Di Napoli, R. In StatPearls [Internet] (StatPearls Publishing, 2020).
Sungnak, W. et al. SARS-CoV-2 entry factors are highly expressed in nasal epithelial cells together with innate immune genes. Nat. Med. https://doi.org/10.1038/s41591-020-0868-6 (2020).
Channappanavar, R. & Perlman, S. Pathogenic human coronavirus infections: causes and consequences of cytokine storm and immunopathology. Semin. Immun. 39, 529–539. https://doi.org/10.1007/s00281-017-0629-x (2017).
Pedersen, S. F. & Ho, Y. C. SARS-CoV-2: a storm is raging. J. Clin. Investig. 130, 2202–2205. https://doi.org/10.1172/jci137647 (2020).
Schaible, B., Schaffer, K. & Taylor, C. T. Hypoxia, innate immunity and infection in the lung. Respir. Physiol. Neurobiol. 174, 235–243. https://doi.org/10.1016/j.resp.2010.08.006 (2010).
Jang, J.-H. et al. Aldehyde dehydrogenase 3A1 protects airway epithelial cells from cigarette smoke-induced DNA damage and cytotoxicity. Free Radic. Biol. Med. 68, 80–86. https://doi.org/10.1016/j.freeradbiomed.2013.11.028 (2014).
Li, L. et al. Angiopoietin-like 4 increases pulmonary tissue leakiness and damage during influenza pneumonia. Cell Rep. 10, 654–663. https://doi.org/10.1016/j.celrep.2015.01.011 (2015).
To, K. K. & Huang, L. E. Suppression of hypoxia-inducible factor 1α (HIF-1α) transcriptional activity by the HIF prolyl hydroxylase EGLN1. J. Biol. Chem. 280, 38102–38107 (2005).
Kron, M., Zimmermann, J. L., Dutschmann, M., Funke, F. & Muller, M. Altered responses of MeCP2-deficient mouse brain stem to severe hypoxia. J. Neurophysiol. 105, 3067–3079. https://doi.org/10.1152/jn.00822.2010 (2011).
Zhang, J. Z., Behrooz, A. & Ismail-Beigi, F. Regulation of glucose transport by hypoxia. Am. J. Kidney Dis. 34, 189–202. https://doi.org/10.1016/s0272-6386(99)70131-9 (1999).
Akman, H. O. et al. Response to hypoxia involves transforming growth factor-beta2 and Smad proteins in human endothelial cells. Blood 98, 3324–3331. https://doi.org/10.1182/blood.v98.12.3324 (2001).
Sperandio, S. et al. The transcription factor Egr1 regulates the HIF-1alpha gene during hypoxia. Mol. Carcinog. 48, 38–44. https://doi.org/10.1002/mc.20454 (2009).
Yan, S.-F., Mackman, N., Kisiel, W., Stern, D. M. & Pinsky, D. J. Hypoxia/hypoxemia-induced activation of the procoagulant pathways and the pathogenesis of ischemia-associated thrombosis. Arterioscler. Thromb. Vasc. Biol. 19, 2029–2035. https://doi.org/10.1161/01.ATV.19.9.2029 (1999).
Niedermeyer, J., Meissner, E. & Fabel, H. Thrombolytic therapy in pulmonary embolism. Indications and therapeutic strategies. Zeitschrift fur die gesamte innere Medizin und ihre Grenzgebiete 48, 332–343 (1993).
Sulemane, S., Baltabaeva, A., Barron, A. J., Chester, R. & Rahman-Haley, S. Acute pulmonary embolism in conjunction with intramural right ventricular thrombus in a SARS-CoV-2-positive patient. Eur. Heart J. Cardiovasc. Imaging https://doi.org/10.1093/ehjci/jeaa115 (2020).
Li, L. et al. Sox9 activation is essential for the recovery of lung function after acute lung injury. Cell. Physiol. Biochem. 37, 1113–1122. https://doi.org/10.1159/000430236 (2015).
Kasinski, A., Dong, X., Khuri, F. R., Boss, J. & Fu, H. Transcriptional regulation of YWHAZ, the gene encoding 14–3-3ζ. PLoS ONE 9, e93480. https://doi.org/10.1371/journal.pone.0093480 (2014).
Noutsios, G. T., Ghattas, P., Bennett, S. & Floros, J. 14–3-3 isoforms bind directly exon B of the 5’-UTR of human surfactant protein A2 mRNA. Am. J. Physiol. Lung Cell Mol. Physiol. 309, L147–L157. https://doi.org/10.1152/ajplung.00088.2015 (2015).
Zhou, Y. et al. Chitinase 3-like 1 suppresses injury and promotes fibroproliferative responses in Mammalian lung fibrosis. Sci. Transl. Med. 6, 240276–240276. https://doi.org/10.1126/scitranslmed.3007096 (2014).
Li, R. et al. Pdgfra marks a cellular lineage with distinct contributions to myofibroblasts in lung maturation and injury response. Elife 7, e36865. https://doi.org/10.7554/eLife.36865 (2018).
Matissek, S. J. & Elsawa, S. F. GLI3: a mediator of genetic diseases, development and cancer. Cell Commun. Signal. 18, 54. https://doi.org/10.1186/s12964-020-00540-x (2020).
Saito, A., Horie, M. & Nagase, T. TGF-β signaling in lung health and disease. Int. J. Mol. Sci. 19, 2460. https://doi.org/10.3390/ijms19082460 (2018).
82Xu, K., Moghal, N. & Egan, S. E. In Notch Signaling in Embryology and Cancer (eds Jörg Reichrath & Sandra Reichrath) 89–98 (Springer, New York, 2012).
Kulkarni, H. S., Liszewski, M. K., Brody, S. L. & Atkinson, J. P. The complement system in the airway epithelium: an overlooked host defense mechanism and therapeutic target?. J. Allergy Clin. Immunol. 141, 1582-1586.e1581. https://doi.org/10.1016/j.jaci.2017.11.046 (2018).
Zhu, J. et al. Cysteinyl leukotriene 1 receptor expression associated with bronchial inflammation in severe exacerbations of COPD. Chest 142, 347–357. https://doi.org/10.1378/chest.11-1581 (2012).
Cloonan, S. M. & Choi, A. M. K. Mitochondria in lung disease. J. Clin. Investig. 126, 809–820. https://doi.org/10.1172/JCI81113 (2016).
Manley, G. C. A., Stokes, C. A., Marsh, E. K., Sabroe, I. & Parker, L. C. DUSP10 negatively regulates the inflammatory response to rhinovirus through interleukin-1β signaling. J. Virol. 93, e01659-e11618. https://doi.org/10.1128/JVI.01659-18 (2019).
Walther, F. J., Gordon, L. M. & Waring, A. J. Advances in synthetic lung surfactant protein technology. Expert Rev. Respir. Med. 13, 499–501. https://doi.org/10.1080/17476348.2019.1589372 (2019).
Wright, J. R. Immunoregulatory functions of surfactant proteins. Nat. Rev. Immunol. 5, 58–68. https://doi.org/10.1038/nri1528 (2005).
Han, S. & Mallampalli, R. K. The role of surfactant in lung disease and host defense against pulmonary infections. Ann. Am. Thorac. Soc. 12, 765–774. https://doi.org/10.1513/AnnalsATS.201411-507FR (2015).
Yamano, G. et al. ABCA3 is a lamellar body membrane protein in human lung alveolar type II cells. FEBS Lett. 508, 221–225. https://doi.org/10.1016/s0014-5793(01)03056-3 (2001).
Skokic, F. et al. Surfactant replacement therapy in influenza A H1N1. Pediatr. Infect. Dis. J. 29, 387. https://doi.org/10.1097/INF.0b013e3181cf2eaa (2010).
Czyzewski, A. M. et al. Effective in vivo treatment of acute lung injury with helical, amphipathic peptoid mimics of pulmonary surfactant proteins. Sci. Rep. 8, 6795. https://doi.org/10.1038/s41598-018-25009-3 (2018).
Bales, M. J. & Timpe, E. M. Respiratory stimulant use in chronic obstructive pulmonary disease. Ann. Pharmacother. 38, 1722–1725. https://doi.org/10.1345/aph.1E039 (2004).
Tazawa, R. et al. Inhaled GM-CSF for pulmonary alveolar proteinosis. N. Engl. J. Med. 381, 923–932. https://doi.org/10.1056/NEJMoa1816216 (2019).
Kaiser, L. et al. Impact of oseltamivir treatment on influenza-related lower respiratory tract complications and hospitalizations. Arch. Intern. Med. 163, 1667–1672. https://doi.org/10.1001/archinte.163.14.1667 (2003).
Barrett, T. et al. NCBI GEO: archive for functional genomics data sets—update. Nucleic Acids Res. 41, D991–D995. https://doi.org/10.1093/nar/gks1193 (2012).
Kauffmann, A., Gentleman, R. & Huber, W. Array quality metrics—a bioconductor package for quality assessment of microarray data. Bioinformatics 25, 415–416 (2009).
Smyth, G. K. limma: linear models for microarray data. In Bioinformatics and Computational Biology Solutions Using R and Bioconductor. Statistics for Biology and Health. (eds Gentleman, R., Carey, V. J., Huber, W., Irizarry, R. A., Dudoit, S.) https://doi.org/10.1007/0-387-29362-0_23 (Springer, New York, NY, 2005).
Flicek, P. et al. Ensembl 2008. Nucleic Acids Res. 36, D707–D714 (2007).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. Roy. Stat. Soc.: Ser. B (Methodol.) 57, 289–300 (1995).
Andrews, S. Babraham Bioinformatics (Babraham Institute, Cambridge, 2010).
Trapnell, C., Pachter, L. & Salzberg, S. L. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25, 1105–1111. https://doi.org/10.1093/bioinformatics/btp120 (2009).
Lander, E. S. et al. Initial sequencing and analysis of the human genome. Nature 409, 860–921. https://doi.org/10.1038/35057062 (2001).
Hansen, K. D., Brenner, S. E. & Dudoit, S. Biases in Illumina transcriptome sequencing caused by random hexamer priming. Nucleic Acids Res. 38, e131. https://doi.org/10.1093/nar/gkq224 (2010).
Hubbard, T. J. P. et al. Ensembl 2007. Nucleic Acids Res. 35, D610–D617 (2007).
Liao, Y., Smyth, G. K. & Shi, W. The Subread aligner: fast, accurate and scalable read mapping by seed-and-vote. Nucleic Acids Res. 41, e108. https://doi.org/10.1093/nar/gkt214 (2013).
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106–R106. https://doi.org/10.1186/gb-2010-11-10-r106 (2010).
Romero, P. et al. Computational prediction of human metabolic pathways from the complete human genome. Genome Biol. 6, R2. https://doi.org/10.1186/gb-2004-6-1-r2 (2004).
Kanehisa, M. & Goto, S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Consortium, E. P. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74. https://doi.org/10.1038/nature11247 (2012).
Han, H. et al. TRRUST v2: an expanded reference database of human and mouse transcriptional regulatory interactions. Nucleic Acids Res. 46, D380-d386. https://doi.org/10.1093/nar/gkx1013 (2018).
Lachmann, A. et al. ChEA: transcription factor regulation inferred from integrating genome-wide ChIP-X experiments. Bioinformatics (Oxford, England) 26, 2438–2444. https://doi.org/10.1093/bioinformatics/btq466 (2010).
Huang, H.-Y. et al. miRTarBase 2020: updates to the experimentally validated microRNA-target interaction database. Nucleic Acids Res. 48, D148–D154. https://doi.org/10.1093/nar/gkz896 (2019).
Tong, Z., Cui, Q., Wang, J. & Zhou, Y. TransmiR v2.0: an updated transcription factor-microRNA regulation database. Nucleic Acids Res. 47, 253–258. https://doi.org/10.1093/nar/gky1023 (2019).
Medvedeva, Y. A. et al. EpiFactors: a comprehensive database of human epigenetic factors and complexes. Database 2015, doi:https://doi.org/10.1093/database/bav067 (2015).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504. https://doi.org/10.1101/gr.1239303 (2003).
Szklarczyk, D. et al. STRING v11: protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 47, D607–D613. https://doi.org/10.1093/nar/gky1131 (2019).
Wishart, D. S. et al. DrugBank: a comprehensive resource for in silico drug discovery and exploration. Nucleic Acids Res. 34, D668-672. https://doi.org/10.1093/nar/gkj067 (2006).
Jourquin, J., Duncan, D., Shi, Z. & Zhang, B. GLAD4U: deriving and prioritizing gene lists from PubMed literature. BMC Genom. 13(Suppl 8), S20–S20. https://doi.org/10.1186/1471-2164-13-S8-S20 (2012).
Acknowledgements
We are grateful to Alexandra Elaine Rader from Department of Biochemistry and Molecular Genetics, University of Illinois at Chicago, USA for critically reading and proofreading of the manuscript.
Funding
This project was not associated with any internal or external source of funding.
Author information
Authors and Affiliations
Contributions
A.B.M.M.K.I. conceived the project, designed the workflow and performed the analyses. Both authors interpreted the results and wrote the manuscript. Both authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Islam, A.B.M.M.K., Khan, M.AAK. Lung transcriptome of a COVID-19 patient and systems biology predictions suggest impaired surfactant production which may be druggable by surfactant therapy. Sci Rep 10, 19395 (2020). https://doi.org/10.1038/s41598-020-76404-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-76404-8
This article is cited by
-
SCARLET (Supplemental Citicoline Administration to Reduce Lung injury Efficacy Trial): study protocol for a single-site, double-blinded, placebo-controlled, and randomized Phase 1/2 trial of i.v. citicoline (CDP-choline) in hospitalized SARS CoV-2-infected patients with hypoxemic acute respiratory failure
Trials (2024)
-
Superior anti-pulmonary viral potential of Natrialba sp. M6-producing surfactin and C50 carotenoid pigment with unveiling its action modes
Virology Journal (2023)
-
A randomized controlled trial of nebulized surfactant for the treatment of severe COVID-19 in adults (COVSurf trial)
Scientific Reports (2023)
-
Bioinformatics and system biology approach to identify potential common pathogenesis for COVID-19 infection and osteoarthritis
Scientific Reports (2023)
-
SARS CoV-2 spike protein variants exploit DC-SIGN/DC-SIGNR receptor for evolution and severity: an in-silico insight
VirusDisease (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.