Abstract
Inborn errors of metabolism are genetic conditions that can disrupt intermediary metabolic pathways and cause defective absorption and metabolism of dietary nutrients. In an Australian Kelpie breeding population, 17 puppies presented with intestinal lipid malabsorption. Juvenile dogs exhibited stunted postnatal growth, steatorrhea, abdominal distension and a wiry coat. Using genome-wide association analysis, an associated locus on CFA28 (Praw = 2.87E−06) was discovered and validated in a closely related population (Praw = 1.75E−45). A 103.3 kb deletion NC_006610.3CFA28:g.23380074_23483377del, containing genes Acyl-CoA Synthetase Long Chain Family Member 5 (ACSL5) and Zinc Finger DHHC-Type Containing 6 (ZDHHC6), was characterised using whole transcriptomic data. Whole transcriptomic sequencing revealed no expression of ACSL5 and disrupted splicing of ZDHHC6 in jejunal tissue of affected Kelpies. The ACSL5 gene plays a key role in long chain fatty acid absorption, a phenotype similar to that of our affected Kelpies has been observed in a knockout mouse model. A PCR-based diagnostic test was developed and confirmed fully penetrant autosomal recessive mode of inheritance. We conclude the structural variant causing a deletion of the ACSL5 gene is the most likely cause for intestinal lipid malabsorption in the Australian Kelpie.
Similar content being viewed by others
Introduction
Long chain fatty acids (LCFA) are the most abundant fats in mammals and play a key role in the canine (Canis lupus familiaris) diet. Pancreatic lipases are largely responsible for the hydrolysis of triglycerides into glycerol and fatty acids, which are absorbed across the brush border of jejunal enterocytes. Activation of fatty acids is the first step in intracellular metabolism of LCFA. The process involves the conjugation of fatty acids with coenzyme-A (CoA) and is catalysed by a group of enzymes called Acyl-CoA synthetases (ACS)1,2,3. Thirteen homologous ACS genes that activate LCFA have been annotated in mammals and cluster in three different gene families: acyl-CoA synthetase long chain (ACSL), acyl-CoA synthetase bubblegum (ACSBG) and fatty acid transport proteins (FATP)1,2,3. The full extent of each gene on normal intestinal absorption of LCFA is unknown. Animal models with heritable phenotypes of intestinal lipid malabsorption provide an opportunity to elucidate this.
The Kelpie is an iconic dog (Canis lupus familiaris) breed established in the late nineteenth century for its natural working ability and resilience in the extreme weather conditions of Australia4. Since inception, the breed has become divided into two separate breeding populations maintained by different pedigree registries5. Dogs selected primarily for strong working ability are known as the Australian Working Kelpie (AWK). The Australian Kelpie (AK) is selected according to a conformation breed standard and is registered by the Australian National Kennel Council. The Federation Cytological International recognizes both breed varieties enabling the populations to co-mingle but, in Australia, the two are maintained as separate breeding populations and are genetically distinct; most notably in genes influencing morphology and behaviour4,5.
We describe an inherited intestinal lipid malabsorption (OMIA 002226-9615) in the AK. Unlike cases of similar disorder hereditary selective ileal cobalamin malabsorption6,7,8,9,10,11, AK show no signs of lethargy and have clear evidence of fat in faeces (steatorrhea). Based on the observed familial segregation the observed phenotype is a fully penetrant autosomal recessive inherited metabolic disorder. The aim of the current study is to provide insight into the molecular genetic aetiology of intestinal lipid malabsorption in the AK.
Materials and methods
Ethics
The research described conforms to the recommendations from the Australian Code for the Care and Use of Animals for Scientific Purposes. Animal ethics approval was granted by the Animal Ethics Committee at the University of Sydney (approval numbers 2015/902 and 2018/1449).
Animal selection and phenotype selection
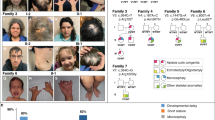
Related juvenile AK dogs were observed to exhibit stunted postnatal growth and intestinal lipid malabsorption. Affected individuals remain a third to one half the size of their littermates during development and mature so that adult dogs are smaller in stature and exhibit persistent intolerance to a fatty diet. Starting in 2011, 17 of 319 puppies, from 45 litters, were born at one Australian kennel presenting with identical clinical features (Fig. 1). As neonates, affected puppies are indistinguishable from their littermates, but rapidly show clinical signs of polyphagia, failure to thrive, stunted growth (around one-third to one-half of the size of their siblings—Fig. 2a), yellowish poorly digested loose and pulpy faeces (Fig. 2b), increased faecal volume, and frequent defecation. Once affected puppies are transferred to a solid diet with digestive enzyme supplementation, faecal consistency normalises. From around six months of age, most affected Kelpies appear to outgrow the characteristic clinical presentation. However, the dogs remain smaller in stature than their siblings, consistently produce more voluminous faeces than age-matched dogs, and their intolerance to high-fat foods persists throughout their lives.
Simplified pedigree of an AK family showing links among affected individuals with hereditary intestinal lipid malabsorption. Females and males are indicated by circles and squares, respectively. Filled symbols indicate affected samples, half-filled symbols represent carriers of the disease allele based on autosomal recessive inheritance. Offspring from a single litter are represented by a line descending from a horizontal connection between parent symbols. A triangle has been used to designate multiple samples (N) from a single litter that are not affected or suspected to be carriers based on recessive inheritance. Litters that included zero affected samples have not been included. Affected samples highlighted in blue were included in the study. Diagnostic testing found all samples to be homozygous for the disease-associated variant.
Side by side comparisons of affected Kelpie and unaffected littermate size and faecal matter. (a) Affected Kelpie at 10 weeks (right) with his littermate. Size and musculature of the affected pup is in stark contrast. (b) Pale poorly digested faeces from an affected Kelpie (right) in comparison with an unaffected littermate (left). There is a significant difference in colour and consistency between the two samples.
This study involved 265 Kelpies. Samples were made up of 35 AK (10 cases and 25 controls), 225 AWK (225 controls), and 5 international Kelpies (one case and four controls). Cases in this study represent dogs that adhered to the described clinical presentation. Cases were easily recognised through signs of ill thrift, faecal appearance (steatorrhea), and stunted growth when compared to littermates. Seven cases from the originally described kennel have been included in this study. Two samples from separate kennels were reported as dams of affected pups. They were included as control samples and treated as obligate carriers when observing results.
Biological samples were collected as whole blood in EDTA tubes or buccal cells using cheek swabs. Genomic DNA was extracted using the PureLink Genomic DNA Mini Kit (Invitrogen, Carlsbad, CA, USA) or submitted as EDTA blood to the genotyping service provider on Whatman Flinders Technology Associates (FTA) cards, supplied by the genotyping service. Genotyping array data for 255 samples was obtained from the CanineHD BeadChip (Illumina, San Diego, CA, USA) by Neogen (Lincoln, NE, USA).
A full post-mortem was conducted at the Veterinary Pathology Diagnostic Services (University of Sydney, Camperdown, NSW, Australia) on a 17-week-old affected AK pup that was euthanized with approval by the owner on welfare grounds. A thorough examination was conducted on tissue of the lung, spleen, liver, heart, major cardiac vessels, lymph nodes, thyroid gland, kidney, bone marrow, pancreas, small intestine (duodenum, jejunum and ileum), brain and spinal cord.
Genome-wide association study (GWAS)
To detect and validate signals associated with malabsorption in the Kelpie population two case–control GWAS were performed using Plink 1.9 (--assoc)12. Quality control of genotypic data was conducted on 25 AK, five internationally bred Kelpies, and 225 AWK. Single Nucleotide variants (SNVs) were excluded if they exhibited a call rate of less than 90% (--geno) or a low minor allele frequency < 10% (--maf). Pairwise identity by decent was calculated (--genome) to detect and remove duplicated or highly related individuals. Population stratification was visualised using a multidimensional scaling (MDS) plot with two dimensions (--mds). One sample from each pair with a pairwise identity by decent > 0.7 was excluded. This was done to control for inflation resulting from cryptic relatedness and population stratification. Population stratification in the preliminary GWAS was determined by the genomic inflation factor based on the median Chi-squared statistic. The primary GWAS was conducted using 30 Kelpies, including 25 AK and five internationally bred Kelpies. Both groups show evidence of carrying the studied trait; reflected in our dataset. To control for the testing of multiple hypotheses, genome-wide significant and suggestive thresholds were Bonferroni-corrected, 5 × 10–7 (Bonferroni cut-off of α = 0.05, n = 99,326) and 1 × 10–5 (Bonferroni cut-off of α = 1.0, n = 99,326), respectively. Reported P-values are chi-square allelic test P-values as calculated in Plink. The 200 most associated markers from the unstratified preliminary GWAS were taken forward to a second analysis that added 225 control dogs from the closely related population of AWK.
Confirmation of deletion by polymerase chain reaction (PCR)
A large segment of consecutive uncalled array markers was observed only in cases suggesting the presence of a large deletion in these animals. Primers designed using primer313,14,15 were used to detect the presence of the deletion. The novel deletion was confirmed through amplification of the last coding exons in impacted genes by PCR. Where no amplification was observed, to gauge the size of the deletion, further primers were designed to amplify the preceding exon. Alternatively, where amplification was witnessed, we designed primers in the gene’s untranslated region (UTR). A total of seven primers were designed (Table S1). PCR was carried out in total volume of 20 μl using AmpliTaq Gold 360 Master Mix (Applied Biosystems, Foster City, CA, USA) according to the manufacturer’s protocol. Fragments were evaluated on a 1.5% agarose gel. Briefly, the PCR conditions were heat activation for 10 min (mins) at 95 degrees Celsius (°C); and 30 cycles of 30 s (s) at 95 °C, 58 °C and 72 °C for denaturation, annealing and extension respectively. The process concluded with a final elongation step at 72 °C for 10 min.
RNA sequencing, alignment and variant detection
In order to gauge if mapped candidate genes were influencing the observed phenotype, whole transcriptomic sequencing was conducted on whole tissue of the jejunum collected during post-mortem. Using Invitrogen TRIzol Reagent, RNA was extracted according to the manufacturer’s protocol. Total RNA sequencing (RNAseq) was performed on Illumina NovaSeq S1 using the TrueSeq Stranded RNA RiboZero Gold (h/m/r) kit at the Ramaciotti Centre for Genomics (University of New South Wales, Kensington, NSW, Australia). A total of 142,658,896, 100 base pair (bp) paired end reads were generated. Tissue matched transcriptomic sequence data for the Labrador retriever JEJUNUM_LABR (Accession identifier: SRR3727723) were obtained from the Sequence Read Archive (SRA) in Genbank (https://www.ncbi.nlm.nih.gov/sra/).
Quality control for the raw paired-end reads was performed with FastQC v0.11.8 (https://www.bioinformatics.babraham.ac.uk/projects/fastqc/) and visualised using MultiQC16. Raw RNAseq reads were mapped to the canine reference genome (CanFam3.1) using STAR aligner v2.7.0e basic options17. Reads surrounding the candidate region were visualised and extracted for a genome-guided de novo transcriptome assembly using Trinity v2.8.318. The distribution of read data was calculated across known gene features using RSeQC v3.0.119.
STAR aligned bam files and Trinity constructed fasta sequences were visualised in Integrative Genomics Viewer v2.8.220. Variants in the affected Kelpie transcript were compared with those of the tissue matched Labrador retriever. Alternate transcript splicing was visualised using a sashimi plot created using ggsashimi21. The minimum read coverage at splice junctions was set to 15 reads to reduce background hybridisation signals.
Multiplex PCR assay design for deletion
Custom primers were designed to capture a disease-associated variant identified in the sequenced AK using primer3. A three-primer multiplex PCR was designed to detect the structural variant (Table S1). The multiplex PCR includes a forward primer, 37 bp upstream of the disease-associated variant, and two reverse primers, one 140 bp downstream from the start of the variant and another ~ 103.7 kb downstream of the forward primer. PCR was carried out in total volume of 20 μl as previously described. A total of 19 Kelpie samples from affected populations were assessed using this method, including nine cases and ten controls. This encompassed two cases and eight controls also utilised in the GWAS analysis. Six of the controls were known to come from families that have produced affected puppies.
Equipment and settings
All images have been formatted for publishing using Adobe Photoshop 2020 (v21.1.1). Images that have been cropped have been done so to improve clarity and conciseness. As such, all images correctly represent the original data. If an electrophoretic gel images has been cropped, it is stated in the figure legend and an original image has been provided in “Supplementary materials”.
Results
Post-mortem results
Post-mortem was conducted on a young, 17-week old, female Kelpie. The dog was received in excellent post-mortem condition immediately following euthanasia. The affected Kelpie pup showed signs of malnutrition including decreased body condition (body condition score 2/5), atrophied musculature and depleted subcutaneous adipose tissue stores. Histological examination of the small intestine showed evidence of mild non-specific chronic enteritis including focal ileal ulceration, rare crypt abscesses in the ileum and colon, and possible crypt fusion in the jejunum.
Genome-wide association study (GWAS)
After frequency and genotype pruning, 99,326 SNVs remained in the analysis. Three cases and 27 control Kelpies were available for the primary analysis. Of these, four controls and one case were bred outside Australia. By MDS the AK and International populations clustered closely and so were treated analytically as one population (Fig. S1a). When AWK were included in the MDS the principal Kelpie population and AWK clustered separately (Fig. S1b).
A preliminary GWAS was performed in the closely clustered Kelpie populations with affected samples (AK and internationally bred Kelpies). The quantile–quantile plot shows limited inflation and the genomic inflation factor was 1.23 (Fig. 3a). GWAS revealed a suggestive association with intestinal lipid malabsorption on canine chromosome 28 (CFA28) (best Praw = 2.87E−6) (Fig. 3b). Six SNVs within a three megabase (Mb) region (28:24,521,377–26,556,336; 2.03 Mb) passed the suggestive genome-wide significance threshold and were in strong linkage with the index SNV (r2 > 0.93). In the validation analysis, 225 AWK controls were added to the leading dataset. When analysing the top 200 associated SNVs in the primary GWAS in the extended cohort, 52 SNVs passed genome-wide significance (Fig. 3c). Of these SNVs, 21 (40.3%) were located on CFA28: 20 that clustered within a four Mb region (28: 24,030,090–27,194,500; 3.16 Mb) including 16 (30.7%) that matched the expected GT frequency for a recessively inherited trait (Table S2). The top SNV from the preliminary analysis remained the strongest in the validation set (best Praw = 1.75E−45).
QQ-plot and Manhattan plots from intestinal lipid malabsorption case–control GWAS. (a) QQ-plot of Data shown in the Manhattan plot. The observed distribution of the test statistic closely follows the expected. (b) Manhattan plot showing the negative log P-value between individual markers. 6 SNVs highlighted in red on CFA28 passed a suggestive threshold (1 × 10–5, dashed line). No SNVs achieved a Genome wide significant score (5 × 10–7, solid line). The top 200 SNVs from this association were were taken forward to a second analysis. (c) Manhattan plot demonstrating genotype association to intestinal lipid malabsorption. 52 SNVs highlighted in red pass the significance threshold of P < 5 × 10–7 (solid line), while 13 SNVs in yellow passed a suggestive threshold of P < 1 × 10–5 (dashed line). The most significant region of interest is noted on CFA28, where the strongest SNV from the preliminary analysis persisted in the validation set (28:24521377 best Praw = 1.75E−45).
Confirmation of deletion by PCR
Within the associated locus, we identified a region of nine consecutive SNVs spanning marker CFA28:23,370,822 (BICF2P674000) to CFA28:23,493,334 (BICF2P338375) 122.5 kb, where all but two SNVs were consistently uncalled in cases but not controls (Table S3), suggesting the presence of a deleted segment. Within the putatively deleted segment, three genes and one pseudogene were identified, being Tectorin beta (TECTB), Acyl-CoA Synthetase Long Chain Family Member 5 (ACSL5), Zinc Finger DHHC Domain-Containing Protein 6 (ZDHHC6) and pseudogene Guanylate Cyclase 2G (GUCY2GP). The gene ACSL5 represented a strong regional candidate for disease. The orientation of the genes within the region of uncalled markers were positioned so that the last coding exon of each gene aligned with the edges of the putative deletion (Fig. S2a). Using seven primer pairs, we confirmed the presence of a deletion in the affected Kelpies between 101.6 kb and 105.2 kb (Fig. S2b). The PCR confirmed a complete loss of GUCY2GP and ACSL5 and partial loss of ZDHHC6. RNAseq data was used to validate this result.
Variant detection and RNA expression
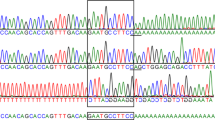
RNAseq data were inspected in Integrative Genomics Viewer. Read distributions indicated RNAseq data had underlying DNA contamination with an equal portion of reads aligning to introns or intergenic regions compared to exons, 34.9% and 38.1% respectively. DNA from the AK with intestinal lipid malabsorption harboured a 103.3 kb deletion, NC_006610.3CFA28:g.23380074_23483377del (CanFam 3.1; Fig. S3), involving the complete loss of ACSL5, pseudogene GUCY2GP and omitting exons 7–10 of ZDHHC6 (Fig. 4). RNAseq data demonstrated no detectable expression of GUCY2GP, ACSL5 and ZDHHC6 exons beyond the breakpoint of the observed deletion. A further gene, TECTB, located outside the deleted region had no observable expression compared with that of the Labrador retriever jejunum. Gene expression of ZDHHC6 and ACSL5 in the control jejunum was consistent with the reference transcript. GUCY2GP and TECTB were not expressed in either case or control. In the AK, expression of novel exons as a result of cryptic splicing were observed 148.4 kb downstream from the ZDHHC6 gene. The alternate splicing event was captured by 152 reads. A consensus sequence was produced using a genome guided de-novo assembly with Trinity (Data S1).
Sashimi plot of RNAseq data for CanFam3 genomic coordinates CFA28:23320000-23500000. The coverage for each alignment track is plotted as a bar graph, the Y axis represents read counts. Arcs are supported exon junctions and reads split across the junction (junction depth). Below the plots are the gene annotations for corresponding genomic coordinates. The figure illustrates RNAseq data from the jejunum of two samples; A case sample (AK Australian Kelpie) and control (LR Labrador retriever). Underlying DNA contamination can be seen in the AK highlighted by the low read count across the genomic region. A 103.3 kb deletion is seen in the AK, illustrated with a transparent box. The gap in the AK includes GUCY2GP, ACSL5, ZDHHC6 and a Long non-coding RNA (lncRNA). In the AK, expression of novel exons can be seen 148.4 kb downstream from the ZDHHC6 gene, the junction is supported by 152 reads. A consensus sequence produced using a genome guided de-novo assembly with Trinity is included in the gene track as ZDHHC6_AK_DEL.
Multiplex PCR assay for deletion
A multiplex PCR test was customised to implicate the associated variant in affected individuals as well as detect carriers of the variant through allele specific amplification. Primer 1 and 3 were designed to amplify the region spanning the disease-associated variant. The variant removes 103,303 bp between the primers and results in a 414 bp fragment in carrier and affected individuals. Primer 2 anneals shortly downstream from the start of the deleted region so that wild type dogs produce a 177 bp fragment while dogs homozygous for the variant show no amplification. Animals that are heterozygous for the disease-associated variant produce both fragments. Of 19 samples tested, nine were homozygous for the deletion, all of which exhibited signs of disease (Fig. S4). In the controls, six samples were homozygous wild type and four were heterozygous for the variant. Dogs heterozygous for the variant were asymptomatic but came from families known to produce offspring with the disease phenotype.
Discussion
Inborn errors of metabolism (IEM) are genetic disorders resulting from defects in biochemical pathways that can have a profound effect on an animal’s overall health22,23. IEM affecting intermediary metabolic pathways are often recognised through clinical signs such as failure to thrive, hypotonia and functional decompensation22,23. Increased prevalence of IEM among specific breeds has previously been observed22,23,24. Frequently reported metabolic disorders clinically similar to intestinal lipid malabsorption are hereditary selective ileal cobalamin malabsorption and exocrine pancreatic insufficiency. Both are IEM that present with failure to thrive and persistent diarrhea11,25,26, however the AK presents earlier (before six weeks of age), show no signs of lethargy and have clear evidence of fat in faeces (steatorrhea). Here we present an IEM affecting lipid absorption in the AK resulting from the deletion of ACSL5 and partial loss of ZDHHC6.
Characterisation of the genetic factors associated with IEM is of strong interest for improving canine welfare and improving our understanding of the genomic control of metabolism. Genes influencing the phenotype described in this study, ACSL5 and ZDHHC6, have not been previously implicated in naturally occurring disease models. Long chain acyl-CoA synthetases are major enzymes in fatty acid metabolism27,28,29,30,31,32. In human and rodent studies, variation in the ACSL gene family are often associated with diet induced metabolic and body composition phenotypes27,31,33,34,35,36,37,38. ACSL5, essential for lipid metabolism and fat deposition in carnivores39, is a principal candidate for the observed phenotype in the AK. ACSL genes have already been implicated in canine body composition phenotypes, with variation in ACSL4 associated with heavy weight dogs40.
The clinical phenotype associated with absent expression of ACSL5 in the jejunal tissue of affected AK puppy is consistent with a knockout (KO) mouse model, including delayed fat absorption and a reduced fat mass27. KO mice exhibited additional increased lean mass and energy expenditure, as well as improved insulin sensitivity; traits not observed or tested in our cases. The results of the mouse KO study contradicted an earlier ACSL5 knockout study, which showed little effect on long-term dietary LCFA absorption and weight gain, likely compensated by residual ACSL activity41. Long chain fatty acid absorption occurs largely through the jejunum where LCFA are absorbed across the brush border of jejunal enterocytes. ACSL5 is expressed in brown adipose tissue, small intestine, liver27,28,42,43,44 and is the primary activator of dietary LCFA in the jejunum41. Expression, synthesis and activity of ACSL5 is connected to the state of villus architecture, epithelial homeostasis and enterocyte apoptosis45,46,47. The relatively improved health status of affected AK at maturity may imply an important role of ACSL5 during early development. The extreme effects identified in immature AK may be partially offset by other ACSL genes as they reach full size or may be linked with a transition to a solid diet.
Following absorption of LCFA in enterocytes, they undergo re-esterification before transportation and storage is possible. Previous research in rodent studies has implicated ACSL5 in fat absorption during the re-esterification of dietary fats3,27,28,48. In the present study it has been noted that once affected Kelpies are on a solid diet with enzyme supplementation, dogs continue to present with a low body condition score. While AK display ongoing sensitivity to dietary lipids into adulthood it remains unconfirmed if their smaller size is a result of persistent intestinal lipid malabsorption or stunted early development.
Further to the complete loss of ACSL5, the genomic deletion resulted in the partial deletion and cryptic splicing event downstream of the last translated exon of ZDHHC6. ZDHHC6 plays a role in posttranslational modification (palmitoylation) of proteins, which can contribute to protein function and regulation beyond underlying genomic architecture. Differences in the palmitoylation of proteins involved in fat and carbohydrate transport and signalling may compromise digestion. Articles reviewing the biological effects of protein palmitoylation have anticipated a functional role in lipid and glucose metabolism49,50, though ZDHHC6 is not currently implicated. ZDHHC6 localises in the endoplasmic reticulum and is reported to be involved in the palmitoylation of five protein targets51,52,53,54,55,56. Within the context of existing research neither ZDHHC6 nor proteins palmitoylated by ZDHHC6 are expected to play a major role in lipid digestion. However, novel roles and targets of palmitoylation are frequently reported and the list of proteins that undergo palmitoylation is constantly growing57 (https://swisspalm.epfl.ch/). It is possible that other key substrates influencing the observed phenotype in the AK are not yet reported and AK harbouring the disease-associated variant may be a unique tool in furthering our current understanding of post-translational modification.
TECTB and GUCY2GP were not expressed in either the case or control RNAseq samples. The genomic region containing the TECTB transcript falls outside the observed variant. It is unlikely that gene expression is altered in appropriate tissue samples. Mice studies indicated that GUCY2G plays a role in jejunal integrity58. However, GUCY2G is a known pseudogene in humans and was suggested to be under purifying selection in the dog59. Conversely, Ensembl genebuild predicts the transcript is non-protein coding (Gene identifier: ENSCAFG00000010908), and recent canine gene catalogue observing ten tissue types reported no expression across all samples and replicates60. The gastrointestinal tract was not reported in the catalogue but a lack of expression in the Labrador retriever control supports the concept of GUCY2GP as a pseudogene, indicating no involvement in the observed phenotype.
Therapies to overcome deficit in ACSL5 function are currently unknown and were not assessed in this research. In humans, therapies for disorders disrupting lipid digestion and absorption, involve removing lipids from the diet or replacing them with those that bypass the genetic block61,62. The disorder described in this study chiefly impacts the metabolism of LCFA. Some human studies have demonstrated positive effects of medium-chain triglyceride formulation (MCT) on individuals suffering from long chain fatty acid disorders63,64,65, however the use of MCT in canine research is restricted66,67,68,69. Auxiliary research into therapeutic options especially during early development is necessary.
Results of the multiplex-PCR were consistent with a fully-penetrant autosomal recessive disorder. Results reported here are not indicative of breed-wide prevalence rates as dogs included in this study originated from a small group of Australian kennels. However, the presence of the deletion in international samples suggests that the variant allele is globally dispersed. To obtain comprehensive prevalence parameters, randomised and wide scale testing is required.
In conclusion we presented a novel deletion of ACSL5, causing hereditary intestinal lipid malabsorption in the Australian Kelpie dog breed. ACSL5 plays an important role in long chain fatty acid storage and metabolism. The improved health of affected individuals with age implies that genetic compensation of this gene beyond neonatal development is possible. This research identifies the first spontaneous animal model to validate key mouse knockout model findings previously reported. The AK model presents a unique opportunity to improve gaps in our understanding of ACSL5. A simple genetic test has been developed and validated to identify dogs harbouring the described variant. International testing of Australian Kelpies is warranted to obtain better estimates of global prevalence. At this time the disorder is presumed to be restricted to a single breed.
Data availability
The dataset used in the current study is available at Figshare https://doi.org/10.6084/m9.figshare.12380564.
References
Watkins, P. A. & Ellis, J. M. Peroxisomal acyl-CoA synthetases. Biochim. Biophys. Acta BBA Mol. Basis Disease 1822, 1411–1420. https://doi.org/10.1016/j.bbadis.2012.02.010 (2012).
Watkins, P. A., Maiguel, D., Jia, Z. & Pevsner, J. Evidence for 26 distinct acyl-coenzyme A synthetase genes in the human genome. J. Lipid Res. 48, 2736–2750. https://doi.org/10.1194/jlr.M700378-JLR200 (2007).
Mashek, D. G., Li, L. O. & Coleman, R. A. Long-chain acyl-CoA synthetases and fatty acid channeling. Future Lipidol. 2, 465–476. https://doi.org/10.2217/17460875.2.4.465 (2007).
Arnott, E. R. et al. Strong selection for behavioural resilience in Australian stock working dogs identified by selective sweep analysis. Canine Genet. Epidemiol. 2, 6. https://doi.org/10.1186/s40575-015-0017-6 (2015).
Chew, T., Willet, C. E., Haase, B. & Wade, C. M. Genomic characterization of external morphology traits in kelpies does not support common ancestry with the Australian Dingo. Genes. https://doi.org/10.3390/genes10050337 (2019).
Fyfe, J. C. et al. Inherited selective cobalamin malabsorption in Komondor dogs associated with a CUBN splice site variant. BMC Vet. Res. 14, 418. https://doi.org/10.1186/s12917-018-1752-1 (2018).
Fyfe, J. C. et al. An exon 53 frameshift mutation in CUBN abrogates cubam function and causes Imerslund-Gräsbeck syndrome in dogs. Mol. Genet. Metab. 109, 390–396. https://doi.org/10.1016/j.ymgme.2013.05.006 (2013).
Fyfe, J. C., Hemker, S. L., Venta, P. J., Stebbing, B. & Giger, U. Selective intestinal cobalamin malabsorption with proteinuria (Imerslund-Gräsbeck syndrome) in juvenile Beagles. J. Vet. Intern. Med. 28, 356–362. https://doi.org/10.1111/jvim.12284 (2014).
Gold, A. J., Scott, M. A. & Fyfe, J. C. Failure to thrive and life-threatening complications due to inherited selective cobalamin malabsorption effectively managed in a juvenile Australian shepherd dog. Can. Vet. J. 56, 1029–1034 (2015).
He, Q. et al. Amnionless function is required for cubilin brush-border expression and intrinsic factor-cobalamin (vitamin B12) absorption in vivo. Blood 106, 1447–1453. https://doi.org/10.1182/blood-2005-03-1197 (2005).
Kather, S., Grützner, N., Kook, P. H., Dengler, F. & Heilmann, R. M. Review of cobalamin status and disorders of cobalamin metabolism in dogs. J. Vet. Intern. Med. 34, 13–28. https://doi.org/10.1111/jvim.15638 (2020).
Chang, C. C. et al. Second-generation PLINK: Rising to the challenge of larger and richer datasets. GigaScience. https://doi.org/10.1186/s13742-015-0047-8 (2015).
Kõressaar, T. et al. Primer3_masker: Integrating masking of template sequence with primer design software. Bioinformatics 34, 1937–1938. https://doi.org/10.1093/bioinformatics/bty036 (2018).
Koressaar, T. & Remm, M. Enhancements and modifications of primer design program Primer3. Bioinformatics 23, 1289–1291. https://doi.org/10.1093/bioinformatics/btm091 (2007).
Untergasser, A. et al. Primer3—New capabilities and interfaces. Nucleic Acids Res. 40, e115–e115. https://doi.org/10.1093/nar/gks596 (2012).
Ewels, P., Magnusson, M., Lundin, S. & Käller, M. MultiQC: Summarize analysis results for multiple tools and samples in a single report. Bioinformatics 32, 3047–3048. https://doi.org/10.1093/bioinformatics/btw354 (2016).
Dobin, A. et al. STAR: Ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21. https://doi.org/10.1093/bioinformatics/bts635 (2013).
Grabherr, M. G. et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 29, 644–652. https://doi.org/10.1038/nbt.1883 (2011).
Wang, L., Wang, S. & Li, W. RSeQC: Quality control of RNA-seq experiments. Bioinformatics 28, 2184–2185. https://doi.org/10.1093/bioinformatics/bts356 (2012).
Thorvaldsdóttir, H., Robinson, J. T. & Mesirov, J. P. Integrative Genomics Viewer (IGV): High-performance genomics data visualization and exploration. Brief. Bioinform. 14, 178–192. https://doi.org/10.1093/bib/bbs017 (2012).
Garrido-Martín, D., Palumbo, E., Guigó, R. & Breschi, A. ggsashimi: Sashimi plot revised for browser- and annotation-independent splicing visualization. PLoS Comput. Biol. 14, e1006360. https://doi.org/10.1371/journal.pcbi.1006360 (2018).
Sewell, A. C., Haskins, M. E. & Giger, U. Inherited metabolic disease in companion animals: Searching for nature’s mistakes. Vet. J. 174, 252–259. https://doi.org/10.1016/j.tvjl.2006.08.017 (2007).
Koeberl, D. D., Pinto, C., Brown, T. & Chen, Y. T. Gene therapy for inherited metabolic disorders in companion animals. ILAR J. 50, 122–127. https://doi.org/10.1093/ilar.50.2.122 (2009).
Dandrieux, J. R. S., Noble, P. J. M., Halladay, L. J., McLean, L. & German, A. J. Canine breed predispositions for marked hypocobalaminaemia or decreased folate concentration assessed by a laboratory survey. J. Small Anim. Pract. 54, 143–148. https://doi.org/10.1111/jsap.12039 (2013).
German, A. J. Exocrine pancreatic insufficiency in the dog: Breed associations, nutritional considerations, and long-term outcome. Topics Companion Animal Med. 27, 104–108. https://doi.org/10.1053/j.tcam.2012.04.004 (2012).
Jergens, A. E. & Simpson, K. W. Inflammatory bowel disease in veterinary medicine. Front. Biosci. (Elite Edition). 4, 1404 (2012).
Bowman, T. A. et al. Acyl CoA synthetase 5 (ACSL5) ablation in mice increases energy expenditure and insulin sensitivity and delays fat absorption. Mol. Metab. 5, 210–220. https://doi.org/10.1016/j.molmet.2016.01.001 (2016).
Mashek, D. G., Li, L. O. & Coleman, R. A. Rat long-chain acyl-CoA synthetase mRNA, protein, and activity vary in tissue distribution and in response to diet. J. Lipid Res. 47, 2004–2010. https://doi.org/10.1194/jlr.M600150-JLR200 (2006).
Mashek, D. G., McKenzie, M. A., Van Horn, C. G. & Coleman, R. A. Rat long chain Acyl-CoA synthetase 5 increases fatty acid uptake and partitioning to cellular triacylglycerol in McArdle-RH7777 cells. J. Biol. Chem. 281, 945–950. https://doi.org/10.1074/jbc.M507646200 (2006).
Poppelreuther, M. et al. The N-terminal region of acyl-CoA synthetase 3 is essential for both the localization on lipid droplets and the function in fatty acid uptake. J. Lipid Res. 53, 888–900. https://doi.org/10.1194/jlr.M024562 (2012).
Teodoro, B. G. et al. Long-chain acyl-CoA synthetase 6 regulates lipid synthesis and mitochondrial oxidative capacity in human and rat skeletal muscle. J. Physiol. 595, 677–693. https://doi.org/10.1113/JP272962 (2017).
Wu, M., Cao, A., Dong, B. & Liu, J. Reduction of serum free fatty acids and triglycerides by liver-targeted expression of long chain acyl-CoA synthetase 3. Int. J. Mol. Med. 27, 655–662. https://doi.org/10.3892/ijmm.2011.624 (2011).
Adamo, K. B. et al. Peroxisome proliferator-activated receptor γ 2 and Acyl-CoA synthetase 5 polymorphisms influence diet response. Obesity 15, 1068–1075. https://doi.org/10.1038/oby.2007.630 (2007).
Ellis, J. M. et al. Adipose acyl-CoA synthetase-1 directs fatty acids toward beta-oxidation and is required for cold thermogenesis. Cell Metab. 12, 53–64. https://doi.org/10.1016/j.cmet.2010.05.012 (2010).
Killion, E. A. et al. A role for long-chain acyl-CoA synthetase-4 (ACSL4) in diet-induced phospholipid remodeling and obesity-associated adipocyte dysfunction. Mol. Metab. 9, 43–56. https://doi.org/10.1016/j.molmet.2018.01.012 (2018).
Privette, J. D., Hickner, R. C., MacDonald, K. G., Pories, W. J. & Barakat, H. A. Fatty acid oxidation by skeletal muscle homogenates from morbidly obese black and white American women. Metabolism 52, 735–738. https://doi.org/10.1016/S0026-0495(03)00034-9 (2003).
Rajkumar, A. et al. Acyl-CoA synthetase long-chain 5 genotype is associated with body composition changes in response to lifestyle interventions in postmenopausal women with overweight and obesity: A genetic association study on cohorts Montréal-Ottawa New Emerging Team, and Complications Associated with Obesity. BMC Med. Genet. 17, 56. https://doi.org/10.1186/s12881-016-0320-4 (2016).
Rajkumar, A. et al. ACSL5 genotype influence on fatty acid metabolism: A cellular, tissue, and whole-body study. Metabolism 83, 271–279. https://doi.org/10.1016/j.metabol.2018.03.019 (2018).
Zhao, C. et al. Adaptive evolution of the ACSL gene family in Carnivora. Genetica 147, 141–148. https://doi.org/10.1007/s10709-019-00057-3 (2019).
Plassais, J. et al. Analysis of large versus small dogs reveals three genes on the canine X chromosome associated with body weight, muscling and back fat thickness. PLoS Genet. 13, e1006661. https://doi.org/10.1371/journal.pgen.1006661 (2017).
Meller, N., Morgan, M. E., Wong, W. P., Altemus, J. B. & Sehayek, E. Targeting of Acyl-CoA synthetase 5 decreases jejunal fatty acid activation with no effect on dietary long-chain fatty acid absorption. Lipids Health Dis. 12, 88–88. https://doi.org/10.1186/1476-511X-12-88 (2013).
Glick, B. S. & Rothman, J. E. Possible role for fatty acyl-coenzyme A in intracellular protein transport. Nature 326, 309–312. https://doi.org/10.1038/326309a0 (1987).
Kang, M.-J. et al. A novel arachidonate-preferring acyl-CoA synthetase is present in steroidogenic cells of the rat adrenal, ovary, and testis. Proc. Natl. Acad. Sci. 94, 2880. https://doi.org/10.1073/pnas.94.7.2880 (1997).
Oikawa, E. et al. A novel Acyl-CoA synthetase, ACS5, expressed in intestinal epithelial cells and proliferating preadipocytes1. J. Biochem. 124, 679–685. https://doi.org/10.1093/oxfordjournals.jbchem.a022165 (1998).
Gassler, N. et al. Expression of acyl-CoA synthetase 5 reflects the state of villus architecture in human small intestine. J. Pathol. 202, 188–196. https://doi.org/10.1002/path.1504 (2004).
Gassler, N. et al. Regulation of enterocyte apoptosis by Acyl-CoA synthetase 5 splicing. Gastroenterology 133, 587–598. https://doi.org/10.1053/j.gastro.2007.06.005 (2007).
Gassler, N. et al. Molecular characterisation of non-absorptive and absorptive enterocytes in human small intestine. Gut 55, 1084–1089. https://doi.org/10.1136/gut.2005.073262 (2006).
Bu, S. Y. & Mashek, D. G. Hepatic long-chain acyl-CoA synthetase 5 mediates fatty acid channeling between anabolic and catabolic pathways. J. Lipid Res. 51, 3270–3280. https://doi.org/10.1194/jlr.M009407 (2010).
Blaskovic, S., Blanc, M. & van der Goot, F. G. What does S-palmitoylation do to membrane proteins?. FEBS J. 280, 2766–2774. https://doi.org/10.1111/febs.12263 (2013).
Spinelli, M., Fusco, S. & Grassi, C. Nutrient-dependent changes of protein palmitoylation: Impact on nuclear enzymes and regulation of gene expression. Int. J. Mol. Sci. https://doi.org/10.3390/ijms19123820 (2018).
Sandoz, P. A. et al. The architecture of the endoplasmic reticulum is regulated by the reversible lipid modification of the shaping protein CLIMP-63. bioRxiv. https://doi.org/10.1101/431106 (2018).
Abrami, L. et al. Identification and dynamics of the human ZDHHC16-ZDHHC6 palmitoylation cascade. Elife 6, e27826. https://doi.org/10.7554/eLife.27826 (2017).
Fairbank, M., Huang, K., El-Husseini, A. & Nabi, I. R. RING finger palmitoylation of the endoplasmic reticulum Gp78 E3 ubiquitin ligase. FEBS Lett. 586, 2488–2493. https://doi.org/10.1016/j.febslet.2012.06.011 (2012).
Fredericks, G. J. et al. Stable expression and function of the inositol 1,4,5-triphosphate receptor requires palmitoylation by a DHHC6/selenoprotein K complex. Proc. Natl. Acad. Sci. 111, 16478. https://doi.org/10.1073/pnas.1417176111 (2014).
Lakkaraju, A. K. K. et al. Palmitoylated calnexin is a key component of the ribosome–translocon complex. EMBO J. 31, 1823–1835. https://doi.org/10.1038/emboj.2012.15 (2012).
Senyilmaz, D. et al. Regulation of mitochondrial morphology and function by stearoylation of TFR1. Nature 525, 124. 10.1038/nature14601. https://www.nature.com/articles/nature14601#supplementary-information (2015).
Blanc, M. et al. SwissPalm: Protein palmitoylation database. F1000Res 4, 261–261. https://doi.org/10.12688/f1000research.6464.1 (2015).
Lo, H.-C., Yang, R.-B. & Lee, C.-H. Guanylyl cyclase-G modulates Jejunal apoptosis and inflammation in mice with intestinal ischemia and reperfusion. PLoS ONE 9, e101314. https://doi.org/10.1371/journal.pone.0101314 (2014).
Young, J. M., Waters, H., Dong, C., Fülle, H.-J. & Liman, E. R. Degeneration of the olfactory guanylyl cyclase D Gene during PRIMATE evolution. PLoS ONE 2, e884. https://doi.org/10.1371/journal.pone.0000884 (2007).
Hoeppner, M. P. et al. An improved canine genome and a comprehensive catalogue of coding genes and non-coding transcripts. PLoS ONE 9, e91172. https://doi.org/10.1371/journal.pone.0091172 (2014).
Goetzman, E. S. Advances in the understanding and treatment of mitochondrial fatty acid oxidation disorders. Curr. Genet. Med. Rep. 5, 132–142. https://doi.org/10.1007/s40142-017-0125-6 (2017).
Spiekerkoetter, U. et al. Current issues regarding treatment of mitochondrial fatty acid oxidation disorders. J. Inherit. Metab. Dis. 33, 555–561. https://doi.org/10.1007/s10545-010-9188-1 (2010).
Behrend, A. M. et al. Substrate oxidation and cardiac performance during exercise in disorders of long chain fatty acid oxidation. Mol. Genet. Metab. 105, 110–115. https://doi.org/10.1016/j.ymgme.2011.09.030 (2012).
Gillingham, M. B., Scott, B., Elliott, D. & Harding, C. O. Metabolic control during exercise with and without medium-chain triglycerides (MCT) in children with long-chain 3-hydroxy acyl-CoA dehydrogenase (LCHAD) or trifunctional protein (TFP) deficiency. Mol. Genet. Metab. 89, 58–63. https://doi.org/10.1016/j.ymgme.2006.06.004 (2006).
van Eerd, D. C. D. et al. Management of an LCHADD patient during pregnancy and high intensity exercise. In JIMD Reports, Vol. 32, 95–100. https://doi.org/10.1007/8904_2016_561 (Springer, Berlin, Heidelberg, 2017).
Beynen, A. C., Kappert, H. J., Lemmens, A. G. & Van Dongen, A. M. Plasma lipid concentrations, macronutrient digestibility and mineral absorption in dogs fed a dry food containing medium-chain triglycerides. J. Animal Physiol. Animal Nutr. 86, 306–312. https://doi.org/10.1046/j.1439-0396.2002.00387.x (2002).
Matulka, R. A., Thompson, D. V. M. L. & Burdock, G. A. Lack of toxicity by medium chain triglycerides (MCT) in canines during a 90-day feeding study. Food Chem. Toxicol. 47, 35–39. https://doi.org/10.1016/j.fct.2008.06.080 (2009).
Rutz, G. M., Steiner, J. M., Bauer, J. E. & Williams, D. A. Effects of exchange of dietary medium chain triglycerides for long-chain triglycerides on serum biochemical variables and subjectively assessed well-being of dogs with exocrine pancreatic insufficiency. Am. J. Vet. Res. 65, 1293. https://doi.org/10.2460/ajvr.2004.65.1293 (2004).
Cotter, R., Taylor, C. A., Johnson, R. & Rowe, W. B. A metabolic comparison of a pure long-chain triglyceride lipid emulsion (LCT) and various medium-chain triglyceride (MCT)-LCT combination emulsions in dogs. Am. J. Clin. Nutr. 45, 927–939. https://doi.org/10.1093/ajcn/45.5.927 (1987).
Acknowledgements
The authors acknowledge the Sydney Informatics Hub, a Core Research Facility, of the University of Sydney for access to the High Performance Computing cluster ‘Artemis’ as well as University of Sydney Veterinary Pathology Diagnostic Services for their assistance with conducting a full post-mortem and histological study for use within this research. The PRJ-010413 Grant was received by corresponding author C. M. Wade from AgriFutures Australia (https://www.agrifutures.com.au/). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
All authors contributed to the conceptualisation of the research and progression of the overall aims. The primary investigation, including experimental work, data analysis, validation and visualisation, was conducted by M.J.O. under the supervision of C.M.W. and N.J.B. T.C. provided valuable advice on the data analysis. C.M.W., N.J.B. and M.S. were involved in the provision of resources. M.J.O. drafted the original manuscript. All authors were involved in editing and refining the manuscript. The final product has been approved by all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
O’Brien, M.J., Beijerink, N.J., Sansom, M. et al. A large deletion on CFA28 omitting ACSL5 gene is associated with intestinal lipid malabsorption in the Australian Kelpie dog breed. Sci Rep 10, 18223 (2020). https://doi.org/10.1038/s41598-020-75243-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-75243-x
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.