Abstract
Integrin activation is associated with conformational regulation. In this study, we developed a system to evaluate conformational changes in α4β7 integrin. We first inserted the PA tag into the plexin-semaphorin-integrin (PSI) domain of β7 chain, which reacted with an anti-PA tag antibody (NZ-1) in an Mn2+-dependent manner. The small GTPase Rap1 deficiency, as well as chemokine stimulation and the introduction of the active form of Rap1, Rap1V12, enhanced the binding of NZ-1 to the PA-tagged mutant integrin, and increased the binding affinity to mucosal addressing cell adhesion molecule-1 (MAdCAM-1). Furthermore, we generated two kinds of hybridomas producing monoclonal antibodies (mAbs) that recognized Mn2+-dependent epitopes of β7. Both epitopes were exposed to bind to mAbs on the cells by the introduction of Rap1V12. Although one epitope in the PSI domain of β7 was exposed on Rap1-deficienct cells, the other epitope in the hybrid domain of β7 was not. These data indicate that the conversion of Rap1-GDP to GTP exerts two distinct effects stepwise on the conformation of α4β7. The induction of colitis by Rap1-deficient CD4+ effector/memory T cells suggests that the removal of constraining effect by Rap1-GDP on α4β7 is sufficient for homing of these pathogenic T cells into colon lamina propria (LP).
Similar content being viewed by others
Introduction
Lymphocyte adhesion and migration are important for the generation and execution of immune and inflammatory responses. Integrins are a family of α/β heterodimeric adhesion receptors that transmit signals bi-directionally across the plasma membrane1,2,3. In the multistep leukocyte adhesion cascade, selectins generally mediate rolling, and integrins mediate subsequent arrest. In contrast, the gut homing integrin α4β7 mediates leukocyte rolling and arrest in vivo4. MAdCAM-1, a ligand for α4β7, is constitutively expressed in postcapillary venules of intestinal lamina propria (LP) and acts as a key addressin for intestinal homing5. Therefore, the adhesive activity of α4β7 directly reflects the ability of cells to move to the mucosal tissues of the intestine.
Regulation of T-cell trafficking by both Rap1-GTP and -GDP is a key control mechanism of the lymphocyte adhesion cascade6. Rap1-GTP recruits downstream effectors, such as RAPL (regulator of cell polarization and adhesion enriched in lymphoid tissues), which binds integrin α chain, and RIAM (RAP1-interacting adapter molecule) and talin which bind integrin β chain7,8,9. Rap1-GDP suppresses lymphocyte rolling behaviors via activation of LOK (lymphocyte-orientated kinase) and phosphorylation of ERM (ezrin, radixin and moesin)10.
The T cell number in mesenteric lymph nodes is important for mucosal tolerance. Integrin activation by Rap1-GTP plays an important role in the circulation of naive T (TN) cells, whereas Rap1-GDP in resting TN and effector/memory T (TEM) cells limits rolling behaviors in blood vessels and retards lymphocyte homing10. Therefore, Rap1 deficiency leads to lymphopenia and the generation of pathogenic TEM cells in lymph nodes. Furthermore, it facilitates homing of TEM cells into the colon, which exacerbates spontaneous T-cell-dependent colitis and tubular adenomas10. Excess infiltration of TEM cells by Rap1 deficiency points to the involvement of Rap1-GDP in the regulation of the activation of α4β7.
Binding of α4β7 to MAdCAM-1 with high affinity is critical step for lymphocyte arrest. The regulation of the ligand-binding affinity is associated with conformational rearrangement of the integrin molecule11. Previous studies showed that integrin extracellular domains existed in distinct global conformational states that differed in their affinity for ligands: a low-affinity bent conformation with a close headpiece and a high-affinity extended conformation with an open head piece12,13,14. The equilibrium among these different states is regulated by integrin inside-out signaling and extracellular stimuli, such as divalent cations. Compared with the low-affinity state in Ca2+/Mg2+, the removal of Ca2+ or the addition of Mn2+ results in a marked increase in ligand-binding of almost all integrins15. There are few conventional activation-specific antibodies to recognize only activated α4β716,17.
In this study, to explore the effects of Rap1 deficiency on the conformation of α4β7, we developed an epitope grafting system to detect conformational changes in α4β7 by inserting a PA tag into the PSI domain of the β7 chain. The PA tag was a 12-residue peptide (GVAMPGARDDVV) derived from human podoplanin, which is recognized by NZ-118. We also prepared one rat mAb (G3 mAb), in which the epitope was located in the PSI domain, as was the case with the grafting site of the PA tag. Using NZ-1 or G3 mAb, we revealed that Rap1 deficiency, as well as chemokine stimulation, induced the active conformation of α4β7, suggesting that Rap1-GDP restricts conformation of α4β7 in an inactive state. The other rat mAb (H3 mAb), in which the epitope was located in the hybrid domain of the β7 chain, recognized the active conformation of α4β7 on Rap1V12-expressing cells, but not that on Rap1-deficient cells. These data suggested that Rap1-GTP further promoted conformational change leading to high-affinity α4β7. Consistent with these data, Rap1-deficient CD4+ TEM cells (pathogenic T cells), which home to colon LP in a α4β7-MAdCAM-1-dependent manner and induce colitis, exhibited increased expression of the epitope recognized by G3 mAb but not H3 mAb. Thus, constraining effect by Rap1-GDP on α4β7 presumably contributes to suppress excess infiltration of pathogenic T cells into colon LP and prevent the development of colitis.
Results
Development of a system to measure the active conformation of α4β7 by inserting a PA tag
α4β7 showed low-affinity state to MAdCAM-1 in Ca2+/Mg2+, and the addition of Mn2+ increased the binding affinity of α4β7 to MAdCAM-1 (Fig. 1a). To probe the conformational state of α4β7 using the PA-tag-NZ-1 system (Fig. 1a), a proB cell line (BAF cells), in which the β7 chain was knocked-out using CRISPR/Cas9-mediated genome editing, and β7 chains cDNA which were inserted PA tag into 4 locations (PAins 1: 23/24, PAins 2: 29/30, PAins 3: 427/428, PAins 4: 431/432) were introduced into BAF cells (Fig. 1b,c). These insertion mutants of β7-expressing cells were stained with NZ-1 or FIB504 (conventional mAb against mouse/human β7, which recognizes the binding sites of α4β7 with MAdCAM-1) in the presence of 1 mM Ca2+/Mg2+ or 0.5 mM Mn2+, and analyzed by flow cytometry. All mutants of β7 were approximately equally expressed on the cell surface (Fig. 1c). PA expression on the surface of PAins2-expressing cells (PAins2 cells) was reduced in the low-affinity α4β7 with the bent conformation in the presence of Ca2+/Mg2+ and exhibited an eightfold increase in the high-affinity α4β7 with the extended conformation in the presence of Mn2+ (Fig. 1d). In the cells expressing other insertion mutants of β7, PA was exposed to be recognized by NZ-1 in the bent conformation, the same as in the extended conformation of β7 (Fig. 1d). These data showed that PAins2 in the PSI domain of β7 was an appropriate PA tag insertion design.
Detection of conformational changes in α4β7 using a PA tag. (a) (Left) A model of the conformational states of the extracellular domain of α4β7: bent form with a closed headpiece in low affinity and extended form with an open headpiece in high affinity. (Right) The binding of wild-type (wt) BAF cells to MAdCAM-1 in the presence of 1 mM Ca2+/Mg2+, or 0.5 mM Mn2+. The IMF (Mean Fluorescence Intensity) of binding to MAdCAM-1-Fc was normalized to the IMF of anti-β7 (FIB504). The IMF is presented as the fold-increase relative to that of wt cells in the presence of 1 mM Ca2+/Mg2+ values of 1. Data represent the mean ± SE of three independent experiments. *P < 0.005, versus in the presence of Ca2+/Mg2+. (b) (Upper) Systematic overview of the gene targeting strategy. The position of the single-guide RNA target site of exon 3 is indicated by an underline. (Lower) Flow cytometry profiles of anti-β7 of control or knockout cells using CRISPR/Cas9-mediated genome editing is shown. (c) (Left) Insertion sites of the PA tag. The PA tag sequence was inserted at the indicated sites of the β7 sequence in the designed mutants (PAins 1–4). (Right) Flow cytometry profiles of anti-β7 of BAF cells transfected with each designed mutant. (d) (Left) The binding of NZ-1 to each transfectant in the presence of 1 mM Ca2+/Mg2+, or 0.5 mM Mn2+. The IMF of binding of each transfectant to NZ-1 was normalized to the IMF of anti-β7 and is presented as the fold-increase relative to that of each transfectant in the presence of 1 mM Ca2+/Mg2+ values of 1. Data represent the mean ± SE of three independent experiments. (Right) Flow cytometry profiles of NZ-1 of PAins1 and PAins2 in the presence of 1 mM Ca2+/Mg2+, or 0.5 mM Mn2+. *P < 0.001, versus in the presence of Ca2+/Mg2+.
As exogenous addition of antibodies that preferentially bind to the extended conformation can shift the equilibrium toward the high-affinity state of integrins, they often activate integrins from outside the cell. Therefore, we confirmed the conformational change in α4β7 in the presence of Mn2+ using the Fv-clasp of NZ-1. The Fv-clasp of NZ-1, an artificially designed small antibody fragment of 37 kDa, was used as a reporter of conformational change, as it did not affect the equilibrium between the high- and low-affinity states19. Using Fv-clasp of NZ-1, the surface expression of PA tag exhibited a 11-fold increase in the presence of Mn2+, compared to that in the presence of Ca2+/Mg2+ (Fig. 2), indicating that the surface expression of PA precisely reflects the active conformation of α4β7.
The Fv-clasp of NZ-1 recognized a conformational change in α4β7. (Left) Flow cytometry profiles of FIB504 or the Fv-clasp of NZ-1 of PAins2-expressing cells in the presence of 1 mM Ca2+/Mg2+, or 0.5 mM Mn2+. (Right) The binding of the Fv-clasp of NZ-1 to PAins2-expressing cells in the presence of 1 mM Ca2+/Mg2+, or 0.5 mM Mn2+. The IMF of binding of PAins2-expressing cells to the Fv-clasp of NZ-1 in the presence of 0.5 mM Mn2+ was normalized to the IMF of anti-β7 (FIB504) and is presented as the fold-increase relative to that in the presence of 1 mM Ca2+/Mg2+ values of 1. Data represent the mean ± SE of three independent experiments. *P < 0.003, versus in the presence of Ca2+/Mg2+.
Rap1 deficiency induced a conformational change in α4β7
A conformational equilibrium between bent (low-affinity) and extended (high-affinity) states of integrin was found to be regulated by inside-out signaling such as Rap1. Using PAins2 cells, we examined the effects of Rap1 on the conformational state of α4β7. To this end, we introduced the GTP-binding form of Rap1, Rap1V12, Rap1 GTPase activating protein (GAP), Spa-1, and knocked down of Rap1a/b in PAins2 cells (Fig. 3a). We confirmed that CXCL12-induced Rap1 activation was inhibited in Spa-1-expressing PAins2 cells (Fig. 3b).
The conformation of α4β7 was regulated by Rap1. (a) Immunoblot of the total cell lysates from the cells introduced with control, T7-Rap1V12, flag-Spa-1-expressing, or Rap1a/b-knockdown cells with anti-T7, flag, or Rap1 antibodies. (b) GTP-bound Rap1 was analyzed by a pull-down assay using GST-Ral-GDS-RBD. Control or SPA-1-expressing cells were stimulated with 100 nM of CXCL12 at the indicated times, lysed, and subjected to a pull-down assay. Bound Rap1 and total Rap1 were detected by immunoblotting using an anti-Rap1 antibody. (c) Ligand binding affinity to soluble MAdCAM-1-Fc. The binding of each transfectant to soluble MAdCAM-1-Fc in the presence or absence of CXCL12. The IMF of soluble MAdCAM-1 binding was normalized to the IMF of anti-β7 (FIB504) and is presented as the fold-increase relative to that of unstimulated control cell values of 1. Data represent the mean ± SE of three independent experiments. *1P < 0.001, versus unstimulated cells, *2P < 0.001, versus unstimulated control cells, *3P < 0.006, versus CXCL12-stimulated control cells, *4P < 0.05, versus unstimulated control cells. (d) The binding of NZ-1 to each transfectant in the presence or absence of CXCL12. The IMF of NZ-1 binding was normalized to the IMF of anti-β7 and is presented as the fold-increase relative to that of unstimulated control cell values of 1. Data represent the mean ± SE of three independent experiments. *1P < 0.001, versus unstimulated control cells, *2P < 0.001, versus unstimulated control cells, *3P < 0.001, versus CXCL12-stimulated control cells, *4P < 0.001, versus unstimulated control cells.
Next, we examined the binding activity of α4β7 on each transfectant to soluble MAdCAM-1 in the presence or absence of CXCL12. As shown in Fig. 3c, CXCL12 stimulation elevated the binding of α4β7 on control cells to soluble MAdCAM-1, indicating that chemokine stimulation shifted the equilibrium of α4β7 toward high-affinity state. As expected, overexpression of Rap1V12 increased the binding of α4β7 to soluble MAdCAM-1, without CXCL12 stimulation (Fig. 3c). The inhibition of Rap1 activation by overexpression of Spa-1 suppressed CXCL12-dependent increase in the binding of α4β7 to MAdCAM-1 (Fig. 3c). Knockdown of Rap1 also significantly increased the binding activity, but the effect was weak compared to that of Rap1V12-expressing cells (Fig. 3c). These data suggest that Rap1-GDP locks α4β7 in the low-affinity state and that Rap1-GTP further promotes an equilibrium toward high-affinity state of α4β7.
Subsequently, we examined changes in the surface expression of the PA tag in each transfectant. CXCL12 stimulation exhibited a 1.3-fold increase in PA surface expression, and overexpression of Spa-1 completely inhibited PA surface expression induced by CXCL12 stimulation (Fig. 3d), indicating that the conversion of Rap1-GDP to GTP is necessary for the surface expression of PA. Overexpression of Rap1V12 significantly promoted PA surface expression (Fig. 3d). PA surface expression was higher in Rap1-deficient cells as compared with that in Rap1V12-expressing cells (Fig. 3d), suggesting that the loss of Rap1-GDP induced a conformational change in α4β7 and that this change is different from the Rap1V12-induced conformation of α4β7. These results indicate that Rap1-GDP suppresses conformational changes to active form of α4β7.
In addition, talin is reported to bind Rap1-GTP and integrin, and trigger integrin activation20. Therefore, we examined the effect of the knockdown (KD) of talin. The abundance of talin protein in talin KD cells was reduced to 5% of control cells (Fig. S1a). As shown in Fig. S1b, the silencing of talin reduced basal surface expression of PA, whereas CXCL12 increased surface expression of PA in talin-KD cells at a same proportion as control cells. This result suggests that talin is a basic cytoskeletal component necessary for active conformation of α4β7, rather than a downstream effector molecule of chemokine-mediated signaling.
Identification and characterization of rat mAbs to detect Rap1-dependent conformational changes in α4β7
To establish hybridomas producing mAbs that exclusively reacted with α4β7 in an Mn2+-dependent manner, immunogenic N-terminal amino acids (1–458) of β7-MBP fusion protein were injected into rats. A hybridoma producing rat mAb G3 (γ2/κ) for Mn2+-dependent conformation of α4β7 was established. As shown in Fig. 4a, the G3 epitope was almost not detected in the low-affinity α4β7 with a bent conformation in the presence of Ca2+/Mg2+ but increased 4.8-fold in the high-affinity α4β7 with the extended conformation in the presence of Mn2+.
G3 mAb recognized the active conformation of α4β7. (a) (Upper) (left) Flow cytometry profiles of G3 mAb of wild-type (wt) BAF cells in the presence of 1 mM Ca2+/Mg2+, or 0.5 mM Mn2+. (right) The IMF of binding to G3 mAb was normalized to the IMF of anti-β7 (FIB504) and is presented as the fold-increase relative to that of wt BAF cells in the presence of 1 mM Ca2+/Mg2+ values of 1. Data represent the mean ± SE of three independent experiments. *P < 0.001, versus Ca2+/Mg2. (Lower) (left) Flow cytometry profiles of G3 of Jurkat cells expressing human β7 in the presence of 1 mM Ca2+/Mg2+, or 0.5 mM Mn2+. (right) The IMF of binding to G3 mAb was normalized to the IMF of anti-β7 and is presented as the fold-increase relative to that of Jurkat cells in the presence of 1 mM Ca2+/Mg2+ values of 1. Data represent the mean ± SE of three independent experiments. *P < 0.001, versus Ca2+/Mg2+. (b) (Left) (upper) Mapping of G3 epitope. G3 mAb reactivity was determined by co-expressing chimeric murine β1/β7 subunits or ∆1–19 murine β7 with endogenous α4 in β7-knockout BAF cells in the presence of 0.5 mM Mn2+ using flow cytometry. (lower) The amino acid sequence of 1–19 a.a. of murine, human β7 and murine β1. The red characters indicate the candidates recognized by G3 mAb. (right) Flow cytometry profiles of the transfectants with anti-β7 or G3 in the presence of 0.5 mM Mn2+. (c) The binding of G3 mAb to α4β7 on the cells introduced with control, Rap1V12, Spa-1-expressing, or Rap1a/b-knockdown cells in the presence or absence of CXCL12. The IMF of binding to G3 mAb was normalized to the IMF of anti-β7 and is presented as the fold-increase relative to that of unstimulated control cell values of 1. Data represent the mean ± SE of three independent experiments. *1P < 0.001, versus unstimulated cells, *2P < 0.03, versus unstimulated control cells, *3P < 0.002 versus CXCL12-stimulated control cells, *4P < 0.001, versus unstimulated control cells.
Next, we tested the cross-reactivity of the G3 mAb with human α4β7 using Jurkat cells transfected with human β7. The surface expression level of β7 was determined in the Jurkat cells using FIB504 (Fig. 4a). G3 epitope expression in the Jurkat cells was extremely low in the presence of Ca2+/Mg2+ and increased fivefold in the presence of Mn2+ (Fig. 4a). To identify the epitope of G3, we constructed murine β1/β7 chimeras and the deletion mutant (∆1–19) of β7. These mutants were co-expressed with endogenous α4 in β7-knockout BAF cells, and the surface expression level was confirmed by the immunostaining with FIB504 (Fig. 4b). As shown in Fig. 4b, the deletion of N-terminal 1–19 a.a. of β7, which does not exist in β1, let G3 mAb lose the reactivity to murine β7. Thus, flow cytometric analysis of these transfectants showed that the β7 segment 1–19 a.a. located in the PSI domain was required for binding of G3 mAb to β7 (Fig. 4b), which was close to the PA grafting site (Fig. 1c). As expected, G3 epitope expression increased 1.2-fold with CXCL12 stimulation (Fig. 4c). Overexpression of Rap1V12 enhanced the expression of G3 epitope to 1.9-fold (Fig. 4c). Rap1-deficiency also increased the expression of G3 epitope to 2.8-fold (Fig. 4c). Consistent with the results using the PA-tag-NZ-1 system, these data indicate that Rap1-GDP locks the conformation of α4β7 in inactive state.
We also established a hybridoma producing rat mAb H3 (γ2/κ) to detect Mn2+-dependent conformation of α4β7. As shown in Fig. 5a, the expression of H3 epitope was almost not detected in the low-affinity of α4β7 with the bent conformation in the presence of Ca2+/Mg2+ and increased 33-fold in the presence of Mn2+, suggesting that H3 mAb recognized the high-affinity α4β7. Subsequently, we explored the cross-reactivity of H3 mAb with human α4β7 using Jurkat cells transfected with human β7. H3 epitope on Jurkat cells was not expressed in the presence of Mn2+ (Fig. 5a), indicating that H3 mAb did not recognize the active conformation of human β7. To identify the epitope of H3, we constructed murine/human β7 chimeras. These chimeras were co-expressed with endogenous α4 in β7-knockout BAF cells, and the surface expression level was confirmed by immunostaining with FIB504 (Fig. 5b). Flow cytometric analysis of these transfectants showed that the β7 segment 373–393 a.a. located in the hybrid domain was required for binding of H3 mAb to β7 (Fig. 5b). As expected, the expression of H3 epitope also increased 1.4-fold with CXCL12 stimulation (Fig. 5c). Overexpression of Rap1V12 enhanced the expression of H3 epitope to 3.7-fold (Fig. 5c). However, Rap1 deficiency did not increase the expression of H3 epitope with or without CXCL12 stimulation (Fig. 5c). These data indicate that the expression of H3 epitope requires Rap1-GTP.
H3 mAb recognized the active conformation of α4β7. (a) (Upper) (left) Flow cytometry profiles of H3 mAb of wt BAF cells in the presence of 1 mM Ca2+/Mg2+, or 0.5 mM Mn2+. (right) The IMF of binding to H3 mAb was normalized to the IMF of anti-β7 (FIB504) and is presented as the fold-increase relative to that of wt BAF cells in the presence of 1 mM Ca2+/Mg2+ values of 1. Data represent the mean ± SE of three independent experiments. *P < 0.001, versus Ca2+/Mg2+. (Lower) (left) Flow cytometry profiles of H3 of Jurkat cells expressing human β7 in the presence of 0.5 mM Mn2+. (right) Normalized IMF of binding to H3 mAb is presented as the fold-increase relative to that of Jurkat cells in the presence of 1 mM Ca2+/Mg2+ values of 1. Data represent the mean ± SE of three independent experiments. (b) (Left) (upper) Mapping of H3 epitope. H3 mAb reactivity was determined by co-expressing chimeric murine/human β7 subunits with endogenous α4 in β7-knockout BAF cells in the presence of 0.5 mM Mn2+ using flow cytometry. (lower) The amino acid sequence of 373–393 a.a. of murine and human β7. The red characters indicate the candidates recognized by H3 mAb. The blue characters indicate the corresponding human amino acid sequence. (right) Flow cytometry profiles of the transfectants with anti-β7 or H3 mAb in the presence of 0.5 mM Mn2+. (c) The binding of H3 mAb to α4β7 on the cells introduced with control, Rap1V12, Spa-1-expressing, or Rap1a/b-knockdown cells in the presence or absence of CXCL12. The IMF of binding to H3 mAb was normalized to the IMF of anti-β7 and is presented as the fold-increase relative to that of unstimulated control cell values of 1. Data represent the mean ± SE of three independent experiments. *1P < 0.02, versus unstimulated cells, *2P < 0.005, versus unstimulated control cells, *3P < 0.05 versus CXCL12-stimulated control cells, *4P < 0.05 versus CXCL12-stimulated control cells.
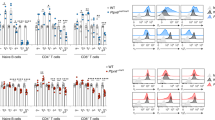
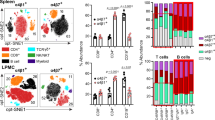
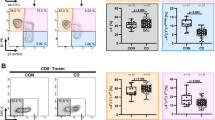
In our previous paper10, we demonstrated that T cell-specific Rap1-deficient mice developed severe colitis with infiltration of CD4+ TEM cells into colon LP and that adoptive transfer of these cells into normal mice induced colitis. Previous study also demonstrated that α4β7-MAdCAM-1-dependent rolling was significantly promoted in Rap1-deficient CD4+ TEM cells10. In the present study, the injection of a MAdCAM-1 inhibitory mAb into T cell-specific Rap1-deficient mice prevented the development of colitis (Fig. 6a, Fig. S3). These findings confirmed that the α4β7-MAdCAM-1 interaction was critical for the development of colitis in T cell-specific Rap1a/b-knockout mice. Therefore, we explored whether a conformational change in α4β7 was observed in pathogenic T cells. Since CCR9 and its ligand CCL25 are found to play essential roles in gut-homing of TEM cells21, we used CCL25 for the stimulation of CD4+ TEM cells. As shown in Fig. 6, G3 epitope significantly increased in pathogenic T cells as compared to that in wild-type TEM cells, although the surface expression of α4β7 was elevated in the pathogenic T cells. CCL25 increased the expression of G3 epitope in control cells but not in pathogenic T cells (Fig. 6b). As expected, the expression of the epitope recognized by H3 mAb was reduced and not induced by CCL25 stimulation in pathogenic T cells, although the addition of Mn2+ induced H3 epitope in these cells at a similar level to that in wild-type TEM cells (Fig. 6c). These data suggest that active conformation in α4β7 induced by Rap1 deficiency is sufficient for the infiltration of the pathogenic T cells into colon LP through rolling and arrest on MAdCAM-1-expressing endothelial cells.
Expression of G3 epitope in colitis-causing CD4+ TEM cells. (a) The administration of anti-MAdCAM-1 inhibited the development of spontaneous colitis in T cell-specific Rap1a/b knockout mice (colitis mice). (Upper) (left) The body weights of colitis mice injected with control or anti-MAdCAM-1 Ab are presented as percentages of the original body weight. (n = 3 in each group). *P < 0.05 versus control Ab. (middle) Histology of intestinal inflammation in a set (#1) of the littermates of colitis mice that received control or anti-MAdCAM-1 Ab (×40). Scale bar, 500 μm. (right) Light microscopic assessment of damages of colitis. (Lower) (left) The density of infiltrated CD4+ T cells in the colon of colitis mice was evaluated using an immunohistological study. *P < 0.02 versus control Ab. (right) Frozen sections of the colon from colitis mice that received control or anti-MAdCAM-1 Ab were stained with anti-CD4 (green) and DAPI (red). Low (×100) Scale bar, 100 μm. (b) The expression of G3 epitope in CD4+ TEM cells. (Upper) The binding of G3 to wt and Rap1-deficient (Rap1KO) CD4+ (CD44+) TEM cells in the presence or absence of CCL25 (left) and with 1 mM Ca2+/Mg2+, or 0.5 mM Mn2+ (right). The IMF of binding to G3 mAb was normalized to the IMF of anti-β7 (FIB504) and is presented as the fold-increase relative to that of unstimulated wt T cell values of 1. Data represent the mean ± SE of three independent experiments. *1P < 0.005, versus unstimulated wt cells. *2P < 0.001, versus unstimulated wt cells. (Lower) Flow cytometry profiles of anti-β7 on unstimulated wt and Rap1KO TEM (CD44+CD62L−) cells, and G3 mAb on β7+-gated wt and Rap1KO TEM. (c) (Upper) The binding of H3 mAb to wt and Rap1 KO CD4+ (CD44+) TEM cells in the presence or absence of CCL25 (left) and with 1 mM Ca2+/Mg2+, or 0.5 mM Mn2+ (right). The IMF of binding to H3 mAb was normalized to the IMF of anti-β7 and is presented as the fold-increase relative to that of unstimulated wt cell values of 1. Data represent the mean ± SE of three independent experiments. *1P < 0.001 versus unstimulated wt cells, *2P < 0.001 versus unstimulated wt cells. (Lower) Flow cytometry profiles of anti-β7 on unstimulated wt and Rap1KO TEM (CD44+CD62L-) cells, and H3 mAb on β7+-gated wt and Rap1KO TEM.
In addition, previous study reports that CCL25 and CXCL10 induces different active conformation of α4β722, but there is no difference between the effects of CCL25 and CXCL10 in the expression of G3 and H3 epitopes on CD4+ TEM cells (Fig. S4).
Discussion
In this study, we developed a sensitive system to probe the conformational states of α4β7 using the insertion of PA tag into the PSI domain of the β7 chain. Using this system, we found that the conformation of α4β7 was regulated by Rap1. To examine the conformational changes of β7 in primary lymphocytes, we isolated two rat mAbs (G3 and H3) against activation-dependent α4β7. G3 mAb recognized the epitope in the PSI domain of β7, and H3 mAb recognized the epitope in the hybrid domain of β7 (Fig. 7a). Both epitopes were hidden in the low-affinity α4β7 with the bent conformation and exposed in the high-affinity α4β7 with the extended conformation induced by the addition of Mn2+. The introduction of Rap1V12 induced the exposure of G3 and H3 epitopes to be recognized by mAbs. However, the expression of G3 epitope was increased by depletion of Rap1-GDP, whereas the conversion to Rap1-GTP was indispensable for the exposure of H3 epitope. Thus, Rap1-GDP and GTP independently regulated the conformation of α4β7 (Fig. 7).
Model for the regulation of the conformation of α4β7 by Rap1. (a) The structure of the α4β7 headpiece. Crystal structure of α4β7 headpiece was derived from PDB database with PDB code of 3V4P4, and created using the PyMOL molecular visualization system. The PSI domain of the β1 headpiece structure is used instead of the PSI domain of β7, and the N-terminal 19 a.a. structure is unknown (surrounded by a dot-line). G3 mAb recognized the epitope in the PSI domain of β7 (red), and H3 mAb recognized the epitope in the hybrid domain of β7 (blue). (b) The model for conformational changes of α4β7. Both G3 (red) and the PA-tag located in PSI domain (green), and H3 (blue) epitopes were all hidden in the low affinity α4β7 with the bent conformation. The surface expression of G3 epitope and PA increased by Rap1 deficiency, which might have induced an “extended-closed form” of α4β7. Rap1-GDP restrained conformational changes in α4β7 and maintained the binding of α4β7 to MAdCAM-1 under conditions of low-affinity binding (the bent form of α4β7). The exposure of H3 epitope was dependent on Rap1-GTP, which was strongly correlated with the binding activity of α4β7 to soluble MAdCAM-1. Rap1-GTP may have induced swing-out of the hybrid domain, which corresponded to the “extended open form” of α4β7, resulting in the binding of α4β7 to MAdCAM-1 with high affinity. (c) Table summarizing the Rap1-binding guanine nucleotide (GTP or GDP) and each antibody (NZ-1, G3, H3)-binding activity (+ or −) in control, Rap1V12 or Spa-1-expressing cells, and Rap1KD cells in the presence or absence (No Stim) of CXCL12. The binding of the antibody to unstimulated control cells was displayed with (−), and the case that significantly increased, was displayed with (+).
The binding of NZ-1 or G3 mAb to the PSI domain of β7 was suppressed by Rap1-GDP, and loss of Rap1-GDP was sufficient for maximal expression of these epitopes (Fig. 7). On the other hand, H3 epitope in the hybrid domain of β7 was not exposed by only deletion of Rap-GDP. The loss of Rap1-GDP had marginal effect on the binding of α4β7 to soluble MAdCAM-1. The overexpression of Rap1V12 as well as the addition of Mn2+, which increased the binding of α4β7 to soluble MAdCAM-1, induced the exposure of H3 epitope (Figs. 3c, 5c). Thus, the surface expression of H3 epitope was strongly correlated with the binding activity of α4β7 to soluble MAdCAM-1. These findings indicate that H3 mAb might detect the swing-out of the hybrid domain in the β7 chain, which is predicted to stabilize the high-affinity conformation23 (Fig. 7). These data suggest that Rap1-GDP restrained the bent conformation in α4β7 and maintain the binding of α4β7 to MAdCAM-1 in low-affinity and that the conversion into Rap1-GTP further facilitated the active conformation of α4β7, resulting in binding of α4β7 to MAdCAM-1 in high-affinity (Fig. 7). As previously reported24,25, conformation-specific antibodies are useful for the elucidation of the functions and regulatory mechanisms of integrin conformation. The combination of G3 and H3 mAbs, which might differentiate extended closed from extended open conformation of α4β7, will contribute to various studies of conformational regulation of α4β7.
The insertion site of PA-tag was between the 29th and 30th in N-terminal of amino acid sequence and G3 epitope was in N-terminal first 19 amino acids. Previous study26,27 report that AP5 is a mAb that recognizes β3 integrin only in the extended conformation. These findings suggest that N-terminal epitope could be used in many or all beta integrins to obtain antibodies that recognize an active conformation. Since there are many kinds of antibodies including N-terminal amino acids of β chain28, it is necessary to clarify whether these antibodies specifically recognize the active conformation of other integrin.
In a previous study, we demonstrated that Rap1-GDP activated LOK and promoted ERM phosphorylation and that the introduction of the active form of LOK or phospho-mimetic ezrin did not prevent conformational changes in α4β7. Although RIAM (Rap1-interacting molecule) and talin are known to induce active conformation of integrins, they are associated with Rap1-GTP but not Rap1-GDP8,9,29. In this study, we suggest that talin is a basic cytoskeletal component involved in active conformation of α4β7. Studies are needed to shed light on what are the downstream effector molecules of Rap1-GDP/GTP and the roles of cytoskeletal proteins such as RIAM and talin in regulation of conformation of α4β7.
Previous paper22 demonstrates that CCL25 and CXCL10 induce different conformation of α4β7, which favors a MAdCAM-1- and VCAM-1-binding, respectively. According to that paper, CCL25 and CXCL10 activates the different signaling pathways which lead to different phosphorylation states of β7 and distinct talin and kindling-3 binding patterns22. On the other hand, there was no difference in surface expression of G3 and H3 epitopes between CCL25 and CXCL10-stimulated T cells (Fig. S4). Therefore, G3 and H3 mAbs did not seem to discriminate the MAdCAM-1 or VCAM-1-binding conformation of α4β7. In this study, the conformation of α4β7 recognized by G3 mAb was suggested to be critical for gut-homing of TEM cells, whereas the physiological significance of Rap1-GTP-dependent active conformation recognized by H3 mAb remains to be solved. It is important to clarify the biological implication of conformational regulation.
Rap1 is indispensable for chemokine-dependent integrin activation and naive lymphocyte recirculation, and its deficiency leads to lymphopenia in secondary lymph nodes10,30. On the other hand, Rap1-deficient TEM cells exhibit enhanced α4β7/MAdCAM-1-dependent rolling and arrest on the endothelium, as well as accelerated homing into colon LP10. In this study, we found that Rap1 deficiency in TEM cells led to a conformational change in α4β7, which promoted α4β7/MAdCAM-1-dependent endothelial transmigration and homing to colon LP10. Furthermore, the Rap1-GTP-dependent high-affinity conformation of α4β7, which was recognized by H3 mAb, was unnecessary for homing of pathogenic T cells into colon LP.
G3 mAb recognized the murine and human active form of α4β7, and the expression of G3 epitope correlated with the infiltration of pathogenic TEM cells into colon LP. Thus, this mAb might be a useful tool to deliver drugs to pathogenic TEM cells. In addition, as the inhibition of the conformational change in α4β7 seemed to be effective in preventing the infiltration of TEM cells into colon LP, the system we developed can be used to screen for drugs to treat colitis. By recognizing conformational changes in α4β7, the system can serve as a useful tool for studies on α4β7 activation mechanisms and the development of new therapies for colitis and subsequent colorectal cancer.
Methods
Mice
All animal experiments were carried out in accordance with Regulations for the Care and Use of Laboratory Animals in Kitasato University, and the protocols used in this study were ethically approved by the Institutional Animal Care and Use Committee for Kitasato University.
Rap1af/f mice containing floxed exons 2–3 of Rap1a, Rap1bf/f mice containing floxed exon 1 of Rap1b were maintained under specific pathogen–free conditions. Those mice were crossed with CD4-Cre mice, yielding mice with T cell-specific deletion of Rap1a/b10.
Cell lines
Ba/F3 cells (BAF cells) and Jurkat cells were cultured as previously reported31. BAF cells were cultured with RPMI1640 medium containing 10% fetal calf serum, 50 mM beta-mercaptoethanol, and 1% WEHI-3 conditional medium as a source of interleukin 3. Jurkat cells were maintained with RPMI1640 medium containing 10% fetal calf serum. All cell lines were tested for mycoplasma contamination by 4′,6-diamidino-2-phenylindole (DAPI) staining with negative results.
Antibodies and reagents
Fluorescence-conjugated anti-mouse CD4, CD44, anti-β7 (FIB504) (BioLegend), anti-Rap1(BD Biosciences), β-actin (Sigma), T7 (MBL) , Flag (Wako), anti-talin (Abcam), APC-conjugated anti-Rat or human IgG, and peroxidase-conjugated goat anti-Rabbit or -mouse IgG (Cell Signaling) were used for flow cytometry and immunoblotting. Anti-MAdCAM-1(MECA-367) (ATCC), G3 and H3 mAb were purified using HiTrap Protein G HP (GE healthcare). Mouse CXCL12, CCL25 and CXCL10 were purchased from R&D Systems. The single-chain Fv of NZ-1 (Fv-clasp) was created by fusing an anti-parallel coiled-coil structure derived from the SARAH domain of human Mst1 kinase to the fragments of VH and VL of NZ-1. NZ-1VH-SARAH and VL-SARAH were separately expressed in E. coli strain BL21, and cultured in standard LB media19. After the cell lysis by sonication, the denatured and solubilized VH-SARAH and VL-SARAH chains were then mixed, and the denaturing reagent was diluted to promote protein folding, and correctly-folded, disulfide-bonded Fv-clasp was purified as previously reported19. Fv-clasp was fluorescently labeled with Alexa Fluor 647 Amine-Reactive Dye (Thermo Fisher Scientific).
DNA constructs and transfection
cDNA encoding murine β7 cDNA was subcloned into a pcDNA3.1 vector. Then, β7 mutants with a PA tag were generated from a pcDNA3.1-murine β7 construct using inverse PCR. The following oligonucleotides and their corresponding complimentary strands were used: for PAins 1: 5′-CCTGACCTGTCTCTGCAGGGCGTTGCCATGCCAGGTGCCGAAGATGATGTGGTGGGATCCTGCCAGCCAGTT-3′; for PAins 2: 5′-GGATCCTGCCAGCCAGTTGGCGTTGCCATGCCAGGTGCCGAAGATGATGTGGTGCCTTCCTGCCAGAAGTGT-3′: for PAins 3: 5′-TGGGTCACTCTTCAAGCTACTGGCGTTGCCATGCCAGGTGCCGAAGATGATGTGGTGCACTGCCTCCCAGAAGCCCAC-3′: and for PAins 4: 5′-CAAGCTACTCACTGCCTCCCAGGCGTTGCCATGCCAGGTGCCGAAGATGATGTGGTGGAAGCCCACGTCCTACGA-3′. The insertion positions are shown in Fig. 1c. To generate expression constructs of the PAins mutants, they were subcloned into an EcoRI/XbaI site of a lentivirus vector (CSII-EF-MCS; a gift from H. Miyoshi, RIKEN, Wako, Japan). A Rap1V12 mutant and Spa-1 were generated as previously described32. The fidelity of all the constructs was verified by sequencing.
Generation of a hybridoma producing mAbs G3 or H3
DNA encoding N-terminal 1–458 amino acids of murine β7 was subcloned into a pMALc2x vector and that vector was transformed into E.coli BL21 competent cells. To synthesize recombinant maltose binding protein (MBP)-β7, BL21 were cultured at 37 °C to reach an OD600 of 0.4–0.6, and then isopropyl β-D-thiogalactopyranoside was added to a concentration of 0.2 mM and incubated at 30 °C for 3 h. The cells were lysed in lysis buffer (0.1% Triton X-100, 20 mM Tris–HCl, 200 mM NaCl, and 1 mM EDTA). MBP-β7 was purified from cell lysates using a pMAL Protein Fusion and Purification System (New England Biolabs).
WKY rats (8 week old) were injected intramuscularly at the tail base with an antigen emulsion containing MBP-β7 and Freund’s complete adjuvant (BD Biosciences). Then, 2 weeks later, lymphocytes were collected from iliac lymph nodes and fused with a murine myeloma cell line, SP2/0, using GenomeONE (ISHIHARA SANGYO)33. Hybridoma clones producing mAbs against the active form of β7 were screened by flow cytometry of BAF cells using the hybridoma supernatant in the presence or absence of Mn2+.
RNA-mediated interference and gene introduction via lentiviral transduction
RNA-mediated interference was used to suppress mouse expression. As previously reported34, a 19-nucleotide -specific sense RNA sequences or a scrambled control RNA sequence of (Rap1a: 5′-GAATGGCCAAGGGTTTGCA-3′, Rap1b: 5′-AGACACTGATGATGTTCCA-3′, and talin: 5′-CGGTGAAGACTATCATGGT-3′) were introduced into BAF cells using a lentivirus vector with GFP (a gift from Dr. Miyoshi H., RIKEN, Wako, Japan) containing the RNAi construct under control of the H1 promoter cassette, respectively. The production and concentration of lentivirus particles were assessed as previously described35. The transduction efficiencies were greater than 90%. A GFP high population was collected by cell sorting and subjected to adhesion assays and immunoblot analysis.
Immunoblot analysis
BAF cells were lysed in buffer (1% Nonidet P-40, 150 mM NaCl, 25 mM Tris–HCl [pH 7.4], 10% glycerol, 2 mM MgCl2, 1 mM phenylmethylsulfonylfluoride, 1 mM leupeptin, and 0.1 mM aprotinin). Cell lysates were subjected to immunoblotting32.
Pull-down assays
Rap1-GTP was pulled down with a glutathione S-transferase (GST)-RBD of RalGDS fusion protein, respectively36. Briefly, 107 cells were lysed in ice-cold lysis buffer (1% Triton X-100, 50 mM Tris–HCl [pH 7.5], 100 mM NaCl, 10 mM MgCl2, 1 mM phenylmethylsulfonyl fluoride, 1 mM leupeptin, and 0.5 mM aprotinin) and incubated for 1 h at 4 °C with GST-fusion proteins coupled to glutathione agarose beads. The beads were washed three times with lysis buffer and subjected to immunoblot analysis using an anti-Rap1 antibody. Immunoblotting of total cell lysates (5 × 104 cells) was also performed.
Assessment of activation epitopes by mAbs staining
Immunofluorescence flow cytometry was performed as described previously31. For NZ-1, G3 or H3 mAbs staining, cells were washed with binding buffer (0.1% BSA, 1 mM CaCl2, 1 mM MgCl2 or 0.1% BSA, 0.5 mM MnCl2 in HBSS), resuspended in 50 μl of the same buffer, and incubated for 30 min at 37 °C with 10 μg/ml of each mAb in the presence or absence of 0.5 μM CXCL12, CCL25 or CXCL10. Mean fluorescence intensities were measured using a Gallios flow cytometry or CytoFLEX (Beckman Coulter).
Generation of β7-deficient BAF cells by the CRISPR/Cas9 system
The guide sequence targeting exon of the mouse β7 was cloned into pX330 (Addgene #42230)37. pX330-U6-Chimeric_BB-CBh-hSpCas9 was a gift from Feng Zhang (Addgene plasmid #42230, https://n2t.net/addgene:42230; RRID: Addgene_42230). pCAG-EGxxFP was used to examine efficiency of the target DNA cleavage by the guide sequence and Cas9 activity. The resultant guide sequence was cloned into GFP expressing plasmid DNA pX458 (Addgene #48138)38. pSpCas9(BB)-2A-GFP (PX458) was a gift from Feng Zhang (Addgene plasmid #48138, https://n2t.net/addgene:48138; RRID: Addgene_48138). The pX458 plasmid was transfected into BAF cells. 24 h after transfection, cells were sorted GFP-high population, followed by limiting dilution. Expression of full length β7 protein in each isolated clone was tested by flowcytometry. The sequence of the primer used were as follows: β7 Exon, Forward: 5′-GGGTCGACGCTGTGGAGTGAGTGAACTG-3′ and Reverse: 5′-GGGAATTCCTCTGAAGCCCAGTGCATTC-3′. Exon of guide sequence: Forward: 5′-CACCCACCTGGTCGCAGCGTGACG-3′ and Reverse: 5′-AAACCGTCACGCTGCGACCAGGTG-3′. Exon of β7 from edited clones was PCR amplified and verified by sequencing.
Epitope mapping of G3 and H3
Human/murine β7, murine β1/β7 chimeras or ∆1–19 murine β7 were constructed using an In-Fusion HD cloning kit (TaKaRa). The In-Fusion HD enzyme premix fuses multiple PCR-generated sequences and linearized vectors efficiently and precisely, utilizing a 20-bp overlap at their ends. This 20-bp-overlap allows complementary base pairs between two pieces of DNA to anneal, leading to fragment joining. Therefore, when individual DNA fragments derived from human β7, murine β1 or β7 were amplified by PCR, a 20-bp overlap was engineered by designing custom primers (Table S1). The objective DNA fragments with a 20-bp overlap were joined into a linearized CSII-EF-MCS-IRES2-venus vector. The constructs were transduced to β7-knockout BAF cells by lentivirus. The binding of G3 or H3 to the BAF cells expressing the chimera β7 was measured in the presence of 0.5 mM Mn2+ by flow cytometry.
Anti-MAdCAM-1 antibody treatment of colitis
T cell-specific Rap1a/b knockout mice aged 4 or 5 weeks were injected intraperitoneally with PBS containing 1 mg of rat IgG or anti-MAdCAM-1 antibody39 on days 0, 7, and 21. Their body weights were measured every 2 days. Pathological or frozen sections were prepared on day 28.
Histological examination
Colon sections were fixed in 10% buffered formalin and embedded in paraffin. Paraffin-embedded colon sections were cut (3 μm), stained with haematoxylin and eosin and examined on an Olympus IX51 light microscope equipped with a CCD (charge-coupled device) camera. Tissues were graded semiquantitatively as described before10,40. Histological grades were assigned in a blinded manner on a scale of 0–5, as follows: grade 0, no changes observed,grade 1, minimal scattered mucosal inflammatory cell infiltrates, with or without minimal epithelial hyperplasia,grade 2, mild scattered to diffuse inflammatory cell infiltrates, sometimes extending into the submucosa and associated with erosions, with mild to moderate epithelial hyperplasia and mild to moderate mucin depletion from goblet cells; grade 3, moderate inflammatory cell infiltrates that were sometimes transmural, with moderate to severe epithelial hyperplasia and mucin depletion; grade 4, marked inflammatory cell infiltrates that were often transmural and associated with crypt abscesses or occasional ulceration, with marked epithelial hyperplasia, mucin depletion; and grade 5, marked transmural inflammation with severe ulceration or loss of intestinal glands.
Immunostaining
Preparation of frozen sections of the colon from colitis mice were performed as described previously. Sections were blocked for 1 h at 20 °C with PBS containing 10% goat serum and 0.1% Triton X-100 and incubated overnight at 4 °C with APC conjugated anti-CD4 antibody. Nuclei were stained with SlowFade Gold antifade reagent with DAPI (invitrogen). Sections were examined on TCS SP8 (Leica).
Measurement of soluble MAdCAM-1 binding.
The binding of mouse MAdCAM-1-Fc to BAF cells was measured as described before10. Cells were suspended in 50 μl binding buffer (0.1% BSA, 1 mM CaCl2, 1 mM MgCl2 or 0.1% BSA, 0.5 mM MnCl2 in HBSS), and 2 × 105 cells/50 μl were then incubated with mouse MAdCAM-1-Fc (30 μg/ml)34. After two washes, samples were incubated for 20 min on ice with APC-conjugated mouse antibody to human IgG Fc-specific antibody (1 μg/ml). Unbound secondary antibody was removed by washing. Mean fluorescence intensities were measured using Gallios flow cytometry (Beckman Coulter).
Statistical analysis
Statistical analysis was performed using two-tailed Student’s t-test. P values less than 0.05 were considered significant.
References
Hogg, N., Patzak, I. & Willenbrock, F. The insider’s guide to leukocyte integrin signalling and function. Nat. Rev. Immunol.11, 416–426 (2011).
Humphries, J. D., Chastney, M. R., Askari, J. A. & Humphries, M. J. Signal transduction via integrin adhesion complexes. Curr. Opin. Cell Biol.56, 14–21 (2019).
Lock, J. G. et al. Clathrin-containing adhesion complexes. J. Cell Biol.218, 2086–2095 (2019).
Yu, Y. et al. Structural specializations of alpha(4)beta(7), an integrin that mediates rolling adhesion. J. Cell Biol.196, 131–146 (2012).
Firrell, J. C. & Lipowsky, H. H. Leukocyte margination and deformation in mesenteric venules of rat. Am. J. Physiol.256, H1667-1674 (1989).
Kinashi, T. & Katagiri, K. Regulation of lymphocyte adhesion and migration by the small GTPase Rap1 and its effector molecule, RAPL. Immunol. Lett.93, 1–5 (2004).
Katagiri, K., Maeda, A., Shimonaka, M. & Kinashi, T. RAPL, a Rap1-binding molecule that mediates Rap1-induced adhesion through spatial regulation of LFA-1. Nat. Immunol.4, 741–748 (2003).
Lafuente, E. & Boussiotis, V. A. Rap1 regulation of RIAM and cell adhesion. Methods Enzymol.407, 345–358 (2006).
Gingras, A. R. et al. Rap1 binding and a lipid-dependent helix in talin F1 domain promote integrin activation in tandem. J. Cell Biol.218, 1799–1809 (2019).
Ishihara, S. et al. Dual functions of Rap1 are crucial for T-cell homeostasis and prevention of spontaneous colitis. Nat. Commun.6, 8982 (2015).
Bachmann, M., Kukkurainen, S., Hytonen, V. P. & Wehrle-Haller, B. Cell adhesion by integrins. Physiol. Rev.99, 1655–1699 (2019).
Schurpf, T. & Springer, T. A. Regulation of integrin affinity on cell surfaces. EMBO J.30, 4712–4727 (2011).
Kim, M., Carman, C. V. & Springer, T. A. Bidirectional transmembrane signaling by cytoplasmic domain separation in integrins. Science301, 1720–1725 (2003).
Xiao, T., Takagi, J., Coller, B. S., Wang, J. H. & Springer, T. A. Structural basis for allostery in integrins and binding to fibrinogen-mimetic therapeutics. Nature432, 59–67 (2004).
Zhang, K. & Chen, J. The regulation of integrin function by divalent cations. Cell Adh. Migr.6, 20–29 (2012).
Qi, J. et al. Identification, characterization, and epitope mapping of human monoclonal antibody J19 that specifically recognizes activated integrin alpha4beta7. J. Biol. Chem.287, 15749–15759 (2012).
Hosen, N. et al. The activated conformation of integrin beta7 is a novel multiple myeloma-specific target for CAR T cell therapy. Nat. Med.23, 1436–1443 (2017).
Fujii, Y. et al. Tailored placement of a turn-forming PA tag into the structured domain of a protein to probe its conformational state. J. Cell Sci.129, 1512–1522 (2016).
Arimori, T. et al. Fv-clasp: An artificially designed small antibody fragment with improved production compatibility, stability, and crystallizability. Structure25, 1611-1622.e4 (2017).
Zhu, L. et al. Structure of Rap1b bound to talin reveals a pathway for triggering integrin activation. Nat. Commun.8, 1744 (2017).
Mora, J. R. et al. Selective imprinting of gut-homing T cells by Peyer’s patch dendritic cells. Nature424, 88–93 (2003).
Sun, H. et al. Distinct chemokine signaling regulates integrin ligand specificity to dictate tissue-specific lymphocyte homing. Dev. Cell30, 61–70 (2014).
Wang, S. et al. Integrin alpha4beta7 switches its ligand specificity via distinct conformer-specific activation. J. Cell Biol.217, 2799–2812 (2018).
Li, J. et al. Conformational equilibria and intrinsic affinities define integrin activation. EMBO J.36, 629–645 (2017).
Nordenfelt, P., Elliott, H. L. & Springer, T. A. Coordinated integrin activation by actin-dependent force during T-cell migration. Nat. Commun.7, 13119 (2016).
Honda, S. et al. Topography of ligand-induced binding sites, including a novel cation-sensitive epitope (AP5) at the amino terminus, of the human integrin beta 3 subunit. J. Biol. Chem.270, 11947–11954 (1995).
Pokharel, S. M. et al. Integrin activation by the lipid molecule 25-hydroxycholesterol induces a proinflammatory response. Nat. Commun.10, 1482 (2019).
Byron, A. et al. Anti-integrin monoclonal antibodies. J. Cell Sci.122, 4009–4011 (2009).
Wynne, J. P. et al. Rap1-interacting adapter molecule (RIAM) associates with the plasma membrane via a proximity detector. J. Cell Biol.199, 317–330 (2012).
Katagiri, K. et al. Crucial functions of the Rap1 effector molecule RAPL in lymphocyte and dendritic cell trafficking. Nat. Immunol.5, 1045–1051 (2004).
Katagiri, K. et al. Rap1 is a potent activation signal for leukocyte function-associated antigen 1 distinct from protein kinase C and phosphatidylinositol-3-OH kinase. Mol. Cell Biol.20, 1956–1969 (2000).
Katagiri, K., Hattori, M., Minato, N. & Kinashi, T. Rap1 functions as a key regulator of T-cell and antigen-presenting cell interactions and modulates T-cell responses. Mol. Cell Biol.22, 1001–1015 (2002).
Sado, Y., Inoue, S., Tomono, Y. & Omori, H. Lymphocytes from enlarged iliac lymph nodes as fusion partners for the production of monoclonal antibodies after a single tail base immunization attempt. Acta Histochem. Cytochem.39, 89–94 (2006).
Ebisuno, Y. et al. Rap1 controls lymphocyte adhesion cascade and interstitial migration within lymph nodes in RAPL-dependent and -independent manners. Blood115, 804–814 (2009).
Katagiri, K., Imamura, M. & Kinashi, T. Spatiotemporal regulation of the kinase Mst1 by binding protein RAPL is critical for lymphocyte polarity and adhesion. Nat. Immunol.7, 919–928 (2006).
Katagiri, K., Shimonaka, M. & Kinashi, T. Rap1-mediated lymphocyte function-associated antigen-1 activation by the T cell antigen receptor is dependent on phospholipase C-gamma1. J. Biol. Chem.279, 11875–11881 (2004).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science339, 819–823 (2013).
Ran, F. A. et al. Genome engineering using the CRISPR-Cas9 system. Nat. Protoc.8, 2281–2308 (2013).
Picarella, D. et al. Monoclonal antibodies specific for beta 7 integrin and mucosal addressin cell adhesion molecule-1 (MAdCAM-1) reduce inflammation in the colon of scid mice reconstituted with CD45RBhigh CD4+ T cells. J. Immunol.158, 2099–2106 (1997).
Chen, H. et al. Extracellular vesicles from apoptotic cells promote TGFbeta production in macrophages and suppress experimental colitis. Sci. Rep.9, 5875 (2019).
Acknowledgements
We would like to thank Ms. N. Saho for technical assistance. This work was supported by Japan Society for the Promotion of Science KAKENHI 16K19163 and 19K07612, Takeda Science Foundation, The Naito Foundation. Kitasato University Research Grant for young researchers. Suzuken Memorial Foundation.
Author information
Authors and Affiliations
Contributions
T.S. and K.K. designed, performed experiments, and wrote the paper. S.I., and R.M. performed the experiments. J.T. contributed to the preparation of essential materials and commented on the experiments and paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sato, T., Ishihara, S., Marui, R. et al. Dissection of α4β7 integrin regulation by Rap1 using novel conformation-specific monoclonal anti-β7 antibodies. Sci Rep 10, 13221 (2020). https://doi.org/10.1038/s41598-020-70111-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-70111-0
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.