Abstract
Intramuscular injection and electroporation of naked plasmid DNA (IMEP) has emerged as a potential alternative to viral vector injection for transgene expression into skeletal muscles. In this study, IMEP was used to express the DUX4 gene into mouse tibialis anterior muscle. DUX4 is normally expressed in germ cells and early embryo, and silenced in adult muscle cells where its pathological reactivation leads to Facioscapulohumeral muscular dystrophy. DUX4 encodes a potent transcription factor causing a large deregulation cascade. Its high toxicity but sporadic expression constitutes major issues for testing emerging therapeutics. The IMEP method appeared as a convenient technique to locally express DUX4 in mouse muscles. Histological analyses revealed well delineated muscle lesions 1-week after DUX4 IMEP. We have therefore developed a convenient outcome measure by quantification of the damaged muscle area using color thresholding. This method was used to characterize lesion distribution and to assess plasmid recirculation and dose–response. DUX4 expression and activity were confirmed at the mRNA and protein levels and through a quantification of target gene expression. Finally, this study gives a proof of concept of IMEP model usefulness for the rapid screening of therapeutic strategies, as demonstrated using antisense oligonucleotides against DUX4 mRNA.
Similar content being viewed by others
Introduction
Electroporation (EP), also named electro-transfection, is a non-viral method allowing an enhanced cellular uptake of various type of exogenous molecules such as DNA, RNA, proteins or chemicals. The procedure is based on the application of short electric pulses that transiently permeabilize cell membrane permitting cellular and nuclear entrance of large particles1,2,3,4. Neumann et al. were the first to report in 1982 that pulsed electric fields could efficiently introduce linear or circular DNA into mouse lyoma cells in culture5. The mechanisms by which EP facilitates DNA transport across cell membrane, cytoplasm and nuclear membranes are still debated. Electric pulses induce electrophoretic forces that facilitate migration of charge-carrying molecules, such as naked DNA plasmids (pDNA), maximizing their interaction with cell membranes6. Cells exposed to an electrical field present a change in transmembrane potential. At a critical threshold value, a re-orientation of membrane phospholipids occurs, leading to the formation of small hydrophilic openings called electropores. These breakdowns are reversible and allow water, ions and membrane-impermeable molecule flow2,3,7,8. By this way, pDNA can enter into cells and its encoded transgene can be expressed. However, recent studies demonstrated the importance of endocytosis pathways (both clathrin- and caveolin-mediated endocytosis) for DNA internalization following EP3,4,9,10, notably in mouse muscles11. Since its discovery, numerous advances were made in the field and EP was applied with success both in vitro and in vivo in various cell and tissue types. Today, EP has many biomedical applications. Most studies are related to anticancer drug delivery, also called electrochemotherapy, using intratumoral injection coupled with EP to enhance cellular uptake of a therapeutic agent presenting high intrinsic toxicity but low plasma membrane permeability12,13,14. Besides its direct clinical application in the field of cancer, EP is broadly used as a gene delivery tool15 that can be applied either for vaccination16,17,18,19,20, immunotherapy (especially in cancer applications21,22), gene therapy23,24,25,26, genome editing27,28 and generation of induced pluripotent stem cells29,30. Number of phase I and II clinical trials are underway or have been completed, demonstrating the safety and efficacy of this procedure3. EP has several advantages compared to other tools for gene delivery. Indeed, pDNA are easy to modify and prepare, inexpensive to produce in large scale, and may be administered multiple times without significant inflammation or immune response. pDNA are injected as “naked” molecules meaning that no additional chemicals are associated, limiting risks and undesirable side effects1,2. Unlike viral vectors, there is no safety considerations1,31. EP procedure is relatively simple and easy to set up and does not need expensive instrumentation. This procedure allows increased levels of gene transfer and expression, nearing those of viral vector, and is applicable from cells transfection to drug and therapeutic gene delivery into living tissues from rodents to humans2,3.
Skeletal muscles constitute attractive targets for gene therapy, due to their accessibility and high vascularization. Notably, skeletal muscles were used for systemic delivery of therapeutic protein such as erythropoietin32, coagulation factors33 or anti-inflammatory cytokines34. Muscle EP protocols, first described in35, have been optimized over time to improve transfection efficacy and transgene expression level1,36,37,38,39,40. In the context of muscle dystrophies, EP methodology was explored as a potential route for treatment. EP was notably reported as efficient to transduce constructs encoding dystrophin in the mdx mouse and dog models of Duchenne Muscular Dystrophy (DMD)41,42. Contrary to DMD, resulting from the loss of dystrophin, Facioscapulohumeral muscular dystrophy (FSHD)43 is a gain of function disease caused by the inappropriate expression in skeletal muscle of DUX4, a gene normally only expressed in germline and early embryogenesis44,45,46,47,48,49,50,51,52,53,54,55. The DUX4 gene encodes a transcription factor that deregulates a large molecular network53,54,55,56,57,58,59,60. However, the precise mechanisms by which DUX4 leads to clinical symptoms still must be clarified. Even though various therapeutic strategies are emerging, there is currently no curative treatment for FSHD. Several drug-based therapies aiming either for muscle improvement (anti-inflammatory approach61, β2-adrenergic agonists62,63,64,65, antioxidants66) or inhibition of DUX4 expression such as mitogen-activated protein kinase inhibitors67 (Losmapimod, Fulcrum therapeutics, NCT04003974 ) are investigated in clinical trials68,69. In parallel, gene therapy has been explored to reduce or avoid DUX4 protein expression and/or activity by controlling D4Z4 locus methylation70,71, or to silence DUX4 mRNA. Among those, antisense oligonucleotides (AOs) and siRNA targeting the DUX4 mRNA and preventing its translation have been developed and successfully tested in vitro and in vivo54,72,73,74,75. Development of in vivo proof-of-concept studies for emerging therapies are now required as a next step towards clinical trials. Several hurdles such as DUX4 toxicity and its stochastic low expression have made the generation of an animal model recapitulating all the pathophysiological aspects of FSHD very challenging. FSHD-like mouse models have now been described, each of them possessing their own advantages and limits59,76,77,78,79,80,81,82. Some of these mouse models allow inducible conditional DUX4 expression, bypassing DUX4 high toxicity during embryonic development and enabling mice to grow up and develop muscular dystrophy78,79,80,83. These models open new ways to investigate molecular mechanisms leading to FSHD symptoms. However, the use of inducible transgenic models is often time consuming and costly. It is especially an issue in the current context of FSHD where high throughput molecule screenings are required to identify new potential therapeutics. In the present study, we describe a convenient in vivo model of DUX4 local muscle expression using an EP procedure. This model is simple, unexpansive, reproducible and associated with an easy read out that facilitates quantitative analysis. Therefore, this model can be useful at the forefront for high throughput therapeutic screening.
Results
Hyaluronidase pre-treatment improves gene expression following naked DNA injection and electroporation in vivo
Mouse TA muscles were injected with the pCMV-lacZ reporter plasmid and then electroporated (IMEP procedure). In order to determine whether muscle pre-treatment with hyaluronidase (which digests hyaluronic acid, a major constituent of the extracellular matrix) could modify gene electroporation efficacy, naked DNA injection was preceded (hIMEP group) or not (IMEP group) by an intramuscular injection (IM) of hyaluronidase. TA muscles were harvested 7 days after injection. The reporter expression level was evaluated by X-gal staining of β-galactosidase activity on cryosections from proximal, medial and distal muscle regions and averaged for each group. As observed in Fig. 1, hyaluronidase pre-treatment significantly improved gene electroporation efficiency as shown by the increased β-galactosidase-positive (β-gal+) muscle area (Fig. 1A–C). Quantification of β-gal+ muscle surface showed a three-fold increase in mouse TA pre-treated with hyaluronidase, with median value of 36.4% (p < 0.001, hIMEP vs IMEP, Fig. 1C). The hIMEP procedure was therefore applied in the next steps of the study.
Hyaluronidase pre-treatment improves β-galactosidase expression in mouse Tibialis Anterior muscle (TA). (A,B) Representative sections of TA electroporated (A) without hyaluronidase pre-treatment (IMEP) or (B) with hyaluronidase pre-treatment 2 h before the electroporation procedure (hIMEP). TA muscles were injected by IMEP or hIMEP with 40 µg of pCMV-lacZ reporter plasmid. TA were harvested 1-week post-injection and cryosections stained with X-gal (blue) and counterstained with Eosin (pink). Scale 500 µm. (C) Percentage of surface area expressing β-galactosidase (β-gal+) quantified by color thresholding using ImageJ. Data are represented as boxplots, ****p < 0.0001 Mann–Whitney Rank Sum Test; n = 4 for each group. The graph was generated using GraphPad Prism 6.01.
TA electroporation with DUX4-expression plasmid induces readily quantifiable muscle lesions
Because of the known DUX4 toxicity in human FSHD muscles, we first checked whether a DUX4 local expression induced by the hIMEP procedure could impact muscle structure in mice. To this aim, the TA muscles were injected and electroporated with pCIneo-DUX4 expression plasmid using the hIMEP procedure. The empty pCIneo plasmid was used as negative control. In both conditions, a simultaneous injection of the pCMV-lacZ reporter vector was performed to facilitate the location of muscle areas having incorporated the transgenes. Mice were sacrificed 1 week later and TA muscles were harvested, quickly frozen, cryosectioned and stained with X-gal to detect reporter gene activity, or with Hematoxylin–Eosin–Heidenhain blue (HEB) for histological evidence of muscle damage.
One-week post-injection, TA muscles electroporated with the control plasmid presented a normal histological structure with peripheral nuclei in geometric fibres surrounded by a thin layer of endomysial extracellular matrix, as shown by HEB staining (Fig. 2A). A restricted area exhibiting some smaller fibres, central nuclei and a slight focal inflammatory infiltrate around the site of injection (β-gal+ area) was sometimes observed (data not shown). X-gal staining in control TA confirms electroporation efficiency as shown by the presence of grouped β-gal+ fibres (Fig. 2A, Box). In contrast, TA muscles electroporated with the DUX4-expression plasmid showed a large damaged region containing atrophic fibres with high size variability, some regenerating fibres characterized by central nuclei, a marked inflammatory infiltrate and an accumulation of conjunctive tissue (fibrosis) (Fig. 2B; SI Fig. S1 online). Interestingly, X-gal staining in adjacent sections showed a lower number of β-gal+ myofibres with a different spatial distribution as compared to TA muscles injected with the control plasmid (Fig. 2B, Box). Indeed, a median value of 2.1% of the total muscle surface was positive for β-gal in the DUX4 expression group against 27.97% in the control group (p < 0.001, Fig. 2C). As expected, lesion area quantification on HEB-colored total cryosections indicated that the percentage of damaged surface area was significantly higher in TA muscles injected with the DUX4-expression plasmid as compared to the control group (p < 0.001) with a median value of 25.1 and 8.5%, respectively (Fig. 2D).
hIMEP of DUX4-expression plasmid induces muscle lesions. (A,B) Representative sections of TA electroporated with 10 µg of pCMV-lacZ and 40 µg of (A) pCIneo or (B) pCIneo-DUX4 plasmids. TA muscles were harvested 1-week post injection and cryosections from distal, medial and proximal regions stained with X-gal to assess β-gal+ areas (small pictures). Adjacent sections were stained with HEB coloration for muscle damages evaluation (large pictures). Scales 50 µm (red) and 100 µm (black). (C,D) Percentage of surface area (C) expressing β-galactosidase (β-gal+) and (D) damaged in mouse TA 1-week post hIMEP using 10 µg of pCMV-lacZ concomitant with 40 µg of pCIneo (Ctrl) or pCIneo-DUX4 (DUX4) plasmids. β-gal+ and lesion area percentage were evaluated on total section stained with X-gal or HEB respectively, and quantified by color thresholding using ImageJ. (E) Recirculation test. Both TA from same mouse were electroporated, one using 40 µg of pCIneo-DUX4, the other saline solution. One week after hIMEP, lesion area was evaluated on HEB stained TA cryosections by color thresholding using ImageJ. All results are presented as boxplots, **p < 0.01 and ****p < 0.0001 Mann–Whitney Rank Sum Test; n = 5 for each group except in (E) n = 2. Graphs were generated using GraphPad Prism 6.01.
We also investigated the plasmid recirculation potential following hIMEP. Indeed, we were wondering whether the recirculation of the DUX4 expression plasmid would be sufficient to cause transfection of the contralateral TA resulting in quantifiable lesions. To this aim, both TA muscles from the same mouse were electroporated, one using pCIneo-DUX4, the other one using saline solution. One week after hIMEP, quantification of the damaged area on HEB-stained cryosections showed no detectable lesion in the saline-injected TA, in contrast to the contralateral TA injected with the DUX4-expression plasmid which exhibited a significant lesion area (Fig. 2E). This suggested that no plasmid recirculation occurred, allowing an independent treatment of both TA from the same animal.
Dose–response analysis
A dose–response analysis was performed by electroporating mouse TA muscles with increasing doses of the DUX4-expression plasmid or control plasmid, concomitantly with the pCMV-lacZ reporter vector. The quantification of muscle damages (HEB staining) was (1) reported either to total muscle section or (2) to the injected area, corresponding to the β-gal+ region. Regardless of the quantification method, no statistical difference was observed following the injection of different doses of the control plasmid (1–20–40 µg pCIneo; one-way ANOVA on the ranks followed by Dunn’s post hoc test, p = 0.959 and 0.153 in total and injected area, respectively, data not shown). The data from these three control groups were therefore pooled in a single control group (Fig. 3). Regarding the muscle damage quantification reported to total section (Fig. 3A), a significant increase of damaged surface between the control group and all DUX4 groups was demonstrated (p < 0.01). Similar results were obtained when the injected area was the only area considered for the quantification (Fig. 3B). However, there was no significant difference in the percentage of damaged area obtained after injection of increasing doses of pCIneo-DUX4 using either quantification system (p = 0.894 and 0.957, reported to total section or to injected area, respectively). Data from the four DUX4 expression groups have thus been grouped (in Fig. 3C) to compare results obtained from both quantification methods. As expected, the percentage of damaged muscle surface area was higher when quantified within the injected region, as compared to the percentage calculated on total muscle section. This difference is highly statistically relevant in the DUX4 group (p < 0.0001) and significant in the control groups (p = 0.01) where the altered muscle area was limited to 7.4% of the total muscle section and to 7.3% when calculated in the injected region (Fig. 3C).
Dose–response of muscle lesion area in mouse TA 1-week post hIMEP procedure. (A,B) Lesion area percentage was evaluated (A) on total cryosection or (B) in the injected area (defined by β-gal+ region) from distal, medial and proximal part of TA electroporated with different doses of control or DUX4-expression plasmid and 10 µg of pCMV-lacZ. Sections were stained with HEB and lesions were quantified by color thresholding using ImageJ. Control groups (Ctrl) include 1-, 20-, 40-µg pCIneo-injected groups as there was no statistical difference among them (one-way ANOVA followed by Dunn’s post hoc test, p = 0.959 and 0.153 for total and injected region quantifications respectively, NS). (C) Comparison of injured area percentages quantified on total section (black) or in injected area (defined by the β-gal+ region) (grey). DUX4 groups include 1-, 5-, 20-, 40-µg pCIneo-DUX4-injected groups as there was no statistical difference among them (one-way ANOVA followed by Dunn’s post hoc test, p = 0.894 and 0.957 for total and injected regions respectively, NS). All results are presented as boxplots, ****p < 0.0001, **p < 0.01, (vs ctrl (A,B) or as indicated (C)). (A,B) One-way ANOVA on the ranks followed by Dunn’s post hoc test; n = 3, except 5 µg DUX4 (n = 4), 40 µg DUX4 (n = 5) and Ctrl (n = 11). (C) Wilcoxon signed rank test; n = 11 (ctrl) or 15 (DUX4). Graphs were generated using GraphPad Prism 6.01.
Distribution of the lesions through Tibialis Anterior muscles
In order to investigate DUX4 expression distribution through TA following hIMEP with pCIneo-DUX4, muscle damages were quantified in 3 different TA regions (proximal, medial and distal region). Since no statistically significant difference was observed between tested doses considering each region individually (One way Anova, p = 0.871, 0.821 and 0.395 in the proximal, medial and distal region, respectively, data not shown), data corresponding to the 1-, 5-, 20-, 40 µg DUX4 plasmid groups were respectively pooled for each muscle region. As illustrated in Fig. 4, at 7 days post-injection, the percentage of damaged muscle area in the proximal and medial part of the TA were not statistically different, with respective medians of 47.11% and 46.69% (p: NS, Friedman Repeated Measures Analysis of Variance on Ranks). Regarding the distal muscle part, the percentage of lesion was significantly lower than in the other muscle regions (p < 0.05) with a median of 25.6% (Fig. 4). Considering these results, the subsequent analyses were based on the proximal and medial TA parts to minimize variability.
Lesion distribution through TA regions. Mouse TAs were electroporated with different doses of pCIneo-DUX4 (1, 5,10, 20 and 40 µg) and 10 µg of pCMV-lacZ. Muscle damaged area 1 week following TA hIMEP was evaluated in the injected area (β-gal+) of HEB stained cryosections from distal, medial and proximal regions of TA (color thresholding, ImageJ). Since no statistical difference was observed between tested doses considering each region individually, data were respectively pooled to form single proximal, medial and distal groups (data not shown, One-way Anova, p = 0.871, 0.821 and 0.395 in proximal, medial and distal region respectively). All data are presented as boxplots, *p < 0.05 and **p < 0.01. Friedman Repeated Measures Analysis of Variance on Ranks; n = 15 for each group. Graphs were generated using GraphPad Prism 6.01.
Time-course analysis
The development of the TA lesions was evaluated 1-, 3- and 7-days after the hIMEP procedure by using 1 µg of the DUX4-expression plasmid and a concomitant injection of pCMV-lacZ reporter. Muscle damages were then evaluated in TA proximal and medial parts by focusing on the injected area (β-gal+ region). At 1- and 3-days after injection, a slight thickening of the extracellular matrix and a straight mark with some mononuclear cells and regenerating fibres (probably corresponding to the injection site) were observed. This led to a global ‘lesion’ percentage defined by HEB staining of 16.3% and 12.9% at 1- and 3-days post-injection, respectively. At 7 days after electroporation, extensive injuries, a marked inflammatory infiltrate and an extracellular matrix expansion were detected. At this time point, damaged surface area represented 47.2% of the injected area (Fig. 5, median value, p < 0.05 7-days vs 1- and 3-days).
Time-course analysis. TA muscles were electroporated with 1 µg of pCIneo-DUX4 and 10 µg of pCMV-LacZ. Lesion area was evaluated 1-, 3- or 7-days after hIMEP in the injected area (β-gal+) on HEB stained cryosections of medial and proximal TA regions (color thresholding, ImageJ). Results are presented as boxplots, *p < 0.05 and **p < 0.01. One-way ANOVA on the ranks followed by Dunn’s post hoc test; n = 3 for each group. Scale 1 mm. The graph was generated using GraphPad Prism 6.01.
Expression of DUX4 and its target genes in hIMEP mice
DUX4 expression at the mRNA level was investigated by 3′RACE in hIMEP treated mice. In addition, mRNA levels of two DUX4 target genes (Wfdc3, Zscan4c), commonly used as biomarkers of DUX4 transcriptional activity78,79,80, were quantified along with the mRNA level of Myog a myogenic marker known to be downregulated in FSHD, contrary to Wfdc3 and Zscan4c84,85,86. To this aim, total mRNAs were extracted from mouse TAs treated by hIMEP (1-, 3- and 7-days) using 1 µg of pCIneo-DUX4 or control plasmid pCIneo concomitant with reporter vector pCMV-lacZ. The 3′RACE products were separated by electrophoresis on agarose gel and 3 distinct product lengths (~ 1,350 bp, ~ 550 bp and ~ 350 bp) were observed. These fragments were only detected in the DUX4 expression group and at all investigated time points. Interestingly, the 550-bp fragment presented with the highest intensity and frequency (8/12 tested samples), suggesting a higher expression level of the corresponding mRNA as compared to the others. No specific fragment was detected in control mice (Fig. 6; SI Fig. S2 online). All PCR products were cloned, sequenced and analyzed to ensure DUX4 specificity. We also confirmed, by in silico analysis, presence of the target sequence for antisense oligonucleotide pLAM3A (− 12 + 13) (described in54,72) in the 550-bp sequences. DUX4 expression was also evaluated at the protein level by immunofluorescence (Fig. 7) confirming the presence of DUX4-positive nuclei, only in TA muscles injected with pCIneo-DUX4 at all investigated time-points. RT-qPCR analysis of DUX4 target gene mRNAs highlighted a statistical increase of Wfdc3 and Zscan4c expression at each time point in the DUX4 group compared to the control group (p < 0.001). However, there was no statistical difference between time points for these genes. Myog mRNA levels were stable in control groups regardless of the time point but significantly decreased at day 1 post hIMEP in the DUX4 expression group. Myog expression then increased over time. However, we could not detect any significant difference between DUX4 and control groups at 3- and 7-days post hIMEP (Fig. 8).
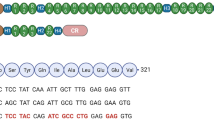
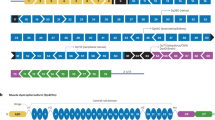
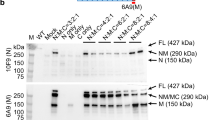
Confirmation of DUX4 mRNA expression by 3′RACE. (A) Nested PCR was used to amplify the 3′ end of DUX4 transcripts from total RNA extracted from TA muscles electroporated with 1 µg of control (left panel) or DUX4-expression pDNA (right panel). Representative cropped gels. Lanes 1 and 2 come from one gel; lanes 3, 4 and 5 come from a second gel ran in parallel. Corresponding full-length gels are available in supplementary information (SI Fig. S2 online). 3′RACE were performed 1-, 3- and 7-days post hIMEP. Wells (−) show negative controls without retro-transcription. Several fragments were detected (~ 1,350 bp, ~ 550 bp and ~ 350 bp; colored arrows) in each time point (not shown), cloned, sequenced and analyzed to confirm DUX4 specificity. (B) Schematic representation (not to scale) of 3′RACE products analysis. In silico alignment confirmed conservation of vPMO [pLAM3A (− 12 + 13)] target sequence (black line).
Confirmation of DUX4 protein expression by Immunofluorescence. Mouse TAs were injected by hIMEP with 10 µg pCMV-lacZ plasmid concomitantly with 40 µg either pCIneo-DUX4 or pCIneo, or with a saline solution (negative control), as indicated. Cryosections were analyzed 1, 3 and 7 days post-injection by immunofluorescence with antibodies directed against either DUX4 (E5-5) (red) or laminin α2 (green) to stain the myofibre basal lamina (for details, see “Methods”). DAPI was used to visualize nuclei (blue). Pictures were taken with a Nikon Eclipse 80i microscope and merged using NIS-Elements software. Scale bar 50 µm.
RTqPCR analyses of DUX4 target genes (Wfdc3, Zscan4c) and Myog expression level in TA muscle. TAs were injected by hIMEP with either 1 µg of control pCIneo DNA, 1 µg of pCIneo-DUX4- or with 50 µl of saline solution and harvested 1-, 3- or 7-days post injection. Total RNA was extracted with Trizol and qPCRs were performed in duplicates using SYBR Green FastStart Essential DNA Green Master. Analyzes were performed with LightCycler 96 software. At each time, results obtained from control plasmid and saline solution injected groups were pooled to form a single control group, as no statistical difference could be highlighted between them for all tested genes (one-way ANOVA on the ranks followed by Dunn’s post hoc test). Results are presented as fold difference to Rplp0 for Wfdc3 and Zscan4c and as relative to control for Myog. All data are presented as boxplots, *p < 0.05, **p < 0.01, ***p < 0.001 and #p < 0.05 vs DUX4 7 days, One-way ANOVA on the ranks followed by Dunn’s post hoc test n = 14 (Ctrl) or 7 (DUX4) (Wfdc3 and Zscan4c) and n = 8 (ctrl) or 4 (DUX4) (Myog). Graphs were generated using GraphPad Prism 6.01.
Preliminary evaluation of antisense oligonucleotide injected into DUX4 hIMEP mouse
Given that the DUX4 hIMEP mouse model was developed in order to easily test the efficiency of therapeutics in vivo, we investigated antisense oligonucleotide pLAM3A (− 12 + 13) (described in54,72) with the vPMO (octa-guanidine conjugated Phosphorodiamidate Morpholino Oligomer) chemistry as a proof-of-concept study. This vPMO silences DUX4 mRNA translation by interfering with intron II splicing. Its efficiency was previously demonstrated in vitro54 and in vivo72 in another mouse model87. To this aim, mouse TA muscles were electroporated using 20 µg of pCIneo-DUX4 plasmid by hIMEP and treated 6 h later with a single intraperitoneal injection of 250 µg of vPMO pLAM3A − 12 + 13. Seven days after injection, a highly significant 2.5-fold decrease of histological lesion area was observed in TA muscles of mice treated with the vPMO, as compared to untreated hIMEP DUX4 mice (p < 0.001) (Fig. 9).
vPMO treatment of mice electroporated with DUX4-expression plasmid decreases TA muscle lesions. TA muscles were injected by hIMEP with 20 µg of pCineo-DUX4 plasmid. Six hours later, mice received either an intraperitoneal injection of 250 µg of vPMO pLAM3A (− 12 + 13) targeting DUX4 mRNA (vPMO group) or no supplemental treatment (DUX4 group). Percentage of lesion area was evaluated 7 days post IMEP by color thresholding using ImageJ on HEB colored cryosections of TA proximal and medial parts. Results are presented as Box plots, ***p < 0,001 by Mann–Whitney Rank Sum Test; n = 4 for each group. The graph was generated using GraphPad Prism 6.01. Black arrows show fibrosis, white arrows atrophic fibres, black circles inflammatory infiltrate. Scales 1 mm (black) 250 µm (green), 100 µm (yellow) and 50 µm (red).
Discussion
Non-viral methods for transgene delivery have received increased attentions thanks to their relative safety, simplicity and cost-effectiveness compared to the administration of viral vectors which were found associated with cytotoxicity, inflammation and immune response1,2. Initially, the main drawback of naked pDNA gene transfer was its low transfection efficiency limiting its use in fundamental research or therapeutic applications. Indeed, as there are no specific DNA transporters at the surface of mammalian cells, naked plasmids can only be internalized by non-specific endocytosis. As this is a slow process, most of the injected DNA is retained into extracellular space and degraded by endogenous DNAses88. Nevertheless, electroporation (EP) was shown to improve transgene uptake and expression both in vitro and in vivo2,3,7. Previous studies have demonstrated the importance of various factors influencing efficacy and safety of pDNA EP in several tissues, inter alia, skeletal muscles. These factors include electrical parameters (intensity, duration, pulse number and frequency), electrode type and geometry, plasmid features including its length and GC content and finally, additional elements such as the host age or tissue permeability (e.g. normal vs DMD muscle41). In the present study, we applied optimized EP parameters according to Mir et al.89 who have demonstrated the influence of voltage-to-distance (between both electrodes) ratio, pulse length, number and frequency for muscle EP in vivo. Their observations suggested that 8 pulses of 200 V/cm, 20 ms and 2 Hz was an optimal combination for high transgene expression in mouse skeletal muscle. Those parameters have been largely and successfully used in other studies1,3,38,40,41,90,91. Different types of electrode can be used, such as plate electrodes, penetrating needle electrodes or multi-electrode arrays2,3,38. Here, we chose to use plate electrodes which have the advantage to apply a large and uniform electrical field92,93, to be less invasive and less damaging than the needle one38,40. Furthermore, the efficiency of gene transfer essentially depends on the effective DNA electrophoresis and distribution in the area where the electrical field is applied. However, the extracellular matrix (ECM) surrounding the myofibre surface limits a homogenous distribution of pDNA and thereby reduces transfection efficiency. Different enzymes modifying connective tissue permeability such as hyaluronidase or collagenase were tested to improve the distribution of naked pDNA33,39,40,94,95,96. Hyaluronidase is a mucolytic enzyme that depolymerizes N-acetyl-hexosamine glycosidic bonds in hyaluronic acid, interfering with its barrier function in the ECM96. Intact hyaluronic acid binds water molecules and forms a gelatin-like matrix that provides a structural support for the surrounding tissue. Hyaluronic acid fragments resulting from hyaluronidase digestion are still able to bind water molecules but fail to form a functional viscous gel. Consequently, the density of glycosaminoglycans is reduced, allowing a better DNA electrophoresis, improving its contact with the plasma membrane, thus increasing the transfection rate95,97. Consistent with previous studies, we observed an increased reporter gene (pCMV-lacZ) expression level when hyaluronidase was injected 2 h before the IMEP procedure. Histological analysis did not reveal any apparent muscle alterations linked to hyaluronidase pre-treatment, consistent with other observations36,39,94,95,96. Moreover, hyaluronidase is currently used in clinical applications96 supporting that it is not by itself harmful for tissue.
In this study, the EP procedure was used in order to develop a simple mouse model of DUX4 local expression in skeletal muscle, with the end purpose of rapid therapeutic testing for FSHD. The development of a mouse model recapitulating the full FSHD phenotype and molecular signature is very challenging due to the need for DUX4 precise expression timeframe in the early embryo98,99. Attempts at the generation of transgenic mouse models revealed that a tight regulation of DUX4 expression was essential to obtain viable mice able to reach adulthood and to develop a dystrophic phenotype76,77. The most recently developed murine models allow a controlled DUX4 expression: two of these are the iDUX4 mouse [2.7]77 and its improved version iDUX4pA78, both doxycycline-regulated. In addition the FLExDUX479,83 and the TIC-DUX480 mice allow conditional expression of human DUX4-fl following Cre recombination. When crossed with the appropriate cre-driver line (tamoxifen-inducible cre line, cre expressed from a muscle-specific promoter), those models allow the appearance of a progressive myopathy, useful for investigations on FSHD pathophysiological mechanisms. However, the use of inducible transgenic models is often time consuming and costly. Therefore, cheap convenient in vivo models, with a local well delineated muscle lesion, although not representing the whole disease complexity, may constitute excellent tools in the frame of initial high-throughput therapeutic screening. Moreover, the IMEP model is customizable. For instance, a construct containing a weaker promoter100 could be used in order to mimic a more “physiological” DUX4 expression level. This type of model would be useful to better understand FSHD pathophysiological mechanisms. In addition, the contribution of modifier genes could be similarly investigated. In the present study, we have demonstrated that a single injection of pCIneo-DUX4 plasmid into mouse TA by hIMEP leads to the development of muscle damage after 7 days. These damages are directly linked to DUX4 expression, as evidenced by the absence of comparable lesions in muscles electroporated in the same conditions with control pDNA. Lesions in muscles injected with DUX4-expression plasmid are characterized by the presence of degenerating fibres with a heterogeneity in size, including numerous small fibres. Large mononuclear cell infiltration, extracellular matrix thickening, fibrosis and central nucleation are also observed as described in other DUX4 studies59,78–80. Conversely, control TAs display a normal histology without dystrophic pattern but some of them showed a thin and straight lesion characterized by fibres with central nuclei, sign of regeneration events most probably due to the needle injury caused by the intra-muscular injection. Since our results suggest an absence of significant systemic dissemination of pDNA following hIMEP, both hindlimbs could be used as independent systems, fitting therefore with the animal ethics principle of reduction. The local character of this myopathy model also presents two other advantages. First, only the injected muscle is impaired, so mice do not show any phenotype59,78,79,80. Second, the localized lesion observed 7 days after administration of DUX4 pDNA by hIMEP facilitates therapeutic evaluation. To this aim, we developed an easy and reproducible quantification method to evaluate the percentage of damaged muscle surface area. To ensure the distinction between healthy and injured area, we improved the basic hematoxylin–eosin histological coloration by adding a step of Heidenhain’s blue counterstaining that results in a high blue-pink contrast, easily distinguishable by automatic color thresholding. As expected, the percentage of damaged surface area is significantly higher in the injected area (β-gal+) than in the total TA section in DUX4 pDNA-injected mice. Interestingly, in the DUX4 group, we noticed a decrease of the β-gal+ surface area and a different spatial distribution of β-gal+ fibres which appear more scattered, as compared to the control group. This could be explained by the DUX4 toxicity. Indeed, since both plasmids are co-injected, it is reasonable to hypothesize that most fibres expressing the LacZ plasmid also express DUX4. Due to its high toxicity, the DUX4 protein likely causes fibre degeneration, sarcolemmal damages and a consecutive loss of β-gal signal.
We were not able to highlight a reduction of muscle damage by decreasing DUX4 plasmid doses. If the absence of dose–response is surprising, different hypotheses may be suggested. First, those results are in agreement with the high myotoxic potential of DUX4, even when initially expressed in a limited number of myonuclei, as described previously either in patients52,55,101,102 or in FSHD animal models78,79,80. As described by Tassin et al.55 and confirmed by Ferreboeuf et al.102, the model of “DUX4 nuclear spreading” and the subsequent amplification of the DUX4-induced cascade could partly explain this phenomenon. Second, direct DUX4-induced deregulation pathways are most probably followed by feedback loop processes generating a self-sustaining system involving e.g. oxidative stress and/or inflammation66,103. By this way, structural muscle damages observed in our study certainly result from pathophysiological processes that have become independent from the initial DUX4 boost.
Regarding spatial distribution of lesions within the TA muscle, we did not find any significant difference in lesion extent between medial and proximal sections, but a lower percentage of damaged surface area was observed in the distal TA part. This result was unexpected given that the injected transgene could in theory diffuse along the entire myofibres. Moreover, as DUX4 is a transcription factor and harbors nuclear localization signals104, it can be transported into several neighboring nuclei in a myotube, as described in previous studies55,102. Parameters such as electrode shape and positioning may be involved in inter-regional variation of hIMEP efficiency. Since TA is not cylinder- but cone-shaped, the distances between each muscle region and the electrodes are not identical along the whole muscle. As the distal TA part is located close to the tendon, the contact with electrode and the resulting electrical field was probably lower in this region, despite the use of a conductive gel. Nevertheless, pDNA may theoretically diffuse through the entire myofibre. However, since pDNA is administrated through a single injection, a dilution phenomenon could explain a lower expression in the regions remote from the injection site55. In addition, a limited diffusion of nuclear proteins in muscle fibres has already been mentioned105,106. The same observations were reported in porcine muscle following EP: in that study transgene expression was limited to the muscle area delineated by the electrical field but no mechanistic hypothesis was proposed107. The DUX4 hIMEP procedure is therefore associated to the rapid development of localized muscle lesions. Although this pattern allows to provide an easy read-out for therapeutic screening, the study of pathophysiological mechanisms underlying consequences of DUX4 muscle expression requires a more scattered distribution pattern of the transgene, closer to the one observed in FSHD muscles. This pattern may be obtained by using another route of delivery of naked pDNA. Indeed, the hydrodynamic injection of pDNA via the saphenous vein, first described by Hagstrom et al.108,109 allowed a more diffuse expression of the transgene in a panel of hindlimb muscles.
Regarding time-course investigations, no damages were observed 1 and 3 days post-hIMEP with DUX4 expression plasmid, while DUX4 mRNA and protein were already detected. Seven days after the hIMEP procedure, a severe muscle lesion had developed. Although the presence of early damages between the 3rd and the 7th day post-injection cannot be excluded, the period of 7 days seems appropriate to obtain quantifiable and well-delineated muscle lesions consecutive to DUX4 expression. It seems also reasonable to hypothesize that a delay is necessary for the establishment of pathophysiological processes which will ultimately overcome skeletal muscle compensatory mechanisms. Since the expression of DUX4 target genes (“DUX4 footprint genes”) is a good marker of DUX4 protein expression and transcriptional activity78,79,80, DUX4 target gene signature was investigated at the mRNA level to confirm the production of a functional DUX4 protein. As expected, at all-time points, Wfdc3 and Zscan4c expressions were induced in DUX4 pDNA-injected TAs as compared to controls. Myog, a marker of muscle regeneration, presented a decreased expression 1-day post-injection. This is similar to results published by other groups showing that Myog expression was decreased by DUX4 expression84,85,86. The progressive increase over time is consistent with histological regeneration features (central nuclei) observed 7-days after IMEP. DUX4 protein level was also investigated and as expected given our expression data on DUX4 mRNA and its target genes, DUX4 protein could be detected at all-time points.
Finally, given the relevance of our mouse model and quantification system in the framework of antisense therapy, a preclinical proof-of-concept study was performed with an antisense oligonucleotide (AO) directed against the DUX4 mRNA54,74. To this aim, we used pLAM3A (− 12 + 13) AO targeting a splice site in the DUX4 mRNA 3′UTR region. This AO was previously synthesized with a 2′OMe chemistry (phosphorothioate backbone) and successfully tested in vitro either on cells overexpressing DUX4 or on FSHD primary myotubes54. The same AO synthetized as a vPMO (octa-guanidine dendrimer-conjugated vivo morpholino) was used in a preliminary study in vivo showing its ability to decrease DUX4 mRNA expressed from an AAV vector injected in mouse TA74. In that study, the vPMO efficiency was monitored through DUX4 mRNA detection and semi-quantification using 3′RACE. In the present study, we first confirmed that the target sequence of this AO was conserved in the DUX4 mRNA detected in mouse TA electroporated with the DUX4-expression plasmid. We then tested this vPMO on the DUX4 hIMEP model and demonstrated it could prevent the development of DUX4-induced histological muscle damages.
In conclusion, we have developed a rapid and easy-to-use mouse model of DUX4 local over-expression. Even if the mouse in which TAs are injected with pCIneo-DUX4 by hIMEP does not recapitulate all the complexity of FSHD pathophysiology, this model has an added value in the forefront of pre-clinical evaluations, particularly in a context in which high throughput therapeutic screening is still necessary. This model could also be applied for the injection of other expression vectors to model various gain-of-function muscular diseases in order to test potential therapeutics.
Methods
Ethics statement
All animal experiments met the Belgian national standard requirements regarding animal care and were conducted in accordance with the Ethics and Welfare Committee of the University of Mons. Protocols were approved by the Ethics and Welfare Committee of the University of Mons (reference number LE016/03).
Animals
Female C57BL/6 mice, aged between 8 and 12 weeks, were purchase from Charles River laboratories (France). Mice were housed in a conventional animal colony and maintained at 35–40% relative humidity with a constant room temperature (21 °C) and natural day/night light cycle (12–12 h). Food and water were provided ad libitum and animals were subjected to an adaptation period of 7 days before starting experiments.
Plasmids
The commercial reporter plasmid pCMV-lacZ encoding the Escherichia coli β-galactosidase protein was used to assess gene electrotransfer efficiency. The pCIneo-DUX4 plasmid encoding the full length DUX4 protein is detailed in110. This plasmid contains the complete DUX4 ORF followed by the pLAM region isolated from the 4q35 fragment of a patient with FSHD. The pCIneo backbone vector from Promega (USA) was used as negative control. Expression plasmids used in this study are driven by cytomegalovirus (CMV) promoter. All DNA plasmids were produced by PlasmidFactory (Bielefeld, Germany) in a research grade quality for pre-clinical and veterinary studies.
Intra-muscular injection and electroporation (hIMEP)
Adult female C57BL/6 mice were anesthetized by inhalation of 4% isoflurane in an induction chamber then maintained with 2% isoflurane using a mouse anesthesia mask. Hind limbs were shaved and 40 µg of hyaluronidase from bovine testes (Sigma-Aldrich, USA) diluted in 20 µl sterile saline buffer (0.9% NaCl, Physiodose, Gilbert Laboratories, France) were injected through the skin into the tibialis anterior (TA) muscle. Mice were then allowed to recover from anesthesia in their cages. After two hours, mice were re-anesthetized by an intra-peritoneal injection of Ketamine 100 mg/kg (Anestketin, Eurovet animal health) and Xylasine 10 mg/kg (Sigma-Aldrich, USA). Each TA was injected, with a Hamilton syringe, through the skin with one dose of naked pDNA (1, 5, 10, 20 or 40 µg) diluted in an equal volume of sterile saline buffer (50 µl). Flat parallel electrodes were placed on each side of the TA and good contact between electrodes and the overlying skin was ensured by use of a conductive gel (Rodisonic, Pannoc, Belgium). A train of eight square-wave pulses at a voltage of 95 V (200 V/cm; voltage to distance ratio calculated on the hind limb thickness mean) and a duration of 20 ms at 500 ms interval (2 Hz) was generated using an EMKA stimulator. After electroporation, animals were transferred back into their cages and were monitored until complete recovery from anesthesia. Mice were daily checked and then sacrificed by an intraperitoneal injection of Nembutal (120 mg/kg, CEVO) 1-, 3- or 7-days post electroporation. For vPMO preliminary test, both mouse TAs were electroporated with 20 µg of pCIneo-DUX4 plasmid. Six hours after the procedure, mice received either an intra-peritoneal injection of 250 µg of vPMO pLAM3A − 12 + 13 (Gene tools, described in54,72) or no supplemental treatment. TA were harvested 7 days after hIMEP.
Tissue preparation and histology
At the indicated euthanasia time points, right and left TAs were removed, embedded in OCT compound (VWR) and frozen in liquid nitrogen-cooled isopentane. Eight µm thick cryostat sections from proximal, medial and distal part of TA were cut using a Leica cryotome and serial sections were colored respectively with X-gal staining (β-gal staining kit, Invitrogen) to assess percentage and localization of muscle area expressing the electrotransfered genes, and with Hematoxylin–Eosin–Heidenhain blue (HEB) to evaluate the percentage of damaged muscle area. HEB coloration consists in a basic Hematoxylin–Eosin coloration followed by a 45-s incubation in Heidenhain’s Blue staining (mix of orange G and Aniline Blue, Sigma-Aldrich, USA), allowing an intense blue labeling of fibrotic fibresand collagenous tissues which improves contrast with healthy myofibres to facilitate muscle lesion quantifications. Slides were then scanned using the NanoZoomer-SQ Digital slide scanner (Hamamatsu Photonics). Images were processed by color thresholding using ImageJ 1.52a software, Rasband, W.S., ImageJ, U. S. National Institutes of Health, Bethesda, MA, USA, https://imagej.nih.gov/ij/, 1997–2018. The blue surface area was measured (threshold parameters: Hue = 35–255/50–255, saturation = 53–255/53–255 for X-gal/HEB staining respectively) and reported to the complete surface section. Measurements were performed on total section and on the injected area defined by X-gal staining (SI Fig. S3 online).
Immunofluorescence
Tissue cryosections were fixed with 4% paraformaldehyde/PBS on ice for 20 min, permeabilized with 0.25% TritonX-100/PBS for 10 min, then incubated with blocking solution (5% normal goat serum (Dako), 2% BSA, 0.01% TritonX-100/PBS) for 30 min. Sections were then incubated with anti-DUX4 (rabbit monoclonal E5-5; 1:200, Abcam) and anti-laminin α (rat monoclonal; 1:100; Sigma-Aldrich) primary antibodies at 4 °C overnight. They were subsequently incubated with secondary antibodies Alexa 555 Goat anti-rabbit IgG (1:500, Biowest) and Alexa 488 Goat anti-rat Ig (1:500, Invitrogen) at room temperature for 1 h. Stained sections were mounted with EverBrite Mounting Medium with DAPI (Biotium) for nuclear staining. Pictures were taken with a Nikon Eclipse 80i microscope and merged using NIS-Elements software.
RNA isolation, 3′RACE and real time quantitative PCR
RNA extraction
TA muscles were trimmed between proximal-medial and medial-distal parts using Leica cryotome (30 sections of 50 µm in both regions). These slices were ground into liquid nitrogen, homogenized in 1 ml of TRIzol reagent (Invitrogen) and RNA was isolated according to the manufacturer’s directions. Total RNA was then treated with DNAse I kit (amplification grade, ThermoFisher).
3′RACE
cDNAs were synthetized using SuperScript III Reverse transcriptase kit (Invitrogen) and the 3′RACE adaptor of the RLM-RACE kit (5′-GCGAGCACAGAATTAATACGACTCACTATAGGTTTTTTTTTTTTN-3′, Ambion). The resulting cDNAs were amplified by nested PCR using PrimeSTAR Max DNA polymerase (Takara-bioJapan). The specific outer primer for DUX4 cDNA amplification was: 5′-AGGCGCAACCTCTCCTAG AAAC-3′ and the inner primer was: 5′-TGGAAGCACCCCTCAGCGAGGAA-3′. Cycling conditions for outer PCR were as follows: initial denaturation step at 98 °C for 3 min, followed by amplification step with 25 cycles of 10 s at 98 °C, 10 s at 58 °C and 5 s at 72 °C, ended by 5 min at 72 °C. Same cycling conditions were used for inner PCR except the last step of the amplification which was 10 s at 72 °C instead of 5 s. The 3′RACE PCR products were analyzed by electrophoresis on 1%-agarose gel and staining with ethidium bromide (gel pictures provided Fig. 6 were made in compliance with the digital image and integrity policies of Scientific Reports). The products were then extracted from agarose band using PCR clean up gel extraction kit (Macherey–Nagel), cloned into pJET1.2/blunt (ThermoFisher), amplified in E. coli and sequenced (Genewiz Inc.) to confirm specific DUX4 mRNA amplification and vPMO targeted sequence conservation.
RT-qPCR
cDNAs were synthetized using Maxima First Strand cDNA Synthesis kit (ThermoFisher). All qPCRs were performed in duplicates using SYBR Green FastStart Essential DNA Green Master (Roche) and following primers (10 µM): Wfdc3 (5′-CTTCCATGTCAGGAGCTGTG-3′, 5′-ACCAGGATTCTGGGACATTG-3′) and Zscan4c (5′-GATTATTGGCCACAGGACAAG-3′, 5′-TCAGGGTGCTGTTCTTTCTG-3′) from76, Myog (5′-GAGACATCCCCCTATTTCTACCA-3′, 5′-GCTCAGTCCGCTCATAGCC-3′) and Rplp0 (5′-TCATCCAGCAG GTGTTCG-3′, 5′-AGCAAGTGGGAAGGTGTAA-3′) custom designed (Eurogentec). Cycling conditions were as follows: initial denaturation step at 95 °C for 10 min, followed by 40 cycles of 15 s at 95 °C and 60 s at 60 °C. qPCR results were analysis with LightCycler 96 software (Roche).
Statistical analyses
Statistical analyses were done using GraphPad Prism version 6.01 for Windows, GraphPad Prism software, La Jolla, CA, USA, www.graphpad.com. Data were tested for normality of distribution, using the Kolmogorov–Smirnov test, and all considered as non-parametric. Differences between experimental groups were statistically evaluated by a Mann Whitney rank sum test (hyaluronidase pre-injection, DUX4 expression and vPMO evaluation), one-way analysis of variance (ANOVA) on the ranks followed by a Dunn’s post hoc test for multiple comparison (dose–response, time–response and qPCR analysis), Friedman repeated ANOVA on the ranks (muscle distribution) or Wilcox on signed rank test (quantification on total vs injected area). Differences were considered statistically significant at a p value < 0.05. All data are represented as boxplot for non-parametric statistical tests (with minimum to maximum whiskers).
References
Sokołowska, E. & Błachnio-Zabielska, A. U. A critical review of electroporation as a plasmid delivery system in mouse skeletal muscle. Int. J. Mol. Sci. 20, 2776. https://doi.org/10.3390/ijms20112776 (2019).
Cervia, L. D. & Yuan, F. Current progress in electrotransfection as a nonviral method for gene delivery. Mol. Pharm. 15, 3617–3624 (2018).
Young, J. L. & Dean, D. A. Electroporation-mediated gene delivery. Adv. Genet. 89, 49–88 (2015).
Rosazza, C., Meglic, S. H., Zumbusch, A., Rols, M.-P. & Miklavcic, D. Gene electrotransfer: A mechanistic perspective. Curr. Gene Ther. 16, 98–129 (2016).
Neumann, E., Schaefer-Ridder, M., Wang, Y. & Hofschneider, P. H. Gene transfer into mouse lyoma cells by electroporation in high electric fields. EMBO J. 1, 841–845 (1982).
Pucihar, G., Kotnik, T., Miklavčič, D. & Teissié, J. Kinetics of transmembrane transport of small molecules into electropermeabilized cells. Biophys. J. 95, 2837–2848 (2008).
Shi, J. et al. A review on electroporation-based intracellular delivery. Molecules 23, 3044. https://doi.org/10.3390/molecules23113044 (2018).
Jakutavičiūtė, M., Ruzgys, P., Tamošiūnas, M., Maciulevičius, M. & Šatkauskas, S. Physical methods for drug and gene delivery through the cell plasma membrane. Adv. Anat. Embryol. Cell Biol. 227, 73–92 (2017).
Antov, Y., Barbul, A., Mantsur, H. & Korenstein, R. Electroendocytosis: Exposure of cells to pulsed low electric fields enhances adsorption and uptake of macromolecules. Biophys. J. 88, 2206–2223 (2005).
Rosazza, C. et al. Endocytosis and endosomal trafficking of DNA after gene electrotransfer in vitro. Mol. Ther. Nucleic Acids 5, e286 (2016).
Markelc, B. et al. Inhibitor of endocytosis impairs gene electrotransfer to mouse muscle in vivo. Bioelectrochemistry 103, 111–119 (2015).
Probst, U., Fuhrmann, I., Beyer, L. & Wiggermann, P. Electrochemotherapy as a new modality in interventional oncology: A review. Technol. Cancer Res. Treat. 17, 1–12. https://doi.org/10.1177/1533033818785329 (2018).
Marty, M. et al. Electrochemotherapy—An easy, highly effective and safe treatment of cutaneous and subcutaneous metastases: Results of ESOPE (European Standard Operating Procedures of Electrochemotherapy) study. Eur. J. Cancer Suppl. 4, 3–13 (2006).
Sersa, G. & Miklavcic, D. Electrochemotherapy of tumours. J. Vis. Exp. https://doi.org/10.3791/1038 (2008).
Heller, R. & Heller, L. C. Gene electrotransfer clinical trials. Adv. Genet. 89, 235–262 (2015).
Gothelf, A. & Gehl, J. What you always needed to know about electroporation based DNA vaccines. Hum. Vaccin. Immunother. 8, 1694–1702 (2012).
Grødeland, G. & Bogen, B. Efficient vaccine against pandemic influenza: Combining DNA vaccination and targeted delivery to MHC class II molecules. Expert Rev. Vaccines 14, 805–814 (2015).
Zhong, Z. et al. Immunogenicity and protection efficacy of a naked self-replicating mRNA-based Zika virus vaccine. Vaccines (Basel) 7, 96. https://doi.org/10.3390/vaccines7030096 (2019).
Williams, M. et al. Enhanced immunogenicity and protective efficacy of a tetravalent dengue DNA vaccine using electroporation and intradermal delivery. Vaccine 37, 4444–4453 (2019).
Albrecht, M. T. et al. Electroporation of a multivalent DNA vaccine cocktail elicits a protective immune response against anthrax and plague. Vaccine 30, 4872–4883 (2012).
Daud, A. I. et al. Phase I trial of interleukin-12 plasmid electroporation in patients with metastatic melanoma. J. Clin. Oncol. 26, 5896–5903 (2008).
Pasquet, L. et al. Pre-clinical investigation of the synergy effect of interleukin-12 gene-electro-transfer during partially irreversible electropermeabilization against melanoma. J. Immunother. Cancer 7, 161 (2019).
Isaka, Y. et al. DNAzyme for TGF-beta suppressed extracellular matrix accumulation in experimental glomerulonephritis. Kidney Int. 66, 586–590 (2004).
Hojman, P., Gissel, H. & Gehl, J. Sensitive and precise regulation of haemoglobin after gene transfer of erythropoietin to muscle tissue using electroporation. Gene Ther. 14, 950–959 (2007).
Gothelf, A., Hojman, P. & Gehl, J. Therapeutic levels of erythropoietin (EPO) achieved after gene electrotransfer to skin in mice. Gene Ther. 17, 1077–1084 (2010).
Ayuni, E. L. et al. In vivo electroporation mediate****d gene delivery to the beating heart. PLoS One 5, e14467 (2010).
Cox, D. B. T., Platt, R. J. & Zhang, F. Therapeutic genome editing: Prospects and challenges. Nat. Med. 21, 121–131 (2015).
Latella, M. C. et al. In vivo editing of the human mutant rhodopsin gene by electroporation of plasmid-based CRISPR/Cas9 in the mouse retina. Mol. Ther. Nucleic Acids 5, e389 (2016).
Son, M.-Y. et al. Generation and characterization of integration-free induced pluripotent stem cells from patients with autoimmune disease. Exp. Mol. Med. 48, e232 (2016).
Okita, K. et al. A more efficient method to generate integration-free human iPS cells. Nat. Methods 8, 409–412 (2011).
Thomas, C. E., Ehrhardt, A. & Kay, M. A. Progress and problems with the use of viral vectors for gene therapy. Nat. Rev. Genet. 4, 346–358 (2003).
Maruyama, H. et al. Long-term production of erythropoietin after electroporation-mediated transfer of plasmid DNA into the muscles of normal and uremic rats. Gene Ther. 8, 461–468 (2001).
Long, Y. C. et al. FVIII gene delivery by muscle electroporation corrects murine hemophilia A. J. Gene Med. 7, 494–505 (2005).
Hanna, E. et al. Intramuscular electroporation delivery of IL-12 gene for treatment of squamous cell carcinoma located at distant site. Cancer Gene Ther. 8, 151–157 (2001).
Aihara, H. & Miyazaki, J. Gene transfer into muscle by electroporation in vivo. Nat. Biotechnol. 16, 867–870 (1998).
Akerstrom, T. et al. Optimizing hyaluronidase dose and plasmid DNA delivery greatly improves gene electrotransfer efficiency in rat skeletal muscle. Biochem. Biophys. Rep. 4, 342–350 (2015).
Hartikka, J. et al. Electroporation-facilitated delivery of plasmid DNA in skeletal muscle: Plasmid dependence of muscle damage and effect of poloxamer 188. Mol. Ther. 4, 407–415 (2001).
Taylor, J. et al. Optimization of ectopic gene expression in skeletal muscle through DNA transfer by electroporation. BMC Biotechnol. 4, 11 (2004).
McMahon, J. M., Signori, E., Wells, K. E., Fazio, V. M. & Wells, D. J. Optimisation of electrotransfer of plasmid into skeletal muscle by pretreatment with hyaluronidase—increased expression with reduced muscle damage. Gene Ther. 8, 1264–1270 (2001).
Molnar, M. J. et al. Factors influencing the efficacy, longevity, and safety of electroporation-assisted plasmid-based gene transfer into mouse muscles. Mol. Ther. 10, 447–455 (2004).
Vilquin, J. T. et al. Electrotransfer of naked DNA in the skeletal muscles of animal models of muscular dystrophies. Gene Ther. 8, 1097–1107 (2001).
Pichavant, C., Chapdelaine, P., Cerri, D. G., Bizario, J. C. S. & Tremblay, J. P. Electrotransfer of the full-length dog dystrophin into mouse and dystrophic dog muscles. Hum. Gene Ther. 21, 1591–1601 (2010).
Statland, J. M. & Tawil, R. Facioscapulohumeral muscular dystrophy. Continuum (Minneap Minn) 22, 1916–1931 (2016).
Wijmenga, C. et al. Location of facioscapulohumeral muscular dystrophy gene on chromosome 4. Lancet 336, 651–653 (1990).
Wijmenga, C. et al. Mapping of facioscapulohumeral muscular dystrophy gene to chromosome 4q35-qter by multipoint linkage analysis and in situ hybridization. Genomics 9, 570–575 (1991).
van Deutekom, J. C. et al. FSHD associated DNA rearrangements are due to deletions of integral copies of a 3.2 kb tandemly repeated unit. Hum. Mol. Genet. 2, 2037–2042 (1993).
Das, S. & Chadwick, B. P. Influence of repressive histone and DNA methylation upon D4Z4 transcription in non-myogenic cells. PLoS One 11, e0160022 (2016).
Lemmers, R. J. L. F. et al. Facioscapulohumeral muscular dystrophy is uniquely associated with one of the two variants of the 4q subtelomere. Nat. Genet. 32, 235–236 (2002).
Lemmers, R. J. L. F. et al. Specific sequence variations within the 4q35 region are associated with facioscapulohumeral muscular dystrophy. Am. J. Hum. Genet. 81, 884–894 (2007).
Lemmers, R. J. L. F. et al. A unifying genetic model for facioscapulohumeral muscular dystrophy. Science 329, 1650–1653 (2010).
Lemmers, R. J. L. F. et al. Digenic inheritance of an SMCHD1 mutation and an FSHD-permissive D4Z4 allele causes facioscapulohumeral muscular dystrophy type 2. Nat. Genet. 44, 1370–1374 (2012).
Snider, L. et al. Facioscapulohumeral dystrophy: incomplete suppression of a retrotransposed gene. PLoS Genet. 6, e1001181 (2010).
Geng, L. N. et al. DUX4 activates germline genes, retroelements and immune-mediators: Implications for facioscapulohumeral dystrophy. Dev. Cell 22, 38–51 (2012).
Vanderplanck, C. et al. The FSHD atrophic myotube phenotype is caused by DUX4 expression. PLoS One 6, e26820 (2011).
Tassin, A. et al. DUX4 expression in FSHD muscle cells: How could such a rare protein cause a myopathy?. J. Cell. Mol. Med. 17, 76–89 (2013).
Feng, Q. et al. A feedback loop between nonsense-mediated decay and the retrogene DUX4 in facioscapulohumeral muscular dystrophy. eLife 4, e04996 (2015).
Shadle, S. C. et al. DUX4-induced dsRNA and MYC mRNA stabilization activate apoptotic pathways in human cell models of facioscapulohumeral dystrophy. PLoS Genet. 13, e1006658 (2017).
Kowaljow, V. et al. The DUX4 gene at the FSHD1A locus encodes a pro-apoptotic protein. Neuromuscul. Disord. 17, 611–623 (2007).
Wallace, L. M. et al. DUX4, a candidate gene for facioscapulohumeral muscular dystrophy, causes p53-dependent myopathy in vivo. Ann. Neurol. 69, 540–552 (2011).
Rickard, A. M., Petek, L. M. & Miller, D. G. Endogenous DUX4 expression in FSHD myotubes is sufficient to cause cell death and disrupts RNA splicing and cell migration pathways. Hum. Mol. Genet. 24, 5901–5914 (2015).
Tawil, R. et al. A pilot trial of prednisone in facioscapulohumeral muscular dystrophy. FSH-DY Group. Neurology 48, 46–49 (1997).
Kissel, J. T. et al. Randomized, double-blind, placebo-controlled trial of albuterol in facioscapulohumeral dystrophy. Neurology 57, 1434–1440 (2001).
van der Kooi, E. L. et al. Strength training and albuterol in facioscapulohumeral muscular dystrophy. Neurology 63, 702–708 (2004).
Payan, C. A. et al. Periodic salbutamol in facioscapulohumeral muscular dystrophy: A randomized controlled trial. Arch. Phys. Med. Rehabil. 90, 1094–1101 (2009).
van der Kooi, E. L. et al. Effects of training and albuterol on pain and fatigue in facioscapulohumeral muscular dystrophy. J. Neurol. 254, 931–940 (2007).
Passerieux, E. et al. Effects of vitamin C, vitamin E, zinc gluconate, and selenomethionine supplementation on muscle function and oxidative stress biomarkers in patients with facioscapulohumeral dystrophy: A double-blind randomized controlled clinical trial. Free Radic. Biol. Med. 81, 158–169 (2015).
Oliva, J. et al. Clinically advanced p38 inhibitors suppress DUX4 expression in cellular and animal models of facioscapulohumeral muscular dystrophy. J. Pharmacol. Exp. Ther. 370, 219–230 (2019).
Hamel, J. & Tawil, R. Facioscapulohumeral muscular dystrophy: Update on pathogenesis and future treatments. Neurotherapeutics 15, 863–871 (2018).
Bao, B., Maruyama, R. & Yokota, T. Targeting mRNA for the treatment of facioscapulohumeral muscular dystrophy. Intractable Rare Dis. Res. 5, 168–176 (2016).
Balog, J. et al. Increased DUX4 expression during muscle differentiation correlates with decreased SMCHD1 protein levels at D4Z4. Epigenetics 10, 1133–1142 (2015).
Campbell, A. E. et al. NuRD and CAF-1-mediated silencing of the D4Z4 array is modulated by DUX4-induced MBD3L proteins. Elife https://doi.org/10.7554/eLife.31023 (2018).
Ansseau, E. et al. Antisense oligonucleotides used to target the DUX4 mRNA as therapeutic approaches in faciosscapulohumeral muscular dystrophy (FSHD). Genes (Basel) 8, 93. https://doi.org/10.3390/genes8030093 (2017).
Chen, J. C. et al. Morpholino-mediated knockdown of DUX4 toward facioscapulohumeral muscular dystrophy therapeutics. Mol. Ther. 24, 1405–1411 (2016).
Wallace, L. M. et al. Pre-clinical safety and off-target studies to support translation of AAV-mediated RNAi therapy for FSHD. Mol. Ther. Methods Clin. Dev. 8, 121–130 (2018).
Wallace, L. M. et al. RNA interference inhibits DUX4-induced muscle toxicity in vivo: Implications for a targeted FSHD therapy. Mol. Ther. 20, 1417–1423 (2012).
Krom, Y. D. et al. Intrinsic epigenetic regulation of the D4Z4 macrosatellite repeat in a transgenic mouse model for FSHD. PLoS Genet. 9, e1003415 (2013).
Dandapat, A. et al. Dominant lethal pathologies in male mice engineered to contain an X-linked DUX4 transgene. Cell. Rep. 8, 1484–1496 (2014).
Bosnakovski, D. et al. Muscle pathology from stochastic low level DUX4 expression in an FSHD mouse model. Nat. Commun. 8, 550 (2017).
Jones, T. & Jones, P. L. A cre-inducible DUX4 transgenic mouse model for investigating facioscapulohumeral muscular dystrophy. PLoS One 13, e0192657 (2018).
Giesige, C. R. et al. AAV-mediated follistatin gene therapy improves functional outcomes in the TIC-DUX4 mouse model of FSHD. JCI Insight 3, 22. https://doi.org/10.1172/jci.insight.123538 (2018).
Mueller, A. L. et al. Muscle xenografts reproduce key molecular features of facioscapulohumeral muscular dystrophy. Exp. Neurol. 320, 113011 (2019).
Zhang, Y. et al. Human skeletal muscle xenograft as a new preclinical model for muscle disorders. Hum. Mol. Genet. 23, 3180–3188 (2014).
Jones, T. I. et al. Transgenic mice expressing tunable levels of DUX4 develop characteristic facioscapulohumeral muscular dystrophy-like pathophysiology ranging in severity. Skelet. Muscle 10, 8 (2020).
Bosnakovski, D. et al. An isogenetic myoblast expression screen identifies DUX4-mediated FSHD-associated molecular pathologies. EMBO J. 27, 2766–2779 (2008).
Sharma, V., Harafuji, N., Belayew, A. & Chen, Y.-W. DUX4 differentially regulates transcriptomes of human rhabdomyosarcoma and mouse C2C12 cells. PLoS One 8, e64691 (2013).
Bosnakovski, D. et al. Low level DUX4 expression disrupts myogenesis through deregulation of myogenic gene expression. Sci. Rep. 8, 16957 (2018).
Ansseau, E. et al. Aberrant splicing in transgenes containing introns, exons, and V5 epitopes: Lessons from developing an FSHD mouse model expressing a D4Z4 repeat with flanking genomic sequences. PLoS One 10, e0118813 (2015).
Levy, M. Y., Barron, L. G., Meyer, K. B. & Szoka, F. C. Characterization of plasmid DNA transfer into mouse skeletal muscle: Evaluation of uptake mechanism, expression and secretion of gene products into blood. Gene Ther. 3, 201–211 (1996).
Mir, L. M. et al. High-efficiency gene transfer into skeletal muscle mediated by electric pulses. Proc. Natl. Acad. Sci. USA 96, 4262–4267 (1999).
Schertzer, J. D., Plant, D. R. & Lynch, G. S. Optimizing plasmid-based gene transfer for investigating skeletal muscle structure and function. Mol. Ther. 13, 795–803 (2006).
Bettan, M. et al. High-level protein secretion into blood circulation after electric pulse-mediated gene transfer into skeletal muscle. Mol. Ther. 2, 204–210 (2000).
Gehl, J. et al. In vivo electroporation of skeletal muscle: Threshold, efficacy and relation to electric field distribution. Biochim. Biophys. Acta 1428, 233–240 (1999).
Gehl, J. & Mir, L. M. Determination of optimal parameters for in vivo gene transfer by electroporation, using a rapid in vivo test for cell permeabilization. Biochem. Biophys. Res. Commun. 261, 377–380 (1999).
Favre, D. et al. Hyaluronidase enhances recombinant adeno-associated virus (rAAV)-mediated gene transfer in the rat skeletal muscle. Gene Ther. 7, 1417–1420 (2000).
Cemazar, M. et al. Hyaluronidase and collagenase increase the transfection efficiency of gene electrotransfer in various murine tumors. Hum. Gene Ther. 23, 128–137 (2012).
Mennuni, C. et al. Hyaluronidase increases electrogene transfer efficiency in skeletal muscle. Hum. Gene Ther. 13, 355–365 (2002).
Eikenes, L., Tari, M., Tufto, I., Bruland, O. S. & de LangeDavies, C. Hyaluronidase induces a transcapillary pressure gradient and improves the distribution and uptake of liposomal doxorubicin (Caelyx) in human osteosarcoma xenografts. Br. J. Cancer 93, 81–88 (2005).
Guo, M. et al. Precise temporal regulation of Dux is important for embryo development. Cell Res. 29, 956–959 (2019).
Percharde, M. et al. A LINE1-nucleolin partnership regulates early development and ESC identity. Cell 174, 391-405.e19 (2018).
Ferreira, J. P., Peacock, R. W. S., Lawhorn, I. E. B. & Wang, C. L. Modulating ectopic gene expression levels by using retroviral vectors equipped with synthetic promoters. Syst. Synth. Biol. 5, 131–138 (2011).
Block, G. J. et al. Wnt/β-catenin signaling suppresses DUX4 expression and prevents apoptosis of FSHD muscle cells. Hum. Mol. Genet. 22, 4661–4672 (2013).
Ferreboeuf, M. et al. DUX4 and DUX4 downstream target genes are expressed in fetal FSHD muscles. Hum. Mol. Genet. 23, 171–181 (2014).
Turki, A. et al. Functional muscle impairment in facioscapulohumeral muscular dystrophy is correlated with oxidative stress and mitochondrial dysfunction. Free Radic. Biol. Med. 53, 1068–1079 (2012).
Corona, E. D., Jacquelin, D., Gatica, L. & Rosa, A. L. Multiple protein domains contribute to nuclear import and cell toxicity of DUX4, a candidate pathogenic protein for facioscapulohumeral muscular dystrophy. PLoS One 8, e75614 (2013).
Ono, T. et al. Limited diffusibility of gene products directed by a single nucleus in the cytoplasm of multinucleated myofibres. FEBS Lett. 337, 18–22 (1994).
Newlands, S. et al. Transcription occurs in pulses in muscle fibers. Genes Dev. 12, 2748–2758 (1998).
Spanggaard, I. et al. Spatial distribution of transgenic protein after gene electrotransfer to porcine muscle. Hum. Gene Ther. Methods 23, 387–392 (2012).
Wooddell, C. I. et al. Dose response in rodents and nonhuman primates after hydrodynamic limb vein delivery of naked plasmid DNA. Hum. Gene Ther. 22, 889–903 (2011).
Hagstrom, J. E. et al. A facile nonviral method for delivering genes and siRNAs to skeletal muscle of mammalian limbs. Mol. Ther. 10, 386–398 (2004).
Gabriëls, J. et al. Nucleotide sequence of the partially deleted D4Z4 locus in a patient with FSHD identifies a putative gene within each 3.3 kb element. Gene 236, 25–32 (1999).
Acknowledgements
This study was funded by the AFM-Téléthon (Association française contre les myopathies), Grant #18916 “DUX4SILIMICE”. A.D. and TH.N. held Ph.D. fellowships from the Belgian F.R.S-FNRS (Fonds de la Recherche Scientifique). We acknowledge Dr. J. Dumonceaux (University College London) for advice on the IMEP procedure. We also acknowledge B. Blairon, and C. Ruelle (UMONS) for technical assistance.
Author information
Authors and Affiliations
Contributions
A.L., A.T., A.B., A.D. and A.E.D. conceived the study, participated in its design and coordination; A.D., T.H.N., A.T., V.J. and E.D. carried out the animal experimentations; A.D., T.H.N., E.A., A.T., A.E.D., V.J., and F.C. participated in the molecular studies; A.D., A.T., A.E.D., A.B. and A.L. drafted the manuscript; A.D. and A.L. performed statistical analysis. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest. However, regarding the experiment involving an antisense oligonucleotide (vPMO pLAM3A −12 + 13), we have to mention that FC, EA and AB are co-inventors on patent US2012225034A1.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Derenne, A., Tassin, A., Nguyen, T.H. et al. Induction of a local muscular dystrophy using electroporation in vivo: an easy tool for screening therapeutics. Sci Rep 10, 11301 (2020). https://doi.org/10.1038/s41598-020-68135-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-68135-7
This article is cited by
-
Transplantation of PSC-derived myogenic progenitors counteracts disease phenotypes in FSHD mice
npj Regenerative Medicine (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.