Abstract
Huge quantities of keratinaceous waste are a substantial and almost totally unexploited protein resource which could be upgraded for use as high value-added products by efficient keratinolytic enzymes. In this study, we found that Bacillus sp. 8A6 can efficiently degrade chicken feather after 24 h growth. According to phylogenetic analysis, the strain (formerly identified as Bacillus pumilus 8A6) belongs to the B. pumilus species clade but it is more closely related to B. safensis. Hotpep predicted 233 putative proteases from Bacillus sp. 8A6 genome. Proteomic analysis of culture broths from Bacillus sp. 8A6 cultured on chicken feathers or on a mixture of bristles and hooves showed high abundance of proteins with functions related to peptidase activity. Five proteases (one from family M12, one from family S01A, two from family S08A and one from family T3) and four oligopeptide and dipeptide binding proteins were highly expressed when Bacillus sp. 8A6 was grown in keratin media compared to LB medium. This study is the first to report that bacterial proteases in families M12, S01A and T3 are involved in keratin degradation together with proteases from family S08.
Similar content being viewed by others
Introduction
A very large amount of animal-derived keratinaceous waste such as feathers, skin, hair, bristles, horns, hooves, claws, nails, beaks, reptilian osteoderm, and fish teeth and slime are generated annually1. Keratin is the third most abundant polymer in nature after cellulose and chitin2. Keratinaceous materials are fibrous proteins composed of recalcitrant polymers with a high degree of cross-linking disulfide bonds, hydrogen bonds, and hydrophobic interactions. The keratin is self-assembled from two polypeptides that form an intermediate filament, sterically mediated by a distinct head and tail structure1. Additionally, post-translational modifications, such as the formation of disulfide bonds, phosphorylation and glycosylation leads to various keratin structures and different degrees of recalcitrance and bio-availabilities3. The recalcitrant structure of keratin is the first line of defense against microbial attacks on animal body parts.
Molyneux4 was the first to isolate a Bacillus sp. that was able to degrade keratin. Lin et al.5 were later the first to purify and characterize keratinase KerA of the S8 protease family from a B. licheniformis strain. Subsequently, several different kinds of bacteria with keratinolytic capability have been isolated and described. Single keratinolytic proteases have been purified from culture broth or recombinantly expressed from single strains. However, keratin cannot be degraded by only one keratinase. The complex and recalcitrant structure needs synergistic interaction of different types of keratinolytic enzymes in order to be effectively decomposed6.
Bacillus species such as B. licheniformis, B. subtilis and B. cereus are well-known for their keratinolytic capability7,8,9. In this study, Bacillus sp. 8A6 (deposited at the Bacillus Genetic Stock Center as “B. pumilus 8A6”; B. pumilus has GRAS status) was found to be an efficient keratin degrader. Based on whole-genome comparisons, the strain studied was found to be more closely related to B. safensis. Until now, however, only a few reports have indicated that B. safensis has keratin degradation capability and no keratin-degrading enzymes have been identified from this species10.
In this study, keratinases from Bacillus sp. 8A6 were identified by bioinformatic analysis using Hotpep11 and by proteome analysis of culture broths of Bacillus sp. 8A6 grown on keratin (chicken feather (CF) or on a mixture of bristles and hooves (BH)). We found five proteases and four oligopeptide and dipeptide binding proteins that were highly up-regulated, i.e. more abundant, when Bacillus sp. 8A6 was grown on keratinaceous substrates. These enzymes are likely to be involved in keratin degradation.
Results
Strain selection
The Bacillus pumilus strain FH9 has exceptional proteolytic enzymes capable of degrading feather waste12 and its hydrolytic capacities makes it suitable for use in production of bioplastic13. Several other strains of B. pumilus are available at the Bacillus Genetic Stock Centre. We chose to use the strain 8A6, which was isolated from a sample taken above the flood line of a stream in a 7 miles long cave in carbonate rock in Kentucky, USA14.
Genome sequencing and de novo assembly
The assembly obtained with k-mer value 59 was selected as the best representation of the genome scaffold based on assessment of the constructed deBrujin graphs (highest number of vertices with an ingoing degree of 1 and an outgoing degree of 1) and assembly statistics (highest N50 value). Because of the high quality of the reads and coverage of around 400 ×, neither additional trimming nor error correction were necessary and did not improve the assembly when tested. The k-mer value of 59 resulted in 10 contigs with a length >=100 nt; the total length, average length and largest contig length were 3,729,788 bp, 372,987 bp, and 975,857 bp, respectively. The N50 was 962,040 bp and 3900 genes were predicted in selected assembly by GeneMarkS. Genome draft completeness was calculated by both BUSCO and CheckM to be 99.6% where 524 out of 526 complete markers and 710 out of 711 complete markers were identified by BUSCO and CheckM, respectively. Duplicated markers were not found, which indicated lack of contamination.
Classification of Bacillus sp. 8A6
Bacillus sp. 8A6 was first published named only to the level of genus, and then was subsequently deposited in the Bacillus Genetic Stock Center as Bacillus pumilus 8A6, based on the 16 S rRNA gene sequence14. In this study, the identified 16 S rRNA gene sequence of this strain was compared with 16 S rRNA sequences identified in 202 assemblies of Bacillus type strain genomes from the NCBI Assembly database. The most similar sequences identified by BLASTn were subsequently used for phylogenetic analysis (Fig. 1). The 16 S rRNA sequence of Bacillus sp. 8A6 formed a distinct clade together with 16 S rRNA sequences from B. safensis Fo-36b, B. pumilus species and B. zhangzhouensis DW5-4. However, species classification of strains related to B. pumilus based on the 16 S rRNA gene is not reliable because the majority of them share over 99.5% of 16 S rRNA gene identity15. Therefore whole-genome in silico comparison using average nucleotide identity (ANI) and Genome-to-Genome distance calculator (GGDC) was applied for more accurate taxonomic placement (Table 1). We found that Bacillus sp. 8A6 is more closely related to the B. safensis Fo-36b strain than to B. pumilus based on significantly higher ANI (98.79%) and digital DNA-DNA hybridization (dDDH) values (89.3), which is higher than the proposed thresholds for species delineation, of 95–96% and 70%, respectively.
Phylogenetic tree (maximum-likelihood) of the full length 16 S rRNA gene sequences identified in Bacillus sp. 8A6 strain (red) and the closest known relatives from type strains. The percentage of replicate trees (>60%) in which the associated taxa clustered together in the bootstrap test (500 replicates) is shown next to the branches.
Proteases prediction by Hotpep analysis in Bacillus sp. 8A6 and B. safensis Fo-36b genomes
Proteases were predicted in both Bacillus sp. 8A6 and B. safensis Fo-36b genomes by Hotpep analysis. All the predicted sequences were supported by CDD search and blasting in the Merops database. Finally, 233 and 235 proteases in Aspartic (A), Cysteine (C), Metallo (M), Asparagine (N), Serine (S), Threonine (T) and Unknown (U) protease/peptidase families were found in Bacillus sp. 8A6 and B. safensis Fo-36b genomes, respectively (Fig. 2). The protease profiles of both strains were similar, especially for serine proteases and metalloproteases. Potential keratinolytic proteases of Bacillus sp. 8A6, B. safensis Fo-36b, B. pumilus SH-B9, B. altitudinis 41KF2b, B. stratosphericus LAMA585 and B. safensis KCTC12796BP were found in 10 protease families (A1, A8, A22, A24A, A25, A28, A31, A36, M03B and S08A) according to functional prediction of keratinolytic proteases from fungi and bacteria (Supplementary Table S1). 16 potential keratinolytic protease sequences were found in the Bacillus sp. 8A6 genome.
Proteomic analysis of keratinolytic protease genes in Bacillus sp. 8A6
Efficient degradation of keratin by Bacillus sp. 8A6
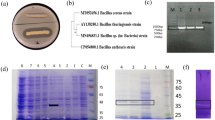
Bacillus sp. 8A6 showed high capability for keratin degradation when grown in liquid culture including keratinaceous substrate. This strain efficiently degraded CF in 24 h at 37 °C when using CF as sole carbon and nitrogen source. Keratinase activity when using CF or BH as substrate was 4,463 and 4,043 U/ml, respectively, while keratinase activity was only 790 U/ml when grown in LB medium. Moreover, protease activity when Bacillus sp. 8A6 was grown in medium with CF or with BH (33,650 and 30,500 U/ml, respectively) was higher than when grown in LB medium (3,917 U/ml). After keratin degradation by Bacillus sp. 8A6, up to 3.79 and 9.55 mg/ml soluble protein were released into the medium from CF and BH, respectively.
MS analysis of protein composition of secretome of Bacillus sp. 8A6
In total, 194, 68, and 33 proteins were identified in culture broth from Bacillus sp. 8A6 grown in LB, BH and CF medium, respectively (Fig. 3). According to the GO ontology comparison of proteins detected in secretome samples (Supplementary Fig. S1), proteins with functions related to peptidase activity, catalytic activity acting on a protein and hydrolase activity in molecular functions showed significantly higher abundance when Bacillus sp. 8A6 was grown in the keratin media than in LB medium (Fig. 4 and Supplementary Fig. S2).
Of the proteins with annotated function, four proteins (derived from gene_2558: GH16_lichenase, gene_2946: xylanase, gene_3650: the substrate-binding component of the oligopeptide-binding protein, and gene_9: the substrate-binding component of an ABC-type dipeptide import system containing the type 2 periplasmic binding fold) were found only in culture broth with BH and CF media. According to the t-test results for culture broth from keratin media against the results for culture broth of LB medium, six proteases (one from family M12, two from family S08A, one from family S01A, one from family T3 and one from family M17) showed statistically significant up-regulation according to the –LOG (P-value) when Bacillus sp. 8A6 was grown on CF (Table 2, Supplementary Table S2). Furthermore, seven proteases (one from family M12, one from family S01A, three from family S08A, one from family M20B and one in family T3) showed statistically significant up-regulation when Bacillus sp. 8A6 was grown on BH medium (Table 3, Supplementary Table S3). Thus, when Bacillus sp. 8A6 was grown on keratinaceous substrates, five proteases (derived from gene_1796 in family M12, gene_3018 in family S01A, gene_3289 and gene_3746 in family S08A, and gene_3552 in family T3) were shown to be highly up-regulated.
In this study we also found that the substrate binding component for oligopeptide and dipeptide, such as gene_3645, was highly up-regulated in the culture broth with CF and BH (Table 4). Furthermore, three other substrate binding components (gene_3650, gene_9 and gene_992) could also be found only in the keratin medium but not in the LB medium (Table 4).
Discussion
Keratinaceous waste streams such as feathers, pig bristles and hooves have high amounts of intermediate filament-forming proteins with specific physicochemical properties. Recently, upgrade not just of lignocellulose and chitin but also of keratinaceous waste has been studied and discussed in relation to the new bioeconomy2,16. Several types of bacteria and fungi have been found to process sets of enzymes needed for keratinolytic capability2. Specialized fungi such as saprotrophs growing on e.g. feather and hooves (as e.g. Onygena corvina)17 or dermatophytic species, have been shown to have a rich diversity of keratinolytic enzymes18. However, the fungal enzymatic keratinolytic process appears to be rather slow, taking a longer time (e.g.> 10 days) than is easily accommodated in an industrial setting of waste conversion17. Interestingly, and in contrast, some bacteria have optimum keratinase activity production at around 30–72 h. These species include B. licheniformis PWD-19, B. licheniformis RG119, Fervidobacterium pennavorans20, Fervidobacterium islandicum AW-121 and Meiothermus taiwanensis WR-223022 which can completely degrade keratin in a short time (24–48 h), excepting B. licheniformis PWD-1 which needs 10 days. However, these species are slower than the Bacillus sp. 8A6 studied here regarding keratin degradation. In this study, we found that Bacillus sp. 8A6 efficiently degraded CF and that the keratinase activity of the culture broth was at its highest within only 24 h of incubation. This strain was found also to produce the highest keratinase activity after only 18 h growth in a 5 L fermenter when using BH as substrate (data not shown). These findings are a motivation for further characterization of the highly efficient keratinolytic proteases from Bacillus sp. 8A6.
In this study we investigated the taxonomy of Bacillus sp. 8A6 by supplementing the 16 S rRNA phylogenetic approach with whole-genome in silico comparisons. The designation of the species within the “B. pumilus” group based solely on ribosomal gene markers is not reliable because most of the group members share high 16 S rRNA gene identity. For example, Espariz et al.15 found that as many as 50% of strains in this group were misclassified. This highlights the necessity of utilizing more detailed phylogenomic and whole-genome based approaches for more specific classification of organisms in the B. pumilus complex. Based on comparison of the Bacillus sp. 8A6 genome reported in this study with available genomes of type strains belonging to the “B. pumilus species group”, we propose that strain B. pumilus 8A6 be renamed as B. safensis 8A6. B. safensis was first identified in 2006 as a contaminant in the spacecraft assembly facility at the Jet Propulsion Laboratory, USA, from which it derived its specific epithet ‘safensis’23. Although B. safensis does not have GRAS status yet, it has been regarded as a safe industrial microorganism with promising biotechnological applications due to its ability to produce various industrial enzymes and secondary metabolites24. Until now, only B. safensis LAU 13 has been reported to have keratinolytic activity10,25. However, specific keratinolytic proteases have not been identified nor characterized from B. safensis.
The Hotpep predicted protease profile of Bacillus sp. 8A6 was similar to B. safensis FO-36b. We also attempted to investigate whether or not Bacillus sp. 8A6 and other Bacillus species have lytic polysaccharide monooxygenases (LPMOs). Notably, LPMO AA11 is widely found in keratin degrading fungi26. Further, AA11 was described from the keratin degrading fungus, O. corvina. Based on detailed study of the keratinolytic capability of O. corvina, it was hypothesized that LPMO AA11 is important for the keratinolytic effect of this fungus2. In contrast, no LPMO protein could be found by HotPep analysis in Bacillus sp. 8A6. Similarly, LPMOs were also not found in the genome of B. safensis Fo-36b, B. pumilus SH-B9, B. aerophilus C772, B. altitudinis 41KF2b, B. stratosphericus LAMA585 or B. safensis KCTC12796BP. The apparent lack of LPMO genes in the genome of keratinolytic bacteria suggests that bacteria may have a different mechanism(s) for keratin degradation than fungi. Notably, the predicted potential keratinolytic proteases belonging to protease families A1, A8, A22, A24A, A25, A28, A31, A36, M03B and S08A in the above bacterial species are also different from those of fungi such as O. corvina, which mainly has proteases with keratinolytic activity from families M03, M28 and S86.
The first keratinase described from B. licheniformis strain were serine protease in family S85. Recently, a broader diversity of keratinolytic bacteria has been found, but most of the identified bacterial keratinolytic proteases were actually in family S827. All the commercial keratinases were classified in the S8 family2. However, not all the proteases in family S8 have high keratinolytic capability. In this study, the two proteases from family S8 showed high up-regulation when Bacillus sp. 8A6 was grown on CF or BH. Some other proteases in family S8 did not show any significant difference when the secretome of Bacillus sp. 8A6 grown in keratin medium and in LB medium was compared.
Only few proteases of M12 family have been found from bacteria since 199528. The M12 metalloproteases from the deep-sea bacterium Myroides profundi D25 was found to have elastinolytic activity and a synergistic role in collagen hydrolysis28. This specific swollen collagen structure property was further confirmed by an optimized scale up process29. The most highly up-regulated M12 metalloprotease of Bacillus sp. 8A6 when grown in keratin medium might indicate that this metalloprotease M12 can modify the structure of the keratin, which as a result then becomes accessible for other keratinolytic proteases.
In this study, a non-keratin-specific trypsin-like protease from family S01A was also found to be abundant in the studied keratinolytic culture broths. In 2001, a new trypsin-like protease was first isolated from a B. licheniformis strain when grown in fermentation broth with milled chicken feather30. However, the function of this trypsin-like protease during keratin degradation has never been clarified. Streptomycetes exfoliatus and S. albidoflavus were found to produce trypsin when grown under limiting carbon, nitrogen and phosphate growth conditions31. The production of trypsin by S. exfoliatus and S. albidoflavus was also positively related to aerial Streptomycete-mycelium formation and growth32. Therefore, the up-regulated trypsin-like protease in keratin-rich medium might be induced by insoluble CF and BH and may further degrade oligopeptides produced by specific keratinolytic proteases. It is also found that multifunctional aminopeptidase A in M17 family has been highly up-regulated when Bacillus sp. 8A6 grown on chicken feather, which might participate for further degrading oligopeptides as the function of the aminopeptidase from O. corvina6.
Feathers, bristles and hooves are cysteine-rich keratin-associated recalcitrant protein structures33. After the feathers, bristles and hooves have been degraded by the keratinolytic proteases, the resulting oligopeptides will also be rich in cysteine, which opens the way for production of e.g. glutathione. The secreted gamma-glutamyl transpeptidase in family T3 from Bacillus sp. 8A6 can hydrolyze glutathione to glutaminate and cysteinyl-glycine, which can be further hydrolyzed to cysteine and glycine. The resulting glutaminate, cysteine and glycine can be taken up by the microbial cell directly. It has been reported that gamma-glutamyl transpeptidase can balance the levels of intracellular cysteine30,34. Grumbt et al.35 have proposed that during keratin degradation by dermatophytes, intracellular cysteine can be metabolized to sulfite via the action of the enzyme cysteine dioxygenase Cdo1. Then the released sulfite further facilitates keratin degradation. Therefore, the up-regulated gamma-glutamyl transpeptidase in the keratin medium not only provides the cell with cysteine but also with, the further metabolized products that may also support the keratin degradation process itself. Disulfide reductases are also well-known for participating the keratin degradation36; however, we did not find disulfide reductases in the secretome when Bacillus sp. 8A6 was grown on CF or BH.
Besides the keratinolytic proteases, the mass spectrometric analyses also identified four oligopeptide and dipeptide substrate binding proteins. These substrate binding proteins were the membrane-associated components of an ATP-binding cassette (ABC) transport system OppABCDEF consisting of five subunits. The substrate-binding components determined the substrate specificity of the transport system. It has been found that this ABC transport system has an important role in supplying bacteria with essential amino acids in a low energy-requiring process37. This system appears to be very important for the keratinolytic microorganism’s uptake of oligopeptides or amino acid from the medium after the keratin degradation18.
On the basis of the comprehensive genome and secretome analysis, we therefore propose the synthetic effects of different enzymes for keratin degradation by Bacillus sp. 8A6: The structure of hard keratin is first weakened by metabolized product sulfite and M12 metalloprotease, and subsequently the loosened structure of keratin is further hydrolyzed by S08A serine proteases. The resulting long peptides are further degraded by a S01A serine protease and the oligopeptides and dipeptides are transported into the cell by an ABC transport system. The secreted family T3 gamma-glutamyl transpeptidase provides the cell with cysteine while additional metabolized products such as sulfite might also support the keratin degradation process. Detailed characterization of each enzymes is part of ongoing research.
In conclusion, this study showed that Bacillus sp. 8A6 strain could efficiently degrade CF and BH, paving the way for further improvements in utilization of keratin-rich materials. Phylogenetic analysis revealed that formerly identified as B. pumilus, strain 8A6 is more properly classified as B. safensis. Hotpep analysis predicted 233 proteases and 16 potential keratinolytic proteases in the Bacillus sp. 8A6 genome. Proteomic analysis of Bacillus sp. 8A6 secretome showed a significant induction of proteins related to peptidase activity by keratin. Five proteases (one from family M12, one from family S01A, two from family S8A and one from family T3) and four oligopeptide and dipeptide binding components were highly expressed both in CF and in BH medium compared to LB medium. This study is the first to suggest the involvement of proteases of families M12, S01A, T3 and S8 in keratin degradation.
Methods
Microorganism and growth conditions
Bacillus sp. 8A6 (formerly identified as Bacillus pumilus 8A6) (wild type isolate, original code GGC-D3, BGSCID 8A6) was obtained from the Bacillus Genetic Stock Center. This strain was grown on LB agar medium. For keratinolytic enzymes production, a single clone of Bacillus sp. 8A6 was inoculated in 100 ml LB medium and incubated at 37 °C for 20 h (OD600: 1.7–2). Harvest of cells was performed by centrifugation at 6000 g for 5 min and the cell pellet was resuspended in 50 ml PBS buffer (pH 7.5). Two ml of the resuspended cells was inoculated into 50 ml mineral keratin medium (3% BH/1% CF, 0.75 g/l NaCl, 1.75 g/l K2HPO4, 0.25 g/l MgSO4·7H2O, 0.055 g/l CaCl2, 0.010 g/l FeSO4·7H2O, 0.005 g/l ZnSO4·7 H2O, 10 mM MOPS, pH 8) or into 50 ml LB medium as negative control. The culture and control were incubated at 37 °C on a rotary shaker (180 rpm) for 24 h.
The pig bristles and hooves mixture (BH), which have been chopped, steam treated (150 °C, 6 bar, 20 min), dried and crushed into smaller particles, were supplied by DAKA, Saria group, Løsning, Denmark. Chicken feathers (CF) were obtained from Rose Poultry (Vinderup, Skovsgaard, Denmark). The concentration of BH and CF in the medium was determined according to Huang et al., manuscript in preparation.
Enzyme analysis and protein determination
Protease activity was analyzed as described6 by using azocasein as substrate (Megazyme) with a small modification. First, 20 μl 1.5% w/v azocasein suspension was mixed with 50 mM sodium carbonate buffer pH9 and 20 μl diluted enzyme in 1.5 ml tubes. The reactions were then conducted at 60 °C for 15 min with constant agitation at 500 rpm (Huang et al., manuscript in preparation). One arbitrary unit (U) of protease activity was defined as the amount of enzyme causing a 0.01 absorbance increase between the sample and control at 405 nm under the assay conditions.
Keratinolytic protease activity was analyzed using azokeratin (kindly provided by Professor Søren Sørensen’s group, Section of Microbiology, University of Copenhagen). 500 µl diluted enzyme was added to 800 µl of 0.01 g/ml azokeratin in 50 mM sodium carbonate buffer pH9. The reaction was carried out at 60 °C and 1000 rpm for 1 h (Huang et al., manuscript in preparation). Then the mixture was centrifuged at 16,000 g for 1 min and 150 μl supernatant was transferred to a microtiter plate. Absorbance was read at 415 nm using a plate reader. A control was prepared using 500 µl 50 mM sodium carbonate buffer pH9 to replace the addition of 500 µl diluted enzyme. One arbitrary unit (U) of keratinolytic protease activity was defined as the amount of enzyme causing a 0.001 absorbance increase between the sample and control at 415 nm under the assay conditions.
Culture broth protein concentration was determined by PIERCE BCA Protein Assay Kit (23225, Thermo Scientific) using BSA as standards.
Genomic DNA extraction and sequencing
Bacillus sp. 8A6 was grown in LB medium for 20 h at 37 °C and 180 rpm and then 1 ml of culture was centrifuged at 5000 g for 10 min. The genomic DNA was extracted using DNeasy Blood & Tissue Kit (Qiagen, Cat No./ID: 69506) according to a protocol for gram-positive bacteria. The quantity and quality of the genomic DNA was checked on a NanoDrop 1000 spectrophotometer (THERMO ECIENTIFIC) and by electrophoresis on 1% agarose gel. The sample was submitted to Macrogen (Korea) for Truseq PCR-free (350 bp) shotgun library preparation and sequencing using an Illumina HiSeq. 2500 instrument (150 bp paired-end sequencing).
De novo genome assembly and open-reading frame prediction
The last bases (101) were removed from all reads. The read-through adapters on the 3’ ends were trimmed by using k-mer based detection implemented in BBDuk (settings: hdist = 1, k = 21, mink = 11, tbo, tpe)38. Reads were quality-trimmed using a Phred algorithm (to a value higher than 20). Assembly was performed using Ray (de Bruijin graph-based assembly) with varying k-mer length (35, 39, 43, 49, 55, 59, 65, 75, 95)39. The constructed library had an average outer distance between reads (insert length of the library) of 385 ± 83. Open reading frames were predicted with GeneMarkS40 with combined GeneMarkS generated (native) and Heuristic model parameters integrated into one model. The genome draft completeness was assessed by BUSCO (v3), with lineage-specific ortholog set for Bacillales (odb9), and CheckM v1.0.9 with lineage-specific marker set for Bacillus genus (UID864). Both approaches use collocated sets of genes that are ubiquitous and single-copy within a phylogenetic lineage41,42.
Protease prediction by Hotpep analysis
The protease sequences were first predicted by Hotpep analysis using the same approach as described for CAZymes11. The predicted protease sequences were then further supported by batch CDD searching in NCBI and blasting in the MEROPS database 12.0 (file “pepunit.lib”)43 (2018, March) using CLC Main Workbench version 7 with default parameters.
Phylogenetic analysis of 16 S rRNA
The phylogenetic analysis was performed as follows. Ribosomal 16 S RNA gene sequences were predicted using barrnap 0.744. One full length 16 S rRNA sequence was identified. The reference database was prepared from 16 S RNA gene sequences identified in 202 assemblies of Bacillus type strains’ genomes from NCBI Assembly database. The identified 16 S gene was submitted to a BLAST analysis against the reference database and 50 hits (>1,000 bp, E value <1e-20) with the highest identities (>95%) were extracted. Reference sequences were de-replicated and together with query sequence aligned to the RF00177 model from the Rfam database using SSU- ALIGN45. SSU-mask was applied for automated probabilistic masking. A maximum-likelihood (ML) phylogenetic tree was constructed using MEGA 7 with the general time-reversible model with correction for among site rate variation (Gamma distributed with Invariant sites, G + I). A bootstrap replication of 500 was used for testing the phylogeny.
Genome to genome comparisons
The genomes of five highly related type strains of Bacillus species (Table 1) were obtained from the NCBI Assembly database (March 2018). These downloaded genomes and the draft genome of Bacillus sp. 8A6 were compared with the GGDC webservice at http://ggdc.dsmz.de (BLAST + as an alignment tool and sum of all identities found in HSPs divided by overall HSP length as a distance measure)46 and the ANI Calculator webservice at https://www.ezbiocloud.net/tools/ani47.
Precipitation of proteins in culture broth supernatant of Bacillus sp. 8A6 for MS analysis
Culture broth of Bacillus sp. 8A6 grown in medium with either CF, BH or LB medium, was harvested by centrifugation at 10000 g for 15 min at 4 °C. The supernatant was filtered (0.2 µm). The extracellular proteins were precipitated as described6. Prior to LC-MS/MS measurements, proteins were in-solution digested using trypsin as described before6. After tryptic digestion, peptides were purified using C18 packed StageTips48 and dried by vacuum centrifugation.
Analysis of proteins by LC-MS/MS
Peptides were reconstituted in 0.1% trifluoroacetic acid/2% acetonitrile solution. Eight microliters of each sample were injected by autosampler and concentrated as well as washed on a trapping column (Pepmap100, C18, 100 μm × 2 cm, 5 μm, THERMO FISHER SCIENTIFIC) with water containing 0.1% formic acid at 800 bar. Afterwards, the peptides were eluted from a separation column (PepmapRSLC, C18, 75 μm×50 cm, 2 μm, THERMO FISHER SCIENTIFIC). Chromatography was performed with 0.1% formic acid in solvent A (100% water) and B (80% acetonitrile, 20% water). Three successive linear gradients with solvent B were run one after another. First, from 5% to 12% within 5 min, next from 12 to 37% over 50 min, and then from 37 to 50% within 5 min, followed by a final one-minute-step gradient to 100% solvent B, which was maintained for 20 min using a nano-high-pressure liquid chromatography system (Easy-nLC 1200, THERMO FISHER SCIENTIFIC). Ionized peptides were measured and fragmented by a Q Exactive mass spectrometer (THERMO FISHER SCIENTIFIC). For an unbiased analysis, continuous scanning of eluted peptide ions was carried out between 400 and 12,000 m/z, automatically switching to MS/MS higher energy collisional dissociation (HCD) mode and 12 MS/MS events per survey scan. For MS/MS HCD measurements, a dynamic precursor exclusion of 30 s, peptide match, and an apex trigger of 2 to 20 s were enabled.
MS data analysis
Protein identification was done with the open-source software MaxQuant (v. 1.5.3.30)49. The label-free quantification (LFQ) algorithm50, IBAQ, and the match between runs feature were activated. Carbamidomethylation of cysteines was defined as fixed modification, and oxidation of methionines as well as N-terminal acetylation was defined as variable modification. The remaining settings were kept at default. This included a maximum peptide and protein false discovery rate of 1% and a minimum of two peptides for LFQ calculation. The predicted proteins of Bacillus sp. 8A6 database after the GeneMarkS prediction as described above was used as a search database in MaxQuant. The mean LFQ per protein was calculated if a protein was quantified in at least two out of three biological replicates. For comparison of relative changes, the LFQ ratios between conditions were formed and log2 transformed. Statistical significances of abundance changes were assessed by t-test (two-tailed, heteroscedastic). Batch CD search51 was used to search for conserved domains and annotation of identified protein. A Venn diagram was created by Venny 2.152. Sequences of proteins identified in secretomes were subjected to annotation with the InterProScan pipeline standalone suite (5.27–66.0) using all possible analyses and databases53. GO annotations for each dataset were subsequently compared with WEGO54.
Data availability
The genome sequence of Bacillus sp. 8A6 has been submitted to the Genbank database under accession number QFZE00000000. The Hotpep predicted protease nucleotide sequences in Bacillus sp. 8A6 genome and the up-regulated protein nucleotide sequences in the secretome when Bacillus sp. 8A6 was grown in keratin media compared to LB medium have been submitted to the Genbank database under accession numbers MH475952-MH476188 (Tables 2, 3, 4 and Supplementary Table S4)
References
McKittrick, J. et al. The structure, functions, and mechanical properties of keratin. JOM. 64, 449–468 (2012).
Lange, L., Huang, Y. & Busk, P. K. Microbial decomposition of keratin in nature-a new hypothesis of industrial relevance. Appl. Microbiol. Biot. 100, 2083–2096 (2016).
Yamada, S., Wirtz, D. & Coulombe, P. A. Pairwise assembly determines the intrinsic potential for self-organization and mechanical properties of keratin filaments. Molecular Biology of the Cell. 13, 382–391 (2002).
Molyneux, G. The digestion of wool by a keratinolytic Bacillus. Aust. J. Biol. Sci. 12, 274–281 (1959).
Lin, X., Lee, C. G., Casale, E. S. & Shih, J. C. Purification and characterization of a keratinase from a feather-degrading Bacillus licheniformis strain. Appl. Environ. Microb. 58, 3271–3275 (1992).
Huang, Y., Busk, P. K., Herbst, F.-A. & Lange, L. Genome and secretome analyses provide insights into keratin decomposition by novel proteases from the non-pathogenic fungus Onygena corvina. Appl. Microbiol. Biot. 99, 9635–9649 (2015).
Kim, J. M., Lim, W. J. & Suh, H. J. Feather-degrading Bacillus species from poultry waste. Process Biochem. 37, 287–291 (2001).
Macedo, A. J. et al. Novel keratinase from Bacillus subtilis S14 exhibiting remarkable dehairing capabilities. Appl. Environ. Microb. 71, 594–596 (2005).
Williams, C. M., Richter, C. S., Mackenzie, J. M. & Shih, J. C. Isolation, identification, and characterization of a feather-degrading bacterium. Appl. Environ. Microb. 56, 1509–1515 (1990).
Lateef, A., Adelere, I. A. & Gueguim-Kana, E. B. Bacillus safensis LAU 13: a new source of keratinase and its multi-functional biocatalytic applications. Biotechnol. Biotec. Eq. 29, 54–63 (2015).
Busk, P. K., Pilgaard, B., Lezyk, M. J., Meyer, A. S. & Lange, L. Homology to peptide pattern for annotation of carbohydrate-active enzymes and prediction of function. BMC Bioinformatics. 18, 214 (2017).
El-Refai, H. A., AbdelNaby, M. A., Gaballa, A., El-Araby, M. H. & Abdel Fattah, A. F. Improvement of the newly isolated Bacillus pumilus FH9 keratinolytic activity. Process Biochem. 40, 2325–2332 (2005).
Ibrahim, M. H. A., Takwa, M. & Hatti-kaul, R. Process for extraction of bioplastic and production of monomers from the bioplastic. United States patent 20170253713, (2017).
Banks, E. D. et al. Bacterial calcium carbonate precipitation in cave environments: A function of calcium homeostasis. Geomicrobiol. J. 27, 444–454 (2010).
Espariz, M., Zuljan, F. A., Esteban, L. & Magni, C. Taxonomic identity resolution of highly phylogenetically related strains and selection of phylogenetic markers by using genome-scale methods: The Bacillus pumilus group case. PLoS One. 11, e0163098–e0163098 (2016).
Gousterova, A. et al. Degradation of keratin and collagen containing wastes by newly isolated thermoactinomycetes or by alkaline hydrolysis. Lett. Appl. Microbiol. 40, 335–340 (2005).
Huang, Y., Busk, P. & Lange, L. Production and characterization of keratinolytic proteases produced by Onygena corvina. Fungal Genom. Biol. 5, 1–7 (2015).
Mercer, D. K. & Stewart, C. S. Keratin hydrolysis by dermatophytes. Med. Mycol. 57, 13–22 (2019).
Priya, R., Rajni, S. & Rani, G. Keratinolytic potential of Bacillus licheniformis RG1: structural and biochemical mechanism of feather degradation. Can. J. Microbiol. 51, 191–196 (2005).
Friedrich, A. B. & Antranikian, G. Keratin degradation by Fervidobacterium pennavorans, a novel thermophilic anaerobic species of the order Thermotogales. Appl. Environ. Microb. 62, 2875–2882 (1996).
Nam, G. W. et al. Native-feather degradation by Fervidobacterium islandicum AW-1, a newly isolated keratinase-producing thermophilic anaerobe. Arch. Microbiol. 178, 538–547 (2002).
Wu, W. L. et al. The discovery of novel heat-stable keratinases from Meiothermus taiwanensis WR-220 and other extremophiles. Sci. Rep. 7, 4658–4658 (2017).
Satomi, M., La Duc, M. T. & Venkateswaran, K. Bacillus safensis sp. nov., isolated from spacecraft and assembly-facility surfaces. Int. J. Syst. Evol. Micr. 56, 1735–1740 (2006).
Lateef, A., Adelere, I. A. & Gueguim-Kana, E. B. The biology and potential biotechnological applications of Bacillus safensis. Biologia. 70, 411–419 (2015).
Lateef, A., Adelere, I. A., Gueguim-Kana, E. B., Asafa, T. B. & Beukes, L. S. Green synthesis of silver nanoparticles using keratinase obtained from a strain of Bacillus safensis LAU 13. Int. Nano Lett. 5, 29–35 (2015).
Busk, P. K. & Lange, L. Classification of fungal and bacterial lytic polysaccharide monooxygenases. BMC Genomics. 16, 368 (2015).
Brandelli, A. Bacterial keratinases: Useful enzymes for bioprocessing agroindustrial wastes and beyond. Food Bioprocess Tech. 1, 105–116 (2008).
Chen, X. et al. Ecological function of myroilysin, a novel bacterial M12 metalloprotease with elastinolytic activity and a synergistic role in collagen hydrolysis, in biodegradation of deep-sea high-molecular-weight organic nitrogen. Appl. Environ. Microb. 75, 1838–1844 (2009).
Shao, X. et al. Culture condition optimization and pilot scale production of the M12 metalloprotease myroilysin produced by the deep-sea bacterium Myroides profundi D25. Molecules. 20, 11891–11901 (2015).
Rozs, M., Manczinger, L., Vágvölgyi, C. & Kevei, F. Secretion of a trypsin-like thiol protease by a new keratinolytic strain of Bacillus licheniformis. FEMS Microbiol. Lett. 205, 221–224 (2001).
Kang, S. G., Kim, I. S., Rho, Y. T. & Lee, K. J. Production dynamics of extracellular proteases accompanying morphological differentiation of Streptomyces albidoflavus SMF301. Microbiology. 141, 3095–3103 (1995).
Kim, I. S. & Lee, K. J. Regulation of production of leupeptin, leupeptin-inactivating enzyme, and trypsin-like protease in Streptomyces exfoliatus SMF13. J. Ferment. Bioeng. 80, 434–439 (1995).
Strasser, B., Mlitz, V., Hermann, M., Tschachler, E. & Eckhart, L. Convergent evolution of cysteine-rich proteins in feathers and hair. BMC Evol. Biol. 15, 82–82 (2015).
Mehdi, K. & Penninckx, M. J. An important role for glutathione and γ-glutamyltranspeptidase in the supply of growth requirements during nitrogen starvation of the yeast Saccharomyces cerevisiae. Microbiology. 143, 1885–1889 (1997).
Grumbt, M. et al. Keratin degradation by dermatophytes relies on cysteine dioxygenase and a sulfite efflux pump. J. Invest. Dermatol. 133, 1550–1555 (2013).
Yamamura, S., Morita, Y., Hasan, Q., Yokoyama, K. & Tamiya, E. Keratin degradation: a cooperative action of two enzymes from Stenotrophomonas sp. Biochem. Bioph. Res. Co. 294, 1138–1143 (2002).
Garault, P., Le Bars, D., Besset, C. & Monnet, V. Three oligopeptide-binding proteins are involved in the oligopeptide transport of Streptococcus thermophilus. J. Biol. Chem. 277, 32–39 (2002).
Bushnell, B. BBMap short read aligner, http://sourceforge.net/projects/bbmap/ (2016).
Boisvert, S., Raymond, F., Godzaridis, É., Laviolette, F. & Corbeil, J. Ray Meta: scalable de novo metagenome assembly and profiling. Genome Biol. 13, R122 (2012).
Besemer, J., Lomsadze, A. & Borodovsky, M. GeneMarkS: a self-training method for prediction of gene starts in microbial genomes. Implications for finding sequence motifs in regulatory regions. Nucleic Acids Res. 29, 2607–2618 (2001).
Parks, D. H., Imelfort, M., Skennerton, C. T., Hugenholtz, P. & Tyson, G. W. CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res. 25, 1043–1055 (2015).
Waterhouse, R. M. et al. BUSCO applications from quality assessments to gene prediction and phylogenomics. Mol. Biol. Evol. 35, 543–548 (2017).
Rawlings, N. D. et al. The MEROPS database of proteolytic enzymes, their substrates and inhibitors in 2017 and a comparison with peptidases in the PANTHER database. Nucleic Acids Res. 46, D624–D632 (2018).
Seemann, T. Victorian bioinformatics consortium, http://www.vicbioinformatics.com/software.barrnap.shtml (2014).
Nawrocki, E. Structural RNA homology search and alignment using covariance models PhD thesis, Washington University in Saint Louis (2009).
Meier-Kolthoff, J. P., Auch, A. F., Klenk, H.-P. & Göker, M. Genome sequence-based species delimitation with confidence intervals and improved distance functions. BMC Bioinformatics. 14, 60 (2013).
Yoon, S. H., Ha, S. M., Lim, J., Kwon, S. & Chun, J. A large-scale evaluation of algorithms to calculate average nucleotide identity. Antonie van Leeuwenhoek. 110, 1281–1286 (2017).
Rappsilber, J., Mann, M. & Ishihama, Y. Protocol for micro-purification, enrichment, pre-fractionation and storage of peptides for proteomics using StageTips. Nat. Protoc. 2, 1896 (2007).
Tyanova, S., Temu, T. & Cox, J. The MaxQuant computational platform for mass spectrometry-based shotgun proteomics. Nat. Protoc. 11, 2301 (2016).
Cox, J. et al. Accurate proteome-wide label-free quantification by delayed normalization and maximal peptide ratio extraction, termed MaxLFQ. Molecular & Cellular Proteomics. 13, 2513–2526 (2014).
Marchler-Bauer, A. et al. CDD: a Conserved Domain Database for the functional annotation of proteins. Nucleic Acids Res. 39, D225–D229 (2011).
Oliveros, J. VENNY. An interactive tool for comparing lists with Venn Diagrams, http://bioinfogp.cnb.csic.es/tools/venny/index.html (2007).
Jones, P. et al. InterProScan 5: genome-scale protein function classification. Bioinformatics. 30, 1236–1240 (2014).
Ye, J. et al. WEGO: a web tool for plotting GO annotations. Nucleic Acids Res. 34, W293–W297 (2006).
Acknowledgements
This study was funded by The Danish Council for Strategic Research, now the Danish Innovation Fund (grant number 1308-00015B, Keratin2Protein). We thank Bo Pilgaard for protease prediction with Hotpep, Chicken Rose Poultry for providing chicken feathers, Daka and Danish Crown for providing pig bristles, milled bristles and hooves.
Author information
Authors and Affiliations
Contributions
Y.H. and L.L. designed the experiment. Y.H. conducted the experiments and analyzed the results. M.L. analyzed the genome sequence. F.H. conducted the LC-MS/MS experiment. P.B. provided Hotpep result. Y.H. wrote the paper, and L.L., Y.H., M.L., F.H. and P.B. revised the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Huang, Y., Łężyk, M., Herbst, FA. et al. Novel keratinolytic enzymes, discovered from a talented and efficient bacterial keratin degrader. Sci Rep 10, 10033 (2020). https://doi.org/10.1038/s41598-020-66792-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-66792-2
This article is cited by
-
Comprehensive insights into the mechanism of keratin degradation and exploitation of keratinase to enhance the bioaccessibility of soybean protein
Biotechnology for Biofuels and Bioproducts (2023)
-
Characterization of high-yield Bacillus subtilis cysteine protease for diverse industrial applications
Brazilian Journal of Microbiology (2023)
-
Multifarious revolutionary aspects of microbial keratinases: an efficient green technology for future generation with prospective applications
Environmental Science and Pollution Research (2022)
-
Feather-Degrading Bacillus cereus HD1: Genomic Analysis and Its Optimization for Keratinase Production and Feather Degradation
Current Microbiology (2022)
-
Optimization of Electroporation Conditions for Bacillus pumilus 3–19 Strain
BioNanoScience (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.