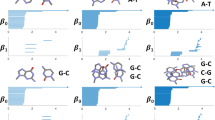
Abstract
It has long been observed that trimethylamine N-oxide (TMAO) and urea demonstrate dramatically different properties in a protein folding process. Even with the enormous theoretical and experimental research work on these two osmolytes, various aspects of their underlying mechanisms still remain largely elusive. In this paper, we propose to use the weighted persistent homology to systematically study the osmolytes molecular aggregation and their hydrogen-bonding network from a local topological perspective. We consider two weighted models, i.e., localized persistent homology (LPH) and interactive persistent homology (IPH). Boltzmann persistent entropy (BPE) is proposed to quantitatively characterize the topological features from LPH and IPH, together with persistent Betti number (PBN). More specifically, from the localized persistent homology models, we have found that TMAO and urea have very different local topology. TMAO is found to exhibit a local network structure. With the concentration increase, the circle elements in these networks show a clear increase in their total numbers and a decrease in their relative sizes. In contrast, urea shows two types of local topological patterns, i.e., local clusters around 6 Å and a few global circle elements at around 12 Å. From the interactive persistent homology models, it has been found that our persistent radial distribution function (PRDF) from the global-scale IPH has same physical properties as the traditional radial distribution function. Moreover, PRDFs from the local-scale IPH can also be generated and used to characterize the local interaction information. Other than the clear difference of the first peak value of PRDFs at filtration size 4 Å, TMAO and urea also shows very different behaviors at the second peak region from filtration size 5 Å to 10 Å. These differences are also reflected in the PBNs and BPEs of the local-scale IPH. These localized topological information has never been revealed before. Since graphs can be transferred into simplicial complexes by the clique complex, our weighted persistent homology models can be used in the analysis of various networks and graphs from any molecular structures and aggregation systems.
Similar content being viewed by others
Introduction
Tri-methylamine N-oxide (TMAO) and urea are organic osmolytes widely existing in the animal metabolisms. Deep-sea organisms use the protein stabilizing effects of TMAO to counteract the high pressure perturbation, while mammalian kidneys use the strong denaturant function of urea1. As a protecting osmolyte, TMAO can counteract the urea protein-denaturing effects. Currently, it is well accepted that urea acts by directly binding to the protein backbones and side chains2. It has very little disturbance on the surrounding water structures. The TMAO’s stabilization is not well understood. It has been suggested that TMAO molecules form complexes with two to three water molecules, and protein stabilization is the result of depletion effects associated with unfavorable interaction of TMAO with protein backbone3. Others suggest that TMAO interacts with polypeptides and protein stabilization is a result of surfactant-like effects of TMAO4. The interaction between TMAO and urea is also not well understood5. Even though it is suggested that the interaction is through the TMAO’s modification of urea-water structures, recent experiments show that the addition of TMAO induces blue shifts in urea’s H-N-H symmetric bending modes, indicating the direct interactions between the two cosolvents6,7. Although great progress has been made in both experimental and theoretical research for urea and TMAO8,9,10,11,12,13,14,15,16,17,18,19,20, a detailed mechanism of their molecular aggregations and the corresponding hydrogen-bonding networks still remain elusive.
Theoretically, graph or network based models21,22,23, especially the spectral graph models and combinatorial graph models, play a key role in the characterization of biomolecular structures, interaction networks, and hydrogen-bonding networks24,25,26,27,28,29,30. The most commonly-used graph-based measurements31 include, node degree, shortest path, clique, cluster coefficient, closeness, centrality, betweenness, Cheeger constant, modularity, graph Laplacian, graph spectral, Erdös number and percolation information. Differential geometry tools32,33,34,35,36,37,38, such as Voronoi diagram, alpha shape, geometric flows, have also been considered to quantitatively characterize the biomolecular structure, surface, volume, cavity, void, tunnel and interface. These models contribute greatly to the better understanding of biological systems.
Recently, a new topological model known as persistent homology has demonstrated a great promise in biomolecular structure, flexibility, dynamics and function analysis39,40,41. Persistent homology based machine learning and deep learning models42 have achieved great successes in protein-ligand binding affinity prediction43,44,45, protein stability change upon mutation46,47 and toxicity prediction48. These topology based machine learning models have constantly achieved some of the best results in D3R Grand challenge49. Motivated by the great success of topological modeling in biomolecules, we have applied persistent homology in the analysis of ion aggregations and hydrogen-bonding networks50. The two types of ion aggregation models, i.e., local clusters and extended ion networks, can be well characterized by our model. Further, we have identified, for the first time, different types of topology for the two hydrogen-bonding network systems50. Moreover, we have studied the osmolyte molecular aggregation and their hydrogen-bonding networks31. Two osmolytes, i.e., TMAO and urea, are found to share very similar topological patterns with the two types of ion systems, i.e., KSCN and NaCl. Particularly, the topological fingerprints for the hydrogen-bonding network from ion systems and osmolyte systems share a great similarity. This indicates that our topological representation can characterize certain intrinsic difference between “structure making” and “structure breaking” systems31. Features from persistent homology can be used as topological descriptors, which have also been used in a range of atomistic water models51, coarse-grained Stillinger-Weber (SW) potential model51, and aqueous solubility modeling52.
More recently, weighted persistent homology (WPH) models have been proposed to incorporate physical, chemical and biological properties into topological modeling53. Essentially, the weight value, which reflects physical, chemical and biological properties, can be assigned to vertices (atom centers), edges (bonds), or higher order simplexes (cluster of atoms), depending on the biomolecular structure, function, and dynamic properties53. In this way, weighted persistent homology can be classified into three major categories, i.e., vertex-weighted54,55,56,57,58, edge-weighted41,44,46,59,60, and simplex-weighted models61,62,63. Among them, the localized (weighted) persistent homology (LPH) and interactive persistent homology (IPH) are found to be of great importance in the classification and clustering of DNA structures and trajectories53, and protein ligand interactions46.
In this paper, for the first time, we apply the localized persistent homology and interactive persistent homology in the study of osmolyte molecular aggregation and their hydrogen-bonding networks. To quantitatively characterize the topological features from LPH and IPH, we propose Boltzmann persistent entropy (BPE). We have revealed that TMAO and urea have very different local topologies. Local network structures are observed in TMAO system. With the concentration increase, the circles within these networks show a huge increase in their total numbers and a sharp decrease in their relative sizes. In contrast, urea shows two distinguishable local topological features, i.e., local clusters around 6 Å and global-scale circle structures at around 12 Å. Further, we have demonstrated that our global-scale IPH based persistent radial distribution function (PRDF) is similar to the traditional radial distribution function (RDF) and can be used to characterize the double layer information. Moreover, a local-scale PRDFs can be generated from our local-scale IPH model. Essentially, in global-scale IPH, each osmolyte molecule interacts with all the water molecules in the system. In local-scale IPH, water molecules are classified into different cells based on the Voronoi diagram of osmolyte molecules. Interactions only happen between a central osmolyte molecule and the surrounding water molecules within its Voronoi cell and between two osmolyte molecules from closest adjacent Voronoi cells. This classification is naturally embedded in the filtration process of IPH analysis. Further, IPH based PBNs and BPEs can be used in studying the interaction patterns between osmolyte molecules and water molecules. Other than osmolyte systems, our weighted persistent homology models can be applied in the analysis of various kinds of networks and graphs from material, chemical, and biological systems.
The paper is organized as follows. A brief introduction of persistent homology and two weighted persistent homology models are given in Section 1.1 and Section 1.2. The methodology and implementation details of LPH and IPH models are discussed in Section 1.3. The main results are presented in Section 2. The LPH based molecular aggregation and hydrogen-bonding networks is discussed in Section 2.1. The IPH based topological features for osmolyte-water interaction networks are discussed in Section 2.2. The paper ends with a conclusion.
Methods and models
In this section, we will give a brief introduction of persistent homology and weighted persistent homology. Three types of persistent functions, including persistent Betti number, persistent entropy and persistent radial distribution function, will be discussed in detail. A general description of the two WPH models, i.e., localized persistent homology and interactive persistent homology, will also be presented.
Persistent homology
The persistent homology, a tool from algebraic topology and computational topology, is proposed to characterize data “shape”64. It has been widely used in data analysis64,65,66,67,68,69,70,71,72,73,74,75,76,77,78,79,80,81 with various developed softwares59,82,83,84,85,86,87 and visualization models88,89,90,91.
Persistent homology can be understood from three different aspects. Firstly, it is the relation between a graph and a simplicial complex. Mathematically, a graph, which is composed of only nodes (0-simplexes) and edges (1-simplexes), is a kind of simplicial complex. A simplicial complex K can viewed as a set of simplexes that satisfy two conditions. Firstly, any face of a simplex from K is also in K. Secondly, the intersection of any two simplexes in K is either empty or a shared face92. Other than 0- and 1-simplexes, it also includes 2-simplexes (solid triangles), 3-simplexes (tetrahedrons), and other higher-dimensional components. Secondly, it is about geometric measurements and topological invariants. In persistent homology, the data is characterized by Betti numbers, including β0, β1, β2 and higher order topological invariants93,94. These measurements are significantly different from previous geometric measurements, like distances, angles, areas, etc. Thirdly, it is the difference between single scale model and multi-scale representation. Essentially, a series of related simplical complexes are considered in persistent homology and they provide a multiscale representation that balances geometry and topology. A more detailed description of its mathematical background can be found in refs. 93,94,95, and its application in molecular biology and ion aggregation systems can be found in refs. 31,41,44,46,50,52.
Geometrically, β0 indicates the number of connected components, β1 corresponds to the number of circles, rings or loops, and β2 represents the number of voids or cavities. The key concept in persistent homology is the filtration93,94. For instance, given a point cloud data, we can associate each point with an identical-sized sphere and assign its radius as the filtration parameter. As the filtration value is increased, these spheres will systematically enlarge and subsequently merge with each other to form simplexes. Roughly speaking, an edge between two points is formed when the two corresponding spheres overlap93,94. A triangle is formed when each of two spheres (of the three corresponding spheres from triangle vertices) overlap. A tetrahedron is formed when each three spheres (of the four corresponding spheres from tetrahedron vertices) overlap92. At each filtration value, all the simplexes, i.e., vertices, edges, triangles, tetrahedrons, form a simplicial complex. From it, topological invariants, i.e., Betti numbers, can be calculated. The persistent homology hierarchically increases the complexity in data representation by systematically incorporating higher order simplices as the filtration proceeds. This enables a multiscale representation of topological invariants from simplicial complexes93,94,95. In this way, a systematic variation of the filtration parameter leads to a series of simplicial complexes at different scales93,94,95. Some topological invariants persist longer in these simplicial complexes, while others disappear quickly as the filtration value is increased. The length of the β1 bar defines the “lifespan” of the topological invariants (circles, loops, etc) and provides a natural geometric measurement93,94,95. More specifically, the lifespan, known as the persistence, measures how “large” are the circles, loops and voids in the system. We denote a filtration value at which a topological invariant formed or killed as birth time and death time respectively. In this way, each topological invariant has a “lifespan” defined by its birth and death time. Essentially, the lifespan provides a geometric measurement of the topological invariant. If we use a one-dimensional bar, which starts at a birth time and ends at a death time, to represent each homology generator, a barcode representation is generated. Figure 1 illustrates the basic topological components, including simplexes, Betti number, filtration process, and persistent barcodes.
The illustration of the basic components in persistent homology. Essentially, persistent homology is based on simplicial complex, which is composed of simplexes. In persistent homology, only topological invariants, known as Betti number, are considered. A series of simplicial complexes are generated through a filtration process. The results from persistent homology are represented as persistent barcodes.
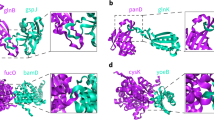
Essentially, simplicial-complex-based persistent homology models are very different from traditional graph or network models. In general, Laplacian matrixes or adjacent matrixes are constructed from graph models and their eigen spectrum information is used in structure characterization. In contrast, persistent homology describes the structure with the topological invariants together with a geometric measurement. Figure 2 illustrates the comparison between persistent homology model and traditional graph models. The two types of models reveal very different topological information of the biomolecular systems.
Illustration of the comparison between graph models and persistent homology for osmolyte molecular aggregation. (a) An ion aggregation system (from MD simulation) with both osmolytes (red balls) and waters (green balls). (b1) The graph or network representation is composed of only vertices (0-simplex) and edges (1-simplex). (b2) The simplicial complex representation has higher order simplexes, including triangles (faces) and tetrahedrons (solids). (c1) The distribution of Eigenvalues from the adjacency matrixes from the graph model. (c2) The persistent barcodes computed from a persistent homology representation.
We use notations ak,j and bk,j to represent birth times and death times of the j-th topological invariant of k-th dimension. The set of k-th dimensional barcodes is denoted as Lk. The persistent Betti number (PBN)66,88,93,96 is defined as \(f(x;{L}_{k})={\sum }_{j}\,{\chi }_{[{a}_{k,j},{b}_{k,j}]}(x)\). We propose two new functions, i.e., Boltzmann persistent entropy and Persistent radial distribution function.
Boltzmann persistent entropy
The persistent entropy has been proposed96,97,98,99 to measure the system disorder. For the k-th dimensional barcodes, it is defined as,
with the probability function,
Even though persistent entropy has been a powerful tool for the characterization of “topological disorder”, its physical meaning is usually unclear, thus hinders its further application in chemical, physical and biological systems. In this paper, we propose a Boltzmann persistent entropy (BPE) based on the Betti energy and Boltzmann distribution. Essentially, we define a Betti energy for the j-th number of k-dimension Betti bar as follows,
Here κ is an integer, η is a scale value with the same unit of the filtration parameter, and α is an energy-related constant value. The probability function is then defined according to the Boltzmann distribution,
Here kB is the Boltzmann constant and T is the thermodynamic temperature. The BPE can then be calculated from Eq. (1). Physically, when a Betti bar has a longer length, it will contribute a larger Betti energy, thus a lower probability. In contrast, a longer Betti bar has a higher probability in the traditional persistent entropy. Note that a long persisting β0 bar always exists in β0 barcodes. In traditional persistent entropy, the probability value for this long persisting bar is exactly equal to 1.0, and persistent entropy is always equal to 0 irrespective of the other β0 bars, if this long persisting bar is considered in persistent entropy. In our PBE, this bar contributes zero Betti energy according to Eq. (3), thus a probability zero. Note that in our calculation below we take α = kBT, η = 1 Å and κ = 2.
Persistent radial distribution function
Based on the β0 barcodes, we propose the persistent radial distribution function (PRDF) as follows,
Here xt is the filtration value when the PBN reduces to one, i.e., only one connected component. The integer N0 is the total number of β0 bars. Essentially, if we consider the global interactive persistent homology, our PRDF will result in the conventional radial distribution function100. On the other hand, If we use the local interactive persistent homology, our PRDF will focus on the interaction within each cell of the Voronoi diagram. A more detailed discussion is given in Section 1.2.2.
Weighted persistent homology
The weighted persistent homology models have been proposed to incorporate physical, chemical and biological properties into topological modeling53. They can also be designed to characterize local topological information and certain special interaction patterns. In this paper, we will focus on two WPH models, i.e., localized persistent homology and interactive persistent homology.
Localized persistent homology
The design of our LPH model is inspired by the great success of element specific persistent homology (ESPH)43,44. Different from all previous topological models, which consider the data/structure as an inseparable system, ESPH decomposes the data/structure into a series of subsets made of certain type(s) of atoms, which have been found to characterize very well various biological properties, such as hydrophobic or hydrophilic interactions43,44,45,46,47,48,49. Moreover, our LPH model is very different from persistent local homology92,101,102,103,104,105. Mathematically, persistent local homology studies the relative homology groups between a topological space and its subspace, while LPH explores the homology groups from local topology. Previously, LPH has been used to characterize local topological features of biomolecular structure or complexes53. In LPH, the structure is decomposed into a series of local domains or regions, that may overlap with each other, and persistent homology analysis is then systematically applied on part (or all) of these local domains or regions. In this paper, our main focus is to characterize the local features, such as ion clustering, double layer and local aggregations, that widely exists in ion or molecular aggregation and hydrogen-bonding networks.
Mathematically, the global persistent homology analysis considers the complete domain, while the localized persistent homology is performed on a local region, subdomain or subspace. Note that topological invariants for the global structure is not simply the addition of all local invariants. Stated differently, topological invariants are usually not additive! In the current paper, we define the subspace as a sphere with radius (Rc). More specifically, a sphere of radius Rc is considered around each molecule (either osmolyte or water molecule) and only the molecules within this sphere are chosen for the localized persistent homology analysis. Figure 3 illustrates the persistent homology analysis performed on molecular dynamics simulation data using two different approaches. Figure 3(a) and a(1) show the osmolyte distribution and their corresponding persistent barcodes obtained from persistent homology analysis. Figure 3(b) depicts the way of selecting local regions. Essentially, an individual molecule is selected and a sphere of radius Rc is drawn around it. All molecules within this enclosure are chosen as its local neighbors. The persistent homology analysis is carried out for all the selected molecules to generate the local persistent barcodes. This procedure is repeated for each molecule in the configuration. The corresponding persistent barcodes are as shown in Fig. 3(b1) to (b3). In essence, each molecule in a given configuration is associated with certain local neighbors which determine its local structure.
Illustration of the global and localized persistent homology analysis. The corresponding persistent barcodes for (a) Global and (b) Local approaches are demonstrated. The global persistent homology considers all the molecules in the simulation box as shown in (a), while localized persistent homology is carried out within each local region shown as grey spheres in (b). The persistent barcodes corresponding to three such local regions are illustrated from (b1)–(b3). In our LPH, we systematically consider all the molecules and generate a local region (of the same size) for each of them.
Interactive persistent homology
The interactive persistent homology (IPH) was proposed to study the interaction between proteins and ligands46. The essential idea is to study the topological invariants of the interaction networks, which are formed between protein atoms and ligand atoms. More specifically, for a protein-ligand complex, an interaction matrix can be built with its elements as the Euclidean distance between two atoms. However, if two atoms come from the same molecule (either protein or ligand), its distance is set to infinity, meaning they will never interact in IPH. In this way, the IPH model can be used in the characterization of the protein-ligand interactions. Actually, IPH based machine learning models are found to deliver the best results in protein-ligand binding affinity prediction46,47,49. Note that it seems to be better called as interaction persistent homology, as the model studies the interactions.
In this section, we use IPH models to characterize the interactions between osmolyte and water molecules. Two different models, i.e., global-scale IPH and local-scale IPH, are considered. In global-scale model, when an osmolyte molecule is selected, the distances (dij) between all the water molecules in the domain to this osmolyte molecule are considered. More specifically, suppose there are Nw number of water molecules, a global-scale IPH matrix of size (Nw + 1) × (Nw + 1) can be constructed between a selected osmolyte molecule and all water molecules as follows,
Here Type(i) is used to tell if the i-th molecule is osmolyte or water, i.e., type of the molecule. If there are Ns number of osmolyte molecule, we can construct a total Ns number of global-scale IPH matrices, with size (Nw + 1) × (Nw + 1). From these matrices, PRDF as in Eq. (4) can be calculated and the average of these PRDFs will characterize the same physical properties as the traditional radial distribution function100.
In local-scale IPH, a similar IPH matrix as in Eq. (5) is considered. But this new IPH matrix is now of size (Nw + Ns) × (Nw + Ns), meaning all distances between water and osmolyte molecules are considered simultaneously. The new IPH matrix based filtration characterizes dramatically different topological information. More specifically, molecules with shorter distances to their neighbors will form connections at earlier stage of the filtration. In this way, a Voronoi diagram will naturally form when water molecules connect to their center osmolyte molecule. Later, Voronoi cells will merge with closest neighbors to become a well-connected entity. The β0 barcodes capture very well the above topological information. And the corresponding PRDFs describe the local interactions within the Voronoi cells. A comparison of the persistent barcodes obtained from persistent homology and interactive persistent homology is illustrated in Fig. 4. It can be seen that they show totally different patterns.
The persistent barcodes obtained using persistent homology and (b) interactive persistent homology. Note that in interactive persistent homology, interactions happen only between two types of different molecules. That is to say, water molecules can only interact with osmolyte molecules and vice versa. No interactions exist between water and water, or between osmolyte and osmolyte. It can be seen that the β1 bars generated from an interactive persistent homology remain persistent forever.
Essentially, each osmolyte molecule can interact directly with all water molecules in global-scale model and the resulting PRDF (from β0 barcodes) characterizes the same physical properties as radial distribution function. In local-scale IPH model, only the interactions between the osmolyte molecule and water molecules in its Voronoi cell, and the Voronoi cell-cell interactions are captured in β0 bars. It should be noticed that the corresponding PRDFs only describe the local interaction information, and they are very different from the traditional radial distribution function. It should also be noticed that the value for our local PRDFs will decrease to zero when the filtration size is large enough.
WPH for osmolyte molecular aggregation and hydrogen-bonding network analysis
The weighted persistent homology models are considered for the study of topological structures of two types of osmolytes, namely, trimethylamine N-oxide (TMAO) and urea. Two models, i.e., localized persistent homology and interactive persistent homology, are used to reveal the local topological features in the ion aggregation, hydrogen-bonding networks and their interactions. Note that only Vietoris-Rips complex is used in all our persistent homology models.
MD simulation and data generation
The molecular trajectory or the time evolution data of the two osmolytes needed for the current work is generated using a molecular dynamics simulation. We consider the same molecular dynamics (MD) setting as in the paper31. More specifically, we consider GROMACS-5.1.2106,107 for the MD simulation. The four point (TIP4P-EW)108 water model is used, and Kast model109 is adopted for TMAO whereas the urea model is from AMBER package110. Two osmolytes with eight different concentrations, from 1 M to 8 M, in pure water are studied, respectively. To construct the initial state, urea/TMAO molecules are randomly distributed using insert-molecules utility in GROMACS, after that 3000 water molecules are inserted randomly into the cubic simulation box. We carry out the equilibration process under NVT conditions (Temperature = 300 K) for 10 ps and then under NPT conditions for 100 ps using 2 fs time step, Berendsen thermostat τ = 0.1 ps) and barostat (τ = 2 ps). LINCS algorithm111 is used for bonds and the angles constriction. Further, we carry out three repeats under NPT conditions for 100 ns with Berendsen thermostat (Temperature = 300 K, τ = 0.1 ps), Parrinello-Rahman barostat (Pressure = 1 bar, τ = 2 ps) and using a time step of 2 fs. The integration of Newton’s equation of motion is done by using a leap-frog algorithm. A cut-off of 1.0 nm is used for both van der Waals (VDW) interaction and short-range electrostatic interaction. Particle mesh Ewald (PME)112 method is employed to deal with the long-range electrostatic interactions. The configuration trajectories are output every 1 ps.
LPH analysis of osmolytes
The localized persistent homology is used to explore the topological fingerprints of molecular aggregation and hydrogen-bonding network at a local scale. In TMAO systems, there are 63, 125, 204, 290, 400, 533, 700 and 887 TMAO molecules with 3000 water molecules from concentration 1 M to 8 M, respectively. In urea systems, there are 60, 120, 192, 267, 352, 450, 555 and 681 urea molecules with 3000 water molecules for concentration 1 M to 8 M respectively. To analyze the local topology in their molecular aggregation and hydrogen-bonding networks, the TMAO and urea molecules are coarse-grained as their nitrogen and carbon atoms, respectively. The water molecules are coarse-grained as their oxygen atoms. Since the configuration data is obtained from an NPT simulation, the size of simulation box is allowed to adjust for each configuration to attain equilibrium conditions. Periodic boundary condition is used in the specification of local domains. For each simulation, we consider 101 frames (or configurations) sampled at equal intervals from the simulation trajectory. Our topological analysis is performed on these 101 frames.
In our LPH model, a local spherical region is defined for each ion using a cutoff radius Rc and the atoms within this enclosure is chosen for analysis. Persistent homology is applied to each of these local regions and the persistent barcodes are computed. We used an open source software Ripser113 for the computation of persistent barcodes. The persistent Betti functions (PBNs) and the Boltzmann persistent entropy (BPEs) are calculated from these barcodes.
IPH analysis of osmolytes
Both global-scale and local-scale IPHs are considered for analyzing the interactions between osmolyte molecules and water molecules. In global-scale IPH, for each osmolyte molecule, we can construct a series of IPH matrixes as in Eq. (5) with the same size of 3001 × 3001, as there are totally 3000 water molecules. From the β0 barcode of the IPH matrices, a single PRDF can be calculated. Further by averaging the PRDFs over all the 101 frames and all osmolyte molecules in each frame, we can obtain the average global-scale PRDF.
In local-scale IPH, for each configuration or frame, an individual IPH can be constructed. Note that the size of the local-scale IPH matrix as in Eq. (5) is (Nw + Ns) × (Nw + Ns), i.e., the total number of osmolyte and water molecules in the simulation. The average local-scale PRDF (or PBN) can be evaluated by averaging their values over all the 101 frames.
Results and discussions
In this section, we systematically study the local topological features and interaction properties of the osmolyte molecular aggregation and their hydrogen-bonding networks. The corresponding PBNs, BPEs and PRDFs are used to quantitatively characterize the intrinsic local topology information.
LPH for molecular aggregation and hydrogen-bonding networks
To facilitate an intuitive understanding of local topological information of molecular aggregation, we demonstrate the persistent barcodes calculated from TMAO and urea systems with a cutoff radius of Rc = 9 Å. More specifically, we consider the last configuration of the MD simulation from four different concentrations. An osmolyte molecule is randomly chosen from the last frame and its neighbouring osmolyte molecules located within the cutoff radius Rc = 9 Å are selected. Persistent homology analysis is then applied on these molecules. The results from TMAO and urea systems are demonstrated in Fig. 5. The indexes (a) and (b) corresponds to TMAO and urea respectively. The subscripts 1–4 indicates the four different concentrations considered, i.e., 2 M, 4 M, 6 M, and 8 M respectively. In both TMAO and urea, the total number of β0 bars roughly increase with concentration (M), indicating the aggregation of neighboring molecules with the concentration. The β1 bars also seem to appear more and more frequently with the increase in concentration.
The local persistent barcodes for TMAO and urea aggregation. TMAO and urea molecules are coarse-grained as their nitrogen and carbon atoms. Subfigures (a1 to a4) represent the results from TMAO with concentration 2 M, 4 M, 6 M, and 8 M, respectively. Subfigures (b1 to b4) represent the results from urea with concentration 2 M, 4 M, 6 M, and 8 M, respectively. The barcodes are generated from a randomly picked molecule at the last frame of the simulation. A cutoff radius of 9 Å is used. Roughly speaking, both β0 and β1 bars tend to increase with the concentration.
The results shown in Fig. 5 are based on a randomly chosen molecule in the last frame of the simulation trajectories and can not characterize the overall behavior very well. To have a better comparison, we consider the ensemble average. Meaning, for each frame, the local persistent barcode from each molecule is calculated and then averaged. It should be noticed that we use the periodic boundary condition to include all the “neighboring” molecules. This process is repeated for all the 101 frames in each trajectory. We represent each persistent barcode as their PBN and BPEs. These PBNs are then averaged over all the frames and all the molecules in each frame to generate a single PBN for each simulation or trajectory. The BPEs are averaged over all the molecules in each frame, so that a total 101 BPEs are obtained from each simulation.
Figure 6 shows the β1 PBNs obtained for the TMAO and urea system at eight different concentrations, from 1 M to 8 M, using three different cutoff radii. The indexes (a) and (b) corresponds to TMAO and urea respectively. The subscripts 1–3 indicates the three different cutoff radii namely., Rc = 9 Å, 12 Å and 15 Å respectively. The corresponding global persistent homology analysis for TMAO and urea is shown in (a4) and (b4) respectively. As stated above, β1 bars represent the ring, circle and loop structures in the system. For TMAO system, at each cutoff radius, the peak value of the local β1 PBNs systematically increases with the concentration, indicating that more and more circle structures are generated. At the same time, the position of these peak values shifts from around 13 Å to 7 Å, which implies a systematic decrease in the size of these circles. When we consider larger cutoff radii, similar topological patterns are observed. However, the peak values of PBNs from lower concentration systems increase much faster, even though all PBN peak values increase with the cutoff radius. This result indicates that for a lower concentration system, there exists large-sized topological features which can not be well characterized by LPH with a small cutoff radius. For urea system, their PBNs have a dramatically different behavior in comparison with TMAO. Roughly speaking, there are two types of peak for urea system, especially the urea system at lower concentrations. One type of peak is located around 5 Å, and the other is around 10 Å to 12 Å. The peak at 5 Å appears even at very low concentrations and its magnitude keeps increasing with the concentration rise. The shape of this peak is much narrower than that of TMAO PBNs. The second type of peak can only be distinctly observed at lower concentrations. It has much smaller magnitude compared with that of the first type of peak.
The comparison of average β1 PBNs for (a) TMAO and (b) urea at eight different concentrations from 1 M to 8 M. Subfigures (a1 to a3) are results from TMAO with three different cutoff radii namely., Rc = 9 Å, 12 Å and 15 Å, respectively. Subfigures (b1 to b3) are results from urea with three different cutoff radii namely., Rc = 9 Å, 12 Å and 15 Å, respectively. The β1 PBNs are averaged over all the frames and all molecules in each frame. Subfigures (a4) and (b4) are the PBNs obtained from a global persistent homology analysis. It can be seen that, TMAO and urea show dramatically different local topological characteristics.
From Fig. 6, we can also see that TMAO and urea demonstrate dramatically different local topological characteristics. Essentially, TMAO shows a regular local network structure. The size and total number of the circle structures from these networks consistently decrease and increase with the concentration, respectively. In contrast, urea shows a cluster-like local aggregations. Urea molecules form local clusters, whose size stays relatively consistent but the total number consistently increases with concentration. More interestingly, if we compare our LPH results with the ones from persistent homology analysis of the whole osmolyte systems31, as in Fig. 6(a4) and (b4), we can find some unique similarities and differences. Essentially, the general pattern of PBNs at lower filtration values has less changes and remains relatively stable, while PBNs at larger filtration values change more dramatically. Stated differently, the LPH focuses more on the local topological information and systematically attenuates the influence from global topological features.
Other than the PBNs, we can also calculate BPEs from the LPH barcodes and use them to characterize the “topological regularity”. Figure 7 demonstrates the β1 BPEs for both TMAO and urea at eight concentrations and three cutoff radii as stated above. Note that for each simulation or trajectory, we consider 101 configurations or frames which generates 101 β1 BPEs. It can be seen that, at a small cutoff radius, the BPE values from 1 M concentration is almost all zeros, meaning that there is almost no circle structures at local scale. This is consistent with the PBN profile in Fig. 6. Further, the average BPE value increases systematically with the concentration for both TMAO and urea. However, the BPE variance shows a very different behavior. With the concentration increase, the TMAO BPE variance systematically decreases, while urea BPE variance consistently increases. These results are also consistent with our findings from persistent homology analysis of the whole system31 and is also presented here for clarity. Essentially, with the concentration increase, all osmolyte systems become topologically more and more disordered. However, the variation of topological regularity for each trajectory decreases in the TMAO system but increases in the urea system. The BPE are found to be consistent with the global persistent homology analysis as shown in Fig.7(a4) and (b4) for TMAO and Urea respectively.
The comparison of average persistent entropies for (a) TMAO and (b) urea at eight different concentrations from 1 M to 8 M. Subfigures (a1 to a3) are results from TMAO with three different cutoff radii namely., Rc = 9 Å, 12 Å and 15 Å, respectively. Subfigures (b1 to b3) are results from urea with three different cutoff radii namely., Rc = 9 Å, 12 Å and 15 Å, respectively. Subfigures (a4) and (b4) are the PEs for TMAO and urea obtained from global persistent homology analysis, respectively. The BPEs are averaged over all the molecules in each frames, thus a total 101 BPEs are obtained for each simulation. It can be seen that, for a small cutoff radius of Rc = 9 Å, both TMAO and urea BPEs at 1 M are almost all zero. Further, the average BPEs for both systems increase with the concentration, but their BPE variances have very different properties. The TMAO BPE variance decreases with concentration while urea BPE variance increases.
To have an intuitive understanding of the inner topological differences between TMAO and urea molecular aggregation, we generate simplicial complexes from the last frame of the simulation data of TMAO and urea at highest concentration (8 M). For a better visualization, we consider the value of filtration parameter r to be 5 Å, 6 Å, 7 Å and 8 Å, and plot only the 2-simplexes (triangles) and 3-simplexex (tetrahedrons). The results are illustrated in Fig. 8. It can be seen that, TMAO molecules are more evenly distributed, while urea molecules tend to concentrate into clusters. Topologically, evenly-distributed molecules will generate more “large” circle structures (longer bars in β1 barcodes), while local clustering contributes more small circles (shorter bars in β1 barcodes).
The comparison of the simplicial complexes from TMAO and urea molecule aggregation at 8 M concentration. The subfigures (a1 to a4) are for TMAO and subfigures (b1 to b4) are for TMAO. The filtration for (a1 to a4) is 5 Å, 6 Å, 7 Å and 8 Å, respectively. Note that we only plot the 2-simplexes and 3-simplexes for better visualization. The same setting is used for urea systems in b1 to b4.
LPH based topological features of hydrogen-bonding networks
In our hydrogen-bonding network analysis, we consider the topological features for water molecules at a local scale. Similar to osmolyte systems, The LPH analysis is carried out for each water molecule along with its neighbours located within a cutoff radius Rc. For each frame, we systematically go over all the 3000 water molecules and calculate 3000 local persistent barcodes. Again periodic boundary condition is considered to include all “neighboring” water molecules. The process is repeated over all the 101 frames in each trajectory. A single β1 PBN is generated for each simulation by averaging β1 PBNs over all the 101 frames and all the 3000 water molecules in each frame. The β1 BPEs are averaged over the 3000 water molecules in each frame, so that a total 101 β1 BPEs are obtained from each simulation. Two cutoff radii, i.e, Rc = 7 Å and Rc = 9 Å, are considered in our LPH analysis.
Figure 9 shows the comparison of average β1 PBNs for TMAO and urea hydrogen-bonding networks. Figure 9(a1) and (a2) indicates that, for TMAO system, the PBNs have a peak value located at around 3.5 Å. With the concentration increase, the peak value of TMAO PBNs gradually decreases. In the meantime, there is a consistent rise of the PBN values in the range from around 4.5 Å to 7.0 Å. Even though all PBNs significantly increase with the cutoff radius, the general PBN profile pattern from eight different concentrations is highly consistent. Similar to TMAO, Fig.9(b1) and (b2) shows that, urea PBNs also have a peak value at filtration value 3.5 Å. The peak value slightly decreases with the concentration increase. Further, the general PBN profile pattern from eight different concentrations shares a remarkable similarity at two different local scales, even though the PBN peak values are systematically increased.
The comparison of average β1 PBNs for hydrogen bonding networks of (a) TMAO and (b) urea using two different cutoff radii at eight different concentrations from 1 M to 8 M. Subfigures (a1 to a2) are results from TMAO hydrogen bonding networks with two different cutoff radii namely., Rc = 7 Å and 9 Å, respectively. Subfigures (b1 to b2) are results from urea hydrogen bonding networks with two different cutoff radii namely., Rc = 7 Å and 9 Å, respectively. Subfigures (a3) and (b3) are the corresponding PEs obtained from global persistent homology analysis. The coarse-grained representation of water as its oxygen atom is considered. The indexes (a) and (b) corresponds to TMAO and urea respectively. The PBNs are averaged over all the molecules and configuration numbers. It can be seen that, TMAO and urea show very different topological characteristics.
From Fig. 9, we can see that TMAO and urea hydrogen-bonding networks demonstrate dramatically different local topological characteristics. For TMAO hydrogen-bonding networks, with the concentration increase, there is a systematic decrease of small-sized circle structures as well as an increase of relatively large-sized circle structures. For urea hydrogen-bonding networks, there is only a slight decrease of small-sized circle structures and no significant increase of large-sized circle structures. More interestingly, if we compare our LPH results with the ones from the whole hydrogen-bonding network in both ion and osmolyte systems31,50, we can see that there exists a great similarity in their PBNs. Essentially, TMAO and urea hydrogen-bonding networks show two types of topological behaviors. With the concentration increase, TMAO molecules tend to destroy the local hydrogen-bonding networks, resulting in a significant increase of the large circle structures. In contrast, the urea molecules have a much less impact on the hydrogen-bonding networks.
The persistent entropy from the LPH barcodes can also be used to characterize the “topological regularity” of hydrogen-bonding networks. Figure 10 demonstrates the β1 BPEs for both TMAO and urea hydrogen-bonding networks at eight concentrations and two cutoff radii. The indexes (a) and (b) denote TMAO and urea systems respectively, at eight different concentrations from 1 M to 8 M. The subscripts 1–2 indicates the cutoff radii Rc = 7 Å and Rc = 9 Å respectively. Similar to molecular aggregation analysis, for each simulation, we consider 101 configurations or frames and generate 101 β1 BPEs. It can be seen that, the average BPE value for both TMAO and urea hydrogen-bonding networks decreases with the concentration increase. The same pattern is observed at two local scales. Topologically, these results indicate that both the hydrogen-bonding networks become more and more regular and lattice-like with concentration increase. Note that molecular aggregation has a totally different topological behavior, their BPE value systematically increases with the concentration. More interestingly, the urea BPE variance is significantly larger than that of TMAO and consistently increases with the concentration. This is exactly the same as in the urea aggregation system.
The comparison of average β1 BPEs for hydrogen-bonding networks from (a) TMAO and (b) urea systems using two different cutoff radii at eight different concentrations from 1 M to 8 M. Subfigures (a1 to a2) are results from TMAO hydrogen bonding networks with two different cutoff radii namely., Rc = 7 Å and 9 Å, respectively. Subfigures (b1 to b2) are results from urea hydrogen bonding networks with two different cutoff radii namely., Rc = 7 Å and 9 Å, respectively. The BPEs are averaged over all the water molecules in each frames, thus a total 101 BPEs are obtained for each simulation. It can be seen that, the average BPE decreases with the concentration for both TMAO and urea. However, the BPE variance for urea systematically increases. (a3) and (b3) are the PEs obtained from a global persistent homology analysis.
In summary, we have used LPH models to explore the osmolyte molecular aggregation and their hydrogen-bonding networks. Essentially, we segregate osmolyte molecules from water molecules, and study their local topological features separately. In the next section, we will focus on the interaction between osmolyte molecules and water molecules and characterize the topology of their interaction networks.
Figure 11 illustrates the simplicial complexes of the hydrogen-bonding networks from TMAO and urea. They are generated from the last frame of the simulation data of TMAO and urea at highest concentration (8 M). We consider the value of filtration parameter r to be 3 Å, 4 Å, 5 Å and 6 Å, and plot only the 2-simplexes (triangles) and 3-simplexex (tetrahedrons), for a better visualization. It can be seen that, similar to the results in Fig. 8, water molecules in TMAO systems are more evenly distributed, while water molecules in urea sytems tend to concentrate into clusters. Topologically, evenly-distributed water molecules will generate more “large” circle structures (longer bars in β1 barcodes), while local clustering contributes more small circles (shorter bars in β1 barcodes). For the above analysis, it can be noticed that persistent barcode provides a unique way of analyzing the inner topological structures of the systems.
The comparison of the simplicial complexes from hydrogen-bonding networks from TMAO and urea molecules at 8 M concentration. The subfigures (a1 to a4) are for TMAO and subfigures (b1 to b4) are for TMAO. The filtration values for (a1 to a4) is 3 Å, 4 Å, 5 Å and 6 Å, respectively. Note that we only plot the 2-simplexes and 3-simplexes for better visualization. The same setting is used for urea systems in b1 to b4.
IPH based topological features for osmolyte-water interaction network
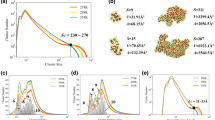
Figure 12 shows the comparison of global-scale and local-scale PRDFs for both TMAO and urea systems. The indexes (a) and (b) represents TMAO and urea respectively. The subscripts 1–2 corresponds to the global-scale and local-scale PRDFs respectively. Both global-scale and local-scale PRDFs are normalized with the number density of the oxygen atom. The number density in global-scale is calculated by considering the number of oxygen atoms averaged over all the spheres around each ion with radius rmax. The value of rmax is half the box size. In the local-scale, the number density is simply the number of oxygen atoms divided by the volume of the simulation box for a given concentration. Essentially, the global-scale PRDFs are identical to the traditional radial density functions. It can be seen that both TMAO and urea have two very obvious peaks, one located at around 4 Å and the other located at around 7 Å. However, their behaviors are dramatically different. For TMAO, the first peak value consistently increases with the concentration while the second peak value decreases with the concentration. The change of the TMAO peak values are relatively small, especially for the second peak. In contrast, both peaks of urea PRDFs vary greatly with concentration change. In local-scale IPH, PRDF values converge quickly to zero when the filtration value is larger than 12 Å, which is dramatically different from the situation in global-scale PRDFs when their values converge to 1 at large filtration value. However, the first peak of local-scale PRDFs has similar pattern as the global-scale ones. The TMAO peak value increases with concentration, while urea peak value decreases with concentration. Moreover, at the region of filtration value from 5 Å and 10 Å, which is the region for the second peak of global PRDFs, the TMAO PRDF values decrease much faster than those of urea. When the concentration is larger than 5 M, nearly all TMAO PRDF values drops to zero, while urea PRDF still remains largely positive.
The comparison of global-scale and local-scale PRDFs for (a) TMAO and (b) urea. Subfigures (a1 to a2) are results from global-scale and local-scale PRDFs of TMAO systems, respectively. Subfigures (b1 to b2) are results from global-scale and local-scale PRDFs of urea systems, respectively. The PRDF of N-O is examined in the case of TMAO and C-O for the urea osmolyte. It can be seen that, the first peak value of the TMAO PRDFs increases with the concentration, while the first peak value of urea PRDFs decreases with the concentration.
To have a better understanding of the local-scale PRDFs, we check the PBNs and PEs from the local-scale IPH. Figure 13 demonstrates β0 PBNs for TMAO and urea. The indices (a) and (b) represents TMAO and urea respectively. The subscripts 1–2 corresponds to the PBNs and PEs respectively. The β0 PBNs are directly related to PRDFs. It can be seen that indeed the TMAO β0 PBNs decrease much faster than those of TMAO at the filtration region from 5 Å to 10 Å, consistent with our observations in local-scale PRDFs. Further, we study the corresponding BPEs. It can be seen in Fig. 13, that the average BPE values for both local-scale IPH models increase with the concentration. More interestingly, the BPE variance for TMAO decreases with the concentration, while that for urea systematically increases with the concentration.
The comparison of average β0 PBNs and BPEs for hydrogen-bonding networks of (a) TMAO and (b) urea systems. Subfigures (a1 to a2) are TMAO β0 PBNs and β0 BPEs, respectively. Subfigures (b1 to b2) are urea β0 PBNs and β0 BPEs, respectively. The N-O pair interaction network is considered for analysis. The comparison of average β0 BPEs from the IPH analysis of TMAO and urea systems. For each configuration, a BPE value can be calculated, thus a total 101 BPEs are obtained for each simulation. It can be seen that, the average BPE increases with the concentration for both TMAO and urea. However, the BPE variance for urea systematically increases.
Conclusion
In this paper, we use the weighted persistent homology to study the topological properties for osmolyte molecular aggregation and their hydrogen-bonding networks at a local scale. Two different models, i.e., localized persistent homology (LPH) and interactive persistent homology (IPH), are considered. We propose Boltzmann persistent entropy (BPE) to quantitatively characterize the topological features from LPH and IPH, together with persistent Betti number (PBN). Based on persistent barcodes, we have proposed the persistent radial distribution function (PRDF). It has been found that the global-scale PRDF will reduce to traditional radial distribution function. While local-scale PRDFs can efficiently characterize the local interactions within the Voronoi cells. We will consider MD simulations, including high pressure conditions, solution with both urea and TMAO, and protein with TMAO or urea, in our future works to fully elucidate the mechanisms for protein stabilization. Note that since any graph can be constructed into a simplicial complex (through clique complex), our weighted persistent homology models can be used in the analysis of graph and network structures from material, chemical and biological systems114,115,116.
Data availability
Our codes are available at https://www.ntu.edu.sg/home/xiakelin/WPH_Osmolytes.zip.
References
Sahle, C. J., Schroer, M. A., Juurinen, I. & Niskanen, J. Influence of TMAO and urea on the structure of water studied by inelastic X-ray scattering. Physical Chemistry Chemical Physics 18(24), 16518–16526 (2016).
Ganguly, P., Boserman, P., van der Vegt, N. F. & Shea, J. E. Trimethylamine N-oxide counteracts urea denaturation by inhibiting protein–urea preferential interaction. Journal of the American Chemical Society 140(1), 483–492 (2017).
Hunger, J., Ottosson, N., Mazur, K., Bonn, M. & Bakker, H. J. Water-mediated interactions between trimethylamine-N-oxide and urea. Physical Chemistry Chemical Physics 17(1), 298–306 (2015).
Liao, Y. T., Manson, A. C., DeLyser, M. R., Noid, W. G. & Cremer, P. S. Trimethylamine N-oxide stabilizes proteins via a distinct mechanism compared with betaine and glycine. Proceedings of the National Academy of Sciences 114(10), 2479–2484 (2017).
Ganguly, P., van der Vegt, N. F. & Shea, J. E. Hydrophobic association in mixed urea–TMAO solutions. The journal of physical chemistry letters 7(15), 3052–3059 (2016).
Xie, W. J. et al. Large hydrogen-bond mismatch between TMAO and urea promotes their hydrophobic association. Chem 4(11), 2615–2627 (2018).
Zetterholm, S. G. et al. Noncovalent interactions between Trimethylamine N-Oxide (TMAO), urea, and water. The Journal of Physical Chemistry B 122(38), 8805–8811 (2018).
Bandyopadhyay, D., Mohan, S., Ghosh, S. K. & Choudhury, N. Molecular dynamics simulation of aqueous urea solution: is urea a structure breaker? The Journal of Physical Chemistry B 118(40), 11757–11768 (2014).
Baskakov, I. & Bolen, D. W. Forcing thermodynamically unfolded proteins to fold. Journal of Biological Chemistry 273(9), 4831–4834 (1998).
Baskakov, I. V. et al. Trimethylamine N-oxide-induced cooperative folding of an intrinsically unfolded transcription-activating fragment of human glucocorticoid receptor. Journal of Biological Chemistry 274(16), 10693–10696 (1999).
Ganguly, P., Hajari, T., Shea, J. E. & van der Vegt, N. F. A. Mutual exclusion of urea and trimethylamine N-oxide from amino acids in mixed solvent environment. The journal of physical chemistry letters 6(4), 581–585 (2015).
Idrissi, A. et al. The effect of urea on the structure of water: A molecular dynamics simulation. The Journal of Physical Chemistry B 114(13), 4731–4738 (2010).
Meersman, F., Bowron, D., Soper, A. K. & Koch, M. H. J. An X-ray and neutron scattering study of the equilibrium between trimethylamine N-oxide and urea in aqueous solution. Physical Chemistry Chemical Physics 13(30), 13765–13771 (2011).
Panuszko, A., Bruzdziak, P., Zielkiewicz, J., Wyrzykowski, D. & Stangret, J. Effects of urea and trimethylamine-N-oxide on the properties of water and the secondary structure of hen egg white lysozyme. The Journal of Physical Chemistry B 113(44), 14797–14809 (2009).
Paul, S. & Patey, G. N. Structure and interaction in aqueous urea-trimethylamine-N-oxide solutions. Journal of the American Chemical Society 129(14), 4476–4482 (2007).
Rezus, Y. L. A. & Bakker, H. J. Effect of urea on the structural dynamics of water. Proceedings of the National Academy of Sciences 103(49), 18417–18420 (2006).
Rosgen, J. & Jackson-Atogi, R. Volume exclusion and H-bonding dominate the thermodynamics and solvation of trimethylamine-N-oxide in aqueous urea. Journal of the American Chemical Society 134(7), 3590–3597 (2012).
Rossky, P. J. Protein denaturation by urea: slash and bond. Proceedings of the National Academy of Sciences 105(44), 16825–16826 (2008).
Tseng, H. C. & Graves, D. J. Natural methylamine osmolytes, trimethylamine N-oxide and betaine, increase tau-induced polymerization of microtubules. Biochemical and biophysical research communications 250(3), 726–730 (1998).
Uversky, V. N., Li, J. & Fink, A. L. Trimethylamine-N-oxide-induced folding of α-synuclein. FEBS letters 509(1), 31–35 (2001).
dos Santos, V. M. L., Moreira, F. G. B. & Longo, R. L. Topology of the hydrogen bond networks in liquid water at room and supercritical conditions: a small-world structure. Chemical physics letters 390(1), 157–161 (2004).
Oleinikova, A., Smolin, N., Brovchenko, I. & Geiger, A. and Roland Winter. Formation of spanning water networks on protein surfaces via 2D percolation transition. The journal of physical chemistry B 109(5), 1988–1998 (2005).
Radhakrishnan, T. P. & Herndon, W. C. Graph theoretical analysis of water clusters. The Journal of Physical Chemistry 95(26), 10609–10617 (1991).
Bakó, I. et al. Hydrogen bond network topology in liquid water and methanol: a graph theory approach. Physical Chemistry Chemical Physics 15(36), 15163–15171 (2013).
Bakó, I., Megyes, T., Bálint, S., Grósz, T. & Chihaia, V. Water–methanol mixtures: topology of hydrogen bonded network. Physical Chemistry Chemical Physics 10(32), 5004–5011 (2008).
Choi, J. & Cho, M. Ion aggregation in high salt solutions. II. spectral graph analysis of water hydrogen-bonding network and ion aggregate structures. The Journal of chemical physics 141(15), 154502 (2014).
Choi, J. & Cho, M. Ion aggregation in high salt solutions. IV. graph-theoretical analyses of ion aggregate structure and water hydrogen bonding network. The Journal of chemical physics 143(10), 104110 (2015).
Choi, J. & Cho, M. Ion aggregation in high salt solutions. VI. spectral graph analysis of chaotropic ion aggregates. The Journal of chemical physics 145(17), 174501 (2016).
Choi, J. H., Lee, H., Choi, H. R. & Cho, M. Graph theory and ion and molecular aggregation in aqueous solutions. Annual review of physical chemistry 69, 125–149 (2018).
da Silva, J. A. B., Moreira, F. G. B., dos Santos, V. M. L. & Longo, R. L. On the hydrogen bond networks in the water–methanol mixtures: topology, percolation and small-world. Physical Chemistry Chemical Physics 13(14), 6452–6461 (2011).
Meng, Z. Y., Anand, D. V., Lu, Y. P., Wu, J. & Xia, K. L. Persistent homology analysis of osmolyte molecular aggregation and their hydrogen-bonding networks. Physical Chemistry Chemical Physics 21, 21038–21048 (2019).
Bates, P. W. & Wei, G. W. and Shan Zhao. Minimal molecular surfaces and their applications. Journal of Computational Chemistry 29(3), 380–91 (2008).
Cazals, F., Proust, F., Bahadur, R. P. & Janin, J. Revisiting the voronoi description of protein–protein interfaces. Protein Science 15(9), 2082–2092 (2006).
Chalikian, T. V. & Breslauer, K. J. Thermodynamic analysis of biomolecules: a volumetric approach. Current opinion in structural biology 8(5), 657–664 (1998).
Edelsbrunner, H. & Koehl, P. The geometry of biomolecular solvation. Combinatorial and computational geometry 52, 243–275 (2005).
Feng, X., Xia, K. L., Tong, Y. Y. & Wei, G. W. Multiscale geometric modeling of macromolecules II: lagrangian representation. Journal of Computational Chemistry 34, 2100–2120 (2013).
Petřek, M., Košinová, P., Koča, J. & Otyepka, M. MOLE: a voronoi diagram-based explorer of molecular channels, pores, and tunnels. Structure 15(11), 1357–1363 (2007).
Smolin, N. et al. TMAO and urea in the hydration shell of the protein snase. Physical Chemistry Chemical Physics 19(9), 6345–6357 (2017).
Wang, B. & Wei, G. W. Object-oriented persistent homology. Journal of Computational Physics 305, 276–299 (2016).
Xia, K. L., Feng, X., Tong, Y. Y. & Wei, G. W. Persistent homology for the quantitative prediction of fullerene stability. Journal of Computational Chemsitry 36, 408–422 (2015).
Xia, K. L. & Wei, G. W. Persistent homology analysis of protein structure, flexibility, and folding. International journal for numerical methods in biomedical engineering 30(8), 814–844 (2014).
Pun, C. S. Xia, K. L. and S. X. Lee. Persistent-homology-based machine learning and its applications–a survey. arXiv preprint arXiv:1811.00252 (2018).
Cang, Z. X. and Wei. G. W. Integration of element specific persistent homology and machine learning for protein-ligand binding affinity prediction. International journal for numerical methods in biomedical engineering, page 10.1002/cnm.2914 (2017).
Cang, Z. X. & Wei, G. W. TopologyNet: Topology based deep convolutional and multi-task neural networks for biomolecular property predictions. PLOS Computational Biology 13(7), e1005690 (2017).
Nguyen, D. D., Xiao, T., Wang, M. L. & Wei, G. W. Rigidity strengthening: A mechanism for protein–ligand binding. Journal of chemical information and modeling 57(7), 1715–1721 (2017).
Cang, Z. X., Mu, L. & Wei, G. W. Representability of algebraic topology for biomolecules in machine learning based scoring and virtual screening. PLoS computational biology 14(1), e1005929 (2018).
Cang, Z. X. & Wei, G. W. Analysis and prediction of protein folding energy changes upon mutation by element specific persistent homology. Bioinformatics 33(22), 3549–3557 (2017).
Wu, K. D. and Wei, G. W. Quantitative toxicity prediction using topology based multi-task deep neural networks. Journal of chemical information and modeling, page https://doi.org/10.1021/acs.jcim.7b00558 (2018).
Nguyen, D. D. et al. Wei. Mathematical deep learning for pose and binding affinity prediction and ranking in D3R Grand Challenges. Journal of computer-aided molecular design 33(1), 71–82 (2019).
Xia, K. L. Persistent homology analysis of ion aggregations and hydrogen-bonding networks. Physical Chemistry Chemical Physics 20(19), 13448–13460 (2018).
Steinberg, L., Russo, J. & Frey, J. G. A new topological descriptor for water network structure. Journal of cheminformatics 11(1), 48 (2019).
Pirashvili, M. et al. Improved understanding of aqueous solubility modeling through topological data analysis. Journal of cheminformatics 10(1), 54 (2018).
Meng, Z. Y., Anand, D. V., Lu, Y. P., Wu, J. & Xia, K. L. Weighted persistent homology for biomolecular data analysis. Scientific Report 10(1), 1–15 (2020).
G. Bell, A. Lawson, J. Martin, J. Rudzinski, and C. Smyth. Weighted persistent homology. arXiv preprint arXiv:1709.00097 (2017).
Buchet, M., Chazal, F., Oudot, S. Y. & Sheehy, D. R. Efficient and robust persistent homology for measures. Computational Geometry 58, 70–96 (2016).
H. Edelsbrunner. Weighted alpha shapes, volume 92. University of Illinois at Urbana-Champaign, Department of Computer Science (1992).
Guibas, L., Morozov, D. & Mérigot, Q. Witnessed k-distance. Discrete & Computational Geometry 49(1), 22–45 (2013).
Xia, K. L. & Wei, G. W. Multidimensional persistence in biomolecular data. Journal Computational Chemistry 36, 1502–1520 (2015).
Binchi, J., Merelli, E., Rucco, M., Petri, G. & Vaccarino, F. jholes: A tool for understanding biological complex networks via clique weight rank persistent homology. Electronic Notes in Theoretical Computer Science 306, 5–18 (2014).
Petri, G., Scolamiero, M., Donato, I. & Vaccarino, F. Topological strata of weighted complex networks. PloS one 8(6), e66506 (2013).
Dawson, R. J. M. Homology of weighted simplicial complexes. Cahiers de Topologie et Géométrie Différentielle Catégoriques 31(3), 229–243 (1990).
Ren, S. Q., Wu, C. Y. & Wu, J. Weighted persistent homology. Rocky Mountain Journal of Mathematics 48(8), 2661–2687 (2018).
C. Y. Wu, S. Q. Ren, J. Wu, and K. L. Xia. Weighted (co) homology and weighted laplacian. arXiv preprint arXiv:1804.06990 (2018).
Carlsson, G., Ishkhanov, T., Silva, V. & Zomorodian, A. On the local behavior of spaces of natural images. International Journal of Computer Vision 76(1), 1–12 (2008).
Bendich, P., Edelsbrunner, H. & Kerber, M. Computing robustness and persistence for images. IEEE Transactions on Visualization and Computer Graphics 16, 1251–1260 (2010).
Carlsson, G. Topology and data. Am. Math. Soc 46(2), 255–308 (2009).
Di Fabio, B. & Landi, C. A Mayer-Vietoris formula for persistent homology with an application to shape recognition in the presence of occlusions. Foundations of Computational Mathematics 11, 499–527 (2011).
Frosini, P. & Landi, C. Persistent Betti numbers for a noise tolerant shape-based approach to image retrieval. Pattern Recognition Letters 34(8), 863–872 (2013).
M. Gameiro et al. Topological measurement of protein compressibility via persistence diagrams. preprint (2013).
D. Horak, S Maletic, and M. Rajkovic. Persistent homology of complex networks. Journal of Statistical Mechanics: Theory and Experiment, 2009(03):P03034 (2009).
Kasson, P. M. et al. Persistent voids a new structural metric for membrane fusion. Bioinformatics 23, 1753–1759 (2007).
Lee, H., Kang, H., Chung, M. K., Kim, B. & Lee, D. S. Persistent brain network homology from the perspective of dendrogram. Medical Imaging, IEEE Transactions on 31(12), 2267–2277 (Dec 2012).
Liu, X., Xie, Z. & Yi, D. Y. A fast algorithm for constructing topological structure in large data. Homology, Homotopy and Applications 14, 221–238 (2012).
Mischaikow, K., Mrozek, M., Reiss, J. & Szymczak, A. Construction of symbolic dynamics from experimental time series. Physical Review Letters 82, 1144–1147 (1999).
Niyogi, P., Smale, S. & Weinberger, S. A topological view of unsupervised learning from noisy data. SIAM Journal on Computing 40, 646–663 (2011).
D. Pachauri, C. Hinrichs, M. K. Chung, S. C. Johnson, and V. Singh. Topology-based kernels with application to inference problems in alzheimer’s disease. Medical Imaging, IEEE Transactions on, 30(10):1760–1770, Oct (2011).
Rieck, B., Mara, H. & Leitte, H. Multivariate data analysis using persistence-based filtering and topological signatures. IEEE Transactions on Visualization and Computer Graphics 18, 2382–2391 (2012).
Silva V. D. and Ghrist R. Blind swarms for coverage in 2-d. In In Proceedings of Robotics: Science and Systems, page 01 (2005).
G. Singh et al. Topological analysis of population activity in visual cortex. Journal of Vision, 8(8) (2008).
Wang, B., Summa, B., Pascucci, V. & Vejdemo-Johansson, M. Branching and circular features in high dimensional data. IEEE Transactions on Visualization and Computer Graphics 17, 1902–1911 (2011).
Yao, Y. et al. Topological methods for exploring low-density states in biomolecular folding pathways. The Journal of Chemical Physics 130, 144115 (2009).
Dionysus: the persistent homology software. Software available at http://www.mrzv.org/software/dionysus.
U. Bauer, M. Kerber, and J. Reininghaus. Distributed computation of persistent homology. Proceedings of the Sixteenth Workshop on Algorithm Engineering and Experiments (ALENEX) (2014).
B. T. Fasy, J. Kim, F. Lecci, and C. Maria. Introduction to the R package TDA. arXiv preprint arXiv:1411.1830 (2014).
C. Maria. Filtered complexes. In GUDHI User and Reference Manual. GUDHI Editorial Board (2015).
Vidit N. Perseus: the persistent homology software. Software available at http://www.sas.upenn.edu/vnanda/perseus.
A Tausz, M Vejdemo-Johansson, and H Adams. Javaplex: A research software package for persistent (co)homology. Software available at http://code.google.com/p/javaplex (2011).
Bubenik, P. Statistical topological data analysis using persistence landscapes. The Journal of Machine Learning Research 16(1), 77–102 (2015).
Bubenik, P. & Kim, P. T. A statistical approach to persistent homology. Homology, Homotopy and Applications 19, 337–362 (2007).
Ghrist, R. Barcodes: the persistent topology of data. Bulletin of the American Mathematical Society 45(1), 61–75 (2008).
Mischaikow, K. & Nanda, V. Morse theory for filtrations and efficient computation of persistent homology. Discrete and Computational Geometry 50(2), 330–353 (2013).
Munkres J. R. Elements of algebraic topology. CRC Press (2018).
Edelsbrunner, H., Letscher, D. & Zomorodian, A. Topological persistence and simplification. Discrete Comput. Geom. 28, 511–533 (2002).
Zomorodian, A. & Carlsson, G. Computing persistent homology. Discrete Comput. Geom. 33, 249–274 (2005).
Carlsson, G., Zomorodian, A., Collins, A. & Guibas, L. J. Persistence barcodes for shapes. International Journal of Shape Modeling 11(2), 149–187 (2005).
Chintakunta, H., Gentimis, T., Gonzalez-Diaz, R., Jimenez, M. J. & Krim, H. An entropy-based persistence barcode. Pattern Recognition 48(2), 391–401 (2015).
Merelli, E., Rucco, M., Sloot, P. & Tesei, L. Topological characterization of complex systems: Using persistent entropy. Entropy 17(10), 6872–6892 (2015).
Rucco, M., Castiglione F., Merelli E., and Pettini M. Characterisation of the idiotypic immune network through persistent entropy. In Proceedings of ECCS 2014, pages 117–128. Springer (2016).
Xia, K. L., Li, Z. M. & Mu, L. Multiscale persistent functions for biomolecular structure characterization. Bulletin of mathematical biology 80(1), 1–31 (2018).
Chandler, D. Introduction to Modern Statistical Mechanics. Oxford University Press (1987).
Ahmed, M., Fasy, B. T., and Wenk, C. Local persistent homology based distance between maps. In Proceedings of the 22nd ACM SIGSPATIAL International Conference on Advances in Geographic Information Systems, pages 43–52. ACM (2014).
Bendich, P., Cohen-Steiner, D., Edelsbrunner, H., Harer, J., and Morozov, D. Inferring local homology from sampled stratified spaces. In Foundations of Computer Science, 2007. FOCS’07. 48th Annual IEEE Symposium on, pages 536–546. IEEE (2007).
Bendich, P., Gasparovic, E., Harer, J., Izmailov, R., and Ness, L. Multi-scale local shape analysis and feature selection in machine learning applications. In Neural Networks (IJCNN), 2015 International Joint Conference on, pages 1–8. IEEE (2015).
Bendich P., Wang B., and Mukherjee S. Local homology transfer and stratification learning. In Proceedings of the twenty-third annual ACM-SIAM symposium on Discrete Algorithms, pages 1355–1370. SIAM (2012).
Fasy, B. T. and Wang, B. Exploring persistent local homology in topological data analysis. In Acoustics, Speech and Signal Processing (ICASSP), 2016 IEEE International Conference on, pages 6430–6434. IEEE (2016).
Abraham, M. J. et al. GROMACS: High performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1, 19–25 (2015).
Berendsen, H. J. C., van der Spoel, D. & van Drunen, R. GROMACS: a message-passing parallel molecular dynamics implementation. Computer physics communications 91(1-3), 43–56 (1995).
Horn, H. W. et al. Development of an improved four-site water model for biomolecular simulations: TIP4P-Ew. The Journal of chemical physics 120(20), 9665–9678 (2004).
Kast, K. M., Brickmann, J., Kast, S. M. & Berry, R. S. Binary phases of aliphatic n-oxides and water: Force field development and molecular dynamics simulation. The Journal of Physical Chemistry A 107(27), 5342–5351 (2003).
Pearlman, D. A. et al. AMBER, a package of computer programs for applying molecular mechanics, normal mode analysis, molecular dynamics and free energy calculations to simulate the structural and energetic properties of molecules. Computer Physics Communications 91(1-3), 1–41 (1995).
Hess, B., Bekker, H., Berendsen, H. J. C. & Fraaije, J. G. E. M. Lincs: a linear constraint solver for molecular simulations. Journal of computational chemistry 18(12), 1463–1472 (1997).
Essmann, U. et al. A smooth particle mesh Ewald method. The Journal of chemical physics 103(19), 8577–8593 (1995).
Ulrich B. Ripser: efficient computation of Vietoris-Rips persistence barcodes. Preprint 1908.02518, (2019).
Giusti, C., Ghrist, R. & Bassett, D. S. Two’s company, three (or more) is a simplex. Journal of computational neuroscience 41(1), 1–14 (2016).
Hiraoka, Y. et al. Hierarchical structures of amorphous solids characterized by persistent homology. Proceedings of the National Academy of Sciences 113(26), 7035–7040 (2016).
Saadatfar, M., Takeuchi, H., Robins, V., Francois, N. & Hiraoka, Y. Pore configuration landscape of granular crystallization. Nature communications 8, 15082 (2017).
Acknowledgements
This work was supported in part by Nanyang Technological University Startup Grant M4081842 and Singapore Ministry of Education Academic Research fund Tier 1 RG31/18, Tier 2 MOE2018-T2–1–033.
Author information
Authors and Affiliations
Contributions
K.X. and V. A. contributed algorithm design and wrote the main manuscript. M.Z. reviewed the manuscript and algorithms. Y.G. provided simulation data.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Anand, D.V., Meng, Z., Xia, K. et al. Weighted persistent homology for osmolyte molecular aggregation and hydrogen-bonding network analysis. Sci Rep 10, 9685 (2020). https://doi.org/10.1038/s41598-020-66710-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-66710-6
This article is cited by
-
Hybrid Topological Data Analysis and Deep Learning for Basal Cell Carcinoma Diagnosis
Journal of Imaging Informatics in Medicine (2024)
-
Persistent Dirac for molecular representation
Scientific Reports (2023)
-
Steady and ranging sets in graph persistence
Journal of Applied and Computational Topology (2023)
-
Using persistent homology as preprocessing of early warning signals for critical transition in flood
Scientific Reports (2021)
-
Higher-order structure of polymer melt described by persistent homology
Scientific Reports (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.