Abstract
Babesia microti is an intraerythrocytic parasite and the primary causative agent of human babesiosis. It is transmitted by Ixodes ticks, transfusion of blood and blood products, organ donation, and perinatally. Despite its global public health impact, limited progress has been made to identify and characterize immunodominant B. microti antigens for diagnostic and vaccine use. Using genome-wide immunoscreening, we identified 56 B. microti antigens, including some previously uncharacterized antigens. Thirty of the most immunodominant B. microti antigens were expressed as recombinant proteins in E. coli. Among these, the combined use of two novel antigens and one previously described antigen provided 96% sensitivity and 100% specificity in identifying B. microti antibody containing sera in an ELISA. Using extensive computational sequence and bioinformatics analyses and cellular localization studies, we have clarified the domain architectures, potential biological functions, and evolutionary relationships of the most immunodominant B. microti antigens. Notably, we found that the BMN-family antigens are not monophyletic as currently annotated, but rather can be categorized into two evolutionary unrelated groups of BMN proteins respectively defined by two structurally distinct classes of extracellular domains. Our studies have enhanced the repertoire of immunodominant B. microti antigens, and assigned potential biological function to these antigens, which can be evaluated to develop novel assays and candidate vaccines.
Similar content being viewed by others
Introduction
Babesia microti, an intraerythrocytic parasite belonging to the phylum Apicomplexa, is the primary causative agent of human babesiosis1. It is transmitted by Ixodes ticks but also can be transmitted by transfusion of blood, blood products, solid organ donation, and perinatal transmission2,3,4,5,6,7. Human B. microti infection occurs worldwide but is most commonly reported in the United States and is endemic in the Northeast and upper Midwest1. The number of tick-borne and transfusion-transmitted cases of babesiosis in the United States has risen dramatically in the past two decades5,8. New cases of babesiosis have been reported throughout the world, including a new Babesia endemic area in Northeast China due to Babesia venatorum9. Asymptomatic B. microti infection is not uncommon in healthy residents of B. microti-endemic regions and among blood donors. On Block Island, Rhode Island, about a fifth of healthy adults infected with B. microti are asymptomatic10. In a comprehensive study in 60,512 healthy blood donors in endemic areas in northeastern United States, 0.38% of the donors were shown to carry B. microti DNA or B. microti antibody in their bloodstream11. Asymptomatically infected individuals may transmit the infection through the blood supply if they donate blood. Death occurs in about a fifth of individuals receiving contaminated blood transfusion5. Similar fatality rates have been reported in asplenic patients and in patients suffering from cancer with or without asplenia and/or rituximab therapy. The latter group of patients often require hospital admission and experience persistent relapsing disease that may last for more than a year11,12,13. Fatality estimates for B. microti infection have ranged from <1% to 2% in the general population and 3% to 9% in hospitalized babesiosis patients14,15,16,17,18,19. Fatality rates among asplenic European patients with Babesia divergens infection have reached as high as 42% but more recently have decreased as a result of improved adjunctive therapy19,20,21.
Although the full genome sequence for B. microti was published in 201222,23, there is still a scarcity of well-characterized, immunodominant B. microti antigens for use in diagnostic assays and vaccine development. Antibody testing using the Indirect Fluorescence Antibody assay (IFA) has been used in conjunction with B. microti PCR as an effective blood donor screening approach to prevent transfusion transmitted babesiosis11,24. Earlier efforts using cDNA expression libraries have led to the identification of novel B. microti immunodominant antigens which have proven useful as diagnostic markers for infection25,26,27. The availability of full genome sequence for B. microti22,23 allows the genome-wide screening to identify novel immunodominant B. microti antigens and to perform genetic and biological characterization of these antigens. Such investigations may not only contribute to the development of superior diagnostic assays but also provide a better understanding of Babesia pathogenesis and phylogenetic relationships among different Babesia species/strains.
In this work, we generated B. microti cDNA phage display libraries to identify a battery of novel immuno-dominant B. microti antigens, searched for structural features that could predict their biological function and cellular localization, and corrected the annotation and phylogenetic classification of some previously discovered antigens. Additionally, we have evaluated the use of these antigens (BmSERA1, BmMCFRP1 and BmPiβS1) in an enzyme-linked immunosorbent B. microti antibody assay (BmELISA). Our study significantly expands the pool of well-characterized B. microti antigens as potential diagnostic biomarkers and vaccine candidates and demonstrates their utility as screening targets in a highly sensitive and specific diagnostic antibody assay.
Results
Collection of sera from Babesia microti-infected patients and healthy control study subjects
Sera were obtained from 28 babesiosis patients from Nantucket, MA, an island that is highly endemic for babesiosis. Each patient had symptoms consistent with babesiosis. B. microti infection was confirmed in patients by thin blood smear and/or B. microti PCR testing. Sera were collected one to three weeks after the onset of symptoms in 21 patients and four to nine weeks in seven patients. Negative control sera were obtained from 23 healthy study subjects who were enrolled in a serosurvey on Block Island, RI and tested negative for B. microti antibody.
Genome-wide immunoscreening of B. microti cDNA phage library to detect B. microti antigens
To conduct a genome-wide search for immunodominant B. microti antigens, a cDNA library in the phage expression system was constructed to potentially represent the complete proteome expressed during the asexual blood stage of parasites isolated from infected DBA/2 mice. The foreign protein is incorporated into the virion as gIIIp fusion, which retains infectivity and displays the foreign peptide in an immunologically accessible form. The average size of the cDNA fragments fused to the gIII gene (M13 phage gene encoding gIIIp surface protein) was 400 bp (small fragments: 50–300 bp and large fragments: 300–1000 bp). Following characterization, we obtained a library size of more than 106 fusion phages, each displaying a B. microti peptide. Given the B. microti genome size of ~ 6.5 Mbp and 70% of the genes interrupted by short introns of 20–25 bp, this library should theoretically represent 100% coverage of the entire open reading frame of B. microti. The M13 phage display library expressing the B. microti transcriptome was screened with a pool of seven sera obtained from babesiosis patients. Serum samples were obtained from patients between 1 and 9 weeks after the onset of symptoms, but in most cases one to three weeks after the onset of symptoms. Following two rounds of panning, a total of 960 clones were isolated and amplified via PCR, before being subjected to nucleotide sequencing. The gene sequences thus obtained were aligned to the B. microti genome (www.piroplasmadb.org) which led to identification of 56 B. microti antigens that were recognized by antibodies generated against these antigens. Of the 56 proteins, 11 were predicted to be secreted or cell-surface proteins. The annotations, protein domain(s) and the phyletic patterns of these antigens where relevant are presented in Table 1.
Immunodominant B. microti antigens and their genetic variability analysis
To identify the most promising diagnostic biomarkers, 30 of the 56 B. microti antigens with the highest antibody reactivity in a phage-ELISA (against pool of seven babesiosis patient sera as indicated above; data not shown) were selected for further antigenic characterization. To assess the possible use of the 30 B. microti antigens as diagnostic biomarkers, their cDNAs were cloned with a hexa-histidine tag at the N-terminus and were expressed in E. coli.
The expressed region for each identified antigen is shown in Supplementary Figure S1. The recombinantly expressed proteins were purified to homogeneity using affinity chromatography (data not shown). The availability of whole genome sequence of B. microti22 has made feasible in silico analyses of these identified immunodominant antigens. We find that the 30 most immunodominant antigens identified from the phage library immunoscreening do not show any preferential clustering and are distributed across all four chromosomes of B. microti. This indicates that the antigens identified are not preferentially drawn from the proteins encoded by the highly variable sub-telomeric regions22,23.
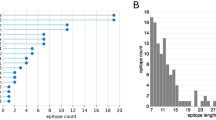
We also analyzed the genetic variability in the 30 most immunoreactive antigens by determining the non-synonymous (dN)/synonymous (dS) substitutions in relation to the recently published 41 B. microti genome sequences28. The dN/dS ratio quantifies selection pressure acting on a protein coding region. A dN/dS ratio of more than one indicates that the protein is under immune pressure and leads to changes in amino acids. In our dataset, 11 out of 30 B. microti proteins have dN/dS ratio of >1 suggesting selective pressure on immunodominant proteins for divergence (Fig. 1). More than half of the most immunoreactive proteins (18/30) have a dN/dS ratio of <1 suggesting a conserved nature of this subset of antigens. One of these 30 proteins has a dN/dS ratio of exactly one, raising the possibility that this gene might be under relaxed selection.
Heat map showing replicates of phages displaying 56 B. microti protein sequences as gIIIp fusion isolated following immunoscreening. The green color indicates the peptide domain isolated with highest frequency while other clones are represented by different intensities of red from highest to lowest frequency. The bar graph on the right shows dN/dS ratio to represent the presence of gene polymorphism as analyzed by aligning the sequences from the 41 B. microti isolates available on Piroplasmadb.org.
The sensitivity of B. microti recombinant immunodominant antigens for detection of antibody
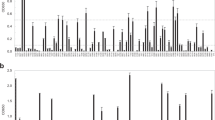
The seven sera that had been used to identify antigens by phage display were screened in an ELISA that used the 30 most immunoreactive recombinant B. microti proteins as capture antigens. Of this screening, we further selected 19 highest immunoreactive antigens for more in- depth evaluation as capture antigen in ELISA of using sera from 28 babesiosis patients and 15 negative controls (Supplementary Table S1 and Fig. 2). The sensitivity of novel immunodominant antigens in the BmELISA varied greatly and there was a clear antigenic hierarchy. No single recombinant antigen could recognize antibody in all 28 serum samples tested. On the basis of reactivity, these 19 recombinant antigens were organized as high reactivity (reactivity against >75% of the patient sera), medium reactivity (reactivity against >50% but ≤75% of the patient sera), and low reactivity (reactivity against ≤50% of the patient sera). The three most reactive B. microti antigens were antigenically characterized by protein mass spectrometry and SDS-PAGE (see Supplementary text, Supplementary Figure S2 and S3).
Reactivity of recombinant B. microti proteins against sera from B. microti-infected patients in BmELISA. Nineteen recombinant B. microti proteins were tested against 28 human sera positive for B. microti antibodies. Signal/cutoff is calculated by dividing the individual absorbance (A450nm) with the mean optical density reading for the human serum samples negative for antibodies to B. microti (n = 15) + 2 standard deviations of the mean. Circles represent the reactivity of each patient serum against individual recombinant protein. Immunoreactivity was classified as high, medium and low based on having the highest percentage of positives over the signal/cut-off value above 1.
Babesia microti antibody ELISA (BmELISA) using three immunodominant proteins
We further determined the sensitivity of the three immunoreactive B. microti antigens – BmSERA1, BmMCFRP1 and BmPiβS1 individually and in a combination of three antigens in a BmELISA tested against serum samples from 28 babesiosis patients and 23 normal human serum samples. In addition, we also compared the sensitivity of these three antigens to a previously described immunodominant antigen – BmBAHCS126,29. The reactivity of individual antigens in BmELISA is as follows: BmSERA1 (24/28: 86%), BmMCFRP1 (23/28: 82%), BmPiβS1 (22/28: 79%), and BmBAHCS1 (27/28: 96%). A combination of BmSERA1, BmMCFRP1 and BmPiβS1 gave a sensitivity of 96% (27/28). When BmBAHCS1 was included, the 4 antigen combination was able to recognize all 28 (100%) of the babesiosis patient samples (Table 2), thus markedly improving overall sensitivity compared to the single-antigen ELISA formats. All of the 23/23 (100%) serum samples from healthy individuals were negative in both single- or multi-antigen BmELISA format indicating a high level of specificity.
Bioinformatics analyses, annotation, and biological function of immunodominant antigens
Sequence analysis
We next performed a detailed sequence analysis of all 56 immunoreactive proteins identified in our study. Eleven of the 56 proteins are predicted to be cell surface or secreted proteins (Table 1). Of the 56 proteins, the seven proteins with highest immunoreactivity which is defined as having the highest percentage of positives over the signal/cut-off value above 1 (Fig. 2), six are predicted to be secreted based on sequence analysis. This suggests that our screen was successful in identifying proteins that are most likely to provoke an antibody response by virtue of being extracellularly exposed to the host immune cells. Of these 11 secreted or surface proteins five are members of protein families that have been reported as antigenic in other strains of B. microti or other Babesia species. Through a comprehensive sequence searches against the complete panel of publicly available eukaryotic genome sequences in the Genbank database we observed that a total of 11 of the immunoreactive proteins recovered in our screen showed a phyletic pattern restricted uniquely to piroplasmida or more broadly to apicomplexa and related alveolates. These include some of the proteins with highest immunoreactivity suggesting that they are likely to serve as specific diagnostic markers with no risk of cross reactivity with conserved proteins from other organisms. These findings do not discount the presence of intracellular B. microti antigens that are capable of eliciting an effective protective response when presented to host immune cells.
Annotation of the most immunodominant proteins
Some of the most immunodominant proteins identified in the B. microti Peabody strain were cell surface proteins (BmR1_03g00690 [BmEGF1], BmR1_02g04285 [BmMCFRP1], BmR1_04g08155 [BmSERA1], BmR1_03g04855 [BmPiβS1], BmR1_03g00785 [BmBAHCS1], and BmR1_04g06300) except BmR1_01g03455 which was found to be an AP2-superfamily transcription factor typical of apicomplexa. Five of these proteins namely, BmR1_03g00690, BmR1_02g04285, BmR1_04g08155, BmR1_03g04855, and BmR1_03g00785 were subjected to an in-depth sequence analysis to obtain a better understanding of their evolutionary history and potential functional features. BmR1_03g00690 was found to contain four copies of the epidermal growth factor (EGF) domains (Fig. 3). Accordingly, we named it BmEGF1. EGF domains have been found in wide range of cytoadherence proteins from a variety of microbes in apicomplexa, such as MSP1, MSP4, MSP5, MSP8 and MSP10, which are merozoite surface proteins of Plasmodium falciparum30. The presence of EGF domains in BmR1_03g00690 was not previously known and this protein is unrelated to any EGF domain proteins from other Apicomplexa parasites, including other piroplasms. The EGF domains in this protein show certain features related to the pattern of conserved cysteines that are shared with specific EGF domains found in certain Giardia intestinalis and animal secreted proteins (Fig. 3). This suggests that this EGF domain protein is a unique surface protein of B. microti, which might have been acquired relatively recently, either from an animal source or from some other parasitic eukaryote.
Multiple Alignment of (A) BAHCS family (B) PiβS family (C) EGF domain containing BmR1_03g00690 family. Representative multiple sequence alignments of the families are shown with a 95%, 90% and 90% consensus, respectively. The sequences are denoted by their gene names or gene IDs, species names, and accession numbers. Proteins with repeats are shown with underscore followed by the repeat number. The amino acid range is shown at the beginning and end of the sequence. The numbers within the alignment represent poorly conserved inserts of the given number of amino acid residues that are not shown. The predicted secondary structure or the crystal structure are shown with orange cylinders representing helices and green arrows representing beta sheets. The coloring is based on the consensus of the whole family alignment: “h” is for hydrophobic residues (ACFILMVWY [yellow]); “l” represents the aliphatic subset of the hydrophobic class [ILV (yellow]); “s” represents small residues (ACDGNPSTV (green)); “u” represents the tiny subclass of small residues (GAS [green]); “p” represents polar residues (CDEHKNQRST [blue]); “c” represents the charged subclass of polar residues (DEHKR [pink]); “—“ represents the negative subclass of charged residues (DE); “o” represents alcoholic residues (ST[orange]); and “b” represents big residues (KFILMQRWYE [gray]). Any absolutely conserved residue is labeled and shaded red.
The gene of a second immunodominant protein, BmR1_02g04285 (BmMCFRP1), is currently improperly predicted, where a 5′ exon containing the signal peptide has been omitted (Genbank ID: XP_021338378.1). This protein contains a conserved domain that is also found in two copies in the paralogous protein BMR1_04g07535 from B. microti. These two proteins in turn are homologous of the Maltese cross form-related antigen (N1–15/BMN1–15 protein; Genbank ID: BAC07532.1) that was identified in two distinct isolates (the B. microti Munich and MN1 strains; Table 3). They are thought to provide protection against B. microti infection in mice, although immunization with a control baculovirus with no B. microti antigen also induced some level of immunity31,32. Interestingly, there is extensive sequence divergence between the published Maltese cross form-related antigens from the Munich and MN1 strains and the cognate in the deposited B. microti R1 genome, suggesting that this protein and its paralogs might be polymorphic in B. microti.
A third immunodominant B. microti cell surface protein BmR1_04g08155 (BmSERA1) is a 946-amino acid protein, which was erroneously annotated as having “homologies with serine-repeat antigen 4”. This annotation is unsupported by sequence analysis and arises from improper masking of low complexity sequence. This protein has a previously described homolog in the Munich strain of B. microti named BmP94 and was reported to have antigenic properties consistent with those identified in the current study33. Remarkably, comparison of BmR1_04g08155 with this homolog protein suggests that BmR1_04g08155 is extremely fast-evolving even between these two strains, with a sequence identity of about 43%. This is much higher than the sequence divergence for other available proteins of these two strains (~95–98% identity). This divergence could be due to selection exerted by differences in the erythrocytes of potentially distinct reservoir hosts of the two strains. Alternatively, these findings might indicate that the divergence is an evolutionary response to variable immune pressure of different hosts, consistent with its character as an antigenic secreted/cell surface protein.
Annotation of two highly immunodominant proteins of BMN group
The other two highly immunodominant proteins, BmR1_03g04855 and BmR1_03g00785 are members of the so called “BMN” class of antigenic proteins, which are shared by different B. microti strains and B. rodhaini. Some of these antigens (SA5–1–1, SA26 and SA17) were first identified in B. rodhaini in 198834 and several subsequent studies in B. microti have identified them as antigenic proteins26,31. These studies have not fully defined the evolutionary relationships of these proteins, however, resulting in incomplete understanding of their nomenclature as reported in the literature26,31,35,36. Our analysis of these proteins helps to further clarify their evolutionary relationships (BmR1_03g04855 and BmR1_03g00785). Our sequence analysis shows that the proteins which have been considered BMN antigens do not constitute a monophyletic group and should not be classified as belonging to a unified “BMN family” because all members currently annotated as BMN proteins are not monophyletic. Instead our analysis shows that the majority can be classified into two largely evolutionarily unrelated groups of BMN proteins. The first of these groups includes the previously characterized BMN1–10, N1–10, BMN1–4, BMN1–3B, BMN1–8, and BMN1–11 proteins from the B. microti MN1 strain, the IRA protein from the B. microti Gray strain and the Br-1 and Br-2 proteins from the B. rodhaini Japan strain. The second major group is comprised of BMN1–2, BMN1–3, BMN1–6, BMN1–7, BMN1–9, BMN1–13, BMN1–4, MN-10, N1–21 from the B. microti MN1 strain, BmSA1 from the B. microti Gray strain, BmP32 from the B. microti Munich strain, MSA1 and MSA2 from B. rodhaini Australia strain and Br-1, p25, and p26 from the B. rodhaini Japan strain. The equivalent versions of these two groups from the B. microti R1 strain are provided in Table 3. Beyond these, the proteins BMN1–17 and BMN1–20 are paralogous proteins that are unrelated to any of the above groups. Likewise, the Maltese cross form-related antigen (BMN1–15/N1–15) and BmR1_02g04285 (BmMCFRP1) define a distinct family unrelated to the other BMN1 antigens (see above). Therefore, we recommend that the BMN antigens be treated as distinct groups as per their conserved domains and evolutionary relationships described here (Fig. 4, Table 3).
Phylogenetic tree of the (A) BAHCS family (B) BmPiβS1, and other families of proteins. Strain specific clades are shown with colored branches and boxes. B. microti MN1 strain is colored blue, the B. microti R1 strain is colored red, B. microti gray strain is colored green, and the B. rodhaini japan strain is colored light blue. The proteins are denoted by their gene names or gene ids, species names, and accession numbers.
Our analysis shows that BmR1_03g04855 from the B. microti R1 strain belongs to the first of the major BMN groups (i.e. BMN1–10 and its relatives). B. microti R1 has a total of 10 members of this group (Table 3). These correspond in large part to the B. microti proteins termed BMN1 in the work by Silva et al.37. Analysis of these proteins shows that they are characterized by the presence of a conserved domain which might be present in one to five copies per protein, with a single copy in BmR1_03g04855. Secondary-structure-prediction based on the alignment of this domain showed that it contains an N-terminal region with 8 conserved β-strands followed by a C-terminal region with multiple cysteines. The N-terminal region is likely to adopt a β-sandwich fold, whereas the C-terminal region is likely to adopt a disulfide bond-supported structure. Interestingly, iterative sequence profile analysis identified proteins with a divergent version of this domain outside of Babesia in a group of secreted proteins in Theileria. While this family is expanded across Theileria, it is particularly abundant in the horse-parasitic species T. equi (~460 members); however, it is present in fewer numbers in T. annulata, T. orientalis, and T. parva. Because it is present in both the piroplasms, we accordingly named this domain the piroplasm β-strand (PiβS) domain (Fig. 3, Supplementary Fig. S4). Accordingly, we named BmR1_03g04855 as BmPiβS1. Given that the PiβS family is inferred to have been ancestrally present in the piroplasms, it is likely that it has played a key role in host-parasite dynamics of the entire piroplasm lineage. Importantly, our phylogenetic analysis of the PiβS domain in the genus Babesia shows that its evolution is dominated by lineage-specific expansions (Fig. 4). Notably, the versions in B. rodhaini appear to have radiated entirely independently of those from B. microti. Moreover, even within B. microti we found clades exclusively or predominantly containing R1 strain or MN1 strain proteins (Fig. 4).
Our analysis shows that BmR1_03g00785 from our screen belongs to the second major BMN group (prototyped by BMN1–2, BMN1–9 etc.). These correspond in large part to the B. microti proteins termed BMN2 in the work by Silva et al.37. We find that this group is characterized by a distinctive α-helical domain with 9 α-helices. Further, these proteins terminate at the C-terminus in a hydrophobic GPI-anchor sequence. This suggests that all these proteins are anchored to the cell membrane with α-helical domain lying proximal to the membrane. This domain is thus far found only in the genus Babesia. Nine proteins containing this domain are present in the B. microti R1 strain. Accordingly, we named this protein as Babesia α-helical cell surface (BAHCS) domain. Unlike the PiβS domain, the BAHCS domain is always found in a single copy in a protein. Thus, we named BmR1_03g00785 as BmBAHCS1. In a single protein, Br-1 from B. rodhaini and its cognate BMN1–4 from the B. microti MN1 strain, it is combined with an N-terminal PiβS domain in the same protein. Notably, phylogenetic analysis of the BAHCS domain shows a similar evolutionary trend as the PiβS domain, with independent expansions in B. rodhaini and B. microti followed by notable inter-strain radiations in the later species (Fig. 4).
In summary, our analyses suggest that the immunodominant cell surface antigens that we identified have been evolving rapidly in phylogenetically closely related organisms through independent lineage-specific expansions. Such a pattern is the hallmark of an arms race with the host and has been observed before in the case of other apicomplexan surface proteins such as the rifin-like38 and the var/DBL1 superfamilies in Plasmodium falciparum, and the vir/yir superfamilies in P. vivax/P. yoelii30. This suggests that the PiβS and BAHCS domain families are likely to be expressed on the cell-surface at the interface with host immune cells. The dynamic evolution suggests that the lineage-specific expansions are a positively selected response against the host immunity targeting them.
Localization of three immunodominant proteins (BmSERA1, BmMCFRP1, and BmPiβS1) using an immunofluorescence antibody assay (IFA)
To assign a biological function to the three most immunodominant B. microti proteins; we investigated the localization of BmSERA1, BmMCFRP1 and BmPiβS1 proteins on intraerythrocytic B. microti parasites. We used immunofluorescence microscopy and polyclonal antibodies generated in mice against the respective recombinant proteins. Anti-BmSERA1 antibody showed expression of BmSERA1 on the surface of the late stage merozoite which is a tetrad structure (Fig. 5A), suggesting a possible role of this protein in invasion of B. microti parasites into fresh RBCs after their release from infected RBCs. The distribution of BmMCFRP1 when captured using anti-BmMCFRP1 polyclonal antibody, indicate expression on a free merozoite and on a parasite that is in the process of invading an RBC. Interestingly, we also noted a distinct scattered and punctate pattern of expression on the surface of infected RBCs. At the same time these speckled patterns are also detected in the adjacent uninfected RBCs (Fig. 5B), suggesting a possible dual function as a membrane bound and as a secreted protein that binds uninfected RBCs. The immunofluorescence signal obtained with anti-BmPiβS1 antibody shows high level expression on the surface of infected RBCs harboring tetrad form parasites and as a diffuse punctate pattern which appears to be inside the RBC cytoplasm (Fig. 5A), suggesting a role as a transported protein which is anchored in the infected RBC membrane. No non-specific reactivity was observed using antibodies against the CFA/IFA adjuvant alone formulation.
(A) The fluorescent image shows localization of BmSERA1, BmMCFRP1 and BmPiβS1 in B. microti infected mouse red blood cells (see results). Green channel (Alexa 488) images correspond to the location of B. microti parasite proteins. Red channel images are cell membrane stained by PKH26 and Blue channel shows DAPI stained nuclei. Merged images contain the overlay of green, red and blue channels. As observed, sera from mice immunized with Freund’s adjuvant only (negative control) shows DAPI stained nuclei demonstrating B. microti infected RBCs without a traceable green signal, confirming specificity of the assay. Sera collected from mice immunized with BmSERA1, BmMCFRP1 and BmPiβS1 in Freund’s Complete Adjuvant show specific localization of the B. microti proteins as membrane bound on the parasite surface (anti-BmSERA1) or on the surface of infected RBCs (anti-BmMCFRP1 and anti-BmPiβS1). Scale bar indicates 1 µm. (B) I. Free B. microti merozoite located near the uninfected RBC expressed BmMCFRP1 (arrow, green). Note that plasma membrane of B. microti and RBC were stained by PKH26 dye (red). Counterstaining by DAPI indicates B. microti nuclei (blue). II. The B. microti merozoite found inside RBCs showed negative staining for anti-BmMCFRP1. The apical region (entry site) of RBCs showed strong positive staining for BmMCFRP1 (arrow, green). In addition, the surface of infected and surrounding uninfected RBCs showed punctate pattern of BmMCFRP1 (arrowheads). Scale bar indicates 2 μm.
Genetic diversity of B. microti immunodominant antigens (BmSERA1, BmMCFRP1, and BmPiβS1) using sequence analysis
We sought an in-depth study of the distribution and natural dynamics of the polymorphisms in the three highly reactive B. microti immunodominant antigens – BmSERA1, BmPiβS1 and BmMCFRP1- in endemic populations where diversity is driven by naturally acquired immunity and other biological and ecological factors. To accomplish this, we aligned the nucleotide sequences coding for the respective antigens with the sequences from the 41 B. microti isolates available in the Piroplasmadb.org. The full length BmSERA1, BmPiβS1 and BmMCFRP1 have a total of 72 (2841 nucleotides), 11 (816 nucleotides) and 20 (531 nucleotides) non-synonymous substitutions showing 2.5%, 1.3% and 3.7% chance of substitution, respectively. The nucleotide variation reported here as non-synonymous substitutions are mostly derived from the B. microti Russia-1995 strain, which is more variable at the genomic level than the strains from the continental United States28. When analysis for nucleotide polymorphism was limited to the continental United States isolates, the extent of non-synonymous substitutions was reduced to 23 (2841 nucleotides), 0 (816 nucleotides) and 0 (531 nucleotides) showing 0.8%, 0% and 0% chance of substitution, respectively for BmSERA1, BmPiβS1 and BmMCFRP1. We separately analyzed the amino acid sequences in BmSERA1 from the four United States isolates, which showed a minimal antigenic variation (Two from Minnesota isolates: MN-1: 1.7% and MNB010: 2.5%; one from Wisconsin: W17: 1.1%; and one from New England/North Dakota: ND-11: 0%) suggesting the diagnostic value of this molecule. Overall, our data indicate a very limited or absent genetic polymorphism among immunodominant B. microti antigens in the United States.
Discussion
We utilized a phage display library encoding fragments of the B. microti transcriptome to identify antigenic proteins that were recognized by antibodies generated after natural infection in humans. We have identified 56 B. microti antigens using two cycles of affinity panning, that exhibited strong reactivity with anti-B. microti antibodies in patients living in a highly endemic area in the northeastern United States. We produced 30 recombinantly expressed immunoreactive B. microti antigens and, based on immunodominance, 19 of these were extensively screened in ELISA. Three of the most immunodominant of these antigens - BmSERA1, BmMCFRP1 and BmPiβS1 were selected for sensitivity and specificity studies in Bm-ELISA. They exhibited 86%, 82% and 79% reactivity, respectively when probed against sera from babesiosis patients. A combination of 3 antigens gave an improved sensitivity of 96% (27/28) (Table 2). Notably, in earlier studies, the BmBAHCS1 (BmR1_03g00785; named BMN1–9 by Lodes et al.26, BmSA1 by Luo et al.29, and BmGPI12 by Cornillot et al.36) allowed the detection of 100% of babesiosis serum samples tested, demonstrating the diagnostic value of this antigen. Our results clearly indicate that the discovery of these novel antigens has enhanced the repertoire of immunodominant B. microti molecules that could be explored to develop highly sensitive and specific B. microti antibody and antigen capture assays for diagnosis of acute babesiosis, blood donor screening, and support of vaccine development. Availability of these antigens also paves the way to develop multiantigen-based B. microti detection assays that would offset the potential sensitivity loss due to geographical antigenic polymorphism and antigenic drift over time.
Analyses of the 30 most highly immunoreactive B. microti antigens revealed that the genes encoding these proteins are distributed throughout the four B. microti chromosomes, suggesting a lack of clustering of genes coding for immuno-dominant antigens to a specific genomic locus. Of the 30 antigens, 11 may be subjected to immune pressure, defined by the dN/dS ratio >1 (Fig. 1). Furthermore, minimal or no genetic diversity was observed in the three most immunodominant antigens - BmSERA1, BmMCFRP1 and BmPiβS1 when sequence analyses were performed with 38 B. microti isolates from the continental United States28. These results indicate a high degree of genetic stability in B. microti strains circulating in the United States. They are consistent with an earlier report that B. microti strains in the continental United States represent minimal diversity as a geographical group, but largely differ from the published Russian strain28. The possible reasons for limited genetic diversity among B. microti isolates may include limited host specificity in invasion and survival in red blood cells and a lack of recombination events during the sexual cycle in tick-vectors.
Further sequence and bioinformatics analyses of the five most immunodominant B. microti antigens - BmEGF1, BmMCFRP1, BmSERA1, BmPiβS1 and BmBAHCS1 has better revealed their domain structure, evolutionary relationships, and putative biological functions. Our sequence analysis led to a correction in annotaion of the BmSERA1 gene which was erroneously classified as homologous with serine-repeat antigen 4. A notable contribution of this study is the finding that the major group of the BMN family antigens are not monophyletic as currently annotated, but rather can be classified into two evolutionary unrelated groups of BMN proteins (see the results section). Based on our analyses, we strongly recommend that henceforth the BMN antigens be treated as distinct groups per their conserved domains and evolutionary relationships (Fig. 4, Table 3).
The localization studies by immunofluorescence microscopy further confirmed the findings of the bioinformatics analyses. The localization study revealed that BmSERA1 and BmMCFRP1 are surface exposed and secreted antigens, respectively, making them a potential immune target (Table 1). Further, BmSERA1 is distributed in a pattern that is similar to that observed in the merozoite surface protein (MSP) family in malaria parasites, Plasmodium falciparum39 and Plasmodium vivax40. MSPs in Plasmodium parasite have been shown to be required for the initial attachment of merozoites to RBCs for invasion. It is therefore possible that BmSERA1 may play a role in the invasion process. Given its expression pattern, BmMCFRP1 may be associated with trafficking of proteins to the surface or involved in organelle or membrane formation, as seen in Plasmodium and other apicomplexan parasites. In P. falciparum, binding of erythrocyte-binding antigen 175 (EBA-175) has been shown to prime the erythrocyte surface by altering the biophysical nature of target cell and, thereby reduces the energy barrier to invasion41. Thus, it is feasible that binding of BmMCFRP1 to an uninfected RBC functions in a similar fashion as P. falciparum EBA-175 that may serve as a gateway to invasion and new infection. Interestingly, the bright continuous fluorescent signal of BmPiβS1 is comparable to that of Plasmodium proteins, including knob-associated histidine-rich protein (KAHRP), ring-infected erythrocyte surface antigen (RESA), and the subtelomeric variant open reading frame protein (STEVOR)42,43,44,45.
In summary, our unbiased genome-wide screening for immuno-dominant B. microti antigens has led to the discovery of 56 novel antigens that are attractive diagnostic and vaccine targets. These antigens are available for evaluation to develop higher sensitivity antigen/antibody based detection assays and as vaccine candidates in experimental models of B. microti. Finally, our bioinformatics analyses and immunofluorescence studies allowed us to better define the structural features and evlolutionary relationships and a finer classification of the B. microti BMN family antigens and assign potential biological functions to the most immunodominant B. microti antigens.
Materials and methods
B. microti parasites and construction of B. microti cDNA phage library
B. microti (Franca) Reichenow Peabody strain46 was obtained from the American Type Culture Collection (ATCC, VA). Female DBA/2NCr mice were injected with B. microti parasites and the parasite-infected red blood cells (RBCs) were isolated at 15–20% parasitemia. All mouse experiments were performed at White Oak animal care facility in compliance to the guidelines of White Oak Consolidated Animal Care and Use Committee (WOC ACUC). The animal study protocol used for the mouse experiments was approved by the WOC ACUC (protocol # 2009–03).
B. microti parasites were harvested by lysing the infected RBCs with Sarkosyl buffer (10 mM Tris-HCl [pH 7.5], 10 mM EDTA, 10 mM NaCl, 0.5% N-lauroylsarcosine sodium salt [Sigma-Aldrich, MO]). B. microti RNA was prepared using TriZol reagent (Life Technologies, NY), followed by chloroform extraction and precipitation that was performed with isopropyl alcohol and ethanol. Complimentary DNA (cDNA) encompassing the open reading frames was prepared from the B. microti RNA using the SMART® cDNA library construction kit (Clontech laboratories Inc., CA) following manufacturer’s instructions. Briefly, the first strand cDNA was synthesized from the B. microti RNA using a modified oligo (dT) primer and SMART IV oligo primers. The second-strand cDNA was made using Long Distance (LD) PCR conditions. The double stranded cDNA (~10 µg) product was subjected to controlled fragmentation using sonication (Sonic Dismembrator; Thermo Fisher Scientific, MA) to generate small (50–300 bp) and large (300–1000 bp) cDNA library fragments which were separated by agarose gel electrophoresis. These cDNA fragments were dephosphorylated and polished to obtain blunt ended fragments to be ligated into Sma I (CCC^GGG) digested M13-derived phage vector. The ligation products were transformed into Escherichia coli TG1 cells (Agilent technologies, MD) and selected for recombinants (tetr) on tetracycline plates. Transformed cells were cultured at 37 °C with shaking at 250 rpm in 100 ml of 2XYT broth containing 5 µg/ml tetracycline for ~16 h. The recombinant lysogenic phages displaying fusion protein domains were recovered from the supernatant and the phage titer was determined. The cDNA inserts were expressed as NH2-terminal fusion to the gIIIp surface protein of the M13 phage. Both small (50–300 bp) and large fragment (300–1000 bp) B. microti libraries yielded 106 independent clones, as established by limiting dilution of the transformed bacterial cells. Forty-eight clones were randomly picked from each library; PCR amplified using phage specific primers; and sequenced to determine the random distribution and diversity of the B. microti genome libraries.
Genome-wide immunoscreening of B. microti cDNA phage library to detect B. microti antigens
A pool of seven babesiosis patient sera (anti-B. microti IFA titer >1:500) collected from babesiosis patients from Nantucket, MA were used for panning of the B. microti libraries. To minimize non-specific reactivity, pooled babesiosis sera were incubated on ultraviolet-killed M13K07 phage-coated petri dishes. For the affinity panning of the phage library, 96-well microwell plates (Immulon 4 HBX; Thermo Fisher Scientific, MA) were coated overnight at 4 °C with 1 µg per well of goat anti-human IgG Fcγ antibodies in phosphate-buffered saline (PBS; 10 mM Na2HPO4, 1.8 mM KH2PO4, 2.7 mM KCl, 137 mM NaCl, pH 7.4). After three washings with PBST (PBS [pH 7.4] containing 0.1% Tween 20 [Sigma-Aldrich, MO]), 5% bovine serum albumin (BSA fraction V, Sigma-Aldrich, MO) in PBST was added to the wells to block the unoccupied reactive sites. The preadsorbed babesiosis patient sera from the earlier step was added to the wells and incubated for 1 h at room temperature (RT). Wells were washed three times with PBST, and 108 phages from the B. microti library were added for 1 h at RT. Non-adherent phages were removed by nine washes with PBST followed by three washes with PBS. The adherent phages were eluted by the addition of 0.1 N Glycine.HCl (pH 2.2), 100 µl per well for 10 min at 37 °C. The eluate was immediately neutralized by the addition of 2 M Tris (pH unadjusted). The eluate was simultaneously titrated and amplified for the next round of panning in log phase (OD600nm ~0.8) E. coli TG1 cells. For the phage amplification, the phage infected TG1 cells were incubated at 37 °C for 90 min without shaking followed by dilution with 10 ml of 2XYT medium containing 5 µg/ml tetracycline and incubated at 37 °C, with shaking at 250 rpm for ~16 h. Phage supernatants were collected after centrifugation and one more round of panning was carried out. Phage titration plates were used for harvesting/selecting the colonies and performing PCR amplification with subsequent sequencing to establish the identity of the cloned insert. A total of 960 phage clones were sequenced using phage specific primers. The sequences obtained after Sanger’s di-deoxy sequencing (performed at the FDA center facility) were analyzed by the NCBI BLAST to identify the B. microti protein that each encodes. Finally, the sequencing reads were aligned to the target sequence in the MacVector program.
Phage ELISA to select B. microti antigens with highest antibody affinity
The reactivity of affinity-selected phage supernatants with babesiosis patient sera was measured by ELISA. The wells of 96-well microwell plates (Immulon 4 HBX, Thermo Scientific, MA) were coated with 50 ng/well of anti-M13 phage antibody (GE Healthcare, Piscataway, NJ) and blocked with 5% skim-milk (Bio Rad, CA) PBST (0.5% Tween-20). Subsequently, phage supernatants of the selected clones were added to each well and incubated for 1 h at room temperature. Next, serially diluted sera (in 5% skim-milk PBST) were added and incubated at room temperature for 1 h. The bound antibodies were probed with horseradish peroxidase-conjugated goat anti-human IgG antibodies (Jackson Immunoresearch Laboratories, PA), and the enzymatic activity was revealed by incubating the plates with chromogenic substrate, ABTS (KPL, Inc., MD). The genes encoding the domains with high ELISA reactive phage clones were selected for cloning into an E. coli expression system.
Recombinant expression and purification of B. microti antigens and generation of antibodies against recombinant B. microti antigens
Expression of recombinant protein was accomplished by amplifying either the full-length gene or a domain of it as predicted by the theoretical antigenicity index using Immune Epitope Database and Analysis Resource (IEDB, funded by National Institute of Allergy and Infectious Diseases, NIH). The putative signal and transmembrane sequences were identified using SignalP 4.1 Server and TMHMM Server v. 2.0, respectively, and excised from the constructs encompassing the protein regions selected for the recombinant expression. The PCR-amplified product was cloned into NotI and AscI (NEB, MA) restriction site in a pET11a vector (MERCK, Germany) which was modified to include a NH2-terminal hexa-histidine tag to facilitate purification. The protein expression was carried out in E. coli BL-21 (λDE3) cells with the IPTG induction. Induced E. coli cells were lysed with the BugBuster Protein Extraction Reagent (EMD Millipore, MA) and the soluble proteins were purified from the supernatant on HisTrap column (GE Healthcare life sciences, PA). The insoluble proteins were purified by lysing the cell pellet with a combination of lysozyme and sonication, followed by buffer (50 mM Tris-HCl [pH 8.0)], 20 mM EDTA) washing 4–6 times to obtain pure inclusion bodies (IBs). The insoluble protein in the IBs was denatured in the solubilization buffer (100 mM Tris-HCl [pH 8.0], 2 mM EDTA, 6 M Guanidine HCl) before refolding under controlled redox condition in the renaturation buffer (100 mM Tris-HCl (pH 8.0), 2 mM EDTA, 500 mM L-Arginine HCl, 0.9 mM oxidized Glutathione). The refolded protein was dialyzed against a gradient of urea and finally brought into 20 mM Tris-HCl (pH 8.0) buffer and purified on HisTrap column. The purified recombinant proteins were quantified using Bradford’s reagent (Sigma-Aldrich, MO). The degree of purity of recombinant proteins was determined on SDS-PAG stained with Simply Blue Safestain. Mass-Spectrometry analysis of the purified recombinant B. microti proteins was performed to confirm their identity.
Female Balb/c mice (5–6 weeks old) were purchased from Jackson Laboratories (Bar Harbor, MA) and were housed at Center for Biologics Evaluation and Research (CBER) animal care facility in compliance with the guidelines of CBER Animal Care and Use Committee. Mice (5 per group) were immunized three times with 50 µg of purified recombinant BmSERA1, BmMCFRP1 and BmPiβS1 per mouse subcutaneously in Freund’s adjuvant (Complete Freund’s adjuvant (Sigma-Aldrich, MO) for the primary dose. The initial dose was followed by two booster doses in Incomplete Freund’s adjuvant) at 3-week intervals. Serum samples were collected two weeks after the third immunization and stored at −20 °C until use.
ELISA
Individual recombinant B. microti antigens - BmSERA1, BmMCFRP1 and BmPiβS1 and a combination of these antigens were coated overnight (~16 h) on a 96-well Immulon 4HBX ELISA plate in PBS at 50 ng/well. Plates were washed with PBST (PBS with 0.1% Tween-20) and blocked with blocking buffer (5% skim milk PBS with 0.5% Tween-20) for 2 h at 37 °C. This was followed by washing with PBST. One hundred-fold diluted sera in blocking buffer were added to the wells and plates were incubated for 1 h at 37 °C, followed by PBST washing and then incubating with 1/10,000 diluted HRP conjugated goat anti-human IgG and IgM antibody (Jackson Immunoresearch Laboratories, PA) for an additional 1 h at 37 °C. Plates were washed six times with PBST and three times with PBS and then incubated with 50 µl per well of SureBlue Reserve TMB (KPL Inc., MD) substrate solution for an additional 10 min at room temperature. The reaction was stopped using 50 µl per well of stop solution (0.16 M H2SO4, Thermo Fisher Scientific, MA). The plates were read at 450 nm using a plate reader (SpectraMax384, Molecular devices, CA). The ELISA cutoff value was determined as follows: The mean+2 Standard Deviation (S.D.) of the negative human samples were used to determine the Cutoff Ratio. The Signal to Cutoff Ratio was then calculated by dividing the Absorbance450 (A450) of each test sample by the Cutoff Ratio. A test sample was considered as positive if the A450 > 1.
Localization of three immunodominant proteins using an immunofluorescence antibody assay (IFA)
The cellular localization of native BmSERA1, BmMCFRP1, and BmPiβS1 proteins on B. microti parasites was determined by IFA. The B. microti infected RBCs grown in DBA/2 mice were washed and the RBC membrane was stained with PKH26 red fluorescent cell membrane labeling kit (Sigma-Aldrich, MO) as per manufacturer’s instructions. The stained blood cells were fixed with 3% paraformaldehyde-10 mM PIPES [piperazine-N,N’-bis (2-ethanesulfonic acid)] buffer (pH 6.4)-PBS for 30 min at room temperature, smeared on poly-L-lysine-coated glass slides, permeabilized for 10 min at room temperature with 0.25% Triton X-100-PBS, washed three times in PBS, and blocked in 5% normal goat serum (NGS)-PBS for 1 h at room temperature47. The slides were incubated with anti-BmSERA1, anti-BmMCFRP1 and anti-BmPiβS1 mouse serum diluted 1:100 in 2% NGS-PBS for 1 h at room temperature, washed three times in PBS, and the intra-erythrocytic parasites were counterstained for 1 h at room temperature with goat anti-mouse IgG (H + L) conjugated to Alexa Fluor 488 (Thermo Fisher Scientific, MA). The nucleus was stained with 4’, 6-diamidino-2-phenylindole (DAPI; Sigma-Aldrich, MO). The slides were washed three times with PBS in the dark, briefly swirled in distilled water, and mounted with fluoromount-G slide mounting medium (Electron microscopy sciences, VWR, PA) and sealed with a coverslip. The slides were observed with Leica TCS_SP8 DMI6000 confocal microscope system (Leica Microsystems, Germany) with a 100 × 1.4-numerical-objective (NA) oil objective lens. Images were acquired at a resolution of 1,024 by 1,024 pixels and stored in tif format for further analysis. Huygens (Scientific Volume Imaging, Hilversum, Netherlands) and Imaris (Bitplane AG, Zurich, Switzerland) software was used for deconvolution and image analysis48.
Genetic diversity of B. microti immune-dominant antigens using sequence analysis
The presence of single nucleotide polymorphisms (SNPs) in BmSERA1 (Gene ID: BmR1_04g08155), BmPiβS1 (Gene ID: BmR1_03g04855) and BmMCFRP1 (Gene ID: BmR1_02g04285) was studied by aligning the nucleotide sequences coding for the respective antigens with the 41 B. microti isolates available on Piroplasmadb.org.
Protein sequence searches were performed using the iterative PSI-BLAST program against the NCBI non-redundant (NR) protein database. Multiple sequence alignments were then constructed for the individual domains using Kalign2 or Muscle, followed by manual adjustments based on profile-profile comparisons, secondary structure and structural alignments. Similarity-based clustering for both classification and culling of nearly identical sequences was performed using the BLASTCLUST program (ftp://ftp.ncbi.nih.gov/blast/documents/blastclust.html). Signal peptides and transmembrane helices were predicted using SignalP 3.0 and TMHMM v1.0 programs, respectively. Phylogenetic trees for individual families were constructed using multiple alignments of the respective domains with the FastTree program.
Data availability
All data generated or analyzed during this study are included in this published article (and its Supplementary Information files).
References
Vannier, E. & Krause, P. J. Human babesiosis. N Engl J Med 366, 2397–2407, https://doi.org/10.1056/NEJMra1202018 (2012).
Spielman, A. Human babesiosis on Nantucket Island: transmission by nymphal Ixodes ticks. Am J Trop Med Hyg 25, 784–787 (1976).
Mintz, E. D., Anderson, J. F., Cable, R. G. & Hadler, J. L. Transfusion-transmitted babesiosis: a case report from a new endemic area. Transfusion 31, 365–368 (1991).
Perdrizet, G. A. et al. Babesiosis in a renal transplant recipient acquired through blood transfusion. Transplantation 70, 205–208 (2000).
Herwaldt, B. L. et al. Transfusion-associated babesiosis in the United States: a description of cases. Ann Intern Med 155, 509–519, https://doi.org/10.7326/0003-4819-155-8-201110180-00362 (2011).
Joseph, J. T. et al. Vertical transmission of Babesia microti, United States. Emerg Infect Dis 18, 1318–1321, https://doi.org/10.3201/eid1808.110988 (2012).
Brennan, M. B. et al. Transmission of Babesia microti Parasites by Solid Organ Transplantation. Emerg Infect Dis 22, https://doi.org/10.3201/eid2211.151028 (2016).
Linden, J. V. et al. Transfusion-transmitted and community-acquired babesiosis in New York, 2004 to 2015. Transfusion 58, 660–668, https://doi.org/10.1111/trf.14476 (2018).
Fang, L. Q. et al. Emerging tick-borne infections in mainland China: an increasing public health threat. Lancet Infect Dis 15, 1467–1479, https://doi.org/10.1016/S1473-3099(15)00177-2 (2015).
Krause, P. J. et al. Increasing health burden of human babesiosis in endemic sites. Am J Trop Med Hyg 68, 431–436 (2003).
Moritz, E. D. et al. Screening for Babesia microti in the U.S. Blood Supply. N Engl J Med 375, 2236–2245, https://doi.org/10.1056/NEJMoa1600897 (2016).
Rosner, F., Zarrabi, M. H., Benach, J. L. & Habicht, G. S. Babesiosis in splenectomized adults. Review of 22 reported cases. Am J Med 76, 696–701, https://doi.org/10.1016/0002-9343(84)90298-5 (1984).
Krause, P. J. et al. Persistent and relapsing babesiosis in immunocompromised patients. Clin Infect Dis 46, 370–376, https://doi.org/10.1086/525852 (2008).
Mareedu, N. et al. Risk Factors for Severe Infection, Hospitalization, and Prolonged Antimicrobial Therapy in Patients with Babesiosis. Am J Trop Med Hyg 97, 1218–1225, https://doi.org/10.4269/ajtmh.17-0146 (2017).
Menis, M. et al. Babesiosis Occurrence among the Elderly in the United States, as Recorded in Large Medicare Databases during 2006-2013. PLoS One 10, e0140332, https://doi.org/10.1371/journal.pone.0140332 (2015).
Kletsova, E. A., Spitzer, E. D., Fries, B. C. & Marcos, L. A. Babesiosis in Long Island: review of 62 cases focusing on treatment with azithromycin and atovaquone. Ann Clin Microbiol Antimicrob 16, 26, https://doi.org/10.1186/s12941-017-0198-9 (2017).
Hatcher, J. C., Greenberg, P. D., Antique, J. & Jimenez-Lucho, V. E. Severe babesiosis in Long Island: review of 34 cases and their complications. Clin Infect Dis 32, 1117–1125, https://doi.org/10.1086/319742 (2001).
White, D. J. et al. Human babesiosis in New York State: Review of 139 hospitalized cases and analysis of prognostic factors. Arch Intern Med 158, 2149–2154, https://doi.org/10.1001/archinte.158.19.2149 (1998).
Hildebrandt, A., Gray, J. S. & Hunfeld, K. P. Human babesiosis in Europe: what clinicians need to know. Infection 41, 1057–1072, https://doi.org/10.1007/s15010-013-0526-8 (2013).
Zintl, A., Mulcahy, G., Skerrett, H. E., Taylor, S. M. & Gray, J. S. Babesia divergens, a bovine blood parasite of veterinary and zoonotic importance. Clin Microbiol Rev 16, 622–636, https://doi.org/10.1128/cmr.16.4.622-636.2003 (2003).
Hunfeld, K. P., Hildebrandt, A. & Gray, J. S. Babesiosis: recent insights into an ancient disease. Int J Parasitol 38, 1219–1237, https://doi.org/10.1016/j.ijpara.2008.03.001 (2008).
Cornillot, E. et al. Sequencing of the smallest Apicomplexan genome from the human pathogen Babesia microti. Nucleic Acids Res 40, 9102–9114, https://doi.org/10.1093/nar/gks700 (2012).
Cornillot, E. et al. Whole genome mapping and re-organization of the nuclear and mitochondrial genomes of Babesia microti isolates. PLoS One 8, e72657, https://doi.org/10.1371/journal.pone.0072657 (2013).
Young, C. et al. Preventing transfusion-transmitted babesiosis: preliminary experience of the first laboratory-based blood donor screening program. Transfusion 52, 1523–1529, https://doi.org/10.1111/j.1537-2995.2012.03612.x (2012).
Homer, M. J. et al. A polymorphic multigene family encoding an immunodominant protein from Babesia microti. J Clin Microbiol 38, 362–368 (2000).
Lodes, M. J. et al. Serological expression cloning of novel immunoreactive antigens of Babesia microti. Infect Immun 68, 2783–2790 (2000).
Nishisaka, M. et al. Characterisation of the gene encoding a protective antigen from Babesia microti identified it as eta subunit of chaperonin containing T-complex protein 1. Int J Parasitol 31, 1673–1679 (2001).
Lemieux, J. E. et al. A global map of genetic diversity in Babesia microti reveals strong population structure and identifies variants associated with clinical relapse. Nat Microbiol 1, 16079, https://doi.org/10.1038/nmicrobiol.2016.79 (2016).
Luo, Y. et al. Identification and characterization of a novel secreted antigen 1 of Babesia microti and evaluation of its potential use in enzyme-linked immunosorbent assay and immunochromatographic test. Parasitol Int 60, 119–125, https://doi.org/10.1016/j.parint.2010.11.001 (2011).
Anantharaman, V., Iyer, L. M., Balaji, S. & Aravind, L. Adhesion molecules and other secreted host-interaction determinants in Apicomplexa: insights from comparative genomics. Int Rev Cytol 262, 1–74, https://doi.org/10.1016/S0074-7696(07)62001-4 (2007).
Homer, M. J. et al. Identification and characterization of putative secreted antigens from Babesia microti. J Clin Microbiol 41, 723–729 (2003).
Yokoyama, N. et al. Roles of the Maltese cross form in the development of parasitemia and protection against Babesia microti infection in mice. Infect Immun 71, 411–417 (2003).
Ooka, H. et al. Babesia microti: molecular and antigenic characterizations of a novel 94-kDa protein (BmP94). Exp Parasitol 127, 287–293, https://doi.org/10.1016/j.exppara.2010.06.018 (2011).
Snary, D. & Smith, M. A. Sequence homology of surface membrane proteins of Babesia rodhaini. Mol Biochem Parasitol 27, 303–312 (1988).
Cao, S. et al. Identification and characterization of an interspersed repeat antigen of Babesia microti (BmIRA). Exp Parasitol 133, 346–352, https://doi.org/10.1016/j.exppara.2012.12.015 (2013).
Cornillot, E. et al. A targeted immunomic approach identifies diagnostic antigens in the human pathogen Babesia microti. Transfusion 56, 2085–2099, https://doi.org/10.1111/trf.13640 (2016).
Silva, J. C. et al. Genome-wide diversity and gene expression profiling of Babesia microti isolates identify polymorphic genes that mediate host-pathogen interactions. Sci Rep 6, 35284, https://doi.org/10.1038/srep35284 (2016).
Gardner, M. J. et al. Chromosome 2 sequence of the human malaria parasite Plasmodium falciparum. Science 282, 1126–1132 (1998).
Kadekoppala, M. & Holder, A. A. Merozoite surface proteins of the malaria parasite: the MSP1 complex and the MSP7 family. Int J Parasitol 40, 1155–1161, https://doi.org/10.1016/j.ijpara.2010.04.008 (2010).
Jiang, J., Barnwell, J. W., Meyer, E. V. & Galinski, M. R. Plasmodium vivax merozoite surface protein-3 (PvMSP3): expression of an 11 member multigene family in blood-stage parasites. PLoS One 8, e63888, https://doi.org/10.1371/journal.pone.0063888 (2013).
Koch, M. et al. Plasmodium falciparum erythrocyte-binding antigen 175 triggers a biophysical change in the red blood cell that facilitates invasion. Proc Natl Acad Sci USA 114, 4225–4230, https://doi.org/10.1073/pnas.1620843114 (2017).
Rug, M., Prescott, S. W., Fernandez, K. M., Cooke, B. M. & Cowman, A. F. The role of KAHRP domains in knob formation and cytoadherence of P falciparum-infected human erythrocytes. Blood 108, 370–378, https://doi.org/10.1182/blood-2005-11-4624 (2006).
Tiburcio, M. et al. Specific expression and export of the Plasmodium falciparum Gametocyte EXported Protein-5 marks the gametocyte ring stage. Malar J 14, 334, https://doi.org/10.1186/s12936-015-0853-6 (2015).
Niang, M., Yan Yam, X. & Preiser, P. R. The Plasmodium falciparum STEVOR multigene family mediates antigenic variation of the infected erythrocyte. PLoS pathogens 5, e1000307, https://doi.org/10.1371/journal.ppat.1000307 (2009).
Niang, M. et al. STEVOR is a Plasmodium falciparum erythrocyte binding protein that mediates merozoite invasion and rosetting. Cell Host Microbe 16, 81–93, https://doi.org/10.1016/j.chom.2014.06.004 (2014).
Ruebush, M. J. & Hanson, W. L. Susceptibility of five strains of mice to Babesia microti of human origin. J Parasitol 65, 430–433 (1979).
Moitra, P. et al. Expression, Purification, and Biological Characterization of Babesia microti Apical Membrane Antigen 1. Infect Immun 83, 3890–3901, https://doi.org/10.1128/IAI.00168-15 (2015).
Mahajan, B. et al. Centrins, cell cycle regulation proteins in human malaria parasite Plasmodium falciparum. J Biol Chem 283, 31871–31883, https://doi.org/10.1074/jbc.M800028200 (2008).
Acknowledgements
We thank Bryan Grabias for the careful review of the manuscript and the FDA Intramural Research Program and the Gordon and Llura Gund Foundation for providing financial support for this study.
Author information
Authors and Affiliations
Contributions
N.V. and S.K. conceived the idea. N.V., A.P., E.E, H.Z., S.W., T.L. and P.K. contributed to data collection. N.V., V.A., S.B., L.A., K.G. and S.K. led the analysis and interpretation of the data. N.V., L.A., K.G. P.K. and S.K. wrote the first draft of the paper. All authors reviewed and approved the final draft of the paper prior to submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Verma, N., Puri, A., Essuman, E. et al. Antigen Discovery, Bioinformatics and Biological Characterization of Novel Immunodominant Babesia microti Antigens. Sci Rep 10, 9598 (2020). https://doi.org/10.1038/s41598-020-66273-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-66273-6
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.