Abstract
The world is in the midst of an antimicrobial resistance crisis, driving a need to discover novel antibiotic substances. Using chemical cues as inducers to unveil a microorganism’s full metabolic potential is considered a successful strategy. To this end, we investigated an inducible antagonistic behavior in multiple isolates of the order Bacillales, where large inhibition zones were produced against Ralstonia solanacearum only when grown in the presence of the indicator triphenyl tetrazolium chloride (TTC). This bioactivity was produced in a TTC-dose dependent manner. Escherichia coli and Staphylococcus sp. isolates were also inhibited by Bacillus sp. strains in TTC presence, to a lesser extent. Knockout mutants and transcriptomic analysis of B. subtilis NCIB 3610 cells revealed that genes from the L-histidine biosynthetic pathway, the purine, pyrimidine de novo synthesis and salvage and interconversion routes, were significantly upregulated. Chemical space studied through metabolomic analysis, showed increased presence of nitrogenous compounds in extracts from induced bacteria. The metabolites orotic acid and L-phenylalaninamide were tested against R. solanacearum, E. coli, Staphylococcus sp. and B. subtilis, and exhibited activity against pathogens only in the presence of TTC, suggesting a biotransformation of nitrogenous compounds in Bacillus sp. cells as the plausible cause of the inducible antagonistic behavior.
Similar content being viewed by others
Introduction
Since the discovery of penicillin1 and its subsequent large-scale production and introduction as a therapeutic agent2, disease control in both animals and plants has been mainly achieved by the use of natural products (NPs)3,4. Antimicrobials constitute a large fraction of these NPs, used to combat animal infections of bacterial origin and to control fungal or bacterial plant diseases5,6. Applications other than disease control have resulted in excessive and indiscriminate use of antimicrobials, with reports suggesting that around 80% of antimicrobial use is for non-therapeutic purposes in the US7,8 and 70% in the European Union countries9. Even though microbial resistance to bioactive NPs occurs naturally10, the excessive use of antibiotics selects resistance elements in bacterial populations and makes them available to be captured by previously antibiotic-sensitive pathogens11. This is believed to have precipitated the present-day global crisis of antimicrobial resistance (AMR). This crisis has become so severe that the World Health Organization (WHO) and the Food and Agriculture Organization (FAO) from the United Nations (UN) have started synergistic initiatives to counteract the growing appearance of resistant strains by promoting research programs worldwide12,13. This underscores the importance of discovery of novel antibiotic substances and antimicrobial technologies for use against pathogens of clinical, veterinary and agricultural importance14.
Antibiotics have been historically discovered and sourced from microbes4,6, with the soil microbiome being a well-known source of antimicrobial-producing bacteria and fungi. In many bacteria, significant portions of the genome are devoted to the production of NPs, also known as secondary metabolites, with some bacterial isolates having the potential to produce as many as 50 secondary metabolites6. These metabolites were traditionally considered to have a natural defense role for the producing organism, being mostly produced under stressful circumstances to selectively inhibit the growth of competitors15. However, recent research suggests that there may be alternative or additional roles for antimicrobials. For example, they may act as signaling or quorum sensing molecules and influence regulation of gene expression in natural ecosystems16,17. The probable functional diversity of these metabolites indicates that we need a broader understanding regarding the ecological context in which these small molecules are made, how they function in natural settings, and/or how they evolved16.
In modern drug discovery and development programs, most compounds currently in the clinical pipeline are modifications of existing classes of antibiotics and represent only short-term solutions to the growing resistance problem. There is a growing interest in the discovery of new families of compounds with novel structures and mechanisms of action that can be used to treat drug resistant pathogens18. This interest has been aided by the emergence of a set of innovative technical platforms and methodologies for discovering novel antimicrobials of microbial origin4,18. Among these platforms, new cultivation techniques to explore the diverse but yet to be cultured bacteria in soil are a major focus. Cultivation of bacteria is highly biased toward a few phylogenetic groups, many of which exclude currently underexplored bacterial lineages likely to hold novel biosynthetic gene clusters (BGCs) and thus, many biochemical pathways remain to be discovered19. The new cultivation approaches have yielded encouraging results, leading to the discovery of two promising new structures, lassomycin20 from Lentzea kentuckyensis and teixobactin21 from Eleftheria terrae. An alternative and complementary approach is the investigation of silent BGCs and their activation through chemical cues and microbial interactions22. This method has its grounds in the greater metabolic capacity of microorganisms reflected in their genome sequences than that which is expressed in traditional fermentation conditions. A term for this greater capacity is “OSMAC” (one-strain many compounds) and it suggests the activation of unexpressed BGCs through varying growth conditions or cues from other microbes22,23. These cues can be of natural origin if they are produced by a neighboring, interacting strain through co-culture, or external and of synthetic origin, achieved by adding individual or mixtures of compounds to the culture media24. The recent discovery of Amycomicin25 exemplifies the power of chemical enhancement and microbial interactions for unveiling novel metabolites. Further exploration of the total chemical space of those interactions, through metabolomics, can yield a complete spectrum of novel chemistries produced by a given strain during interactions or under inducing conditions4.
Here, we describe a system to find bioactive molecules in Bacillales, by causing an alteration in the electron and carbon flow within metabolic pathways using an unusual but well-known chemical cue, triphenyl tetrazolium chloride (TTC), at sub-inhibitory doses. This alteration in culture media triggers an induced antimicrobial activity in the Bacillales. TTC is a mild antibiotic for Gram-positive bacteria and a redox indicator and mediator compound26 which readily enter cells, where is reduced by NADPH-dependent oxidoreductases and dehydrogenases into insoluble TPF (triphenyl formazan)27,28.
We show that sub-inhibitory doses of TTC induce the production of compounds by Bacilliales which are active against plant pathogens of high economic importance in inter-tropical zones, such as R. solanacearum which attacks over 50 plant families29, and against mammalian pathogens such as E. coli and Staphylococcus sp. These compounds are produced by a plausible biotransformation mechanism between TTC and nitrogenous compounds overexpressed in Bacillus sp.
Results
TTC induces antimicrobial activity in several species of Bacillales against R. solanacearum in a dose dependent manner
R. solanacearum has been routinely grown on BGTA medium for pathogenomic studies30 while King B, LB and TSA media, have often been used to screen antagonist bacteria against R. solanacearum31. These media all contain complex nutrients but differ in the content of TTC salt. Here BGTA medium was used to evaluate antibacterial activity of 100 isolates from different species of the Bacillales order. Surprisingly, all isolates were inhibitory toward R. solanacearum. To determine whether antimicrobial activity was associated with TTC amendment, different isolates (Supplementary Table S1) were tested by the agar plug diffusion test against R. solanacearum. In general, all bacterial strains produced inhibition zones when grown in medium supplemented with TTC (BGTA medium) but not without TTC (BGA medium) with the exception of B. amyloliquefaciens EA-CB0959 (Table 1). Inhibition zones in the presence of TTC ranged from 7.3 to 26.8 mm, with B. cereus, B. pumilus, B. altitudinis, B. licheniformis, B. thuringiensis, B. coagulans and A. palidus (Table 1) showing highest antimicrobial activity. Using other tetrazolium salts (INT: 2-(4-iodophenyl)-3-(4-nitrophenyl)-5-phenyl-2H-tetrazolium chloride; NBT: Nitrotetrazolium blue, XTT: Sodium 2,3,-bis(2-methoxy-4-nitro-5-sulfophenyl)-5-[(phenylamino)-carbonyl]-2H-tetrazolium) did not produced significant inhibitory activity (Supplementary Fig. S1). Because these results might suggest that TTC sensitized R. solanacearum, an antibiogram was performed in the presence and absence of TTC. None of the antibiotics (ampicillin, erythromycin, gentamicin, kanamycin B, norfloxacin, penicillin, rifampicin, streptocycline, sulfamethoxazole) generated larger inhibition halos under TTC conditions (Supplementary Fig. S2), indicating that TTC does not sensitize the pathogen but induces the production of antimicrobial compounds in strains from the Bacillales.
To test if TTC induced antimicrobial activity in isolates from other taxonomic families such as Burkholderiales, Pseudomonadales, Enterobacteriales, various strains were tested in plug diffusion tests. Inhibition zones were significantly larger in S. marcescens UA-1538 and B. cepacea UA-1541; reduced in D. tsuruhatensis UA-1537, and TTC had no effect on P. putida UA-0095 and H. seropedicae UA-1542 (Supplementary Table S2). Compared to Gram-negative microorganisms, the inducible activity obtained in strains from the Bacillales was higher and more broadly spread across different species.
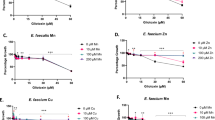
The size of the inhibition zones was found to be TTC-dose dependent for some strains of Bacillales (Fig. 1b–d) and not dependent on the presence of the target pathogen nor related to nutrient deprivation (Supplementary Fig. S3). Strains B. cereus EA-CB1047, B. pumilus EA-CB0009, B. subtilis NCIB 3610 and B. subtilis EA-CB0015, showed an increase in the inhibition zones that were dependent of the TTC dose applied, reaching a maximum between 50 to 100 mg/L (Fig. 1b–d), while B. amyloliquefaciens EA-CB0959 inhibition zones were not dose-dependent (Fig. 1e). Above 200 or 400 mg/L of TTC, inhibition zones of B. cereus were reduced (Fig. 1b), due to impaired growth at those concentrations (Supplementary Fig. S4a). In contrast, R. solanacearum growth was not affected by low-medium (0–100 mg/L) or high (200–400 mg/L) TTC concentrations (Supplementary Fig. S5a). Furthermore, neither B. cereus, B. pumilus nor R. solanacearum growth was impaired in the presence of high concentrations of the TTC salt reduction product TPF (from 0–400 mg/L) (Supplementary Figs. S4b, S4d, S5b). These results altogether suggest that TTC induces the expression of antimicrobial compounds in Aerobic Endospore Forming Bacteria (AEFB) against R. solanacearum and possibly against Gram-negative pathogens.
Effect of TTC concentration on inducible antagonism production by Bacillus sp. strains. (a) Inducible antagonism phenomenon in the presence of TTC of B. cereus EA-CB1047 (Left BGA, right BGTA medium). Effect of different TTC concentrations on the size of inducible inhibition zones produced by (b) B. cereus EA-CB1047, (c) B. pumilus EA-CB0009, (d) B. subtilis EA-CB0015 (■) and B. subtilis NCIB 3610 (□), and (e) B. amyloliquefaciens EA-CB0959. Interval bars represent standard errors of the mean (n = 3). Bars sharing capitalized or non-capitalized letters do not differ statistically in their values, determined by ANOVA followed by a Tukey test (for b–d■ and e) and by Kruskal-Wallis methods and post-hoc Dunn test for multiple comparisons (for d□). B. cereus EA-CB1047 P-value = 4.34 × 10−11, B. pumilus EA-CB0009 P-value = 1.89 × 10−13, B. subtilis EA-CB0015 P-value = 1.19 × 10−10, B. subtilis NCIB 3610 P-value = 8.62 × 10−2, B. amyloliquefaciens EA-CB0959 P-value = 3.30 × 10−3).
TTC inducible activity works against other target bacterial species
The spectrum of activity of the inducible antagonism produced by B. cereus EA-CB1047 and B. subtilis NCIB 3610, was determined against isolates representing diverse genera. Inducible activity was observed against Cupriavidus necator for B. subtilis NCIB 3610 (Fig. 2a); and Staphylococcus sp. and E. coli for B. cereus EA-CB1047 (Fig. 2b), albeit to a lesser extent than against R. solanacearum.
Antagonistic activity of B. subtilis NCIB 3610 and B. cereus EA-CB1047 against target bacterial strains in the presence and absence of TTC. (a) Effect of TTC on the antagonistic activity of B. subtilis NCIB 3610 and (b) B. cereus EA-CB1047 against several target strains. Intervals represent standard errors of the mean (n = 3). +I denotes 50 mg/L TTC amendment in the medium. −I denotes the absence of TTC from the medium. * denotes a statistically significant difference (P < 0.05) between the conditions −I and +I for each antagonistic strain, according to Student t-test ((a) R. solanacearum AW1 P-value = 1.72 × 10−3, C. necator H16 P-value = 2,53 × 10−3, (b) Staphylococcus sp. G1 P-value = 2.93 × 10−3, E. coli DH5α P-value = 1.26 × 10−5; R. solanacearum AW1 P-value = 5.37 × 10−3).
TTC induces antimicrobial activity of Bacillus sp. strains in liquid co-cultures and mono-cultures
To investigate whether TTC-induced antimicrobial activity was restricted to solid cultures, liquid co-cultures were established (Supplementary Fig. S6). We found that in the presence of TTC, the population of R. solanacearum decreased 8.3-fold after the onset of stationary phase for B. cereus. Similar results were found for B. subtilis NCIB 3610 strain (data not shown). Such a pronounced decrease in the population was not observable for R. solanacearum in control cultures where TTC was not present. The production of inducible antimicrobial activity in liquid medium, detected through the decrease of R. solanacearum colony forming units at approximately 14 h of culture, indicates that inducible compounds can be produced both in liquid and solid media when supplemented with TTC. Although cell-free supernatants of liquid cultures obtained from monocultures of B. cereus EA-C1047 grown in the presence of TTC showed activity when sampled between 12–14 h post-inoculation (data not shown), inducible activity was always enhanced on solid media, thus all further tests were done in agar plates, including the extraction of inducible compounds from inhibition zones for metabolomic analysis.
B. subtilis carrying mutations in biosynthetic genes for characterized natural products are not defective in producing inducible inhibition zones
Different B. subtilis NCIB 3610 knockout mutant strains defective in the production of well-known antibiotics or in major regulatory pathways (Supplementary Table S3) were tested through the agar plug diffusion test against R. solanacearum AW1 and EAP-009 (Fig. 3). No strain was found to be defective, but significantly larger inhibition zones were produced by B. subtilis NCIB 3610 mutants spo0A::erm, kinA::mls, kinB::kan, kinC::cat. The gene products of spo0A, kinA, kinB, kinC or downstream expression cascades and metabolic routes regulated by these genes, may be involved in the repression of the inducible antagonism in the WT strain.
Inhibition zones produced by knockout mutants of B. subtilis NCIB 3610 against R. solanacearum in BGTA medium. (a) B. subtilis NCIB 3160 mutants against R. solanacearum AW1, (b) B. subtilis NCIB 3160 mutants against R. solanacearum EAP-009. Intervals represent standard errors of the mean (n = 2). Bars sharing non-capitalized letters do not differ statistically in their values, by ANOVA followed by a Tukey test. ((a) P-value = 1.21 × 10−4, (b) P-value = 2.98 × 10−3).
Gene-expression profiling of B. subtilis NCIB 3610 cells in presence of TTC shows an upregulation of central genes involved in nitrogen metabolism
Transcriptomic profiling obtained by total mRNA-seq and Nanostring both showed expression changes in genes involved in nitrogen metabolism. Specifically, RNA-seq data showed six genes were differentially expressed by TTC-induced B. subtilis NCIB 3610 cells relative to controls (log2 of fold-ratio from 4.7 to 5.2), across the three bioinformatic methodologies used (Fig. 4a, Supplementary Tables S4a and S4b). These differentially expressed genes in B. subtilis NCIB 3610 were in the major his biosynthetic operon (hisB, hisD, hisG, hisH, hisE) (Fig. 4a). Results of transcriptomic profiling obtained from Nanostring for B. subtilis NCIB 3610 (Fig. 4b) showed significant upregulation of genes related to nitrogen metabolism through pyrimidine and purine de novo synthesis and salvage and interconversion routes (pyrP, pyrC, purF, xpt), as well as an onset on the DNA damage-machinery (mutS), which is related to DNA repair during oxidative stress. On the other hand, downregulated genes were related to exopolysaccharide synthesis (epsA) and membrane fluidity and stability (sqhC). Consistent with data from mutant studies (above), no induction of genes linked to well-known natural products of B. subtilis (bacillaene (pks), sublancin (sunA), surfactin (srfAA), plispastatin (ppsA), subtilisin (aprE) and bacilysin (bacA)) was observed using either technique. Furthermore, when testing the effect of siderophore production by the chromeazurol S (CAS) assay, no increments were found on the size of the inducible inhibition zones (Supplementary Fig. S7).
Overall results for transcriptomic profiling indicate that the two transcriptomic analysis methodologies performed for B. subtilis NCIB 3610 cells growing in the presence of TTC converge towards the same induced metabolic route: purine and pyrimidine biosynthesis. All upregulated metabolic pathways are interconnected, with L-histidine and other amino acids such as alanine, glutamate and aspartate being direct precursors for biosynthesis of purine and pyrimidine nitrogenous bases.
TTC causes alterations in central nitrogen metabolism in B. subtilis NCIB 3610
The effect of TTC on the metabolic profile of B. subtilis NCIB 3610 is represented in a complex network generated by Cytoscape32 using mass spectrometry (MS) data, which were initially analyzed and grouped through the GNPS platform33 (Fig. 5). Individual features (precursor ions) present in each treatment are denoted as single nodes within the spectral network, and each node contains one or more MS2 spectra which were determined to be identical by the algorithm applied through GNPS platform33. The edges connecting nodes denote structural relatedness. The resulting network originally contained 1265 total nodes, distributed in a specific fashion among treatments (Supplementary Table S5).
Aggregated and refined spectral network of the metabolomic profile of B. subtilis NCIB 3610 extracts. Complete spectral network composed of nodes representing features associated with B. subtilis NCIB 3610: active extracts obtained from inducible inhibition zones (blue nodes), extracts from non-induced control (purple nodes), compounds shared between these two extracts (cyan nodes) and other extracts and absolute controls: TTC in 50% methanol (red nodes) and precursor ions shared between control extracts (BGTA medium and 50% methanol solution) and active extracts (grey nodes). These grey nodes denote abundance by more than 2-fold of the precursor ion in active extracts than in control. This figure was created using Cytoscape Version 3.7.1., an open source software platform for visualizing complex networks and integrating these with any type of attribute data. URL: https://cytoscape.org/.
In the complete network (Fig. 5), several nodes were found to present hits when screened against natural products libraries. These clusters report the presence of surfactins, bacilysin, glycyl-leucine, cyclo-(Ala-4-Hyp), diglycosyl diacylglycerol and N-acetylprimaquines, fengycins, siderophores, bacillibactin, desferrioxamine-related compounds, amino acids, dipeptides and cyclic dipeptides, and N-acetyl-phenylalanyl-valine as well as cell wall components like N-acetyl D-galactosamine and diglycosyl diacylglycerol (Fig. 5, Supplementary Table S6). Interestingly, nitrogenous compounds like amino acids, dipeptides and cyclic dipeptides reported higher spectral counts in the active extract than in the non-induced control extract, as did the N-acetyl primaquines, suggesting an alteration in the nitrogenous intermediates in active extracts. The network includes a pair cluster corresponding to the inducer compound TTC, including a node with the exact inducer compound molecular mass (334.1 + [H]+ = 335.125).
Of particular interest is the subnetwork-1 (Fig. 5) which is made up of 82 nodes representing structurally related unidentified compounds which had 3-fold or higher spectral counts in the active extracts than in the non-induced control. Final selection of nodes for data mining included those in subnetwork-1 which were unique to the active extract or that had at least a 3-fold increase in spectral counts in the active extract. In addition, unclustered features from the larger network classified as unique or only present in the active extract (blue nodes) were included, for a total of 28 selected features (Supplementary Table S7).
Analysis of data mining results suggests that the abundant compounds include selected functional groups associated with intermediates or final products of the nitrogen metabolism biosynthetic routes (Supplementary Table S8). A selection of nitrogen metabolism intermediates and end products which possess the detected functional groups is displayed in Table 2. The list includes compounds such as L-histidine and orotic acid which transcriptomic analysis suggested may be induced. All compounds were validated by comparing the MS2 fragmentation pattern of each potential compound to the one of the associated unidentified feature from the network (query).
Compounds of nitrogen metabolism inhibit pathogens sensitive to inducible antagonism, in the presence of TTC
Metabolites from Table 2, specifically L-arginine, L-histidine, imidazole, orotic acid, L-phenylalanylamide, and 1,1-dimethylguanidine were tested at concentrations of 100 µM, 100 mM and 1 M against R. solanacearum EAP-009, and strains of Staphylococcus sp., E. coli, B. subtilis NCIB 3610, Xanthomonas sp. and S. enterica (Table 3). R. solanacearum growth was inhibited by all six compounds in the presence of TTC at concentrations of 1 M, and by orotic acid at 100 mM. In the absence of TTC, R. solanacearum was only inhibited by L-arginine and 1,1-dimethylguanidine at concentration of 1 M. Staphylococcus sp. cells were sensitive to five compounds, excluding 1,1-dimethylguanidine, at 1 M in the presence of TTC (Table 3). E. coli DH5α was the least sensitive to the nitrogenous compounds, being inhibited only by L-arginine in both the presence and absence of TTC, and by imidazole and L-phenylalaninamide at concentrations of 1 M with TTC present. B. subtilis cells were inhibited by all tested compounds with the exception of orotic acid and L-phenylalaninamide. These two compounds became of special interest by being active against the pathogens but not against the producing strain. From results (Table 3), it can be concluded that the inhibitory activity of these compounds either occurs only the presence of TTC or it is significantly higher when the inducer compound is present.
Although tested compounds showed activity in high concentrations (100 mM−1M) which would not be suitable for in vivo applications, these nitrogenous substances alone are probably not the direct product of the inducible phenomenon. Instead, these results suggest that an enzymatic biotransformation of nitrogenous compounds (i.e. orotic acid, L-phenylalaninamide) is occurring with TTC or its reduced product TPF, which might be an incomplete reaction, meaning that reactants are not completely converted to products. Thus, smaller quantities of the nitrogenous products might be needed to observe bioactivity in an in vivo system. These products which result from reaction with TTC/TPF are suggested to have bioactivity against R. solanacearum and against other target pathogens (e.g. Staphylococcus sp., and E. coli), and are being studied in detail to elucidate their complete chemical structure.
Discussion
Isolates from the order Bacillales, which includes the genus Bacillus, are well known to produce bioactive substances, with some of these compounds being extensively studied for their in vitro and in vivo biological activities including anti-fungal, anti-inflammatory and anti-tumor properties34,35. But can novel compounds, with diverse and exceptional properties, still be found in widely explored microbial orders such as actinomycetes36 and Bacillales34?. A promising research avenue is to study the chemistry of microbes within their environments and while interacting with chemical signals37, in order to increase the likelihood of finding novel metabolites active against antibiotic resistant pathogens, as well as to inhibit crop pathogens5. The research presented here exemplifies this type of alternative platform for discovering active biomolecules. Results here indicate that Bacillales strains generate inducible antagonistic compounds against R. solanacearum, E. coli and Staphylococcus sp. (Figs. 1 and 2) by a process of induction and a possible biotransformation (Table 3). Using wild isolates and type strains (Supplementary Table S1) belonging to several species from the Bacillales order, this research represents innovation in natural product investigations, specifically the capacity of TTC to change the redox balance within the cells and act as an elicitor of bioactivity in the Bacillales.
The production of inducible inhibition zones by Bacillus sp. against R. solanacearum occurs in a TTC dose-dependent fashion (Fig. 1), a phenomenon previously observed in several antibiosis-inducible species when using chemical signals, specially antibiotics24, acting as elicitors or inducers of otherwise silent BGCs in microbial genomes. Various reports support the role of the chemical environment surrounding bacteria as a trigger of metabolic responses, with the term OSMAC illustrating how the metabolic output of a strain can be finely tuned or stimulated in the presence of natural or synthetic signals22,23,24. Some of the most recently reported novel bioactive substances have been isolated either from a chemical interaction with a neighboring microbial strain or natural signal25 or by enrichment cultivation methods which concentrate the chemical ecological signals and allow the growth of otherwise non-cultivatable bacteria21,38. Tetrazolium salts, particularly the widely used compound TTC, have not been reported as inducers of antimicrobial behavior, and thus this study reveals a novel platform of induction of bioactive substances in microbial strains.
TTC does not induce antibacterial activity by every bacterium. The inducible inhibition zones observed in this study, produced by bacterial isolates in the presence of TTC, are more strongly produced by species from the Bacillales order (Table 1, Fig. 1), than by species from other taxonomic families as Burkholderiales, Pseudomonadales, Enterobacteriales (Supplementary Table S2). These results indicate an intrinsic capacity of Bacillus sp. and related species to be more susceptible to metabolite induction than other taxonomical orders tested, with respect to the production of these specific antimicrobial substances in the presence of TTC. Among bacteria there is a high diversity of dehydrogenases and respiratory systems39, and it is possible that not all dehydrogenase systems are capable of using TTC to the same extent, becoming toxic to some bacteria at high concentrations (>500 mg/L)26 while allowing an enhanced capacity of Bacillus sp. to metabolize TTC. This could occur since TTC readily enters the cells, mostly driven by the partial negative charge inside cell membranes due to membrane potential phenomenon27, and its intracellular reduction is fundamentally done by membrane bound respiratory chain dehydrogenases. It was then hypothesized that Reactive Oxygen Species (ROS), well known for their cell damaging effects in bacteria40, could be overproduced due to the enhanced reductive activity occurring inside Bacillus sp. cells and thus be responsible for affecting R. solanacearum growth. However, complementation of culture media with antioxidants compounds (α-tocopherol, uric acid, ascorbic acid, L-glutathione) did not alter the size of inhibitions zones produced by B. cereus EA-CB 1047 and B. subtilis NCIB 3610 against R. solanacearum (Supplementary Fig. S8).
Four other traits of the inducible antagonism observed in the presence of TTC support the conclusion that a chemical interaction with TTC is the triggering mechanism for producing the observed antibiosis: first, the induction is TTC dose-dependent (Fig. 1), with the largest inducible inhibition zones observed within the range of 50–100 mg/L for all Bacillus species tested. Second, TTC does not sensitize the pathogen R. solanacearum to any from a wide range of antibiotic classes (Supplementary Fig. S2). Third, the antagonism test using polyamide membranes shows that nutrient depletion is not the cause of observed inhibition zones and excludes a possible microbial interaction with the target pathogen (Supplementary Fig. S3). Finally, when knockout mutant stains from B. subtilis NCIB 3610 were tested for inducible antibiosis capacity (Fig. 3) none was repressed in the production of TTC-induced inhibition zones. Given that the majority of these knockout mutants were in genes related to biosynthesis of known natural products such as surfactins and bacilysin, it can be suggested that these known compounds are not directly responsible for the observed growth impairment of target strains and are apparently not being induced, and thus TTC likely induces production of other inhibitory compounds.
When the system was investigated using gene expression analysis, a significant upregulation of the nitrogen metabolism (Fig. 4, Supplementary Tables S4a and S4b) was observed from two different methodologies employed: total RNA-seq and Nanostring. This observation does not correlate with other transcriptomic findings for antimicrobial-producing Bacillales strains: for example, Yang et al.41 studied antibiosis activity of B. subtilis induced by a neighboring B. cereus strain and found that most overexpressed genes were related to glycolysis and the TCA cycle. Another study looked at a plant-growth-promoting (PGP) strain of B. subtilis (OKB105) from a transcriptomics perspective during interaction with its host plant, where surfactin production is a key factor in its protective effects. That study revealed several overexpressed metabolic routes, specifically carbohydrate metabolism, processes of transport/binding and lipoproteins and RNA regulation42.
Among some of the most interesting results of the present research is the convergence of results between the two transcriptomic approaches, RNA-seq and Nanostring, which both indicate that the L-histidine, purine and pyrimidine biosynthetic routes were highly upregulated in B. subtilis NCIB 3610 cells in inducible conditions (Fig. 4, Supplementary Table S4). L-histidine and other amino acids such as alanine, glutamate and aspartate are direct precursors for purine and pyrimidine nitrogenous base biosynthesis. This can support the transcriptomic results which suggest a common deviation of much of the cell’s pool of excess of nitrogen to these biosynthetic routes, possibly resulting from TPF accumulation. Another possible scenario for this diversion of the cell nitrogen metabolism is an upregulation of amino acid and nitrogenous base biosynthesis, specifically through the his biosynthetic operon and purine and pyrimidine salvage and interconversion routes (pyrP, pyrC, purF, xpt), due to an excess of reducing equivalents NAD+ and NADP+ in their oxidized form, produced due to high reduction rates of TTC into TPF.
Besides these findings, gene-expression analysis (Fig. 4) supports results from experiments with knockout mutants and validates the hypothesis that previously characterized natural products are not directly responsible for the inducible antagonism. The expression of known NP biosynthetic genes was not significantly altered by the presence of TTC. There is strong evidence to support the conclusion that characterized NPs produced by Bacillus are not the compounds responsible for the inducible inhibition zones produced by Bacillus sp. cells in the presence of TTC.
Metabolomic data in subnetwork-1 are supportive of the interpretation of transcriptomic data. The metabolomic data converge on detection of unique or more numerous spectral counts of nitrogenous compounds in extracts from inducible inhibition zones (Fig. 5). The detailed analysis from these nodes, coupled with transcriptomic analysis information, led to the selection of compounds such as L-arginine, L-histidine, imidazole, orotic acid, phenylalaninamide and 1,1-Dimethylguanidine for antagonism tests (Table 2, Table 3). Results from the antagonism tests of these compounds show that most of the metabolites are only active in the presence of TTC and are acting as antimicrobials when in high concentrations. Above values of 100 mM, most of them, which include precursors, intermediates and end-products of biosynthetic routes from nitrogen metabolism, are active against both plant and mammalian pathogens (R. solanacearum, E. coli and Staphyloccous sp.) (Table 3). Of special interest are orotic acid and L-phenylalaninamide, which both showed activity against the sensitive pathogens but not against the producing strain B. subtilis. Results (Table 3) highlight the role of orotic acid, which has previous reports of acting as an antibacterial agent43, and has a central role as a precursor of pyrimidine nucleotides44.
From overall results we conclude that the overproduction of nitrogen metabolism intermediates is related to the process of intracellular TTC reduction and TPF accumulation. Foundations are set to further explore an enzymatic biotransformation of these nitrogenous intermediates with TTC or TPF, resulting in biologically relevant compounds which are acting as the inducible antimicrobials. Further work, suggesting a plausible mechanism of inhibition, is a priority for this research. Work is currently ongoing in this area by analyzing the genomes of the target pathogen R. solanacearum in the search for genes which could code for targets of the compounds, and by determination of the identity and structure of the inhibitory molecule or group of molecules within this pool of bio-transformed nitrogenous compounds.
Materials and Methods
Microorganisms
Aerobic Endospore Forming Bacteria (AEFB) isolated from Musa sp. plants (Humboldt Institute Collection No. 191), B. subtilis NCIB 3610 (wild type, WT), B. subtilis SMY, B. subtilis PY79 and knockout strains of NCIB 3610 (Supplementary Tables S1 and S3) were used in the different inducible antagonism trials. These strains were stored in TSB (trypticase soy broth, Merck, Germany) with 20% v/v glycerol at −80 °C and activated in TSA (trypticase soy agar, Merck) or LB agar (Luria Bertani Agar, Basingstoke, Oxoid, England) for 48 h at 30 °C before use. Strains R. solanacearum EAP-009 (GenBank accession N° KU603426), Serratia marcescens EAD-005 (GenBank accession N° KU603427)45 and R. solanacearum AW146, all were stored in BG medium47 plus 20% v/v glycerol at −80 °C and activated at 30 °C for 72 h in BGA medium (BG plus 18 g/L agar (BD, Ontario, Canada)). BG medium was composed of 10 g/L special peptone (Oxoid), 1 g/L casamino acids (BD), 1 g/L yeast extract (Merck) and 5 g/L glucose (Merck). Pseudomonas putida UA-0095, Xanthomonas sp. UA-1539, B. cepacea UA-1541, S. marcescens UA-1538, Salmonella sp. ATCC 14028, Staphylococcus sp. G and E. coli DH5α (Supplementary Table S1) were stored in TSB (trypticase soy broth, Merck) with 20% v/v glycerol at −80 °C and activated in TSA or LB agar for 48 h at 30 °C before use.
Inducible antagonism tests
BGTA medium (BGA supplemented with 50 mg/L of TTC (Merck, Darmstadt, Germany)), was used for inducible antagonism tests. BGA medium, without TTC48 served as a control. Two methods of antagonism tests49 were employed: (i) agar plug diffusion test, to evaluate the inducible antagonism capacity of different bacterial strains against target strains, and (ii) Well-diffusion test, to determine the inducible antagonism capacity of strains belonging to B. subtilis NCIB 3610 knockout mutants against two strains of R. solanacearum, EAP-009 and AW1 and the antibiotic activity of extracts. Briefly, for the agar plug diffusion test, 100 μL of the target strain at 1 × 106 CFU/mL were plated onto BGTA and BGA plates. In another version, the target strain was submerged into molten BGTA or BGA at a desired final concentration and mixed well. A 0.5 cm plug (diameter) containing the antagonistic strain grown in the appropriate medium (50% TSA, LB) was placed on top of the target strain. After 48 h of incubation at room temperature (22 ± 1 °C), the radius size of inhibition zones were recorded. For the well diffusion test, wells were opened with a sterile Pasteur pipette in BGTA and BGA plates containing the target strain, either submerged or plated, followed by the addition of 50 μL of an overnight inoculum of the antagonistic strain grown at 22 °C and 150 rpm. After incubation at 22 °C for 48 h, the radius of inhibition zones was registered in two equidistant points from the center of the plug or well and an average value was obtained. In each plate, three plugs or wells were made for each treatment and three replicates (plates) per treatment were used. Each experiment was independently repeated two or three times. Variations in the amendments to the culture media, such as tetrazolium salts, TTC concentration, and antioxidants, were done accordingly to evaluate different conditions as outlined in the relevant places in the text.
Transcriptome analysis
The effect of TTC on the transcriptome profile of Bacillus sp. cells was performed by total RNA sequencing (RNA-seq) and Nanostring platform for B. subtilis NCIB 3610. For RNA-seq, RNA was extracted from submerged cultures of B. subtilis NCIB 3610 grown either in BGT or BG for 14 h at 22 °C and 150 rpm. At 14 h of culture, 500 μL were withdrawn and treated with RNAprotect cell reagent (Qiagen, Netherlands) and then every step of the RNeasy RNA isolation kit (Qiagen, Netherlands) was followed. Total RNA libraries were prepared with kit for Illumina NEBNext (New England Biolabs, MA, USA) and sequenced at TUCF Genomics (Tufts University Core Facility, Boston, MA, USA). Data analysis was carried using an automatic pipeline in Slurm Workload Manager 18.08.1. Differential expression analysis was performed through three approaches: Tuxedo suite, specifically the CuffDiff tool50, R-package DE-seq51 and EDGE-pro52 in order to calculate differential gene expression (DE) through the normalized parameter RPKM (read per million reads per million bases sequenced). Fold change was calculated by the ratio of the RPKM parameter in the presence of TTC vs absence of TTC, and its significance by the statistic p-adj value. Statistical analysis and data visualization prior to data analysis was done using PCA (principal component analysis) and Venn diagrams. PCA was done to asses replicates dispersion and Venn diagrams to find common DE genes across methodologies. Reported values consist of log 2 scale of fold ratios.
For Nanostring, total RNA was extracted from well diffusion assays in BGTA plates and control BGA, from three biological replicates per treatment using RNeasy RNA isolation kit (Qiagen, Netherlands). The experiment was repeated two times independently. Counts for each gene of B. subtilis NCIB 3610 were subjected to a technical normalization considering the counts obtained for positive control probe sets. After the technical normalization, a biological normalization was done using housekeeping genes def, folB, glpF, gsaB, ptsG included in the CodeSet used, via the Nanostring nCounter analysis system (Nanostring technologies, USA)53. We applied student’s t-test to evaluate the significance of gene expression differences between the TTC supplementation and control using companion software nSolver package (Nanostring technologies, USA). Relative over or under-expression of each gene across different groups was calculated in log2 scale of fold ratio and the level of statistical significance was defined as p ≤ 0.0554. Each experiment was independently repeated two times for both methodologies (Nanostring and RNA-seq).
Metabolomic analysis
The effect of TTC on the metabolic profile of induced B. subtilis NCIB 3610 cells was determined by UHPLC/MS. Active and controls extracts were obtained by methanolic extraction of 20 g of agar from inhibition zones by a methodology based on the studies by Liu et al.55 and Watrous et al.56 were agar extractions were performed. Briefly, 20 g of agar cut from control and antagonism plates were mixed with 80 mL of methanol in sterile glass flasks, sonicated with 30% amplitude for 20 min and agitated by pulse-vortexing every 20 min for 4 h. The solution was then filtered by 0.2 μm polyamide filter and the solvent evaporated 10X. Control extracts were obtained by extracting 20 g from B. subtilis NCIB 3610 grown on BGA plates with no presence of TTC and no R. solanacearum inoculation (non-induced control). Negative controls included: media control, obtained by extracting 20 g of BGTA with methanol, inducer control (solution of 50 mg/L of TTC in methanol) and methanol control (50% of aqueous methanol solution). Activity evaluation of each extract followed prior to metabolomic analysis, through agar well diffusion method, using R. solanacearum AW1 as target strain.
Metabolomic data acquisition
Two replicates of each treatment or control were subjected to UHPLC/MS analysis. The equipment for acquiring metabolomic data consisted of an UHPLC system (Dionex UltiMate 3000, ThermoFisher, USA) coupled to a mass spectrometer (MS, Thermo Scientific Q-Exactive Quadrupole-Orbitrap mass spectrometer with a Heated Electrospray Ionization (HESI) source). Sample was injected (5 µL) into a C18 column (50 mm × 2.1 mm, 2.2 µm, Thermo Scientific AcclaimTM RSLC) at 0.4 mL/min and 30 °C. Separation was carried by a gradient of solvent A (water + 0.1% formic acid) and solvent B (methanol + 0.1% formic acid), starting at 10% B for 0–1.5 min, then increasing from 10–100% from 1.5–11.5 min, then at 100% B from 11.5–14.5 min, lowering from 100–10% from 14.5–15 and ending at 10% B from 15–18 min. Peak detection was done at 214 nm on a universal diode array detector, followed by injection into the coupled MS equipment. MS analyses were done in a positive ion mode, with a scan range of 150–2000 m/z. Full scan parameters were: resolution of 70,000 full width at half-maximum (FWHM), automatic gain control (AGC) target of 3 × 106 ions and a maximum ion injected time (IT) of 100 ms. The injection capillary temperature was 300 °C, sweep gas was N2 and spray voltage was 3.5 kV. MS/MS parameters were: resolution of 17.500 FWHM, AGC target of 1 × 105 ions, maximum IT of 50 ms, quadrupole isolation window of 4.0 m/z, normalized collision energy (NCE) of 30%. Tandem MS was acquired using the data-dependent Top5 method considering precursor ion abundance. Data obtained from UHPLC and MS was visualized using Xcalibur software.
Bioinformatic analysis and spectral networking
Data files (.mzXML) were uploaded to the Mass Spectrometry Interactive Virtual Environment (MassIVE) data repository for further analysis through Global Natural Products Social Molecular Networking (GNPS) web platform33. Conditions for GNPS analysis are detailed in Supplementary Table S5. Spectral network (Fig. 5) was generated using data from GNPS and cytoscape. Further network curation and annotation was done manually, through literature reports and AntiSmash analysis57 (Supplementary Table S6); the majority of control nodes were removed, leaving only the ones in which the spectral counts from the active extracts outnumbered 3 fold or more the detected spectra from control extracts. m/z values of sub-network-1 (Fig. 5) nodes were extracted manually and used as queries on molecular weight search on Chemspider58 and PubChem59 databases using a source code (Supplementary Table S8).
Candidate compounds testing
Different compounds derived from metabolomic analysis were tested against target strains, R. solanacearum, Staphylococcus sp., E. coli, Xanthomonas sp., S. enterica (Table S1). Specifically, 1 M stocks of: imidazole (Sigma-Aldrich, OH, USA), uric acid (Sigma-Aldrich, OH, USA), L-histidine (Sigma-Aldrich, OH, USA), L-arginine (Sigma-Aldrich, OH, USA), Uracil (Sigma-Aldrich, OH, USA), 1,1-dimethylguanidine (Sigma-Aldrich, OH, USA) orotic acid (Sigma-Aldrich, OH, USA), L-phenylalanylvaline (Sigma-Aldrich, OH, USA) were prepared in appropriate solvents (sterile distilled water, 50% aqueous methanol for 1,1-dimethylguanidine) and filtered through a polyamide 0.2 µm pore-size syringe filter (Sartorius, Gottingen, Germany). Well-diffusion antagonism tests were prepared with each compound reaching final concentrations of 100 µM, 100 mM and 1 M.
Statistical analysis
All datasets for each experiment were analyzed either in R60 or by a Student t-test (TTEST function) when having two condition datasets. As a first step for R package analysis, datasets were tested for their fitting to ANOVA (analysis of variance) assumptions: independence of observations, normal distribution of residuals and homoscedasticity or homogeneity in variance of residuals. If assumptions were met, a one-way ANOVA analysis followed, with an α = 0.05 (95% family-wise confidence level), calling statistically significant differences across means of treatments if P-value ≤ 0.05. Post-hoc analysis consisted of a Tukey honestly significance difference test in order to find specific differences among groups. If ANOVA assumptions were not met, a Kruskal-Wallis non-parametric test was performed, and homogeneous groups were found if P-value ≤ 0.05, by applying Dunn post-hoc method.
Data availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Fleming, A. Classics in infectious diseases: on the antibacterial action of cultures of a Penicillium, with special reference to their use in the isolation of B. influenzae by Alexander Fleming, Reprinted from the British Journal of Experimental Pathology 10:226–236, 1929. Rev Infect Dis. 2, 129–139 (1980).
Davies, J. & Davies, D. Origins and evolution of antibiotic resistance. Microbiol Mol Biol Rev. 74, 417–433, https://doi.org/10.1128/MMBR.00016-10 (2010).
Demain, A. L. From natural products discovery to commercialization: a success story. J Ind Microbiol Biotechnol. 33, 486–495, https://doi.org/10.1007/s10295-005-0076-x (2006).
Harvey, A. L., Edrada-Ebel, R. & Quinn, R. J. The re-emergence of natural products for drug discovery in the genomics era. Nat Rev Drug Discov. 14, 111–129, https://doi.org/10.1038/nrd4510 (2015).
McManus, P. S., Stockwell, V. O., Sundin, G. W. & Jones, A. L. Antibiotic use in plant agriculture. Annu Rev Phytopathol. 40, 443–465, https://doi.org/10.1146/annurev.phyto.40.120301.093927 (2002).
Demain, A. L. Importance of microbial natural products and the need to revitalize their discovery. J Ind Microbiol Biotechnol 41, 185–201, https://doi.org/10.1007/s10295-013-1325-z (2014).
Van Boeckel, T. P. et al. Global trends in antimicrobial use in food animals. Proc Natl Acad Sci USA 112, 5649–5654, https://doi.org/10.1073/pnas.1503141112 (2015).
Laxminarayan, R., Van Boeckel, T. & Teillant, A. The Economic Costs of Withdrawing Antimicrobial Growth Promoters from the Livestock Sector. (OECD Publishing, 2015).
European Medicines Agency & European Surveillance of Veterinary Antimicrobial Consumption. Sales of veterinaryantimicrobial agents in 30 European countries in 2016. (2018).
D’Costa, V. M. et al. Antibiotic resistance is ancient. Nature 477, 457–461, https://doi.org/10.1038/nature10388 (2011).
Brown, E. D. & Wright, G. D. Antibacterial drug discovery in the resistance era. Nature 529, 336–343, https://doi.org/10.1038/nature17042 (2016).
Food and Agriculture Organization FAO. 25 (2016).
World Health Organization WHO. Global Action Plan on Antimicrobial Resistance. 28 (2015).
Wilkinson, K. et al. Infectious diseases of animals and plants: an interdisciplinary approach. Philos Trans R Soc Lond B Biol Sci 366, 1933–1942, https://doi.org/10.1098/rstb.2010.0415 (2011).
Waksman, S. A. Antagonistic Relations of Microorganisms. Bacteriol Rev. 5, 231–291 (1941).
Traxler, M. F. & Kolter, R. Natural products in soil microbe interactions and evolution. Nat Prod Rep. 32, 956–970, https://doi.org/10.1039/c5np00013k (2015).
Davies, J., Spiegelman, G. B. & Yim, G. The world of subinhibitory antibiotic concentrations. Curr Opin Microbiol. 9, 445–453, https://doi.org/10.1016/j.mib.2006.08.006 (2006).
Wright, G. D. Opportunities for natural products in 21(st) century antibiotic discovery. Nat Prod Rep. 34, 694–701, https://doi.org/10.1039/c7np00019g (2017).
Overmann, J., Abt, B. & Sikorski, J. Present and future of culturing bacteria. Annu Rev Microbiol. 71, 711–730, https://doi.org/10.1146/annurev-micro-090816-093449 (2017).
Gavrish, E. et al. Lassomycin, a ribosomally synthesized cyclic peptide, kills Mycobacterium tuberculosis by targeting the ATP-dependent protease ClpC1P1P2. Chem Biol. 21, 509–518, https://doi.org/10.1016/j.chembiol.2014.01.014 (2014).
Ling, L. L. et al. A new antibiotic kills pathogens without detectable resistance. Nature. 517, 455–459, https://doi.org/10.1038/nature14098 (2015).
Reen, F. J., Romano, S., Dobson, A. D. & O’Gara, F. The sound of silence: Activating silent biosynthetic gene clusters in marine microorganisms. Mar Drugs. 13, 4754–4783, https://doi.org/10.3390/md13084754 (2015).
Bode, H. B., Bethe, B., Höfs, R. & Zeeck, A. Big effects from small changes: possible ways to explore nature’s chemical diversity. Chembiochem 3, 619–627, 10.1002/1439-7633(20020703)3:7<619::AID-CBIC619>3.0.CO;2-9 (2002).
Seyedsayamdost, M. R. High-throughput platform for the discovery of elicitors of silent bacterial gene clusters. Proc Natl Acad Sci USA 111, 7266–7271, https://doi.org/10.1073/pnas.1400019111 (2014).
Pishchany, G. et al. Amycomicin is a potent and specific antibiotic discovered with a targeted interaction screen. Proc Natl Acad Sci USA 115, 10124–10129, https://doi.org/10.1073/pnas.1807613115 (2018).
Weinberg, E. D. Selective inhibition of microbial growth by the incorporation of triphenyl tetrazolium chloride in culture media. J Bacteriol. 66, 240–242 (1953).
Berridge, M. V., Herst, P. M. & Tan, A. S. Tetrazolium dyes as tools in cell biology: new insights into their cellular reduction. Biotechnol Annu Rev. 11, 127–152, https://doi.org/10.1016/S1387-2656(05)11004-7 (2005).
Mattson, A. M., Jensen, C. O. & Dutcher, R. A. Triphenyltetrazolium chloride as a dye for vital tissues. Science. 106, 294–295, https://doi.org/10.1126/science.106.2752.294-a (1947).
Denny, T. P. In Plant-Associated Bacteria (ed S.S. Gnanamanickam.) 573–644. (Springer, 2006).
Clough, S. J., Schell, M. A. & Denny, T. P. Evidence for involvement of a volatile extracellular factor in Pseudomonas solanacearum virulence gene expression. Mol Plant-Microbe In. 7, 621–630, https://doi.org/10.1094/mpmi-7-0621 (1994).
Lemessa, F. & Zeller, W. Screening rhizobacteria for biological control of Ralstonia solanacearum in Ethiopia. Biol Control. 42, 336–344 (2007).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 13, 2498–2504, https://doi.org/10.1101/gr.1239303 (2003).
Wang, M. et al. Sharing and community curation of mass spectrometry data with Global Natural Products Social Molecular Networking. Nat Biotechnol. 34, 828–837, https://doi.org/10.1038/nbt.3597 (2016).
Aleti, G., Sessitsch, A. & Brader, G. Genome mining: Prediction of lipopeptides and polyketides from Bacillus and related Firmicutes. Comput Struct Biotechnol J. 13, 192–203, https://doi.org/10.1016/j.csbj.2015.03.003 (2015).
Hamdache, A., Lamarti, A., Aleu, J. & Collado, I. G. Non-peptide metabolites from the genus Bacillus. J Nat Prod. 74, 893–899, https://doi.org/10.1021/np100853e (2011).
Wilson, M. C. et al. An environmental bacterial taxon with a large and distinct metabolic repertoire. Nature. 506, 58–62, https://doi.org/10.1038/nature12959 (2014).
Rutledge, P. J. & Challis, G. L. Discovery of microbial natural products by activation of silent biosynthetic gene clusters. Nat Rev Microbiol. 13, 509–523, https://doi.org/10.1038/nrmicro3496 (2015).
Zipperer, A. et al. Human commensals producing a novel antibiotic impair pathogen colonization. Nature. 535, 511–516, https://doi.org/10.1038/nature18634 (2016).
Smith, J. J. & McFeters, G. A. Mechanisms of INT (2-(4-iodophenyl)-3-(4-nitrophenyl)-5-phenyl tetrazolium chloride), and CTC (5-cyano-2,3-ditolyl tetrazolium chloride) reduction in Escherichia coli K-12. J. Microbiol Methods. 29, 161–175, https://doi.org/10.1016/S0167-7012(97)00036-5 (1997).
Zhao, X. & Drlica, K. Reactive oxygen species and the bacterial response to lethal stress. Curr Opin Microbiol. 21, 1–6, https://doi.org/10.1016/j.mib.2014.06.008 (2014).
Yeo, I. C., Lee, N. K., Yang, B. W. & Hahm, Y. T. RNA-seq analysis of antibiotic-producing Bacillus subtilis SC-8 in response to signal peptide PapR of Bacillus cereus. Appl Biochem Biotechnol. 172, 580–594, https://doi.org/10.1007/s12010-013-0516-4 (2014).
Xie, S. et al. Transcriptome profiling of Bacillus subtilis OKB105 in response to rice seedlings. BMC Microbiol. 15, 21, https://doi.org/10.1186/s12866-015-0353-4 (2015).
Andrade-Domínguez, A., Salazar, E., Vargas-Lagunas, M. E. C., Kolter, R. & Encarnación, S. Eco-evolutionary feedbacks drive species interactions. ISME J. 8, 1041–1054, https://doi.org/10.1038/ismej.2013.208 (2014).
O’Donovan, G. A. & Neuhard, J. Pyrimidine metabolism in microorganisms. Bacteriol Rev. 34, 278–343 (1970).
Villegas-Escobar, V. et al. Lipopeptides from Bacillus sp. EA-CB0959: Active metabolites responsible for in vitro and in vivo control of Ralstonia solanacearum. Biol Control. 125, 20–28, https://doi.org/10.1016/j.biocontrol.2018.06.005 (2018).
Denny, T. P., Makini, F. W. & Brumbley, S. M. Characterization of Pseudomonas solanacearum Tn5 mutants deficient in extracellular polysaccharide. Mol Plant Microbe In. 1, 215–223 (1988).
Boucher, C. A., Barberis, P. A., Trigalet, A. P. & Demery, D. A. Transposon mutagenesis of Pseudomonas solanacearum isolation of tn-5-induced avirulent mutants. J. Gen Microbiol. 131, 2449–2458 (1985).
Kelman, A. The relationship of pathogenicity of Pseudomonas solanacearum to colony appearance in a tetrazolium medium. Phytopathology. 44, 693–695 (1954).
Balouiri, M., Sadiki, M. & Ibnsouda, S. K. Methods for in vitro evaluating antimicrobial activity: A review. J Pharm Anal. 6, 71–79, https://doi.org/10.1016/j.jpha.2015.11.005 (2016).
Trapnell, C. et al. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat Protoc. 7, 562–578, https://doi.org/10.1038/nprot.2012.016 (2012).
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol 11, R106, https://doi.org/10.1186/gb-2010-11-10-r106 (2010).
Magoc, T., Wood, D. & Salzberg, S. L. EDGE-pro: Estimated degree of gene expression in prokaryotic genomes. Evol Bioinform Online. 9, 127–136, https://doi.org/10.4137/EBO.S11250 (2013).
nanoString Technologies. in MAN-C0011-04 (ed. nanoString Technologies) (2017).
Brumbaugh, C. D., Kim, H. J., Giovacchini, M. & Pourmand, N. NanoStriDE: normalization and differential expression analysis of NanoString nCounter data. BMC Bioinformatics. 12, 479, https://doi.org/10.1186/1471-2105-12-479 (2011).
Liu, W. T. et al. Imaging mass spectrometry of intraspecies metabolic exchange revealed the cannibalistic factors of Bacillus subtilis. Proc Natl Acad Sci USA 107, 16286–16290, https://doi.org/10.1073/pnas.1008368107 (2010).
Watrous, J. et al. Mass spectral molecular networking of living microbial colonies. Proc Natl Acad Sci USA 109, E1743–1752, https://doi.org/10.1073/pnas.1203689109 (2012).
Medema, M. H. et al. antiSMASH: rapid identification, annotation and analysis of secondary metabolite biosynthesis gene clusters in bacterial and fungal genome sequences. Nucleic Acids Res. 39, W339–346, https://doi.org/10.1093/nar/gkr466 (2011).
Pence, H. E. & Williams, A. ChemSpider: An Online Chemical Information Resource. J Chem Educ. 87, 1123–1124, https://doi.org/10.1021/ed100697w (2010).
Kim, S. et al. PubChem substance and compound databases. Nucleic Acids Res. 44, D1202–1213, https://doi.org/10.1093/nar/gkv951 (2016).
R: A language and environment for statistical computing [Online access: http://www.R-project.org/] (Vienna, Austria, 2014).
Acknowledgements
Colciencias Scholarship for doctoral studies call 567 of 2012 for Laura Sierra’s support and tuition; Universidad EAFIT, Association of Banana Producers of Colombia AUGURA and Colciencias (contract number 0555–2013 and FP44842-473-2016) for the financial support of this research. RutaN international Mobility grant call of 2016. Dr. Roberto Kolter for allowing the summer research visit at his laboratory; Nick Lyons (Harvard medical School, Kolter Lab), for his advice on selecting and providing B. subtilis NCIB 3610 mutant strains, Nanostring sets and obtained data analysis; Nicolás Franco-Sierra for his support with RNA-seq and chemical compounds data mining bioinformatic tools and tests; Merlin Andrea Valencia for her support with candidate compounds testing. This research was made possible by the Subscribe Contract Number 166 and 139 with the Ministerio de Medio Ambiente y Desarrollo Territorial of Colombia in the categories “Contrato de Acceso a Recursos Genéticos y Productos Derivados para Investigación Científica” and “Contrato de Acceso a Recursos Genéticos y Productos Derivados con Fines Comerciales” respectively.
Author information
Authors and Affiliations
Contributions
Laura Sierra-Zapata, performed the experiments, analysis, rationale and strategy of the research outline and wrote the manuscript. Javier C. Álvarez and Magally Romero-Tabarez were members of Laura’s advisory committee and provided fundamental advice and strategy to biochemical test and transcriptomic analysis, respectively. Mark.W Silby was the advisor during all transcriptomic experimental data acquisition, analysis and rationale. Matthew F. Traxler, Scott W. Behie and Rita de Cassia Pessotti provided the strategy, training, analysis of data acquired and rationale of metabolomics analysis, as well being part of the manuscript writing. Valeska Villegas-Escobar was the discoverer of the phenomenon, strategist of the general outline of the research along with Laura, wrote the manuscript as well as being the principal investigator (PI) and advisor of Laura’s doctoral thesis.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sierra-Zapata, L., Álvarez, J.C., Romero-Tabarez, M. et al. Inducible Antibacterial Activity in the Bacillales by Triphenyl Tetrazolium Chloride. Sci Rep 10, 5563 (2020). https://doi.org/10.1038/s41598-020-62236-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-62236-z
This article is cited by
-
Towards the sustainable discovery and development of new antibiotics
Nature Reviews Chemistry (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.