Abstract
The black and gold howler monkey (Alouatta caraya) is a neotropical primate threatened by habitat loss and capture for illegal trade in Argentina. Using multilocus microsatellite genotypes from 178 A. caraya individuals sampled from 15 localities in Argentina, we built a genotype reference database (GRDB). Bayesian assignment methods applied to the GRDB allowed us to correctly re-assign 73% of individuals to their true location of origin and 93.3% to their cluster of origin. We used the GRDB to assign 22 confiscated individuals (17 of which were reintroduced), and 3 corpses to both localities and clusters of origin. We assigned with a probability >70% the locality of origin of 14 individuals and the cluster of origin of 21. We found that most of the confiscated individuals were assigned to one cluster (F-Ch-C) and two localities included in the GRDB, suggesting that trafficked A. caraya primarily originated in this area. Our results reveal that only 4 of 17 reintroduced individuals were released in sites corresponding to their cluster of origin. Our findings illustrate the applicability of genotype databases for inferring hotspots of illegal capture and for guiding future reintroduction efforts, both of which are essential elements of species protection and recovery programs.
Similar content being viewed by others
Introduction
Similar to other countries, wildlife illegal hunting and trade are threats to Argentinian wildlife. Confiscated and surrendered animals from trafficking are transported to rehabilitation centres, and the return of these confiscated animals to the wild receives strong support from the public. Ideally, trafficked animals would be reintroduced into the population that they were extracted from or translocated to another suitable site within the species’ original range. Although translocations are considered a good option for conservation and a solution to trafficked animals1, they might be detrimental for the animal and/or the environment if choices over where to return individuals to their natural habitats are not properly based on scientific evidence2,3,4. For example, when significant genetic structure exists within the species in question, translocations may inadvertently lead to admixture of distinct evolutionary lineages and act to homogenize existing diversity and biogeographic patterns instead of protect them5. However, the genetic consequences of translocations have seldom been studied6, and in Latin America, the development of biodiversity management and conservation plans as part of public policy have not yet taken advantage of newly developed genetic techniques to inform translocation policy and decisions7. Conservation genetics can help strengthen the links between scientists and decision makers and improve reintroduction and translocation policy.
Molecular genetic studies using individuals of known origin allow researchers to calculate levels of differentiation among populations and to assess population structure within a species in the wild8,9,10. Based on this, the populations studied may be grouped into clusters based on genetic similarity. Once such an assessment of differentiation between populations/localities or clusters is obtained, genetic assignment analysis of individuals of unknown provenience can be performed to identify the likely locality or cluster to which they belong – i.e., to infer their geographical sites of origin. This identification, made in advance of any translocation attempt, enables researchers to estimate the genetic diversity introduced into the environment when translocations and/or reintroductions occur. Genetic assignment to a geographic area of origin is possible only when a suitable genotype reference database (GRDB) is available to compare results. The construction of a GRDB represents a requirement and, ideally, should include the largest possible sample across a species’ distribution range. Moreover, the data quality must be extremely reliable, because future determinations will rely on these data.
Geographic origin determination using simple tandem repeats markers (STRs, also known as “microsatellites”) has already been applied to several species, such as bobcats11, tortoises12, African elephants for the ivory trade13,14, mouflons15, bears16, salmon17, timber18, palm trees19, and macaws10. These studies allowed researchers to: (1) accurately discriminate among ivory trafficked from elephants from the four major regions of Africa13,14, (2) identify poaching of the protected Sardinian mouflon, where a suspected poacher found in possession of a carcass initially claimed that it was from a sheep from his flock15, (3) identify farmed Atlantic salmon escapes representing a threat to the genetic integrity of natural populations17, (4) develop a genetic reference database characterising the populations of origin for illegally logged timber, which creates market disadvantages for products from sustainable forestry18, (5) determine that illegally-traded palm species in Brazil were obtained from several sites showing that there is no single target locality used by poachers19, and (6) differentiate among macaws from 3 geographic regions of Brazil and identify the most probable regions of origin of 24 confiscated individuals10. These approaches underscore the importance and potential of molecular analyses for wildlife management and for identification of species to prevent illegal trafficking.
Alouatta caraya is the most commonly trafficked primate species found in the illegal pet trade in Argentina. Trade in this species appears to be internal as there are no records of cross-national trafficking. This trafficking is reflected by the number of individuals confiscated during control and inspection operations20,21,22. Official records from the National Wildlife Surveillance and Control Directorate show that 10 individuals of A. caraya were confiscated and/or voluntarily surrendered in the last 4 years, although the number of unofficial cases is higher. Alouatta caraya is globally categorized as “Near Threatened” by the IUCN23, but is considered “Vulnerable” in the red list of Argentina24 due to population reduction, decreased area of occupancy and/or quality of habitat, increased exploitation due to hunting or illegal traffic (pet trade), and the effects of pathogens and parasites. Currently, Argentina’s native forests are rapidly being replaced as a result of growing urbanization, industrialization, and large-scale clearing for agricultural purposes21,22. This process of transforming native subtropical forests is clearly evident in northern Argentina where five species of primates are found (Alouatta caraya, Alouatta guariba clamitans, Aotus azarai, Sapajus nigritus and Sapajus cay). Of these five, both Alouatta species are of great epidemiological importance since they are highly sensitive to the Yellow Fever virus (YFV). Alouatta caraya show high mortality when infected by the virus. Thus, this species serves as an early epidemiological sentinel, allowing the establishment of control and prevention measures24,25,26,27.
Alouatta caraya has been the focus of several population genetic studies of dispersal patterns, kinship, reproductive success, and phylogeography, among other topics28,29,30,31. In a recent study, ten nuclear microsatellites were used to generate a DNA genetic reference database characterising the southernmost populations of this species9 (Fig. 1). Genetic clustering of the studied populations identified a number of distinct genetic groups considered as clusters or management units (MUs)9. One of these clusters consists of populations inhabiting different localities in the Atlantic Forest in the littoral zone bordering Brazil (Misiones cluster9). As part of the monitoring program for the epidemiological surveillance of YFV and other arboviruses in non-human primates (NHP), the number of genotypes included in the reference database for this area was increased with samples collected during 2017 and 2018.
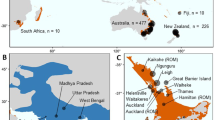
(a) Map of the 15 localities included in the database of A. caraya genotypes in Argentina. Maps show (1 to 10) previously published and (11 to 15) newly sampled localities. Color-coded circles indicate the three genetic clusters identified using the structure analysis in the present study: blue: P-RP cluster, red: F-Ch-C cluster, and green: M-RU cluster. The complete names of sampling sites are listed in Table 1. Black squares indicate the sites were corpses of A. caraya were found. (b) Map showing the distribution range of A. caraya. Black stars show the location of the rescue centres included in this study. Black triangles represent the reintroduction sites.
Here, we present the results of genetic analyses designed to estimate the origin of 22 A. caraya confiscated from the illegal trade and subsequently housed at the Güirá-Oga rescue centre (Güirá-Oga) in Puerto Iguazú, Misiones and the rescue centre Estación Zoológica Experimental Granja La Esmeralda (Esmeralda) in Santa Fe, Argentina, as well as 3 individuals found dead in cities in northern Argentina. We aimed to infer the geographic site of origin of these animals by comparing the specimens’ genetic profiles with the profiles included in a regional GRDB for A. caraya.
This study underscores the use of STR-based genetic databases as a tool for identifying the origin of individual animals, an outcome with application to understanding patterns of illegal wildlife trade, for assessing whether prior translocations in northern Argentina had indeed occurred into likely areas of origin, and to provide a reliable tool for future translocation/reintroduction program protocols.
Results
Genotypes of 143 individuals from 10 localities (Locs) were available from previous studies9,28. To these, we added newly derived and complete genotypes at 10 microsatellite loci obtained from fecal samples of 39 individuals from 5 new localities. Two fecal samples were collected from each of these individuals, and the genotype assignment for each individual at each locus was replicated twice (in the case of heterozygotes) or four times (in case of homozygotes) in order to minimize possible genotyping errors due to allelic dropout. We likewise constructed complete 10-locus genotypes for all 25 additional individual samples of unknown provenience for genetic assignment. The complete Genotype database obtained from all localities and clusters of A. caraya studied in this investigation is available at: https://doi.org/10.5281/zenodo.3660723, and the genotypes of all the assigned individuals are presented as Supplementary Table S1.
We did not observe evidence of scoring errors due to stuttering, large allele dropout, or null alleles for any locus in any population in our screening with the program Micro-Checker v2.2.332.
Using Arlequin v 3.533, we did not observe evidence of linkage between any pair of loci (P > 0.05). Significant deviation from Hardy-Weinberg equilibrium was only detected by GenAlEx v 6.534 and Arlequin v 3.533 softwares for the marker D8S165 in Loc 9 (Piñalito Province Park). Significant evidence of inbreeding (inbreeding coefficient: FIS = 0.29) was already found for this population in a previous study9. The numbers of different alleles, effective and private alleles, observed heterozygosity (Ho), expected heterozygosity (He) and unbiased expected heterozygosity (uHe) are presented in Table 1.
Analysis using the software Structure v.2.3.435 and applying the method described by Evanno36 to a dataset comprising genotypes already collected for A. caraya in Oklander et al.9 plus the new localities sampled here showed that the most likely number of different genetic clusters was three (K = 3, Fig. 2). This new analysis resulted in the disappearance of one previously published cluster (EBCO cluster7, K = 4, Fig. 2), which in our expanded dataset now clusters with the localities of Chaco National Park, Chaco (Loc 4), Guaycolec, Formosa (Loc 5), and San Alonso, Corrientes (Loc 6, Fig. 2). Accordingly, the complete set of 15 locations sampled were grouped into 3 distinct clusters or regions (Fig. 1). Cluster 1 includes two localities (Locs 1 and 2) from Paraguay-Isla Rio Paraná (P-RP); cluster 2 includes four localities (Locs 3 to 6) from Formosa-Chaco-Corrientes (F-Ch-C); and finally, cluster 3 includes nine localities (Locs 7 to 15) from Misiones-Rio Uruguay (M-RU).
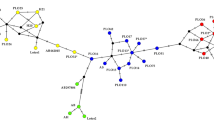
(a) Structure analysis of clusters in A. caraya9 (K = 4): blue: P-RP cluster 1, red: EBCO cluster 2, yellow: F-Ch-C cluster 3, and green: M-RU cluster 4. (b) Structure analysis incorporating samples from the new localities (11 to 15) (K = 3) sampled in the present study: blue: P-RP cluster 1, red: F-Ch-C cluster 2, and green: M-RU cluster 3. Individuals are represented by vertical lines (y-axis) coloured in proportion to their membership coefficients in each cluster and grouped into populations of samples and separated with a black line. Complete names of populations are listed in Table 1.
Aiming to evaluate the efficiency of the software GeneClass237 for assigning individuals to their localities or clusters of origin, we first tested the individuals included in the complete database whose origins was known. The software correctly assigned 73% of individuals in the database (Quality index 68.73%) when separated according to the 15 localities and 93.3% (Quality index 89.23%) when separated according to the three clusters. We then used GeneClass2 to assign the genotypes of each of the 22 confiscated individuals and 3 corpses of unknown origin to both localities and clusters in the GRDB (Table 2). We established a threshold value to consider the assignment to be reliable if it was higher than 70%38,39 (Table 2).
For the 25 individuals analysed, 9 individuals’ assignment values were below the threshold for assignment to a specific locality. The lack of assignment was expected because not all known populations of A. caraya are represented in the database.
Of the 15 confiscated individuals with assignment values above the threshold, almost half of them (seven) most likely came from Loc 3, two from Loc 2, two from Loc 5, two from Loc 6, one from Loc 1 and one from Loc 13. Additionally, of the 3 corpses found in cities, we could only assign one them, found in Loc 13, to Loc 5, therefore showing a different likely site of origin from the place where was found (Table 2).
As a means to identify the approximate origins of the individuals who were not assigned to locations, our next step was to evaluate if these individuals could be assigned to one of the three broader clusters described above (Fig. 2). Assignment to clusters allowed the detection of the possible region of origin of 6 additional individuals, one showed a value very close to the threshold level (69.5%), and 1 remained unassignable (Table 2).
We assigned more than half of the confiscated individuals (14) to 1 of the 3 clusters (F-Ch-C). Four were assigned to the P-PR cluster and 3 to the M-RU cluster.
There was a great deal of variation in the extent to which individuals housed at particular rehabilitation centre actually came from nearby clusters. Specifically, the centre Güirá-Oga, in Misiones province, housed only 2 individuals from the nearby cluster (M-RU where they were reintroduced), while 11 belonged to the cluster F-Ch-C and other 4 to the P-RP clusters (Table 2). Of the 5 confiscated animals housed at Esmeralda, only 4 could be assigned to clusters. Three of them likely came from the nearby F-Ch-C cluster and one from the M-RU cluster. The confiscation sites of these 5 individuals were registered, and the animals that clustered in F-Ch-C (individuals Esmeralda 2, and 4, Supplementary Table S1) were confiscated in Santa Fe province, while the other two (Esmeralda 1 and 5) were confiscated in provinces that do not belong within the natural distribution of A. caraya (Supplementary Table S1). Regarding the 3 corpses found in cities, we could assign one that belonged to a cluster in the same region of Argentina where the city lies, and another one that belonged to a different cluster (F-Ch-C, Table 2). In summary, of all 25 analysed individuals, 15 were inferred to have come from sites within the F-Ch-C cluster.
Finally, of the 17 individuals reintroduced (12 in Misiones and 5 in Santa Fe) only four were reintroduced into a site in their cluster of origin (1 into Misiones, 3 into Santa Fe, Table 2, Supplementary Table S1).
Discussion
Molecular genetic studies have allowed the identification of species confiscated from the illegal wildlife trade, as well as traded animal, timber, and wood products. These applications are even used for species identification from limited samples of body parts (e.g., teeth, feathers, processed tusks), allowing a reliable assessment of the effects of exploitation and the conservation needs of species that would be impossible otherwise. This is the case of the genetic assignment of 28 ivory samples from different elephant populations in Africa14 between 1996 and 2014 that resulted in the identification of two major poaching hotspots, or determination of the species of dried shark fins being sold in Asian and Mediterranean commercial markets40, allowing the monitoring of trade for conservation assessment. Moreover, a study conducted in South Africa revealed that products labelled as “game meat” belonged to domestic species in 76.5% of cases41. Thus, molecular analyses are helpful for poaching detection, traffic route identification, and other crimes involving wildlife.
In a study on tortoises12, where researchers also developed a GRDB and correctly assigned 90% of the individuals in that database to their population of origin, the lack of assignment of confiscated individuals was attributed to the fact that they came from different locations than the sampling sites included in the GRDB. Nevertheless, the researchers were able to determine that all the confiscated individuals came from the same population. This other approach of genetic assignment of living animals (mostly confiscated or in captivity) shows how genetic tools can be used by wildlife managers to identify the most probable regions of origin of individuals as well as to determine the genetic appropriateness of potential recipient populations when designing reintroduction projects. Translocations have been used to mitigate population decline and restore locally extinct populations6. In these cases, genetic data are necessary to guide the selection of populations of origin to which translocated individuals should be released and subsequently evaluate the success of the restoration2,42,43.
In this study, the first application of the GRDB of howler monkeys, our results indicate that the most likely origins of most of the confiscated and surrendered individuals were from the areas around Locs 2 and 3, close to the Argentina-Paraguay border (Fig. 1). Therefore, the largest number of illegally trafficked A. caraya originated in this area.
This area is also the location of northeastern Argentina’s largest cities, Chaco and Corrientes, and National Highway 12, the main highway connecting these cities with Buenos Aires. The illegal sale of A. caraya has been reported at several locations along this highway21,22. This information supports a possible animal trafficking route that begins in northeastern Argentina and ends in Buenos Aires, where the majority of confiscations occur (10 of 22, Supplementary Table S221,22). Importantly, most of the confiscations and surrenders occurred in cities outside the normal distribution of the species (17 of 22, Supplementary Table S2), indicating that these animals are not only opportunistically captured by locals, but that these animals are intentionally transferred to urban centres. This example illustrates how genetic analysis helps trace wildlife trafficking routes and hotspots and thus aids in the planning and implementation of more effective control measures.
On the other hand, 15 of the 17 animals that arrived at the rescue centre Güirá-Oga, in Misiones, were assigned to either to the F-Ch-C or P-RP clusters; twelve of these individuals were subsequently reintroduced near this rescue centre on Isla Palacio, where the endemic genetic variation belongs to the M-RU cluster; thus, translocation and reintroduction resulted in the injection of genetic variation from animals belonging to different genetic clusters. The 5 individuals that arrived at Esmeralda were also reintroduced in a protected area of General Obligado, Santa Fe (Fig. 1). Although nearby localities are not sampled in the database, we would expect that of our sample areas, genetic variation in the liberation area would be most similar to F-Ch-C, similar to the southernmost area of the distribution of A. caraya. Of these five reintroduced individuals, three belonged to the same cluster and only one belonged to the M-RU cluster; therefore, translocation and reintroduction of these animals also introduced non-local genetic variability, albeit in a lower proportion.
Our findings highlight the importance of conducting genetic studies prior to the liberation of rescued animals. These results also raise the concern of establishing rehabilitation centres servicing each of the three described clusters that could be considered as management units for A. caraya if the goal is to reintroduce animals into their native populations. Based on this, future work could consist of the genetic assignment of all the individuals that are going to be part of translocation programs, as well as the extension of the GRDB in areas where releases are scheduled. In this way, the genetic variability that would be entering a reintroduction site could be evaluated and possible restoration analysis would be possible afterwards.
Conservation genetics is generally not yet well integrated with other efforts in conservation policies. In Latin America, the practical application of genetic principles for the management of threatened species and in the development and implementation of conservation plans should be emphasized7. One possible explanation for this disconnect may be that knowledge obtained from scientific research is often not communicated effectively to the field practitioners and/or the authorities who formulate and enact policies.
As shown in the present study, concrete and measurable genetic data represent a very effective tool to help establish and enforce adequate legislation to curb the loss of biodiversity, generate conservation guidelines, and develop population management strategies that include translocation and reintroduction projects.
Methods
Sampling for GRDB
The GRDB for A. caraya used in this study was built using a previously complied database for many locations in Argentina that contained 143 individuals9. The number of individuals in the initial database was increased by adding individuals sampled during a monitoring program for the epidemiological surveillance of YFV and other arboviroses in NHP from several sites in Misiones province in 2017 and 2018 (Table 1 and Fig. 1). We sampled 34 individuals from five newly sampled localities (Locs 11 to 15 in Fig. 1) as well as three individuals from Loc 8 and five individuals from Loc 9, which were resampled localities. In total, these 42 individuals sampled corresponded to 39 new individuals (3 wound up being duplicates of previously sampled in the resampled localities, 1 individual in Loc 8 and 2 in Loc 9; Table 1). Summarizing, we used 178 individuals sampled from 15 localities in the overall GRDB. This included 139 from our previous study9 (we excluded 4 individuals from 2 localities that shared an allele at each locus and were thus considered first-order relatives) and 39 new individuals. Table 1 summarizes the number of individuals, geographical coordinates of sampling locations, and type of samples analysed.
Sampling for genetic assignments
Twenty-five samples were collected for individual genetic assignment. We received hair samples from 17 howler monkeys arriving at Güirá-Oga in 2017. Twelve of these individuals were later reintroduced into a protected area in Isla Palacio at 25°53′32″S, 54°24′38″W (Fig. 1). We also received five tissue samples from monkeys arriving at Esmeralda. All these individuals were later reintroduced into a protected area in General Obligado, Santa Fe at 28°00′12.7″S, 59°32′42.09″W (Fig. 1). A detailed description of these individuals is provided in Supplementary Table S2. Finally, we analysed three tissue samples from monkeys found dead by local authorities in Apóstoles, Posadas, and San Antonio, all in Misiones province in northern Argentina (Fig. 1).
DNA extraction
Two separate faecal samples per individual were stored at room temperature in 50 ml screw-top tubes containing solid NaCl44 until DNA extraction (three months to one year later). DNA was extracted from faeces using the QIAamp DNA Stool Mini Kit (QIAGEN, Valencia, USA), according to the manufacturer’s protocols with slight modifications. DNA was extracted from tissue and hair samples using standard SDS/Proteinase K digestion followed by phenol: chloroform (1 to 1 volume ratio) organic extraction and Microcon P-100 counter-dialysis filters45.
Microsatellite amplification
Ten microsatellites, developed for A. caraya or other primates and previously used in studies of A. caraya population genetics, were amplified from each sample: AC14, AC17, AC45, TGMS1, TGMS2, D8S165, D17S804, LL1118, LL157 and AB0746,47,48,49. Genotyping PCRs were performed in a final volume of 25 µl using 5–10 ng of DNA template for tissue samples or 5 µl of the extraction pool from stool samples and included 20 mM Tris–HCl, 50 mM KCl, 1.5 mM MgCl2, 0.2 mM each dNTP, 1 U of GoTaq DNA polymerase (Promega, Madison, USA), 1 pmol of each forward primer bearing an M13 tail, 4 pmol of each reverse primer, and 4 pmol of M13 labelled with a fluorescent dye (6‐FAM) on its 5′ end following recommendations from previous studies48,50. All amplifications were performed in a Gen Amp ABI 9700 machine (Thermo Fisher, Palo Alto, USA). PCR products labelled with different fluorochromes were combined and the amplicons separated by electrophoresis on an ABI PRISM 310 Genetic Analyzer (Thermo Fisher, Palo Alto, USA). Alleles were manually scored by performing a visual inspection of electropherograms after developing the bin panel for each locus in GeneMapper ID-X v. 1.2 (Thermo Fisher, Palo Alto, USA) using HD400-ROX as internal size standard. For DNA extracted from stool samples, PCR and sizing was repeated twice (in the case of a heterozygous genotype call) or four times (in case of a homozygous genotype call) to minimize possible genotyping errors due to allelic dropout51,52. We recorded an allele only if it was observed at least twice in different amplifications from the same DNA extract. All amplification assays included negative controls.
Ethics statement
This study was carried out in strict accordance with Argentinean laws for research on NHP and following the recommendations of ‘Principles for the Ethical Treatment of Primates’ of the American Society of Primatologists (available at: https://www.asp.org/society/resolutions/EthicalTreatmentOfNonHumanPrimates.cfm). We received specific approval to conduct this study by the Consejo Nacional de Investigaciones Científicas y Técnicas (CONICET) of Argentina (no. 11420110100322CO). Sampling permits for all the locations of the original database previously complied were already published7. Additional specific sampling permits for the new samples presented in this study were obtained from the Ministry of Ecology, Misiones province, Argentina (Permit number: 9910-00086/17) and from the Ministry of Production Santa Fe province, Argentina (Permit Number: GT 13605). Faecal collection was conducted without capturing animals and therefore does not cause any harm to the studied species.
Statistical analysis
Genotypes were screened for null-alleles and to discriminate between errors in allele frequency estimates caused by null-alleles, allele dropout or stutter bands using Micro-Checker v2.2.332. Numbers of different alleles, effective and private alleles, observed heterozygosity (Ho), expected heterozygosity (He), unbiased expected heterozygosity (uHe) and inbreeding coefficient were computed with the software GenAlEx v6.534 for each locus and population. Deviations from Hardy-Weinberg equilibrium (HWE) were assessed by employing an exact test and FIS inbreeding coefficient using Arlequin v 3.5 software33. Allelic richness was calculated for each locus in a population using the equation: equation:
where Ni represents the number of alleles of type i among the 2N genes, and n is sample size, using the software, Fstat v2.9.453.
The new complete set of samples collected was analysed using non-spatial Bayesian clustering with the Structure v.2.3.435 program. A series of 20 independent runs per K (ranging from 2 to 6) was conducted using the admixture model with correlated allele frequencies, sampling locations as a prior (LOCPRIOR), and 500,000 Monte Carlo-Markov iterations after a burn-in of 50,000 replicates. The data analysis procedure was further refined using Clump software54 and a bar plot was constructed with the Disrupt software55. The most likely number of K was identified using the method described by Evanno36.
Assignment tests give the probability of an individual’s multilocus genotype of belonging to reference populations/locations or clusters56.
We used Bayesian methods computed by the software GeneClass237 to assign the origin of confiscated individuals of unknown origin into the 15 potential locations sampled and into the different three clusters described here using a leave one-out procedure, excluding self-assignment and being 0.05 the assignment threshold score (or p-value, set by the program for default). After testing all the combinations of approaches presented by the program, we chose the Bayesian criteria described by Rannala & Mountain57, resulting in a higher quality index and the highest number of correctly assigned individuals when tested against the database. This program is suitable for our study because it does not assume that all populations of origin have been sampled.
For each individual, the best matching location and cluster were sorted and a score in percentage was obtained. The score of an individual, i, in a population, T, is computed as follows:
with Li, T the likelihood value of the individual i in the population T.
Once this probability of assignment of each individual to a certain location or cluster was obtained, we established a threshold value above which assignment was considered reliable. The criteria were that the scoring for a given population was superior to 70%. This threshold was based on our correctly obtained re-assignments of 73% of the database individuals to their locality and in previous works on assignments for fishes and birds33,39.
Data availability
The genotypic reference database (GRDB) for A. caraya presented in this study is available at: https://doi.org/10.5281/zenodo.3660723.
References
Polak, T. & Saltz, D. Reintroduction as an ecosystem restoration technique. Conserv. Biol. 25, 424–424 (2011).
Germano, J. M. et al. Mitigation-driven translocations: are we moving wildlife in the right direction? Front. Ecol. Environ. 13, 100–105 (2015).
Pérez, I. et al. What is wrong with current translocations? A review and a decision‐making proposal. Front. Ecol. Environ. 10, 494–501 (2012).
Seddon, P. J., Strauss, W. M. & Innes J. Animal Translocations: What are they and why do we do them? In Reintroduction Biology (eds. Ewen, J. G., Armstrong, D. P., Parker, K. A. & Seddon, P. J.) 1–32 (Blackwell Publishing Ltd, 2012).
Gippoliti, S., Cotterill, F. P. D., Zinner, D. & Groves, C. P. Poor taxonomy and genetic rescue are possible co-agents of silent extinction and biogeographic homogenization among ungulate mammals. J. Integr. Biogeog. 33, 41–54 (2018).
Jahner, J. et al. The genetic legacy of 50 years of desert bighorn sheep translocations. Evol. Appl. 12, 198–213 (2019).
Torres-Florez, J. P. et al. The coming of age of conservation genetics in Latin America: what has been achieved and what needs to be done. Conserv. Genet. 19, 1–15 (2018).
Cerda, J. M., López, F., Palacios‐Fuentes, P. & Ojeda, F. P. Genetic structure analysis of Girella laevifrons populations in central Chile. J. Appl. Ichthiol. 35, 1147–1153 (2019).
Oklander, L. I., Miño, C. I., Fernández, G., Caputo, M. & Corach, D. Genetic structure in the southernmost populations of black-and-gold howler monkeys (Alouatta caraya) and its conservation implications. Plos One 12, e0185867 (2017).
Presti, F. T., Guedes, N. M. R., Antas, P. T. Z. & Miyaki, C. Y. Population Genetic Structure in Hyacinth Macaws (Anodorhynchus hyacinthinus) and Identification of the Probable Origin of Confiscated Individuals. J. Heredity 106, 491–502 (2015).
Millions, D. G. & Swanson, B. J. An application of Manel’s model: detecting bobcat poaching in Michigan. Wildl. Soc. Bull. 34, 150–155 (2006).
Schwartz, T. S. & Karl, S. A. Population genetic assignment of confiscated gopher tortoises. J. Wildl. Manag. 72, 254–259 (2008).
Wasser, S. K. Assigning African elephant DNA to geographic region of origin: applications to the ivory trade. Proc. Natl. Acad. Sci. USA 101, 14847–14852 (2004).
Wasser, S. K. et al. Combating the illegal trade in African elephant ivory with DNA forensics. Conserv. Biol. 22, 1065–1071 (2008).
Lorenzini, R., Cabras, P., Fanelli, R. & Carboni, G. L. Wildlife molecular forensics: identification of the Sardinian mouflon using STR profiling and the Bayesian assignment test. Forensic Sci. Int. Genet. 5, 345–349 (2011).
Andreassen, R. et al. A forensic DNA profiling system for Northern European brown bears (Ursus arctos). Forensic Sci. Int. Genet. 6, 798–809 (2012).
Glover, K. A., Hansen, M. M. & Skaala, O. Identifying the source of farmed escaped Atlantic salmon (Salmo salar): bayesian clustering analysis increases accuracy of assignment. Aquaculture 290, 37–46 (2009).
Degen, B. et al. Verifying the geographic origin of mahogany (Swietenia macrophylla, King) with DNA-fingerprints. Forensic Sci. Int. Genet. 7, 55–62 (2013).
Nazareno, A. G. & dos Reis, M. S. Where did they come from: genetic diversity and forensic investigation of the threatened palm species Butia eriospatha. Conserv. Genet. 15, 441–452 (2014).
Bertonatti, C. El comercio de primates en la República Argentina. Neotrop. Primates 3, 35–37 (1995).
MAYDS Informe del Estado de Ambiente. https://www.argentina.gob.ar/sites/default/files/mayds_informe_estado_ambiente_2016_baja_1_0.pdf. Presidencia de la Nación, Argentina 2016.
MAYDS MAYDS Informe del Estado de Ambiente. https://www.argentina.gob.ar/sites/default/files/informedelambiente2017.pdf. Presidencia de la Nación, Argentina 2017.
Oklander, L. I. et al. Alouatta caraya. in Categorización 2019 de los mamíferos de Argentina según su riesgo de extinción Lista Roja de los mamíferos de Argentina. http://cma.sarem.org.ar/es/especie-nativa/alouatta-caraya (eds SAREM) (SAYDS – SAREM, 2019).
Almeida, M. A. B. et al. Surveillance for Yellow Fever Virus in Non-Human Primates in Southern Brazil, 2001–2011: A Tool for Prioritizing Human Populations for Vaccination. PLoS NTD 8, e2741 (2014).
Butcher, L. V. Monitoring of Alouatta monkeys as an early warning system in yellow fever surveillance. In Studies on the natural history of yellow fever in Trinidad (ed. Tikasingh, E. S.) 59–62 (Port of Spain, CAREC Monograph Series, 1, 1991).
Rifakis, P. M., Benitez, J. A., De-la-Paz-Pineda, J. & Rodriguez-Morales, A. J. Epizootics of yellow fever in Venezuela (2004–2005): an emerging zoonotic disease. Ann. N. Y. Acad. Sci. 1081, 57–60 (2006).
Strode, G. K. Yellow Fever (McGraw-Hill, 1951).
Oklander, L. I., Kowalewski, M. M. & Corach, D. Genetic consequences of habitat fragmentation in black-and-gold howler (Alouatta caraya) population from Northern Argentina. Int. J. Primatol. 31, 813–832 (2010).
Oklander, L. I., Peker, S. & Kowalewski, M. M. The situation of field primatology in Argentina: recent studies, status and priorities in A Primatologia no Brasil (eds Miranda J. M. D. & Hirano Z. M. B.) 31–50 (UFPR/SBPr, 2011).
Oklander, L. & Corach, D. Kinship and dispersal patterns in Alouatta caraya inhabiting continuous and fragmented habitats of Argentina in Primates in fragments: complexity and resilience (eds Marsh L. K. & Chapmans C. A.) 399–412 (Springer, 2013).
Oklander, L. I., Kowalewski, M. M. & Corach, D. Male reproductive strategies in black and gold howler monkeys (Alouatta caraya). Am. J. Primatol. 76, 43–55 (2014).
Van Oosterhout, C., Hutchinson, W. F., Wills, D. P. M. & Shipley, P. MICRO-CHECKER: Software for identifying and correcting genotyping errors in microsatellite data. Mol. Ecol. Notes 4, 535–538 (2004).
Excoffier, L. & Lischer, H. E. L. ARLEQUIN suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol. Ecol. Resour. 10, 564–567 (2010).
Peakall, R. & Smouse, P. E. GenAlEx 6.5: genetic analysis in Excel. Population genetic software for teaching and research-an update. Bioinformatics 28, 2537–2539 (2012).
Pritchard, J. K., Stephens, M. & Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 155, 945–959 (2000).
Evanno, G., Regnaut, S. & Goudet, J. Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol. Ecol. 14, 2611–2620 (2005).
Piry, S. et al. GENECLASS2: A Software for Genetic Assignment and First-Generation Migrant Detection. J. Heredity 95, 536–539 (2004).
Rolli, J., Girardet, S., Monachon, C. & Richard, C. Microsatellite Analysis of Perch (Perca fluviatilis) and its Genetic Authentication of Geographical Localization. CHIMIA Int. J. Chem. 68, 726–731 (2014).
Dominguez, M., Pizzarello, G., Atencio, M., Scardamaglia, R. & Mahler, B. Genetic assignment and monitoring of yellow cardinals. J. Wildl. Manag. 83, 1336–1344 (2019).
Shivji, M. et al. Genetic Identification of Pelagic Shark Body Parts for Conservation and Trade Monitoring. Conserv. Biol. 16, 1036–1047 (2002).
D’Amato, M. E., Alechine, E., Cloete, K. W., Davison, S. & Corach, D. Where is the game? Wild meat products authentication in South Africa: a case study. Investig. Genet. 4, 6 (2013).
Mijangos, J. L., Pacioni, C., Spencer, P. B. & Craig, M. D. Contribution of genetics to ecological restoration. Mol. Ecol. 24, 22–37 (2015).
Bouzat, J. L. et al. Beyond the beneficial effects of translocations as an effective tool for the genetic restoration of isolated populations. Conser. Genet. 10, 191–201 (2009).
Oklander, L. I., Marino, M., Zunino, G. E. & Corach, D. Preservation and extraction of DNA from feces in howler monkeys (Alouatta caraya). Neotr. Primates 12, 59–63 (2004).
Green, M. R. & Sambrook, J. Molecular Cloning: A Laboratory Manual, 4th ed. (Cold Spring Harbor Laboratory Press, 2012).
Di Fiore, A. & Fleischer, R. C. Microsatellite markers for woolly monkeys (Lagothrix lagotricha) and their amplification in other New World primates (Primates: Platyrrhini). Mol. Ecol. Notes 4, 246–249 (2004).
Gonçalves, E. C., Silva, A., Barbosa, M. S. R. & Schneider, M. P. C. Isolation and characterization of microsatellite loci in Amazonian red‐handed howlers Alouatta belzebul (Primates, Plathyrrini). Mol. Ecol. Notes 4, 406–408 (2004).
Oklander, L. I., Zunino, G. E., Di Fiore, A. & Corach, D. Isolation, characterization and evaluation of 11 autosomal STRs suitable for population studies in black and gold howler monkeys Alouatta caraya. Mol. Ecol. Notes 7, 117–120 (2007).
Tomer, Y., Greenberg, D. A., Concepcion, E., Ban, Y. & Davies, T. F. Thyroglobulin is a thyroid specific gene for the familial autoimmune Thyroid diseases. J. Clin. Endocrinol. Metab. 87, 404–407 (2002).
Schuelke, M. An economic method for the fluorescent labeling of PCR fragments. Nat. Biotechnol. 18, 233–234 (2000).
Surridge, A. K., Smith, A. C., Buchanan-Smith, H. M. & Mundy, N. I. Single-copy nuclear DNA sequences obtained from noninvasively collected primate feces. Am. J. Primatol. 56, 185–90 (2002).
Taberlet, P. et al. Reliable genotyping of samples with very low DNA quantities using PCR. Nucleic Acids Res. 24, 3189–94 (1996).
Goudet, J. Fstat (ver. 2.9.4), a program to estimate and test population genetics parameters. Available from http://www.unil.ch/izea/softwares/fstat.html Updated from Goudet 1995 (2003).
Jakobsson, M. & Rosenberg, N. CLUMPP: A Cluster Matching and Permutation Program for Dealing with Label Switching and Multimodality in Analysis of Population Structure. Bioinformatics 23, 1801–1806 (2007).
Rosenberg, N. A. Distruct: a program for the graphical display of population structure. Mol. Ecol. Notes 4, 137–138 (2004).
Iyengar, A. Forensic DNA analysis for animal protection and biodiversity conservation: a review. J. Nat. Conserv. 22, 195–205 (2014).
Rannala, B. & Mountain, J. L. Detecting immigration by using multilocus genotypes. Proc. Natl. Acad. Sci. USA 94, 5 (1997).
Acknowledgements
We greatly appreciate the collaboration with the Ministerio de Ecología y Recursos Sustentables de la Provincia de Misiones, specifically Patricia Sandoval and Cristina Buhler, and the rescue centers Güirá-Oga and Estación Zoológica Experimental Granja La Esmeralda, particularly Jorge Anfuso, Agustin Anzoátegui, Rocio Rodriguez, Antonio Sciabarrasi Bagilet and Pablo Siroski. We thank Silvana Peker and Ricardo Negreira from Secretaría de Gobierno de Ambiente y Desarrollo Sustentable de la Nación Argentina. We are grateful to Sam Shanee and Anthony Di Fiore for providing comments on the manuscript. This study was supported by CONICET grants to LO and by DNA Fingerprinting Service (SHDG), School of Pharmacy and Biochemistry, University of Buenos Aires, Argentina. LO, MC and DC are members of the Carrera de Investigator (CONICET-Argentina).
Author information
Authors and Affiliations
Contributions
L.I.O. and D.C. originally formulate the idea, L.I.O. and A.S. conducted fieldwork, L.I.O. and M.C. performed laboratory analyses, L.I.O., M.C. and D.C. performed statistical analyses and L.I.O., M.C. and A.S. prepared figures and tables. All authors wrote and reviewed manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Oklander, L.I., Caputo, M., Solari, A. et al. Genetic assignment of illegally trafficked neotropical primates and implications for reintroduction programs. Sci Rep 10, 3676 (2020). https://doi.org/10.1038/s41598-020-60569-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-60569-3
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.