Abstract
The ‘Abundant-Centre Hypothesis’ is a well-established but controversial hypothesis stating that the abundance of a species is highest at the centre of its range and decreases towards the edges, where conditions are unfavourable. As genetic diversity depends on population size, edge populations are expected to show lower intra-population genetic diversity than core populations, while showing high inter-population genetic divergence. Here, the genetic implications of the Abundant-Centre Hypothesis were tested on two coastal mussels from South Africa that disperse by means of planktonic larvae, the native Perna perna and the invasive Mytilus galloprovincialis. Genetic structure was found within P. perna, which, together with evidence from Lagrangian particle simulations, points to significant reductions in gene flow between sites. Despite this, the expected diversity pattern between centre and edge populations was not found for either species. We conclude that the genetic predictions of the Abundant-Centre Hypothesis are unlikely to be met by high-dispersal species with large population sizes, and may only become evident in species with much lower levels of connectivity.
Similar content being viewed by others
Introduction
The Abundant-Centre Hypothesis (ACH) states that species perform better at the centre of their range, where high densities are observed, and that densities decline towards the range edges1,2. Both biotic and abiotic factors determine the success of a species within its realised niche2, and it is expected that population abundance, reproductive output and recruitment will decline towards the periphery of its distribution, due to the less favourable conditions prevailing there2,3. Patterns of population genetic structure across a species’ range are, in turn, expected to be influenced by these ecological effects. Because genetic diversity depends on population size, it is predicted that populations at the periphery of the species’ range will show lower intra-population diversity and higher inter-population divergence when compared to centre populations2,3,4. These differences are attributed to reduced gene flow, bottleneck effects, habitat fragmentation and random genetic drift in smaller populations5. The reduced diversity and increased genetic differentiation of edge populations may translate into lower adaptive potential particularly at the periphery of a species’ range6,7, which suggests that maladaptation may cause these populations to be demographic sinks.
Several recent studies have raised doubts about the generality of these demographic and genetic expectations. A review found that there is limited empirical evidence for the ACH, with support from only 39% of the reviewed studies, which covered a wide range of genetic and demographic predictions derived from the hypothesis8. Similarly, another review assessing population genetic structure across the geographical ranges of 115 species of plants and animals showed that, on average, only 64% of studies detected a decline in genetic diversity towards the range margins, although there was a geographic bias as the majority of studies concerned the northern range limits of northern hemisphere species9. Taken together, these studies show that species abundance, demography and genetic diversity appear to be more variable than predicted by the ACH, as they are determined by the interaction of multiple fluctuating contemporary environmental drivers plus historical effects9,10.
Along the South African coastline, rocky shores are a very common habitat type. They constitute nearly 60% of the coastline and span four major biogeographic regions11,12, making common rocky shore fauna particularly suitable for studying the ACH. We tested the predictions of the ACH for the dominant rocky shore mussels in this region11, the invasive Mytilus galloprovincialis and the native Perna perna. The latter is represented by two distinct regional genetic lineages12, effectively providing edge and centre populations for three model organisms with similar biology. We hypothesised that centre populations of the three mussels would have greater genetic diversity than edge populations and tested this by comparing genetic diversity for the two population types.
Materials and Methods
Study species and their ranges
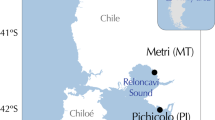
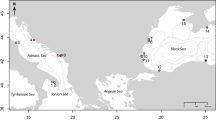
Perna perna has a wide range from the warm-temperate south coast of South Africa to the tropical coastline of southern Mozambique on the east coast11. Due to the cold upwelled water of the Benguela Current on the west coast, P. perna does not occur west of the Cape of Good Hope, but it re-emerges in Namibia, and its range stretches along the west coast of Africa all the way to the Mediterranean Sea13. Previous genetic work using mitochondrial DNA (mtDNA) sequences and microsatellite loci of P. perna identified a phylogeographic break on the south-east coast of South Africa, where the ranges of a western and an eastern lineage overlap along 200 km of coastline, from approximately Port Alfred to Haga Haga14,15 (Fig. 1). A subsequent study showed a non-sister relationship for the two lineages, possibly reflecting an Indo-Pacific origin followed by dispersal into the Atlantic through the Tethys seaway, independent southward expansion along the western and eastern shores of the African continent and recent secondary contact on the south-east coast of South Africa16.
Sampling sites for the western (circles) and eastern (pentangle) mtDNA COI lineages of Perna perna, and of Mytilus galloprovincialis (triangles). Centre (black) and edge (grey) sites are indicated for each lineage/species. The map was created in StepMap (https://www.stepmap.de/) and modified in InkScape 0.92.4 (https://inkscape.org).
The other dominant mussel in South Africa, the Mediterranean mussel Mytilus galloprovincialis, is a particularly successful marine invader that has established itself on all continents except Antarctica17. In South Africa, M. galloprovincialis is found in cooler waters west of the Cape of Good Hope where P. perna is absent. Both species co-exist on the south coast, while only P. perna occurs along the subtropical and tropical east coast.
For both the western and eastern lineages of P. perna, sites on the south-east coast where the ranges of these lineages overlap constitute edge populations, and the same region also constitutes the range periphery of M. galloprovincialis. On the south-west coast, the western lineage of P. perna has a second edge population (Fig. 1 and Table 1).
Assignment of sites as either centre or edge populations was decided a priori based on previous intensive field surveys14,15,18. These identified the area between approximately Port Alfred to Haga Haga as being located in a biogeographical transition zone and thus edge habitat, while Cape Agulhas (Fig. 1) was identified as the location that separated core populations on the south coast from western edge populations.
Estimation of mussel cover
To confirm that pre-defined edge populations had smaller population densities than core populations, percentage mussel cover at specific sites was estimated with ImageJ software19 using unpublished data from earlier fieldwork. Mussel cover was measured between January and April 2013 by haphazardly placing quadrats (25 × 25 cm; N = 20) onto mussel beds. Site M3 (Fig. 1) was surveyed in 2011, and for site P8, which is difficult to access, the survey was conducted at a nearby location (Ballito). No data were available for site P9.
Estimates of cover for the two lineages of P. perna at sites P5 and P6 could only be estimated indirectly as the two lineages are morphologically indistinguishable without genetic identification. Thus, the percentage cover of each lineage was estimated by using the relative proportions of individuals from each lineage from a previous mtDNA-based population genetic study14.
Genetic analyses
Genetic diversity in P. perna was assessed using data from mitochondrial DNA sequences (COI) and a suite of microsatellites loci. Only microsatellites were assessed for the invasive M. galloprovincialis since its COI diversity is very low14. Moreover, mitochondrial DNA evolves comparatively slowly and would only reveal historical demographic information from the species’ native European habitat14.
Fifty adult mussels (shell length > 4 cm) were collected from each site in November 2014, and mantle tissue weighing 20–30 mg was dissected from each mussel, preserved in 100% ethanol and stored at –20 °C. Specimens of P. perna and M. galloprovincialis were collected from edge and centre populations as shown in Fig. 1 and Table 1.
Whole genomic DNA was extracted using the CTAB method20. From each specimen, a piece of tissue the size of a match head was excised and placed into a labelled 1.5 ml Eppendorf tube and left to dry. One ml of CTAB buffer (2 g of cetyltrimethyl ammonium bromide powder, 1 g of polyvinyl pyrrolidone powder, 100 ml of 1 M Tris, 28 ml of 5 M NaCl, 4 ml of 0.5 M EDTA and filled up to 100 ml with deionised water) and 10 µl of Proteinase K were then added. The tubes were kept at 55 °C for three hours (or at 37 °C overnight) and occasionally shaken until tissues had dissolved completely, whereupon 500 µl of a 1:24 isoamylalcohol-chloroform mixture was added, and tubes shaken vigorously for 30 seconds. The tubes were then centrifuged at 13 000 rpm for 5 min, whereafter 500 µl of the supernatant was transferred to newly labelled 1.5 ml Eppendorf tubes. Next, 500 µl of isopropanol was added to the tubes, shaken, and the mixture was centrifuged for 10 min at 13 000 rpm. Tubes were then decanted, and DNA pellets washed twice with 700 µl of 70% ethanol, and centrifuged at 13 000 rpm for 10 minutes. All liquid was removed from the tubes, and DNA pellets were left to dry completely at room temperature for 30 min. 20 µl of dilute TE (Tris-EDTA) buffer (0.1 mM Tris-Cl [pH 7.6], 0.001 mM EDTA) was added to each Eppendorf tube, vortexed and stored in the freezer at −20 °C.
The mtDNA was amplified with the universal primers LCO1490 (5′-GGT CAA CAA ATC ATA AAG ATA TTGG -3′) and HCO2198 (5′-TAA ACT TCA GGG TGA CCA AAA AAT CA-3′)21. PCR reactions were performed in reaction volumes of 20 µl containing the following reagents: 2 µl of DNA template, 1.2 µl of 25 mM MgCl2 and 2 µl of 10x PCR buffer (Promega), 0.64 µl of 1 mM dNTP mixture (Sigma-Aldrich), 0.5 µl of forward and reverse primers (10 mM), 0.24 µl of BSA, 0.16 µl of Super-Therm Taq polymerase (5 units/ml, Separation Scientific SA), and 12.7 µl of distilled water. PCR amplifications were performed on a Multigene Optimax Thermal Cycler (Labnet) using the following profile: an initial denaturation step (94 °C for 2 min), 40 cycles of denaturation (94 °C for 30 sec), annealing (50 °C for 40 sec), and extension (72 °C for 1 min), and a final extension step (72 °C for 10 min). Gel electrophoresis was performed to confirm DNA amplification. PCR products were then purified with the Wizard SV Gel and PCR Clean-Up System (Promega), cycle sequenced both in the forward and reverse directions using Big Dye Terminator v3.1 Cycle Sequencing Kit (Applied Biosystems), and visualized on an ABI 3730 Genetic Analyser.
Microsatellite genotyping of P. perna and M. galloprovincialis was carried out using a set of 10 and nine loci, respectively, with markers previously reported to conform to the expectations of Linkage Disequilibrium and Hardy-Weinberg equilibrium (Table S1). A reaction volume of 18 µl of a PCR solution was used. Each tube contained the following reagents: 2 µl DNA template, 1.5 µl of 25 mM MgCl2, 2 µl of 10x PCR buffer (Promega), 2 µl 1 µM dNTP mixture (Sigma-Aldrich), 0.4 µl of forward and reverse primers (10 mM), as well as 0.6 µl of universal fluorescently labelled M13 primer, 0.24 µl of BSA, 8.7 µl of deionised water, and 0.16 µl of Super-Therm Taq polymerase (5 units/ml, Separation Scientific SA). The PCR amplification of P. perna was performed on a GeneAmp 9700 thermocycler (PE Applied Biosystems) under the following conditions: an initial denaturation step (95 °C for 5 min), 35 cycles of denaturation (95 °C for 30 sec), annealing at a primer-specific annealing temperature (Table S1) for 30 sec and extension (72 °C for 40 sec), and a final extension step (72 °C for 20 min). For M. galloprovincialis, amplification was performed under the following conditions: an initial denaturation step (94 °C for 2 min), 30 cycles of denaturation (92 °C for 1 min), annealing at a primer-specific annealing temperature (Table S1) for 1 min and extension (72 °C for 30 sec), followed by a final extension at 72 °C for 10 min.
A post-PCR multiplexing technique was employed for both species, where loci were amplified separately and then pooled for genotyping, as previously described for P. perna22. The PCR products were analysed using an ABI PRISM 3730 DNA Analyser (Applied Biosystems), with GeneScan Liz 500 (Applied Biosystems) as size standard.
Mitochondrial DNA sequences of P. perna were aligned and edited in MEGA723. For the microsatellite data, GENEIOUS microsatellite plug-in 1.424 was used to analyse, score and bin alleles. Two researchers confirmed the scoring of alleles independently. The quality of the microsatellite data was assessed by exploring the occurrence of null alleles and by calculating departures from Linkage Disequilibrium (LD) and Hardy-Weinberg equilibrium (HWE). The proportion of null alleles per locus was calculated in FreeNA25, while LD and HWE were calculated in Arlequin v3.5.2.226. Likelihood ratio tests for LD between all pairs of loci were calculated for each sampling location, with 1000 permutations to test for significance, two initial conditions for the expectation-maximisation algorithm, and Bonferroni correction27 for multiple tests. Exact tests for HWE were conducted by specifying 106 steps in the Markov chain and 105 dememorisation steps.
Genetic differentiation
The evolutionary relationships between the COI sequences of P. perna were visualized by constructing a median-joining network27 in PopART 1.728. Samples that were not part of the species’ dominant lineage in the region where they were found were excluded from subsequent analyses (1 individual from site P2 and 1 individual from site P3); in both cases these were eastern haplotypes that were present at sites that were dominated by western haplotypes and thus reflect long-distance migration. To assess genetic differentiation within the ranges of individual lineages that may indicate reduced gene flow between centre and edge sites, population pairwise ΦPT was calculated using GenAlEx 6.50129, and Bonferroni correction was applied to account for multiple comparisons.
STRUCTURE 2.3.430 was used to assess genetic structuring in the microsatellite data of both P. perna and M. galloprovincialis, with both the admixture and ‘locprior’ models invoked to infer the ancestry of individuals. The program was run on the Lengau supercomputer at the CSIR Centre for High Performance Computing in Rondebosch, South Africa, by specifying up to five genetic clusters (K = 1–5) with a burn-in of 1,000,000 iterations followed by 10,000,000 MCMC iterations. Each run was repeated ten times with different starting seeds to confirm that the program was run sufficiently long for the results to be consistent. Evanno’s ΔK31 was calculated with STRUCTURE HARVESTER32 to identify the best-supported number of distinct genetic clusters comprising each species. After confirming with CLUMPAK33 that all ten independent runs per K converged on the same solution, a barplot was created from one of the Q-matrices for P. perna with STRUCTURE PLOT 2.034. Further regional genetic subdivision in P. perna was explored by analysing western and eastern sites separately, using the same settings as above. GenAlEx was used to calculate pairwise FST values, as well as G”ST, a genetic structure statistic similar to FST that is particularly suitable for microsatellites35. Bonferroni correction was applied as described above. The program FreeNA was used to determine whether null alleles affected pairwise FST, with significant differences between corrected and uncorrected estimates determined by calculating 95% confidence intervals that were based on 1000 bootstrap replications.
Evolutionary relationships between the lineages were also visualized using a minimum spanning network that was obtained from the R package Poppr 2.3.036. Average Bruvo’s distance37 over all loci in a population was calculated, the model for missing data was set to average addition/loss and the repeat lengths of each locus were entered.
Estimates of genetic diversity
For the COI data, we used Arlequin to calculate site-specific haplotype diversity h38 and nucleotide diversity π39. When h is high, this implies that a population has a large number of haplotypes, whereas a high π indicates that the haplotypes present in a population are very different from each other. Both are typical of large populations, and it was expected that h and π would be greater for centre sites than for edge sites. To determine whether these diversity indices differed between pairs of sites, we used the R package genetic_diversity_diffs 1.0.540 and specified 1000 iterations to calculate Bonferroni-corrected P-values.
To assess genetic diversity for the microsatellite data, we calculated allelic richness (Ar) in the R package diveRsity41. Compared to the more commonly used expected heterozygosity (HE), this statistic is more likely to be affected by stochastic processes in small populations at the range edge, and for that reason is more sensitive in detecting differences between centre and edge sites9. Significant differences in the magnitude of Ar between sites were identified by calculating 95% confidence intervals based on 1000 bootstrap replications, with non-overlapping confidence intervals between two sites implying significant differences. As it is not possible to calculate confidence intervals for Ar in diveRsity when rarefaction (correction for differences in sample size) is applied, we reduced the number of individuals at all sites to the number at the site with the smallest sample size by excluding individuals with the most missing data, which was 41 individuals for P. perna. No reduction was necessary for M. galloprovincialis because all sites had 48 individuals.
Lagrangian particle simulations
To provide an independent estimate of the dispersal potential of P. perna and M. galloprovincialis, Lagrangian particle simulation was performed. The simulation followed implementations used previously18,42,43, and used data assembled from the Hybrid Coordinate Ocean Model (HYCOM), a product delivering ocean current fields on a daily basis (spatial resolution of 0.08°, approx. 6–9 km). This is a model forced by heat flux, precipitation, wind stress and wind speed that is able to resolve meandering currents, eddies, filaments and fronts44, i.e., important mesoscale oceanographic processes required to accurately simulate passively dispersing larvae18,45.
The region of simulation comprised ~2750 km of coastline that slightly exceeded the range from which genetic samples were obtained, from St. Helena Bay (South Africa; 32.75° S, 17.85° E) in the west to Ponta Dobela (Mozambique; 26.25° S, 32.95° E) in the east. A high-resolution polygon representing global landmasses46 was used to define source/sink locations 1 km apart. From these points, individual particles simulating pelagic mussel larvae were released on a daily basis, and allowed to drift at the surface of the virtual environment for up to 30 days18 until they eventually ended up on the shore, or were lost in the open ocean. The position of each particle was determined every hour using the bilinear interpolation of HYCOM’s ocean velocity fields. Individual trajectories were aggregated to matrices representing the probability of connectivity between every pair of coastal locations, by dividing the number of times a particle released from location i reached location j, by the total number of particles released from location i.
To account for inter-annual variability in ocean flows, the simulations were run individually for a period of 10 years (2003 to 2012), and a final asymmetrical connectivity matrix between all pairs of coastal locations was produced by averaging the annual matrices. Overall, a total of 2,697,350 particles were released.
The potential connectivity between the locations sampled for genetics was determined with direct probability estimates from the final asymmetrical matrix, and by considering stepping-stone probability estimates. For the latter approach, network analysis was implemented, with vertices being locations and edges the probabilities of asymmetrical connectivity47. Floyd–Warshall’s algorithm, which minimises the overall sum of log‐transformed probabilities, was used to estimate the shortest path between sampled locations, from which probabilities of connectivity were determined by a product function. Finally, clusters of locations with higher connectedness were determined by finding the optimal partitioning for maximizing the modularity index (i.e., goodness-of-fit) over all possible partitions. Dispersal simulations and network analyses were performed using the R packages igraph48, dismo v1.1-449, parallel v3.6.1 (R-core), raster v0.4650, and vegan v2.5-651.
Ethical approval and informed consent
A research permit (RES2014/12) issued jointly by the Department of Environmental Affairs and the Department of Agriculture, Forestry and Fisheries was used to sample mussels from the South African coastline.
Results
Estimation of mussel cover
Mussel cover was on average 0.13% at edge sites of the western lineage, increasing to 58.47% at centre sites (Table 2). Centre sites of the eastern lineage had even greater mussel cover, on average 72.59%. Based on a neighbour-joining phylogram of mtDNA sequences14, the two lineages were equally represented at site P5, while at site P6, 30% of the mussels were assigned to the western and 70% to the eastern lineage. Given mussel cover of 62.2% and 52.1% at sites P5 and P6 respectively, the resulting cover for each lineage was estimated to be 33.1% (P5) and 15.6% (P6) for the western lineage, and 33.1% (P5) and 36.4% (P6) for the eastern lineage. Centre locations of Mytilus galloprovincialis also had higher cover than edge locations (58.3% and 0.23%, respectively).
Genetic analyses
Genetic structure
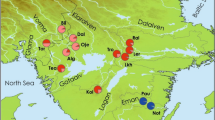
The two COI lineages of Perna perna were recovered as distinct genetic clusters (Fig. 2), with some evidence for migration (some typical western haplotypes were also found in the east, and eastern haplotypes were found in the west). Eastern edge individuals clustered throughout the network, but in both lineages, those individuals were limited to certain clusters.
A median-joining network constructed from COI haplotypes of Perna perna with PopART 1.7 (http://popart.otago.ac.nz/). Haplotypes are represented by circles whose size is proportional to their frequency. Vertical lines represent the number of nucleotide differences between haplotypes, and colours represent sampling sites.
For both lineages of P. perna, significant ΦPT values were found for several pairs of sites (Tables S2 and S3), but these did not conform to the pre-defined centre and edge groupings. In the western lineage, there was a clear geographical division into western sites (edge sites P1, P2 and centre site P3) vs. eastern sites (centre site P4 and edge site P5). In the eastern lineage, the northernmost site (centre site P9) was distinct from all other sites.
FreeNA analyses indicated that null alleles were quite common for some microsatellite loci (Tables S4 and S5), although this had no significant effect on estimates of genetic structure (see next paragraph). Tests for Linkage Disequilibrium (LD) between microsatellite loci of P. perna were significant in several cases, although no pair of loci was consistently in LD at all sites (Table S6). In contrast, very few tests of M. galloprovincialis loci were significant (Table S7). Departures from Hardy-Weinberg equilibrium, which were mostly heterozygote deficits, were also much more common for P. perna (Table S8) than for M. galloprovincialis (Table S9). Such results are often found in studies on high-dispersal marine invertebrates52. Although the difference between the two mussel species is difficult to explain, it may be related to complex small-scale selective forces in P. perna that result in skewed sex ratios, reducing the chances of random mating53 and potentially producing localized Wahlund effects54 where migrants from adjacent but different habitats have reduced fitness55. As analyses of microsatellite data produced results that were very similar to those from the mtDNA data, and null alleles did not significantly affect genetic structure (see next paragraph), we decided not to exclude any loci from subsequent analyses.
FST and G”ST values (Table S10) were significantly different between most pairs of sites even after Bonferroni correction. The only pairs of sites showing non-significant differences were found in the same region (west or east), but no clear trend was evident between centre and edge sites within a particular region. A comparison of FST values corrected for null alleles with the uncorrected FST values showed that these were not significantly different, as 95% confidence intervals overlapped in all cases (Fig. S1).
STRUCTURE analyses revealed that the best number of distinct clusters based on Evanno’s ΔK was not 2 (western and eastern lineage) but 3 (Fig. S2), with most individuals from site P5 being part of a distinct cluster (Fig. 3a). STRUCTURE analyses using only western sites (P1–P4) or eastern sites (P6–P9) further revealed that the westernmost edge site (P1) was distinct from other western sites (P2-P4), while no further subdivision was evident in the east (Fig. 3b).
Barplots of Perna perna microsatellite data constructed using STRUCTURE 2.3.4. Each vertical bar represents a single individual, and colours indicate the association of these with genetic clusters; (a) result for all nine sites (P1-P9) and three clusters; (b) results for western and eastern sites separately for two clusters, excluding site P5. Association of individuals with centre and edge sites is shown below the barplots. The figures were constructed in STRUCTURE PLOT 2.0 (http://omicsspeaks.com/strplot2/) and modified in InkScape 0.92.4 (https://inkscape.org).
The minimum-spanning network for the microsatellite data of P. perna (Fig. S3) did not recover distinct western and eastern evolutionary lineages, although trends are evident (e.g., the cluster on the left mostly comprises eastern sites, and that on the top right has particularly large numbers of western sites).
Evanno’s ΔK method supported two genetic clusters of M. galloprovincialis (Fig. S4). However, this seems to be a result of the method being unable to provide a result for K = 1. A STRUCTURE barplot did not identify any spatial genetic structure but assigned each individual to both clusters (Fig. S4), and there is thus no evidence for more than one genetic cluster. This conclusion is supported by the minimum-spanning network for this species, which showed no spatial trends (Fig. S5), and by the fact that, with a single exception, FST and G”ST values were not significantly different for all pairs of sites (Table S11). Again, FST values corrected for null alleles were not significantly different from uncorrected values (Fig. S1).
Comparisons of genetic diversity indices
On the basis of the genetic structure results for P. perna, several adjustments to the a priori assignment of sites to either centre or edge populations shown in Fig. 1 were considered necessary. First, site P5 could not be clearly assigned to either western or eastern lineages, and for that reason, diversity indices were either reported separately for the individuals associated with each lineage (mtDNA data), or it was treated as a distinct site (microsatellite data). Second, site P6, which was previously identified as being located in a contact area of the two lineages14 (Fig. 1, Table 2), an arrangement that was also supported by the Lagrangian modelling results (see below), had only eastern COI haplotypes in the present study. It was also not clearly distinct from eastern sites based on the microsatellite data (Fig. 3b), and for that reason was treated as an edge population of the eastern lineage.
For the western lineage of P. perna, significant differences in haplotype diversity (h) were found in three cases (Table S12), but these were no longer significant after Bonferroni correction and thus likely false positives (Fig. 4a). Nucleotide diversity (π) was mostly lower for the eastern lineage than for the western lineage, but no significant differences between centre and edge sites of the same lineage were identified (Fig. 4b, Table S13).
Comparisons of COI-based genetic diversity indices between centre (grey bars) and edge (white bars) sites of the western and eastern lineages of the brown mussel, Perna perna: a) haplotype diversity h and b) nucleotide diversity π. Values are means and error bars are standard deviations. Please see Tables S6 and S7 for details.
Allelic richness (Ar) based on the microsatellite data is reported together for the two lineages of P. perna because they were not as clearly differentiated as was the case for the mtDNA data (Figs. 3 and S3). This statistic was distinctly larger for the native P. perna than for the invasive M. galloprovincialis (Fig. 5). In this case, there were no clear differences in genetic diversity between the two lineages of P. perna. Ninety-five percent confidence intervals overlapped for all the centre and edge sites of both species, which implies no significant differences.
Comparisons of microsatellite-based allelic richness (Ar) between centre (grey bars) and edge (white bars) sites of the western and eastern lineages of the brown mussel, Perna perna and the invasive Mytilus galloprovincialis. Error bars are 95% confidence intervals, and overlap between these indicates that genetic diversity indices are not significantly different. In this case, individuals of P. perna could not always be readily assigned to either the western or the eastern lineage, and site P5 (which was distinct from western and eastern sites, Fig. 3a) is shown in black.
Lagrangian simulations
The particle simulations over the 10-year period showed a sharp decline of potential connectivity with distance (Fig. S6), with high probabilities for larval retention near the parent habitat (matrix diagonals in Fig. 6a) and mean traveled distances of 59.35 ± 69.86 km in 4.97 ± 5.21 days (maximum 1002.81 km; Table S14; Mov S1). When considering direct connectivity, only two pairs of locations sampled for genetics were potentially connected by ocean currents (P2 to P1 and P4 to P3; Fig. 6a, Table S15), but when considering a stepping-stone scenario, all pairs were potentially connected, albeit with skewed probabilities with increasing distance (Table S16). This scenario recovered three main clusters with higher connectedness that, overall, conformed well to the regional clusters identified using genetic methods: a western cluster including locations P1 to P4, as well as M1 and M2; a central cluster including locations P5, P6, M3 and M4; and an eastern cluster including locations P7 to P9 (Fig. 6b). The main differences compared to the genetic results are thus a) the distinctness of the originally defined eastern edge sites (P5 and P6) from eastern centre sites (P7–P9) in P. perna and b) a clear division into centre (M1 and M2) and edge (M3 and M4) sites in M. galloprovincialis.
Pairwise connectivity inferred with Lagrangian particle simulations; (a) connectivity matrix between all coastal locations in South Africa averaged for a 10-year period; (b) connectivity and clustering between locations from which genetic samples were collected (considering the stepping-stone scenario) created in InkScape 0.92.4 (https://inkscape.org). Thicker lines indicate higher probabilities of connectivity (please refer to Tables S15 and S16).
Discussion
Originally developed in the context of population genetics56, the Abundant-Centre Hypothesis (ACH) has subsequently been extended to include ecological aspects of species, such as physiological performance57. Empirical studies do not always support this hypothesis, and it is useful to consider the definition of an edge population. Two definitions have been suggested56: populations whose size fluctuates dramatically and which have a high likelihood of becoming extinct, and populations whose members are sparse and physiologically stressed. Such stress could be due to environmental conditions or competitive interactions58. This makes the important point that marginal populations are not necessarily found at the periphery of a species’ distribution59, and this in turn is compatible with the observation that species’ ranges need not contract from the periphery to the centre60. In terms of testing the ACH, coastlines have the advantage of being essentially one-dimensional, making it relatively easy to identify central and peripheral populations. The South African coastline, apart from being much more linear than many other coastlines, has the additional advantage that it encompasses a wide range of climatic and oceanographic conditions that define at least four marine biogeographic provinces12. These are separated by biogeographical transition zones that represent the range edge in numerous species61 and that are so distinct environmentally that they can drive the evolution of genetic lineages endemic to these transition zones62,63. On this basis, we have made the assumption that populations near the species’ distributional limits are indeed marginal56. The clear spatial arrangement of centre and edge regions theoretically offers an ideal setting that facilitates multiple tests of the ACH. The rejection of the hypothesis in two evolutionary lineages of the native mussel Perna perna and in the invasive Mytilus galloprovincialis provides insights into the conditions that prevent species from developing the genetic patterns expected to result from the ACH.
Genetic structure results for mtDNA and microsatellite data of P. perna were similar in that both markers identified the western and eastern lineages. However, smaller-scale genetic structure did not clearly conform to the pre-defined divisions into center and edge populations, and also differed between markers. The western COI lineage could be further subdivided into a western and an eastern group (with a boundary between sites P3 and P4) whose ranges did not match the biogeography-based division into centre and edge sites, but which may nonetheless reflect a thermal gradient in this area64,65. A similar result was found for the microsatellite data, although the location of the boundary differed (it was located between sites P1 and P2) and may reflect marker-specific differences in lineage sorting66 or selection67. Similarly, the distinctness of the COI haplotypes from the northernmost site (P9), which was not found with microsatellites, matches the location of previously reported phylogeographic breaks that separate subtropical from tropical fauna61,68,69,70, and may thus also reflect differential adaptation across a temperature gradient.
In terms of genetic diversity, there were clear trends between species or lineages, but none were related to differences between centre and edge sites. Genetic diversity indices were higher in western P. perna than in eastern P. perna (mtDNA data), and higher in the microsatellite data of P. perna than in those of M. galloprovincialis. As mentioned earlier, no mtDNA data were generated for the latter species because its genetic diversity is very low14. In addition, given the slow mutation rate of this marker, its genetic diversity likely still reflects the species’ demographic history in the northern hemisphere prior to its introduction to southern Africa. However, the microsatellite data were also considerably less variable than in P. perna, possibly because of a reduction in diversity during the founder event(s) that occurred at the beginning of the species’ invasion of southern Africa in the 1970s.
A reduction in genetic diversity towards the range edge expected under the ACH is predicted on the basis of a combination of smaller population size and increased spatial isolation from the centre of the range71. Hence, even when the abundance of mussels is lower in edge populations, reduced genetic diversity is only expected when migration from the centre is so low that some alleles are so rare in the periphery that they are not represented in a reasonably large genetic sample.
A recent study on two species of the gastropod genus Crepidula with very different dispersal potential identified support for the ACH for both species72. Although the absolute reduction in genetic diversity that was found for marginal populations was relatively small, this implies that dispersal capability is irrelevant. In our case, all three taxa have relatively long-lived planktonic larvae and are expected to have a high capacity for dispersal, although empirical evidence suggests that the numbers of larvae undergoing long-range dispersal on this coast may not be high in terms of ecological connectivity73. Accordingly, previous research using mtDNA sequence data of P. perna identified surprisingly well-developed genetic structure within the western lineage that was driven by coastal topography55 and even resulted in sex-biased genetic structure between bays and the open coast53. In the present study, we identified numerous examples of genetic structure that imply some degree of spatial isolation, but the data from neither genetic marker supported division into groups of centre or edge sites.
The Lagrangian modelling allowed the assignment of locations to three distinct clusters with high within-group and lower between-group connectedness, the clusters largely coinciding with the biogeography-linked regional patterns of genetic structure found for P. perna. Interestingly, the modelling results supported the distinctness of the originally-defined edge populations of both species, even though their distinctness was not strongly supported by the present genetic data. While a 30-day dispersal period may potentially generate connectivity over a distance of up to 1000 km, connectivity events on average were in the order of only ~60 km in ~5 days43. Nevertheless, the possibility of year-to-year stepping-stone connectivity47, together with the mussels’ high reproductive output and long pelagic larval phases, may facilitate the spread of genes across each species’ entire range.
The rejection of the genetic expectations of the ACH, despite: (a) lower population densities of edge populations, (b) clear departures from the expectations of panmixia in P. perna, and (c) reduced larval connectivity between some of the originally-defined centre and edge populations identified using Lagrangian modelling, implies that the genetic consequences of the ACH may only become evident when levels of connectivity are so low that they manifest at the evolutionary (rather than ecological) timescale. In addition, population sizes even of edge populations may be sufficiently large to prevent diversity-reducing factors that affect small populations (e.g. genetic bottlenecks). Hence, occasional larval exchange between large central and peripheral populations of South African marine mussels may prevent geographic genetic divergence, and be sufficient to nullify the conditions expected to underpin an ACH effect. We suggest that the genetic predictions of the ACH are unlikely to be supported by species with high dispersal capabilities such as those with planktonic, wind- or vector-driven dispersal of propagules.
Data availability
DNA sequences have been submitted to GenBank (accession numbers MN720954–MN721291). Microsatellite data and an animation of larval dispersal are available from figshare (https://doi.org/10.6084/m9.figshare.8910812).
References
Brown, J. H. On the relationship between abundance and distribution of species. Am. Nat. 124, 255–279 (1984).
Brown, J. H., Mehlman, D. W. & Stevens, G. C. Spatial variation in abundance. Ecology 76, 2028–2043 (1995).
Gilman, S. E. The northern geographic range limit of the intertidal limpet Collisella scabra: a test of performance, recruitment, and temperature hypotheses. Ecography 29, 709–720 (2006).
Levy, E. et al. Contrasting influences of geographic range and distribution of populations on patterns of genetic diversity in two sympatric Pilbara acacias. Plos One 11, e0163995 (2016).
Provan, J. & Maggs, C. A. Unique genetic variation at a species’ rear edge is under threat from global climate change. Proc. R. Soc. B Biol. Sci. 279, 39–47 (2012).
Kirkpatrick, M. & Barton, N. H. Evolution of a species’ range. Am. Nat. 150, 1–23 (1997).
Kawecki, T. J. & Holt, R. D. Evolutionary consequences of asymmetric dispersal rates. Am. Nat. 160, 333–347 (2002).
Sagarin, R. D. & Gaines, S. D. The ‘abundant centre’ distribution: to what extent is it a biogeographical rule? Ecol. Lett. 5, 137–147 (2002).
Eckert, C. G., Samis, K. E. & Lougheed, S. C. Genetic variation across species’ geographical ranges: The central-marginal hypothesis and beyond. Mol. Ecol. 17, 1170–1188 (2008).
Zardi, G. I. et al. Closer to the rear edge: ecology and genetic diversity down the core-edge gradient of a marine macroalga. Ecosphere 6, 1–26 (2015).
Bally, R., McQuaid, C. D. & Brown, A. C. Shores of mixed sand and rock: an unexplored marine ecosystem. S. Afr. J. Sci. 80, 500–5003 (1984).
Lombard, A. T. Marine component of the National Spatial Biodiversity Assessment for the development of South Africa’s National Biodiversity Strategic and Action Plan. 101 pp.-101 pp (2004).
Schurink, V. E. & Griffiths, C. C. L. Marine mussels of southern Africa - their distribution patterns, standing stocks, exploitation and culture. J. Shellfish Res. 9, 75–85 (1990).
Zardi, G. I., McQuaid, C. D., Teske, P. R. & Barker, N. P. Unexpected genetic structure of mussel populations in South Africa: indigenous Perna perna and invasive Mytilus galloprovincialis. Mar. Ecol. Prog. Ser. 337, 135–144 (2007).
Zardi, G. I. et al. Intraspecific genetic lineages of a marine mussel show behavioural divergence and spatial segregation over a tropical/subtropical biogeographic transition. BMC Evol. Biol. 15, 100 (2015).
Cunha, R. L. et al. Wider sampling reveals a non-sister relationship for geographically contiguous lineages of a marine mussel. Ecol. Evol. 4, 2070–2081 (2014).
McQuaid, C. D., Porri, F., Nicastro, K. R. & Zardi, G. I. Simple, scale-dependent patterns emerge from very complex effects - an example from the intertidal mussels Mytilus galloprovincialis and Perna perna. in Oceanography and Marine Biology: An Annual Review (eds. Hughes, R. N., Hughes, D. J., Smith, I. P. & Dale, A. C.) vol. 53, 127–156 (Taylor & Francis, 2015).
Assis, J. et al. Oceanographic conditions limit the spread of a marine invader along southern African shores. Plos One 10, e0128124 (2015).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Doyle, J. CTAB total DNA isolation. in Molecular techniques in taxonomy 283–293, https://doi.org/10.1007/978-3-642-83962-7_18 (1991).
Folmer, O., Black, M., Hoeh, W., Lutz, R. & Vrijenhoek, R. DNA primers for amplification of mitochondrial cytochrome c oxidase subunit I from diverse metazoan invertebrates. Mol. Mar. Biol. Biotechnol. 3, 294–299 (1994).
Coelho, N. C., Zardi, G. I., Pearson, G. A., Serrão, E. A. & Nicastro, K. R. Characterization of ten highly polymorphic microsatellite loci for the intertidal mussel Perna perna, and cross species amplification within the genus. BMC Res. Notes 5, 558 (2012).
Kumar, S., Stecher, G. & Tamura, K. MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 33, 1870–1874 (2016).
Kearse, M. et al. Geneious Basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28, 1647–1649 (2012).
Chapuis, M.-P. & Estoup, A. Microsatellite null alleles and estimation of population differentiation. Mol. Biol. Evol. 24, 621–631 (2007).
Excoffier, L. & Lischer, H. E. L. Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol. Ecol. Resour. 10, 564–567 (2010).
Bandelt, H. J., Forster, P. & Röhl, A. Median-joining networks for inferring intraspecific phylogenies. Mol. Biol. Evol. 16, 37–48 (1999).
Leigh, J. W. & Bryant, D. POPART: full-feature software for haplotype network construction. Methods Ecol. Evol. 6, 1110–1116 (2015).
Peakall, R. & Smouse, P. E. GenAlEx 6.5: Genetic analysis in Excel. Population genetic software for teaching and research-an update. Bioinformatics 28, 2537–2539 (2012).
Pritchard, J. K., Stephens, M. & Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 155, 945–959 (2000).
Evanno, G., Regnaut, S. & Goudet, J. Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol. Ecol. 14, 2611–2620 (2005).
Earl, D. A. & vonHoldt, B. M. Structure Harvester: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv. Genet. Resour. 4, 359–361 (2012).
Kopelman, N. M., Mayzel, J., Jakobsson, M., Rosenberg, N. A. & Mayrose, I. Clumpak: a program for identifying clustering modes and packaging population structure inferences across K. Mol. Ecol. Resour. 15, 1179–1191 (2015).
Ramasamy, R. K., Ramasamy, S., Bindroo, B. B. & Naik, V. G. STRUCTURE PLOT: a program for drawing elegant STRUCTURE bar plots in user friendly interface. SpringerPlus 3, 431 (2014).
Meirmans, P. G. & Hedrick, P. W. Assessing population structure: F ST and related measures. Mol. Ecol. Resour. 11, 5–18 (2011).
Kamvar, Z. N., Tabima, J. F. & Grünwald, N. J. Poppr: an R package for genetic analysis of populations with clonal, partially clonal, and/or sexual reproduction. PeerJ 2, e281 (2014).
Bruvo, R., Michiels, N. K., D’Souza, T. G. & Schulenburg, H. A simple method for the calculation of microsatellite genotype distances irrespective of ploidy level. Mol. Ecol. 13, 2101–2106 (2004).
Nei, M. Molecular evolutionary genetics. (Columbia University Press, 1987).
Nei, M. & Li, W. H. Mathematical model for studying genetic variation in terms of restriction endonucleases. Proc. Natl. Acad. Sci. USA 76, 5269–5273 (1979).
Alexander, A. et al. What influences the worldwide genetic structure of sperm whales (Physeter macrocephalus)? Mol. Ecol. 25, 2754–2772 (2016).
Keenan, K., McGinnity, P., Cross, T. F., Crozier, W. W. & Prodöhl, P. A. diveRsity: An R package for the estimation and exploration of population genetics parameters and their associated errors. Methods Ecol. Evol. 4, 782–788 (2013).
Assis, J. et al. Past climate changes and strong oceanographic barriers structured low-latitude genetic relics for the golden kelp Laminaria ochroleuca. J. Biogeogr. 45, 2326–2336 (2018).
Cunha, R. L. et al. Drivers of Cape Verde archipelagic endemism in keyhole limpets. Sci. Rep. 7, 41817 (2017).
Chassignet, E. P. et al. The HYCOM (HYbrid Coordinate Ocean Model) data assimilative system. J. Mar. Syst. 65, 60–83 (2007).
Lett, C. et al. A Lagrangian tool for modelling ichthyoplankton dynamics. Environ. Model.Softw. 23, 1210–1214 (2008).
Haklay, M. & Weber, P. OpenStreetMap: User-generated street maps. IEEE Pervasive Comput. 7, 12–18 (2008).
Buonomo, R. et al. Habitat continuity and stepping-stone oceanographic distances explain population genetic connectivity of the brown alga Cystoseira amentacea. Mol. Ecol. 26, 766–780 (2017).
Csardi, G. & Nepusz, T. The igraph software package for complex network research. InterJournal Complex Syst. 1695, 1–9 (2006).
Hijmans, R. J., Phillips, S., Leathwick, J., Elith, J. & Hijmans, M. R. J. dismo: species distribution modeling, http://CRAN.R-project.org/ package=dismo (2017).
Lamigueiro, O. P. & Hijmans, R. Package ‘rasterVis’ (2019).
Oksanen, J. et al. Package ‘vegan’ (2019).
Addison, J. A. & Hart, M. W. Spawning, copulation and inbreeding coefficients in marine invertebrates. Biol. Lett. 1, 450–453 (2005).
Teske, P. R., Papadopoulos, I., Barker, N. P. & McQuaid, C. D. Mitochondrial DNA paradox: sex-specific genetic structure in a marine mussel – despite maternal inheritance and passive dispersal. BMC Genet. 13, 45–45 (2012).
Wahlund, S. Zusammensetzung von Populationen und Korrelationserscheinungen vom Standpunkt der Vererbungslehre aus betrachtet. Hereditas 11, 65–106 (1928).
Nicastro, K. R., Zardi, G. I., McQuaid, C. D., Teske, P. R. & Barker, N. P. Coastal topography drives genetic structure in marine mussels. Mar. Ecol. Prog. Ser. 368, 189–195 (2008).
Soulé, M. E., Yang, S. Y., Weiler, M. G. W. & Gorman, G. C. Island lizards: the genetic-phenetic variation correlation. Nature 242, 191 (1973).
Abeli, T., Gentili, R., Mondoni, A., Orsenigo, S. & Rossi, G. Effects of marginality on plant population performance. J. Biogeogr. 41, 239–249 (2014).
Péron, G. & Altwegg, R. Departures from the energy-biodiversity relationship in South African passerines: are the legacies of past climates mediated by behavoural constraints on dispersal? Plos One 10, e0133992 (2015).
Chardon, N. I., Cornwell, W. K., Flint, L. E., Flint, A. L. & Ackerly, D. D. Topographic, latitudinal and climatic distribution of Pinus coulteri: geographic range limits are not at the edge of the climate envelope. Ecography 38, 590–601 (2015).
Pironon, S., Villellas, J., Morris, W. F., Doak, D. F. & García, M. B. Do geographic, climatic or historical ranges differentiate the performance of central versus peripheral populations? Glob. Ecol. Biogeogr. 24, 611–620 (2015).
Teske, P. R., Von der Heyden, S., McQuaid, C. D. & Barker, N. P. A review of marine phylogeography in southern Africa. S. Afr. J. Sci. 107, 1–11 (2011).
Teske, P. R. et al. Implications of life history for genetic structure and migration rates of southern African coastal invertebrates: planktonic, abbreviated and direct development. Mar. Biol. 152, 697–711 (2007).
Teske, P. R. et al. Thermal selection as a driver of marine ecological speciation. Proc. R. Soc. B Biol. Sci. 286, 20182023 (2019).
Bolton, J. J. Marine phytogeography of the Benguela upwelling region on the west coast of Southern Africa: a temperature dependent approach. Bot. Mar. 29, 251–256 (1986).
van den Hoek, C. The distribution of benthic marine algae in relation to the temperature regulation of their life histories. Biol. J. Linn. Soc. 18, 81–144 (1982).
Lynch, J. A., Olesnicky, E. C. & Desplan, C. Regulation and function of tailless in the long germ wasp Nasonia vitripennis. Dev. Genes Evol. 216, 493–498 (2006).
Stoeckle, M. Y. & Thaler, D. S. DNA barcoding works in practice but not in (neutral) theory. PloS One 9, e100755 (2014).
Teske, P. R., Winker, H., McQuaid, C. D. & Barker, N. P. A tropical/subtropical biogeographic disjunction in southeastern Africa separates two evolutionarily significant units of an estuarine prawn. Mar. Biol. 156, 1265–1275 (2009).
Teske, P. R. et al. ‘Nested’ cryptic diversity in a widespread marine ecosystem engineer: a challenge for detecting biological invasions. BMC Evol. Biol. 11, 176–176 (2011).
Jooste, C. M., Oliver, J., Emami-Khoyi, A. & Teske, P. R. Is the Wild Coast in eastern South Africa a distinct marine bioregion? Helgol. Mar. Res. 72, 6 (2018).
Vucetich, J. A. & Waite, T. A. Spatial patterns of demography and genetic processes across the species’ range: null hypotheses for landscape conservation genetics. Conserv. Genet. 4, 639–645 (2003).
Cahill, A. E. & Levinton, J. S. Genetic differentiation and reduced genetic diversity at the northern range edge of two species with different dispersal modes. Mol. Ecol. 25, 515–526 (2016).
McQuaid, C. D. & Phillips, T. E. Limited wind-driven dispersal of intertidal mussel larvae: In situ evidence from the plankton and the spread of the invasive species Mytilus galloprovincialis in South Africa. Mar. Ecol. Prog. Ser. 201, 211–220 (2000).
Acknowledgements
This study was funded by the Foundation for Science and Technology (FCT) of Portugal through the projects IF/01413/2014/CP1217/CT0004 and UID/Multi/04326/2019, and in the scope of the transitional norm – DL57/2016/CP1361/CT0035. Additional funding was provided by the National Research Foundation (NRF) of South Africa (grant number 64801), and by the University of Johannesburg (FRC/URC grant to P.R.T.). The NRF is gratefully acknowledged for granting an MSc bursary to N. Ntuli. Bioinformatics support was provided by the CSIR Centre for High Performance Computing (CHPC), Rondebosch, South Africa.
Author information
Authors and Affiliations
Contributions
G.I.Z. and K.R.N. designed the study and estimated mussel cover; N.N.N. and P.R.T. collected the samples; N.N.N. generated the genetic data; N.N.N., G.I.Z., K.R.N. and P.R.T. analysed the genetic data; J.A. conducted dispersal simulations; N.N.N., G.I.Z., K.R.N., C.D.M., J.A. and P.R.T. wrote the paper. All authors read and approved the final version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ntuli, N.N., Nicastro, K.R., Zardi, G.I. et al. Rejection of the genetic implications of the “Abundant Centre Hypothesis” in marine mussels. Sci Rep 10, 604 (2020). https://doi.org/10.1038/s41598-020-57474-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-57474-0
This article is cited by
-
Multidisciplinary estimates of connectivity and population structure suggest the use of multiple units for the conservation and management of meagre, Argyrosomus regius
Scientific Reports (2024)
-
Photosynthetic response to a winter heatwave in leading and trailing edge populations of the intertidal red alga Corallina officinalis (Rhodophyta)
Acta Oceanologica Sinica (2024)
-
Genetic diversity and gene expression diversity shape the adaptive pattern of the aquatic plant Batrachium bungei along an altitudinal gradient on the Qinghai–Tibet plateau
Plant Molecular Biology (2023)
-
Phenotypic Plasticity in Sargassum Forests May Not Counteract Projected Biomass Losses Along a Broad Latitudinal Gradient
Ecosystems (2023)
-
Habitat characterizations and suitability analysis for conservation implications of Gymnosphaera gigantea (Wall. ex Hook.) S.Y.Dong: a threatened tree fern
Arabian Journal of Geosciences (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.