Abstract
Sarcoidosis is a disorder characterized by granulomatous inflammation of unclear etiology. In this study we evaluated whether veterans with sarcoidosis exhibited different plasma metabolomic and metallomic profiles compared with civilians with sarcoidosis. A case control study was performed on veteran and civilian patients with confirmed sarcoidosis. Proton nuclear magnetic resonance spectroscopy (1H NMR), hydrophilic interaction liquid chromatography mass spectrometry (HILIC-MS) and inductively coupled plasma mass spectrometry (ICP-MS) were applied to quantify metabolites and metal elements in plasma samples. Our results revealed that the veterans with sarcoidosis significantly differed from civilians, according to metabolic and metallomics profiles. Moreover, the results showed that veterans with sarcoidosis and veterans with COPD were similar to each other in metabolomics and metallomics profiles. This study suggests the important role of environmental risk factors in the development of different molecular phenotypic responses of sarcoidosis. In addition, this study suggests that sarcoidosis in veterans may be an occupational disease.
Similar content being viewed by others
Introduction
Sarcoidosis is a granulomatous entity of unknown etiology. The incidence of sarcoidosis has been well studied in the American civilian population and ranges between 11 in 100,000 in Caucasians to up to 36 in 100,000 in African American populations1. Although not well studied, the incidence of sarcoidosis among veterans is estimated to be even higher2. Sarcoidosis has protean manifestations including ophthalmic, joint, skin, and liver involvements, but most often involves the lung, affecting almost 90% of cases. In some individuals it can lead to serious disability of affected organs such as pulmonary fibrosis, cirrhosis, blindness, and may be fatal. Unfortunately, there is no available noninvasive biomarker to facilitate the diagnosis nor predict progression of disease.
Metabolomics is a method to identify and measure small molecules – metabolites – which may provide a way to study and monitor disease progression. In addition, has the potential to differentiate various stages of a particular disorder, provide means for a more accurate diagnosis and possibly stratification of prognosis3,4. Two of the more commonly used analytical platforms for metabolomics studies are proton nuclear magnetic resonance (1H-NMR) spectroscopy, a very robust and consistent method, and hydrophilic interaction liquid chromatography mass spectrometry (HILIC-MS), an extremely sensitive method5. Metabolomics has helped to identify potential biomarkers for the diagnosis and prognosis of infectious and non-infectious respiratory conditions such as asthma6, chronic obstructive pulmonary disease (COPD)7,8, influenza9, pneumonia10, tuberculosis11, and acute respiratory distress syndrome (ARDS)12,13.
Metallomics targets various elements and metal ions that participate in many biological pathways in close relationship with proteins and metabolites14. Inductively coupled plasma-mass spectrometry (ICP-MS) is a highly sensitive technique used to detect and quantify elements in the periodic table in biological fluids15.
The veteran population comprise a particular group of subjects with a unique history of exposure to hazardous materials, including gun smoke, jet fuel, air pollution (burn pit smoke, dust) and occupational hazards (asbestos, lead)16. Because of this, we sought to evaluate whether veterans with sarcoidosis exhibit different metabolomics and metallomics profiles compared with civilians with sarcoidosis. Furthermore, we evaluated whether veterans with sarcoidosis have a different metabolomic and metallomic profile compared to other veterans with a non-sarcoidosis pulmonary condition, (COPD).
Methods
Patient enrollment
We performed a case-control study on veterans (n = 13) and civilians (n = 30) with confirmed pulmonary sarcoidosis. Sarcoidosis was defined as the presence of clinical signs and symptoms of pulmonary sarcoidosis or the presence or history of bilateral hilar lymphadenopathy, along with biopsy-proven sarcoid-like granulomas in pulmonary samples, with the exclusion of other granulomatous conditions, including mycobacterial infection.
Airflow obstruction was defined per American Thoracic Society and European Respiratory Society (ATS/ERS) guidelines with an FEV1/FVC ratio less than lower limit of normal (24). In order to further analyze the metabolomic profiles of veterans without sarcoidosis and with similar exposure history, we randomly selected 35 veterans with COPD as controls, matched by race, gender, and deployment history. Civilians were recruited from the University of Miami Sarcoidosis Program and veterans from the Miami Veterans Administration Sarcoidosis Program.
This retrospective study was conducted in accordance with Helsinki Declaration and the study was been approved by the University of Miami’s Institutional Review Board (No.20150612). Written informed consent was obtained from all participants.
1H-NMR spectroscopy and metabolites profiling
Plasma samples were obtained from all patients and analyzed by 1H-NMR spectroscopy. Details of sample preparation have been described in the Supplementary Appendix (Supp. 1). One dimensional 1H-NMR was performed using a 600 MHz Bruker Ultrashield Plus 1H-NMR spectrometer (Bruker BioSpin Ltd., Canada). Details of the 1H-NMR analysis have previously published9 and provided in the Supp. 1. To quantify the metabolite concentrations, DSS (4,4-dimethyl-4-silapentane-1-sulfonic acid) was used as an internal standard. Multivariate statistical analysis models were developed to identify metabolites involved in the discrimination. ChenomX 7.1 was used to identify and quantify metabolites17.
HILIC-MS and metabolites profiling
Plasma samples were also analyzed by liquid chromatography mass spectroscopy (LC-MS) using the Q Exactive HF Hybrid Quadrupole-Orbitrap Mass Spectrometer, Thermo-Fisher). Chromatography was performed using a 2.1 mm × 100 mm long Syncronis HILIC (thermos-Fisher) LC column packed in-house with 3 µm porous Hypercarb particles.
The elution gradient of solvent B (%) (acetonitrile with 0.1% formic acid) over time was: 95% for 2 min, 85%–95% for 5 min, 5–80% for 3 min, and 5% for 5 min, 5–95% for 2 min and then held at 95% for last 3 min against solvent A (20 mM ammonium formate pH 3.0 in H2O). MS conditions were as follows: HESI-II temperature 325 °C, auxiliary gas flow 10 units, sheath gas flow 25, spray voltage ± 2.50 kV, capillary temperature 275 °C, and S-lens RF level 60%, auxiliary gas heater temperature 275 °C for negative ion mode. To acquire mass spectra, mass scan parameters were set up as follows: runtime 20 mins, full MS scan type, resolution 240,000, AGC target 3e6, maximum IT 200 ms and scan range 70–1000 m/z. Maven software, an open source software, was used for processing metabolomics data obtained by LC-MS18.
ICP-MS analysis of metallome profiles
The plasma metallome was assessed using inductively coupled plasma mass spectrometry (ICP-MS). Details of the sample preparation is also described in the Supp. 1. ICP-MS analysis was performed using a PlasmaQuant® MS Elite (Analytik Jena, Jena, Germany) spectrometer on plasma samples. A standard SeronormTM serum samples was used as quality control samples and provided a way to translate the elemental count of the ICP-MS to semi quantitative. Integrated collision reaction cell (iCRC) mode was applied for analysis of plasma using a single continuous method. To attenuate all polyatomic interference, hydrogen gas was added to the iCRC skimmer. Both iCRC and non-iCRC modes were applied for analysis, thus the isotopes (different forms of an element with ‘different number of neutrons) 51V, 75As, 78Se and 90Zr were quantified by both iCRC mode and non-iCRC mode. 24Mg, 56Fe and 51Cr were quantified only in iCRC mode and non-iCRC mode was used for all the remaining isotopes.
Data analysis
Both univariate and multivariate data analyses were applied to the extracted information from the metabolomic datasets. Univariate analyses such as, ANOVA and T-test were used as complementary methods to the multivariate analysis in order to provide useful information on metabolomics profiles and on metabolite individually. MetaboAnalyst19 and MetaBox20 were used for univariate data analyses. Data were not preprocessed for the univariate analyses.
Multivariate data analysis was applied to reduce the complexity of metabolomics data and for data mining. Principal component analysis (PCA) was performed to find outliers, trends, and similarities, using datasets derived from the plasma samples for the evaluation of interrelations and groupings of metabolomics data between veteran and civilians with pulmonary sarcoidosis and veterans with COPD.
Partial least square-discriminant analysis (PLS-DA) and orthogonal partial least square-discriminant analysis (OPLS-DA) are two supervised multivariate data analyses methods for classification and discrimination between groups by maximizing separation using most differentiating variables responsible for class discriminations. PLS-Da and OPLS-DA were used to separate different populations and to establish metabolomic profiles for each population based on the most differentiating metabolites. Q2Y (goodness of validation) and cross validation ANOVA (CV-ANOVA) parameters were considered for predictability and significance of separation of study populations using metabolomics profiles21,22. Q2Y > 0.3 and 0.5 were evaluated as acceptable and good models for human samples. P value ≤ 0.05 was considered a significant model. The highest Q2Y was considered in choosing the OPLS-DA model. S-plot and coefficient plot were applied to find the most significant metabolites that contributed to the separation as potential variables in the study. S-plot was used to extract the putative biomarkers, that have both high reliability and magnitude. Compounds with covariance >0.1 were considered important metabolites23. SIMCA-P v15.0.2 (Umetrics AB, Umeå, Sweden) and MetaboAnalyst 3.0 software were used for multivariate analyses. Data were normalized (median), transformed (log) and auto-scaled for multivariate analyses.
Prediction test
Prediction test was used to obtain sensitivity, specificity and area under the curve for receiver operating (AUROC) parameters. The prediction test was carried out by splitting the population of the study into a training and a predicting set. The predicting group was created by randomly selecting 25% of the samples. A misclassification table was obtained for all discriminant analysis models to measure sensitivity and specificity using SIMCA-P v15.0.2 software. AUROC was calculated using Graph Pad Prism v 3.03.
Pathway analysis
Parallel pathway analyses were performed using MetaboAnalyst 3.024, a free web-based tool, and Cytoscape 3.6.025 using selected discriminative metabolites in the separation of veteran sarcoidosis from civilian cohorts. Reactome, an open-source, open access, pathway database was also used to explore biochemical networks26. The list of discriminative metabolites was chosen according to the best OPLS-DA model which had higher predictability (Q2Y).
Results
Patient characteristics
Of the 78 subjects who were consented for the study, 43 had sarcoidosis (13 veterans and 30 civilians) and 35 were veterans with COPD. Demographic characteristics are summarized in Table 1. Out of the 43 subjects enrolled with sarcoidosis, 10 subjects (33%) in the civilian group and 1 subject (8%) in the veteran group were female (P = 0.107). There were no significant differences in age, race or lung function between participants. The difference in the mean (SD) CPI score of civilian and veteran subjects were 19.1 (21.3) and 29.2 (18.8), respectively (P = 0.0148).
Among COPD subjects, the mean (SD) age was 63.5 (4.7), which was significantly older than veterans with sarcoidosis (P = 0.038). However, there was no difference in race and gender between these two groups.
Metabolomics study
Metabolite Identification
Using ChenomX 1H-NMR Suite 7.127, one dimensional 1H-NMR analysis resulted in the identification of 55 metabolites including sugars, organic acids, amino acids, and volatile organic compounds. Using the Maven software, 103 variables were identified in a targeted approach to the HILIC-MS analysis and consisted of sugars, amino acids, organic acids, acylcarnitines and their derivatives.
Principal component analysis (PCA) clearly showed clustering between veteran and civilian sarcoidosis and COPD control
Unsupervised PCA demonstrated clustering among the three groups using the 55 and 103 metabolites detected by 1H-NMR and HILIC-MS, respectively (Fig. S1A,B). However, PCA showed a a higher degree of separation between veteran and civilian sarcoidosis cohorts reflecting two distinct metabolomic profiles (Fig. S2A,B). The 6-component PCA models had an R2X = 0.483 and 0.456 for 1H-NMR and HILIC-MS, respectively, demonstrating a relative high variation of metabolites among the samples. Further analysis revealed very good clustering between civilian sarcoidosis and veterans with COPD, whereas the two veteran cohorts were roughly clustered (Figs. S3 and S4).
Veterans with sarcoidosis have a different metabolomic profile compared to civilians with sarcoidosis
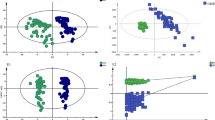
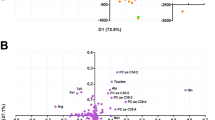
Multivariate data analysis of metabolomic profiles using OPLS-DA detected that the veteran sarcoidosis cohort was significantly different from the civilian sarcoidosis cohort. The results provided a highly predictive and significant model to separate the two cohorts particularly for HILIC-MS (Q2Y = 0.607, p = 2.8 × 10−7) than 1H-NMR spectroscopy (Q2Y = 0.454, p = 9.9 × 10−5) (Fig. 1A,B). Forty-one metabolites contributed to the differentiation of metabolic profiles between civilians and veterans with sarcoidosis for each of the two analytical approaches (Fig. S5A,B). Six metabolites were detected in both HILIC-MS and 1H-NMR to be significantly different between civilian and veteran sarcoidosis subjects: arginine, glutamine, creatinine, glycine, taurine, and methionine.
Shows OPLS-DA generated from 1H-NMR (A) and HILIC-MS (B) data, demonstrating a significant difference between the veteran and civilian subjects with pulmonary sarcoidosis with a higher predictive power for HILIC-MS (Q2Y = 0.607, p = 2.8 × 10−7) than 1H-NMR spectroscopy (Q2Y = 0.454, p = 9.9 × 10−5).
The performances of the discriminative models are summarized in Table 2. Most differentiating metabolites were selected for the OPLS-DA models to obtain higher predictability (Q2Y) using a variable importance in projection (VIP) approach. S-plot revealed that taurine, sucrose, n-methyl-d-aspartic acid, ll-2–6-diaminoheptanedioate, hypoxanthine, ethanolamine, alpha-hydroxyisobutyric acid, sn-glycerol 3-phosphate, 2-oxoisocaporate, 4-hydroxybutyrate, acetone histidine, isoleucine, isopropanol, methionine and beta-alanine were the most plausible biomarkers to differentiate between the two sarcoidosis cohorts in both datasets (Fig. S6A,B). This study also showed that a targeted analysis using HILIC-MS could be a better discriminative method because of its higher Q2 and the fact that includes all 103 identified metabolites (Fig. S7). Permutation test confirmed the validation of R2 and Q2 values of each predictive model (OPLS-DA) using 200 repetitions (Fig. S8).
Unpaired T-tests on the sarcoidosis populations were performed using 55 and 103 metabolites identified by 1H-NMR and HILIC-MS, respectively. The T-test analysis on the 1H-NMR data showed 8 metabolites that were significantly (p < 0.05) different between the veteran and civilian cohorts with 6 having a significant low false discovery rates (FDR) (<0.05) (Table S1 & Fig. 2). T-tests on the HILIC-MS dataset showed 38 metabolites differentially expressed (p < 0.05), from which 24 metabolites had significant FDR (p < 0.05) (Table S2 & Fig. 3). These results confirm that the two sarcoidosis populations are metabolomically different.
Veterans and civilians with sarcoidosis have different metabolomic profiles compared to veterans with COPD
MVA analysis also showed very distinct metabolic profiles for the sarcoidosis civilian cohort and the COPD cohort for both HILIC-MS (Q2 = 0.724) and 1H-NMR (Q2 = 0.713) (Figs. S9B and S10B). However, a smaller difference occurred between the two veteran cohorts. The predictability (Q2Y = 0.408 and 0.46) and significant difference (p = 0.00012 and 0.0001) for both 1H-NMR and HILIC-MS methods were considerably lower between the two veteran cohorts (Table 2) as shown in Figs. S9A and S10A. Also, a univariate approach using unpaired t-test proved that the metabolite alterations were more significant between civilian sarcoidosis and veterans with COPD than between the two veterans’ groups (Table S3–6).
Metallomics study
PCA of the three study groups show significant clustering
In a targeted approach, 33 trace elements were quantified in the plasma samples of the three cohorts (civilians with sarcoidosis, veterans with sarcoidosis and veteran controls with COPD) using ICP-MS techniques. PCA analysis (Fig S11) shows a very clear separation of veteran patients with sarcoidosis from civilians with sarcoidosis. Similar to metabolomics results, this discrepancy was more noticeable between civilian sarcoidosis vs. COPD than veteran sarcoidosis vs. COPD (Figs. S12 and S13).
Veteran sarcoidosis showed significant different metallomic profile compared to civilian sarcoidosis
OPLS-DA analysis revealed a very good, highly predictive (Q2Y = 0.688) and significant (p = 6.003 × 10−6) model to discriminate between the two sarcoidosis groups based on the 33 elements (Fig. 4). The coefficient plot (Fig. S14) depicts differences in 33 elements between both cohorts showing increased rubidium (85Rb), gallium (71Ga), nickel (60Ni), cesium (133Cs), arsenic (75As), barium (137Ba), aluminum (127Al) and mercury (202Hg) and decreased boron (11Ba), antimony (121Sb), cadmium (114Cd) bromine (79Br), cobalt (59Co), Calcium (44Ca), selenium (77Se), strontium (88Sr), palladium (105Pd), magnesium (24Mg) and lead (208Pb) in the plasma samples of the veterans with sarcoidosis compared to the civilian cohort. Unpaired t-test, a univariate approach, showed 18 elements that were significantly different (p < 0.05) between the two sarcoidosis groups, including 10 elements with significant FDR (<0.05) (Table S7). It has been shown that other elements such as iron (56Fe) and manganese (55Mn) were significantly increased in the plasma of veteran sarcoidosis compared to civilian cohort. Permutation test confirmed the validation of R2 and Q2 values of each predictive model (OPLS-DA) using 200 repetitions.
Metallomics profiles were significantly different between subjects with sarcoidosis and COPD
OPLS-DA analyses showed closer metallomics profiles between the two veteran groups compared to the civilian cohort. Utilizing the 33 elements, OPLS-DA analysis was highly predictive (Q2Y = 0.81) in separating civilian sarcoidosis subjects from veterans with COPD, while it was a less predictive discriminant model (Q2Y = 0.53) in separating veteran sarcoidosis subjects from veterans with COPD, using 29 elements (Fig. S15A,B). Additionally, individual t-tests provided more elements that were significantly (p < 0.05) changed when comparing civilian subjects with sarcoidosis versus veterans with COPD compared to veterans with sarcoidosis versus veterans with COPD (Tables S8 and S9). Overall, our metallomics results were remarkably similar to our metabolomics studies, likely reflecting different pathophysiological mechanisms of sarcoidosis in veterans compared to civilians.
Correlation of metabolomic and metallomic profiles and sarcoidosis cohort status and clinical composite physiological index (CPI)
PLS-regression showed a strong relationship between predictive metabolomics profiles in distinguishing the veteran and civilian cohorts with sarcoidosis. R2 values were 0.95 and 0.91 for the HILIC-MS and the 1H-NMR metabolomic data, respectively (Fig. S16). The relation was weak between metabolomics profiles and CPI as an independent variable (R2 = 0.55 for HILIC-MS and R2 = 0.77 for 1H-NMR). R2 values for the regression were obtained using the PLSR method based on the most important metabolites obtained by the OPLS-DA analysis. Moreover, logistic regression showed that CPI is not a good predictive marker to discriminate between the two sarcoidosis cohorts (Table 1).
A similar PLS-regression analysis on metallomics profiles demonstrated a weaker correlation (R2 = 0.80) than metabolomics in discriminating between the sarcoidosis cohorts (Fig. S17). There was not a strong relationship between metallomics data in discriminating sarcoidosis from COPD.
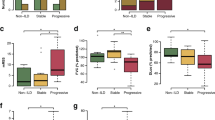
Relation of Metabolomics profiles and radiologic sarcoidosis stage
Subjects with stage 4 sarcoidosis different from other stages in their metabolomics profile (HILIC-MS data) with a high degree of predictability and a statistic significant difference (Q2Y = 0.522, p = 0.00019) (Fig. 5). Twenty-nine metabolites contributed in the differentiation of the radiologic sarcoidosis stages. Coefficient plots that illustrate the relative correlation of metabolites characteristic of sarcoidosis stage 4 vs. stages 1–3 is shown in Fig. S18. Univariate analysis showed only 7 metabolites that significantly (p < 0.05) differed between stages 1–3 and stage 4, which were already noted in the multivariate analysis (Table S10).
Integration of 1H-NMR and HILIC-MS datasets
The 1H-NMR and HILIC-MS metabolomics datasets were further integrated using a normalized and block scaled method. Although there were several overlapped metabolites, both methods used different approaches for metabolite identification. The OPLS-DA model obtained, showed a slight improvement in separating the two sarcoidosis cohorts (Q2Y = 0.626) based on the 72 metabolites compared to the HILIC-MS dataset alone. Similar results were obtained for the discrimination between veterans with sarcoidosis and COPD, with a Q2Y = 0.693 and Q2Y = 0.843, respectively. Therefore, the combination of the two datasets further illustrates the metabolomic similarity of the two veteran cohorts when compared with the civilian sarcoidosis group.
Pathway analysis
We applied parallel pathways analysis by MetaboAnalyst 3.0 and Cytoscape 3.6.0 using most differentiating metabolites obtained by the HILIC-MS and 1H-NMR platforms. Table 3 summarizes the details of the significant (p < 0.05) biochemical pathways with impact factor >1.0 after integration of both datasets. Using Cytoscape, the most important upregulated and downregulated biochemical pathways between sarcoidosis cohorts was graphically illustrated. We found that pathways involved in the transport of inorganic cations/anions and amino acids/oligopeptides, creatine metabolism, amino acid synthesis, and interconversion (transamination), amino acid transport across the plasma membrane, transport of glycerol from adipocytes to the liver by aquaporins, glycerophospholipid catabolism, and GABA biosynthesis, metabolism, and catabolism could be the most relevant biological networks that differ between veterans and civilians with sarcoidosis. A more detailed pathway analysis for both individual 1H-NMR and HILIC-MS datasets is shown in the supplementary section. (Tables S12 and S13).
Discussion
This study shows that veterans with sarcoidosis have distinct metabolomic and metallomic profiles when compared to civilians with sarcoidosis. Since these populations have different environmental and occupational exposures, our findings agree with the notion that sarcoidosis is an abnormal inflammatory condition in response to many (still unidentified) triggers that may have their own particular pathophysiologic signature. Not surprisingly, the metabolic profiles among the two veteran cohorts studied (sarcoidosis and COPD) showed more similarity between them compared to civilians with sarcoidosis.
A number of reviews and research studies have addressed the impact of “exposomes” in altering metabolomics signatures and the role of metabolomics in characterizing particular environmental factors28,29,30,31, but to the best of our knowledge, no study has experimentally demonstrated this relation clearly when related to environmental exposure and sarcoidosis. The application of metabolomics in studying sarcoidosis has been reported previously. Lactate, acetate and N-butyrate is increased in saliva and 3-hydroxybutyrate, acetoacetate, carnitine, cystine, homocysteine, pyruvate, and trimethylamine N-oxide are some metabolites increased in the serum samples of patients with sarcoidosis compared to normal controls, while serum methanol, butyrate in saliva and isoleucine, glutamine and succinate are decreased32,33. We previously showed LC-MS-based metabolomics could differentiate fibrosing from non-fibrosing pulmonary sarcoidosis as well as differentiate subjects with composite phycological index (CPI) > 40 versus those that had CPI < 4034.
In this study we used metabolomics as a tool to characterize the exposome and investigate the molecular fingerprints and key cellular processes that are likely activated by exposure to the environment. The differences in metabolite alterations were readily detectable by both 1H-NMR and HILIC-MS detection methods, but most notably by the latter. Veterans with sarcoidosis have unique exposures and this may dictate the development of a particular type of sarcoidosis. For example, Gulf War Illness (GWI) is a syndrome described in deployed military to the Persian Gulf region exposed to particular environmental threats such as dust storms35. It is a multisystem disease diagnosed based on the Kansas criteria (presence of 3 or more of following chronic symptoms: fatigue, sleep problems, pain syndrome, neurologic and cognitive symptoms, gastrointestinal, respiratory and skin symptoms)35. Targeted metabolomics studies have shown GWI is a unique metabolic syndrome that is characterized with increased sphingolipids and phosphatidylcholines and decreased purines and endocannabinoids36, a specific metabolic fingerprint different to other conditions with similar symptomatology such as myalgic encephalomyelitis/chronic fatigue syndrome (ME/CFS)36. Furthermore, the lipid profile of GWI can be replicated in rat and mouse models exposed to a nerve gas antidote, pyridostigmine bromide37,38. On the other hand, mounting evidence has indicated that environmental exposure such airborne inorganic dust could be risk factors for sarcoidosis39 and that the nature of the exposure can impact the frequency and clinical phenotypes of the disease40. In mouse models, exposure to silica dust leads to metabolomic alterations after lung inflammation41. Similarly, we present here significant differences in metabolomic profiles in 2 sarcoidosis populations with different exposures.
One of the significant metabolomic changes observed was in amino acids. Compared to civilians, veterans with sarcoidosis exhibited lower concentration in most amino acids analyzed molecules important in the induction of anti-inflammatory mechanisms and reducing stress response in inflammation42,43. Amino acids such as glutamine, phenylalanine, tryptophan, arginine and cysteine play a vital role in immunomodulation, particularly through the T-cell proliferation and activation44. Veterans also exhibited an increased concentration in some anti-inflammatory biomarkers such as taurine, hypoxanthine, glucosamine, reflecting a high degree of interactions between both anti-inflammatory and pro-inflammation mechanisms.
Exposure to metallic environmental factors, including occupational and infectious causes, have been associated with sarcoidosis45,46. For example, exposure to beryllium, aluminum, rare earth elements, and titanium have been associated with sarcoidosis in mine, manufacturer, and agriculture workers46,47,48,49. We found significantly higher in the plasma concentration of rubidium, gallium, nickel, and cesium in the plasma of veterans compared to civilians with sarcoidosis. Veterans deployed to the Middle East have particular exposures to heavy metal and trace elements. For example, Pb, Zn, Cd, Ni, and Cr contamination of seawater, food, and soils due to the Gulf War and oil spills have been reported50,51. Military operations by themselves have added another potential source of contamination of water and soil in the gulf, During the two Gulf Wars in the 1990s, millions of hectares were contaminated due to spilling of millions of oil barrels and burning of hundreds of oil wells and is estimated that 340 tons of uranium was depleted in the first Gulf War52. Accordingly, we have shown that the plasma level of uranium in veterans with sarcoidosis is higher than was observed in the civilian cohort. Furthermore, nickel contamination from different sources such as batteries can have effect on lung and antioxidant system in the form of water-soluble nickel phosphate53. Nickel was also higher in the veteran sarcoidosis group.
The lung is the first line of exposure that is exposed to essential or non-essential/toxic elements and potentially toxic metal-related nanomaterials. Nanoparticles generated from dust storms or gun powders may deposit in the lungs and direct activated alveolar macrophages toward the pulmonary interstitium causing a more severe immune response than larger particles54. Metalloids become localized and interact with pulmonary cells and tissues or are dispersed all over the body by passing biological barriers55. Metal ions have significant biological effects in different metabolic pathways56,57. For example, radioactive Cesium 137 (137Cs) has been associated with reduced lung function in children in the Chernobyl disaster58.
Dust storms in deployment areas represent an additional exposure hazard due to their high contents of silica, aluminum, and rare earth elements59,60. The association of silica exposure and development of sarcoidosis was studied by Rafnsson et al.61, who found that people who were exposed to dust had up to a 5 times higher risk of development of pulmonary sarcoidosis when compared to a normal population (9.3 vs. 0.5–2.7 in 100,000, respectively). Dust storms also contain particles such as, calcium oxide (CaO) and magnesium oxide (MgO), and oxides of sodium and potassium (Na2O and K2O) as well as silicon dioxide (SiO2), aluminum oxide (Al2O3), iron (Fe2O3) and titanium (TiO2) oxides and trace elements such as, zirconium (Zr), strontium (Sr), rubidium (Rb)62. Our data showed that Mg, Ca, Al, Ti, and Fe is increased in veterans with sarcoidosis compared to the civilian sarcoidosis population. Investigations on dust in the Iraq desert revealed that the quartz particle is surrounded by calcium carbonate containing various elements such as aluminum, iron, uranium, nickel, cobalt, copper, lead, chromium, strontium, tin, manganese, zinc, barium, arsenic, and vanadium63. Accordingly, we observed an increase in most of the aforementioned elements in the plasma samples of veteran sarcoidosis. Of note, cadmium, known to be involved in the pathogenesis of pulmonary carcinogenesis64, was not higher among the veterans studied.
We also noted that veterans with sarcoidosis have higher concentrations of elements that regulate biologic processes in immune cells. For instance, lanthanum (a rare earth element) can compete with calcium in different proteins65,66. This substitution can alter the function of annexin A11 (ANXA11), an intracellular metalloprotein that carries 5 calcium ions. It has been proposed that alteration of ANXA11 is associated with pulmonary fibrosis in sarcoidosis67. In this way, sarcoidosis in veterans may be considered an occupational disease.
Another interesting finding noted in veterans with sarcoidosis was with pathway analysis, which showed that GABA synthesis and metabolism might be downregulated in veterans compared to civilian with sarcoidosis, with both precursor metabolites of GABA (glutamate or glutamine) and breakdown metabolites of GABA (succinate semialdehyde) were decrease in veterans. GABA can modulate the immune response via secretion of cytokines, activation, proliferation and migration of immune cells68.
The current study has two major limitations. A relatively low sample size in our study may lead to a lack of precision and the results have not been validated in an external validation cohort. To partially overcome the limitations, we applied 3 different methods to validate the findings with higher resolution. The very similar results obtained using three different techniques, 1H-NMR, HILIC-MS and ICP-MS, serves as an important internal validation of our results.
Using metabolomics and metallomics, this study provides relevant evidence for the role of potential environmental risk factors for different molecular phenotypic responses of sarcoidosis. More investigation will be needed to identify biochemical pathways that link between environmental risk factors and a disease phenotype as well as metal-related metabolomics perturbations in human diseases. We conclude then that this comprehensive metabolite and metallomic profiling clearly shows distinct metabolomic profiles between two sarcoidosis populations exposed to different environmental factors. Metabolic fingerprints can be a powerful and useful tool for sarcoidosis phenotyping in a more accurate manner than the current clinical approach. Furthermore, characterization of metabolic profiles is a promising platform to better understand the underlying pathophysiological mechanisms, establish the importance of environmental exposures and determine disease severity, prognosis, and response to treatment.
References
Rybicki, B. A., Major, M., Popovich, J. Jr., Maliarik, M. J. & Iannuzzi, M. C. Racial differences in sarcoidosis incidence: a 5-year study in a health maintenance organization. Am J Epidemiol 145, 234–241 (1997).
Becher, F. et al. A strategy for liquid chromatography/tandem mass spectrometric assays of intracellular drugs: application to the validation of the triphosphorylated anabolite of antiretrovirals in peripheral blood mononuclear cells. J Mass Spectrom 38, 879–890, https://doi.org/10.1002/jms.500 (2003).
Oliver, S. G., Winson, M. K., Kell, D. B. & Baganz, F. Systematic functional analysis of the yeast genome. Trends Biotechnol 16, 373–378 (1998).
Heijne, W. H., Kienhuis, A. S., van Ommen, B., Stierum, R. H. & Groten, J. P. Systems toxicology: applications of toxicogenomics, transcriptomics, proteomics and metabolomics in toxicology. Expert Rev Proteomics 2, 767–780, https://doi.org/10.1586/14789450.2.5.767 (2005).
Banoei, M. M. et al. Metabolomics in critical care medicine: a new approach to biomarker discovery. Clinical and Inestigative Medicine 37, E363–376 (2014).
Ho, W. E. et al. Metabolomics reveals altered metabolic pathways in experimental asthma. Am J Respir Cell Mol Biol 48, 204–211, https://doi.org/10.1165/rcmb.2012-0246OC (2013).
Ubhi, B. K. et al. Metabolic profiling detects biomarkers of protein degradation in COPD patients. Eur Respir J 40, 345–355, https://doi.org/10.1183/09031936.00112411 (2012).
Ubhi, B. K. et al. Targeted metabolomics identifies perturbations in amino acid metabolism that sub-classify patients with COPD. Molecular Bio Systems 8, 3125–3133, https://doi.org/10.1039/C2MB25194A (2012).
Banoei, M. M. et al. Plasma metabolomics for the diagnosis and prognosis of H1N1 influenza pneumonia. Crit Care 21, 97, https://doi.org/10.1186/s13054-017-1672-7 (2017).
Seymour, C. W. et al. Metabolomics in pneumonia and sepsis: an analysis of the GenIMS cohort study. Intensive Care Med 39, 1423–1434, https://doi.org/10.1007/s00134-013-2935-7 (2013).
Weiner, J. III. et al. Biomarkers of inflammation, immunosuppression and stress with active disease are revealed by metabolomic profiling of tuberculosis patients. Plos One 7, e40221, https://doi.org/10.1371/journal.pone.0040221 (2012).
Bos, L. D. J. et al. Exhaled breath metabolomics as a noninvasive diagnostic tool for acute respiratory distress syndrome. European Respiratory Journal 44, 188–197, https://doi.org/10.1183/09031936.00005614 (2014).
Evans, C. R. et al. Untargeted LC-MS metabolomics of bronchoalveolar lavage fluid differentiates acute respiratory distress syndrome from health. J Proteome Res 13, 640–649, https://doi.org/10.1021/pr4007624 (2014).
Haraguchi, H. Metallomics: the history over the last decade and a future outlook. Metallomics: integrated biometal science 9, 1001–1013, https://doi.org/10.1039/c7mt00023e (2017).
Profrock, D. & Prange, A. Inductively coupled plasma-mass spectrometry (ICP-MS) for quantitative analysis in environmental and life sciences: a review of challenges, solutions, and trends. Applied spectroscopy 66, 843–868, https://doi.org/10.1366/12-06681 (2012).
Olenick, M., Flowers, M. & Diaz, V. J. US veterans and their unique issues: enhancing health care professional awareness. Adv Med Educ Pract 6, 635–639, https://doi.org/10.2147/AMEP.S89479 (2015).
Weljie, A. M., Dowlatabadi, R., Miller, B. J., Vogel, H. J. & Jirik, F. R. An Inflammatory Arthritis-Associated Metabolite Biomarker Pattern Revealed by1H NMR Spectroscopy. Journal of Proteome Research 6, 3456–3464, https://doi.org/10.1021/pr070123j (2007).
Clasquin, M. F., Melamud, E. & Rabinowitz, J. D. LC-MS Data Processing with MAVEN: A Metabolomic Analysis and Visualization Engine. Current protocols in bioinformatics/editoral board, Andreas D. Baxevanis… [et al.] 0 14, Unit14.11-Unit14.11, https://doi.org/10.1002/0471250953.bi1411s37 (2012).
Xia, J., Sinelnikov, I. V., Han, B. & Wishart, D. S. MetaboAnalyst 3.0–making metabolomics more meaningful. Nucleic Acids Res 43, W251–257, https://doi.org/10.1093/nar/gkv380 (2015).
Wanichthanarak, K., Fan, S., Grapov, D., Barupal, D. K. & Fiehn, O. Metabox: A Toolbox for Metabolomic Data Analysis, Interpretation and Integrative Exploration. Plos One 12, e0171046, https://doi.org/10.1371/journal.pone.0171046 (2017).
Eriksson, L., Byrne, T., Johansson, E., Trygg, J. & Wikström, C. Multi- and Megavariate Data Analysis Basic Principles and Applications, Third revised edition (Umetrics Academy, 2013).
Eriksson, L., Trygg, J. & Wold, S. CV-ANOVA for significance testing of PLS and OPLS® models. Journal of Chemometrics 22, 594–600 (2008).
Galindo‐Prieto, B., Eriksson, L. & Trygg, J. Variable influence on projection (VIP) for orthogonal projections to latent structures (OPLS). Journal of Chemometrics 28, 623–632 (2014).
Xia, J. & Wishart, D. S. Using MetaboAnalyst 3.0 for Comprehensive Metabolomics Data Analysis. Current protocols in bioinformatics 55, 14.10.11–14.10.91, https://doi.org/10.1002/cpbi.11 (2016).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome research 13, 2498–2504, https://doi.org/10.1101/gr.1239303 (2003).
Sidiropoulos, K. et al. Reactome enhanced pathway visualization. Bioinformatics 33, 3461–3467, https://doi.org/10.1093/bioinformatics/btx441 (2017).
Weljie, A. M., Newton, J., Mercier, P., Carlson, E. & Slupsky, C. M. Targeted profiling: quantitative analysis of 1H NMR metabolomics data. Anal Chem 78, 4430–4442, https://doi.org/10.1021/ac060209g (2006).
Deng, P. et al. Application of metabolomics to characterize environmental pollutant toxicity and disease risks. Reviews on environmental health 34, 251–259, https://doi.org/10.1515/reveh-2019-0030 (2019).
Vermeulen, R. The Use of High-Resolution Metabolomics in Occupational Exposure and Health Research. Annals of work exposures and health 61, 395–397, https://doi.org/10.1093/annweh/wxx016 (2017).
Walker, D. I. et al. Metabolomic assessment of exposure to near-highway ultrafine particles. Journal of exposure science & environmental epidemiology 29, 469–483, https://doi.org/10.1038/s41370-018-0102-5 (2019).
Yan, Q. et al. Maternal serum metabolome and traffic-related air pollution exposure in pregnancy. Environment international 130, 104872, https://doi.org/10.1016/j.envint.2019.05.066 (2019).
Geamanu, A., Gupta, S. V., Bauerfeld, C. & Samavati, L. Metabolomics connects aberrant bioenergetic, transmethylation, and gut microbiota in sarcoidosis. Metabolomics: Official journal of the Metabolomic Society 12, https://doi.org/10.1007/s11306-015-0932-2 (2016).
Duchemann, B. et al. Nuclear magnetic resonance spectroscopic analysis of salivary metabolome in sarcoidosis. Sarcoidosis, vasculitis, and diffuse lung diseases: official journal of WASOG 33, 10–16 (2016).
Mirsaeidi, M. et al. Plasma metabolomic profile in fibrosing pulmonary sarcoidosis. Sarcoidosis, vasculitis, and diffuse lung diseases: official journal of WASOG 33, 29–38 (2016).
Steele, L. Prevalence and patterns of Gulf War illness in Kansas veterans: association of symptoms with characteristics of person, place, and time of military service. American journal of epidemiology 152, 992–1002, https://doi.org/10.1093/aje/152.10.992 (2000).
Naviaux, R. K. et al. Metabolic features of Gulf War illness. PLoS One 14, e0219531, https://doi.org/10.1371/journal.pone.0219531 (2019).
Abdullah, L. et al. Chronic elevation of phosphocholine containing lipids in mice exposed to Gulf War agents pyridostigmine bromide and permethrin. Neurotoxicology and teratology 40, 74–84, https://doi.org/10.1016/j.ntt.2013.10.002 (2013).
Emmerich, T. et al. Phospholipid profiling of plasma from GW veterans and rodent models to identify potential biomarkers of Gulf War Illness. PLoS One 12, e0176634, https://doi.org/10.1371/journal.pone.0176634 (2017).
Catinon, M. et al. Sarcoidosis, inorganic dust exposure and content of bronchoalveolar lavage fuid: The minasarc pilot study. Sarcoidosis Vasculitis and Diffuse Lung Diseases 35, 327–332 (2018).
Brito-Zeron, P., Kostov, B., Superville, D., Baughman, R. P. & Ramos-Casals, M. Geoepidemiological big data approach to sarcoidosis: geographical and ethnic determinants. Clinical and experimental rheumatology (2019).
Hu, J. Z. et al. Metabolomics in lung inflammation: A high-resolution 1H NMR study of mice exposedto silica dust. Toxicology Mechanisms and Methods 18, 385–398 (2008).
Chaudhary, K. & Madaio, M. P. Amino acid limitation stress response in inflammation. Translational Cancer Research 5, 220–222 (2016).
McGaha, T. L. et al. Amino acid catabolism: a pivotal regulator of innate and adaptive immunity. Immunological reviews 249, 135–157, https://doi.org/10.1111/j.1600-065X.2012.01149.x (2012).
Sikalidis, A. K. Amino Acids and Immune Response: A Role for Cysteine, Glutamine, Phenylalanine, Tryptophan and Arginine in T-cell Function and Cancer? Pathology & Oncology Research 21, 9–17, https://doi.org/10.1007/s12253-014-9860-0 (2015).
Sawahata, M. & Sugiyama, Y. An epidemiological perspective of the pathology and etiology of sarcoidosis. Sarcoidosis, vasculitis, and diffuse lung diseases: official journal of WASOG 33, 112–116 (2016).
Newman, K. L. & Newman, L. S. Occupational causes of sarcoidosis. Curr Opin Allergy Clin Immunol 12, 145–150, https://doi.org/10.1097/ACI.0b013e3283515173 (2012).
Newman, L. S. Metals that cause sarcoidosis. Semin Respir Infect 13, 212–220 (1998).
Redline, S., Barna, B. P., Tomashefski, J. F. Jr. & Abraham, J. L. Granulomatous disease associated with pulmonary deposition of titanium. Br J Ind Med 43, 652–656 (1986).
Skelton, H. G. III. et al. Zirconium granuloma resulting from an aluminum zirconium complex: a previously unrecognized agent in the development of hypersensitivity granulomas. J Am Acad Dermatol 28, 874–876 (1993).
Freije, A. M. Heavy metal, trace element and petroleum hydrocarbon pollution in the Arabian Gulf: Review. Journal of the Association of Arab Universities for Basic and Applied Sciences 17, 90–100, https://doi.org/10.1016/j.jaubas.2014.02.001 (2015).
Alaani, S., Tafash, M., Busby, C., Hamdan, M. & Blaurock-Busch, E. Uranium and other contaminants in hair from the parents of children with congenital anomalies in Fallujah. Iraq. Conflict and Health 5, 15, https://doi.org/10.1186/1752-1505-5-15 (2011).
Brown, M. Toxicological assessments of Gulf War veterans. Philos Trans R Soc Lond B Biol Sci 361, 649–679, https://doi.org/10.1098/rstb.2006.1825 (2006).
Das, K. K., Das, S. N. & Dhundasi, S. A. Nickel, its adverse health effects & oxidative stress. The Indian journal of medical research 128, 412–425 (2008).
Oberdorster, G. Pulmonary effects of inhaled ultrafine particles. International archives of occupational and environmental health 74, 1–8 (2001).
Gomez-Ariza, J. L., Jahromi, E. Z., Gonzalez-Fernandez, M., Garcia-Barrera, T. & Gailer, J. Liquid chromatography-inductively coupled plasma-based metallomic approaches to probe health-relevant interactions between xenobiotics and mammalian organisms. Metallomics: integrated biometal science 3, 566–577, https://doi.org/10.1039/c1mt00037c (2011).
Li, Y. F., Gao, Y., Chai, Z. & Chen, C. Nanometallomics: an emerging field studying the biological effects of metal-related nanomaterials. Metallomics: integrated biometal science 6, 220–232, https://doi.org/10.1039/c3mt00316g (2014).
Li, Y.-F. et al. In Pure and Applied Chemistry 80 2577 (2008).
Svendsen, E. R. et al. Reduced Lung Function in Children Associated with Cesium 137 Body Burden. Annals of the American Thoracic Society 12, 1050–1057, https://doi.org/10.1513/AnnalsATS.201409-432OC (2015).
Abdeen, Z. et al. Spatial and temporal variation in fine particulate matter mass and chemical composition: the Middle East Consortium for Aerosol Research Study. ScientificWorldJournal 2014, 878704, https://doi.org/10.1155/2014/878704 (2014).
Lyles, M. et al. Medical Geology: Dust Exposure and Potential Health Risks in the Middle East. (2013).
Rafnsson, V., Ingimarsson, O., Hjalmarsson, I. & Gunnarsdottir, H. Association between exposure to crystalline silica and risk of sarcoidosis. Occup Environ Med 55, 657–660 (1998).
Schweitzer, M. D. et al. Lung health in era of climate change and dust storms. Environ Res 163, 36–42, https://doi.org/10.1016/j.envres.2018.02.001 (2018).
Kalinich, J. F. & Kasper, C. E. Are Internalized Metals a Long-term Health Hazard for Military Veterans? Public Health Reports 131, 831–833, https://doi.org/10.1177/0033354916669324 (2016).
Joseph, P. Mechanisms of cadmium carcinogenesis. Toxicology and applied pharmacology 238, 272–279, https://doi.org/10.1016/j.taap.2009.01.011 (2009).
Lettvin, J. Y., Pickard, W. F., McCulloch, W. S. & Pitts, W. A Theory of Passive Ion Flux through Axon Membranes. Nature 202, 1338–1339 (1964).
Curtis, M. J., Quastel, D. M. & Saint, D. A. Lanthanum as a surrogate for calcium in transmitter release at mouse motor nerve terminals. J Physiol 373, 243–260 (1986).
Mirsaeidi, M. et al. Annexin A11 is associated with pulmonary fibrosis in African American patients with sarcoidosis. Sarcoidosis Vasc Diffuse Lung Dis 33, 418–422 (2016).
Jin, Z., Mendu Sk Fau - Birnir, B. & Birnir, B. GABA is an effective immunomodulatory molecule. Amino Acids, 45, 87–94 (2013).
Acknowledgements
Authors would think all our patients and veterans for participating in this study. This work was supported in part by Health Resources and Services Administration contract 234-2005-37011C, and Mallinckrodt Pharmaceuticals for research grant support. The funder (Mallinckrodt Pharmaceuticals) was not involved in the study design, collection, analysis, interpretation of data, the writing of this article or the decision to submit it for publication.
Author information
Authors and Affiliations
Contributions
M.M.B. contributed to metabolomics and metallomics profiling analysis, data collection, statistical analysis interpretation and drafting the manuscript. I.I. contributed to samples collection and drafting the manuscript. R.D.B. contributed to metallomic analysis, manuscript revision and interpretation. M.C. contributed to manuscript revision and interpretation. H.J.V. contributed to manuscript revision and interpretation. B.W.W. contributed to manuscript revision and interpretation. M.M. contributed to the study concept and design, interpretation, drafting the manuscript and approved the final version. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Banoei, M.M., Iupe, I., Bazaz, R.D. et al. Metabolomic and metallomic profile differences between Veterans and Civilians with Pulmonary Sarcoidosis. Sci Rep 9, 19584 (2019). https://doi.org/10.1038/s41598-019-56174-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-56174-8
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.