Abstract
Radical-Induced Cell Death1 (RCD1) functions as a cellular hub interacting with intrinsically disordered transcription factor regions, which lack a well-defined three-dimensional structure, to regulate plant stress. Here, we address the molecular evolution of the RCD1-interactome. Using bioinformatics, its history was traced back more than 480 million years to the emergence of land plants with the RCD1-binding short linear motif (SLiM) identified from mosses to flowering plants. SLiM variants were biophysically verified to be functional and to depend on the same RCD1 residues as the DREB2A transcription factor. Based on this, numerous additional members may be assigned to the RCD1-interactome. Conservation was further strengthened by similar intrinsic disorder profiles of the transcription factor homologs. The unique structural plasticity of the RCD1-interactome, with RCD1-binding induced α-helix formation in DREB2A, but not detectable in ANAC046 or ANAC013, is apparently conserved. Thermodynamic analysis also indicated conservation with interchangeability between Arabidopsis and soybean RCD1 and DREB2A, although with fine-tuned co-evolved binding interfaces. Interruption of conservation was observed, as moss DREB2 lacked the SLiM, likely reflecting differences in plant stress responses. This whole-interactome study uncovers principles of the evolution of SLiM:hub-interactions, such as conservation of α-helix propensities, which may be paradigmatic for disorder-based interactomes in eukaryotes.
Similar content being viewed by others
Introduction
Most functional proteins fold into well-defined structures. However, 30–50% of eukaryotic proteins contain large regions of intrinsic disorder (ID)1,2 allowing function without well-defined folds3,4,5,6,7,8. Intrinsically disordered proteins (IDPs) and -regions (IDRs) often participate in protein-protein interaction networks (interactomes), which govern key functions such as transcription and cell-cycle regulation9,10,11,12. Accordingly, transcription factors (TFs) contain high fractions of functionally essential IDRs13,14. IDPs evolve faster and are more permissive to substitutions than folded proteins15,16. However, sites within IDRs with secondary structure propensities are evolutionary more constrained than sites within secondary structures17. Such regions may coincide with short linear motifs (SLiMs), originally discovered as islands of taxonomic conservation within rapidly evolving regions18,19. SLiMs are regularly gained and lost during evolution making them useful tools for determining clade-specificity19,20. While SLiMs are not limited to IDRs, molecular recognition features (MoRFs) represent a concept developed for disorder-based interactions. MoRFs are structure-prone regions located within long IDRs, which may undergo disorder-to-order transitions upon binding21. Evolutionary studies of the p53 TF family have shown that MoRFs are more conserved than their surrounding regions22. Although several bioinformatics studies have addressed the evolution of IDPs, SLiMs, and MoRFs, most of these studies lack biophysical support and perspectives on the whole interactome.
The plant hub-protein Radical-Induced Cell Death 1 (RCD1) represents a suitable model for studies of interactions between folded hub domains and SLiMs in disordered targets, and allows translation from in vitro to the organismal level23. RCD1 is a member of the plant-specific Similar to Rcd One (SRO) family. SRO proteins contain a Poly (ADP-Ribose) Polymerase (PARP) domain and an RCD1-SRO-TAF4 (RST) domain24, and some also have a WWE domain (Fig. 1A). Arabidopsis thaliana RCD1 (AtRCD1, with At referring to the species-specific origin), uses its helical RST domain (Fig. 1B), a member of the αα-hub family25, for interactions with many TFs26. AtRCD1 plays roles in hormone signaling, responses to reactive oxygen species and other abiotic stress factors, immunity and development26,27,28,29. It interacts with numerous TFs (Fig. 1C)26,30, including Dehydration-Responsive Element-Binding protein (DREB) 2 A, implicated in abiotic stress responses31. Recent studies suggest that AtRCD1 negatively regulates its interaction partners. Thus, downregulation of AtRCD1 or loss of the RCD1-interacting site of DREB2A is required for proper DREB2A function under stress conditions32, and inactivation of RCD1 resulted in increased expression of Arabidopsis No-apical-meristem, Arabidopsis transcription activation factor, Cup-shaped cotyledon (ANAC)013 and ANAC017-regulated genes from the mitochondrial dysfunction stimulon33.
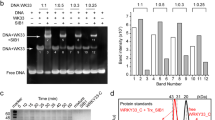
RCD1 structure and interactions. (A) Domain structure (WWE, PARP, and RST domains) of AtRCD1. (B) Sequence alignment of AtRCD1-RST with the most related RST domain from barley, soybean, and moss. α-helix positions in AtRCD1-RST are shown at the top. Top alignment: Conservation of residues with colouring in accordance with percentage identity (Jalview; darker blue is more conserved). Middle top alignment: Colouring in accordance with number of contacts (6 Å cut-off) to other structural elements (helix 1–4 or loop 1–2) in the AtRCD1-RST structure (PDB 5OAO). Dark green is three, green two, and light green one contact. The β3-position of the αL-β4 loop motif is grey. Middle bottom alignment: Residue groups that are in contact in the AtRCD1-RST structure (PDB 5OAO;5N9Q) and appear to co-evolve in same colour. The β3-position of the αL-β4 loop motif is grey, Gly555 in dark grey. Bottom alignment: Conserved residues that cannot be explained by fold conservation are highlighted. Conserved residues are red, while conservative substitutions are light red. (C) Left: TF-interactome of AtRCD1 from the STRING database. Names of TFs, which have been shown to bind the RST domain30, are shown. Right: Expanded AtRCD1:TF interactome predicted based on the expanded SLiM [ED].{1,2}[^RK][YF].{1,4}[^RK][DE]([LIVMF]|.L). (D) Structure of the AtRCD1 RST domain (light grey; PDB 5OAO) in modelled complex with AtDREB2A(255–272) (dark grey; PDB 5OAP)25. AtRCD1-RST key residues (R551, R560, I563) for AtDREB2A(255–272) binding and the α-helix stabilizing hydrophobic staple motif residues, F259 and L264, of AtDREB2A(255–272), are highlighted. (E) NMR solution structure of AtRCD1-RST (light grey; PDB 5N9Q) and models (light grey) of the barley, soybean, and moss RST domains generated using I-TASSER70. Left: Superposition of structure and models with residues having three or two contacts to other structural elements in dark green sticks and green lines, respectively. Middle: Superposition of structure and models with conserved residues not explained by fold conservation shown as sticks. Fully conserved residues are red while positions with conservative substitutions are light red. Right: Structure of AtRCD1-RST (light grey; PDB 5N9Q) with same residue highlights as the superpositions (left and middle).
Recently, the consensus motif [ED].{1,2}[^RK][YF].{1,4}[^RK][DE]L was identified as the RCD1-binding SLiM (hereafter referred to as RBS) positioned in IDRs of RCD1 target TFs30. Interestingly, different structures, containing this motif, may be formed in the TFs upon association with the RST domain of RCD1. While α-helix is induced in DREB2A (Fig. 1D), no helical structure has been observed in ANAC046 and ANAC013 upon complex formation30 suggestive of structural plasticity in the RCD1-interactome.
In this study, the evolution of the RCD1:TF interactome was analyzed. This revealed evolutionary conservation of RCD1, the RBS and the TF order-disorder patterns in land plants over a period of 480 million years. Both cross-species interactions and molecular co-adaptation in high-affinity RCD1:TF interactions were observed. Interruption of conservation in moss DREB2, which does not contain the SLiM, may reflect differences in plant stress responses. Our whole-interactome approach, spurred by ID profiles and motif based interactions in a molecularly intriguing interactome, is readily transferrable to other ID-based interactomes.
Results
Taxonomic conservation of the SRO family
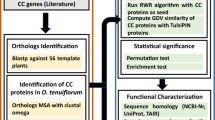
Ortholog identification, which is generally accepted as a proxy for identification of genes with similar functions in different species34, was used for analysing the evolution of the SRO:TF interactome. Sequences were obtained from PLAZA, developed specifically for comparative genomics in plants with many paralogs34,35. Since SRO proteins have been identified only in land plants, these were the focus of this study (Fig. 2A; Supplementary Table S1)36,37. The moss Physcomitrella patens was the only non-vascular plant included. Selaginella moellendorffii (Fig. 2A; Supplementary Table S1) represents lycophytes, whereas Cycas micholitzii, Ginkgo biloba, Gnetum montanum and Picea abies represent gymnosperms (Fig. 2B). In addition, Amborella trichopoda, a living ancestor of the sister lineage to all other living angiosperms, Oryza sativa ssp. japonica, Zea mays, and Hordeum vulgare representing different monocot clades (Fig. 2C), and eight dicots were included. Comparison of Arabidopsis thaliana and Arabidopsis lyrata reveals recent evolutionary changes, whereas the other dicots Eucalyptus grandis, Populus trichocarpa, Fragaria vesca, Vitis vinifera, Gossypium Raimondii, and Glycine max were selected to represent diversity. SRO proteins have been identified in all of these species, except six, which were analysed here (Supplementary Table S2). Several sequences obtained using Integrative Orthology Viewer were incomplete, and only sequences with PARP and RST domains were used for further analysis. The number of identified SRO proteins varied from none for Gnetum montanum to four for Amborella trichopoda, Zea mays, and Ginkgo biloba, and Gnetum montanum was, therefore, not included in this study (Fig. 2B; Supplementary Table S2). Searching pico-PLAZA 2.0, which represents eukaryotic microorganisms, confirmed that SRO proteins are specific to land plants.
Phylogenetic relationships of plants analysed for RCD1-interacting motif in transcription factors with the domain structure shown. (A) Evolution of the plant kingdom showing splitting of land plants into nonvascular and vascular plants, and further into seedless and seed plants. (B) Phylogenetic relationship of gymnospermea (non-flowering) species (modified from PLAZA34,35,72). (C) Phylogenetic relationship of angiospermea (flowering) species (modified from PLAZA34,35,72). Species included in this study are highlighted in red. (D) Domains and conserved sequence patterns for NAC013/016/017, DREB2, NAC046, bZIP23, BBX8 and STO TFs. The DBDs are grey, sequence patterns conserved in land plants are black, while sequence patterns conserved specifically in angiosperms are hatched. The positions of the RBS are marked by a red arrow. Black lines below boxes indicate regions where ELMs overlap with the SLiM. TM: transmembrane region. The figures are not drawn to scale.
The structure of AtRCD1-RST consists of four α-helices organized in an L-glove and with a super-secondary αα-hairpin structure as its base formed by helices 2 and 325. Molecular modelling using the AtRCD1-RST solution structure as template indicated that the RST domain structure is conserved from moss to different flowering plants (barley and soybean). From sequence alignments of Arabidopsis, moss, barley and soybean RST domains in combination with analysis of the tertiary structure of the AtRST domain25,33,38 it is clear that the residues that make up the central hydrophobic core responsible for the RST-fold are highly conserved (Fig. 1B, middle top). Here, a LIVYLIV motif (At: L-x3-I-x3-V-x11-Y-x16-L-x2-I-V) is responsible for the central core at the closed end of the helix 1–3 assembly (Fig. 1B, middle top;1E, left), and the fold is further supported by a 3xPhe-1xLeu motif at the open end and conserved hydrophobic residues of helix 4 contacting the LIVYLIV network (Fig. 1B, middle top; 1E, left). The sequences align without any insertions or deletions and several residue positions that are in contact in the AtRST structure have co-evolved across the species (Fig. 1B, middle bottom). The stabilizing β3-position of the αL-β4 loop motif of the fundamental αα-hairpin super-secondary structure39 is maintained across the species but alternates between different, favoured residues for this motif position (Met541 in Arabidopsis, Leu in moss, Ile in soybean and barley). Gly555 (residue numbering relative to AtRCD1), the one-residue loop responsible for the angle between helix 3 and 4, is likewise conserved across the species. Taken together, these findings indicate that the RST fold is highly conserved across the species.
It is also clear that there are conserved residues that are not immediately explained by structural conservation (Fig. 1B, bottom). Besides a Pro-motif in the disordered N-terminus, these mainly localize to helix 3 and 4 and are mostly Arg (543, 551) or Arg/Lys (537, 539, 548), but also Glu/Asp (545), Asp (556), Ile/Val (547, 563) and Gln (567). Nonetheless, Lys/Arg537, Lys/Arg548 and Asp556 are charge neutralized by substitutions with Gln, Gln and Ala, respectively, in moss. When plotted onto the structure of AtRST and the models, these residues localize to the rim of the hydrophobic L-glove pocket of the structure where DREB2A binds to the hub (Fig. 1E, middle), which could indicate that they influence binding affinity and specificity.
Identification of TF orthologs and inparalogs
For analysis of evolutionary conservation of the RCD1-interactome, orthologs and inparalogs of the ten Arabidopsis TFs with a functional SLiM30 were identified and aligned. Since the Arabidopsis NAC proteins ANAC013, ANAC016 and ANAC017 (with A in ANAC referring to the Arabidopsis origin) and the Arabidopsis DREB2 proteins AtDREB2A, AtDREB2B and AtDREB2C, respectively, are closely related only one multiple protein sequence alignment was created for each homology group. The sequences generally align well in the DNA-binding domains (DBDs), whereas sequence variation outside these is consistent with ID30,40. However, conserved sequence patterns, which can be regarded as homolog signatures, were found throughout the IDRs (Fig. 2D).
The alignment of the NAC013/016/017 TFs revealed a conserved DBD and various C-terminal regions corresponding to disordered transcriptional activation domains40. Nine short conserved regions (A1–9) were apparent with the RBS mapping to A4 (Fig. 2D). The SLiM was identified in three of the four moss, the three gymnosperm and all of the dicot homologs, and for monocots in one maize, rice (reverse orientation) and barley homolog (Fig. 3A). Although it was lacking from one of the rice and Amborella sequences, these contained a motif with isoleucine substituting leucine conferring to the motif [ED].{1,2}[^RK][YF].{1,4}[^RK][DE][LI]. A search in the eukaryotic linear motif (ELM) database revealed a docking motif for PIKK kinases in 15 dicot homologs, which overlaps with the SLiM.
Conservation of the RBS. (A–F) Sequence alignments based on the experimentally verified RBS from the Arabidopsis TFs ANAC013 (AT1G32870), ANAC016 (AT1G34180), ANAC017 (AT1G34190), AtDREB2A (AT5G05410), AtDREB2B (AT3G11020), AtDREB2C (AT2G40340), ANAC046 (AT3G04060), AtbZIP23 (AT2G16770), AtBBX8 (AT5G48250), and AtSTO (AT1G06040)30 with homologs from different species. The gene names of experimentally verified binders of AtRCD1-RST are underlined. The aligned regions do not represent the exact peptides used in the experimental studies (Table 2). The figure shows the names without the prefix A or At, specifically referring to NAC TFs of Arabidopsis origin, since sequences from many different plant species are shown. The amino acid residues are coloured according to the degree of conservation (Jalview; darker blue is more conserved). For all alignments, SLiM residues are marked with black asterisks in the top and bottom of the alignment based on the experimentally verified binders from each family30. For ANAC087 (At5g18270), the alternative putative SLiM variant is marked with red asterisks. The sequences were obtained from the PLAZA platform34,35,72 for the following species: Physcomitrella patens (PP), Selaginella moellendorffii (SM), Amborella trichopoda (ATR), Oryza sativa ssp. Japonica (OS), Zea mays (ZM), Arabidopsis lyrata (AL), Arabidopsis thaliana (AT), Eucalyptus grandis (EG), Fragaria vesca (FV), Glycine max (GM), Gossysium raimondii (GR), Populus trichocarpa (PT), Vitis vinifera (VV), Ginkgo biloba (GBI), Cycas micholitzii (CMI), Gnetum montanum (GMO) and Picea abies (PAB). The sequences of the plant species Hordeum vugare (HV, BAJ or AAO) were obtained by BLAST homology searches. Extended alignments including non-binders can be found in Supplementary Fig. S1.
In the DREB2 TFs, the DBD is flanked by IDRs with six conserved regions (B1–6; Fig. 2D). In addition to being present in the AtDREB2s30, the SLiM was found in the Amborella ortholog, one of the two Norway spruce orthologs as well as in most of the dicot orthologs (Fig. 3B). It was not identified in the DREB2 TFs from moss, and for monocots, it was only found in barley, explaining the inability of rice OsDREB2A to interact with OsSRO1c41. For several homologs, most of the core residues are present, and in three cases [DE].L terminates the motif suggestive of motif expansion to [ED].{1,2}[^RK][YF].{1,4}[^RK][DE](L|.L). Thus, although the existence of the SLiM is unlikely for non-vascular DREB2s, it is conserved in seed plants.
In the NAC046 TFs, the SLiM is located in the most C-terminal conserved region, C3 (Fig. 2D). It is conserved in the ortholog most closely related to ANAC046, AL3G03610, but it is not generally conserved (Fig. 3C). A tryptophan residue appears to be conserved among the NAC homologs, but Trp332 in ANAC046 was shown to have a less significant effect on RCD1-binding than the RBS-defining residues. Therefore, the conserved tryptophan pattern may reflect the presence of an overlapping activation motif of transcription as previously suggested42. The RBS is present in one of the moss homologs and in homologs from cotton and eucalyptus, but located in the region between C2 and C3 (Supplementary Fig. S1). In this region, ten sequences differ from the SLiM only by the absence of the acidic residue in the first SLiM position, thus conferring to. {1,2}[^RK][YF][ED]L. In total, sequence patterns identical or similar to the SLiM are observed in 13 of 27 NAC046 homologs within or close to C3.
For the bZIP23 TFs with five conserved regions, the SLiM maps to D2 (Fig. 2D) in six of the plant species, and [ED].{1,2}[^RK][YF].{1,4}[^RK][DE]([LI]|.L) is present in bZIP23 TFs from several species including moss and spikemoss, whereas other homologs have motifs described by.{1,2}[^RK][YF].{1,4}[^RK][DE]L or [ED].{1,2}[^RK][YF].{1,4}[^RK][DE][FIMV] (Fig. 3D). Thus, the SLiM and single residue variants were identified in all species except Norway spruce. The phosphorylation site ELM MOD_CK1_1 maps to D2 of the bZIP23 TFs.
Four conserved regions were identified for the BBX8 TFs with the SLiM mapping to E2 (Fig. 2D). In all dicot BBX8 TFs, the SLiM is bi-directional because of conservation of an additional leucine (Fig. 3E). It is lacking from three moss homologs and the Norway spruce homolog which nonetheless have the expanded SLiM [ED].{1,2}[^RK][YF].{1,4}[^RK][DE][LIM], suggestive of conservation across land plants.
The SLiM maps to F3 in the STO TFs (Fig. 2D). It is present in half of the moss STOs, but missing from the gymnosperm STOs (Fig. 3F). However, several of these contain the expanded SLiM [ED].{1,2}[^RK][YF].{1,4}[^RK][DE][LIV]. The N-terminal conserved region of the bipartite F3 region is specific for angiosperm STO TFs, whereas a second conserved region is located at the C-termini in mosses and gymnosperms (Supplementary Fig. S1). The degradation ELM DEG_COP1_1 is present in most of the sequences and overlaps with the SLiM.
Non-binding motifs
AtMYB91 and AtDREB2A contain a motif that meets the criteria of the RBS, but is not able to bind RCD1-RST30. In AtMYB91, the non-binding motif is present at the N-terminal border of a conserved region in five additional MYB91 homologs, but for the rest of the homologs, the SLiM is absent. Similarly, the non-binding motif of AtDREB2A is located in a non-conserved region. Therefore, and importantly, non-binding SLiMs differ from binding-competent SLiMs by being non-conserved (Supplementary Fig. S1). Thus, conservation can be used as an additional filter for the identification of functional RBS regions.
Conservation of ID patterns and MoRFs
Disorder-order patterns have been suggested to be more constrained than sequences43, and for different NAC sub-groups the ID profiles are well conserved44. The ID profiles of the NAC013/016/017, DREB2 and NAC046 TFs were analyzed using IUPRED, which allows disorder prediction for multiple sequences45. In the TFs, the DBDs have similar ID-profiles reflecting structured domains (Fig. 4A–C). In contrast, the profiles vary for the IDRs, although similarities were also apparent. The RBS maps to a dip in the NAC013/016/017, DREB2 and NAC046 disorder profiles suggestive of local structure propensities. The long disordered NAC046 C-termini with sparse sequence motifs (Fig. 2D) and secondary structures42 provide a conserved flexible platform for interactions mediated by the RBS at the very C-termini. About half of the conserved regions of the three TF groups also map to dips in the disorder profiles. The TFs were also analyzed for MoRF conservation. For the NAC013/016/017 and the NAC046 TFs, a MoRF was predicted for the RBS region in more than 50% of the TFs. However, MoRFs were not generally conserved for the motif regions. To conclude, similarities were observed in the ID-profiles of the long IDRs with the RCD1-binding SLiM mapping to regions with local structure propensity.
ID and MoRF predictions for RCD1-interacting TFs. (A–C) IUPRED ID predictions for the NAC013/016/017, DREB2 and NAC046 homologs made from Jalview alignments of the sequences included in Fig. 3. The IUPRED scores of the homologues protein sequences are plotted for each alignment position by the minimum and maximum (grey curves), the median (blue curve) and the interquartile range as the (red) area between the 1. and 3. quartiles. The threshold of 0.5 for ID is marked by a line. Blue bars represent positions for which MoRFs were predicted for more than 50% of the aligned homologs using MoRFpred. Black bars show the positions of conserved sequence motifs. Orange and grey bars show the positions of the RBS and DBD, respectively.
Expanding the RBS
The results indicate that the RBS, [ED].{1,2}[^RK][YF].{1,4}[^RK][DE]L, is conserved among the NAC013/016/017, DREB2, STO and BBX8 homologs with some exceptions. It is missing from the moss DREB2 TFs, whereas the lack of spikemoss homologs and SLiMs may be explained by the quality of the sequence data for this organism. For the NAC046 and bZIP23 TFs, the SLiM is only present in a few species, and it is not completely conserved for any of the TF groups. However, most of its core residues are conserved, and three alternative SLiM versions (variant 1–3, referred to as RBS-V1 - V3) (Table 1) are apparent. These differ from the original RBS only at one position and are in accordance with the SLiM-discriminatory features of a central aromatic residue, pI value below 4.530 and conserved location in the parent protein. RBS-V1 lacks the acidic residue in the first SLiM position, in RBS-V2 the C-terminal leucine is substituted with phenylalanine, isoleucine, methionine or valine, and in RBS-V3 an additional residue is inserted before the C-terminal leucine. The original SLiM was identified in 99 homologs and is more frequent than the alternative versions. However, variant 1 is most frequent among the NAC046 TFs, while variant 2 and 3 are most frequent among the bZIP23 TFs. The frequent alternative motifs are all described by the expanded motif [ED].{1,2}[^RK][YF].{1,4}[^RK][DE]([LIVMF]|.L). This SLiM, which would increase the number of TFs in the RCD1-interactome from 21 to 166 (Fig. 1C; Supplementary Table S3), is found across all lineages of land plants indicating conservation of the pattern. Although DREB2 and STO TFs were identified in microorganisms, these TFs lack the SLiM in accordance with a main role of the SLiM being RCD1-binding.
Experimental verification of the expanded RBS
RBS-V2 (Table 1) with isoleucine instead of leucine in the last SLiM position is found frequently among the TF homologs (Fig. 3). Since isoleucine destabilizes α-helices, this substitution could affect RCD1-binding of the NAC and DREB2 TFs differently, as they likely differ in their bound structure30. α-helical conformation was predicted for residues 261 to 270 of AtDREB2A(244–272), and the Leu264 to Ile substitution resulted in only a slight overall decrease in helical propensity (Fig. 5A). A similar observation was made by circular dicroism (CD) spectroscopy in studies using trifluoroethanol (TFE) to probe for helical propensity (Fig. 5B). In buffer, a low population of α-helix was measured for both peptides, but addition of TFE to 10% had a slightly larger α-helix inducing effect on AtDREB2A(244–272) than on AtDREB2A(244–272; L264I). At 40% TFE, both spectra displayed the characteristics of α-helix, moving the minima towards 222 and 208 nm (Fig. 5B) indicating that the isoleucine variant is able to form α-helix to a similar extent as the wildtype peptide. For ANAC013(254–274)30 and ANAC013(254–274;L266I) the predicted level of α-helical conformation was low (Fig. 5A), and CD spectra suggested that the two peptides contain similar levels of α-helical structure (Fig. 5B).
Experimental verification of SLiM variants. (A) Left: Helicity per residue predicted by Agadir for AtDREB2A(244–272) and AtDREB2A(244–272; L264I) (top) and sequence of AtDREB2A(244–272) (bottom) with the N’ and N4 residues of the hydrophobic staple motif marked and the α-helix formed in complex with RCD1 underlined. Right: Helicity per residue predicted by Agadir for ANAC013(254–274; L266I). (B) Far-UV CD spectra of 15‒20 µM TF peptide as indicated above the spectra in 10 mM Na2HPO4/NaH2PO4, pH 7.0, and 0–40% (v/v) TFE. Data for ANAC013(254–274) has been reported previously30. (C) Representative ITC measurements, here shown for the RCD1-RST(499–572) interactions with AtDREB2A(244–272), AtDREB2A(244–272;L264I), and AtANAC013(254–274;L266I). The AtANAC013(254–274):AtRCD1-RST(499–572) interaction was published previously30. AtRCD1-RST(499–572) was titrated into the TF fragments. In each panel, the upper portion shows baseline-corrected raw data from the titration, and the lower portion shows the normalized integrated binding isotherms together with the fitted binding curves. The data were fitted to a “one set of sites” binding model. Parameters obtained from the non-linear fits are presented in Table 2.
Isothermal titration calorimetry (ITC) was used to analyse how Leu→Ile substitution affects binding to RCD1. AtDREB2A(244–272) has high affinity (Kd 16 nM) for AtRCD1-RST(499–572) and was not significantly affected by the change of Leu264 to Ile (Table 2; Fig. 5C). This is in accordance with structure analysis indicating that Asp260 forms the N-cap (first α-helix position) of the AtDREB2A α-helix, which contains a hydrophobic staple motif46 with N’ and N4 interacting and stabilizing the α-helix by strengthening capping (Figs. 1D; 5A). In addition to Leu and Ile, Phe and Val, which are frequent in position N4 (Figs. 3B; 5A), also mediate α-helix stabilization by staple motif formation46 suggestive of evolutionary conservation of coupled folding and binding for DREB2 association with RCD1. Leu266 to Ile substitution in ANAC013(254–274) resulted in an approximately 20-fold decrease in affinity (Kd 223 nM) (Fig. 5C; Table 2), suggesting that β-branching may result in steric hindrance in the RCD1-ANAC013 complex. However, both the AtDREB2A and the ANAC013 variants bound RCD1-RST(499–572) strongly, confirming that isoleucine, and other hydrophobic residues, are allowed at this last SLiM position, as suggested by the SLiM variants (Table 1).
Conservation of RCD1-binding in functionally redundant NAC TFs
For the NAC046 TFs, SLiM conservation was less clear than for the other TF groups, with the RBS-V1.{1,2}[^RK][YF].{1,4}[^RK][DE][LI] suggested from the alignment (Fig. 3C; Table 1). Focusing on the NAC046 TFs in Arabidopsis, this motif is present in ANAC087 (At5g18270) (Fig. 3C), the inparalog most closely related to ANAC04640. Both ANAC046 and ANAC087 are induced by abiotic stress40,42 and these genes redundantly control the onset of cell death execution47. ANAC087(315–335) bound AtRCD1-RST(499–572) with a relatively weak affinity corresponding to Kd 1751 nM (Table 2). This was also the case for ANAC046:RCD1-RST association (Kd 609 nM)42 and in both cases entropy significantly contributed to binding. To address if the RST domain uses the same residues for binding of ANAC087 as for AtDREB2A (Fig. 1D)25, binding of ANAC087(315–335) to the double substitution variant AtRCD1-RST(499–572;R551Q;R560Q) was analyzed. R551 is conserved without having an essential role in the RST-fold, while R560 varies (Fig. 1B,E). No binding was observed for ANAC087 (Table 2) as in the case of AtDREB2A25. The single substitution variants AtRCD1-RST(499–572;I563Q) and AtRCD1-RST(499–572;R560Q), with no and compromised AtDREB2A(255–272) binding, respectively (Fig. 1D)25, did not bind ANAC087(315–335). Similar to R551, I563 is chemically conserved despite no structural involvement in the RST fold, thus suggesting that other functional constraints warrant conservation (Fig. 1B,E). In conclusion, ANAC087 binds RCD1 and shares key RCD1-target residues with AtDREB2A.
Evolutionary conservation of the RCD1-TF system
Evolutionary conservation of the RCD1-TF system was also addressed, both by predictions and experimentally. Predictions of the α-helix propensities of the SLiM regions from different species suggested a range of α-helix propensities for the DREB2 TFs (Fig. 6A). Up to 47% α-helix was predicted for the SLiM region in barley HvDRF1.1, and CD experiments suggested a slightly higher intrinsic population of α-helix in HvDRF1.1(255–273) (16%) than in the AtDREB2A (12%) (Fig. 5B). Several DREB2A SLiM regions have the same level of α-helix propensity as AtDREB2A (max. 20%), and some SLiM regions such as those from soybean, GM02G42960 and AtDREB2B and AtDREB2C, which bound AtRCD130, have lower α-helix propensities (max 4%). Stress-associated HvDRF1.148 is the only barley DREB2 homolog with a SLiM (Fig. 3B). HvDRF1.1(255–273) bound AtRCD1-RST(499–572) with Kd 450 nM (Table 2) suggesting that the DREB2A-binding site of a dicot RST domain can accommodate DREB2A from an evolutionary distant monocot. The barley ortholog of ANAC046, HvNAC013(176–346), which in the yeast two-hybrid system bound HvRCD1-RST49, also bound AtRCD1-RST(499–572), further indicating that features of the RCD1-TF interactome are evolutionary constrained. No binding of HvDRF1.1(255–273) to HvHis6-RCD1-RST(467–579) was detected. This may be explained by the lack of a positively charged residue in the position corresponding to Arg-560 of AtRCD1 (Fig. 1B,C), which is required for its association with AtDREB2A25. The low α-helix propensities of ANAC013 and ANAC04630 are evolutionary conserved among the NAC013/016/017 and NAC046 TFs (Fig. 6B,C).
Evolution of the RCD1:TF system. (A–C) Helicity per residue predicted by Agadir for the DREB2, NAC013/016/017 and NAC046 SLiM regions from different plant species. (D) Model of the RCD1:TF interactome based on the structures of the AtRST domain (PDB 5OAO) and AtDREB2A(255–272) (PDB 5OAP) from the AtRCD1-RST:AtDREB2A complex. The different RCD1 interacting TFs compete for binding to RCD1. The TFs have different helical propensities and may form different structures in their bound states. Complexes with ANAC013 and ANAC046 were modelled using the CABS-dock server. The basic residues R551 and R560 and the hydrophobic residue I563, which are of key importance to TF-binding, are highlighted as blue and orange sticks, respectively. The schematic cartoon displays the typical binding interface between RCD1-RST and the TF activation domains. Activation domains are enriched in acidic and hydrophobic residues that interact with basic and hydrophobic contacts within the RCD1-RST binding pocket. The cartoon is based on the modelled complex between AtRCD1-RST and AtDREB2A25.
Conservation of the RCD1:TF system was also analysed by testing for conserved interactions within other species. The RCD1:TF interactome is most conserved in dicots, especially eucalyptus and soybean (Eucalyptus grandis and Glycine max, respectively) (Figs. 2C, 3), with the SLiM found in all TF groups analysed. The RST domain from the soybean RCD1 ortholog, GmRCD1, in which AtRCD1 key residues for AtDREB2A binding are conserved (Fig. 1B), indeed bound both AtDREB2A(244–272) and ANAC013(254–274) with affinities in the mid-nanomolar range (Table 2). When using one of two soybean DREB2 TFs containing the SLiM (Fig. 3B) in BLAST searches for Arabidopsis proteins, AtDREB2A appeared as a top hit. The GM02G42960 peptide GmDREB2A(273–290), bound GmRCD1-RST(483–583) with an affinity of Kd approximately 2500 nM and the Arabidopsis RST domain, AtRCD1-RST(499–572), with an affinity of Kd approximately 400 nM (Table 2).
Discussion
This study demonstrates that the RBS of the ID-based RCD1:TF interactome arose in the land plant lineage 480 mill. years ago (Fig. 2A) and is evolutionary highly conserved. TF from microorganisms do not contain the SLiM further linking it to interactions with RCD1, also only found in land plants37,41. Thus, the RCD1:interactome has evolved to cope with land plant life. Confirming their important functions, 13 of 15 Arabidopsis thaliana TFs and their SLiMs are conserved in Arabidopsis lyrata (Fig. 3), which cluster with Arabidopsis thaliana (Fig. 2C). The SLiM is highly conserved in flowering plants (angiosperms), and for most TF groups the SLiM - especially the expanded [ED].{1,2}[^RK][YF].{1,4}[^RK][DE]([LIVMF]|.L) version - was also traced to the ancient sister seed plant clade, gymnosperms (Fig. 2B). It was also identified in all TF groups, except DREB2 and BBX8, from moss. This likely reflects a different defense system in mosses, although the expression of a couple of AP2/ERF transcription factors is affected by abiotic stress in mosses50. Structurally, this could be connected to the charge neutralization of Lys/Arg537, Lys/Arg548 and Asp556 (AtRCD1 numbering) by substitution with Gln, Gln and Ala, respectively, in the moss RST domain (Fig. 1B). Conservation of the SLiM over a large taxonomic range indicates that it is constrained and therefore functional51.
Because of their small size and low sequence complexity, SLiMs easily evolve de novo19. The RBS was identified in six unrelated TF families30 making de novo evolution at multiple occasions followed by convergence likely suggestive of common regulatory tasks52. Accordingly, the interactions of AtDREB2A, ANAC013, ANAC016, and ANAC017 with AtRCD1 result in down-regulation of their target genes32,33 pointing to a general regulatory function of the RBS.
The TFs have several conserved sequence regions outside their DBDs (Fig. 2D). Most of these are present in species across different taxa of land plants, but some are specific for angiosperms suggestive of functions related to higher plants. For all TF groups, the SLiM maps to a conserved region and overlaps with an ELM in NAC013/016/017 (A4), bZIP23 (D2) and STO (F3) (Fig. 2D). The degron, DEG_COP1_1, present in the STO TFs, is interesting because it is the site for interactions with the E3 ubiquitin ligase COP1 mediating regulated TF degradation53. Overlapping SLiMs reflect the conformational adaptability of IDRs allowing interactions with several distinct binding partners. The ID profiles of NAC013/016/017, DREB2, and NAC046 TFs from different taxa revealed complex patterns but also group-specific characteristic (Fig. 4). That the SLiMs map to dips in the disorder profiles indicate conserved local structure propensities though only a few predicted MoRFs appear to be conserved among the NAC013/016/017, DREB2 and NAC046 TFs (Fig. 4). ID-based dendrograms have been suggested as tools for studies of distant relationships among proteins43, and such use is supported by the ID-profiles shown in Fig. 4.
The SLiM is least conserved in the NAC046 TFs. ANAC087 is expressed by the same abiotic stress factors as ANAC04640, and the two NAC TFs are functionally redundant47. ANAC087(315–335), which contains the SLiM variant RBS-V1 (Table 1; Fig. 3C), was able to associate with AtRCD1-RST(499–572) (Table 2). Although its RCD1-affinity is relatively low, motif contexts may affect binding affinity significantly30,42. Furthermore, the RCD1-interaction partners AtDREB2A, ANAC013, ANAC087 and ANAC046 are induced by various abiotic stress parameters28,31,40,47, which will affect in vivo binding and competition. Together this suggests that additional NAC046 TFs compete for binding to RCD1.
Based on our results, the SLiM was expanded from [ED].{1,2}[^RK][YF].{1,4}[^RK][DE]L to [ED].{1,2}[^RK][YF].{1,4}[^RK][DE]([LIVMF]|.L). Introducing the Leu264→Ile substitution in AtDREB2A(244–272), mapping to an internal α-helix position in RCD1-bound DREB2A25, did not affect binding affinity significantly, although isoleucine is relatively destabilizing in α-helices. This may be explained by helix capping (Fig. 5A) and thermodynamics (Table 2). Enthalpy contributed less to the affinity of AtDREB2A(244–272;L264I) than of AtDREB2A(244–272), suggestive of sub-optimal binding geometry. However, this was compensated by entropy, possible due to less loss in entropy upon binding for the substituted peptide. In contrast, 25-fold decrease in affinity was measured for the corresponding substitution in ANAC013(254–274). The small contribution of entropy to binding of the wildtype peptide was changed into an unfavorable contribution upon substitution, indicating that the conformationally restricted isoleucine may be a disadvantage to a dynamic RCD1:ANAC013 complex30. Nonetheless, the SLiM variants are prospectively binding-competent putatively expanding the TF-interactome considerably (Fig. 1C).
Evolutionary conservation of the RCD1:TF interactome was indicated by interspecies cross-binding with Arabidopsis and soybean RCD1-RST and DREB2A being interchangeable interaction partners, and with HvDRF1.1 binding AtRCD1-RST (Table 2) suggestive of conserved interaction interfaces and properties. However, species-specific co-evolution of the RST:TF interfaces was also apparent from comparison of the AtDREB2A(244–272) and ANAC013(254–274) association with Arabidopsis versus soybean RCD1-RST. The affinities for the soybean RST domain were lower than for the Arabidopsis domain with 25-fold and 58-fold decreases for AtDREB2A(244–272) and ANAC013(254–272), respectively. In both cases, this was due to a decrease in binding enthalpy reflecting sub-optimal inter-species interfaces. Such pronounced species-specific co-adaptation was not observed for the GmDREB2A(273–290):GmRCD1-RST(483–583) interaction, which may, however, not represent the best molecular match among their different paralogs. Although our study revealed molecular co-adaptation as exemplified by the high affinity AtRCD1-RST:AtDREB2A complex, several parameters, such as SLiM context30,54, pH55 and partner dynamics55 may also influence affinity.
Intriguingly, helix folding-upon-binding was detected for AtDREB2A, but to a lesser extent for ANAC013 and not at all for ANAC04630. Accordingly, the α-helix propensities of the DREB2 SLiM regions were higher than those of the NAC013/NAC016/NAC017 and the NAC046 SLiM regions (Fig. 6A–C), further supportive of convergent evolution. Thus, molecular heterogeneity previously suggested for the RCD1:TF interactions seems to be evolutionary conserved. In contrast, binding to other αα-hub domains, such as the PAH domains of the transcriptional regulator Sin3, fully depend on coupled folding and binding with α-helix formation in the TFs25,56. AtRCD1-RST was able to bind the HvDRF1.1 SLiM, which has a higher intrinsic propensity for α-helix than the other DREB2A SLiM regions (Figs. 5B; 6A). The mechanism of DREB2A association with RCD1, e.g. conformational selection or induced fit, remains unknown. However, the results suggest that pre-formation of α-helix does not hinder complex formation and vice versa may also not be an advantage.
As for TF activation domains binding co-activators, negatively charged and hydrophobic residues are used by TF IDRs for RCD1 binding30,42,57. Likewise, the RCD1-RST domain shows characteristics typical of activation domain-interacting domains. Thus, it has a positively charged patch and a hydrophobic binding pocket58. The DREB2A, ANAC013, and ANAC046 genes showed similar expression patterns in response to various stress treatments28. The question is whether the corresponding proteins share RCD1 binding site despite their different bound conformations or form dynamic complexes (Fig. 6D) as reported for TF interactions with Mediator coactivator subunits59,60,61. RCD1-RST(499–572) mutant analysis (Table 2) indicate that ANAC087 targets the same interface on RCD1 as AtDREB2A25 despite its low α-helix propensity (Fig. 6C) and may explain why Arg551 and Ile563 are conserved without contributing to the RST core structure (Fig. 1B,E). Interestingly, HvRCD1 has a valine in the position corresponding to Arg560 of AtRCD1 (Fig. 1B), which is important for TF-binding, and HvRCD1 did not bind HvDRF1.1 (Table 2). Furthermore, both electrostatic and hydrophobic interactions are involved in RCD1 interactions with the DREB2A and NAC TFs25,30,46 (Table 2). Together these data lead to the current RCD1:TF binding model (Fig. 6D) which points out the structural plasticity of the RCD1:TF interactome. Additionally, this whole-interactome study opens new scientific avenues addressing co-evolution of structure and dynamics in interactomes, evolution-based interactome expansion, ID:hub interaction mechanisms based on whole-interactome strategies, and in vivo competition for TF binding to RCD1.
Methods
Bioinformatics
Inparalogs and orthologs were identified using Integrative Orthology Viewer at PLAZA, which integrates four complementary methods34,35. Dicots PLAZA 4.0, Monocots PLAZA 4.0, Gymno PLAZA 1.0, and Pico-PLAZA 2.0 were used to obtain data from a wide range of plant species. The protein sequences are named by the PLAZA identifier codes which are specific for the Dicot, Monocot and Gymno PLAZA platforms. In addition, BLAST homology searches62 were performed to obtain full-length proteins from Hordeum vulgare (barley). The WWE (PS50918) and PARP (PS51059) domains were identified by ScanProsite63 which scans the submitted sequences against the signatures from the Prosite database. The RST domain (PF12174) was identified by SMART64 in the ‘normal’ mode which allows SMART to search for domains by scanning the homologues sequences against the whole SMART database. In addition to the protein domains annotated in SMART, the search for Pfam domains was included. Multiple protein sequence alignments were created using Clustal Omega65 and evaluated using Transitive Consistency Score. For the multiple sequence alignments, minor manual adjustments were made in Jalview45. The TFs were analyzed for ID-conservation using IUPRED66, which allows disorder prediction for multiple sequences imported from Jalview. The median as well as the first and third quartiles were calculated for the IUPRED scores at each alignment position in order to show the interquartile range together with minimum and maximum values. Eukaryotic Linear Motif (ELM)67 was used to search for documented eukaryotic SLiMs, and α-helix propensity was analyzed using Agadir68. MoRFpred69 was used for prediction of MoRFs counting four or more residues. The non-redundant protein database was searched for additional transcription factors putatively binding RCD1 using two different BLAST programs, PHI-BLAST and PSI-BLAST62.
Modelling
The structures of HvRST, GmRST and PpRST were modelled using the I-TASSER70 webserver (https://zhanglab.ccmb.med.umich.edu/I-TASSER/) with standard settings and no constraints. Complexes between AtRST and either ANAC013 or ANAC046 were modelled using the CABS-dock server71 (http://biocomp.chem.uw.edu.pl/CABSdock/), with the structure of complexed AtRST (PDB 5OAO) and the peptide sequences of ANAC013(254–274) or ANAC046(319–338) as input.
Cloning, heterologous expression and protein purification
The cDNA encoding Glycine max (soybean) RCD1 (ID XP_003516978) residues 483–583 was obtained from ATUM and cloned into pET-15b. The RST domain was purified as previously described for RST domains42,49. ANAC087 (ID AT5G18270) and AtDREB2A (ID AT5G05410) cDNAs were obtained from the REGINA collection (Paz-Ares and the REGIA Consortium 2002). Gene-specific primers, encoding a tobacco etch virus cleavage site, were used for PCR-based amplification and the fragments cloned into pGEX-4T-1 (GE Healthcare) to produce GST-tagged proteins. The constructs were verified by sequencing, and site-specific mutations were introduced using the QuikChange mutagenesis kit (Stratagene). The GST-tagged fusion proteins were expressed in Escherichia coli strain BL21-(DE3) at 37 °C, induced by 0.5 mM isopropyl-D-thiogalactopyranoside, harvested after 3 h, resuspended in 20 mM Tris-HCl, pH 8.0, 100 mM NaCl, and sonicated. After centrifugation for 15 min at 36.000 × g, the supernatant was incubated at 4 °C for one h with glutathione-Sepharose 4B (Sigma) resin. Bound GST-tagged recombinant protein was cleaved from the GST tag using tobacco etch virus protease by incubation overnight in resuspension buffer containing 1 mM EDTA, 5 mM DTT and 1.0 µg of tobacco etch virus/0.1 mg protein. To remove the protease after cleavage, the disordered fragments were heated at 72 °C for 10 min and centrifuged at 20,000 × g for 10 min. Salt was removed from the peptides by freeze-drying before resuspension in 0.1% trifluoroacetic acid and purification on a Vydac C18 column (Grace) equilibrated in 20% ethanol, 0.1% trifluoroacetic acid and eluted in a linear gradient from 20 to 100% ethanol. Purified peptides were analyzed by MALDI-TOF (Autoflex Bruker) mass spectrometry and SDS-PAGE. N-terminally acetylated and C-terminally amidated peptides of HvDRF1.1(255–273) (ID AAO27885.1) and GmDREB2A(273–290) (ID XP_028217593.1/PLAZA ID GM02G42960) were obtained from TAG Copenhagen A/S. Plasmids encoding ANAC013 (ID AT1G32870) residues 254–274, HvNAC013 (ID AK376297.1) residues 176–346, and HvRCD1 (ID FR846236) residues 467–579 were from previous work, and the peptides and protein were expressed and purified as described30,49.
CD spectroscopy
Far-UV CD spectra of peptides were recorded in 10 mM Na2HPO4/Na2PO4, pH 7.0, at 15–20 µM and increasing amounts of TFE (0–40%; v/v) as indicated in the figure legends. Details of the recordings were as in42.
Isothermal titration calorimetry (ITC)
ITC was used to determine the thermodynamic parameters, the dissociation constant (Kd), the stoichiometry (N), and the binding enthalpy change (ΔH) from which the binding Gibbs free energy change (ΔG) and the binding entropy change (ΔS) were calculated. The experiments were performed in a MicroCal ITC200 microcalorimeter (GE Healthcare). Protein samples were dialyzed against 50 mM Hepes, pH 7.4, 100 mM NaCl, centrifuged at 20,000 × g for 10 min, and degassed for 10 min by stirring under vacuum. The protein concentration in the sample cell was 10–30 µM and the titrant concentrations in the syringe were 100–300 µM. A total of 18 injections separated by 180 s and with a duration of 4 s each of 2 µl titrant was injected into the sample at 25 °C. The Origin 7 software package (MicroCalTM) was used for fitting the data to a “one set of sites” binding model. Standard errors for ΔH, Kd, and N were obtained from Origin when fitting the data. All experiments were repeated at least three times. The heat of dilution was subtracted from the raw data by performing a titration of titrant against buffer or by subtracting the dilution enthalpy obtained when the enthalpy change had reached a constant level.
References
Ward, J. J., Sodhi, J. S., McGuffin, L. J., Buxton, B. F. & Jones, D. T. Prediction and functional analysis of native disorder in proteins from the three kingdoms of life. J. Mol. Biol. 337, 635–645 (2004).
Xue, B., Dunker, A. K. & Uversky, V. N. Orderly order in protein intrinsic disorder distribution: disorder in 3500 proteomes from viruses and the three domains of life. J. Biomol. Struct. Dyn. 30, 137–149 (2012).
Dunker, A. K. et al. Intrinsically disordered protein. J. Mol. Graph. Model. 19, 26–59 (2001).
Wright, P. E. & Dyson, H. J. Intrinsically unstructured proteins: re-assessing the protein structure-function paradigm. J. Mol. Biol. 293, 321–331 (1999).
Tompa, P. Intrinsically unstructured proteins. Trends Biochem. Sci. 27, 527–533 (2002).
Uversky, V. N. What does it mean to be natively unfolded? Eur. J. Biochem. 269, 2–12 (2002).
Borgia, A. et al. Extreme disorder in an ultrahigh-affinity protein complex. Nature 555, 61–66 (2018).
Uversky, V. N., Gillespie, J. R. & Fink, A. L. Why are “natively unfolded” proteins unstructured under physiologic conditions? Proteins 41, 415–427 (2000).
Wright, P. E. & Dyson, H. J. Intrinsically disordered proteins in cellular signalling and regulation. Nat. Rev. Mol. Cell Biol. 16, 18–29 (2015).
Staby, L. et al. Eukaryotic transcription factors: paradigms of protein intrinsic disorder. Biochem. J. 474, 2509–2532 (2017).
Iakoucheva, L. M., Brown, C. J., Lawson, J. D., Obradovic, Z. & Dunker, A. K. Intrinsic disorder in cell-signaling and cancer-associated proteins. J. Mol. Biol. 323, 573–584 (2002).
Dunker, A. K., Brown, C. J., Lawson, J. D., Iakoucheva, L. M. & Obradovic, Z. Intrinsic disorder and protein function. Biochemistry 41, 6573–6582 (2002).
Liu, J. et al. Intrinsic disorder in transcription factors. Biochemistry 45, 6873–6888 (2006).
Minezaki, Y., Homma, K., Kinjo, A. R. & Nishikawa, K. Human transcription factors contain a high fraction of intrinsically disordered regions essential for transcriptional regulation. J. Mol. Biol. 359, 1137–1149 (2006).
Light, S., Sagit, R., Sachenkova, O., Ekman, D. & Elofsson, A. Protein expansion is primarily due to indels in intrinsically disordered regions. Mol. Biol. Evol. 30, 2645–2653 (2013).
Brown, C. J., Johnson, A. K. & Daughdrill, G. W. Comparing models of evolution for ordered and disordered proteins. Mol. Biol. Evol. 27, 609–621 (2010).
Ahrens, J., Dos Santos, H. G. & Siltberg-Liberles, J. The Nuanced Interplay of Intrinsic Disorder and Other Structural Properties Driving Protein Evolution. Mol. Biol. Evol. 33, 2248–2256 (2016).
Glotzer, M., Murray, A. W. & Kirschner, M. W. Cyclin is degraded by the ubiquitin pathway. Nature 349, 132–138 (1991).
Davey, N. E., Cyert, M. S. & Moses, A. M. Short linear motifs - ex nihilo evolution of protein regulation. Cell Commun. Signal. 13, 43 (2015).
Holt, L. J. et al. Global analysis of Cdk1 substrate phosphorylation sites provides insights into evolution. Science 325, 1682–1686 (2009).
Oldfield, C. J. et al. Coupled folding and binding with alpha-helix-forming molecular recognition elements. Biochemistry 44, 12454–12470 (2005).
Xue, B., Brown, C. J., Dunker, A. K. & Uversky, V. N. Intrinsically disordered regions of p53 family are highly diversified in evolution. Biochim. Biophys. Acta 1834, 725–738 (2013).
Kragelund, B. B., Jensen, M. K. & Skriver, K. Order by disorder in plant signaling. Trends Plant Sci. 17, 625–632 (2012).
Jaspers, P., Brosche, M., Overmyer, K. & Kangasjarvi, J. The transcription factor interacting protein RCD1 contains a novel conserved domain. Plant Signal. Behav. 5, 78–80 (2010).
Bugge, K. et al. Structure of Radical-Induced Cell Death1 Hub Domain Reveals a Common alphaalpha-Scaffold for Disorder in Transcriptional Networks. Structure. 26, 734–746 (2018).
Jaspers, P. et al. Unequally redundant RCD1 and SRO1 mediate stress and developmental responses and interact with transcription factors. Plant J. 60, 268–279 (2009).
Teotia, S. & Lamb, R. S. The paralogous genes RADICAL-INDUCED CELL DEATH1 and SIMILAR TO RCD ONE1 have partially redundant functions during Arabidopsis development. Plant Physiol 151, 180–198 (2009).
Brosche, M. et al. Transcriptomics and functional genomics of ROS-induced cell death regulation by RADICAL-INDUCED CELL DEATH1. PLoS. Genet. 10, e1004112 (2014).
Wirthmueller, L. et al. Arabidopsis downy mildew effector HaRxL106 suppresses plant immunity by binding to RADICAL-INDUCED CELL DEATH1. New Phytol. 220, 232–248 (2018).
O’Shea, C. et al. Structures and Short Linear Motif of Disordered Transcription Factor Regions Provide Clues to the Interactome of the Cellular Hub Protein Radical-induced Cell Death1. J. Biol. Chem. 292, 512–527 (2017).
Sakuma, Y. et al. Dual function of an Arabidopsis transcription factor DREB2A in water-stress-responsive and heat-stress-responsive gene expression. Proc. Natl. Acad. Sci. USA 103, 18822–18827 (2006).
Vainonen, J. P. et al. RCD1-DREB2A interaction in leaf senescence and stress responses in Arabidopsis thaliana. Biochem. J. 442, 573–581 (2012).
Shapiguzov,A. et al. Arabidopsis RCD1 coordinates chloroplast and mitochondrial functions through interaction with ANAC transcription factors. Elife. 8 (2019).
Van Bel, M. et al. Dissecting plant genomes with the PLAZA comparative genomics platform. Plant Physiol 158, 590–600 (2012).
Proost, S. et al. PLAZA 3.0: an access point for plant comparative genomics. Nucleic Acids Res. 43, D974–D981 (2015).
You, J. et al. The SNAC1-targeted gene OsSRO1c modulates stomatal closure and oxidative stress tolerance by regulating hydrogen peroxide in rice. J. Exp. Bot. 64, 569–583 (2013).
Jaspers, P. et al. The RST and PARP-like domain containing SRO protein family: analysis of protein structure, function and conservation in land plants. BMC. Genomics 11, 170 (2010).
Tossavainen, H., Hellman, M., Vainonen, J. P., Kangasjarvi, J. & Permi, P. (1)H, (13)C and (15)N NMR chemical shift assignments of A. thaliana RCD1 RST. Biomol. NMR Assign. 11, 207–210 (2017).
Engel, D. E. & DeGrado, W. F. Alpha-alpha linking motifs and interhelical orientations. Proteins 61, 325–337 (2005).
Jensen, M. K. et al. The Arabidopsis thaliana NAC transcription factor family: structure-function relationships and determinants of ANAC019 stress signalling. Biochem. J. 426, 183–196 (2010).
You, J., Zong, W., Du, H., Hu, H. & Xiong, L. A special member of the rice SRO family, OsSRO1c, mediates responses to multiple abiotic stresses through interaction with various transcription factors. Plant Mol. Biol. 84, 693–705 (2014).
O’Shea, C. et al. Protein intrinsic disorder in Arabidopsis NAC transcription factors: transcriptional activation by ANAC013 and ANAC046 and their interactions with RCD1. Biochem. J. 465, 281–294 (2015).
Mahani, A., Henriksson, J. & Wright, A. P. Origins of Myc proteins–using intrinsic protein disorder to trace distant relatives. PLoS. One. 8, e75057 (2013).
Stender, E. G., O’Shea, C. & Skriver, K. Subgroup-specific intrinsic disorder profiles of Arabidopsis NAC transcription factors: Identification of functional hotspots. Plant Signal. Behav. 10, e1010967 (2015).
Waterhouse, A. M., Procter, J. B., Martin, D. M., Clamp, M. & Barton, G. J. Jalview Version 2–a multiple sequence alignment editor and analysis workbench. Bioinformatics. 25, 1189–1191 (2009).
Munoz, V., Blanco, F. J. & Serrano, L. The hydrophobic-staple motif and a role for loop-residues in alpha-helix stability and protein folding. Nat. Struct. Biol. 2, 380–385 (1995).
Huysmans, M. et al. NAC Transcription Factors ANAC087 and ANAC046 Control Distinct Aspects of Programmed Cell Death in the Arabidopsis Columella and Lateral Root Cap. Plant Cell 30, 2197–2213 (2018).
Xue, G. P. & Loveridge, C. W. HvDRF1 is involved in abscisic acid-mediated gene regulation in barley and produces two forms of AP2 transcriptional activators, interacting preferably with a CT-rich element. Plant J. 37, 326–339 (2004).
Kjaersgaard, T. et al. Senescence-associated barley NAC (NAM, ATAF1,2, CUC) transcription factor interacts with radical-induced cell death 1 through a disordered regulatory domain. J. Biol. Chem. 286, 35418–35429 (2011).
Khraiwesh, B. et al. Genome-wide expression analysis offers new insights into the origin and evolution of Physcomitrella patens stress response. Sci. Rep. 5, 17434 (2015).
Krystkowiak, I. & Davey, N. E. SLiMSearch: a framework for proteome-wide discovery and annotation of functional modules in intrinsically disordered regions. Nucleic Acids Res. 45, W464–W469 (2017).
Van Roey, K. et al. Short linear motifs: ubiquitous and functionally diverse protein interaction modules directing cell regulation. Chem. Rev. 114, 6733–6778 (2014).
Holm, M., Hardtke, C. S., Gaudet, R. & Deng, X. W. Identification of a structural motif that confers specific interaction with the WD40 repeat domain of Arabidopsis COP1. EMBO J. 20, 118–127 (2001).
Berlow, R. B., Dyson, H. J. & Wright, P. E. Hypersensitive termination of the hypoxic response by a disordered protein switch. Nature 543, 447–451 (2017).
Jemth, P. et al. Structure and dynamics conspire in the evolution of affinity between intrinsically disordered proteins. Sci. Adv. 4, eaau4130 (2018).
Sahu, S. C. et al. Conserved themes in target recognition by the PAH1 and PAH2 domains of the Sin3 transcriptional corepressor. J. Mol. Biol. 375, 1444–1456 (2008).
Erkine, A. M. ‘Nonlinear’ Biochemistry of Nucleosome Detergents. Trends Biochem. Sci. 43, 951–959 (2018).
Ravarani, C. N. et al. High-throughput discovery of functional disordered regions: investigation of transactivation domains. Mol. Syst. Biol. 14, e8190 (2018).
Brzovic, P. S. et al. The acidic transcription activator Gcn4 binds the mediator subunit Gal11/Med15 using a simple protein interface forming a fuzzy complex. Mol. Cell 44, 942–953 (2011).
Tuttle, L. M. et al. Gcn4-Mediator Specificity Is Mediated by a Large and Dynamic Fuzzy Protein-Protein Complex. Cell Rep. 22, 3251–3264 (2018).
Currie, S. L. et al. ETV4 and AP1 Transcription Factors Form Multivalent Interactions with three Sites on the MED25 Activator-Interacting Domain. J. Mol. Biol. 429, 2975–2995 (2017).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
de Castro, E. et al. ScanProsite: detection of PROSITE signature matches and ProRule-associated functional and structural residues in proteins. Nucleic Acids Res. 34, W362–W365 (2006).
Schultz, J., Milpetz, F., Bork, P. & Ponting, C. P. SMART, a simple modular architecture research tool: identification of signaling domains. Proc. Natl. Acad. Sci. USA 95, 5857–5864 (1998).
Sievers, F. et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 7, 539 (2011).
Dosztanyi, Z., Csizmok, V., Tompa, P. & Simon, I. The pairwise energy content estimated from amino acid composition discriminates between folded and intrinsically unstructured proteins. J. Mol. Biol. 347, 827–839 (2005).
Dinkel, H. et al. The eukaryotic linear motif resource ELM: 10 years and counting. Nucleic Acids Res. 42, D259–D266 (2014).
Munoz, V., Cronet, P., Lopez-Hernandez, E. & Serrano, L. Analysis of the effect of local interactions on protein stability. Fold. Des 1, 167–178 (1996).
Disfani, F. M. et al. MoRFpred, a computational tool for sequence-based prediction and characterization of short disorder-to-order transitioning binding regions in proteins. Bioinformatics. 28, i75–i83 (2012).
Yang, J. et al. The I-TASSER Suite: protein structure and function prediction. Nat. Methods 12, 7–8 (2015).
Kurcinski, M., Jamroz, M., Blaszczyk, M., Kolinski, A. & Kmiecik, S. CABS-dock web server for the flexible docking of peptides to proteins without prior knowledge of the binding site. Nucleic Acids Res. 43, W419–W424 (2015).
Vandepoele, K. et al. pico-PLAZA, a genome database of microbial photosynthetic eukaryotes. Environ. Microbiol. 15, 2147–2153 (2013).
Acknowledgements
This study was supported by Danish Research Councils Grant 4181–00344 (to B. B. K. and K. S.) and the Novo Nordisk Foundation Challenge grant NNF18OC0033926 (to B.B.K. and K.S.).
Author information
Authors and Affiliations
Contributions
Conceptualization: L.F.C. and K.S.; Experiments: L.F.C., L.S., K.B., C.O.S.; Writing: L.F.C., L.S., K.B., B.B.K. and K.S.; Funding acquisition: B.B.K. and K.S.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Christensen, L.F., Staby, L., Bugge, K. et al. Evolutionary conservation of the intrinsic disorder-based Radical-Induced Cell Death1 hub interactome. Sci Rep 9, 18927 (2019). https://doi.org/10.1038/s41598-019-55385-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-55385-3
This article is cited by
-
Molecular switching in transcription through splicing and proline-isomerization regulates stress responses in plants
Nature Communications (2024)
-
Evolutionary fine-tuning of residual helix structure in disordered proteins manifests in complex structure and lifetime
Communications Biology (2023)
-
A dialogue between generations
Nature Plants (2022)
-
Spatially expressed WIP genes control Arabidopsis embryonic root development
Nature Plants (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.