Abstract
Neuroblastoma is the most common extracranial solid tumor found in children and survival rate is extremely meager. HDAC8, a class I zinc-dependent enzyme, is a potential drug target for treatment of neuroblastoma and T cell lymphoma. Most of the HDAC8 inhibitors discovered till date contains a hydroxamic acid group which acts as a zinc binding group. The high binding affinity to the zinc and other ions results in adverse effects. Also, the non-selective inhibition of HDACs cause a variety of side effects. The objective of this is to identify structurally diverse, non-hydroxamate, novel, potential and selective HDAC8 inhibitors. A number of five featured pharmacophore hypotheses were generated using 32 known selective HDAC8 inhibitors. The hypotheses ADDRR.4 were selected for building 3D QSAR model. This model has an excellent correlation coefficient and good predictive ability, which was employed for virtual screening of Phase database containing 4.3 × 106 molecules. The resultant hits with fitness score >1.0 were optimized using in-silico ADMET (absorption, distribution, metabolism, excretion, and toxicity) and XP glide docking studies. On the basis of pharmacophore matching, interacting amino acid residues, XP glide score, more affinity towards HDAC8 and less affinity towards other HDACs, and ADME results five hits- SD-01, SD-02, SD-03, SD-04 and SD-05 with new structural scaffolds, non-hydroxamate were selected for in vitro activity study. SD-01 and SD-02 were found to be active in the nanomolar (nM) range. SD-01 had considerably good selectivity for HDAC8 over HDAC6 and SD-02 had marginal selectivity for HDAC6 over HDAC8. The compounds SD-01 and SD-02 were found to inhibit HDAC8 at concentrations (IC50) 9.0 nM and 2.7 nM, respectively.
Similar content being viewed by others
Introduction
HDACs are one of the most important classes of post-translational regulators that are responsible for deacetylation of lysine residues in histone and non-histone substrates. To date, 18 types of HDACs have been identified and classified into four broad classes: class I, II (further classified as IIa and IIb), III, and IV (Fig. S2). HDAC8, a class I zinc-dependent HDACs, which localizes to either the nucleus or the cytoplasm, typically induces histone deacetylation and represses gene transcription1. HDACs are identified as potential therapeutic targets due to their involvement in various diseases like cancer, inflammation, neurological disorders and infections2,3. In cancer, HDACs are either deregulated, over expressed, or interact with transcription factors4. HDAC8 may be the potential drug target for the treatment of minimal residual disease in neuroblastoma and malignancies such as T-cell lymphoma and acute myeloid leukemia5,6. FDA has approved suberoylanilide hydroxamic acid (vorinostat) for treatment of cutaneous T-cell lymphoma (CTCL) after multiple clinical trials in 20067,8 and romidepsin (cyclic peptide) in 20099,10. Many other HDAC inhibitors like belinostat11, panobinostat12, pracinostat13 have been approved by US-FDA and chidamide have been approved by China-FDA14 for treatment of cancer and several are in clinical trials15.
A number of structurally diverse HDAC8 inhibitors discovered so far are known to be hydroxamic acids16,17,18,19. Hydroxamic acid group binds with zinc, which often creates metabolic and pharmacokinetic troubles. Also, many hydroxamates are unstable in vivo, and on hydrolysis give mutagenic hydroxylamine20. Hydroxamic acid also showed a strong chelating ability with zinc and hence it lack selectivity21. Inhibition of several HDACs simultaneously confers greater toxicity and long term side effects. Therefore discovery of isoform-selective HDACs, improve therapeutic potential22. The highly conserved active site of HDACs family members makes it difficult to design isoform-selective inhibitors19. Thus, the discovery of potential, novel scaffolds and selective HDAC8 inhibitors besides currently existing hydroxamic acid is a necessity.
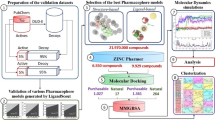
The combined ligand-based and structure-based approach is very important in modern drug discovery for searching potential lead molecules23,24,25,26,27. In the last few decades, the virtual screening tool was employed to identify novel lead molecules with diverse structural features28,29,30. The number of pharmacophore-based virtual screenings for identification of HDAC8 inhibitors against a commercial and in-house database of compounds has been reported to be small31,32 and the number of 3D QSAR model use for selective HDAC8 inhibitors has been reported to be very limited. This study aimed to find out novel non-hydroxamic acid, selective HDAC8 inhibitors using cost-effective and rapid in silico approach, a combination of pharmacophore-based virtual screening, molecular docking, ADMET (absorption, distribution, metabolism, excretion, and toxicity) properties and evaluation of in vitro HDAC8 and HDAC6 inhibitory activity of identified hits.
Results and Discussions
The number of five-feature pharmacophore hypotheses were eleven, generated with a combination of three chemical features i.e. hydrogen-bond acceptor (A), hydrogen bond donor (D) and aromatic ring (R). The pharmacophore hypotheses ADDRR.4, AADRR.4, AADDR.12, AADDR.11, AADDR.15, AADDR.16, AADDR.14, AAADR.20, AAADR.19, AAADR.24, AAADR.23 were generated from 32 known selective HDAC8 inhibitors (Fig. 1). The 32 inhibitors were classified into eight series on the basis of their structural symmetry are series A–E, five and series F, which contains three diverse structures are shown in Fig. S1. Out of these pharmacophore hypotheses, three top-scoring hypotheses- ADDRR.4 (survival score: 5.139, survival-inactive: 3.298) AADRR.4 (survival score: 4.939, survival-inactive: 3.177), and AADDR.12 (survival score: 4.310, survival-inactive: 2.588) were selected for building 3D QSAR models. Out of three derived 3D QSAR models, ADDRR.4 was found to be statistically significant and the statistical parameters of the 3D QSAR model were R2 (squared correlation coefficient): 0.9995, with a root-mean-square error (RMSE) of 0.3023, the good F value of 8732.3. Its predictive correlation coefficient (Q2) which is 0.6626. (Table 1). The 3D QSAR model consisted of a spatial arrangement of five chemical features (Fig. 2). The predictive correlation coefficient confirmed the robustness and predictive ability of the model. The QSAR results showed that the observed activity of training and test set molecules were very close to phase predicted activity (Fig. 3 and Table S2). The predictive ability of the model has further been validated using known selective HDAC8 inhibitors 1’−22’ retrieved from the literature (Tables S1). The comparison of phase predicted pIC50 values of 22 external selective HDAC8 inhibitors from their corresponding experimental pIC50 (Fig. 4), further supports that the predictive ability of the model was excellent. The best predictive 3D QSAR model built from AADRR.4 was used for virtual screening of Phase database. Pharmacophore-based virtual screening resulted in 500 hits fitness score >1.0 from Phase database.
PHASE generated best pharmacophore model ADDRR.4 of selective HDAC8 inhibitors illustrating hydrogen bond acceptor (A2, pink), hydrogen bond donor (D3, D4; sky) and aromatic ring (R7, R8; orange) with their angles (a) are shown by green lines and distance (b) are shown purple lines. The twenty three training set inhibitors were used for this model generation.
QickProp predicts physically significant and the pharmaceutically relevant properties of organic molecules by comparing 95% of known drugs. According to Lipinski’s rule of five for drug-like molecules the molecular weight should be <500, octanol-water partition coefficient (QPlogPo/w) should <5.0, hydrogen bond donor groups (donorHB) should be <5.0, and hydrogen bond acceptor groups should be ≤10. The predicted descriptors for all the five selected hits obeyed the Lipiniski’s rule of five. The QPlogPo/w and water solubility (QPlog S) which are essential in the evaluation of adsorption and distribution of drugs and the range of these parameters for five selected hits were 1.05–2.997 and −2.797–3.66, respectively, which were also in the acceptable range. The other important parameters are apparent Caco-2 cell permeability in nm/sec (QPPCaco) and apparent MDCK cell permeability in nm/sec (QPPMDCK) and the values were 47.626–973.33 and 43.612–2256.642, respectively. For all the five selected hits the values of QPPCaco and QPPMDCK are in the acceptable range but for potential HDAC8 inhibitor, these values for SD-01 were 973.33 and 2256.642, respectively which is great. The important ADME properties of finally identified five hits lie with acceptable range are listed in Table 2 and the remaining properties listed in Table S3.
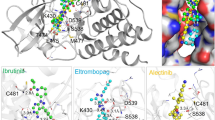
RMSD values of different HDAC isoforms selected for molecular docking were within the acceptable range (Table 3). Superposition of docked co-ligands on its originally bound native conformation of co-ligand indicated that the quality of reproduction of a co-ligands (i.e. crystallographic) binding pose by a computational method was very good (Fig. S3). Therefore these HDAC low RMSD isoforms were used for molecular docking studies. The molecular docking studies of known selective HDAC8 inhibitors 1–32, and hits resulted after ADME filtration was performed to measure the XP glide score and also to find out interacting active site amino acid residues of HDAC8. The 2D ligand interaction diagram of 32 docked inhibitors found in the Supplementary Table S4 and analysis of 2D ligand interaction diagram are shown in Table S5. The docking analysis of 32 known selective HDAC8 inhibitors showed that the most interacting active site functionalities were divalent Zn+2 ion, GLY-151, PHE-152, TYR-306, PHE-208, HIS-142 including some other less interacting residues (Fig. 5). The range of XP glide scores of 32 known inhibitors were –8.1 to –11.4. The ADME filtered hits on XP glide docking with HDAC8 and best docked 20 hits with XP glide score of >9.0 were further docked with other HDACs (1, 2, 3, 4, 6). Based on the high binding affinity towards HDAC8 and less binding affinity towards other HDACs, five hits were identified as selective HDAC8 inhibitors for in vitro activity. The XP glide score of five selected inhibitors for different HDACs are shown in Table 4 and the 2D ligand interaction diagram of selected hits SD-01, SD-02, SD-03, SD-04, and SD-05 with different HDACs are shown in Fig. S4a– S4f. The XP glide score >9.0 of top five selected hits (Figs. 6–10) also indicated that the hits are selective towards HDAC8. The XP glide score of top five hits with other HDACs is poor compare to HDAC8 and are listed in Table 4. The most interacting active site amino acid residues of five selective HDAC8 inhibitors were Zn+2, GLY-151, PHE-152, HIE-180, TYR-306 including some other less interacting amino acid residues (Fig. 5). These interactions were very similar to 32 known inhibitors.
Docking poses of compound SD-01 ((a) 2D ligand interaction diagram like hydrogen bond donor, hydrogen bond acceptor, π-π stacking, (b) 3Dligandinteractions likehydrogen bond donor,hydrogen bond acceptor, π-π stacking, (c) 3D hydrophobic interactions). The 2D interactions are depicted with different colors: pi-pi (green line), hydrogen bond (violet line) for 3D interaction hydrogen bond (purple line), pi-pi (doted sky line).
Docking poses of compound SD-02 ((a) 2D ligand interaction diagram like hydrogen bond donor, hydrogen bond acceptor, π-π stacking, (b) 3Dligandinteractions likehydrogen bond donor, hydrogen bond acceptor, π-π stacking, (c) 3D hydrophobic interactions). The 2D interactions are depicted with different colors: pi-pi (green line), hydrogen bond (violet line) for 3D interaction hydrogen bond (purple line), pi-pi (doted sky line).
Docking poses of compound SD-03 ((a) 2D ligand interaction diagram like hydrogen bond donor, hydrogen bond acceptor, π-π stacking, (b) 3Dligandinteractions like hydrogen bond donor,hydrogen bond acceptor, π-π stacking, (c) 3D hydrophobic interactions). The 2D interactions are depicted with different colors: pi-pi (green line), hydrogen bond (violet line) for 3D interaction hydrogen bond (purple line), pi-pi (doted sky line).
Docking poses of compound SD-04 ((a) 2D ligand interaction diagram like hydrogen bond donor, hydrogen bond acceptor, π-π stacking, (b) 3Dligandinteractions like hydrogen bond donor, hydrogen bond acceptor, π-π stacking, (c) 3D hydrophobic interactions). The 2D interactions are depicted with different colors: pi-pi (green line), hydrogen bond (violet line) for 3D interaction hydrogen bond (purple line), pi-pi (doted sky line).
Docking poses of compound SD-05 ((a) 2D ligand interaction diagram like hydrogen bond donor, hydrogen bond acceptor, π-π stacking, (b) 3Dligandinteractions like hydrogen bond donor, hydrogen bond acceptor, π-π stacking, (c) 3D hydrophobic interactions). The 2D interactions are depicted with different colors: pi-pi (green line), hydrogen bond (violet line) for 3D interaction hydrogen bond (purple line), pi-pi (doted sky line).
The fitness score of SD-01, SD-02, SD-03, SD-04, and SD-05 were 1.113, 1.529, 1.203, 1.294, and 1.84,1 respectively (Fig. 11). The fitness score and pharmacophore matched structures of hits indicated that all the five hits matched five pharmacophores (ADDRR.4). The pharmacophores of SD-01 were A2 which matched with amide oxygen and it binds with Zn+2 (2.01 Å) and TYR-306 (2.26 Å) in the active site. The other two pharmacophores D3 and D4 matched with two hydrogens of the amide group and one of them binds with GLY-151 (1.75 Å). The other two pharmacophores R7 and R8 matched with thiazole ring and 3-methoxy phenyl ring respectively of inhibitor. The pharmacophore R8 had π-π interactions with the aromatic ring of PHE-152. Other than these common pharmacophore interactions, two π-π interactions of 2-aminothiophenol ring of the inhibitor with the aromatic ring of PHE-208 and PHE-152 were observed (Fig. 6. SD-01a,01b). The hits enclosed it with hydrophobic residues PHE-208, PHE-152, LEU-308, TRP-141, TYR-306, PRO-35 and interactions with almost three rings of hits is shown in Fig. 6. SD-01c.
The pharmacophore A2 of SD-02 matched with acetoxy hydrazide carbonyl oxygen, which binds with Zn+2 (2.05 Å) of the active site. The other two pharmacophores were D3 and D4 matched with hydrogens of hydrazine -NH2 group and one of them binds with HIS-142 (2.27 Å) and -NH- hydrogen binds with GLY-151 (2.30 Å). The pharmacophores R7 and R8 were matched with the two rings of benzofuran. The pharmacophore R8 had two π-π interactions with HIE-180 and PHE-208 aromatic rings. The inhibitor also enclosed with hydrophobic PHE-208, PHE-252, TYR-306, TYR-100. All the interactions of SD-02 are shown in Fig. 7. SD-2a, SD-2b, and SD-2c.
In SD-03, the common pharmacophore D3 and D4 matched with amide hydrogens and A2 matched with amide oxygen. The pharmacophore A2 binds with Zn+2 (2.13 Å) and D3, or D4 binds with GLY-151. The flavone ring matched with R7, R8, and R8 had π-π interactions with HIE-180 aromatic ring. Other than these pharmacophores, there were a hydrogen bonding interactions of benzoate ester oxygen of hits with LYS-202 and benzoate ring had π-π interactions with PHE-207. The inhibitor enclosed with hydrophobic amino acid residues PHE-152, PHE-207, PHE-208, and TYR-306 and had interactions with the R8. All the interactions of SD-03 are shown in Fig. 8. SD-03a, SD-03b, and SD-03c.
In SD-04 the hydrogen bond acceptor pharmacophore A2 matched with amide oxygen and hydrogen bond donor pharmacophore D3 and D4 matched with amide hydrogens. Pharmacophore A2 of the inhibitor binds with Zn+2 (2.20 Å) and TYR-306 (2.01 Å). The pharmacophore D3 or D4 binds with GLY-151 (1.98 Å). Pharmacophores R7 and R8 matched with furan ring and thiazole ring respectively, and R7 had a π-π interaction with the HIE-180 ring. Other than these pharmacophore interactions there exists one hydrogen bond acceptor interaction of furan oxygen with HIE-180. All the interactions of SD-04 are shown in Fig. 9. SD-04a, SD-04b, and SD-04c.
In SD-05, pharmacophore A2 matched with an oxygen atom of urea part of inhibitor and binds with TYR-306 (2.02 Å) and Zn+2 (2.03 Å). The two hydrogen bond donor pharmacophores D3 and D4 matched with hydrogens of -NH- group of substituted urea and both binds with GLY-151. The two pharmacophore R7 and R8 matched with two aromatic rings of the naphthyl group. The ring R7 had π-π interaction with PHE-152 and R8 had two π-π interactions with PHE-152 and PHE-208. The hits had a hydrophobic enclosure with TYR-100, PHE-208, PHE-152, and TYR-306. The interactions of SD-05 are shown in Fig. 10. SD-05a, SD-05b, and SD-05c. The 3D interactions of five inhibitors showed that the common pharmacophores A2, D3, D4, R7 and R8 had good interactions with active site amino acid residues in most cases.
The generated E-pharmacophore of best selective known HDAC8 inhibitor PCI-34051 was used for matching of five selected hits by selecting all four features matching option. In SD-01, A3 superimposed on amide oxygen, A1 matched with 3-methoxy phenyl oxygen group, R8 and R9 matched with thiazole ring and 3-methoxyphenyl ring, respectively. All the four pharmacophores of selected PCI-34051 matched with SD-01 (Fig. 12). In SD-03, pharmacophore A3 superimposed on amide oxygen and A1 superimposed on benzoate oxygen (Fig. 12). The pyran-4-one of flavone matched with R8 and benzoate ester ring matched with R9. The SD-03 also matched all the four E-pharmacophores of PCI-34051. The other three compounds did not matched all the four E-pharmacophore of PCI-34051. Therefore the matching options were reduced to three features for other hits, which resulted in SD-02, SD-04 and SD-05 matching with three pharmacophores. The pharmacophore A1 of SD-02 matched with acetoxy hydrazide carbonyl oxygen and R8 superimposed on furan ring of benzofuran. The pharmacophore A3 matched with the carbonyl oxygen of ester group and R9 had no matching (Fig. 12). The pharmacophore A1 of SD-04 and R8 of SD-05 did not match but other pharmacophores were well matched (Fig. 12).
These results showed that SD-01 and SD-03 matched all the pharmacophoric features of the E-pharmacophores and therefore may be more selective than the other three. Based pharmacophore matching, XP glide score, interacting amino acid residues, and ADME properties, virtual screening hits SD-01, SD-02, SD-03, SD-04, and SD-05 were selected for in vitro activity study against HDAC6 and HDAC8.
The structures of all the five identified inhibitors are new and non-hydroxamic acid as confirmed by PubChem structure search, as the activity of these compounds has not been predicted earlier for HDAC8 inhibitory activity.
In vitro HDAC inhibition activity
The in vitro HDAC inhibitory activity results clearly showed that compound SD-01, SD-02, and SD-05 have potential HDAC8 inhibitory activities (Table 5). SD-01 is more selective towards HDAC8 over HDAC6 and SD-02 has marginal selectivity towards HDAC6 over HDAC8. The results demonstrated thatthe experimental results were similar to in silico results. The purity of the best two compounds, SD-01 and SD-02 was 94.543 and 97.630%, respectively.
The fitness score of all the five selected hits were >1.0 and almost all the five pharmacophoric features A2, D3, D4, R7 and R8 matches the selected hits are shown in Fig. 11. Among the selected hits XP glide score of SD-01 was highest –10.2 for HDAC8 and its experimental IC50 value was 9.0 nm. The XP glide score of SD-01 was –4.5 and –8.0 for two pockets zinc finger domain and catalytic domain 2 respectively, of HDAC6 and its IC50 value was >100 nm. The XP glide score of SD-02 was –9.3 for HDAC8 and –5.6 and –9.2 respectively, for zinc finger domain (5WPB) and catalytic domain 2 (5WGI) of two HDAC6 pockets. The experimental activity of SD-02 was 2.7 nM and 0.62 nM respectively, for HDAC8 and HDAC6. The glide score for SD-05 was –8.0 and –9.3 respectively, for HDAC6 catalytic domain 2 and HDAC8. The experimental activities were 30.86 nM and 41.6 nM respectively, for HDAC6 and HDAC8. The hit SD-02 has more affinity towards the catalytic domain 2 rather than zinc finger domain. There was a good symmetry between XP glide scores and experimental activities of the best three hits SD-01, SD-02 and SD-05. The E-pharmacophores A1, A3 (hydrogen bond acceptors), R8 and R9 (ring aromatic) generated form HDAC8 and selective HDAC8 inhibitor (PCI-34051) complex, matched nicely with SD-01 (Fig. 12). Therefore, SD-01 is considerably more selective towards HDAC8 compared to HDAC6. The superimposed docked poses of all selected hits on crystallographic bound co-ligand TSN of HDAC8 (PDB ID: 1T64) in the active site showed that except SD-04, the other hits bind in a similar binding pose with TSN (Fig. S5) and experimental activity of SD-04 was low.
Experimental
The selected virtual screening hits were obtained from Enamine and their structures were further confirmed by mass spectra (Fig. S8a–S8e) and purity of the compound was confirmed by analytical HPLC (Fig. S7a–S7e). The mass spectra of SD-01: HRMS (ES+), exact calc. mass for C19H20N3O2S2 [M + H]+, 386.0997. Found m/z 386.0988. The mass spectra of SD-02: HRMS (ES+), exact calc. mass for C15H19N2O5 [M + H]+, 307.1294. Found m/z 307.1288. The mass spectra of SD-03: HRMS (ES+), exact calc. mass for C19H16NO7 [M + H]+, 370.0927. Found m/z 370.0919. The mass spectra of SD-04: HRMS (ES+), exact calc. mass for C13H10N2O2SCl [M + H]+, 293.0152. Found m/z 293.0146. The mass spectra of SD-05: HRMS (ES+), exact calc. mass for C14H16N2O3Na [M + Na]+, 283.1059. Found m/z 283.1051.
The purity of compounds SD-01, SD-02, SD-03, SD-04, and SD-05 were 94.543, 97.630, 93.014, 83.471 and 80.034%, respectively. The HPLC chromatogram is available in supporting information Fig. S7a– S7e.
HDAC6 and HDAC8 activity assay
HDAC 6 and 8 activities were measured according to the manufacturer’s protocol from Sigma (Cat# CS1010). Briefly, to 5 ng of HDAC6 or 8 enzymes, compounds (1, 10, 50 and 100 nM) were added and incubated in the presence of substrate (Boc-Lys(Ac)-AMC) for 30 min at 30 °C. One μL developer was added to this mix and further incubated for 15 min and the fluorescent group liberated from the cleaved substrate, which is proportional to the deacetylase activity was measured using a fluorescence plate reader at Ex-350 nm; Em-440 nm. Trichostatin A was used as a reference inhibitor. The Perkin Elmer multimode reader, model-Enspire 2300 was used for fluorescence plate reader.
On the basis of in silico ligand, structure-and ADME filtration tools, a series of five HDAC8-selective non-hydroxamate hits SD-01, SD-02, SD-03, SD-04, and SD-05 were identified from the commercial database. In vitro HDAC inhibitory studies of identified leads demonstrated that SD-01 has potential inhibitory activity and good selectivity for HDAC8 over HDAC6. SD-02 has potential activity but marginal selectivity for HDAC6. The results of the present studies encouraged us for further studies on SD-01 to develop a potential candidate for isoform selective HDAC8 drug for the treatment of HDAC8-overexpressed cancers. The interesting facts are all the identified hits are structurally diverse and are from non-hydroxamic acid series. The common pharmacophore information: one hydrogen bond acceptor, two hydrogen bond donor and two ring aromatic and their interactions with active site amino acid residues and molecular docking results of 32 known selective HDAC8 inhibitors which were used in the present study may be useful for further design of isoform-selective HDAC8 inhibitors. The ADME properties revealed that the identified hits can be further developed as good oral drug candidates. These in-vitro identified leads may be used for designing anti-cancer chemo-therapeutics related to over expression of HDAC8 and HDAC6. Further in vitro study on other HDACs will be carried out in a future. This method may be useful for identification of isoform-selective HDACs inhibitors and the E-pharmacophore matched hits may be more selective towards HDACs isoform.
Materials and Methods
Dataset
Thirty two selective HDAC8 inhibitors have been chosen from the existing literature4,33,34,35 with inhibitory activity data (IC50 in μM) (Fig. 1). The IC50 values of the compounds were converted to their pIC50 values consisting of some high active, medium active and low active molecules. The distribution of activity data of known inhibitors and the number of compounds is shown in Fig. 13 which confirms the data span over 4 order magnitudes (4.469–8.000). The Phase database, a database of commercially available compounds contains 4.3 × 106 compounds (only first conformer) with a unique identifier, CACPD2011aCode was chosen for virtual screening36. The crystal structure of HDAC1 (PDB ID: 4BKX: resolution: 3.0 Å)37, HDAC2 (3MAX, 2.05 Å)38, HDAC3 (4A69, 2.06 Å)39, HDAC4 (2VQM, 1.8 Å)40, HDAC6 (5WPB, 1.55 Å)41, HDAC8 (1T64, 1.90 Å)42 was retrieved from RCSB Protein Data Bank (http://www.rcsb.org/pdb/) which was used for molecular docking study. All the computational calculations were performed on HP Z820 Workstation with CentOS 6.3.
Identified hits
The hits SD-01: C19H19N3O2S2, [2-((2-(((2-(3-methoxyphenyl)thiazol-4-yl)methyl)amino)phenyl)thio)acetamide, Phase Databank ID: CACPD2011a-0001983707, Enamine ID: T6550042], SD-02: C15H18N2O5 [isopropyl 5-(2-hydrazineyl-2-oxoethoxy)-2-methylbenzofuran-3-carboxylate, Phase Databank ID: CACPD2011a-0001289047, Enamine ID: Z374510610], SD-03: C19H15NO7 [methyl 4-((7-(2-amino-2-oxoethoxy)-4-oxo-4H-chromen-3-yl)oxy)benzoate, Phase Databank ID: CACPD2011a-0001268860, Enamine ID: Z374511496], SD-04: C13H9ClN2O2S [2-(4-(5-chlorobenzofuran-2-yl)thiazol-2-yl)acetamide, Phase Databank ID: CACPD2011a-0001271129, Enamine ID:T6218428], SD-05:C14H16N2O3 [1-methoxy-3-((6-methoxynaphthalen-2-yl)methyl)urea, Phase Databank ID: CACPD2011a-0000738295, Enamine ID: T5848596] were purchased from ENAMINE Ltd., 78 Chervonotkatska Street, 02660 Kyiv, Ukraine. The 1H-NMR spectra of best three hits SD-01, SD-02 and SD-03 are attached (Fig. S6a, S6b and S6c).
Ligand preparation
All the structures of the inhibitors were drawn using 2D sketcher of Maestro 9.6 and then converted to their corresponding 3D structures. These inhibitors were geometrically refined using Ligprep module of Schrodinger43. During ligand preparation, the OPLS_2003 force field was used for energy minimization. Ligprep generates single, low energy, 3D structures, retaining its original state of chiralities and ionization for each input structure.
Protein preparation
The X-ray crystal structures of different PDB IDs of HDACs namely 4BKX, 3MAX, 4A69, 2VQM, 5WPB, 1T64 were prepared using ‘Protein Preparation Wizard’ workflow in Maestro 11.4 (Schrodinger Inc.). During protein preparation, water molecules were removed and then hydrogens were added to the protein and co-ligand. The energy of the complexes was then minimized until the RMSD between the minimized structure and the starting structure reached 0.30 Å, using the OPLS_2003 force field. The receptor grid box of 15 Å cube was generated by selecting the co-ligand of the active site except for HDAC1. The HDAC1 grid box was generated selecting active site amino acids ASP-264, ASP-176, and HIS-17835. The receptor grid of HDAC8 (PDB ID: 1T64) used in the present study was prepared earlier32.
Generation of common pharmacophore hypothesis
Phase module of Schrodinger44 provides six inbuilt pharmacophore features: hydrogen bond acceptor (A), hydrogen bond donor (D), hydrophobic group (H), negatively charged group (N), positively charged group (P), and aromatic ring (R). The common pharmacophores were identified using a tree-based partitioning technique which groups together similar pharmacophores according to their inter-site distances. Five featured pharmacophore hypotheses were selected and subjected to stringent scoring function analysis. The set of the generated hypotheses with their scoring values are summarized in Table S6. The distance and angle of pharmacophore features for all hypotheses are listed in the supplementary material Table S7 and Table S8. The best hypotheses ADDRR.4 consisting of five features are: one hydrogen bond acceptors (A2), two hydrogen bond donors (D3, D4), and two aromatic rings (R7 and R8) are shown in Fig. 2.
The building of 3D QSAR models
The PHASE provides the option of building a 3D QSAR model with the selected pharmacophore hypothesis. Top ranking hypotheses based on survival score (representing a weighted combination of the vector, site, volume scores) and survival – inactive score, ADDRR.4, AADRR.4, and AADDR.12, were subjected to 3D QSAR model building. During the model building randomly selected 70% molecules were kept in training set and atom based model was generated by keeping 1 Å grid spacing. Thus the maximum of PLS factors which can be used is N/5, where N symbolizes the number of ligands present in the training set. The model with a PLS factor four were considered as the best statistical models. The 3D QSAR models generated from hypotheses ADDRR.4 has admirable statistical parameters (Table 1) and was used for virtual screening of the database.
Validation of 3D QSAR model
Validation of 3D QSAR model is a crucial part, especially when the model is used for virtual screening or prediction of activity for external data set45. The performance of a model was measured by its predictive ability. The experimental activity of external selective HDAC8 inhibitors (1′–22′) retrieved from literature4,6,17,34,46,47,48 with their structures and predicted their activity using built 3D QSAR model (Fig. 4). These molecules were not included in the model building but used for model validation. The chemical features of the ligand structures are mapped to a cubic 3D grid by the 3D QSAR models.
Virtual screening of the database
Pharmacophore-based virtual screening (VS) of the database has been used to retrieve selective HDAC8 inhibitors. The 3D QSAR model built from pharmacophore ADDRR.4 was used to search the Phase database. The virtual screening workflow for the identification of HDAC8 inhibitors is presented in Fig. 14. The compounds of the database matching with minimum of four pharmacophoric features were identified through the ligand pharmacophore mapping process. The number of 500 top scored hits with fitness score ≥ 1.0 resulted from this step were subjected for ADME filtration.
ADME study
The QikProp 3.549, a prediction program designed by Prof. William L. Jorgensen was used to calculate ADME properties. It is a quick, accurate and predicts physically significant descriptors and pharmaceutically relevant properties of organic compounds. It has been widely used as a filter for molecules that would be expected to be further developed in drug design programs. QikProp provides ranges of values for comparing particular molecular properties of organic molecules with those of 95% of recognized drugs. The number of 500 hits resulted from virtual screening were filtered to find out a drug like hits and ADME properties of five finally selected hits with their acceptable range are listed in Table 2.
Validation of docking
The validation of the docking was measured by calculating the root mean square deviation (RMSD). Superimposing the original crystallographic bound conformation of co-ligand and its predicted docked conformations were expressed as RMSD. Lower the value of RMSD, higher the accuracy of docking and RMSD values less than 1.5 Å or 2.0 Å depending on ligand size are considered to have performed successfully50. Amongst the several HDACs, the lowest RMSD PDB ID of each HDACs was selected for molecular docking studies.
Molecular docking studies
The hits from ADME filtration and 32 known selective HDAC8 inhibitors were subjected for molecular docking study with HDAC8 to predict the binding affinity towards HDAC8 using Glide module of Schrodinger51,52,53,54. The best 20 hits with XP glide score >9.0 were further subjected to molecular docking studies with HDAC1, HDAC2, HDAC3, HDAC4, and HDAC6 to identify isoform selectivity of these hits.
E-Pharmacophoregeneration
The best selective known HDAC8 inhibitor PCI-34051(1) [N-hydroxy-1-(4-methoxybenzyl)-1H-indole-6-carboxamide] docked with a previously prepared grid of 1T64 proteins and energy-optimized structure-based pharmacophore (E-pharmacophore) were generated by selecting protein-PCI-34051 docked complex as per protocol55,56. The generated pharmacophore of PCI-34051 could effectively map all the important pharmacophoric features of PCI-34051- A1, A3, R7, and R9. The pharmacophore A1 and R9 superimposed on the oxygen atom of the methoxy group and benzene ring of methoxybenzyl group respectively whereas R7 and A3 superimposed on indole five member ring and hydroxamic acid carbonyl oxygen respectively, of PCI-34051. These generated pharmacophores were used for matching the finally selected five HDAC8 inhibitors.
References
Wang, L. T. et al. Aryl hydrocarbon receptor regulates histone deacetylase 8 expression to repress tumor suppressive activity in hepatocellular carcinoma. Oncotarget. 8, 7489–7501, https://doi.org/10.18632/oncotarget.9841 (2017).
Falkenberg, K. J. & Johnstone, R. W. Histone deacetylases and their inhibitors in cancer, neurological diseases and immune disorders. Nat Rev Drug Discov. 13, 673–691, https://doi.org/10.1038/nrd4360 (2014).
Zwergel, C., Stazi, G., Valente, S. & Antonello, M. Histone Deacetylase Inhibitors: Updated Studies in Various Epigenetic-Related Diseases. J Clin Epigenet. 2, 1–15, https://doi.org/10.21767/2472-1158.100015 (2016).
Chakrabarti, A. et al. HDAC8: a multifaceted target for therapeutic interventions. Trends Pharmacol. Sci. 36, 481–492, https://doi.org/10.1016/j.tips.2015.04.013 (2015).
Oehme, I., Deubzer, H. E., Lodrini, M., Milde, T. & Witt, O. Targeting of HDAC8 and investigational inhibitors in neuroblastoma. Expert Opin. Investig. Drugs. 18, 1605–1617, https://doi.org/10.1517/14728220903241658 (2009).
Tang, G. et al. Identification of a Novel Aminotetralin Class of HDAC6 and HDAC8 Selective Inhibitors. J. Med. Chem. 57, 8026–8034, https://doi.org/10.1021/jm5008962 (2014).
Mann, B. S., Johnson, J. R., Cohen, M. H., Justice, R. & Pazdur, R. FDA approval summary: vorinostat for treatment of advanced primary cutaneous T-cell lymphoma. The Oncologist. 12, 1247–1252, https://doi.org/10.1634/theoncologist.12-10-1247 (2007).
Iwamoto, M. et al. Clinical pharmacology profile of vorinostat, a histone deacetylase inhibitor. Cancer Chemother. Pharmacol. 72, 493–508, https://doi.org/10.1007/s00280-013-2220-z (2013).
FDA, Office of Oncology Drug Products. “What’s New from the Office of Oncology Drug Products” http://www.fda.gov/AboutFDA/CentersOffices/CDER/ucm093885.htm, https://doi.org/10.1093/jnci/djq030 (2010).
Bertino, E. M. & Otterson, G. A. Romidepsin: a novel histone deacetylase inhibitor for cancer. Expert Opin. Investig. Drugs. 20, 1151–1158, https://doi.org/10.1517/13543784.2011.594437 (2011).
Lee, H. Z. et al. FDA approval: belinostat for the treatment of patients with relapsed or refractory peripheral T-cell lymphoma. Clin. Cancer Res. 21, 2666–2670, https://doi.org/10.1158/1078-0432.CCR-14-3119 (2015).
Laubach, J. P., Moreau, P., San-Miguel, J. F. & Richardson, P. G. Panobinostat for the Treatment of Multiple Myeloma. Clin Cancer Res. 21, 4767–73, https://doi.org/10.1158/1078-0432.CCR-15-0530 (2015).
Quintás-Cardama, A. et al. Therapy with the histone deacetylase inhibitor pracinostat for patients with myelofibrosis. Leukemia Res. 36, 1124–1127, https://doi.org/10.1016/j.leukres.2012.03.003 (2012).
Lu, X., Ning, Z., Li, Z., Cao, H. & Wang, X. Development of chidamide for peripheral T-cell lymphoma, the first orphan drug approved in China. Intractable Rare Dis. Res. 5, 185–191, https://doi.org/10.5582/irdr.2016.01024 (2016).
Tan, J., Cang, S., Ma, Y., Petrillo, R. L. & Liu, D. Novel histone deacetylase inhibitors in clinical trials as anti-cancer agents. J Hematol Oncol. 3, 1–13, https://doi.org/10.1186/1756-8722-3-5 (2010).
Singh, R. K. et al. Kinetic and thermodynamic rationale for suberoylanilide hydroxamic acid being a preferential human histone deacetylase 8 inhibi tor as compared to the structurally similar ligand, trichostatin a. Biochemistry. 52, 8139–8149, https://doi.org/10.1021/bi400740x (2013).
Krennhrubec, K., Marshall, B. L., Hedglin, M., Verdin, E. & Ulrich, S. M. Design and evaluation of ‘Linkerless’ hydroxamic acids as selective HDAC8 inhibitors. Bioorg. Med. Chem. Lett. 17, 2874–2878, https://doi.org/10.1016/j.bmcl.2007.02.064 (2007).
Balasubramanian, S. et al. A novel histone deacetylase 8 (HDAC8)-specific inhibitor PCI-34051 induces apoptosis in T-cell lymphomas. Leukemia. 22, 1026–1034, https://doi.org/10.1038/leu.2008.9 (2008).
Thangapandian, S., John, S., Lee, Y., Kim, S. & Lee, K. W. Dynamic structure-based pharmacophore model development: a new and effective addition in the histone deacetylase 8 (HDAC8) inhibitor discovery. Int. J. Mol. Sci. 12, 9440–9462, https://doi.org/10.3390/ijms12129440 (2011).
Suzuki, T. et al. Novel histone deacetylase inhibitors: design, synthesis, enzyme inhibition, and binding mode study of SAHA-based non-hydroxamates. Bioorg. Med. Chem. Lett. 13, 4321–4326, https://doi.org/10.1016/j.bmcl.2003.09.048 (2003).
Feng, T. et al. Novel N-hydroxyfurylacrylamide-based histone deacetylase (HDAC) inhibitors with branched CAP group (Part 2). Bioorg. Med. Chem. 21, 5339–5354, https://doi.org/10.1016/j.bmc.2013.06.009 (2013).
Shen, J. et al. A kinome-wide RNAi screen identifies ALK as a target to sensitize neuroblastoma cells for HDAC8-inhibitor treatment. Cell Death Differ. 25, 2053–2070, https://doi.org/10.1038/s41418-018-0080-0 (2018).
Murumkar, P. R., Zambre, V. P. & Yadav, M. R. Development of predictive pharmacophore model for in silico screening, and 3D QSAR CoMFA and CoMSIA studies for lead optimization, for designing of potent tumor necrosis factor alpha converting enzyme inhibitors. J. Comput. Aided Mol. Des 24, 143–156, https://doi.org/10.1007/s10822-010-9322-z (2010).
Almerico, A. M., Tutone, M. & Lauria, A. 3D-QSAR pharmacophore modeling and in silico screening of new Bcl-xl inhibitors. Eur. J. Med. Chem. 45, 4774–4782, https://doi.org/10.1016/j.ejmech.2010.07.042 (2010).
Pasha, F. A., Muddassar, M., Neaz, M. M. & Cho, S. J. Pharmacophore and docking-based combined in-silico study of KDR inhibitors. J. Mol. Graph. Model. 28, 54–61, https://doi.org/10.1016/j.jmgm.2009.04.006 (2009).
Al-Sha’er, M. A. & Taha, M. O. Discovery of novel CDK1 inhibitors by combining pharmacophore modeling, QSAR analysis and in silico screening followed by in vitro bioassay. Eur. J. Med. Chem. 45, 4316–4330, https://doi.org/10.1016/j.ejmech.2010.06.034 (2010).
Tang, H. et al. Novel inhibitors of human histone deacetylase (HDAC) identified by QSAR modeling of known inhibitors, virtual screening, and experimental validation. J. Chem. Inf. Model. 49, 461–476, https://doi.org/10.1021/ci800366f (2009).
Wang, J. et al. Pharmacophore-Based Virtual Screening and Biological Evaluation of Small Molecule Inhibitors for Protein Arginine Methylation. J. Med. Chem. 55, 7978–7987, https://doi.org/10.1021/jm300521m (2012).
Chen, C. et al. Drug Design, Combining structure-based pharmacophore modeling, virtual screening, and in silico ADMET analysis to discover novel tetrahydro-quinoline based pyruvate kinase isozyme M2 activators with antitumor activity. Drug Des. Devel. Ther. 8, 1195–1210, https://doi.org/10.2147/DDDT.S62921 (2014).
Vidler, L. R. et al. Discovery of Novel Small-Molecule Inhibitors of BRD4 Using Structure-Based Virtual Screening. J. Med. Chem. 56, 8073–8088, https://doi.org/10.1021/jm4011302 (2013).
Thangapandian, S., John, S., Sakkiah, S. & Lee, K. W. Ligand and structure based pharmacophore modeling to facilitate novel histone deacetylase 8 inhibitor design. Eur. J. Med. Chem. 45, 4409–4417, https://doi.org/10.1016/j.ejmech.2010.06.024 (2010).
Debnath, T., Majumdar, S., Kalle, A. M., Aparna, V. & Debnath, S. Identification of potent histone deacetylase 8 inhibitors using pharmacophore-based virtual screening, three-dimensional quantitative structure–activity relationship, and docking study. Res. Rep. Med. Chem. 5, 21–39, https://doi.org/10.2147/RRMC.S81388 (2015).
Cao, G. P. et al. QSAR modeling to design selective histone deacetylase 8 (HDAC8) inhibitors. Arch. Pharm. Res. 39, 1356–1369, https://doi.org/10.1007/s12272-015-0705-5 (2016).
Kulandaivelu, U. et al. Hydroxamates of para-aminobenzoic acid as selective inhibitors of HDAC8. Bioorg. Chem. 57, 116–120, https://doi.org/10.1016/j.bioorg.2014.08.005 (2014).
Pidugu, V. R., Yarla, N. S., Pedada, S. R., Kalle, A. M. & Satya, A. K. Design and synthesis of novel HDAC8 inhibitory 2,5-disubstituted-1,3,4-oxadiazoles containing glycine and alanine hybrids with anti-cancer activity. Bioorg. Med. Chem. 24, 5611–5617, https://doi.org/10.1016/j.bmc.2016.09.022 (2016).
Debnath, S., Debnath, T., Majumdar, S., Arunasree, M. K. & Aparna, V. A combined pharmacophore modeling, 3D QSAR, virtual screening, molecular docking, and ADME studies to identify potential HDAC8 inhibitors. Med. Chem. Res. 25, 1–17, https://doi.org/10.1007/s00044-016-1652-5 (2016).
Millard, C. J. et al. The structure of HDAC1 in complex with the dimeric ELM2-SANT domain of MTA1 from the NuRD complex. Mol. Cell. 51, 57–67, https://doi.org/10.2210/pdb4BKX/pdb (2013).
Skene, R. J. & Jennings, A. J. Crystal Structure of Human HDAC2 complexed with an N-(2 aminophenyl) benzamide. RCSB Protein Data Bank. https://doi.org/10.2210/pdb3MAX/pdb (2010).
Watson, P. J., Fairall, L., Santos, G. M. & Schwabe, J. W. Structure of HDAC3 bound to co-repressor and inositol tetraphosphate. Nature. 481, 335–40, https://doi.org/10.1038/nature10728 (2012).
Bottomley, M. J., et al Structure of HDAC4 catalytic domain bound to a hydroxamic acid inhibitor. RCSB Protein Data Bank, https://doi.org/10.2210/pdb2VQV/pdb (2008).
Harding, R. J. et al. Structural Genomics Consortium (SGC). Crystal structure of fragment 3-(3-(pyridin-2-ylmethoxy)quinoxalin-2-yl)propanoic acid bound in the ubiquitin binding pocket of the HDAC6 zinc-finger domain. RCSB Protein Data Bank, https://doi.org/10.2210/pdb5WPB/pdb (2017).
Somoza, J. R. et al. Structure of human HDAC8 complexed with Trichostatin A. RCSB Protein Data Bank, https://doi.org/10.2210/pdb1T64/pdb (2004).
Versatile generation of accurate 3D molecular models. LigPrep, version 2.6, Schrödinger, LLC, https://www.schrodinger.com/ligprep (2019).
Dixon, S. L. et al. PHASE: a new engine for pharmacophore perception, 3D QSAR model development, and 3D database screening: 1. Methodology and preliminary results. J. Comput. Aided Mol. Des. 20, 647–671, https://doi.org/10.1007/s10822-006-9087-6 (2006).
Veerasamy, R. et al. Validation of QSAR models – strategies and importance. Int. J. Drug Des. Discov. 2, 511–519 (2011).
Suzuki, T. et al. Rapid Discovery of Highly Potent and Selective Inhibitors of Histone Deacetylase 8 Using Click Chemistry to Generate Candidate Libraries. J. Med. Chem. 55, 9562–9575, https://doi.org/10.1021/jm300837y (2012).
Suzuki, T. et al. Design, Synthesis, and Biological Activity of NCC149 Derivatives as Histone Deacetylase 8-Selective Inhibitors. Chem. Med. Chem. 9, 657–664, https://doi.org/10.1002/cmdc.201300414 (2014).
Tang, W., Luo, T., Greenberg, E. F., Bradner, J. E. & Schreiber, S. L. Discovery of histone deacetylase 8 selective inhibitors. Bioorg. Med. Chem. Lett. 21, 2601–2605, https://doi.org/10.1016/j.bmcl.2011.01.134 (2011).
Rapid ADME predictions of drug candidates. Qik Prop, Version 3.5, Schrodinger, LLC, New York, https://www.schrodinger.com/qikprop (2013).
Hevener, K. E. et al. Validation of Molecular Docking Programs for Virtual Screening against Dihydropteroate Synthase. J. Chem. Inf. Model. 9, 444–460, https://doi.org/10.1021/ci800293n (2009).
A complete solution for ligand-receptor docking. Glide, version 6.0, Schrödinger, LLC, New York, NY, https://www.schrodinger.com/glide (2013)
Friesner, R. A. et al. Glide: a new approach for rapid, accurate docking and scoring. 1. Method and assessment of docking accuracy. J. Med. Chem. 47, 1739–1749, https://doi.org/10.1021/jm0306430 (2004).
Halgren, T. A. et al. Glide: a new approach for rapid, accurate docking and scoring. 2. Enrichment factors in database screening. J. Med. Chem. 47, 1750–1759, https://doi.org/10.1021/jm030644s (2004).
Friesner, R. A. et al. Extra precision glide: docking and scoring incorporating a model of hydrophobic enclosure for protein-ligand complexes. J. Med. Chem. 49, 6177–6196, https://doi.org/10.1021/jm051256o (2006).
Salam, N. K., Nuti, R. & Sherman, W. Novel Method for Generating Structure-Based Pharmacophores Using Energetic Analysis. J. Chem. Inf. Model. 49, 2356–2368, https://doi.org/10.1021/ci900212v (2009).
Loving, K., Salam, N. K. & Sherman, W. Energetic analysis of fragment docking and application to structure-based pharmacophore hypothesis generation. J. Comput. Aided Mol. Des. 23, 541–554, https://doi.org/10.1021/ci900212v (2009).
Acknowledgements
The authors are thankful to the Department of Biotechnology, New Delhi, Govt. of India for providing financial support [F.No.BT/327/NE/TBP/2012] for this research work. The authors are also thankful to the CSIR-IICT National MolBank facility for HDAC inhibitory activity screening the molecules.
Author information
Authors and Affiliations
Contributions
S.D. designed the study, analysed the results, performed the data analysis, wrote the manuscript and produced the figures. T.D. was responsible for the pharmacophoe building, docking, wrote the manuscript. S.B. conceived the experiments, data analysis and wrote the manuscript. S.M. performed the spectral analysis, wrote the manuscript, A.M.K. conceptualized study, performed bioassay analysis and wrote the manuscript, V.A. conceptualized study, conceived modeling analysis and manuscript preparation.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supporting Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Debnath, S., Debnath, T., Bhaumik, S. et al. Discovery of novel potential selective HDAC8 inhibitors by combine ligand-based, structure-based virtual screening and in-vitro biological evaluation. Sci Rep 9, 17174 (2019). https://doi.org/10.1038/s41598-019-53376-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-53376-y
This article is cited by
-
Flexible Hybrid Table Recognition and Semantic Interpretation System
SN Computer Science (2023)
-
Identification of potent HDAC 2 inhibitors using E-pharmacophore modelling, structure-based virtual screening and molecular dynamic simulation
Journal of Molecular Modeling (2022)
-
Artificial intelligence to deep learning: machine intelligence approach for drug discovery
Molecular Diversity (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.