Abstract
The Gram-positive bacterium Listeria monocytogenes occurs ubiquitously in the environment and infects humans upon ingestion. It encodes four PadR-like repressors, out of which LftR has been characterized previously and was shown to control gene expression in response to the antibiotic aurantimycin produced by other environmental bacteria. To better understand the PadR regulons of L. monocytogenes, we performed RNA-sequencing with mutants of the other three repressors LadR, LstR and Lmo0599. We show that LadR is primarily responsible for the regulation of the mdrL gene, encoding an efflux pump, while LstR and Lmo0599 mainly regulate their own operons. The lstR operon contains the lmo0421 gene, encoding a homolog of the RodA/FtsW protein family. However, this protein does not possess such functionality, as we demonstrate here. The lmo0599 operon contains two additional genes coding for the hypothetical trans-membrane proteins lmo0600 and lmo0601. A striking phenotype of the lmo0599 mutant is its impaired growth at refrigeration temperature. In light of these and other results we suggest that Lmo0599 should be renamed and propose LltR (listerial low temperature regulator) as its new designation. Based on the nature of the PadR target genes we assume that these repressors collectively respond to compounds acting on the cellular envelope.
Similar content being viewed by others
Introduction
The Gram-positive bacterium Listeria monocytogenes is the causative agent of listeriosis, which is one of the most serious foodborne bacterial infections in humans. The bacterium occurs ubiquitously in the environment and infects humans after consumption of contaminated food. Uncooked and ready-to-eat foods pose the highest risk of infection, the latter because L. monocytogenes is able to grow at refrigeration temperatures1,2. Fatality rates of listeriosis are remarkably high compared to other foodborne bacterial pathogens3, and, hence, the control of L. monocytogenes in food is of utmost importance for the food-processing industry. L. monocytogenes frequently enters the food chain due to its wide-spread presence in the soil, in surface waters, on plants and in the gut of various animals4. Aggravating this situation, the bacterium has a profound capacity to resist many conditions used to prevent food spoilage. It is not only able to grow at 4 °C, but it can also grow at high salt concentrations, accepts a wide pH range for growth and tolerates anti-microbial compounds of cold smoke5,6. L. monocytogenes readily forms biofilms on glass, plastic and steel surfaces7, often complicating effective disinfection of food-processing plants. Moreover, isolates of L. monocytogenes frequently are resistant against commonly used disinfectants such as benzalkonium chloride8,9 and such benzalkonium-resistant isolates have caused big outbreaks in the past10,11. Thus, an improved understanding of the ecology, survival strategies and stress responses of L. monocytogenes is important to reduce the entry and of the bacterium into the food chain and its persistence in food-processing plants.
Among the diverse molecular mechanisms employed by bacteria to sense and respond to environmental stresses are the PadR-type transcriptional regulators. The eponymous protein for this class of repressors is the phenolic acid decarboxylase repressor PadR of the firmicute bacterium Pediococcus pentosaceus12. It activates the expression of phenolic acid decarboxylase (PadA) in response to the exposure to toxic phenolic acids. PadA then converts the toxic phenolic acids into less toxic products, thereby conveying resistance to high levels of phenolic acids12,13. Another well-studied PadR-type repressor, LmrR from Lactococcus lactis, activates expression of the lmrCD multi drug efflux pump genes upon exposure to small toxic compounds like the antibiotic daunorubicin14,15. Under non-inducing conditions, LmrR represses lmrCD transcription by blocking the PlmrCD promoter15,16. Binding of effector molecules induces conformational changes in LmrR causing relieve of repression14,17,18 and compound excretion through LmrCD19.
In L. monocytogenes, the PadR-type repressor LftR controls the expression of the lieAB genes encoding another antibiotic efflux pump20. Recently, we demonstrated that this efflux pump is expressed when L. monocytogenes comes into contact with aurantimycin A21, a depsipeptide antibiotic with very potent bactericidal activity against Gram-positive bacteria and produced by soil-dwelling Streptomyces aurantiacus22. Thus, LftR and LieAB likely promote survival of L. monocytogenes when it comes in contact with S. aurantiacus in the soil, its natural reservoir21, from where it may eventually enter the food chain.
Next to LftR, three additional PadR-like repressors are encoded in the L. monocytogenes EGD-e genome: LadR, LstR and Lmo0599. LadR controls production of the multi drug resistance pump MdrL23, associated with benzalkonium chloride resistance24,25. LstR was linked to heat resistance26 and Lmo0599 has not been characterized until now. In order to better understand the function of PadR-like regulators in L. monocytogenes, we here have identified the regulons of the three mentioned repressors by RNA-Seq and studied the function of their primary effector genes in genetic and functional experiments. In light of the results obtained we suggest to rename Lmo0599 as LltR (listerial low temperature regulator).
Results
Identification of the LadR, LstR and LltR regulons
In order to determine the target genes of LadR, LstR and LltR, the ladR and lstR genes were removed from the genome by allelic exchange and lltR was replaced with a non-functional copy carrying the L49A R51A L52A triple mutation in its helix-turn-helix motif. This triple mutation (lltR*) was designed to prevent binding of LltR to its recognition site. Next, transcriptomes of the L. monocytogenes wild type strain EGD-e and its isogenic ΔladR, ΔlstR and lltR* mutants were analyzed by RNA sequencing. This revealed massive (~150-fold) derepression of the mdrL gene in cells lacking LadR (Table 1), which is in good agreement with previous work demonstrating that mdrL expression is repressed by LadR23. MdrL is a multidrug efflux transporter and supposedly transports compounds like ethidium bromide, cefotaxime and other antibiotics out of the cell27. Moderate induction of the lmo1618-1617 operon28 was also observed in the ΔladR mutant (Table 1), as described previously29. This bicistronic transcription unit comprises the lmo1618 gene coding for a MarR-type regulator followed by lmo1617 encoding the multidrug resistance transporter MdrM. MdrM is involved in the secretion of the signal molecule cyclic diadenosine monophosphate (c-di-AMP), which activates the host cytosolic surveillance pathway during intracellular passages29,30. The expression of other genes was not affected, indicating that LadR specifically regulates the expression its three target genes, among which mdrL appears to be the primary one.
Deletion of lstR caused strong (~140-fold) overexpression of the lmo0421 and lmo0423 genes (Table 1), which form a tricistronic operon together with lstR (lmo0423-lstR-lmo0421)28. While the lmo0421 gene is one of six L. monocytogenes RodA/FtsW paralogs, it is unclear whether it is also involved in peptidoglycan chain polymerization31,32. The lmo0423 gene shares homology with ECF-type sigma factors (27.9% identity to Bacillus subtilis σV) and has been designated σC 26. All three genes of this operon have been implicated in heat stress response26 and σC is also important during cold adaptation33. Only few possible target genes are known for this sigma factor34,35 and its main target is the lmo0423-lstR-lmo0421 operon itself 26. The two divergently transcribed and uncharacterized lmo0420 and lmo0419 genes located downstream of the sigC operon are also considerably overexpressed in the ΔlstR mutant (Table 1). Transcriptional read-through could be an explanation for this. The remaining LstR-affected genes, among which is the lmo0416 gene coding for a putative transcriptional regulator, show lower fold changes and it remains unclear whether these are direct or secondary effects.
A similar hierarchy in fold changes was observed for genes de-repressed in the lltR* mutant. Here, the lltR-lmo0600-lmo0601 operon28 was massively overexpressed (~100- to 150-fold), most likely due to relief of auto-repression through LltR. Derepression of lmo0602, located downstream of this operon and transcribed in the same direction could be due to transcriptional read-through. Besides this, six mildly overexpressed genes were found, among those the liaIH operon, encoding phage shock proteins36, as well as the lmo1636 and lmo1637 genes coding for components of a putative ABC transporter (Table 1).
Promoter fragments controlled by LadR, LstR and Lm0599
To verify the data obtained by RNA-seq, fragments upstream of the mdrL, sigC and lltR genes, all between 200 and 370 bp in length and comprising the start codons of each gene, were fused in frame to the lacZ gene. These promoter-lacZ fusions were then introduced into wild type and mutant backgrounds lacking their cognate PadR-like repressors. Only background β-galactosidase activities, which did not exceed the values obtained with strain LMSH16 carrying a promoter-less lacZ gene for control, were observed for two of the three lacZ fusions in wild type cells. An exception was the PsigC promoter that resulted in a roughly fourfold higher β-galactosidase activity in wild type cells when compared to the strain with the promoter-less lacZ (Fig. 1a). This increased background activity might be explained by the presence of three promoters in front of the sigC-lstR-lmo0421 operon, including one σC-dependent promoter26. Activity of the PsigC-promoter increased 77-fold in the absence of lstR, indicating that induced transcription of the sigC-lstR-lmo0421 operon in the ΔlstR mutant is due to increased promoter activity. Likewise, β-galactosidase activity driven by the PmdrL promoter increased in the absence of ladR, even though only 17-fold, and activity of the PlltR promoter increased over 50-fold in the lltR* mutant (Fig. 1a). Taken together, the activities of the three tested promoters increase substantially in the absence of their cognate PadR repressor proteins. This indicates that LadR, LstR and LltR repress initiation of transcription from the PmdrL, PsigC and PlltR promoters, respectively. We tested induction of all three promoter-lacZ fusions in disc diffusion assays on X-Gal containing agar plates with rhodamine 6G, a known inducer of mdrL expression in L. monocytogenes LO2823, however, induction was not observed (data not shown). Likewise, none of the three promoter-lacZ fusions was induced by ethidium bromide, several antibiotics (penicillin G, fosfomycin, cycloserine, chloramphenicol, tetracycline, spectinomycin, ampicillin, kanamycin, nalidixic acid) or selected disinfectants (benzalkonium, tert-butylhydroquinone, acriflavine).
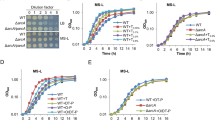
Activity of LadR-, LstR- and LltR-dependent promoters. (a) β-galactosidase activity in strains carrying lacZ fusions of the PmdrL, PlltR and PsigC promoters. Strains LMSH10 (PmdrL-lacZ), LMSH11 (ΔladR PmdrL-lacZ), LMSH14 (PlltR-lacZ), LMSH15 (lltR* PlltR-lacZ), LMSH12 (PsigC-lacZ) and LMSH13 (ΔlstR PsigC-lacZ) were grown in BHI broth at 37 °C to mid-logarithmic growth phase and β-galactosidase activity was determined. The experiment was repeated three times and average values and standard deviations are shown. Asterisks indicate significant differences (P < 0.05). (b) Scheme illustrating gene arrangement at the ladR, lstR and lltR loci. Promoters are either adopted from experimental data26,28 or predicted using the bprom algorithm56. Promoter fragments used for construction of promoter-lacZ fusions are indicated.
Effect of PadR repressors on in vitro and in vivo growth of L. monocytogenes
Next, we analyzed the contribution of the three PadR-like repressor proteins to growth in batch culture and inside eukaryotic cells. All three mutant strains grew like wildtype at 37 °C (Fig. 2a), 30 °C or 42 °C (data not shown). In contrast, strain LMSH3 carrying the mutated lltR* allele was nearly unable to grow at 6 °C, while the other two mutants showed wild type-like growth (Fig. 2b). This growth defect of the lltR* mutant was also evident during growth on BHI agar plates, but could be complemented by ectopic expression of a native lltR allele (Fig. 2c), demonstrating that the lltR* mutation and no secondary site mutation was the cause of the cold-sensitive phenotype.
Growth of L. monocytogenes ladR, lstR and lltR mutants. (a) L. monocytogenes strains EGD-e (wt), LMSH1 (ΔladR), LMSH2 (ΔlstR) and LMSH3 (lltRL49A R51A L52A) were grown in BHI broth at 37 °C. (b) Growth of the same set of strains at 6 °C. Growth curves were repeated three times and average values and standard deviations are shown. (c) Complementation of the cold-sensitive growth defect of the lltR mutant. L. monocytogenes strains EGD-e (wt), LMSH3 (lltRL49A R51A L52A) and the complemented strain LMSH42 (+lltR) were grown for six weeks on BHI agar containing 1 mM IPTG at 6 °C.
The contribution of the three PadR-like repressors to intracellular growth was then studied using the J774 mouse macrophage infection model. This showed that the ΔladR, ΔlstR and lltR* mutant strains were phagocytosed as the wild type strain and that their intracellular growth was unaffected (Fig. S1). Apparently, these proteins are not important during infection under the tested conditions.
Identification of the effector gene of the lltR operon
In addition to the LltR repressor itself, the lltR operon encodes two more genes28. Immediately downstream of lltR is lmo0600, encoding a multi-spanning integral membrane protein of unknown function that contains a DUF1700 domain37. Further downstream there is lmo0601 coding for a hypothetical exoprotein containing a DUF4097 domain37, which is annotated as a possible structural element of bacterial adhesins38. We wondered whether the cold-sensitive growth phenotype of the lltR* mutant results from overexpression of lmo0600, lmo0601 or both genes. To study this, both genes were individually deleted in the lltR* mutant background and the ability of the resulting mutants to grow at 6 °C was then tested on BHI agar plates. This revealed that deletion of lmo0600 restored normal growth in the lltR* Δlmo0600 double mutant, whereas the lltR* Δlmo0601 was as impaired to grow at 6 °C as the lltR* single mutant strain (Fig. 3). This shows that overexpression of the transmembrane protein Lmo0600 is detrimental for growth at low temperatures and emphasizes the importance of LltR in repression of lmo0600 transcription for growth at refrigeration temperature.
Analysis of the effector gene of the sigC operon
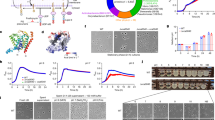
Of particular interest among the PadR-regulated genes is the third gene of the sigC operon, lmo0421, which represents the effector gene of this operon and encodes an uncharacterized protein with reasonable similarity to FtsW/RodA proteins. Recent evidence showed that these proteins constitute peptidoglycan glycosyltransferases31,32. L. monocyctogenes encodes six FtsW/RodA proteins in total39, and among these, Lmo0421 represents a non-canonical homologue that clusters separate from RodA and FtsW proteins (Fig. 4a). We wondered whether lmo0421 could substitute for any of the two FtsW-like or any of the three RodA-like proteins of L. monocytogenes when overexpressed. To study this, we first constructed a novel lstR mutant (strain LMSH40), in which three critical amino acids in the conserved operator recognition site of LstR were replaced by alanines (lstR L90A L92A L93A, designated lstR*). In this mutant, the overall architecture of the sigC operon remains intact, however the PsigC promoter is de-repressed to the same degree as observed in a ΔlstR deletion mutant (Fig. 4b). Interestingly, expression of the sigC operon was completely dependent on σC, as inactivation of the sigC gene in wild type and in lstR* backgrounds reduced activity of the PsigC-lacZ reporter even below that of strain LMSH12 (wt PsigC-lacZ, P > 0.01). Apparently, LstR and σC jointly control expression of the sigC operon in inverse directions. Despite lmo0421 overexpression in lstR inactivated cells, effects on the sensitivity of the lstR* or lstR* Δlmo0421 mutant against antibiotics affecting different steps in cell wall biosynthesis were not detected (Fig. 4c). Alterations in cell wall ultrastructure were also not observed, questioning a possible role of Lmo0421 in cell wall biosynthesis (Fig. S2).
Susceptibility of L. monocytogenes lstR and lmo0421 mutants against antibiotics targeting peptidoglycan biosynthesis. (a) UPGMA tree of the six L. monocytogenes and the three B. subtilis FtsW/RodA homologues. Legend: Substitutions per site. (b) Effect of the lstR L90A L92A L93A mutation and σC inactivation on the activity of the PsigC promoter. L. monocytogenes strains LMSH16 (wt, lacZ) LMSH12 (wt, PsigC-lacZ), LMSH13 (ΔlstR PsigC-lacZ), LMSH63 (lstR* PsigC-lacZ), LMSH96 (sigC*; PsigC-lacZ) and LMSH97 (lstR* sigC* PsigC-lacZ) were grown in BHI broth at 37 °C to mid-logarithmic growth phase and β-galactosidase activity was determined. The experiment was repeated three times and average values and standard deviations are shown. Asterisks indicate significant differences (t-test, P < 0.01). (c) L. monocytogenes strains EGD-e (wt), LMSH39 carrying the lstRL90A L92A L93A mutation (lstR*) and LMSH40 additionally lacking the ftsW/rodA homologue lmo0421 (lstR* Δlmo0421) were tested in filter disc susceptibility assays using penicillin G (1 mg/ml), vancomycin (20 mg/ml), moenomycin (1.6 mg/ml), bacitracin (40 mg/ml), fosfomycin (20 mg/ml) or cycloserine (30 mg/ml). Tests were repeated three times and average values and standard deviations are shown.
In order to test whether Lmo0421 could substitute for FtsW1 or FtsW2, we then introduced plasmids allowing insertional disruption of ftsW1 (lmo1071) and ftsW2 (lmo2688) into the wild type and the lstR* mutant. These plasmids are maintained as extrachromosomal replicons at permissive temperature (30 °C) and, consequently, strains transformed with these vectors grow on erythromycin-containing BHI plates at 30 °C (data not shown). However, plasmid replication is blocked and only clones that have integrated the plasmid into the chromosome can grow in the presence of erythromycin at non-permissive temperature (42 °C, Fig. 5a,b). In good agreement with previous results39, L. monocytogenes wild type tolerated disruption of ftsW2 at 42 °C, whereas ftsW1 could not be inactivated by plasmid integration, illustrating essentiality of ftsW1 and dispensability of ftsW2. Importantly, this gene essentiality pattern was not changed in the lstR* mutant, clearly demonstrating that overproduced Lmo0421 cannot compensate for the loss of FtsW1 function (Fig. 5c). In order to test whether Lmo0421 can functionally replace one of the three RodA proteins, we first deleted the rodA2-rodA1 genes in the wild type and the lstR* mutant. These strains were then transformed with a plasmid that allows insertional disruption of rodA3. While rodA3 can be readily inactivated in wild type and the lstR* mutant, where the rodA2-rodA1 genes are still present, this is not possible in the ΔrodA2-rodA1 strain. This result confirms previous findings showing that at least one of the three RodA proteins is required for viability of L. monocytogenes39. However, disruption of rodA3 was also not possible in the lstR* ΔrodA2-rodA1 mutant, indicating that one of the RodA homologs is required for viability of L. monocytogenes even when Lmo0421 is overexpressed. It is important to note that disruption of ftsW2 is not tolerated in ΔrodA2-rodA1 strains because plasmid insertion into ftsW2 would separate rodA3 from the promoter of the operon upstream of lmo268928. Taken together, we conclude that Lmo0421 has neither FtsW nor RodA functionality under these conditions.
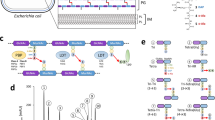
Lmo0421 cannot take over the function of any other FtsW/RodA protein. (a) Scheme showing genetic arrangement of the six ftsW/rodA genes in L. monocytogenes. (b) Scheme illustrating the way of insertional disruption chosen to inactivate the ftsW1, ftsW2 and rodA3 genes. (c) Insertional disruption of the ftsW1, ftsW2 and rodA3 genes in L. monocytogenes strains EGD-e (wt), LMSH39 (lstR*), LMSH67 (ΔrodA2-rodA1) and LMSH68 (lstR* ΔrodA2-rodA1). Temperature sensitive plasmids designed to disrupt the ftsW1 (pSAH66), ftsW2 (pSAH68), or rodA3 genes (pSAH67) or in Campbell-type integration events were forced to integrate into their respective target gene in the different strain backgrounds during growth on BHI agar plates containing erythromycin at 42 °C. Colony formation indicates chromosomal plasmid integration and target gene disruption as depicted in panel B. Please note that all strains can grow on BHI/erythromycin plates at 30 °C.
Discussion
Shared and specific features of the four PadR-type transcriptional repressors of L. monocytogenes begin to emerge. The three repressors studied here specifically control transcription of a small set of genes, usually comprising one to two affected transcription units per repressor. Their target genes are strongly repressed under standard growth conditions in wild type and de-repressed roughly 150-fold in the respective repressor mutants. This regulation patterns suggests that they are disadvantageous during exponential growth but highly beneficial under specific conditions. Among the four, LftR is the strongest repressor as it represses its target promoters about 450-fold21. Another shared feature is the presence of negative feedback loops in the gene expression control circuits. LftR21, LltR and LstR (this work) are autoregulatory by repressing their own genes. Whether LadR represses transcription of its own gene could not be decided based on our RNA-Seq data since the monocistronic ladR transcript is simply absent in the ΔladR mutant. However, the ladR and mdrL promoters are in close proximity23, so that control of both genes by a single LadR operator seems possible. These negative feedback loops switch off transcription when the inducing molecules or conditions are no longer present.
With σC, a second, but positive feedback loop is enmeshed in the control circuit of the sigC operon. This creates the possibility to integrate a second signal so that the sigC operon would only be fully induced when LstR relieves its repression and σC is activated. Provided that σC is not completely active during exponential growth, this would suggest that we only observe an intermediate level of sigC-lstR-lmo0421 transcription in the lstR mutant. σC is important for growth of the L. monocytogenes strain 10403 S at low33 and high temperatures26, but we could not observe similar effects in the L. monocytogenes EGD-e background (Fig. 2b and data not shown). L. monocytogenes σC shares certain homology with extracytoplasmic function (ECF) sigma factors and is the only sigma factor of this type in strain EGD-e26, but it is not known how exactly σC contributes to transcription of its operon. Remarkably, the sigC operon is not present in the entire L. monocytogenes population and only found in strains of phylogenetic lineage II40, to which EGD-e and 10403 S belong. Consistent with this observation and in good agreement with previous results26,35, σC does only activate its own promoter, so that lmo0421 is the only target gene of σC and LstR. ECF sigma factors respond to signals that attack the integrity of the membrane and the cell wall, such as antimicrobial peptides and lytic enzymes41. RodA/FtsW enzymes act in peptidoglycan biosynthesis as glycosyltransferases mediating elongation of peptidoglycan chains31,32,42. According to our experiments, Lmo0421 cannot take over the function of any of the other five RodA/FtsW enzymes and its overproduction in an lstR* background did not affect susceptibility against moenomycin that inhibits glycosyltransferase activity in penicillin binding proteins43. This latter observation suggests that Lmo0421 does not contribute to peptidoglycan transglycosylation under these conditions. However, Lmo0421 may functionally replace one of the other FtsW/RodA proteins under more specific conditions, for example when the FtsW/RodA glycosyltransferases are inhibited by more specific drugs. FtsW/RodA inhibitors are not known, but just recently, a supposedly inhibitory molecule of so far unknown structure has been discovered in a natural compound library screen with a B. subtilis mutant devoid of all transglycosylase activity mediated by penicillin binding proteins31. This compound (preliminary designation 654/A) or related substances might be recognized by LstR (and/or σC) under conditions where the house-keeping RodA/FtsW enzymes are chemically inactivated, leading to production of Lmo0421 as a 654/-resistant back-up protein. Interestingly, compound 654/A like the inductor of LftR, aurantimycin A, is produced by a soil-dwelling Streptomyces strain21,31. However, in order to test this hypothesis the identity of 654/A must be elucidated first. Alternatively, Lmo0421 might use chemically modified lipid II as substrate, which may be produced to confer resistance to antimicrobial peptides44. Regardless of these considerations, a role for Lmo0421 in cell wall biosynthesis is supported by a spontaneous lmo0421 mutation found in a stable L. monocytogenes L-form strain that lacks a cell wall45.
An inducer for LltR is presently not known. Rhodamine 6G was found to induce mdrL transcription in the background of L. monocytogenes LO28 suggesting that LadR senses rhodamine 6G. However, we cannot confirm this for strain EGD-e carrying the PmdrL-lacZ fusion in agar diffusion test (data not shown). Moreover, rhodamine dyes are of synthetic compounds and thus, other naturally occurring LadR effector molecules must exist. Based on the above mentioned considerations we speculate that these molecules, which need to be identified in future work, might be of streptomycetes origin. This would then be another common feature of listerial PadR-like repressors.
Materials and Methods
Bacterial strains and growth conditions
All strains used in this study are listed in Table 2. L. monocytogenes was generally cultivated in BHI broth or on BHI agar plates at 37 °C if not stated otherwise. Where required, antibiotics and supplements were added at the following concentrations: erythromycin (5 µg mL−1), kanamycin (50 µg mL−1) and X-Gal (100 µg mL−1). Escherichia coli TOP10 was used as standard cloning host46.
General methods, manipulation of DNA and oligonucleotide primers
Transformation of E. coli and isolation of plasmid DNA was performed according to standard methods46. Preparation of electro-competent L. monocytogenes cells and transformation of L. monocytogenes were done as described elsewhere47. Restriction and ligation of DNA was carried out as detailed in the manufacturer’s instructions. For restriction free modification of plasmids an altered version of the original QuikChange mutagenesis protocol was employed48. All primer sequences are listed in Table 3. Antibiotic susceptibility assays were recorded using filter discs soaked with solutions of antibiotics as indicated. L. monocytogenes colonies were grown over night in BHI broth and used to swab-inoculate BHI agar plates. Filter discs soaked with antibiotics were placed on top of the agar surface and the plates were incubated at 37 °C overnight.
Construction of plasmids and strains
Plasmid pSAH1 was constructed for deletion of the ladR gene. To this end, fragments up- and downstream of ladR were amplified by PCR using the primers SAH32/SAH33 and SAH34/SAH35 and both fragments were fused together by splicing by overlapping extension PCR (SOE-PCR) with SAH32/SAH35 as the primers. The resulting fragment was ligated into pMAD using BamHI/EcoRI.
Plasmid pSAH3 was constructed for removal of lstR. To this end, regions up- and downstream of lstR were amplified with the primers SAH038/SAH039 and SAH036/SAH037, respectively. These fragments were fused together by SOE-PCR and the resulting fragment was cloned into pMAD using BamHI/NcoI. An unwanted duplication of around 50 bp directly after the BamHI restriction site was removed by digesting the plasmid with BamHI and subsequent self-ligation, finally yielding pSAH3.
Plasmid pSAH5 was constructed for the introduction of the lltRL49A R51A L52A triple mutation (lltR*) into the chromosome and was obtained in two steps. First, the lltR gene was amplified by PCR from EGD-e chromosomal DNA using the primer pair SAH52/SAH53 and cloned into pMAD using EcoRI/NcoI, resulting in plasmid pSAH4. The L49A R51A L52A exchanges were then introduced into pSAH4 by quikchange mutagenesis using SAH58/SAH59 as the mutagenic primers.
Plasmid pSAH33 was generated to introduce the lstRL90A L92A L93A (lstR*) into the chromosome. To this end, fragments upstream and downstream of the region to be mutated were amplified with primers SAH178/SAH177 and SAH176/SAH175 (SAH177 and SAH176 introduced the lstR* mutations), both fragments were combined by SOE-PCR and the resulting fragment was introduced into pMAD by restriction-free cloning.
Plasmid pSAH32 facilitates lmo0421 deletion and was obtained by amplification of fragments up- and downstream of lmo0421 using the primers SAH178/SAH181 and SAH180/SAH179, respectively. Both fragments were fused together in a SOE-PCR and the resulting fragment was cloned into pMAD using EcoRI/NcoI. Plasmid pSAH32 was then used as the template in a quick change PCR using the primers SAH176/SAH177 to introduced the lstR* mutation, yielding pSAH34.
In order to remove lmo0600 from lltR* cells, plasmid pSAH45 was constructed. To this end, fragments up- and downstream of lmo0600 were amplified in PCRs with SAH212/SAH213 and SAH214/SAH215 and chromosomal DNA of strain LMSH3 (lltR*) as the template, respectively, and joined in a SOE-PCR, the product of which was inserted into pMAD by restriction-free cloning.
Plasmid pSAH46 was generated to remove the lmo0601 gene. Here, lmo0601 up- and downstream fragments were PCR amplified with SAH216/SAH217 and SAH218/SAH219, respectively, joined by SOE-PCR and inserted into pMAD by restriction-free cloning.
Plasmid pSAH62 was designed for removal of the rodA2-rodA1 genes. For this, fragments up- and downstream to the rodA2-rodA1 cluster were amplified with SAH262/SAH261 and SAH260/SAH259, spliced together by SOE-PCR and introduced into pMAD by restriction-free cloning.
Plasmid pSAH69 was constructed by amplification of the 5′- and 3′-halves of sigC using the primer pairs SAH256/SAH255 and SAH254/SAH253, respectively, their subsequent joining by SOE-PCR and the cloning of the obtained fragment into pMAD using restriction-free cloning. SAH255 and SAH254 introduced a premature stop codon at the 39th base pair triplet of sigC followed by a KpnI site (sigC*).
Derivatives of pMAD designed for gene deletions were transformed into the respective L. monocytogenes recipient strains and genes were deleted as described elsewhere49. All gene deletions were confirmed by PCR.
For insertional disruption of ftsW1, rodA3 and ftsW2, plasmids pSAH66, pSAH67 and pSAH68, respectively, were constructed. To this end, internal gene fragments were amplified by PCR using primers SAH257/SAH258 (ftsW1), SAH269/SAH270 (rodA3) and SAH263/SAH264 (ftsW2) and inserted into pMAD by restriction-free cloning. Plasmids pSAH66-67 were then introduced into L. monocytogenes strains by electroporation and transformants were selected on BHI agar containing erythromycin at 30 °C.
For construction of promoter-lacZ fusions, fragments carrying the PsigC, PmdrL and PlltR promoters were amplified by PCR using the primer pairs SAH109/SAH110, SAH113/SAH114 and SAH115/SAH116, respectively, and introduced into pBP117 by restriction-free cloning50, resulting in plasmids pSAH12, pSAH14 and pSAH15, respectively.
Plasmid pSAH37 for IPTG-dependent expression of lltR was generated by amplification of lltR using primers SAH185/SAH186 and the insertion of the resulting fragment into pIMK3 by restriction-free cloning.
Derivatives of pBP117 and pIMK3 were introduced into L. monocytogenes strains by electroporation and selected on BHI agar plates containing kanamycin. Plasmid insertion at the attB site of the tRNAArg locus was verified by PCR.
mRNA isolation
mRNA was isolated as described previously21. Briefly, cells from 25 ml of a culture grown in BHI broth (OD600 of ~0.8) was mixed with 25 ml ice cold killing buffer (20 mM Tris-HCl pH 7.5, 5 mM MgCl2, 20 mM NaN3) and harvested by centrifugation after 5 min incubation on ice.
RNA extraction followed the protocol of Gertz et al.51 with modifications as described18. 10 µg total RNA were digested with DNAse using the RNase-Free DNase Set (Qiagen) and then purified using RNA clean & concentrator columns (Zymo Research) for purification for RNA molecules longer than 200 nucleotides. RNA quality was assessed using Agilent Bioanalyzer RNA Nano chips. rRNA was depleted using the Ribo-Zero Bacteria Kit (Illumina), 2 µg purified RNA was treated with 10 µl Ribo-ZeroRemoval Solution and pelleted by ethanol precipitation. RNA concentrations were determined in a Qubit® fluorometer.
Library preparation and sequencing
RNA libraries were prepared using the TruSeq® Stranded mRNA Kit as described35. RNA transcripts were quantified by quasi-mapping of the reads to the L. monocytogenes EGD-e cDNA (Listeria_monocytogenes_egd_e.ASM19603v1.cds.all.fa.gz), provided by the Ensembl Genomes server52, using the Salmon software53. Average expression from three biological replicates of the mutant divided by the average expression from three biological replicates of the wildtype gave the differential expression ratio. Log2-transformed transcript counts from three biological replicates were then used to calculate P values using Students t-test. Significantly differentially expressed genes were defined as having a P-value less than 0.01, an absolute differential expression factor of more than 2 and an expression level of at least 10 TPM.
β-galactosidase reporter assays
Reporter strain cultures were grown in BHI broth at 37 °C until an OD600 of 0.5–0.6. Cells were pelleted, washed once with 500 µl H2O and then resuspended in 1.2 ml Z-Buffer (60 mM Na2HPO4, 40 mM NaH2PO4, 10 mM KCl, 1 mM MgSO4, 20 mM 2-mercaptoethanol). Cells were lysed by sonification and cellular debris was removed by centrifugation. Protein content was determined using Roti®-Nanoquant. Samples were diluted in Z buffer to a final volume of 1 ml and incubated at 30 °C for 10 minutes. The reaction was started by addition of 200 µl ONPG solution (4 mg/ml in Z-Buffer) and stopped by addition of 500 µl 1 M Na2CO3 as soon as the first sample turned clearly yellow. Absorption was measured at 420 nm and Miller units (MU) were calculated.
Infection experiments
Experimental infections were carried out as described earlier54. Briefly, 3 × 105 J774.A1 mouse ascites macrophages (ATCC) were seeded into the wells of a 24 multi well plate and cultivated in 1 ml high glucose DMEM medium (4.5 g/l glucose, 110 mg/l sodium pyruvate, 584 mg/l L-glutamine) supplemented with 10% fetal calf serum (FCS) for one day at 37 °C in a 5% CO2 atmosphere. 5 × 104 bacteria from overnight cultures were resuspended in 1 ml of fresh DMEM without FCS and this inoculum was used to infect the J774 cells during an incubation step of 1 h at 37 °C. Next, the wells were washed once with PBS and all extracellular bacteria were killed during another 1 h incubation step in DMEM (without FCS) containing 40 µg/ml gentamicin. The wells were covered with fresh DMEM (without FCS) containing 10 µg/ml gentamicin after one more PBS wash step, and then incubated at 37 °C in a 5% CO2 atmosphere. Sampling was performed at various time points by lysing the cells in 1 ml of ice-cold PBS containing 0.1% Triton X-100. Serial dilutions were plated on BHI agar plates in order to count the recovered bacterial colonies.
Electron microscopy
Scanning electron microscopy and ultrathin section transmission electron microscopy were performed essentially as described earlier55.
Data Availability
RNA sequencing raw files are available at the NCBI Geo Server (https://www.ncbi.nlm.nih.gov/geo/) under study accession numbers: GSE129904 (ladR), GSE129909 (lstR) and GSE129910 (lltR).
References
Walker, S. J., Archer, P. & Banks, J. G. Growth of Listeria monocytogenes at refrigeration temperatures. Journal of Applied Bacteriology 68, 157–162, https://doi.org/10.1111/j.1365-2672.1990.tb02561.x (1990).
Schlech, W. F. III Foodborne listeriosis. Clinical infectious diseases: an official publication of the Infectious Diseases Society of America 31, 770–775, https://doi.org/10.1086/314008 (2000).
Charlier, C. et al. Clinical features and prognostic factors of listeriosis: the MONALISA national prospective cohort study. Lancet Infect Dis 17, 510–519, https://doi.org/10.1016/S1473-3099(16)30521-7 (2017).
Ivanek, R., Grohn, Y. T. & Wiedmann, M. Listeria monocytogenes in multiple habitats and host populations: review of available data for mathematical modeling. Foodborne Pathog Dis 3, 319–336, https://doi.org/10.1089/fpd.2006.3.319 (2006).
Bucur, F. I., Grigore-Gurgu, L., Crauwels, P., Riedel, C. U. & Nicolau, A. I. Resistance of Listeria monocytogenes to stress conditions encountered in food and food processing environments. Frontiers in microbiology 9, 2700, https://doi.org/10.3389/fmicb.2018.02700 (2018).
Rorvik, L. M. Listeria monocytogenes in the smoked salmon industry. Int J Food Microbiol 62, 183–190 (2000).
Borucki, M. K., Peppin, J. D., White, D., Loge, F. & Call, D. R. Variation in biofilm formation among strains of Listeria monocytogenes. Applied and Environmental Microbiology 69, 7336–7342, https://doi.org/10.1128/aem.69.12.7336-7342.2003 (2003).
Mereghetti, L., Quentin, R., Marquet-Van Der Mee, N. & Audurier, A. Low sensitivity of Listeria monocytogenes to quaternary ammonium compounds. Appl Environ Microbiol 66, 5083–5086 (2000).
Aase, B., Sundheim, G., Langsrud, S. & Rorvik, L. M. Occurrence of and a possible mechanism for resistance to a quaternary ammonium compound in Listeria monocytogenes. Int J Food Microbiol 62, 57–63 (2000).
Kovacevic, J. et al. Tolerance of Listeria monocytogenes to quaternary ammonium sanitizers is mediated by a novel efflux pump encoded by emrE. Appl Environ Microbiol 82, 939–953, https://doi.org/10.1128/AEM.03741-15 (2016).
Elhanafi, D., Dutta, V. & Kathariou, S. Genetic characterization of plasmid-associated benzalkonium chloride resistance determinants in a Listeria monocytogenes strain from the 1998-1999 outbreak. Appl Environ Microbiol 76, 8231–8238, https://doi.org/10.1128/AEM.02056-10 (2010).
Barthelmebs, L., Lecomte, B., Divies, C. & Cavin, J. F. Inducible metabolism of phenolic acids in Pediococcus pentosaceus is encoded by an autoregulated operon which involves a new class of negative transcriptional regulator. J Bacteriol 182, 6724–6731 (2000).
Gury, J., Barthelmebs, L., Tran, N. P., Divies, C. & Cavin, J. F. Cloning, deletion, and characterization of PadR, the transcriptional repressor of the phenolic acid decarboxylase-encoding padA gene of Lactobacillus plantarum. Appl Environ Microbiol 70, 2146–2153 (2004).
Madoori, P. K., Agustiandari, H., Driessen, A. J. & Thunnissen, A. M. Structure of the transcriptional regulator LmrR and its mechanism of multidrug recognition. EMBO J 28, 156–166, https://doi.org/10.1038/emboj.2008.263 (2009).
Agustiandari, H., Lubelski, J., van den Berg van Saparoea, H. B., Kuipers, O. P. & Driessen, A. J. LmrR is a transcriptional repressor of expression of the multidrug ABC transporter LmrCD in Lactococcus lactis. J Bacteriol 190, 759–763, https://doi.org/10.1128/JB.01151-07 (2008).
Agustiandari, H., Peeters, E., de Wit, J. G., Charlier, D. & Driessen, A. J. LmrR-mediated gene regulation of multidrug resistance in Lactococcus lactis. Microbiology 157, 1519–1530, https://doi.org/10.1099/mic.0.048025-0 (2011).
Takeuchi, K., Tokunaga, Y., Imai, M., Takahashi, H. & Shimada, I. Dynamic multidrug recognition by multidrug transcriptional repressor LmrR. Scientific reports 4, 6922, https://doi.org/10.1038/srep06922 (2014).
van der Berg, J. P., Madoori, P. K., Komarudin, A. G., Thunnissen, A. M. & Driessen, A. J. Binding of the lactococcal drug dependent transcriptional regulator LmrR to its ligands and responsive promoter regions. PLoS One 10, e0135467, https://doi.org/10.1371/journal.pone.0135467 (2015).
Lubelski, J. et al. LmrCD is a major multidrug resistance transporter in Lactococcus lactis. Mol Microbiol 61, 771–781, https://doi.org/10.1111/j.1365-2958.2006.05267.x (2006).
Kaval, K. G., Hahn, B., Tusamda, N., Albrecht, D. & Halbedel, S. The PadR-like transcriptional regulator LftR ensures efficient invasion of Listeria monocytogenes into human host cells. Frontiers in microbiology 6, 772, https://doi.org/10.3389/fmicb.2015.00772 (2015).
Hauf, S. et al. Aurantimycin resistance genes contribute to survival of Listeria monocytogenes during life in the environment. Mol Microbiol, accepted, https://doi.org/10.1111/mmi.14205 (2019).
Gräfe, U. et al. Aurantimycins, new depsipeptide antibiotics from Streptomyces aurantiacus IMET 43917. Production, isolation, structure elucidation, and biological activity. J Antibiot (Tokyo) 48, 119–125 (1995).
Huillet, E., Velge, P., Vallaeys, T. & Pardon, P. LadR, a new PadR-related transcriptional regulator from Listeria monocytogenes, negatively regulates the expression of the multidrug efflux pump MdrL. FEMS Microbiol Lett 254, 87–94, https://doi.org/10.1111/j.1574-6968.2005.00014.x (2006).
Jiang, X. et al. MdrL, a major facilitator superfamily efflux pump of Listeria monocytogenes involved in tolerance to benzalkonium chloride. Appl Microbiol Biotechnol, https://doi.org/10.1007/s00253-018-9551-y (2018).
Romanova, N. A., Wolffs, P. F., Brovko, L. Y. & Griffiths, M. W. Role of efflux pumps in adaptation and resistance of Listeria monocytogenes to benzalkonium chloride. Appl Environ Microbiol 72, 3498–3503, https://doi.org/10.1128/AEM.72.5.3498-3503.2006 (2006).
Zhang, C., Nietfeldt, J., Zhang, M. & Benson, A. K. Functional consequences of genome evolution in Listeria monocytogenes: the lmo0423 and lmo0422 genes encode SigmaC and LstR, a lineage II-specific heat shock system. J Bacteriol 187, 7243–7253, https://doi.org/10.1128/JB.187.21.7243-7253.2005 (2005).
Mata, M. T., Baquero, F. & Perez-Diaz, J. C. A multidrug efflux transporter in Listeria monocytogenes. FEMS Microbiol Lett 187, 185–188 (2000).
Toledo-Arana, A. et al. The Listeria transcriptional landscape from saprophytism to virulence. Nature 459, 950–956, https://doi.org/10.1038/nature08080 (2009).
Crimmins, G. T. et al. Listeria monocytogenes multidrug resistance transporters activate a cytosolic surveillance pathway of innate immunity. Proc Natl Acad Sci USA 105, 10191–10196, https://doi.org/10.1073/pnas.0804170105 (2008).
Woodward, J. J., Iavarone, A. T. & Portnoy, D. A. c-di-AMP secreted by intracellular Listeria monocytogenes activates a host type I interferon response. Science 328, 1703–1705, https://doi.org/10.1126/science.1189801 (2010).
Emami, K. et al. RodA as the missing glycosyltransferase in Bacillus subtilis and antibiotic discovery for the peptidoglycan polymerase pathway. Nat Microbiol 2, 16253, https://doi.org/10.1038/nmicrobiol.2016.253 (2017).
Meeske, A. J. et al. SEDS proteins are a widespread family of bacterial cell wall polymerases. Nature 537, 634–638, https://doi.org/10.1038/nature19331 (2016).
Chan, Y. C. et al. Contributions of two-component regulatory systems, alternative sigma factors, and negative regulators to Listeria monocytogenes cold adaptation and cold growth. J Food Prot 71, 420–425 (2008).
Mujahid, S., Orsi, R. H., Boor, K. J. & Wiedmann, M. Protein level identification of the Listeria monocytogenes sigma H, sigma L, and sigma C regulons. BMC Microbiol 13, 156, https://doi.org/10.1186/1471-2180-13-156 (2013).
Chaturongakul, S. et al. Transcriptomic and phenotypic analyses identify coregulated, overlapping regulons among PrfA, CtsR, HrcA, and the alternative sigma factors sigmaB, sigmaC, sigmaH, and sigmaL in Listeria monocytogenes. Appl Environ Microbiol 77, 187–200, https://doi.org/10.1128/AEM.00952-10 (2011).
Jordan, S., Junker, A., Helmann, J. D. & Mascher, T. Regulation of LiaRS-dependent gene expression in Bacillus subtilis: Identification of inhibitor proteins, regulator binding sites, and target genes of a conserved cell envelope stress-sensing two-component system. J Bacteriol 188, 5153–5166, https://doi.org/10.1128/JB.00310-06 (2006).
Renier, S., Micheau, P., Talon, R., Hebraud, M. & Desvaux, M. Subcellular localization of extracytoplasmic proteins in monoderm bacteria: rational secretomics-based strategy for genomic and proteomic analyses. PLoS One 7, e42982, https://doi.org/10.1371/journal.pone.0042982 (2012).
Marchler-Bauer, A. et al. CDD/SPARCLE: functional classification of proteins via subfamily domain architectures. Nucleic Acids Res 45, D200–D203, https://doi.org/10.1093/nar/gkw1129 (2017).
Rismondo, J., Halbedel, S. & Gründling, A. Cell shape and antibiotic resistance is maintained by the activity of multiple FtsW and RodA enzymes in Listeria monocytogenes. bioRxiv, 589911, https://doi.org/10.1101/589911 (2019).
Zhang, C. et al. Genome diversification in phylogenetic lineages I and II of Listeria monocytogenes: identification of segments unique to lineage II populations. J Bacteriol 185, 5573–5584 (2003).
Helmann, J. D. Bacillus subtilis extracytoplasmic function (ECF) sigma factors and defense of the cell envelope. Curr Opin Microbiol 30, 122–132, https://doi.org/10.1016/j.mib.2016.02.002 (2016).
Taguchi, A. et al. FtsW is a peptidoglycan polymerase that is functional only in complex with its cognate penicillin-binding protein. Nat Microbiol, https://doi.org/10.1038/s41564-018-0345-x (2019).
Tamura, T., Suzuki, H., Nishimura, Y., Mizoguchi, J. & Hirota, Y. On the process of cellular division in Escherichia coli: isolation and characterization of penicillin-binding proteins 1a, 1b, and 3. Proc Natl Acad Sci USA 77, 4499–4503 (1980).
Münch, D. & Sahl, H. G. Structural variations of the cell wall precursor lipid II in Gram-positive bacteria - Impact on binding and efficacy of antimicrobial peptides. Biochimica et biophysica acta 1848, 3062–3071, https://doi.org/10.1016/j.bbamem.2015.04.014 (2015).
Studer, P. et al. The Absence of a Mature Cell Wall Sacculus in Stable Listeria monocytogenes L-Form Cells Is Independent of Peptidoglycan Synthesis. PLoS One 11, e0154925, https://doi.org/10.1371/journal.pone.0154925 (2016).
Sambrook, J., Fritsch, E. F. & Maniatis, T. Molecular cloning: a laboratory manual. 2nd edn, (Cold Spring Harbor Laboratory Press, 1989).
Monk, I. R., Gahan, C. G. & Hill, C. Tools for functional postgenomic analysis of Listeria monocytogenes. Appl Environ Microbiol 74, 3921–3934, https://doi.org/10.1128/AEM.00314-08 (2008).
Zheng, L., Baumann, U. & Reymond, J. L. An efficient one-step site-directed and site-saturation mutagenesis protocol. Nucleic Acids Res 32, e115 (2004).
Arnaud, M., Chastanet, A. & Debarbouille, M. New vector for efficient allelic replacement in naturally nontransformable, low-GC-content, gram-positive bacteria. Appl Environ Microbiol 70, 6887–6891, https://doi.org/10.1128/AEM.70.11.6887-6891.2004 (2004).
van den Ent, F. & Löwe, J. RF cloning: a restriction-free method for inserting target genes into plasmids. J Biochem Biophys Methods 67, 67–74, https://doi.org/10.1016/j.jbbm.2005.12.008 (2006).
Gertz, S. et al. Regulation of sigmaB-dependent transcription of sigB and asp23 in two different Staphylococcus aureus strains. Mol Gen Genet 261, 558–566 (1999).
Kersey, P. J. et al. Ensembl Genomes 2018: an integrated omics infrastructure for non-vertebrate species. Nucleic Acids Res 46, D802–D808, https://doi.org/10.1093/nar/gkx1011 (2018).
Patro, R., Duggal, G., Love, M. I., Irizarry, R. A. & Kingsford, C. Salmon provides fast and bias-aware quantification of transcript expression. Nature methods 14, 417–419, https://doi.org/10.1038/nmeth.4197 (2017).
Halbedel, S. et al. A systematic proteomic analysis of Listeria monocytogenes house-keeping protein secretion systems. Molecular & cellular proteomics: MCP 13, 3063–3081, https://doi.org/10.1074/mcp.M114.041327 (2014).
Rismondo, J. et al. Discrete and overlapping functions of peptidoglycan synthases in growth, cell division and virulence of Listeria monocytogenes. Mol Microbiol 95, 332–351, https://doi.org/10.1111/mmi.12873 (2015).
Solovyev, V. V. Solovyev, A Salamov (2011) Automatic Annotation of Microbial Genomes and Metagenomic Sequences. In Metagenomics and its Applications in Agriculture, Biomedicine and Environmental Studies (Ed. Li, R. W.), Nova Science Publishers, p.61–78 (2011).
Acknowledgements
This work was supported by intramural grants of the Robert Koch Institute (to SvH) and a DFG grant (to SvH). We would like to thank Jeanine Rismondo and Angelika Gründling (Imperial College London) for sharing results prior to publication and for helpful discussions. Furthermore, we are grateful to Gudrun Holland and Tobias Hoffmann for excellent technical assistance.
Author information
Authors and Affiliations
Contributions
Sa. H. and L.M. performed the experiments. Sa. H., L.M., S.F. and Sv. H. analysed the data. Sa. H. and Sv. H. wrote the manuscript. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hauf, S., Möller, L., Fuchs, S. et al. PadR-type repressors controlling production of a non-canonical FtsW/RodA homologue and other trans-membrane proteins. Sci Rep 9, 10023 (2019). https://doi.org/10.1038/s41598-019-46347-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-46347-w
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.