Abstract
Long non-coding RNAs (lncRNAs) are known to play a major role in the epigenetic regulation of muscle development. Unfortunately there is little understanding of the mechanisms with which they regulate muscle growth in abalone. Therefore, we used RNA-seq to study the muscle transcriptomes of six Haliotis discus hannai specimens: three large (L_HD group) and three small (S_HD group). We identified 2463 lncRNAs in abalone muscle belonging to two subtypes: 160 anti-sense lncRNAs and 2303 intergenic lncRNAs (lincRNAs). In the L_HD group, we identified 204 significantly differentially expressed lncRNAs (55 upregulated and 149 downregulated), and 2268 significantly differentially expressed mRNAs (994 upregulated and 1274 downregulated), as compared to the S_HD group. The bioinformatics analysis indicated that lncRNAs were relate to cell growth, regulation of growth, MAPK signaling pathway, TGF-β signaling pathway, PI3K-Akt and insulin signaling pathway, which involved in regulating muscle growth. These findings contribute to understanding the possible regulatory mechanisms of muscle growth in Pacific abalone.
Similar content being viewed by others
Introduction
Muscle growth in livestock is very important, as it directly affects meat production. The regulatory mechanisms of muscle growth are complex, and are affected by genetics, nutrition, and the environment1. Of these, genetic factors, including those growth hormone (GH), insulin-like growth factors (IGFs), myogenic regulatory factors (MRFs), myostatin (Mstn), and paired box proteins (Paxs), are the most important1,2. However, studies of the role of non-coding RNA, particularly long noncoding RNA (lncRNA), in the regulation of muscle growth remain scarce.
LncRNAs are RNA molecules longer than 200 nucleotides (nt) that have little or no open reading frame (ORF)3. Compared with mRNAs, lncRNAs are marked by lower expression levels, less conservation, and more variable expression among tissues4,5. Many researches have shown that lncRNAs are relate to various biological processes including cancer, apoptosis, immunity, and development6,7,8. Several studies have also indicated that lncRNAs play a vital role in muscle growth9,10. For example, Lnc133 was highly expressed in the adductor muscle of Pinctada martensii and it could be involved in regulating the cell proliferation of adductor muscles by targeting pm-RhoA11. Most currently identified lncRNAs have been derived from mice and humans12,13,14. Several studies in chickens9, cattle15, pigs16, zebrafish17, and rainbow trout18 have enriched the datasets of animal lncRNA, but little is known about lncRNA in the abalone.
The Pacific abalone, is the most commonly cultivated abalone in China19. Here, we used Illumina HiSeqX sequencing to determine the lncRNA and mRNA expression profiles of two H. discus hannai phenotypes that differ with respect to muscle growth rate. We then used quantitative real-time polymerase chain reactions (qRT-PCR) to compare the expression levels of muscle growth-related genes between these phenotypes. These results increase our knowledge of the molecular mechanisms regulating muscle growth in the abalone.
Results
Identification of candidate lncRNAs
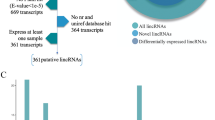
We generated 709,386,602 raw RNA-seq reads (NCBI accession no. SRP126378) from the adductor muscle samples of the three large (L_HD) and three small (S_HD) H. discus hannai specimens. The result of RNA quality was shown in Table 1. After discarding low-quality, adaptor, and poly-N sequences, 688,261,544 clean reads remained. We were able to map between 64.09% and 68.95% of the clean reads in each library to the H. discus hannai reference genome (Supplementary Table S1). Our coding potential analysis identified 2463 lncRNAs (Fig. 1): 2303 lincRNAs (93.5%) and 160 anti-sense lncRNAs (6.5%). We did not identify any intronic lncRNAs.
Genomic characterization of the candidate lncRNAs
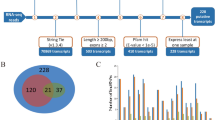
We identified 23,847 mRNAs and 2463 lncRNAs in the adductor muscle samples from the six H. discus hannai specimens. We found that the lncRNAs were less expressed than the mRNAs (Fig. 2a), and the lncRNAs had fewer exons than the mRNAs (Fig. 2b). In addition, in comparison to the mRNAs, most lncRNAs were shorter ORF length (Fig. 2c).
Differential expression (DE) cluster analysis
We obtained 204 lncRNAs (DE-lncRNAs) and 2268 mRNAs (DE-mRNAs) that were significantly differentially expressed between the L_HD and S_HD specimens (P < 0.05; Supplementary Tables S2 and 3). In the L_HD specimens, 55 DE-lncRNAs and 994 DE-mRNAs were upregulated compared to the S_HD specimens, while 149 DE-lncRNAs and 1274 DE-mRNAs were downregulated (Fig. 3a,b). Our heat maps also suggested that lncRNAs (Fig. 3c) and mRNAs (Fig. 3d) were significant expression difference (P < 0.05) between the two groups.
Volcano plots and heat maps of differentially expressed transcripts (P < 0.05). Expression of (a) lncRNAs and (b) mRNAs in large (L_HD) versus small (S_HD) specimens of abalone. Red and green dots indicate up- and down-regulated transcripts, respectively. Hierarchical clustering of differentially expressed (c) lncRNAs and (d) mRNAs. Red rectangles represent upregulated lncRNAs/mRNAs; blue rectangles represent downregulated lncRNAs/mRNAs.
Prediction of the lncRNA target genes
LncRNAs can act in cis to regulate the neighboring genes; or they may function in trans to regulate the expression of genes located in distant domains20. To better understand the functional roles of our identified lncRNAs, we forecasted the targets of lncRNAs. We identified 1727 lncRNAs acting in cis with 5512 mRNAs. Interestingly, several muscle development-related genes including ras homolog family member A (RhoA) and cell division cycle 42 (Cdc42), were targeted by the lncRNAs XLOC_042193 and XLOC_020807, indicating that these muscle growth genes may be cis-regulated by lncRNAs. We identified 327,782 interactions in trans between 2464 lncRNAs and 16,676 mRNAs. Similarly, we observed that several DE-lncRNAs (such as XLOC_031278, XLOC_019246, XLOC_046403, XLOC_021050) acted in trans on muscle growth-related genes (Table 2).
Bioinformatics analysis
Our GO analysis of the DE-target mRNAs regulated in cis by DE-lncRNAs identified 120 significantly terms (P < 0.05). These terms were primarily involved in growth regulation and in biosynthetic-related functions such as glycogen biosynthetic process, regulation of cell growth, insulin-like growth factor binding, and regulation of growth (Fig. 4a). We identified 322 GO terms significantly enriched across the DE-target mRNAs regulated in trans by DE-lncRNAs (P < 0.05). These GO terms encompassed various biological processes, including actin cytoskeleton organization, hexose metabolic process, and regulation of biological process (Fig. 4b).
Analysis of significant GO terms and KEGG pathways for the predicted differentially expressed target mRNAs of our DE-lncRNAs. Significant GO terms for genes (a) cis-regulated and (b) trans-regulated by lncRNAs in L_HD specimens, as compared to S_HD specimens. BP: biological process; MF: molecular function; CC: cellular_component. Significant KEGG pathways for genes (c) cis-regulated and (d) trans-regulated by lncRNAs in L_HD specimens, as compared to S_HD specimens (P < 0.05 is recommended).
The DE-target mRNAs of the DE-lncRNAs regulated in cis were significantly enriched in 82 KEGG pathways. Some of these signaling pathways were concerned with muscle growth, including the MAPK, the FoxO, and the PI3K-Akt signaling pathway (Fig. 4c). Our results therefore indicated that lncRNAs may function in cis on neighboring genes to influence muscle development. Our functional analysis also indicated that DE-target mRNAs in trans were significantly enriched in 103 KEGG pathways. Several of these signaling pathways were associated with muscle growth, including the MAPK, the TGF-β, and the insulin signaling pathway (Fig. 4d).
LncRNA-mRNA interaction network
Our lncRNA-mRNA interaction network results indicated that possible regulatory network interactions were linked to several signaling pathways, including the MAPK, the FoxO, the PI3K-Akt, and the TGF-β signaling pathway. Here, several DE-mRNAs and their corresponding DE-lncRNA regulators were constructed to assess their function with respect to abalone muscle growth (Fig. 5). We found that 59 lncRNAs interacted with five mRNAs in the MAPK signaling pathway (Fig. 5a), while 37 lncRNAs interacted with five mRNAs in the TGF-β signaling pathway (Fig. 5b).
LncRNA-mRNA interaction networks. (a) The MAPK signaling pathway, showing 59 lncRNAs interacting with 5 mRNAs. (b) The TGF-β signaling pathway, showing 37 lncRNAs interacting with 5 mRNAs. All interactions show gene expression in large specimens, as compared to small specimens. Green ovals: downregulated lncRNAs; red ovals: upregulated lncRNAs; green triangles: downregulated genes; red triangles: upregulated genes.
Specific expression of lncRNAs
We found 14 specific lncRNA expressions in the L_HD, particularly XLOC_007603, which has the lowest P value. Genes multiple EGF like domains 10 (Megf10) and bone morphogenetic protein 7 (Bmp7) were targeted by XLOC_007603. We also discovered nine specific lncRNA expressions in the S_HD samples, such as XLOC_004306. Growth hormone secretagogue receptor type 1 (Ghsr) and Actin, both related to growth, were targeted by XLOC_004306. These specific expressed lncRNAs perhaps play crucial roles in abalone muscle growth, although the underlying regulatory mechanisms require further study.
Validation of the transcripts expression by qRT-PCR
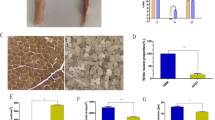
To validate our sequencing results, we selected three upregulated DE-mRNAs, three upregulated DE-lncRNAs, and four downregulated DE-lncRNAs to analyse the expression levels using qRT-PCR. (Fig. 6a). The expression patterns of these DE-lncRNAs and DE-mRNAs were accordance with the sequencing data, suggesting that our RNA-seq data were accurate. Our analysis of the tissue expression patterns of XLOC_033661 and growth differentiation factor 8 (Gdf8) suggested that these were ubiquitously expressed in all examined tissues (Fig. 6b,c).
Relative expression of lncRNAs and mRNAs, quantified with qRT-PCR. (a) Some lncRNAs and mRNAs were tested in the muscle of Haliotis discus hannai. (b) Expression of XLOC_033661 in the mantle, muscle, visceral mass, and gill. (c) Expression of Gdf8 in the mantle, muscle, visceral mass, and gill. Asterisks indicate statistically significant differences between large (L_HD) and small (S_HD) specimens: *P < 0.05; **P < 0.01. Different capital letters indicate significant differences among tissues (P < 0.01).
Discussion
Muscle growth is a complex life activity regulated by the coordinated action of many biological processes. Abalone with different body weights have different growth rates: larger abalones grow faster and smaller abalones grow slower21,22. To clarify the mechanisms underlying muscle growth in Pacific abalone, we used RNA-seq to investigate the discrepancy in mRNA and lncRNA expression patterns between larger and smaller abalone specimens from the same family.
As far as we know, this is the first study of lncRNA expression data in H. discus hannai. Here, we identified 2463 lncRNAs and 23,847 mRNAs. We found that the lncRNAs had fewer exons and were shorter than the mRNAs, consistent with previous studies18,23,24. The average number of exons (mean: 2.6) found in the lncRNAs of H. discus hannai was less than that of zebrafish (mean: 2.8 exons), humans (mean: 2.9 exons), and mice (mean: 3.7 exons)17,24. LncRNAs were also less expressed than the mRNAs, again consistent with previous studies.
LncRNAs act as either cis- or trans-regulatory elements, with either co-localized or co-expressed protein-coding genes as targets10. For example, Linc-MD1, influences muscle development by targeting MAML125. Here, we identified 204 DE-lncRNAs and 2268 DE-mRNAs between the L_HD group and the S_HD group. We also constructed interaction networks between the cis- and trans-acting DE-lncRNAs and their mRNA targets to estimate the function of DE-lncRNAs in the regulation of muscle growth. Some genes have been shown to be connection with muscle growth, including Gdf826,27, kruppel-like factor 5 (Klf5)28, tuberous sclerosis-1 (Tsc1)29, sirtuin 3 (Sirt3)30, myocyte enhancer factor 2 A (Mef2A)31, insulin like growth factor 2 receptor (Igf2R)32, RhoA33, Cdc4234, Megf1035, and myosin heavy chain (Myh)36. Gdf8 (also known as Mtsn) is an important member of the TGF-β superfamily, and functions as a negative regulator of skeletal muscle development and growth37. Our expression analyses suggested that Gdf8 was ubiquitously expressed in all tested tissues, consistent with previous studies27. We found that Gdf8 mRNA was most highly expressed in the muscle and visceral mass, indicating that Gdf8 may play important roles in these tissues. We noticed the highest levels of XLOC_033661 expression in the muscle, indicating that this lncRNA perhaps play a vital role in muscle growth. Mef2A is known to be highly expressed in skeletal muscle, suggesting that it is valuable for skeletal muscle myoblast differentiation38. LncRNA-uc.167 is antisense to the Mef2C gene, and influences P19 cell proliferation and differentiation by regulating Mef2C39. Therefore, we speculate that the lncRNAs XLOC_046403, XLOC_032049, XLOC_002646, XLOC_021050, and XLOC_014032 regulate the muscle growth in H. discus hannai by targeting Mef2A. Similarly, other lncRNAs might affect muscle growth by targeting specific genes.
The results of GO and KEGG pathway analyses could help us understand the mechanisms underlying abalone muscle growth. Moreover, our lncRNA-mRNA interaction network indicated that 59 lncRNAs interacted with 5 mRNAs in the MAPK signaling pathway, and 37 lncRNAs interacted with 5 mRNAs in the TGF-β signaling pathway (Fig. 5a,b). Association of DE-mRNAs and DE-lncRNAs with pathways relevant to growth may partly explain the regulation of muscle development. The MAPK signaling pathway, which includes the p38 MAPK, the extracellular regulated kinase 1 and 2 (ERK1/2), and the Jun NH2-terminal kinase (JNK) pathways, plays a vital role in muscle development40,41. The p38 MAPK though regulating the sequential activation of MRFs and their transcriptional coactivators to control skeletal muscle differentiation42. Mothers against decapentaplegic homolog 3 (Smad3) acts downstream of TGF-β to repress the bHLH domain of MyoD, and thus control myoblast differentiation43. TGF-β/Smad3 stimulated smooth muscle cell (SMC) proliferation is controlled by the PI3K/Akt signaling pathway44. PI3K/Akt is one of the major pathways contributing to skeletal muscle differentiation45. Our results can elucidate key lncRNAs and provide leads to further understand the mechanisms of molluscan muscle growth.
In conclusion, we reported the first lncRNA expression profiles of H. discus hannai using Illumina HiSeqX sequencing technology and identified 2463 lncRNAs. We also found out DE-mRNAs and DE-lncRNAs in slow- and fast- growing specimens of H. discus hannai. We identified lncRNAs acting in cis and trans to target genes (mRNAs). Our bioinformatics analyses suggested that many DE-lncRNAs might influence the regulation of muscle growth in H. discus hannai by affecting target genes. All these findings may help to understand the biological mechanisms controlling muscle growth in the abalone. Nevertheless, the roles of lncRNAs and their target genes analyses need further experiental verification.
Materials and Methods
Experimental sample
A breeding population of H. discus hannai has produced pedigreed offspring; The six H. discus hannai abalones used in this research were obtained from Fuda Aquiculture in Jinjiang, Fujian province, China; all specimens were about 2 years old. Three of the samples were larger (“L_HD” group; mean weight, 95.1 ± 7.7 g; mean muscle weight, 45.5 ± 5.0 g), and three were smaller (“S_HD” group; mean weight, 16.5 ± 1.0 g; mean muscle weight, 7.3 ± 0.8 g). All six specimens of the adductor muscle, mantle, visceral mass, and gill were collected from each abalone, immediately snap-frozen in liquid nitrogen46.
The corresponding author declares that all the methods were approved and perform in agreement with the instructions of the Laboratory Animal Management and Ethics Committee of Xiamen University and that all experimental protocols about abalones were carried out in accordance with the Regulations for the Administration of Affairs Concerning Experimental Animals of Xiamen University. Moreover, all the researcher who perform the animal experiments are trained by attending specifc courses.
RNA isolation and Illumina deep sequencing
The total RNA was isolated from adductor muscle samples taken from each H. discus hannai specimen using TRIzol reagent (Invitrogen, Carlsbad, CA, USA). Then, we checked the purity of the total RNA and assessed its integrity. Approximately 3 µg RNA per sample was used to construct a complementary (cDNA) library. We used a TruSeq PE Cluster Kit v3-cBot-HS with the cBot Cluster Generation System (Illumina, San Diego, CA, USA) to cluster the index-coded sample. The libraries were sequenced on an Illumina HiseqX platform and 150 bp paired-end reads were generated. Raw data were cleaned with in-house Perl scripts. Specifically, our script removed low quality reads, those containing adapter sequences, and those containing poly-N sequences to generate clean reads. At the same time, our script also calculated the Q20, Q30, and GC content of the clean data.
Transcriptome assembly
We used previously generated reference genome and gene model annotation files for H. discus hannai (the files provided by Dr. Weiwei You, Xiamen University, Xiamen). The clean reads were mapped to the H. discus hannai reference genome using TopHat v2.0.947 with default parameters. The mapped reads were assembled with both Scripture (beta2)22 and Cufflinks v2.1.148,49.
Quantification of gene expression level
We calculated the fragments per kilobase (kb) per million reads (FPKMs) for both the lncRNAs and the coding genes using Cuffdiff v2.1.150. We considered transcripts or genes differentially expressed when expression levels were significantly different (adjusted P of <0.05) between the large and small specimens (L_HD and S_HD).
Identification of lncRNAs
We used CPC (0.9-r2)51 and Pfam-scan (v1.3)52 to screen for candidate lncRNAs. Only those transcripts without predicted coding potential were retained. Finally, we selected the candidate lncRNAs predicted by both CPC and Pfam-scan as final lncRNAs for further analyses.
To investigate transcript conservation, we computed phylogenetic models in the Phast (v1.3) package53. Then, we computed the conservation scores of lncRNAs and coding genes using phastCons.
Target gene prediction
LncRNAs acting in cis act on neighboring target genes54,55. To identify these, we searched mRNAs 10 k/100 k up- and down-stream of each lncRNA. LncRNAs acting in trans influence target genes at the expression level. We computed the Pearson’s correlation coefficients both the expression levels of mRNAs and lncRNAs with custom scripts (r > 0.95 or r < −0.95). The lncRNA-mRNA interaction networks of DE-lncRNAs and their corresponding DE-mRNAs were constructed using Cytoscape.
Functional enrichment analysis
To evaluated the functions of the DE-lncRNA, we analyzed GO (Gene Ontology) with the GOseq R package56. We also performed KEGG (http://www.genome.jp/kegg/) analysis on DE-target mRNAs of the DE-lncRNAs using the hypergeometric test in KOBAS57. We considered functions with P < 0.05 significantly enriched.
QRT-PCR
Several genes were chosen for qRT-PCR using gene-specific primers (Supplementary Table S4). Relative gene expression levels were quantified based on β-actin gene expression using the 2−∆∆CT method58.
Statistical analysis
All qRT-PCR data were presented as mean ± standard deviation (SD). The statistical significance was evaluated using SPSS 19.0.
References
Scanes, C. G., Harvey, S., Marsh, J. A. & King, D. B. Hormones and growth in poultry. Poult Sci 63, 2062–2074, https://doi.org/10.3382/ps.0632062 (1984).
Johnston, I. A. et al. Embryonic temperature affects muscle fibre recruitment in adult zebrafish: genome-wide changes in gene and microRNA expression associated with the transition from hyperplastic to hypertrophic growth phenotypes. J Exp Biol 212, 1781–1793, https://doi.org/10.1242/jeb.029918 (2009).
Liao, Q. et al. Large-scale prediction of long non-coding RNA functions in a coding-non-coding gene coexpression network. Nucleic Acids Res 39, 3864–3878, https://doi.org/10.1093/nar/gkq1348 (2011).
Guttman, M. et al. Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature 458, 223–227, https://doi.org/10.1038/nature07672 (2009).
Ponting, C. P., Oliver, P. L. & Reik, W. Evolution and functions of long noncoding RNAs. Cell 136, 629–641, https://doi.org/10.1016/j.cell.2009.02.006 (2009).
Pauli, A., Rinn, J. L. & Schier, A. F. Non-coding RNAs as regulators of embryogenesis. Nat Rev Genet 12, 136–149, https://doi.org/10.1038/nrg2904 (2011).
Batista, P. J. & Chang, H. Y. Long noncoding RNAs: cellular address codes in development and disease. Cell 152, 1298–1307, https://doi.org/10.1016/j.cell.2013.02.012 (2013).
Carpenter, S. et al. A long noncoding RNA mediates both activation and repression of immune response genes. Science 341, 789–792, https://doi.org/10.1126/science.1240925 (2013).
Li, T. et al. Identification of long non-protein coding RNAs in chicken skeletal muscle using next generation sequencing. Genomics 99, 292–298, https://doi.org/10.1016/j.ygeno.2012.02.003 (2012).
Li, Z. et al. Integrated Analysis of Long Non-coding RNAs (LncRNAs) and mRNA Expression profiles reveals the potential role of LncRNAs in skeletal muscle development of the chicken. Front Physiol 7, 687, https://doi.org/10.3389/fphys.2016.00687 (2017).
Zheng, Z. et al. Pm-miR-133 hosting in one potential lncRNA regulates RhoA expression in pearl oyster Pinctada martensii. Gene 591, 484–489, https://doi.org/10.1016/j.gene.2016.06.051 (2016).
Volders, P. J. et al. An update on LNCipedia: a database for annotated human lncRNA sequences. Nucleic Acids Res 43, 4363–4, https://doi.org/10.1093/nar/gkv295 (2015).
Quek, X. C. et al. lncRNAdbv2.0: expanding the reference database for functional long noncoding RNAs. Nucleic Acids Res 43, D168–73, https://doi.org/10.1093/nar/gku988 (2015).
Bu, D. et al. NONCODEv3.0: integrative annotation of long noncoding RNAs. Nucleic Acids Res 40, D210–5, https://doi.org/10.1093/nar/gkr1175 (2012).
Billerey, C. et al. Identification of large intergenic non-coding RNAs in bovine muscle using next-generation transcriptomic sequencing. BMC Genomics 15, 499, https://doi.org/10.1186/1471-2164-15-499 (2014).
Zhao, W. et al. Systematic identification and characterization of long intergenic non-coding RNAs in fetal porcine skeletal muscle development. Sci Rep 5, 8957, https://doi.org/10.1038/srep08957 (2015).
Pauli, A. et al. Systematic identification of long noncoding RNAs expressed during zebrafish embryogenesis. Genome Res 22, 577–591, https://doi.org/10.1101/gr.133009.111 (2012).
Wang, J. et al. Identification and Functional Prediction of Large Intergenic Noncoding RNAs (lincRNAs) in Rainbow Trout (Oncorhynchus mykiss). Mar Biotechnol 18, 271–82, https://doi.org/10.1007/s10126-016-9689-5 (2016).
Luo, X., Ke, C. H. & You, W. W. Estimates of Correlations for Shell Morphological Traits on Body Weight of Interspecific Hybrid Abalone (Haliotis discus hannai and Haliotis gigantea). J Shellfish Res 32, 115–118, https://doi.org/10.2983/035.032.0117 (2013).
Wang, K. C. & Chang, H. Y. Molecular Mechanisms of Long Noncoding RNAs. Mol Cell 43, 904–14, https://doi.org/10.1016/j.molcel.2011.08.018 (2011).
Miranda, D. V., Portilla, M. D. & Escárate, C. G. Characterization of the growth-related transcriptome in California red abalone (Haliotis rufescens) through RNA-Seq analysis. Mar Genom 24, 199–202, https://doi.org/10.1016/j.margen.2015.05.009 (2015).
Choi, M. J., Kim, G. D., Kim, J. M. & Lim, H. K. Differentially-Expressed Genes Associated with Faster Growth of the Pacific Abalone, Haliotis discus hannai. Int J Mol Sci 16, 27520–27534, https://doi.org/10.3390/ijms161126042 (2015).
Guttman, M. et al. Ab initio reconstruction of cell type-specific transcriptomes in mouse reveals the conserved multi-exonic structure of lincRNAs. Nat Biotechnol 28, 503–510, https://doi.org/10.1038/nbt.1633 (2010).
Cabili, M. N. et al. Integrative annotation of human large intergenic noncoding RNAs reveals global properties and specific subclasses. Genes Dev 25, 1915–27, https://doi.org/10.1101/gad.17446611(2015).
Cesana, M. et al. A Long Noncoding RNA Controls Muscle Differentiation by Functioning as a Competing Endogenous RNA. Cell 147, 358–69, https://doi.org/10.1016/j.cell.2011.09.028 (2011).
Carpio, Y. et al. Regulation of body mass growth through activin type IIB receptor in teleost fish. Gen Comp Endocrinol 160, 158–167, https://doi.org/10.1016/j.ygcen.2008.11.009 (2009).
Naipil, C. C., Muñoz, V. V., Valdés, J. A., Molina, A. & Escárate, C. G. RNA interference in Haliotis rufescens myostatin evidences upregulation of insulin signaling pathway. Agri Gene 1, 93–99, https://doi.org/10.1016/j.aggene.2016.07.004 (2016).
Hayashi, S., Manabe, I., Suzuki, Y., Relaix, F. & Oishi, Y. Klf5 regulates muscle differentiation by directly targeting muscle-specific genes in cooperation with MyoD in mice. eLife 5, e17462, https://doi.org/10.7554/eLife.17462 (2016).
Bentzinger, C. F. et al. Differential response of skeletal muscles to mTORC1 signaling during atrophy and hypertrophy. Skeletal Muscle 3, 6, https://doi.org/10.1186/2044-5040-3-6 (2013).
Jing, E. et al. Sirtuin-3 (Sirt3) regulates skeletal muscle metabolism and insulin signaling via altered mitochondrial oxidation and reactive oxygen species production. PNAS 108, 14608–14613, https://doi.org/10.1073/pnas.1111308108 (2011).
Black, B. L. & Olson, E. N. Transcriptional control of muscle development by myocyte enhancer factor-2 (MEF2) proteins. Annu Rev Cell Dev Biol 14, 167–196, https://doi.org/10.1146/annurev.cellbio.14.1.167 (1998).
Elmagd, M. A., Aboalela, H. G., Elnahas, A., Saleh, A. A. & Mansour, A. A. Effects of a novel SNP of IGF2R gene on growth traits and expression rate of IGF2R and IGF2 genes in gluteus medius muscle of Egyptian buffalo. Gene 540, 133–139, https://doi.org/10.1016/j.gene.2014.02.059 (2014).
Carnac, G. et al. RhoA GTPase and Serum Response Factor Control Selectively the Expression of MyoD without Affecting Myf5 in Mouse Myoblasts. Molecular Biology of the Cell 9, 1891–1902, https://doi.org/10.1091/mbc.9.7.1891 (1998).
Meriane, M. et al. Critical Activities of Rac1 and Cdc42Hs in Skeletal Myogenesis: Antagonistic Effects of JNK and p38 Pathways. Molecular Biology of the Cell 11, 2513–2528, https://doi.org/10.1091/mbc.11.8.2513 (2000).
Park, S. Y., Yun, Y., Kim, M. J. & Kim, I. S. Myogenin is a positive regulator of MEGF10 expression in skeletal muscle. Biochem Biophys Res Commun 450, 1631–1637, https://doi.org/10.1016/j.bbrc.2014.07.061 (2014).
Hevrøy, E. M. et al. Myosin heavy chain mRNA expression correlates higher with muscle protein accretion than growth in Atlantic salmon, Salmo salar. Aquaculture 252, 453–461, https://doi.org/10.1016/j.aquaculture.2005.07.003 (2006).
McFarlane, C. et al. Human myostatin negatively regulates human myoblast growth and differentiation. Am J Physiol Cell Physiol 301, 195–203, https://doi.org/10.1152/ajpcell.00012.2011 (2011).
Seok, H. Y. et al. miR-155 Inhibits Expression of the MEF2A Protein to Repress Skeletal Muscle Differentiation. Journal of Biological Chemistry 286, 35339–35346, https://doi.org/10.1074/jbc.M111.273276 (2011).
Song, G. et al. LncRNA-uc.167 influences cell proliferation, apoptosis and differentiation of P19 cells by regulating Mef2c. Gene 590, 97–108, https://doi.org/10.1016/j.gene.2016.06.006 (2016).
Kim, H. J. & Lee, W. J. Ligand-independent activation of the androgen receptor by insulin-like growth factor-I and the role of the MAPK pathway in skeletal muscle cells. Mol Cells 28, 589–93, https://doi.org/10.1007/s10059-009-0167-z (2009).
Keren, A., Tamir, Y. & Bengal, E. The p38 MAPK signaling pathway: a major regulator of skeletal muscle development. Mol Cell Endocrinol 252, 224–30, https://doi.org/10.1016/j.mce.2006.03.017 (2006).
Lluís, F., Perdiguero, E., Nebreda, A. R. & Muñoz-Cánoves, P. Regulation of skeletal muscle gene expression by p38 MAP kinases. Trends Cell Biol 16, 36–44, https://doi.org/10.1016/j.tcb.2005.11.002 (2006).
Liu, D., Black, B. L. & Derynck, R. TGF-β inhibits muscle differentiation through functional repression of myogenic transcription factors by Smad3. Genes & Development 15, 2950–66, https://doi.org/10.1101/gad.925901 (2001).
Suwanabol, P. A. et al. TGF-β and Smad3 modulate PI3K/Akt signaling pathway in vascular smooth muscle cells. Am J Physiol Heart Circ Physiol 302, H2211–9, https://doi.org/10.1152/ajpheart.00966.2011 (2012).
Kaliman, P., Vinals, F., Testar, X., Palacin, M. & Zorzano, A. Phosphatidylinositol 3-kinase inhibitors block differentiation of skeletal muscle cells. J Biol Chem 271, 19146–51, https://doi.org/10.1074/jbc.271.32.19146 (1996).
Huang, J. F., You, W. W., Luo, X. & Ke, C. H. iTRAQ-Based Identification of Proteins Related to Muscle Growth in the Pacific Abalone, Haliotis discus hannai. Int J Mol Sci 18, 2237, https://doi.org/10.3390/ijms18112237 (2017).
Trapnell, C., Pachter, L. & Salzberg, S. L. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25, 1105–11, https://doi.org/10.1093/bioinformatics/btp120 (2009).
Trapnell, C. et al. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat Biotechnol 28, 511–515, https://doi.org/10.1038/nbt.1621 (2010).
Trapnell, C. et al. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat Protoc 7, 562–78, https://doi.org/10.1038/nprot.2012.016 (2012).
Flegel, C., Manteniotis, S., Osthold, S., Hatt, H. & Gisselmann, G. Expression profile of ectopic olfactory receptors determined by deep sequencing. PLoS One 8, e55368, https://doi.org/10.1371/journal.pone.0055368 (2013).
Kong, L. et al. CPC: assess the protein-coding potential of transcripts using sequence features and support vector machine. Nucleic Acids Res 35, W345–9, https://doi.org/10.1093/nar/gkm391 (2007).
Bateman, A. et al. The Pfam Protein Families Database. Nucleic Acids Res 28, 263–6 (2000).
Siepel, A. et al. Evolutionarily conserved elements in vertebrate, insect, worm, and yeast genomes. Genome Res 15, 1034–50, https://doi.org/10.1101/gr.3715005 (2005).
Gomez, J. A. et al. The NeST Long ncRNA Controls Microbial Susceptibility and Epigenetic Activation of the Interferon-gamma Locus. Cell 152, 743–54, https://doi.org/10.1016/j.cell.2013.01.015 (2013).
Lai, F. et al. Activating RNAs associate with Mediator to enhance chromatin architecture and transcription. Nature 494, 497–501, https://doi.org/10.1038/nature11884 (2013).
Young, M. D., Wakefield, M. J., Smyth, G. K. & Oshlack, A. Gene ontology analysis for RNA-seq: accounting for selection bias. Genome Biol 11, R14, https://doi.org/10.1186/gb-2010-11-2-r14 (2010).
Mao, X., Cai, T., Olyarchuk, J. G. & Wei, L. Automated genome annotation and pathway identification using the KEGG Orthology (KO) as a controlled vocabulary. Bioinformatics 21, 3787–3793, https://doi.org/10.1093/bioinformatics/bti430 (2005).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−∆∆CT method. Methods 25, 402–8, https://doi.org/10.1006/meth.2001.1262 (2001).
Acknowledgements
We thank all contributors of the present study. This work was supported by grants from National Natural Science Foundation of China (No. U1605213; 31472277); Earmarked Fund for Modern Agro-industry Technology Research System (No. CARS-49); Key S & T Program of Fujian Province (No. 2016N5012 and 2016NZ0001) and Shandong Province (No. 2016GGH4513).
Author information
Authors and Affiliations
Contributions
J.F.H. conceived of this research, performed data analysis, conducted qRT-PCR validation, and wrote the manuscript; X.L., L.T.Z., Z.K.H. and M.Q.H. participated in the animal experiments, statistical analysis, and surgical processes; W.W.Y. and C.H.K. revised the manuscript. All authors reviewed the final manuscript.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
41598_2018_35202_MOESM1_ESM.pdf
Expression profiling of lncRNAs and mRNAs reveals regulation of muscle growth in the Pacific abalone, Haliotis discus hannai Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Huang, J., Luo, X., Zeng, L. et al. Expression profiling of lncRNAs and mRNAs reveals regulation of muscle growth in the Pacific abalone, Haliotis discus hannai. Sci Rep 8, 16839 (2018). https://doi.org/10.1038/s41598-018-35202-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-35202-z
Keywords
This article is cited by
-
Transcriptome analysis reveals core lncRNA-mRNA networks regulating melanization and biomineralization in Patinopecten yessoensis shell-infested by Polydora
BMC Genomics (2023)
-
Differentiation and Maturation of Muscle and Fat Cells in Cultivated Seafood: Lessons from Developmental Biology
Marine Biotechnology (2023)
-
Transcriptome sequencing reveals the differentially expressed lncRNAs and mRNAs in response to cold acclimation and cold stress in Pomacea canaliculata
BMC Genomics (2022)
-
Long non-coding RNAs in the physiology of aquaculture animals: a perspective update
Reviews in Fish Biology and Fisheries (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.