Abstract
In cerebellar Purkinje cells (PCs) type-1 metabotropic glutamate (mGlu1) receptors play a key role in motor learning and drive the refinement of synaptic innervation during postnatal development. The cognate mGlu5 receptor is absent in mature PCs and shows low expression levels in the adult cerebellar cortex. Here we found that mGlu5 receptors were heavily expressed by PCs in the early postnatal life, when mGlu1α receptors were barely detectable. The developmental decline of mGlu5 receptors coincided with the appearance of mGlu1α receptors in PCs, and both processes were associated with specular changes in CpG methylation in the corresponding gene promoters. It was the mGlu1 receptor that drove the elimination of mGlu5 receptors from PCs, as shown by data obtained with conditional mGlu1α receptor knockout mice and with targeted pharmacological treatments during critical developmental time windows. The suppressing activity of mGlu1 receptors on mGlu5 receptor was maintained in mature PCs, suggesting that expression of mGlu1α and mGlu5 receptors is mutually exclusive in PCs. These findings add complexity to the the finely tuned mechanisms that regulate PC biology during development and in the adult life and lay the groundwork for an in-depth analysis of the role played by mGlu5 receptors in PC maturation.
Similar content being viewed by others
Introduction
Group-I mGlu receptors include mGlu1 and mGlu5 receptor subtypes, which are both coupled to Gq/11 and are predominantly found in the peripheral annulus of postsynaptic densities, where they are linked to Homer proteins and other scaffolding and effector proteins1,2. In spite of these similarities, mGlu1 and mGlu5 receptors differ in their anatomical distribution, with mGlu5 receptors showing a widespread expression pattern in the brain and spinal cord, and mGlu1 receptors being restricted to the olfactory bulb, thalamus, hippocampal dentate gyrus, and cerebellum3,4. In the cerebellum, the mGlu1 receptor is predominantly expressed in PCs, where it plays a crucial role in mechanisms of developmental plasticity and in the control of motor learning and motor coordination5,6,7. Activation of mGlu1 receptors in PC dendrites by the glutamate released from parallel fibers drives the elimination of suprannumerary climbing fibers between the second and third week of postnatal life through a mechanism that involves stimulation of polyphosphoinositide (PI) hydrolysis, activation of protein kinase C (PKC)-γ, and induction of semaphorine-7A8,9. The mGlu1 receptor is also required for the elimination of parallel fiber synapses from dendritic portions in which parallel and climbing fibers overlap, leading to the segregation of climbing and parallel fiber synaptic territories8,10. Genetic deletion of mGlu1 receptors causes a severe impairment of long-term depression (LTD) at parallel fiber-PC synapses associated with a profound defect in the conditioned eyeblink reflex and motor coordination. All these defects are rescued by selective re-introduction of mGlu1α receptors in PCs11,12. Interestingly, abnormalities in the expression/activity of mGlu1 receptors or downstream signaling molecules in PCs have been found in genetic mouse models of type-1, -2 -3, -5, and -14 spinocerebellar ataxias13,14,15,16, in ataxic moonwalker mutant mice17, in mice subjected to experimental autoimmune encephalomyelitis, and in autoptic samples from individuals affected by multiple sclerosis18. Neutralizing autoantibodies directed against mGlu1 receptors have been detected in patients with autoimmune ataxia19,20,21. All these findings suggest that mGlu1 receptors might be targeted by therapeutic intervention in cerebellar disorders7.
As opposed to mGlu1 receptors, mGlu5 receptors are virtually absent in mature cerebellar PCs, and show low expression levels in the adult cerebellar cortex, being mainly localized in Lugaro and Golgi cells22,23. This dampened the interest for the study of mGlu5 receptors in the cerebellum, although changes in cerebellar mGlu5 receptor expression were found in autoptic tissues from individuals affected by psychiatric disorders, autism, and Fragile X-associated tremor/ataxia syndrome24,25,26,27. We were intrigued by the finding that mGlu5 receptor protein levels in the rat cerebellum are higher in the early postnatal life than in the adult life, as opposed to mGlu1α receptor protein levels, which are more abundant in the adulthood than at PND928. This opposite developmental pattern of expression of mGlu1α and mGlu5 receptors was characteristic of the cerebellum and was not observed in other brain regions including the hippocampus, corpus striatum, cerebral cortex, hypothalamus, and olfactory bulb28.
Where precisely the mGlu5 receptor is expressed in the developing cerebellum is unknown, at present. No evidence exists to our knowledge that developing PCs express mGlu5 receptors, although mGlu5 receptor knockdown in the early postnatal life causes a severe impairment in PC maturation29.
We now report that mGlu5 receptors are highly expressed by cerebellar PCs in the first 12 days of postnatal life, and that the developmental decline in mGlu5 receptor expression coincides with the appearance and up-regulation of mGlu1α receptors. In addition, we demonstrate that it is the mGlu1 receptor that down-regulates the expression of mGlu5 receptors during the development of PCs and maintains its suppressing activity in the adult life.
Results
Complementary expression of mGlu1α and mGlu5 receptors in developing cerebellar PCs
We examined mGlu1α receptor expression in cerebellar tissue from mice at 3, 7, 9, 12, 16, and 18 PNDs by combining immunoblot analysis and immunofluorescent staining. mGlu1α receptor protein levels in the cerebellum were very low in the first twelve PND and increased substantially from PND12 to PND16, remaining high at later stages of development (Fig. 1A). The transcript of mGlu1 receptors increased by >2 fold from PND9 to PND18 (Fig. 1B), and this was associated with a trend to a reduction in Grm1 promoter methylation (Fig. 1C). Confocal analysis showed that expression of mGlu1α receptors was faint in calbindin-positive PCs at PND9 and increased substantially at PND16 (Fig. 1D).
Developmental pattern of expression of mGlu1 receptors in the mouse cerebellar PCs. mGlu1α protein levels in the mouse cerebellum at different PNDs are shown in (A). Densitometric values are means ± S.E.M of 4–5 mice/group. *p < 0.001 (One-Way ANOVA + Fisher’s LSD; F(5,20) = 21.8) vs. values obtained in mice at PND18. Grm1 mRNA levels and Grm1 promoter methylation in the cerebellum at PND9 and PND18 are shown in (B) and (C), respectively. Values are means ± S.E.M. of 5 mice/group. *p < 0.001 vs. values at PND9 (Student’s t test; t(7) = −7.96). Confocal microscopy analysis of mGlu1α receptors (green) and calbindin (red) in the cerebellar cortex is shown in (D). Note the faint mGlu1α immunoreactivity in PCs at PND9. Arrowheads indicate mGlu1α receptor immunoreactivity in PCs.
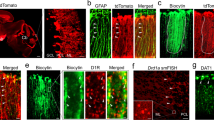
Interestingly, the mGlu5 receptor showed a complementary pattern of expression in the developing cerebellum, with mGlu5 receptor protein levels being high between PND9 and PND12 and decreasing afterwards (Fig. 2A). The transcript of mGlu5 receptors was dramatically reduced from PND9 to PND18 (Fig. 2B), and this was associated with a significant increase in Grm5 gene promoter methylation (Fig. 2C). Confocal microscopy analysis of mGlu5 receptors was performed in two separate experiments at different time points after birth. Developing PCs labelled for carbonic anhydrase-8 (Fig. 2D, upper panel) or calbindin (Fig. 2D, lower panel) expressed mGlu5 receptors at PND7, 9, and 12, but were no longer decorated with mGlu5 receptor antibodies after PND14 (Fig. 2D). After PND14 weak labelling for mGlu5 receptors remained in carbonic anhydrase-8 negative dendritic elements of putative interneurons.
Expression of mGlu5 receptors in cerebellar PCs early after birth. Western blot analysis of mGlu5 receptors in the mouse cerebellum at different PNDs is shown in (A). Densitometric values are means ± S.E.M. of 2–3 mice/group. p = 0.008 vs. values at PND9 (#) or PND12 (*) (One-Way ANOVA + Fisher’s LSD; F(4,9) = 7.01). The β-actin was taken from the same blot but with a lower exposure time. Grm5 mRNA levels and Grm5 promoter methylation in the cerebellum at PND9 and PND18 are shown in (B) and (C), respectively. Values are means ± S.E.M. of 4 mice/group. *p = 0.04 vs. values at PND9 (Student’s t test; t(6) = 2.61), *p = 0.006 vs. values at PND9 (Student’s t test; t(6) = −4.11), for Grm5 mRNA levels and Grm5 promoter methylation, respectively. Confocal microscopy images of mGlu5 receptor immunoreactivity in PCs at different PNDs are shown in D. Images were obtained from two different experiments performed in different laboratories. In the upper panel mGlu5 receptors are in green and type-8 carbonic anhydrase (a marker of PCs) is in red. In the lower panel, mGlu5 receptors are in green and calbindin (another marker of PCs) is in red. Note the disappearance of mGlu5 receptors in PCs after PND12. * indicates PCs; arrowheads indicate mGlu5 receptor immunoreactivity in PCs.
Developmental switch from mGlu1 and mGlu5 receptor signaling in the postnatal cerebellum
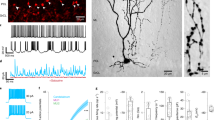
We used cerebellar slices challenged with the mGlu1/5 receptor agonist, DHPG, in the presence of mGlu1 or mGlu5 receptor blockers for the analysis of receptor-stimulated PI hydrolysis. As expected, maximally effective concentrations of DHPG (100 μM) largely increased [3H]inositolmonophosphate (InsP) formation in PND3-12 cerebellar slices. Stimulation of PI hydrolysis decreased at PND16 and became very low at PND21 (Fig. 3A). In the first 12 days of postnatal life, DHPG-stimulated PI hydrolysis was highly sensitive to the mGlu5 receptor negative allosteric modulator (NAM), MPEP, whereas only the mGlu1 receptor NAM, JNJ16259685, was able to antagonize the action of DHPG at PND16 (Fig. 3A). To localize mGlu1 or mGlu5 receptor signaling at cellular level, we assessed intracellular Ca2+ release in PCs from cerebellar slices prepared from PND9, PND17, and PND20. At PND9, DHPG-stimulated Ca2+ release was abrogated by MPEP (Fig. 3B). In contrast, the action of DHPG was antagonized by JNJ16259685, but not by MPEP, at PND17 and PND20 (Fig. 3B).
Developmental pattern of mGlu1 and mGlu5 receptor signaling in the cerebellum and cerebellar PCs. DHPG-stimulated PI hydrolysis in cerebellar slices at different PNDs incubated in the absence or presence of MPEP or JNJ16259685 is shown in (A). Values are means ± S.E.M. of 3–5 determinations. p < 0.001 vs. the respective basal values (*), vs. the respective values obtained with DHPG alone (#), or vs. the respective values obtained with DHPG in combination with MPEP ($) (One-Way ANOVA + Fisher’s LSD; PND3: F(3,8) = 16.74; PND9: F(3,13) = 59.93; PND12: F(3,12) = 37.07; PND16: F(3,12) = 15.97; PND21: F(3,12) = 3.79). DHPG-stimulated intracellular Ca2+ release in cerebellar PCs at different PNDs is shown in (B). Note that MPEP antagonized the action of DHPG at PND9 but was ineffective at PND20. In contrast, the action of DHPG was antagonized by JNJ16259685 at PND17.
mGlu1 receptors drive the developmental decline of mGlu5 receptors in PCs
To examine whether the decline of mGlu5 receptors in PCs was causally related to the appearance of mGlu1α receptors during postnatal development, we used pharmacological and genetic approaches. In a first set of experiments, we treated mice daily with the mGlu1 receptor NAM, JNJ16259685 (2.5 mg/kg, i.p.) during a developmental period that corresponds to the appearance of mGlu1 receptors, i.e., from PND9 to PND16. Pharmacological blockade of mGlu1 receptors during this time window prevented the decline of mGlu5 receptor protein, which was still expressed in PCs at PND16 (Fig. 4A,B). Grm5 gene promoter methylation was reduced in the cerebellum after treatment with JNJ16259685 (Fig. 4C). In addition, MPEP retained the activity of antagonizing DHPG-stimulated PI hydrolysis in cerebellar slices after in vivo treatment with JNJ16259685 (Fig. 4D).
Pharmacological blockade of mGlu1 receptors delayed the developmental decline in mGlu5 receptors in cerebellar PCs. Mice were treated daily with JNJ16259685 (2.5 mg/kg, i.p.) or its vehicle from PND9 to PND16. All measurements were carried out at the end of treatments. Western blot analysis of mGlu5 receptors in the cerebellum is shown in (A). Densitometric values are means ± S.E.M. *p = 0.041, vs. mice treated with vehicle (Student’s t test; t(5) = −2.74). Confocal microscopy analysis of mGlu5 receptors (green) is shown in (B). Note that mGlu5 receptor immunoreactivity is still present in PCs at PND16 only in mice treated with JNJ16259685. Grm5 promoter methylation in mice treated with vehicle or JNJ16259685 is shown in (C), where values are means ± S.E.M. of 3 mice/group. *p = 0.0076 vs. values obtained in mice treated with vehicle (Student’s t test; t(4) = 4.98). DHPG-stimulated PI hydrolysis in cerebellar slices from mice treated with vehicle or JNJ169259685 is shown in (D). Note that MPEP maintained its antagonistic activity in slices from PND16 mice treated with JNJ16259685. Values are means ± S.E.M. of 4 mice/group. (*) vs. the respective basal values or (#) vs. the respective values obtained with DHPG alone (Two-Way ANOVA + Fisher’s LSD; mouse treatment: F(1,24) = 7.13, p = 0.013; slice treatment: F(3,24) = 59.42, p < 0.001; mouse treatment x slice treatment: F(3,24) = 6.05, p = 0.003). Arrowheads indicate mGlu5 receptor immunoreactivity in PCs.
Alternatively, we treated mice with the selective mGlu1 receptor positive allosteric modulator (PAM), Ro0711401 (10 mg/kg, s.c., daily), from PND7 (for Western blot analysis) or PND10 (for functional analysis) to PND12, when expression of mGlu1 receptors in PCs is still low. This treatment accelerated the drop in mGlu5 receptor protein in the cerebellum, which was already reduced at PND12 (Fig. 5A). When DHPG-stimulated PI hydrolysis was monitored in cerebellar slices, MPEP lost its antagonistic activity at PND12 after systemic treatment with Ro0711401 (Fig. 5B). These data suggested that endogenous activation of mGlu1 receptors was responsible for the developmental drop of mGlu5 receptors in PCs.
Pharmacological enhancement of mGlu1 receptors accelerates the developmental decline of mGlu5 receptors in the cerebellum. Mice were treated with Ro0711401 (10 mg/kg, s.c., daily) or its vehicle from PND7 to PND12 in (A) or from PND10 to PND12 in (B). Measurements were carried out at the end of the treatment. Western blot analysis of mGlu5 receptors in the cerebellum is shown in (A). Densitometric values are means ± S.E.M. of 4 mice/group. *p = 0.042 vs. values obtained in mice treated with vehicle (Student’s t test; t(6) = 2.57). DHPG-stimulated PI hydrolysis in cerebellar slices prepared from PND12 mice treated with vehicle or Ro0711401 is shown in (B). Note that MPEP lost its antagonistic activity in slices from mice treated with Ro1711401. Values are means ± S.E.M. of 4 mice/group. (*) vs. the respective basal values or vs. (#) or vs. the respective values obtained with DHPG alone. (Two-Way ANOVA + Fisher’s LSD; mouse treatment: F(1,24) = 38.46, p < 0.001; slice treatment: F(3,24) = 78.76, p < 0.001; mouse treatment x slice treatment: F(3,24) = 20.59, p < 0.001).
To further demonstrate the developmental link between mGlu1 and mGlu5 receptors in PCs, we used conditional mGlu1 receptor knockout (cKO) mice. In these mice, exposure to doxycyclin (added to the drinking water of lactating mothers) from PND0 to PND14 suppressed the expression of mGlu1 receptors in PCs of the offspring (Fig. 6A). Doxycylin-induced knockdown of mGlu1 receptors was associated with a large increase in mGlu5 receptor expression, which was still prominent in the PCs at PND16 (Fig. 6A,B).
mGlu1 receptors drive the developmental decline of mGlu5 receptors in PCs. Conditional mGlu1α receptor knockout mice (cKO mice) were treated with doxycycline (200 μg/ml in drinking water between PND0 to PND14). Western blot analysis of mGlu1α and mGlu5 receptors in the cerebellum at PND16 is shown in (A). Densitometric values are means ± S.E.M. of 4–5 mice/group. *p = 0.017 vs. mGlu1α cKO mice not treated with Doxycycline (Dox−) (Student’s t test; t(7) = −3.10). Confocal microscopy analysis of mGlu5 receptors (green) and calbindin (red) is shown in (B). Arrowheads indicate mGlu5 receptor immunoreactivity in PCs.
mGlu1 receptors retain the suppressive activity on mGlu5 receptor expression in adult PCs
mGlu1α receptor cKO mice were also used to examine whether mGlu1 receptors could maintain the mGlu5 receptor suppressed in PCs also in the life. Chronic treatment with doxycyclin (200 μg/ml in the drinking water) from PND24 to PND76 caused the expected suppression of mGlu1α receptor protein associated with a significant increase in mGlu5 receptor protein levels in the cerebellum (Fig. 7A). Remarkably, cerebellar PCs were highly decorated with mGlu5 receptors after chronic mGlu1α receptor knockdown (Fig. 7B).
mGlu1 receptors retain the suppressive activity on mGlu5 receptor expression in adult PCs. Conditional mGlu1α receptor knockout mice (cKO mice) were treated with doxycycline (200 μg/ml in drinking water between PND24 to PND76). Western blot analysis of mGlu1α and mGlu5 receptors in the cerebellum is shown in (A). Densitometric values are means ± S.E.M. of 3–4 mice per group. *p = 0.026 vs. mGlu1α cKO mice not treated with Doxycycline (Dox−) (Student’s t test; t(5) = −3.12). Confocal microscopy analysis of mGlu5 receptors (green) and calbindin (red) is shown in (B). Arrowheads indicate mGlu5 receptor immunoreactivity in PCs.
Discussion
Our findings demonstrate for the first time that mGlu5 receptors are physiologically expressed by cerebellar PCs in the early stages of postnatal development, and largely mediate group-I mGlu receptor-stimulated PI hydrolysis in the first two weeks of postnatal life. Interestingly, the developmental pattern of expression of mGlu1α and mGlu5 receptors in PCs was specular, with mGlu1α receptors entirely replacing mGlu5 receptors during the first two weeks after birth. Gene methylation analysis demonstrated that the developmental switch between mGlu5 and mGlu1 receptors was driven by an epigenetic mechanism. Of particular interest was the demonstration that the appearance of mGlu1α receptors caused the developmental decline of mGlu5 receptors in PCs. Pharmacological experiments with mGlu1 receptor ligands administered to mice in restricted developmental time windows suggested that endogenous activation of mGlu1 receptors restrained mGlu5 receptor expression by enhancing Grm5 promoter methylation.
These findings raise a number of fundamental questions: (i) why is the mGlu5 receptor highly expressed by PCs in the first twelve days of postnatal life?; (ii) why a robust stimulation of PI hydrolysis is needed during that period?; (iii) why the mGlu1 receptor monopolizes the control of PC physiology by eliminating the mGlu5 receptor after the first two weeks of postnatal life?; and, (iv) are the two receptors different considering that they belong to the same subgroup and are both coupled to PI hydrolysis as a primary signal transduction mechanism?
Hydrolysis of phosphatidylinositol-4,5-bisphosphate generates inositol-1,4,5-trisphosphate (InsP3) and dyacylglycerol (DAG), which stimulates intracellular Ca2+ release and activates protein kinase C (PKC), respectively30. Experiments carried out in heterologous expression systems have shown that mGlu1 and mGlu5 receptors differ in the kinetics of intracellular Ca2+ release in response to receptor activation. Glutamate elicits single-peaked intracellular Ca2+ mobilization in cells expressing mGlu1 receptors and induces Ca2+ oscillations in cells expressing mGlu5 receptors31. This difference relies on a threonine residue of mGlu5 receptor (T845) that is localized in the G-protein interaction domain and is phosphorylated by PKC31. Although PCs are generated before birth, they undergo dramatic morphological changes in a postnatal time window that corresponds to mGlu5 receptor expression, particularly after PND7 when PCs develop their typical dendritic trees32. Dendritic development in rodent PCs proceeds through an initial phase in which the primitive dendritic tree is retracted and multiple filopodia-like processes are formed around the cell body (see Fig. 2D at PND7). After PND7, these processes disappear and the final dendritic tree starts to grow. This second phase of dendritic growth is driven by both intrinsic and extrinsic factors, including the glutamate release from afferent fibers33. Interestingly, a phase of developmental PC death precedes the onset of the second phase of dendritic growth33. Activation of mGlu5 receptors with ensuing stimulation of PI hydrolysis and intracellular Ca2+ oscillations might have a key role in any of these early postnatal events and might provide a survival signal for those Purkinje neurons that undergo early innervation by climbing and parallel fiber. Activation of mGlu5 receptors might also be involved in the early elimination phase of supranumerary climbing fibers (from PND7 to PND15), the underlying mechanism of which is still unknown7,34.
Activation of mGlu1 receptors after PND12 might provide a more localized Ca2+ signal, which, together with PKC-γ activation might be instrumental for the late phase of elimination of supranumerary climbing fibers via the induction of semaphorin-7A or other mechanisms9, as well as for parallel fiber elimination and territory segregation to the distal dendrites of PCs occurring after PND1510. mGlu1 receptors might optimize the signal-to-noise ratio in intracellular Ca2+ release during an active period of synaptic refinement by eliminating a competitor receptor (the mGlu5 receptor) that generates Ca2+ oscillations and waves that may spread across the cytoplasm. In addition, elimination of mGlu5 receptors might serve to deprive PCs of a morphogenetic signal that becomes inappropriate during a period of active synaptic refinement.
Interestingly, mGlu1 receptors maintained the suppressive action on mGlu5 receptors in the adult life, as shown by the reappearance of mGlu5 receptors in PCs after genetic deletion of mGlu1α receptors. At least four functional splice variants of mGlu1 receptors (named mGlu1α or a, -β1 or b, -β2 or f, and -γ or d) have been described35, of which mGlu1α predominates in the cerebellum, and mGlu1α and mGlu1β1 are expressed in PCs36,37. The long C-terminal domain exclusive of mGlu1α receptors allows interaction with Homer proteins38 and better phospholipase-C coupling efficacy39,40 and is also required for perisynaptic targeting of mGlu1 receptors, inositol-1,4,5-trisphosphate-mediated Ca2+ mobilization, elimination of supranumerary climbing fibers, LTD, and motor learning37. We showed that epigenetic regulation of mGlu5 receptor expression is dependent on mGlu1α variant. The role of mGlu1β1 receptor in reguration of mGlu5 receptor would be elucidated by use of mice in which mGlu1β1 transgene is introduced in PCs of mGlu1 receptor knockout mice.
A re-expression of mGlu5 receptors in PCs has been shown in pathological conditions characterized by a reduced expression or activity of mGlu1 receptors in PCs, i.e., in mice modelling type-1 spinocerebellar ataxia (SCA1)15, in mice developing experimental autoimmune encephalomyelitis (EAE)18, and in autoptic cerebellar samples from patients affected by multiple sclerosis18. This is one of the several examples of developmental proteins that are re-expressed in degenerating neurons. At least in SCA1 and EAE mice systemic treatment with mGlu5 receptor PAMs or NAMs did not affect cerebellar motor symptoms, suggesting that re-expressed mGlu5 receptors do not substitute for mGlu1 receptors in the regulation of motor learning and motor coordination. However, we cannot exclude that re-expressed mGlu5 receptors support the survival of degenerating PCs. If so, continuous treatment with a selective mGlu5 receptor PAM could slow the progression of cerebellar disorders by amplifying the otherwise sterile pro-survival program driven by mGlu5 receptors. This interesting hypothesis warrants further investigation.
Materials and Methods
9H-Xanthene-9-carboxylic acid (4-trifluoromethyl-oxazol-2-yl)-amide (RO0711401) was kindly provided by Roche Laboratories (Basel, Switzerland); 2-methyl-6-(phenylethynyl)-pyridine (MPEP), (3,4-dihydro-2H-pyrano[2,3-b]quinolin-7-yl) (cis-4-methoxycyclohexyl) methanone (JNJ16259685), N-cyclobutyl-6-[2-(3-fluorophenyl) ethynyl]-3-pyridine-carboxamide hydrochloride (VU0360172), and (RS)-3,5-dihydoxyphenylglycine (DHPG) were purchased from Tocris Biosciences (Bristol, United Kingdom).
Animals
All mice were kept under environmentally controlled conditions (ambient temperature, 22 °C; humidity, 40%) on a 12 h light/dark cycle with food and water ad libitum. All experiments were carried out according to the European (86/609/EEC) and Italian (D:Lgs. 116/92) guidelines of animal care and the animal welfare committees of the University of Tokyo (P14/117). The experimental protocol was approved by the Italian Ministry of Health (D.M.509/2015-PR).
C57BL/6 male mouse pups were used at postnatal day (PND) 3, 9, 12, 16, 18, 21, the day of birth was designed as PND zero.
Generation of mGlu1−/− mice bearing L7-tTA and TRE-mGlu1α transgenes (mGlu1α receptor cKO mice; C57BL/6 N background) proceeded as described previously41. mGlu1α receptor cKO mice were untreated or treated with 200 µg/ml of Dox (doxycycline hyclate; Sigma, St Louis, MO) in drinking water between 0 and 14 days of age or between 24 and 76 days of age. The Dox-water was delivered in dark bottles to protect Dox from light and changed twice a week. At 16 or 76 days of age, brain samples of these mGlu1α receptor cKO mice were collected. For immunohistochemical analysis, these mice were perfused with 4% paraformaldehyde in 0.1 M phosphate buffer.
RNA isolation, reverse transcription and quantitative real-time PCR
Total RNA from the cerebellum was isolated using the TRIzol reagent (Life Technologies, Monza, Italy) according to the manufacturer’s protocol and retrotranscribed into cDNA by using SuperScript III Reverse Transcriptase (Life Technologies). Real-Time PCR was performed on 20 ng of cDNA by using specific primers and Power SYBR Green Master Mix (Applied Biosystem, Foster City, CA) on an Applied Biosystems Step-One instrument. Thermal cycler conditions were as follows: 10 min at 95 °C, 40 cycles of denaturation (15 sec at 95 °C), and combined annealing/extension (1 min at 60 °C). Real-time PCR was performed by using the following primers:
mGlu1α/β receptors: forward, 5′-CATACGGAAAGGGGAAGTGA-3′; reverse, 5′-AAAAGGCGATGGCTATGATG-3′. mGlu5 receptors: forward, 5′-ACGAAGACCAACCGTATTGC-3′; reverse, 5′-AGACTTCTCGGATGCTTGGA-3′. GAPDH: forward, 5′-CGTCCCGTAGACAAAATGGT-3′; reverse, 5′-TCAATGAAGGGGTCGTTGAT-3′. mRNA levels were calculated from serially diluted standard curves simultaneously amplified with the samples and normalized to GAPDH mRNA levels.
Methylated DNA immunoprecipitation (MeDIP) Analysis
Mouse monoclonal 5-methylcytosine (5-mC) (Diagenode # C15200081-100, lot. GF-003) antibody were used for the immunoprecipitation. Genomic DNA was extracted from mice cerebellum and sonicated, by using Covaris S220 Ultrasonicator, to produce a fragment size of 300–600 bp. After ethanol precipitation 5 μg of sonicated DNA were diluted to 300 µl in TE buffer and heat-denatured at 95 °C for 10 min. Then, 30 μl of sonicated solution were removed and stored at −20 °C to be used to quantify the total amount of promoter before immunoprecipitation (input). The remaining solution was incubated over night at 4 °C with (5-mC monoclonal antibody, 0.5 μl/IP); magnetic beads conjugated to anti-mouse-IgG were used to bind the anti-5mC antibodies, and unbound DNA was removed in the supernatant. The immunoprecipitated DNA was released from the antibody complex by proteinase-K digestion for 2 h at 50 °C. After phenol-chloroform extraction and ethanol precipitation the DNA pellet was resuspended in 20 μl of water. CpG-rich Grm5 and Grm1 gene promoters were measured by Q-PCR with Power SYBR Green Master Mix (Applied Biosystem) on an Applied Biosystems Step-One instrument. Thermal cycler conditions were as follows: 95 °C for 10 min, 42 cycles of (95 °C for 15 seconds, 60 °C for 1 min). Q-PCR was performed by using the following primers: Grm5promoter: forward, 5′-ACCTGCTCTCCAGCTTCTCT-3′ reverse, 5′-GCCTCTTGGTCTCAGGGTTC-3′ Grm1 promoter: forward, 5′-ATGGCCTCCACTCTCTGGAT-3′ reverse 5′-ATCGGAGCCCTCTTCTCAGT-3′. Methylation level was calculated by using the formula % (MeDNA-IP/Total input) = 2[(Ct(20%input) − 3.32) − Ct (MeDNA-IP)] × 100%. Methylation level was confirmed by analysis of methylation enrichment of two promoters, unmethylated and methylated CpG island promoter genes GAPDH and histone H2B type 1-A (TSH2B) (Diagenode Cat# pp-1045-500 and #pp-1042-500).
Western blot analysis
The cerebella were dissected out and homogenized at 4 °C in RIPA buffer containing protease inhibitors cocktail (Merck Millipore, Milano, Italy) for 30 min and an aliquot was used for protein determination.
Equal amounts of proteins (20 μg) from supernatants were separated by 8% SDS polyacrilamide gel at 100 V for 1 hour for the detection of mGlu1α receptors or mGlu5 receptors, using a mini-gel apparatus (Bio-Rad Mini Protean II cell, Milano, Italy). Proteins were than electroblotted on Immuno PVDF membranes (Bio-Rad) for 7 min using Trans Blot Turbo System (Bio-Rad). Filters were washed three times and blocked for 1 hour in Tris-Tween buffered saline (TTBS) containing 5% non-fat dry milk.
The following primary antibodies were used: mouse polyclonal anti-mGlu1α receptor antibody (1:7000, BD Biosciences Milan, Italy, Cat #556389, lot. 3172791); rabbit monoclonal anti-mGlu5 receptor antibody (1:5000, Abcam, Cambridge, UK, Cat #AB76316, lot. GR45647-18). Filters were washed three times with TTBS buffer and then incubated for 1 hour with secondary peroxidase-coupled anti-mouse antibody (1:7000, Millipore Cat #401215) and anti-rabbit antibody (1:7000, Millipore, Cat #401393-2 ML).
Immunostaining was revealed by enhanced chemiluminescence luminosity (Amersham Pharmacia Biotech, Arlington Height, IL). The blots were re-probed with monoclonal anti β-actin antibody (1:50000, Sigma-Aldrich, Cat # A5441, lot. 116M4801V).
Confocal microscopy analysis
Mice were deeply anesthetized and perfused with ice-cold 4% paraformaldehyde, brains were dissected out and equilibrated with 30% sucrose overnight. Cerebellum was sectioned using a Leica cryostat (CM3050). For immunofluorescence analysis, serial sections were incubated with blocking solution (5% normal serum in 0.3% Triton X-100 in PBS) and then with the following antibodies overnight at 4 °C: anti-calbindin (1:500, Abcam, Cat #ab82812, lot. GR110079, and Santa Cruz Biotechnology, Dallas, TX, Cat #sc-7691, lot. G2512), anti-mGlu5 receptor (1:200, Abcam, Cat #AB76316, lot. GR45647-18) and anti-mGlu1α receptor (1:100, BD Biosciences, Cat #556389, lot. 3172791). Sections were also incubated with anti-mGlu5 receptor42 (1 μg/ml) and anti-type-8 carbonic anhydrase43 (1 μg/ml). After washing, sections were incubated with secondary antibodies conjugated with Alexa Fluor 488 (1:200, Molecular Probes, Eugene, OR, Cat #S32354) and Cy3 (1:200, Jackson ImmunoResearch Labs, West Grove, PA, Cat #715-167-003) for 2 hours at room temperature and rinsed in PBS. Finally, sections were mounted with anti-fading agent (Vector Laboratories, Burlingame, CA) and examined with ZEISS 780 confocal laser scanning microscope. We used a 488 nm argon laser to excite alexa488 and 543 HeNe laser to excite Cy3.
Measurement of polyphosphoinositide hydrolysis in cerebellum slices
Group I mGlu receptor-stimulated polyphosphoinositide (PI) hydrolysis was measured in cerebellum slices obtained from PND 3, 9, 12, 16 and 21 mice as described previously44. Briefly, cerebella were dissected aut and sliced (350 × 350 μm) using a McIlwain tissue chopper (Mickle Laboratory Engineering, Guildford, UK) and transferred in Krebs/Henseleit buffer (equilibrated with 95% O2, 5% CO2 to pH 7.4). Forty μl of gravity-packed slices were then incubated for 60 min in 350 μl of Krebs/Henseleit buffer containing 1 μCi of myo-[3H]inositol (18 Ci/mmol, GE Healthcare, Milano, Italy). Slices were incubated with LiCl (10 mM for 10 min) and, after 15 min, slices were challenged with the mGlu1/5 receptor agonist, DHPG, and JNJ16259685 or MPEP, if present, were added 5 min prior to DHPG.
One hour later, the incubation was stopped by the addition of 900 μl of methanol/chloroform (2:1). After further addition of 300 μl of chloroform and 600 μl of water, samples were centrifuged at low speed to facilitate phase separation, and the upper aqueous phase was loaded into Dowex 1-X-8 columns resin (100–200 mesh, formate form; Dow Chemical Company, Midland, MI) for the separation and quantification of [3H]InsP.
Columns were washed twice with water, once with a solution of 5 mM sodium tetraborate and 40 mM sodium formate to elute cyclic InsP and glycerophosphoinositols, and then with 6.5 ml of 0.2 M ammonium formate and 0.1 M formic acid for the elution of [3H]InsP. The remaining aqueous phase and the organic phase were dried under a continuous nitrogen stream, and 0.5 N NaOH was added to each sample.
Calcium imaging analysis
Parasagittal slices from the cerebellum were prepared from C57BL/6J mice at PND9, 17 and 20.
Briefly, animals were anesthetized, the brain including cerebellum dissected out and transferred into an ice-cold solution (artificial cerebro-spinal fluid, ACSF, in mM: 125 NaCl, 2.5 KCl, 2 CaCl2, 1 MgCl2, 25 glucose, pH 7.4 with 95% O2 and 5% CO2). Parasagittal slices (320 μm) were cut from the vermis with a Leica vibratome VT1000S, in an ice-cold solution containing the following (in mM): 130 K-gluconate, 15 KCl, 0.05 EGTA, 20 HEPES, and 25 glucose, with pH adjusted to 7.4 by NaOH, and then kept for 1 min in the solution (in mM): 225 D-mannitol, 2.5 KCl, 1.25 NaH2PO4, 26 NaHCO3, 25 glucose, 0.8 CaCl2, 8 MgCl2, with 95% O2 and 5% CO2.
Slices were recovered in ACSF for 30 min at 34 °C, then loaded for 45 min with the green fluorescent calcium dye OGB1-AM (10 μM) in the presence of 0.02% pluronic, and finally recovered for 45 min at room temperature.
For calcium imaging experiments, slices were perfused in a submerged chamber at a rate of 3–4 ml min-1 with (in mM): 120 NaCl, 2.5 KCl, 1 NaH2PO4, 26 NaHCO3, 1 MgCl2, 2 CaCl2, 10 glucose, pH 7.4 (with 95% O2 and 5% CO2). Oregon Green-loaded Purkinje neurons were visualized upon 488 nm laser excitation under a confocal microscope (TCS-SP5-RS, Leica Microsystems, Germany), and monitored at 1 Hz upon application of mGlu receptor ligands.
References
Xiao, B., Tu, J. C. & Worley, P. F. Homer: a link between neural activity and glutamate receptor function. Curr Opin Neurobiol 10, 370–4 (2000).
Fagni, L., Worley, P. F. & Ango, F. Homer as both a scaffold and transduction molecule. Sci STKE 137 (2002).
Abe, T. et al. Molecular characterization of a novel metabotropic glutamate receptor mGluR5 coupled to inositol phosphate/Ca2+ signal transduction. J Biol Chem 267, 13361–8 (1992).
Shigemoto, R., Nakanishi, S. & Mizuno, N. Distribution of the mRNA for a metabotropic glutamate receptor (mGluR1) in the central nervous system: an in situ hybridization study in adult and developing rat. J Comp Neurol 322, 121–35 (1992).
Aiba, A. et al. Deficient cerebellar long-term depression and impaired motor learning in mGluR1 mutant mice. Cell 79, 377–388 (1994).
Kano, M., Hashimoto, K. & Tabata, T. Type-1 metabotropic glutamate receptor in cerebellar Purkinje cells: a key molecule responsible for long-term depression, endocannabinoid signalling and synapse elimination. Philos Trans R Soc Lond B Biol Sci 363, 2173–86 (2008).
Kano, M. & Watanabe, T. Type-1 metabotropic glutamate receptor signaling in cerebellar Purkinje cells in health and disease. F1000Res 6, 416 (2017).
Kano, M. et al. Persistent multiple climbing fiber innervation of cerebellar Purkinje cells in mice lacking mGluR1. Neuron 18, 71–9 (1997).
Uesaka, N. et al. Retrograde semaphorin signaling regulates synapse elimination in the developing mouse brain. Science 344, 1020–3 (2014).
Ichikawa, R. et al. Territories of heterologous inputs onto Purkinje cell dendrites are segregated by mGluR1-dependent parallel fiber synapse elimination. Proc Natl Acad Sci USA 113, 2282–7 (2016).
Ichise, T. et al. mGluR1 in cerebellar Purkinje cells essential for long-term depression, synapse elimination, and motor coordination. Science 288, 1832–1835 (2000).
Kishimoto, Y. et al. mGluR1 in cerebellar Purkinje cells is required for normal association of temporally contiguous stimuli in classical conditioning. Eur J Neurosci 16, 2416–2424 (2002).
Liu, J. B., Yao, Y. X. & Jiang, W. Inhibitory effects of Group I metabotropic glutamate receptors antagonists on the expression of NMDA receptor NR1 subunit in morphine tolerant rats. Neurosci Lett 452, 268–72 (2009).
Shuvaev, A. N. et al. Mutant PKCγ in spinocerebellar ataxia type 14 disrupts synapse elimination and long-term depression in Purkinje cells in vivo. J Neurosci 31, 14324–34 (2011).
Notartomaso, S. et al. Pharmacological enhancement of mGlu1 metabotropic glutamate receptors causes a prolonged symptomatic benefit in a mouse model of spinocerebellar ataxia type 1. Mol Brain 6, 48 (2013).
Armbrust, K. R. et al. Mutant β-III spectrin causes mGluR1α mislocalization and functional deficits in a mouse model of spinocerebellar ataxia type 5. J Neurosci 34, 9891–904 (2014).
Becker, E. B. et al. A point mutation in TRPC3 causes abnormal Purkinje cell development and cerebellar ataxia in moonwalker mice. Proc Natl Acad Sci USA 106, 6706–11 (2009).
Fazio, F. et al. Switch in the expression of mGlu1 and mGlu5 metabotropic glutamate receptors in the cerebellum of mice developing experimental autoimmune encephalomyelitis and in autoptic cerebellar samples from patients with multiple sclerosis. Neuropharmacology 55, 491–9 (2008).
Sillevis Smitt, P. et al. Paraneoplastic cerebellar ataxia due to autoantibodies against a glutamate receptor. N Engl J Med 342, 21–7 (2000).
Marignier, R. et al. Metabotropic glutamate receptor type 1 autoantibody-associated cerebellitis: a primary autoimmune disease? Arch Neurol 67, 627–30 (2010).
Lopez-Chiriboga, A. S. et al. Metabotropic glutamate receptor type 1 autoimmunity: Clinical features and treatment outcomes. Neurology 86, 1009–13 (2016).
Neki, A. et al. Metabotropic glutamate receptors mGluR2 and mGluR5 are expressed in two non-overlapping populations of Golgi cells in the rat cerebellum. Neuroscience 75, 815–26 (1996).
Négyessy, L. et al. Light and electron microscopic demonstration of mGluR5 metabotropic glutamate receptor immunoreactive neuronal elements in the rat cerebellar cortex. J Comp Neurol 385, 641–50 (1997).
Fatemi, S. H. & Folsom, T. D. Dysregulation of fragile × mental retardation protein and metabotropic glutamate receptor 5 in superior frontal cortex of individuals with autism: a postmortem brain study. Mol Autism 2, 6 (2011).
Fatemi, S. H. & Folsom, T. D. GABA receptor subunit distribution and FMRP-mGluR5 signaling abnormalities in the cerebellum of subjects with schizophrenia, mood disorders, and autism. Schizophr Res 167, 42–56 (2015).
Fatemi, S. H., Folsom, T. D., Rooney, R. J. & Thuras, P. D. mRNA and protein expression for novel GABAA receptors θ and ρ2 are altered in schizophrenia and mood disorders; relevance to FMRP-mGluR5 signaling pathway. Transl Psychiatry 3, e271 (2013).
Pretto, D. I. et al. Reduced excitatory amino acid transporter 1 and metabotropic glutamate receptor 5 expression in the cerebellum of fragile X mental retardation gene 1 premutation carriers with fragile X-associated tremor/ataxia syndrome. Neurobiol Aging 35, 1189–97 (2014).
Casabona, G. et al. Expression and coupling to polyphosphoinositide hydrolysis of group I metabotropic glutamate receptors in early postnatal and adult rat brain. Eur J Neurosci 9, 12–7 (1997).
Catania, M. V. et al. Endogenous activation of group-I metabotropic glutamate receptors is required for differentiation and survival of cerebellar Purkinje cells. J Neurosci 21, 7664–73 (2001).
Berridge, M. J. The Inositol Trisphosphate/Calcium Signaling Pathway in Health and Disease. Physiol Rev 96, 1261–96 (2016).
Kawabata, S. et al. Control of calcium oscillations by phosphorylation of metabotropic glutamate receptors. Nature 383, 89–92 (1996).
Leto, K. et al. Consensus Paper: Cerebellar Development. Cerebellum 15, 789–828 (2016).
Sotelo, C. & Dusart, I. Intrinsic versus extrinsic determinants during the development of Purkinje cell dendrites. Neuroscience 162, 589–600 (2009).
Watanabe, M. & Kano, M. Climbing fiber synapse elimination in cerebellar Purkinje cells. Eur J Neurosci 34, 1697–710 (2011).
Ferraguti, F., Crepaldi, L. & Nicoletti, F. Metabotropic glutamate 1 receptor: current concepts and perspectives. Pharmacol Rev 60, 536–81 (2008).
Fotuhi, M. et al. Differential localization of phosphoinositide-linked metabotropic glutamate receptor (mGluR1) and the inositol 1,4,5-trisphosphate receptor in rat brain. J Neurosci 13, 2001–12 (1993).
Ohtani, Y. et al. The synaptic targeting of mGluR1 by its carboxyl-terminal domain is crucial for cerebellar function. J Neurosci 34, 2702–12 (2014).
Brakeman, P. R. et al. Homer: a protein that selectively binds metabotropic glutamate receptors. Nature 386, 284–8 (1997).
Prézeau, L. et al. Changes in the carboxyl-terminal domain of metabotropic glutamate receptor 1 by alternative splicing generate receptors with differing agonist-independent activity. Mol Pharmacol 49, 422–9 (1996).
Mary, S., Gomeza, J., Prézeau, L., Bockaert, J. & Pin, J. P. A cluster of basic residues in the carboxyl-terminal tail of the short metabotropic glutamate receptor 1 variants impairs their coupling to phospholipase C. J Biol Chem 273, 425–32 (1998).
Nakao, H., Nakao, K., Kano, M. & Aiba, A. Metabotropic glutamate receptor subtype-1 is essential for motor coordination in the adult cerebellum. Neurosci Res 57, 538–543 (2007).
Tanaka, J. et al. Gq protein α subunits Gαq and Gα11 are localized at postsynaptic extra-junctional membrane of cereballar Purkinje cells and hippocampal pyramidal cells. Eur J Neurosci 12, 781–792 (2000).
Patrizi, A. et al. Synapse formation and clustering of neuroligin-2 in the absence of GABAA receptors. Proc Natl Acad Sci USA 105, 13151–13156 (2008).
Nicoletti, F., Iadarola, M. J., Wroblewski, J. T. & Costa, E. Excitatory amino acid recognition sites coupled with inositol phospholipid metabolism: developmental changes and interaction with alpha 1-adrenoceptors. Proc Natl Acad Sci USA 83, 1931–5 (1986).
Author information
Authors and Affiliations
Contributions
S.N. and G.M. designed experiments, performed WB analysis, immunohistochemistry with confocal microscopy, and mouse treatments, and contributed to write the manuscript; H.N. and A.A. performed experiments with conditional mGlu1α receptor knockout mice; P.S. and F.L. performed WB analysis; C.Z. performed experiments of PI hydrolysis in cerebellar slices; Mi. M. performed mouse treatments; M.C. and Ma. M. performed qPCR and MeDIP experiments; M.Z. and G.C. performed calcium imaging experiments; R.G. and M.W. performed immunohistochemical analysis with confocal microscopy; G.B., V.B. and F.N. designed experiments and wrote the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Notartomaso, S., Nakao, H., Mascio, G. et al. mGlu1 Receptors Monopolize the Synaptic Control of Cerebellar Purkinje Cells by Epigenetically Down-Regulating mGlu5 Receptors. Sci Rep 8, 13361 (2018). https://doi.org/10.1038/s41598-018-31369-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-31369-7
This article is cited by
-
Coordination chemogenetics for activation of GPCR-type glutamate receptors in brain tissue
Nature Communications (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.