Abstract
Calmodulin-binding transcription activators (CAMTAs) represent the novel gene family of transcriptional regulators, which play important biological functions. Though, the first ever plant CAMTA gene was evidenced in Nicotiana tabacum in 2002. But, the systematic identification, origin and function of this gene family has not been performed due to the lack of reference genome information until now. Here, we identified 29 CAMTA genes in four Nicotiana species, including thirteen NtabCAMTAs, six NsylCAMTAs, and five NtomCAMTAs and NbenCAMTAs. These CAMTA families were classified into five phylogenetic groups (I-V), among which, the group-IV CAMTAs probably emerged the earliest. The NtabCAMTA family genes have diverse structures, and are randomly localized on five chromosomes and scaffolds. N. tabacum acquired 11 copies of homolog CAMATA genes from the parental genomes of N. tomentosiformis and N. sylvestris, followed by expansion through polyploidization and duplication. The NtabCAMTA genes were differentially expressed in different plant parts, and showed sensitivity towards different abiotic and biotic stresses. Co-expression network analysis revealed that some NtabCAMTA subunits interact with each other, and co-expressed. The current study is the first report presenting a comprehensive overview of Nicotiana CAMTA families, and opens a new avenue for the improvement of the cultivated tobacco.
Similar content being viewed by others
Introduction
Calcium (Ca2+) ions act as ubiquitous secondary messengers for many cellular signaling pathways in eukaryotes1. Ca2+-mediated signal transduction is the key mechanism for transporting signals resulting from different stimuli, hence mediating growth, development and stress response in plants2,3. These nuclear and cytoplasmic Ca2+ signals are detected by different Ca2+-binding proteins such as Calmodulin (CaM), which upon binding to Ca2+, activates and alters the activity of CaM-binding proteins4. Transcription factors (TFs) regulated by Ca2+ or CaM are especially important in this phenomenon. So far, many TFs in plant are reported to interact with CaM5. Among the reported TFs, Calmodulin-binding transcription activators (CAMTAs) represent the latest and novel set of CaM-interacting proteins in plants. NtER1, the first plant and tobacco CAMTA gene was reported to be developmentally regulated and acts as a trigger for senescence and death6. Until now, the CAMTAs have been identified and reported in numerous plant species including Arabidopsis thaliana, rice, grapevine, cabbage and many more7,8,9,10. The plant CAMTA-encoded proteins comprise multiple functional domains, including CG-1, which is named after a partial cDNA clone isolated from parsley encoding a sequence-specific DNA-binding domain11, IPT/TIG (Ig-like, plexins, transcription factors or transcription factor immunoglobulin), ankyrin (ANK) repeats, and calmodulin-binding IQ motifs. These domains take part in protein–protein interactions, CaM binding, nonspecific DNA contacts in TFs and protein dimerization, respectively12,13,14. The Arabidopsis CAMTA family is comprised of six member genes, designated as AtCAMTA1- AtCAMTA615. Latest studies have shown that these genes show quick and differential response to external stimuli, and are crucial for cross-talk between multiple signal transduction pathways involved in stress tolerance6,8,16.
Nicotiana tabacum (common tobacco) is chief commercial/cash crop, cultivated worldwide17,18. Enriched with alkaloid nicotine, tobacco leaves are largely used in cigarettes, cigars, chewing or smoking tobaccos and snuff. N. tabacum is used as a model plant organism and a key tool for plant molecular research, and a source of the BY-2 plant cell line to study primary biological processes19,20. It is also used as a model for plant disease susceptibility, which it shares with other Solanaceae plants including potato, tomato and pepper18. N. tabacum is an allotetraploid specie (2n = 4x = 48), most likely to be originated from a hybridization event (tetraploidization) between S (N. sylvestris) and T (N. tomentosiformis) genomes approximately 200,000 years ago21. N. tabacum therefore has a relatively large genome size (approximately 4500 Mb) compared with other cultivated Solanaceae crops22, and is 50% larger than the human genome.
So far, numerous varieties of tobacco have been domesticated and improved around the world including flue-cured, burley, oriental and cigar23. Similar to N. tabacum, N. benthamiana Domin (wild tobacco) is also an accepted model tobacco specie and has been widely used in experiments related to plant-virus response, protein localization, and plant-based systems for protein expression and purification24,25. Because of its complexity and larger size, the fully annotated reference genome sequence of N. tabacum was not available until now, which left behind large gaps in studying important biological pathways and gene families of tobacco including CAMTA. Taking advantage of the available genome data by Sierro et al.18,26. Edwards et al.27 and China tobacco (Ren et al. unpublished), we used comprehensive bioinformatics and experimental approaches to perform genome-wide identification and characterization of CAMTA gene family in N. tabacum, N. sylvestris, N. tomentosiformis and N. benthaminana species. To elucidate the evolutionary relationship between tobacco and other plants, we comprehensively analyzed the phylogeny between the orthologs of CAMTAs of four Nicotiana species and all plant lineages. Using the available RNA-seq data and Real-time quantitative PCR analysis, we quantified and analyzed the expression profiles of NtabCAMTA family genes during plant growth and development, and stress responses to different biotic and abiotic factors. This study will help to identify novel CAMTA genes for future breeding to improve plant production, quality and stress resistance, and open a new avenue for further elucidation for their roles underlying the signal transduction in tobacco.
Results
Genome-wide identification and domain analyses of CAMTA gene families in Nicotiana
To perform genome-wide identification and obtain the complete overview of CAMTA gene family in four Nicotiana species, a blast search in the tobacco genome sequences dataset was performed using AtCAMTAs as queries. Thirty-five candidate protein sequences were analyzed for the presence of CAMTA-specific conserved domains (CG-1: a sequence-specific DNA-binding domain; IPT/TIG: Ig-like, plexins, transcription factors or transcription factor immunoglobulin; ANK: ankyrin repeats; IQ: calmodulin-binding IQ motifs. As a result, six gene accessions having truncated amino acid sequences and/or lacking specific domains were discarded from analyses (Table S1). Finally, twenty-nine full length CAMTA genes having essential domains were identified from four Nicotiana species, including thirteen from N. tabacum, six from N. sylvestris, and five genes from N. tomentosiformis and N. benthamiana, respectively. These genes were named as NtabCAMTAs, NsylCAMTAs, NtomCAMTAs and NbenCAMTAs, based on their positions in phylogenies (Table 1 and Fig. S1).
The identified Nicotiana CAMTA proteins contained four conserved domains, namely CG-1, ANK repeats, IPT/TIG and IQ motifs, which are characteristic to plant CAMTAs (Fig. 1a). The occurrence of ANK domain varied from 1 to 3, and IQ from 2 to 3. Interestingly, two of N. benthamiana proteins (NbenCAMTA1 and NbenCAMTA2) contained an additional CG-1 domain, which requires further elucidation (Table S2). The multiple sequence alignment exhibited high sequence similarity between 29 CAMTAs, particularly in conserved domain regions. Among the four domains, TIG/IPT is the largest and less conserved (<80%) domain, whereas, CG-1 is interrupted by extra stretch of amino acids from NbenCAMTA1 and NbenCAMTA2 (Fig. S2). We derived stringent consensus motif keys showing the similarities and variances within the conserved domains of Nicotiana CAMTAs. Besides, these motifs can be used to identify and characterize the CAMTA proteins in Nicotiana species in future (Fig. 1b).
Graphical representation of primary domain architecture of Nicotiana CAMTA-encoded proteins (a) and their derived consensus motif keys (b). The four CAMTA-specific domains are: CG-1 (a sequence-specific DNA-binding domain), IPT/TIG (Ig-like, plexins, transcription factors or transcription factor immunoglobulin), ANK (ankyrin) repeats, and calmodulin-binding IQ motifs. The square brackets “[]” indicate the amino acids allowed in this position of motif; “X” represents any amino acid, while round brackets “()” denote the number of amino acids. Letters highlighted as red strikethrough separates the two consecutive domains from each other.
Phylogenetic classification of Nicotiana CAMTAs
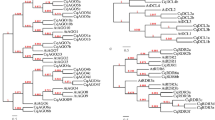
To determine the homology between the Nicotiana CAMTA proteins, a rooted maximum likelihood (ML) phylogenetic tree of the 29 CAMTAs was constructed with 6 AtCAMTAs, using CAMTA proteins from Amborella trichocarpa, Chlamydomonas reinhardtii and Selaginella moellendorffii as outgroup. The inferred rooted tree produced well resolved phylogeny with high bootstrap or Bayesian support, showing that Nicotiana CAMTA family proteins can be classified into five major clusters/groups. Of these five groups, Group I, II, IV and V fall into the phylogenetic classification of Arabidopsis CAMTA family, hence named accordingly. However, two Nicotiana CAMTA proteins (i.e., NtabCAMTA6 and NsylCAMTA3) clustered separately, hence placed into a separate group III (Fig. 2). Group-IV comprised ten CAMTA genes, thus making the largest clade, while, group-III was the smallest clade with two genes only. The best ML scoring rooted tree topology indicated that during evolution Group-IV CAMTAs probably emerged the earliest among all in Nicotiana, while, Group I, II, III and V as the latest respectively (Fig. 2).
Phylogenetic analysis and classification of CAMTA family proteins identified in this study. The analysis involved 36 amino acid sequences, including 29 from 4 Nicotiana species identified in this study, 6 from A. thaliana and single from A. trichocarpa (AmTr_v1.0_scaffold00013.39) as outgroup. The tree with the highest log likelihood (−11525.8253) is shown. The bootstrap values from 1000 resampling are given at each node. Five groups were identified in tobacco CAMTA families, which were named as Group-I, II, III, IV and V. These groups were identified on the basis of Arabidopsis CAMTAs (marked with pink circles), while the members of each CAMTA family are shown with colored diamonds. Evolutionary analyses were conducted in MEGA 6.0.
Evolutionary relationship between Nicotiana and other plant CAMTAs
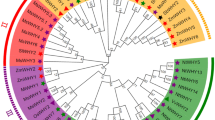
To determine the evolutionary relationship between tobacco and other plants, we comprehensively analyzed the phylogeny between single orthologs of CAMTAs of four Nicotiana species and all plant lineages. For the purpose, we identified 56 CAMTA sequences from different species including green algae, bryophytes, lycophytes, gymnosperms, monocots and dicots, whose protein sequences harbored typical domains and motifs of CAMTA proteins (Table S3).
At least one species from all plant lineages and main groups was selected. The resultant tree contained five major groups with significant bootstrap values, showing the expansion of CAMTA genes in Group I. According to the tree’s topology CAMTAs of all plants share a common ancestor, where the lower plants settled in the basal group. NtabCAMTAs found its place among other Solanaceous plants (S. lycopersicum and S. melongena) in Group I. These analyses also revealed that CAMTA gene family expansion occurred conspicuously greater in eudicots among all plant lineages (Fig. S3). In many previous studies expansion in gene families due to duplications has been discussed28,29,30,31,32,33.
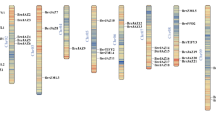
The NtabCAMTA gene family
There are thirteen genes in the genome of cultivated tobacco (NtabCAMTA1-13). The nucleotide lengths of these genes varies between 2658 bp (NtabCAMTA12) and 3324 bp (NtabCAMTA4), while, their encoded protein lengths ranges between 885 (NtabCAMTA12) and 1107 amino acids (NtabCAMTA4) (Table 1). The NtabCAMTA family genes are randomly scattered throughout the N. tabacum genome. As shown in Table 1, seven NtabCAMTA genes are localized on five chromosomes, while five genes are clustered on six scaffolds. Three NtabCAMTA genes are localized on chromosome 2, and chromosomes 3, 6, 9 and 12 has only one gene. Nine of the NtabCAMTA genes are positioned on reverse strands of the chromosomes/scaffolds, and four genes are oriented on forward strand. Alignment between N. tabacum and Arabidopsis CAMTA-encoded proteins showed that NtabCAMTAs have high similarity with AtCAMTAs, especially in CG-1, ANK and IQ regions. In each region, more than 50 residues stretches are conserved, where the differences show the divergence across the two species (Fig. S4). The phylogenetic analyses of NtabCAMTA with Arabidopsis CAMTAs resulted into similar clustering pattern of five groups, where, Group-I contained 2 NtabCAMTA genes, Group-II and V had 3 genes each, Group-III had one gene, while, Group-IV comprised 4 genes (Fig. S5).
The ProtParam tool showed that NtabCAMTA proteins greatly differ in molecular weights (ranging from 99.453 to 124.089.02 kDa), consistent with the number of atoms present. Nearly all of the NtabCAMTA proteins have relatively low isoelectric points (pI < 9), and are hydrophilic. Comparatively, NtabCAMTA4 is the most hydrophilic, while NtabCAMTA6 is the least hydrophilic protein. The aliphatic index showed that most NtabCAMTA proteins are thermostable as other globular proteins. According to the instability index (II), only two proteins (NtabCAMTA4 and NtabCAMTA12) could be classified as stable in the test tubes. In addition, NtabCAMTA proteins, except NtabCAMTA11 and NtabCAMTA12, have more negatively charged residues (aspartic acid/glutamic acid) as compared to positively charged residues (arginine/lysine) (Table 2).
Origin and expansion of NtabCAMTA family genes
The phylogeny between N. tabacum and its ancestors (N. sylvestris and N. tomentosiformis) provided clear picture of how the NtabCAMTA gene family originated and expanded, and to detect the retention and/or loss of CAMTA genes after genome duplication and polyploidization. Since T and S genomes contain 5 and 6 CAMTA genes respectively, which are likely to produce 10, 11 or 12 genes in N. tabacum, if the ancestral genes are to be counted 5 or 6. As of now, there are 13 CAMTA genes in N. tabacum. As shown in Fig. S6, 11 of these NtabCAMTA genes have clear single ancestor in phylogenetic tree, showing that each gene has been acquired from single parent and retained. Two gene pairs (i.e., NtabCAMTA4/NtabCAMTA5 and NtabCAMTA12/NtabCAMTA13), showing high resemblance at nucleotide level (92.12 and 95.24%) as well as at amino acid level (98.64 and 91.69%), seem to be originated by duplication event, which might have occurred after the divergence from parental species (Fig. S6). Two accessions (Ntab0503030and Ntab0966680), which were discarded during preliminary analyses, seem to be redundant copies that probably lost their functions during the long course of evolution. However, this presumption requires experimental validation. Together, these finding suggest that polyploidization, along with gene duplication played important role in expansion of CAMTA gene family in N. tabacum.
Structures of NtabCAMTA family genes and conserved motifs in their-encoded proteins
To characterize the structural diversity of the NtabCAMTA family, exon-intron organization analysis of the individual gene was performed (Fig. 3a). The majority of the NtabCAMTA genes contain twelve or thirteen exons, where NtabCAMTA1, NtabCAMTA10 and NtabCAMTA11 contain highest number of exons (14). Most of the introns in NtabCAMTA genes are in intron phase 0 (77), interrupted by exact triplet codons. Twenty-seven phase-II introns (separated by 2nd and 3rd codons) were observed in NtabCAMTA family genes, where each gene comprise two phase-2 introns, except for NtabCAMTA1, which contain three phase-2 introns. Forty-eight single phase-I introns were detected in NtabCAMTA genes, where each gene contain 3 or 4 single phase-I introns. Several changes in terms of loss/gain of exons, intron phases and their shuffling was observed between the structures of genes belonging to same phylogenetic group, thus adding diversity to both the structures and functions of NtabCAMTAs (Fig. S7). We observed that NtabCAMTA4 and NtabCAMTA12 of the duplicated gene pairs (NtabCAMTA4/5 and NtabCAMTA12/13) have lost their single phase-I intron and exon subsequently, unlike NtabCAMTA5 and NtabCAMTA13 that have similar exon numbers, intron phases and lengths. This suggest that NtabCAMTA4 and NtabCAMTA12 genes arose from NtabCAMTA5 and NtabCAMTA13, which evolved from parental genes. Besides, AtCNGCs comprise more phase-I introns than tobacco homologs, showing that intron loss during the long course of evolution resulted into reduced numbers of introns in NtabCAMTA family genes, particularly in members of Group I-V (Fig. S8).
Schematic diagram representing the structures of NtabCAMTA genes and the distribution of conserved motifs in associated proteins. (a) Gene structures showing the organization of exon and intron structures, and associated intron phases [0, 1 and 2] of 13 NtabCAMTA genes. The NJ phylogenetic tree of CDs is shown on the left side of the figure. (b) The distribution of conserved motifs identified in NtabCAMTA-encoded proteins. Each motif is represented by a colored box, and their names given at the bottom of diagram. The logos and annotations of functionally defined motifs are given in Fig. S9 and Table S4, respectively. The order of motifs corresponds to their positions in protein sequence, however, the length of the boxes does not correspond to the lengths of motifs.
When submitted to online MEME server (Multiple Expectation Maximization for Motif Elicitation), the NtabCAMTA proteins were found to contain at least ten conserved motifs. Among these, seven motifs (motif 1–5, 7 and 8) are part of the known domains, as shown by Pfam codes and WebLogos (Fig. 3b; Fig. S9 and Table S4). Motif 1 and 4 are associated with CG-1 domain; motif 2 is the longest motif of 59 residues that correlate with ankyrin repeat profile or ANK domain; motif 3 is 50 residues long motif associated with IQ domain; motif 5 represent IPT/TIG domain, and contain phosphorylation sites for casein kinase II (CK2), protein kinase C. On other hand, two motifs (i.e., motif 7 and motif 8) are not correlated with known domain in pfam, however, their secondary association can be linked with the MurC (UDP-N-acetylmuramate-alanine ligase [Cell envelope biogenesis, outer membrane]) and xanthine phosphoribosyltransferase, respectively (Table S4). The functionality of the remaining motifs (6, 9 and 10) is still unknown, and awaits further experimental proof.
Potential microRNA target sites in NtabCAMTA transcripts
Identifying the target sites in gene transcripts provide valuable information regarding the role of miRNAs in plant growth, signal transduction pathways and stress responses. Analysis of 164 published tobacco microRNAs34 revealed that NtabCAMTA4 and NtabCAMTA5 contain the target sites for two miRNAs (i.e., nta-miR6164a and nta-miR6164b) (Table 3). Keeping cut-off threshold of 4.5 in the search parameter, which give higher prediction coverage compared to default threshold of 3.0, we identified 8 miRNAs (6 families such as nta-miR159, nta-miR394, nta-miR395, nta-miR477, nta-miR6163 and nta-miR6020b) containing target sites in 7 NtabCAMTA transcripts with the same expectation score (Table S5). The nucleotide lengths of these miRNAs was 21nt, with more stringent cut-off threshold (1.5) showing lower false positive prediction35. These miRNAs are located on the 3′ arm of the stem-loop hairpin structures. The UPE (target accessibility of target site), which is key feature in target identification and exhibit energy required to contact (and cleave) target mRNA, varied from 10.5 (nta-miR6164a) to 16.8 (nta-miR6164a), where lower energy reflects the higher possibility of contact between miRNA and target site. Two miRNAs were found to be involved in cleavage of the target transcript. The nta-miR6164 class of miRNAs has been reported to be involved in wounding and topping stress response in tobacco34.
GO enrichment analysis
Using Blast2GO (v.3.3.5), we were able to assign total of 78 gene ontology (GO) classes to 13 NtabCAMTA genes with blast matches to known proteins in InterPro. Of these, majority were assigned to biological process (35), followed by molecular function (32) and cellular components (10). All genes were found to be integral components of membrane or localized in nucleus (Table S6). These proteins are involved in molecular processes associated with kinase activities, nucleotide, proteins, ions/receptors binding, and the regulation of transferase activities (Table S7). Notably, these NtabCAMTA genes were associated to with GO-terms for numerous biological processes including regulation of gene expression, transport, signal transduction, response to stimulus, anatomical structure development, cell differentiation and other developmental processes (Table S8).
In planta expression of CAMTA genes in tobacco tissues
To get insight into the steady-state expression of NtabCAMTA genes, we utilized the transcriptomic RNA-seq data from Sequence Read Archive in GeneBank (SRP029183) that were reported previously18. The generated RNA-seq data included the expression profiles of seed, leaf (whole leaf, vein and blade), stem, callus, root and flower bud of N. tabacum TN90 at five leaf stages. The final expression data of 13 NtabCAMTA genes was log transformed and illustrated in heatmap (Fig. 4a). Among all, eight NtabCAMTA genes were significantly expressed at relatively higher levels in at least one tissue, including five in leaves, four in roots, leaf blade and stem tissue, two in flower, and single gene was expressed in seed and callus respectively. Five genes namely NtabCAMTA1, NtabCAMTA2, NtabCAMTA6, NtabCAMTA9 and NtabCAMTA10 did not expressed in any tissue. Further investigation revealed that expression of NtabCAMTA3 was up-regulated in all studied plant parts, where it showed maximum level of expression in callus, seed, root, stem, leaf blade and flower bud, demonstrating its importance in tobacco plant growth and development. Among the other genes, NtabCAMTA11 and NtabCAMTA12 showed highest levels of expression in leaf vein, followed by NtabCAMTA13 and NtabCAMTA3 respectively. Post-topping RNA-seq data showed that the members of phylogenetic group-II and group-IV were particularly induced in root and leaf tissues (Fig. 4b). Five genes, including NtabCAMTA3, NtabCAMTA7, NtabCAMTA8, NtabCAMTA10 and NtabCAMTA11 were induced in stem tissues, showing their induction under wounding. Taken together, NtabCAMTA genes exhibited differential expression pattern in different tissues and several genes are induced by wounding in tobacco genome. Higher expression in same tissue indicated their functional conservation, but others point toward their functional diversities.
Expression profiles of NtabCAMTA genes in different plant parts of N. tabacum cultivar: TN90. (a) Normalized expression levels (FPKM log2) at early stage of tobacco growth. (b) Normalized expression levels (FPKM log2) in post-harvested tissues of mature tobacco. The gene names and cluster tree are indicated on the left. The intensity of transcript abundance is indicated with different colors (blue = lower accumulation, black = higher accumulation) and white histograms within the heatmap.
Responses to biotic stress
To get insight into the role of NtabCAMTAs in disease resistance and host stress response, a RT-qPCR analysis was performed on tobacco seedlings (having four fully expanded leaves) exposed to different phytopathogens, using mock inoculum as control. The pathogens used in this experiment included viral pathogens Cucumber mosaic virus-M (M strain of CMV) and Potato virus Y (Mn strain of PVY), and a fungal pathogen black shank or Phytophthora nicotianae36,37. The results showed that the NtabCAMTA genes under study differentially responded to each pathogen understudy at certain time point (Fig. 5). In CMV infected tobacco seedlings, the expression levels of the CAMTA were slightly but significantly repressed at 6 hpi and 24 hpi showing early and late responses. After 6 hpi, the expression levels of eight NtabCAMTA genes were induced and down-regulated compared to control. Only five genes showed down-regulation after 24 hpi. Compared to control, maximum positive response to CMV (fold-change) was noted for group-IV NtabCAMTA genes (Fig. 5a). After inoculation with PVY, the expression of all NtabCAMTA genes was increased after 24 hpi. None of the NtabCAMTA genes showed early response at 6 hpi. Overall, maximum response to PVY-Mn was shown by NtabCAMTA4 that was increased by >2.8-folds after 24 hpi, followed by NtabCAMTA7, which was >2.35-folds up-regulated after 24 hpi compared with control (Fig. 5a).
Results of RT-qPCR analysis showing the expression patterns of CAMTA genes in tobacco plant subjected to external stress at different time points. (a) Biotic stress response to CMV, PVY and BSD (left panel). (b) Abiotic stress response to cold, drought and cadmium (right panel). The names of NtabCAMTA genes are shown with the numbers in X-axis, while Y-axis show the relative expression levels or fold-changes of treatments versus control (ck). Bars with same letter means no significant difference based on LSD test (p ≤ 0.05). For experimental details see Materials and Methods.
Black shank disease (BSD), caused by the oomycete pathogen P. nicotianae, is a devastating root rot disease of tobacco38, causing wilting and yellowing of leaves, along with dark brown to black spots at the base of the plant stem. We calculated the expression levels of NtabCAMTA genes in the leaves of susceptible tobacco variety (i.e., Hong hua Da jin yuan or HD) and resistant cultivar (i.e., Resistance to Black Shank Tobacco or RBST) inoculated with P. nicotianae for 6 or 24 h. The data depicted that most NtabCAMTA genes were early induced by P. nicotianae inoculation. The group-III genes (i.e., NtabCAMTA3, 4 and 5) exhibited almost similar trend of up-regulation in both cultivars, where the intensity of gene expression was higher in N. tabacum cv. HD than the N. tabacum cv. RBST. Moreover, the expression of NtabCAMTA2 gene was significantly induced after 24 hpi in both cultivars. Meanwhile, maximum response was noted for NtabCAMTA3 that was >4-times up-regulated in N. tabacum cv. HD. On other hand, three genes were down-regulated by P. nicotianae inoculation in N. tabacum cv. HD, and nine genes were negatively induced in N. tabacum cv. RBST (Fig. 5a).
Over all the transcription data demonstrated that NtabCAMTAs are amongst the early genes to sense and respond to biotic stress, and their regulation play important roles in plant defense.
Responses to abiotic stress
To assess the role of NtabCAMTA genes in response to abiotic stress, we investigated their expression patterns in the leaves of tobacco seedlings subjected to cold stress at 4 °C for 1–2 days, and drought stress for 2, 4 and 8 days, respectively. Additionally, we quantified expression profiles of NtabCAMTA genes in leaf and root samples of tobacco plants subjected to cadmium stress (Cd: 250 mM) for 1 day. Under cold stress, the expressions of seven genes (i.e., NtabCAMTA3-7, 10 and 11) significantly increased at each time point, and those of five genes decreased after 2 days, compared to control. On other hand, most of the NtabCAMTA genes were up-regulated after 2 day of drought stress. Comparatively, the maximum up-regulation was noted for NtabCAMTA3 that was >9-folds increased after 8 days of drought stress. Among others, group-V NtabCAMTA genes were differentially expressed, depending on the time scale (Fig. 5b).
Under Cd stress, four NtabCAMTA genes were significantly expressed in leaf tissues, while seven genes were expressed in roots. Only one gene (i.e., NtabCAMTA6) was significantly down-regulated in tobacco roots. The remaining five genes (i.e., NtabCAMTA1-3, 8 and 12) did not exhibit significant expressional response to Cd stress in terms of fold-change compared to control. Overall, the expression response of NtabCAMTA genes considerably varied among individual groups, suggesting the importance of the CAMTA genes in the survival of tobacco plants under different abiotic stress conditions (Fig. 5b).
Co-expression network analysis
To enhance the presentation of the dynamical and conservative expression profiling of NtabCAMTA genes, we performed gene co-expression networks analyses for all members of this gene family by means of the RNA-Seq and RT-qPCR analysis data. A global view of co-expression network based on Pearson’s correlation coefficient threshold of 0.75, we found that the expression patterns of 12 NtabCAMTA genes closely correlated (Fig. 6a). Among these, NtabCAMTA12 and 13 independently co-expressed with each other. The NtabCAMTA6 gene, which belong to a separate clade of phylogenetic Group-III, did not show correlation with other NtabCAMTA genes. We further constructed three more networks with Cytoscape to display the relationships between NtabCAMTA genes, which differentially expressed under different conditions (Fig. 6b–d). The expression patterns of eleven NtabCAMTA genes were significantly correlated with each other in different plant tissues (Fig. 6b). Our analysis also indicated that some members of the phylogenetic Groups-IV (i.e., NtabCAMTA7–10) did not co-expressed in biotic and abiotic stress conditions (Fig. 6c,d). On other hand, NtabCAMTA genes belonging to Group-I and IV, co-expressed in group specific manners under abiotic stress (Fig. 6d). Overall, most of these genes displayed similar expression patterns and the results suggested the possibility that NtabCAMTA subunits interact with each other under different conditions.
Co-expression networks of NtabCAMTA genes showing global view of dynamical and conservative expression profiling. (a) Overall co-expression network of NtabCAMTA genes using combined expression data. (b) Co-expression network using in-planta RNA-seq data. (c) Co-expression network under biotic stress. (c) Co-expression network under abiotic stress. The diagrams were prepared using cytoscape. The edge line width represents the Pearson correlation coefficient (r) with the value of “r” ranging between 0.7 to 1. NtabCAMTA genes showing dynamic expressions are not indicated in the diagram.
Discussion
Bioinformatics tools and publicly released genomics data have led to the identification of numerous plant gene families, especially in model plants such as Arabidopsis. Among novel families, the CAMTA gene family has been reported in many plants of agriculture importance7,8,9,10. However, genome-wide identification and annotation of CAMTA genes has not been reported in any of Nicotiana species. In this study, 13 CAMTA family genes were identified in N. tabacum genome, which are distributed onto on five chromosomes/scaffolds and clustered into four phylogenetic groups. In addition, we identified six CAMTA genes in N. sylvestris, and five genes in N. tomentosiformis and N. benthaminana, the wild tobacco. The number of genes in NtabCAMTA family are greater than CAMTA genes in most of the reported crops, such as A. thaliana (6), Oryza sativa (5), Solanum lycopersicum (7), Brassica campestris (8), Vitis vinifera (10), thus making it the largest CAMTA family in plants so far. This is probably due to the larger genome size of N. tabacum of about ~4500 Mb. The NtabCAMTA-encoded proteins are characterized by presence of CG-1, ANK, IQ and/or IPT/TIG domains, which are evolutionarily conserved and characteristic to plant CAMTAs. It is assumed that homologous genes within the same taxonomic/phylogenetic group exhibit similar structural, functional and evolutionary properties, which might help in understanding the role of CAMTA genes in N. tabacum. Here we found that NtabCAMTA-encoded proteins showed high similarity with corresponding NsylCAMTAs, NtomCAMTAs, NbenCAMTAs and AtCAMTAs in terms of domain architectures, amino acid composition and phylogenies. In addition, these proteins share homology with the corresponding orthologues identified in other plant species. As mentioned earlier, N. tabacum is an allotetraploid crossbreed originated from the presumptive parental diploids N. sylvestris and N. tomentosiformis39. We confirmed that most of the NtabCAMTA family genes were descended from parental species, and gene duplication significantly contributed to the expansion of this family40. Our results are corroborated by the findings of Xu et al.41, stating that gene duplication increases the genome content and expands gene function to guarantee optimum adaptability and evolution of plants. Meanwhile, two genes (Ntab0695330 and Ntab0966680) were found to lack functional domains in their sequences, which are probably lost during the long course of evolution. These results are supported by the findings of Liang et al.42, stating that functionally redundant gene copies are often lost during duplication, and only active copies of functional genes are retained.
As in other organisms, transcription factors in plant systems are regulated by different types of miRNAs43. We observed that two NtabCAMTA gene transcripts (i.e., NtabCAMTA4 and NtabCAMTA5) comprised target sites for nta-miR6164a and nta-miR6164b. Former research has shown that these miRNAs are involved in regulation of pathways associated with morphological and metabolic adaptations44, hormone45 and symbiotic nitrogen fixation46. Besides, some of miRNAs, which were predicted with low expectancy score (nta-miR159, nta-miR394, nta-miR395 and miR477) are reported to play important roles in various abiotic and biotic stress responses including: Cd, salt, cold, heat, drought, Fe deficiency, UV-B radiation, hypoxia or oxidative stress, and resistance to powdery mildew infection and tobacco mosaic virus47,48,49,50,51,52,53,54,55,56,57.
The CAMTA transcription factor family play functional role in plant response to several abiotic and biotic stresses, including cold, wounding, drought and pathogens58. Infectious diseases such as PVY, CMV and BSD, along with drought cold and Cd stress have been reported to affect the tobacco yield and production each year59. The detailed analyses of gene expression data in different tissues and stress conditions further clarified the important role of the different CAMTAs in the growth, development and survival of N. tabacum. The expression patterns of NtabCAMTAs were different in different tissues, where most of the genes were expressed in leaf vein and blades. Among them, the transcription level of NtabCAMTA3 was significantly higher than other genes, especially after topping, showing their importance in wound response. Under abiotic stress, NtabCAMTA3 and NtabCAMTA10 had increasing expressions under cold stress, while, others showing differential expression patterns. Our results are consistent with the findings of Doherty et al.60, who evidenced that CG-1 sequence is important for early cold response in AtCAMTA proteins. We noticed that two of N. benthaminana CAMTA genes (NbenCAMTA1 and NbenCAMTA2) comprise two CG-1 domains, therefore, it would be interesting to note if extra domains are functional or contribute to the degree of cold stress tolerance in wild tobacco. Additionally, these NtabCAMTA family genes showed differential responses to viral pathogens CMY, PVY, and black shank pathogen infection, drought and cadmium stress. Finally, different co-expression networks were constructed, which revealed that some of NtabCAMTA subunits interact with each other under different conditions. Multiple studies have shown that certain members of CAMTA family modulate different biotic and abiotic stress responses61,62, which corroborate our findings. Further studies are required to clarify the role and mechanism of differentially expressed NtabCAMTA genes in plant growth and development, and the regulation of signal transduction and stress resistance related pathways.
Conclusion
This work is the first inclusive report about genome-wide identification and systematic characterization of CAMTA gene family in four Nicotiana species with focus on common tobacco. These CAMTA families can be classified into five phylogenetic groups, and Group-IV CAMTAs probably emerged the earliest among all in Nicotiana. There are 13 genes in N. tabacum CAMTA family originating from ancestral genomes of N. sylvestris and N. tomentosiformis. Both polyploidization and duplication events played important role in the expansion of NtabCAMTA family. The available information from bioinformatics analysis can be used in futuristic studies to identify and characterize the CAMTA proteins in Nicotiana species. For example, using stringent consensus motif keys to identify new genes/families in Nicotiana/solanaceous species, construction of protein-protein interaction networks, experimentally validating gene structures and miRNA targets. Furthermore, the expression data of the differentially expressed CAMTA genes (such as NtabCAMTA-3, 6, 7, 10 and 13) can lay the foundation for investigating their molecular regulatory mechanisms, and breeding new cultivars with improved yield, quality and tolerance to abiotic/biotic stress.
Methods
Identification of CAMTA gene family in four Nicotiana species
The reference genome and proteome sequences of tobacco (N. tabacum) variety “K326”, N. sylvestris and N. tomentosiformis available at http://www.tobaccodb.org/were used for annotation of the candidate CAMTA genes. In order to identify these gene families, the DNA and amino acid sequences of six Arabidopsis CAMTA family genes were downloaded from TAIR10 (https://www.arabidopsis.org/), and used as queries to perform homology based search in http://www.tobaccodb.org/, using BLASTN and BLASTP programs respectively with default parameters. Similarly, the 6 AtCAMTA protein sequences were used to search against N. benthaminana genome using TBLASTN at Sol Genomics Network (http://solgenomics.net/). All non-redundant protein sequences of the candidate CAMTA genes were retrieved and subjected to domain analysis by domain analysis programs: Simple Modular Architecture Research Tool (SMART) (http://smart.embl-heidelberg.de/) and the Conserved Domains Database (CDD) (http://www.ncbi.nlm.nih.gov/Structure/cdd/wrpsb.cgi), with the default cut off parameters. Sequences containing CG-1 (PF03859), TIG (PF01833), ANK (PF12796) and IQ (PF00612) domains were recognized as CAMTA proteins. The identified NtabCAMTA, NtomCAMTA, NsylCAMTA and NbenCAMTA genes were named according to their positions in phylogenetic tree. The analysis also included putative orthologous CAMTA genes from other plant species, which were BLASTP searched and downloaded from Phytozome (https://phytozome.jgi.doe.gov/), using NtabCAMTA1-encoded protein as query. Moreover, the sequence data of the identified CAMTA genes for N. tabacum, N. sylvestris and N. tomentosiformis was deposited at the GenBank (https://www.ncbi.nlm.nih.gov/genbank/), which can be accessed (accession numbers: MF142771-MF142783, and MH119945-MH119955) by the readers to retrieve, confirm and reproduce the analysis.
Characterization and physicochemical properties
The information regarding the general characteristics of CAMTA genes and proteins was obtained from the “http://www.tobaccodb.org/”. Amino acid properties and other physicochemical traits such as charge, molecular weight (g mol−1), aliphatic and instability index (II), isoelectric points (pI), grand average of hydropathy (GRAVY) and other properties of a given NtabCAMTA proteins were calculated using the ProtParam tool in the ExPASy web server5. The post-translational modifications sites were predicted by using the ScanProsite tool6.
Sequence alignments and phylogenetic analysis
The multiple sequence alignments for the predicted CAMTA proteins were performed using ClustalX 2.0 program with the default settings63, and viewed by GeneDoc program64. The identified conserved CAMTA-specific domains were manually checked, verified and shaded with DNAMAN software (version 6.0.3.40, Lynnon Corporation). MEGA 6.0 software was used to conduct the evolutionary analyses. Initial tree(s) for the heuristic search were obtained by applying the Neighbor-Joining method to a matrix of pairwise distances estimated using a Jones-Taylor-Thornton (JTT) matrix-based model. Maximum Likelihood method based on the JTT model with bootstrap of 1000 replicates was used to construct the final phylogenetic trees65,66. The trees were drawn to scale, with branch lengths measured in the number of substitutions per site.
Prediction of gene structure, motifs and miRNA target sites
The structures of the CAMTA family genes showing exon-intron organization were determined based on alignments of their coding sequences with the corresponding genomic sequences, and a diagram was obtained using Gene Structure Display Server (GSDS 2.0, http://gsds.cbi.pku.edu.cn/). The conserved motifs in the NtabCAMTA proteins were identified in MEME web server8, keeping the optimal motif width between 6 and 200, and the maximum number of different motifs as 1067. The discovered motifs were annotated with Pfam program (http://pfam.xfam.org/).
For miRNA’s target sites prediction within the NtabCAMTA transcripts, the complete sequence information of all known and published miRNAs of the N. tabacum was obtained from miRBase (http://www.mirbase.org/). The obtained sequences of 164 tobacco miRNAs and NtabCAMTA transcripts were used as input to the psRNATarget server (http://plantgrn.noble.org/psRNATarget/) using default settings and threshold.
Analysis of CAMTA gene expression in tobacco tissues
To explore the expression patterns of NtabCAMTA genes in different tobacco tissues, the Illumina RNA-sequencing data of N. tabacum cultivar: TN90 was downloaded from GenBank archives at https://www.ncbi.nlm.nih.gov/bioproject/PRJNA208209/SRP02918318. There were 36.8 M reads of each RNA sample, from which the low quality reads/adapters were removed, and mapped to the tobacco genes. The gene expression data was normalized by FPKM (fragments per kilobase per million)68. The resulting FPKM values of NtabCAMTA genes were log2 transformed, and the heat maps of the hierarchical clustering were generated and visualized using R language program69.
Growth conditions, sample preparation and stress treatments
Tobacco plants were cultivated in Guiyang County, Guizhou Province, China, under normal growth conditions untill three to five fully expanded leaf stage was reached. For drought stress, N. tabacum cv. hongda seedlings were kept away from water for 2, 4 and 8 days at reduced relative humidity of ~35%. The whole leaf tissues were harvested from control and dehydrated plants after 2, 4 and 8 days respectively.
For cold and cadmium treatments, N. tabacum cv. hongda seedlings were grown on Murashige and Skoog medium70 for three weeks at 25 ± 1 °C with 16 h of light and 8 h of dark. In earlier case the seedlings were exposed to 4 °C of temperature and leaf samples were collected after 1 day, while in later case seedlings were grown on MS medium supplemented with 250 mM of Cd, and the leaf and root samples were harvested.
For viral inoculum preparation. ~0.5 g of systemically infected leaf tissue from CMV/PVY-infected tobacco seedlings were picked and instantly homogenized with kieselguhr in 1 ml water. The viral inoculums were subsequently rub-inoculated onto the top of two leaves of fresh plants having four fully expanded leaves. The mock inoculum was prepared from leaves of the healthy plants and applied in the same way as viral inoculum. Both viral and mock-inoculated plants were sampled after 6 hours and 1 day respectively. The cultivation of BSD pathogen P. nicotianae, and the infection of HD and RBST cultivars were performed using the method described by Scharte et al.71 and Essmann et al.72. After harvesting, each sample was cut into small pieces, followed by immediate storage in liquid nitrogen and subsequent storage at −80 °C until further processing.
RNA extraction and RT-qPCR analysis
Total RNA extraction was performed using TRIzolTM reagent (TransGen) following the manufacturer’s instructions. First-strand cDNA was reverse-transcribed from total RNA using M-MLV reverse transcriptase (Promega) with oligo (dT) as the primer. PCR was performed in a total volume of 10 μL containing 5 μL of 2× SYBR Premix Ex Taq (TaKaRa), 2 μmol L−1 of each gene-specific primer (Table S9), 0.5 μL of the cDNA sample, 0.2 μL of Rox Reference DyeII (TaKaRa) on an ABI StepOne Real-time PCR instrument (Applied Biosystems). The reactions were carried out using the following program: 95 °C for 1 min, 45 cycles of 95 °C for 15 sec, and 60 °C for 34 s. The tobacco actin gene (LOC107788267) was used as an internal reference. Each experiment was performed with three technical replicates. Finally, the 2−ΔΔCt method73 was used to calculate the relative gene expression values, which were subsequently transformed to fold-change and plotted in figures. Student’s t-tests were used to determine significant differences.
Identification of correlated genes and network construction
To determine the co-expression network for NtabCAMTA genes, the FPKM data from in planta, and different stress treatments were collected and used. First, we ranked the correlated genes with Pearson’s correlation coefficient threshold of higher than 0.75. Then, we recalculated the Pearson’s correlation coefficients of the genes and conditions with p-value ≤ 0.05, with the R project (version 3.2.3). Cytoscape (version 2.8.2) software was used to construct co-expression networks between NtabCAMTA and co-expression genes.
Statistical analysis
All the data were subjected to analysis of variance (ANOVA) using computer statistical package (SAS software SAS Institute, Cary, NC). General linear model (GLM) procedure was used to check the significant differences among main treatments. Individual comparisons between mean values were performed by using the least significant differences (LSD) test (p ≤ 0.05). Correlation analysis was performed by CORR procedure and R project.
Change history
10 March 2020
An amendment to this paper has been published and can be accessed via a link at the top of the paper.
References
Wu, M. et al. Genome-wide identification and expression analysis of the IQD gene family in moso bamboo (Phyllostachys edulis). Sci. Rep. 6, 24520, https://doi.org/10.1038/srep24520 (2016).
Takáč, T. et al. Comparative proteomic study of Arabidopsis mutants mpk4 and mpk6. Sci. Rep. 6, 28306, https://doi.org/10.1038/srep28306 (2016).
DeFalco, T. A. et al. Multiple Calmodulin-binding Sites Positively and Negatively Regulate Arabidopsis Cyclic Nucleotide-Gated Channel12. Plant Cell (2016).
Kim, M. C., Chung, W. S., Yun, D.-J. & Cho, M. J. Calcium and calmodulin-mediated regulation of gene expression in plants. Mol Plant. 2, 13–21 (2009).
Galon, Y., Finkler, A. & Fromm, H. Calcium-regulated transcription in plants. Mol Plant. 3, 653–669 (2010).
Yang, T. & Poovaiah, B. W. An early ethylene up-regulated gene encoding a calmodulin-binding protein involved in plant senescence and death. J. Biol. Chem. 275, 38467–38473 (2000).
Choi, M. S. et al. Isolation of a calmodulin-binding transcription factor from rice (Oryza sativa L.). J. Biol. Chem. 280, 40820–40831 (2005).
Finkler, A., Ashery-Padan, R. & Fromm, H. CAMTAs: calmodulin‐binding transcription activators from plants to human. FEBS Lett. 581, 3893–3898 (2007).
Shangguan, L. et al. Identification and bioinformatic analysis of signal responsive/calmodulin-binding transcription activators gene models in Vitis vinifera. Mol. Biol. Rep. 41, 2937–2949 (2014).
Hu, R., Wang, Z., Wu, P., Tang, J. & Hou, X. Identification and abiotic stress analysis of calmodulin-binding transcription activator/signal responsive genes in non-heading Chinese cabbage (Brassica campestris ssp. chinensis Makino). POJ 8, 141–147 (2015).
da Costa e Silva, O. CG-1, a parsley light-induced DNA-binding protein. Plant Mol. Biol. 25, 921–924, https://doi.org/10.1007/bf00028887 (1994).
Aravind, L. & Koonin, E. V. Gleaning non-trivial structural, functional and evolutionary information about proteins by iterative database searches1. J. Mol. Biol. 287, 1023–1040 (1999).
Song, K. et al. The transcriptional coactivator CAMTA2 stimulates cardiac growth by opposing class II histone deacetylases. Cell 125, 453–466 (2006).
Bork, P., Doerks, T., Springer, T. A. & Snel, B. Domains in plexins: links to integrins and transcription factors. Trends Biochem. Sci. 24, 261–263, https://doi.org/10.1016/S0968-0004(99)01416-4 (1999).
Bouché, N., Scharlat, A., Snedden, W., Bouchez, D. & Fromm, H. A novel family of calmodulin-binding transcription activators in multicellular organisms. J. Biol. Chem. 277, 21851–21861 (2002).
Yang, T. & Poovaiah, B. W. A calmodulin-binding/CGCG box DNA-binding protein family involved in multiple signaling pathways in plants. J Biol Chem. 277, 45049–45058 (2002).
Nawaz, Z. et al. Genome-wide identification, evolution and expression analysis of cyclic nucleotide-gated channels in tobacco (Nicotiana tabacum L.). Genomics, https://doi.org/10.1016/j.ygeno.2018.01.010 (2018).
Sierro, N. et al. The tobacco genome sequence and its comparison with those of tomato and potato. Nature communications 5 (2014).
Nagata, T., Nemoto, Y. & Hasezawa, S. Tobacco BY-2 cell line as the “HeLa” cell in the cell biology of higher plants. Int. Rev. Cytol. 132, 1–30 (1992).
Zhang, J., Zhang, Y., Du, Y., Chen, S. & Tang, H. Dynamic metabonomic responses of tobacco (Nicotiana tabacum) plants to salt stress. J. Proteome Res. 10, 1904–1914 (2011).
Leitch, I. J. et al. The ups and downs of genome size evolution in polyploid species of Nicotiana (Solanaceae). Ann. Bot. 101, 805–814 (2008).
Arumuganathan, K., Martin, G. B., Telenius, H., Tanksley, S. D. & Earle, E. D. Chromosome 2-specific DNA clones from flow-sorted chromosomes of tomato. Molecular and General Genetics MGG 242, 551–558 (1994).
Lu, X. et al. Development of DArT markers for a linkage map of flue-cured tobacco. Chin. Sci. Bull. 58, 641–648 (2013).
Goodin, M. M., Zaitlin, D., Naidu, R. A. & Lommel, S. A. Nicotiana benthamiana: its history and future as a model for plant-pathogen interactions. Mol. Plant-Microbe Interact. 21, 1015–1026 (2008).
Baksa, I. et al. Identification of Nicotiana benthamiana microRNAs and their targets using high throughput sequencing and degradome analysis. BMC Genomics 16, 1, https://doi.org/10.1186/s12864-015-2209-6 (2015).
Sierro, N. et al. Reference genomes and transcriptomes of Nicotiana sylvestris and Nicotiana tomentosiformis. Genome Bio. 14, R60 (2013).
Edwards, K. D. et al. A reference genome for Nicotiana tabacum enables map-based cloning of homeologous loci implicated in nitrogen utilization efficiency. BMC Genomics 18, 448, https://doi.org/10.1186/s12864-017-3791-6 (2017).
He, L., Zhao, M., Wang, Y., Gai, J. & He, C. Phylogeny, structural evolution and functional diversification of the plant PHOSPHATE1 gene family: a focus on Glycine max. BMC Evol. Biol. 13, https://doi.org/10.1186/1471-2148-13-10 (2013).
Du, H. et al. Genome-wide identification and evolutionary and expression analyses of MYB-related genes in land plants. DNA Res. 20, 437–448 (2013).
Gomez-Porras, J. L. et al. Phylogenetic analysis of K+ transporters in bryophytes, lycophytes, and flowering plants indicates a specialization of vascular plants. Front. Plant Sci. 3, 167, https://doi.org/10.3389/fpls.2012.00167 (2012).
Sze, H., Geisler, M. & Murphy, A. S. Linking the evolution of plant transporters to their functions. Front. Plant Sci. 4, https://doi.org/10.3389/fpls.2013.00547 (2014).
Nawaz, Z., Kakar, K. U., Saand, M. A. & Shu, Q.-Y. Cyclic nucleotide-gated ion channel gene family in rice, identification, characterization and experimental analysis of expression response to plant hormones, biotic and abiotic stresses. BMC Genomics 15, 1, https://doi.org/10.1186/1471-2164-15-853 (2014).
Saand, M. A. et al. Phylogeny and evolution of plant cyclic nucleotide-gated ion channel (CNGC) gene family and functional analyses of tomato CNGCs. DNA Res. 22, 471–483 (2015).
Tang, S. et al. Identification of wounding and topping responsive small RNAs in tobacco (Nicotiana tabacum). BMC Plant Biol. 12, 28–28 (2012).
Dai, X. & Zhao, P. X. psRNATarget: a plant small RNA target analysis server. Nucleic Acids Res. 39, W155–W159, https://doi.org/10.1093/nar/gkr319 (2011).
Whitham, S. A. et al. Diverse RNA viruses elicit the expression of common sets of genes in susceptible Arabidopsis thaliana plants. Plant J. 33, 271–283 (2003).
Marathe, R., Guan, Z., Anandalakshmi, R., Zhao, H. & Dinesh-Kumar, S. Study of Arabidopsis thalianaresistome in response to Cucumber mosaic virus infection using whole genome microarray. Plant Mol. Biol. 55, 501–520 (2004).
Ji, P., Csinos, A. S., Hickman, L. L. & Hargett, U. Efficacy and application methods of oxathiapiprolin for management of black shank on tobacco. Plant Dis. 98, 1551–1554 (2014).
Volkov, R. A., Borisjuk, N. V., Panchuk, I. I., Schweizer, D. & Hemleben, V. Elimination and rearrangement of parental rDNA in the allotetraploid Nicotiana tabacum. Mol. Biol. Evol. 16, 311–320 (1999).
García-Olmedo, F., Carbonero, P., Aragoncillo, C. & Salcedo, G. Loss of redundant gene expression after polyploidization in plants. Experientia 34, 332–333 (1978).
Xu, G., Guo, C., Shan, H. & Kong, H. Divergence of duplicate genes in exon–intron structure. Proc. Natl. Acad. Sci. 109, 1187–1192 (2012).
Liang, Y. et al. Genome-wide identification, structural analysis and new insights into late embryogenesis abundant (LEA) gene family formation pattern in Brassica napus. Sci. Rep. 6, https://doi.org/10.1038/srep24265 (2016).
Hirayama, T. & Shinozaki, K. Research on plant abiotic stress responses in the post‐genome era: Past, present and future. Plant J. 61, 1041–1052 (2010).
Zhang, Z. et al. Submergence-responsive microRNAs are potentially involved in the regulation of morphological and metabolic adaptations in maize root cells. Ann. Bot. 102, 509–519 (2008).
Reyes, J. L. & Chua, N. H. ABA induction of miR159 controls transcript levels of two MYB factors during Arabidopsis seed germination. Plant J. 49, 592–606 (2007).
Subramanian, S. et al. Novel and nodulation-regulated microRNAs in soybean roots. BMC Genomics 9, 1, https://doi.org/10.1186/1471-2164-9-160 (2008).
Arenas-Huertero, C. et al. Conserved and novel miRNAs in the legume Phaseolus vulgaris in response to stress. Plant Mol. Biol. 70, 385–401 (2009).
He, X., Zheng, W., Cao, F. & Wu, F. Identification and comparative analysis of the microRNA transcriptome in roots of two contrasting tobacco genotypes in response to cadmium stress. Sci. Rep. 6, https://doi.org/10.1038/srep32805 (2016).
Jia, X., Ren, L., Chen, Q.-J. & Li, R. & Tang, G. UV-B-responsive microRNAs in Populus tremula. J. Plant Physiol. 166, 2046–2057 (2009).
Kong, W. W. & Yang, Z. M. Identification of iron-deficiency responsive microRNA genes and cis-elements in Arabidopsis. Plant Physiol. Biochem. 48, 153–159 (2010).
Li, F. et al. MicroRNA regulation of plant innate immune receptors. Proc. Natl. Acad. Sci. 109, 1790–1795 (2012).
Liu, H.-H., Tian, X., Li, Y.-J., Wu, C.-A. & Zheng, C.-C. Microarray-based analysis of stress-regulated microRNAs in Arabidopsis thaliana. RNA 14, 836–843 (2008).
Lu, S., Sun, Y. H. & Chiang, V. L. Stress‐responsive microRNAs in Populus. Plant J. 55, 131–151 (2008).
Moldovan, D. et al. Hypoxia-responsive microRNAs and trans-acting small interfering RNAs in Arabidopsis. J. Exp. Bot., erp296 (2009).
Sunkar, R. & Zhu, J.-K. Novel and stress-regulated microRNAs and other small RNAs from Arabidopsis. Plant Cell 16, 2001–2019 (2004).
Xin, M. et al. Diverse set of microRNAs are responsive to powdery mildew infection and heat stress in wheat (Triticum aestivum L.). BMC Plant Biol. 10, 123 (2010).
Zhou, X., Wang, G. & Zhang, W. UV‐B responsive microRNA genes in Arabidopsis thaliana. Mol. Syst. Biol. 3, 103, https://doi.org/10.1038/msb4100143 (2007).
Reddy, A. S. N., Ali, G. S., Celesnik, H. & Day, I. S. Coping with stresses: roles of calcium-and calcium/calmodulin-regulated gene expression. Plant Cell 23, 2010–2032 (2011).
Riquinho, D. L. & Hennington, E. A. Health, environment and working conditions in tobacco cultivation: a review of the literature. Ciência & Saúde Coletiva 17, 1587–1600 (2012).
Doherty, C. J., Van Buskirk, H. A., Myers, S. J. & Thomashow, M. F. Roles for Arabidopsis CAMTA transcription factors in cold-regulated gene expression and freezing tolerance. Plant Cell 21, 972–984 (2009).
Pandey, N. et al. CAMTA 1 regulates drought responses in Arabidopsis thaliana. BMC Genomics 14, 1, https://doi.org/10.1186/1471-2164-14-216 (2013).
Li, X. et al. Tomato SR/CAMTA transcription factors SlSR1 and SlSR3L negatively regulate disease resistance response and SlSR1L positively modulates drought stress tolerance. BMC Plant Biol. 14, 1, https://doi.org/10.1186/s12870-014-0286-3 (2014).
Larkin, M. A. et al. Clustal W and Clustal X version 2.0. Bioinformatics 23, 2947–2948 (2007).
Nicholas, K. B. & Nicholas, H. B. Jr. GeneDoc: a tool for editing and annotating multiple sequence alignments. Distributed by the author. 1997.
Jones, D. T., Taylor, W. R. & Thornton, J. M. The rapid generation of mutation data matrices from protein sequences. Computer applications in the biosciences: CABIOS 8, 275–282 (1992).
Tamura, K., Stecher, G., Peterson, D., Filipski, A. & Kumar, S. MEGA6: molecular evolutionary genetics analysis version 6.0. Mol. Biol. Evol. 30, 2725–2729 (2013).
Bailey, T. L. et al. MEME SUITE: tools for motif discovery and searching. Nucleic Acids Res. 37, W202–208, https://doi.org/10.1093/nar/gkp335 (2009).
Mortazavi, A., Williams, B. A., McCue, K., Schaeffer, L. & Wold, B. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat. Methods 5, 621 (2008).
Higo, K., Ugawa, Y., Iwamoto, M. & Higo, H. PLACE: a database of plant cis-acting regulatory DNA elements. Nucleic acids research 26, 358–359 (1998).
Murashige, T. & Skoog, F. A revised medium for rapid growth and bio assays with tobacco tissue cultures. Physiol. Plant. 15, 473–497 (1962).
Scharte, J., SCHÖN, H. & Weis, E. Photosynthesis and carbohydrate metabolism in tobacco leaves during an incompatible interaction with Phytophthora nicotianae. Plant, Cell Environ. 28, 1421–1435 (2005).
Essmann, J. et al. RNA interference-mediated repression of cell wall invertase impairs defense in source leaves of tobacco. Plant Physiol. 147, 1288–1299 (2008).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. methods 25, 402–408 (2001).
Acknowledgements
This study was supported by the CNTC Research Programs 110201201006, 110201301004 and 110201503007, and the ZYQK Research Program 201601.
Author information
Authors and Affiliations
Contributions
K.U.K. and Z.N. designed this study and conceptualized the methodology. C.Z. collected the data and perform bioinformatics work along with J.J. and K.U.K. P.C. performed the experiments with assistance from Q.Y.S. K.U.K. and Z.N. analyzed the data and wrote manuscript. Q.Y.S. revised the manuscript. All authors commented on it at every stage. X.l.R. supervised the study.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kakar, K.U., Nawaz, Z., Cui, Z. et al. Evolutionary and expression analysis of CAMTA gene family in Nicotiana tabacum yielded insights into their origin, expansion and stress responses. Sci Rep 8, 10322 (2018). https://doi.org/10.1038/s41598-018-28148-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-28148-9
This article is cited by
-
Genome-wide identification of DCL, AGO, and RDR gene families in wheat (Triticum aestivum L.) and their expression analysis in response to heat stress
Physiology and Molecular Biology of Plants (2023)
-
Characterization of the CqCAMTA gene family reveals the role of CqCAMTA03 in drought tolerance
BMC Plant Biology (2022)
-
Genome‑wide identification of CAMTA gene family members in rice (Oryza sativa L.) and in silico study on their versatility in respect to gene expression and promoter structure
Functional & Integrative Genomics (2022)
-
EgJUB1 and EgERF113 transcription factors as potential master regulators of defense response in Elaeis guineensis against the hemibiotrophic Ganoderma boninense
BMC Plant Biology (2021)
-
Comprehensive genome-wide analysis of calmodulin-binding transcription activator (CAMTA) in Durio zibethinus and identification of fruit ripening-associated DzCAMTAs
BMC Genomics (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.