Abstract
An imbalance between energy uptake and energy expenditure is the most important reason for increasing trends in obesity starting from early in life. Extracellular miRNAs are expressed in all bodily fluids and their expression is influenced by a broad range of stimuli. We examined whether screen time, physical activity and BMI are associated with children’s salivary extracellular miR-222 and miR-146a expression. In 80 children the extracellular fraction of saliva was obtained by means of differential centrifugation and ultracentrifugation. Expression levels of miR-222 and miR-146a were profiled by qPCR. We studied the association between children’s salivary extracellular miRNA expression and screen time, physical activity and BMI using mixed models, while accounting for potential confounders. We found that higher screen time was positively associated with salivary extracellular miR-222 and miR-146a levels. On average, one hour more screen time use per week was associated with a 3.44% higher miR-222 (95% CI: 1.34 to 5.58; p = 0.002) and 1.84% higher miR-146a (95% CI: −0.04 to 3.75; p = 0.055) level in saliva. BMI and physical activity of the child were not significantly associated with either miR-222 or miR-146a. A sedentary behaviour, represented by screen time use in children, is associated with discernible changes in salivary expression of miR-146a and or miR-222. These miRNA targets may emerge attractive candidates to explore the role of these exposures in developmental processes of children’s health.
Similar content being viewed by others
Introduction
A rising prevalence in obesity, both in children and adults has emerged over the last decade1. Obesity fosters the development of a variety of morbidities, among which type 2 diabetes and cardiovascular disease2. The onset of these diseases could occur already early in life, with obesity being an important risk factor3.
An unhealthy lifestyle, with an imbalance between energy intake and expenditure is the predominant cause of obesity. Insufficient energy expenditure can directly be linked to a lack of physical activity4. Although it is well-accepted physical activity is an effective measure to reduce multiple health risk factors, a sedentary lifestyle is frequent in all age groups5. Children spend a substantial amount of time in sedentary pursuits, such as watching television, using the computer and playing videogames. This behaviour not only increases the risk to develop obesity, but also increases the risk to develop metabolic syndrome and cardiovascular disease6.
miRNAs are short (~22 nt long) single-stranded RNA molecules which control gene networks and play a central role in a various (patho)physiological processes7. miRNAs are present both intracellular and in the extracellular environment. Extracellular miRNAs, which are protected from degradation and carried by vesicles and protein aggregates, are stably present in all bodily fluids. Recently, they are recognized as important signal molecules and/or biomarkers in the pathology of various morbidities8. As such, circulating miRNAs are implemented in cardiometabolic disease mechanisms9. Studies focusing on differential miRNA expression patterns in patients with various cardiometabolic disorders compared with healthy subjects showed the importance of miR-222 and miR-146a. In this regard, the inflammatory miR-146a is linked to metabolic and cardiovascular disease9. miR-222 is a cell cycle regulator, which is involved in both physiological and pathological cardiac processes10. The (extracellular) expression profiles of miRNAs in body fluids are suggested as potential biomarkers for cardiometabolic processes. However, these studies either focus on subjects already having a specific metabolic or cardiovascular disorder or on specific interventions in adults. Studies linking miR-146a or miR-222 with lifestyle factors related to cardiometabolic outcomes in childhood are currently lacking. We investigated whether miR-146a and miR-222 in the extracellular fraction of saliva are associated with screen time use, physical activity and BMI in children.
Methods
Study population
This study was based on the COGNAC (COGNition and Air pollution in Children) study11,12,13, which is a panel study with three repeated measures. Between 2011 and 2013, we invited children (grades three to six) from three primary schools in Flanders (Belgium) to participate. 43.4% of all invited children participated in the study. The examinations took place between December 2011 and February 2014 on Monday, Tuesday, Thursday, and Friday between 9:00 a.m. and 2:00 p.m. Of the 334 children within the COGNAC cohort, we randomly selected 80 children, from two participating schools. Saliva samples of the first two study visits, which were approximately 3 months apart, were selected. For each child, all visits were scheduled at the same day of the week and same time point, to rule out diurnal variation. The study was approved by the medical ethical committee of Hasselt University and the Eastern-Limburg Hospital (Belgium) in accordance with the Helsinki Declaration14. Parental informed consent was obtained prior to participation in the study. Information on the socio-economic status (by parental education and occupation), exposure to second-hand smoke through parental smoking and the use of TV and PC screens was accessible by a questionnaire filled out by the parents. Based on the reported weekly TV and PC screen use, we calculated the total screen time as the sum of TV and PC screen use. Based on information of the reported sport activities, out-of-school sport activities were identified and categorized as “none” (i.e. no out-of-school sport activities), “low” (i.e. ≤3 hours per week), “middle” (>3 to <6 hours per week) and high (≥6 hours per week). Passive smoking was defined as exposure to indoor tobacco smoke, when one or more family member(s) smoked inside the house. Height and weight were recorded and body mass index (BMI) was calculated. Overweight and obesity were defined according to international childhood BMI thresholds15.
Saliva extracellular miRNAs
In order to avoid contamination of the samples, children refrained at least 30 minutes from eating, drinking or hygienic procedures prior to saliva donation. Additionally, they rinsed three times with tap water to eliminate possible food residues. Saliva (2 ml) was collected using the Oragene® RNA self-collecting kits (DNA Genotek Inc.) and immediately stabilized by mixing with RNA stabilizer. Within 6 hours, the samples were stored at −20 °C until further analyses.
The extracellular fraction of the saliva was obtained by differential centrifugation and ultracentrifugation, protocol was adapted from Théry et al.16 to integrate the processing of saliva specific to the Oragene collection kits in the procedure. After thawing, the saliva was incubated at 50 °C for one hour. Subsequently, a 1 ml aliquot was incubated at 90 °C for 15 minutes. Next, the debris present in saliva was pelleted by adding 40 µl of neutralizer solution (DNA Genotek Inc.) and centrifuging samples at 1500 × g for 10 minutes. The supernatant was collected and centrifuged at 16000 × g for 20 minutes. Then, the supernatant was ultracentrifuged at 160000 × g for one hour (Optima LE-80 K ultracentrifuge equipped with a ti70 fixed angle rotor; Beckman). The polyallomer tubes for ultracentrifugation were pre-treated with RNAZap (Life Technologies) to remove RNAse activity. After the first ultracentrifugation step, the pellet was resuspended in 1x PBS (pH 7.4) and ultracentrifuged at 160000 × g for one hour. Afterwards, the pellet, containing vesicles and protein aggregates17, was resuspended in RNAse-free water and stored at −80 °C.
Total RNA, including small miRNAs was isolated from the extracellular fraction of saliva using the miRNeasy mini kit (Qiagen), following the manufacturer’s instructions. Samples were spiked with 250 fmol C. elegans miR-39 for normalization of the expression data. 125 ng total RNA was reverse transcribed using looped primers for specific miRNA cDNA synthesis (Megaplex RT primers human pool A & Taqman mircoRNA RT kit; Life Technologies) on a PCR gradient thermal cycler (TC-5000; Techne). For all samples, 1:75 dilutions were made and an equal volume of products was mixed with reagents of the Taqman miRNA assay and the Taqman Fast Advanced mastermix (Life Technologies) for quantification of the miRNAs. qPCR was carried out on an ABI 7900HT sequence detection system (Life Technologies) and thermal cycling was for 10 minutes at 95 °C, followed by 40 cycles of 15 seconds at 95 °C and one minute at 60 °C. All runs were carried out in triplicate and with a no-template control (NTC) on 384-well plates with three inter run calibrators (IRC). Raw qPCR data were analysed using the SDS Relative Quantification Software (version 2.3; Applied Biosystems). Cq values were transformed to a relative quantity to the external spike-in miRNA cel-miR-39 in qbase+ software (Biogazelle).
Statistical analysis
We used SAS (version 9.3, SAS institute Inc., Cary, NC, USA) software for data management and statistical analyses. Demographic characteristics are represented as mean (standard deviation) for continuous variables or number (frequency) for categorical variables. miRNA expression data were log10-transformed to improve normality of the data.
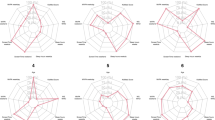
We correlated the total screen time use with BMI (Fig. 1a) and we evaluated the associations of screen time, physical activity and the BMI of the child using linear regression models, taking into account children’s age and gender. Using the LSMEANS option, we studied mean BMI values (with 95% confidence intervals (CI), Fig. 1b) while accounting for gender and age and mean screen time values (with 95% CI, Fig. 1c) for each physical activity category.
Correlation between total screen use, physical activity and body mass index (BMI). (a) Correlation between total screen use and BMI (line represents the linear fit of the correlation with the 95% confidence intervals), (b) Mean (95%CI) BMI in function of the physical activity level category, adjusted for age and gender of the child, (c) Mean (95%CI) total screen time in function of the physical activity level. *p < 0.05.
We evaluated the possible associations between screen time, BMI and physical activity and salivary extracellular miR-222 and miR-146a expression using the MIXED procedure to account for the hierarchical structure of the data (i.e. repeated measures for the miRNAs). This implies correlation between measurements within the same child, but no correlation between different children. The child identifier was included as a random effect nested within the schools in the mixed model. Restricted maximum likelihood estimation (REML) with unstructured autocorrelation was employed to estimate the coefficients and standard errors. A priori selected covariates were added to the model as fixed effects to correct for possible confounding. In models evaluating the associations between screen use/BMI/physical activity and miRNA expression as a dependent variable, the following variables were chosen: age of the child (continuous), gender, maternal education (two categories: up to high school diploma/college or university diploma), passive smoking (yes/no) and the extracellular RNA concentration. Effect estimates of significant covariates were presented based on the model that evaluated the effect of screen time. The effect estimates are represented as a percentage change in extracellular miRNA expression. Q-Q plots of the residuals were checked to test the assumptions of the models. Due to variability and heterogeneity in gene expression profiles, it is difficult to distinguish true gene expression alterations associated with the outcome from passenger signals18. To study the robustness, we applied Monte Carlo simulation (10,000 times) for adjustment of the parameter estimates of the significant predictors of miRNA expression using the PLM procedure. Finally, we evaluated the prediction of the models using receiver-operating characteristics (ROC) plots. The children were stratified according to their reported screen time use with the 75th percentile as a cut-off point (19.5 hours per week). The ROC plots of the models are provided (Supplementary information).
Results
The characteristics of the study population are reported in Table 1. The number of boys and girls included in the study were approximately equal. The average (standard deviation) age of the children was 10 (1) years and BMI averaged 17 (2.4) kg/m². Of the 80 children, 14 (17.5%) were underweight and 10 (12.5%) were overweight. 9 children were exposed to tobacco smoke by parental indoor smoking. The average TV screen use was 9.3 (5.5) hours per week, and average PC screen use was 4.8 (3.7) hours per week. The TV screen use was not correlated with the PC screen use (Pearson correlation coefficient r = 0.11; p = 0.34). Total screen use, defined by the sum of TV and PC use, is on average 14.11 (6.93) hours per week.
Both before (Fig. 1a) and after adjustment of BMI for child’s age and gender, total screen use was positively correlated with children’s BMI. Each one hour increment of screen use per week was associated with a 0.08 kg/m² higher BMI (95% CI: 0.004–0.163; p = 0.04), independent of children’s age and gender. Compared with children without out-of-school physical activities, children with a high activity level have a 2.37 kg/m² lower BMI (95% CI: −4.56–0.17; p = 0.0.4, overall p = 0.16) (Fig. 1b). Physical activity levels were not associated with the total screen time use (overall p = 0.36; Fig. 1c).
The association between total screen time use, BMI and physical activity on salivary extracellular miR-146a and miR-222 expression was evaluated using a mixed model, which took into account the repeated measurements of the miRNAs for each child, while accounting for gender, age, passive smoking, maternal education and the RNA content of saliva (Table 2).
Total weekly screen time use was significantly associated with salivary extracellular miR-146a and miR-222 expression. One hour increase in weekly screen time was associated with 3.44% higher (95% CI: 1.34–5.58; p = 0.002) extracellular miR-222 levels in saliva. One hour increase in weekly screen time was associated with 1.84% higher (95% CI: −0.04–3.75; p = 0.055) levels of salivary extracellular miR-146a. Monte Carlo adjusted confidence intervals and p-values for the associations of the miRNAs with screen time are confirmative and given in Table 3. No significant association between either BMI or physical activity and miR-222 and miR-146a was observed.
In addition to screen time use, salivary extracellular miR-222 was significantly higher in children exposed to passive smoking. On average, passive smoking was associated with 55% higher (95% CI: 1–139; p = 0.044) miR-222 levels in the extracellular fraction of the saliva. Age of the child was inversely associated with both miR-222 and miR-146a. A 1 year increase in age was associated with a 16.11% lower (95% CI: −27.15 to −3.41; p = 0.01) saliva extracellular level of miR-222 and a 15.86% lower (95% CI: −25.95 to −4.37; p = 0.01) saliva extracellular level of miR-146a.
Discussion
Already from childhood onwards, sedentary behaviour is associated with adverse health outcomes19. From both intervention and longitudinal studies it is apparent that watching TV is related to overweight and obesity in children19. A meta-analysis including 170 studies in children, indicated a reduction in BMI with a reduction in the sedentary behaviour19. Due to their established role in developmental processes miRNAs have emerged as attractive candidates to explore the impact of exposures during critical windows of susceptibility7.
In this context the key finding of our current paper is that children’s weekly screen time use is positively associated with BMI and this parallels differential profiles of salivary extracellular miR-146a and miR-222 expression.
The relevance of these miRNAs in the framework of physical (in)activity becomes clear from studies in adults on acute exercise20,21,22 and fitness23, obesity24, atherosclerosis25, and diabetes26,27,28,29,30. For example, after acute exercise a transient up-regulation in circulating levels of miR-146a and miR-222 is observed20,22. Furthermore, this upregulation is also observed short-term after a training session in athletes on a sustained training schedule20. miR-222 can be indicative of a subject’s fitness level based on its positive association with the maximal oxygen uptake (VO2max) in healthy adults23. Compared to normal weight subjects, circulating miR-146a31 and miR-222 levels24 can be alternatively expressed in obese subjects. Many studies report altered expression levels of miR-146a in type 2 diabetes patients26,27,28,29,30,32. The functional significance of changes in the expression of these miRNAs in the extracellular fraction of saliva remains to be elucidated. Extracellular miRNAs could be involved in cell communication, but the process is currently poorly understood. Extracellular miRNAs are promoted as promising biomarkers8, also for aspects related to lifestyle and diet33. Alterations in the (extracellular) miRNA abundance can reflect a dynamic response to metabolic stress in an attempt to re-establish homeostasis or a continuing response involved in disease development.
The findings of this study should be interpreted in the context of its limitations and strengths. As with other studies investigating the effect of screen time use or physical activity, the use of self-reported quantities of screen time and sport activities might give some exposure misclassification. In our study, physical activity was based on the reported out-of-school sport activities. It is important to note, that apart from the out-of-school sport activities other leisure activities contribute to the total physical activity level of the child. Future studies could benefit from the implementation of activity trackers and diaries, to measure physical inactivity in a more objective manner. Second, we only quantified two miRNAs, which is insufficient to draw conclusions on biological mechanisms contributing to the early onset of adult disease. Finally, observational studies cannot prove the utility of a biomarker. Nevertheless, our proof-of-concept findings indicate changes in extracellular miRNAs in relation to childhood life-style factors that are particularly relevant in the development of obesity. miR-222 and miR-146a have been implicated in cardiovascular health outcomes in adults. Our findings add to this by highlighting a correlation between these miRNAs and lifestyle in healthy children, suggesting that their expression profiles might be linked to cardiometabolic processes, already from early-life onwards. Therefore there is a possible role of these miRNAs in translating the impact of sedentary behaviour on our genome, which should be further validated in intervention studies. Gene expression can be highly heterogeneous and this hampers capturing true alterations in gene expression associated with health outcomes or lifestyle factors instead of passenger signals reflecting the heterogeneity18. We confirmed the robustness of our findings by employing a statistical resampling technique. A strength of our study is that we explored a non-invasive matrix by use of saliva which is preferable in population based studies including children.
Conclusion
To conclude, we showed expression of two extracellular miRNAs in saliva is associated with screen time use in children. The alterations in the salivary miRNA signature reflects epigenetic alterations related to an increased sedentary behaviour in school-aged children, which is particularly relevant since these miRNAs are associated with fitness and metabolic health in adults. Further research on this topic is warranted to understand the mechanism underlying the effects early-life inactivity on epigenetic alterations.
References
Ng, M. et al. Global, regional, and national prevalence of overweight and obesity in children and adults during 1980–2013: a systematic analysis for the Global Burden of Disease Study 2013. Lancet 384, 766–781, https://doi.org/10.1016/S0140-6736(14)60460-8 (2014).
Must, A. et al. The disease burden associated with overweight and obesity. Jama 282, 1523–1529 (1999).
Barker, D. J. The developmental origins of adult disease. Journal of the American College of Nutrition 23, 588S–595S (2004).
Hill, J. O., Wyatt, H. R. & Peters, J. C. Energy balance and obesity. Circulation 126, 126–132, https://doi.org/10.1161/CIRCULATIONAHA.111.087213 (2012).
Matthews, C. E. et al. Amount of time spent in sedentary behaviors in the United States, 2003–2004. American journal of epidemiology 167, 875–881, https://doi.org/10.1093/aje/kwm390 (2008).
Mitchell, J. A. & Byun, W. Sedentary Behavior and Health Outcomes in Children and Adolescents. American Journal of Lifestyle Medicine 8, 173–199, https://doi.org/10.1177/1559827613498700 (2014).
Vrijens, K., Bollati, V. & Nawrot, T. S. MicroRNAs as potential signatures of environmental exposure or effect: a systematic review. Environmental health perspectives 123, 399–411, https://doi.org/10.1289/ehp.1408459 (2015).
Etheridge, A., Lee, I., Hood, L., Galas, D. & Wang, K. Extracellular microRNA: a new source of biomarkers. Mutation research 717, 85–90, https://doi.org/10.1016/j.mrfmmm.2011.03.004 (2011).
Parrizas, M. & Novials, A. Circulating microRNAs as biomarkers for metabolic disease. Best practice & research. Clinical endocrinology & metabolism 30, 591–601, https://doi.org/10.1016/j.beem.2016.08.001 (2016).
Ding, S., Huang, H., Xu, Y., Zhu, H. & Zhong, C. MiR-222 in Cardiovascular Diseases: Physiology and Pathology. BioMed research international 2017, 4962426, https://doi.org/10.1155/2017/4962426 (2017).
Vriens, A. et al. Recent exposure to ultrafine particles in school children alters miR-222 expression in the extracellular fraction of saliva. Environmental health: a global access science source 15, 80, https://doi.org/10.1186/s12940-016-0162-8 (2016).
Saenen, N. D. et al. Recent versus chronic exposure to particulate matter air pollution in association with neurobehavioral performance in a panel study of primary schoolchildren. Environment international, https://doi.org/10.1016/j.envint.2016.07.014 (2016).
Provost, E. B. et al. Recent versus chronic fine particulate air pollution exposure as determinant of the retinal microvasculature in school children. Environmental research 159, 103–110, https://doi.org/10.1016/j.envres.2017.07.027 (2017).
World Medical Association Declaration of Helsinki: ethical principles for medical research involving human subjects. Jama 284, 3043–3045 (2000).
Cole, T. J. & Lobstein, T. Extended international (IOTF) body mass index cut-offs for thinness, overweight and obesity. Pediatric obesity 7, 284–294, https://doi.org/10.1111/j.2047-6310.2012.00064.x (2012).
Thery, C., Amigorena, S., Raposo, G. & Clayton, A. Isolation and characterization of exosomes from cell culture supernatants and biological fluids. Current protocols in cell biology Chapter 3, Unit3 22, https://doi.org/10.1002/0471143030.cb0322s30 (2006).
Van Deun, J. et al. The impact of disparate isolation methods for extracellular vesicles on downstream RNA profiling. Journal of extracellular vesicles 3, https://doi.org/10.3402/jev.v3.24858 (2014).
Li, J. et al. Identification of high-quality cancer prognostic markers and metastasis network modules. Nature communications 1, 34, https://doi.org/10.1038/ncomms1033 (2010).
Tremblay, M. S. et al. Systematic review of sedentary behaviour and health indicators in school-aged children and youth. The international journal of behavioral nutrition and physical activity 8, 98, https://doi.org/10.1186/1479-5868-8-98 (2011).
Baggish, A. L. et al. Dynamic regulation of circulating microRNA during acute exhaustive exercise and sustained aerobic exercise training. The Journal of physiology 589, 3983–3994, https://doi.org/10.1113/jphysiol.2011.213363 (2011).
Liu, X. et al. miR-222 is necessary for exercise-induced cardiac growth and protects against pathological cardiac remodeling. Cell metabolism 21, 584–595, https://doi.org/10.1016/j.cmet.2015.02.014 (2015).
Baggish, A. L. et al. Rapid upregulation and clearance of distinct circulating microRNAs after prolonged aerobic exercise. Journal of applied physiology 116, 522–531, https://doi.org/10.1152/japplphysiol.01141.2013 (2014).
Bye, A. et al. Circulating microRNAs and aerobic fitness–the HUNT-Study. PloS one 8, e57496, https://doi.org/10.1371/journal.pone.0057496 (2013).
Hulsmans, M., De Keyzer, D. & Holvoet, P. MicroRNAs regulating oxidative stress and inflammation in relation to obesity and atherosclerosis. FASEB journal: official publication of the Federation of American Societies for Experimental Biology 25, 2515–2527, https://doi.org/10.1096/fj.11-181149 (2011).
Feinberg, M. W. & Moore, K. J. MicroRNA Regulation of Atherosclerosis. Circulation research 118, 703–720, https://doi.org/10.1161/CIRCRESAHA.115.306300 (2016).
Kong, L. et al. Significance of serum microRNAs in pre-diabetes and newly diagnosed type 2 diabetes: a clinical study. Acta diabetologica 48, 61–69, https://doi.org/10.1007/s00592-010-0226-0 (2011).
Baldeon, R. L. et al. Decreased serum level of miR-146a as sign of chronic inflammation in type 2 diabetic patients. PloS one 9, e115209, https://doi.org/10.1371/journal.pone.0115209 (2014).
Rong, Y. et al. Increased microRNA-146a levels in plasma of patients with newly diagnosed type 2 diabetes mellitus. PloS one 8, e73272, https://doi.org/10.1371/journal.pone.0073272 (2013).
Lovis, P. et al. Alterations in microRNA expression contribute to fatty acid-induced pancreatic beta-cell dysfunction. Diabetes 57, 2728–2736, https://doi.org/10.2337/db07-1252 (2008).
Zhong, X. et al. The MicroRNAs in the Pathogenesis of Metabolic Memory. Endocrinology 156, 3157–3168, https://doi.org/10.1210/en.2015-1063 (2015).
Mehta, R. et al. Circulating miRNA in patients with non-alcoholic fatty liver disease and coronary artery disease. BMJ open gastroenterology 3, e000096, https://doi.org/10.1136/bmjgast-2016-000096 (2016).
Nesca, V. et al. Identification of particular groups of microRNAs that positively or negatively impact on beta cell function in obese models of type 2 diabetes. Diabetologia 56, 2203–2212, https://doi.org/10.1007/s00125-013-2993-y (2013).
Rome, S. Use of miRNAs in biofluids as biomarkers in dietary and lifestyle intervention studies. Genes & nutrition 10, 483, https://doi.org/10.1007/s12263-015-0483-1 (2015).
Acknowledgements
The authors thank Michal Kicinski for his contributions during the COGNAC study. The COGNAC study is supported by grants from the European Research Council (ERC-2012-StG310898) and the Flemish Scientific Fund (FWO, 1516112 N/G.0873.11.N.10). AV has a PhD fellowship from Bijzonder Onderzoeksfonds (BOF) Hasselt University. KV has a postdoctoral fellowship of the Research Foundation – Flanders (FWO), 12D7718N. Datasets generated during the study can be obtained through the corresponding author upon reasonable request.
Author information
Authors and Affiliations
Contributions
T.S.N. initiated the COGNAC study and managed funding. E.B.P., N.D.S. and K.V. coordinated and conducted the fieldwork. T.S.N., M.P. and P.D.B. designed the experimental setup. A.V. performed the experiments, analysed the data and wrote the manuscript. O.D.W. contributed in the interpretation of the extracellular miRNAs. All authors revised the first draft of the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Vriens, A., Provost, E.B., Saenen, N.D. et al. Children’s screen time alters the expression of saliva extracellular miR-222 and miR-146a. Sci Rep 8, 8209 (2018). https://doi.org/10.1038/s41598-018-26351-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-26351-2
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.