Abstract
Nucleophosmin (NPM1) is a multifunctional phosphoprotein which plays important roles in diverse biological processes. NPM1 can form homo- or hetero-oligomers through its N-terminal region, and bind DNA and RNA through its C-terminal region. However, the monomer-oligomer distribution of NPM1, and the extent of NPM1 binding and unbinding to RNA in living cells, are not fully understood. In this work, we analysed molecular complexes of NPM1 using size exclusion chromatography. We found that a substantial fraction of NPM1 behaves as an oligomer in HeLa cells. Furthermore, we identified three distinct oligomeric states of NPM1 using molecular characterization techniques such as subcellular localization and RNA binding. Finally, we found that heterozygous expression of a leukemia-associated NPM1 mutant significantly decreases the RNA binding level. Our data demonstrate that size exclusion chromatography provides a powerful tool for analysing NPM1 oligomers.
Similar content being viewed by others
Introduction
Nucleophosmin (NPM1, also known as B23, NO38, or numatrin) was originally identified as a highly phosphorylated protein in the nucleolus1. NPM1 also localizes to the nucleoplasm, and shuttles between the nucleus and the cytoplasm2,3. NPM1 is involved in various biological processes such as ribosome biogenesis4,5,6,7, centrosome duplication3, genome instability8 and apoptosis9,10.
NPM1 forms homo- or hetero-oligomers through its N-terminal region11,12. Disruption of NPM1 oligomers by RNA aptamers results in mislocalization of NPM1 and apoptosis in cancer cells, indicating the importance of NPM1 oligomers13. The C-terminal region of NPM1 comprises two functional domains. One is the nucleic acid-binding domain14 that reportedly binds the G-quadruplex structure of rDNA15, the promoter region of the superoxide dismutase 2 (SOD2) gene16, and 28S, 5.8S, and 5S rRNA in living cells17,18. The second C-terminal domain is a nucleolar localization signal domain19. These two domains are suggested to be structurally related20.
The overexpression of NPM1 has been observed in many types of solid tumors, including gastric21, prostate22, liver23 and colon24. Translocation in the NPM1 gene has been reported for several hematopoietic malignancies25,26; for example, t(2;5)(p23;q35) in 75% of anaplastic lymphoma kinase-positive anaplastic large cell lymphoma27, t(5;17)(q35;q31) in less than 1% of acute promyelocytic leukemia28, and t(3;5)(q25;q35) in less than 1% of acute myeloid leukemia (AML)26. Furthermore, approximately one-third of AML patients harbor frameshift mutations in exon 12 of the NPM1 gene29, resulting in the generation of a nuclear export signal in the C-terminal region of NPM1 and localization of the mutant NPM1 (NPM1c) to the cytoplasm.
Most proteins play physiological roles through temporal interactions with other molecules, or as stable molecular complexes with other proteins, possibly as homo- or hetero-oligomers. Oligomeric proteins exist in at least two different states: monomer and oligomers. A technique is therefore required for isolating monomers and oligomers in order to investigate functional differences between these states. The functions of both N-terminal and C-terminal regions of NPM1 are well-studied. In contrast, details regarding the monomer-oligomer distribution of NPM1 are not fully understood and little is known regarding the extent of NPM1 binding and unbinding to RNA in living cells.
In this study, we isolated NPM1 oligomers and monomer using size exclusion chromatography (SEC) and Western blotting. We show that a substantial fraction of NPM1 in living cells behaves as at least three distinct oligomeric states with different characteristics, such as localization and binding to RNA. Our data demonstrate that the combination of SEC and Western blotting provides a powerful tool for investigating NPM1 oligomers.
Results
Specificity of anti-NPM antibodies
In this study, we used two antibodies that recognize NPM1. One is a commercially available antibody against NPM, clone FC82291. According to the information provided by the supplier, the epitope of FC82291 lies within the 68 amino acids of the C-terminus of NPM1. The other antibody is the clone 9.2–6, which we raised previously30. To obtain more information about the epitope of NPM antibodies, we transfected an expression vector encoding NPM and its deletion mutants tagged with FHG (FLAG, HA and EGFP linked to the N-terminus) to 293T cells (Fig. 1A). As shown in Fig. 1B, the antibody FC82291 recognized NPM1 but did not recognize NPM 1.3, a splice variant of NPM1 which lacks the 37 C-terminal amino acids of NPM1, nor did it recognize NPM1c, an AML patient-related NPM1 mutant in which the 7 C-terminal amino acids of NPM1 are substituted to 11 unrelated amino acids by a frameshift mutation (Fig. 1B, lanes 2–4). Therefore, we used the commercially available antibody FC82291 as an NPM1-specific antibody. On the other hand, the antibody clone 9.2–6 recognized NPM (114–219), a central region of NPM1, as well as full length NPM1, NPM1.3 and NPM1c (Fig. 1B).
Specificity of anti-NPM antibodies. (A) Diagram of the FHG (FLAG-HA-EGFP)-NPM constructs. The amino acid sequences that constitute wild type NPM1 are represented by yellow. NPM1.3: a splice variant of NPM1, lacking the 37 C-terminal amino acids of NPM1, NPM1c: an AML patient-related NPM1 mutant. The 11 C-terminal amino acids which are substituted by frameshift mutation are represented by red. (B) 293T cells were transfected by the expression vectors described in A. After two days, the cells were lysed with SDS-sample buffer and analysed by immunoblot using commercially available anti-NPM (clone FC82291), anti-NPM which we raised previously (clone 9.2–6), or anti-EGFP.
Separation of NPM1 by size exclusion chromatography (SEC)
We first performed simple lysis and fractionation of HeLa cells using Triton X-100 (Fig. 2A). Approximately half of the total NPM1 protein was solubilized using these conditions and half remained insolubilized. This ‘insoluble NPM1’ was retained in the cell pellet following the first centrifugation and was subsequently solubilized by sonication (Fig. 2B). As shown in Fig. 2C, NPM1 localizes at both the nucleoplasm and the nucleolus, as determined by immunostaining involving permeabilization with Triton X-100 followed by cell fixation with formaldehyde (Fig. 2C, upper panel). Conversely, when cells were permeabilized using Triton X-100 prior to fixation to remove the soluble NPM1, no nucleoplasmic NPM1 was detected while nucleolar NPM1 remained (Fig. 2C, lower panel). These results suggest that biochemically fractionated ‘soluble NPM1’ and ‘insoluble NPM1’ mainly represent nucleoplasmic NPM1 and nucleolar NPM1, respectively. Therefore, we analysed soluble NPM1 and insoluble NPM1 by SEC independently. The elution profile of soluble NPM1 from asynchronous HeLa cells showed two peaks, one in fractions 15 to 16 (Group 1), and one in fractions 19 to 21 (Group 2), suggesting that there are at least two populations of nucleoplasmic NPM1 complexes (Fig. 3A). The elution profile of insoluble NPM1 was different from that of soluble NPM1. Whereas the peak in fractions 15 to 16 (Group 3) is also detected in the profile of soluble NPM1, the peak in fractions 23 to 25 (Group 4) was only observed upon fractionation of insoluble NPM1. We found that most of the RNA, which probably largely comprises rRNA, was detected as a single peak in fractions 15 to 16 of both soluble and insoluble cell lysates following SEC. The observed peaks in both the soluble and insoluble fractions coincided with the higher molecular weight peak obtained from the NPM1 fractions (Group 1 and Group 3). RNA molecules in the insoluble fractions were fragmented to some extent, probably due to sonication. Therefore, we suspected that Group 4 NPM1 was a result of sonication. To test this possibility, soluble cell lysates typically prepared without sonication in this study were used to examine the effects of sonication on the elution profiles obtained by SEC (Fig. 3B). When the soluble cell lysates were subjected to SEC without sonication, no peak was observed in fractions 23 to 25 (Fig. 3C, upper panel), whereas sonication of the soluble cell lysates resulted in a peak in fractions 23 to 25, as observed for insoluble NPM1. Therefore, insoluble Group 4 NPM1 detected in fractions 23 to 25 may be an artifact resulting from sonication. These data suggest that nucleolar NPM1 originally provides a single peak in fractions 14 to 16 upon SEC, although NPM1 may not be fully intact due to sonication.
Distribution of NPM1 by simple cell fractionation. (A) Summary of the process of cell fractionation. (B) Distribution of NPM1 in the sample prepared in A. (C) Immunofluorescence of endogenous NPM1. (Upper panel) HeLa cells were fixed with formaldehyde, then permeabilized using Triton X-100. (Lower panel) HeLa cells were permeabilized using Triton X-100, then fixed with formaldehyde. Scale bar, 10 μm.
Elution profiles of endogenous NPM1 in HeLa cells. (A) (Upper panel) Elution profiles of Triton X-100 soluble NPM1 and RNA. (Lower panel) Elution profiles of Triton X-100 insoluble NPM1 and RNA. Endogenous NPM1 was detected by immunoblot. Protein mass standards are indicated above the panel: thyroglobulin (669 kDa), ferritin (440 kDa), aldolase (158 kDa), conalbumin (75 kDa), ovalbumin (44 kDa), and Ribonuclease A (13.7 kDa). RNA was detected by ethidium bromide staining. The four NPM1 groups (Group 1 to Group 4) identified by SEC are underlined. (B) Schematic representation of sample preparation in C. (C) Elution profile of sonication-treated Triton X-100 soluble NPM1 prepared in B.
Association of NPM1 with RNA in high molecular weight fractions
Since NPM1 is an rRNA binding protein, we determined which group of NPM1 associates with RNA. We first used the commercially available monoclonal antibody FC82291. We could not detect the association of NPM1 with RNA using this NPM1 specific antibody (Supplementary Figure 1). The epitope of the antibody is located at the C-terminal end of NPM1, overlapping with the RNA-binding domain of NPM1. These results suggest that the antibody inhibits NPM1 binding to RNA in a competitive manner. Therefore, to investigate the association between NPM1 and RNA, we used the anti-NPM monoclonal antibody 9–2.6, which recognizes the central region of NPM130. As expected, RNA coprecipitated with the Group 1 and Group 3 fractions of NPM1 (Fig. 4A). Next, we investigated the extent to which NPM1 associates with RNA in fractions 15 and 16. To this end, the soluble and the insoluble cell lysates were treated with RNase A prior to analysis by SEC. As shown in Fig. 4B, after RNase A treatment, Group 1 NPM1 (fractions 15 and 16) was completely shifted to fractions corresponding to Group 2 NPM1 (fractions 19 and 20). Similarly, Group 3 NPM1 (fractions 14 to 16) also completely shifted to fractions 19 and 20. These results suggest that almost all of Group 1 and Group 3 of NPM1 associate with RNA in either a direct or indirect manner, and that the majority of cellular NPM1 elutes in fractions 18 to 20 when RNA is removed from the NPM1 complex.
Association of NPM1 with RNA in the high molecular weight fractions. (A) Triton X-100 soluble and insoluble cell lysates were subjected to SEC. The indicated fractions were pooled, then immunoprecipitated with anti-NPM. NPM1 and RNA were detected by immunoblot and ethidium bromide staining, respectively. (B) HeLa cell lysates prepared as in Fig. 2A were treated with or without RNase A, then subjected to SEC. Fractions containing the four NPM1 groups identified in Fig. 3A are underlined.
A substantial fraction of NPM1 is oligomeric in cells
A previous study demonstrated that the bacterially expressed N-terminal region of NPM forms pentamers12, and the ability of NPM1 to form oligomers has been estimated through the interaction between exogenously expressed NPM1 and endogenous NPM131. To gain insight into the oligomeric status of cellular NPM1, we introduced cDNAs for wild type NPM1 and LG mutant (L102A and G105A) NPM1 which does not form oligomers31 into 293T cells. Both cDNAs were fused to the G196 tag sequence encoding five amino acids32. When cells were lysed with Triton X-100-containing buffer, RNA was coprecipitated with G196-NPM1 (WT), but not with G196-NPM1 (LG) (Fig. 5A). In order to detect NPM1-NPM1 interaction independent of RNA, we next performed immunoprecipitation using RNase A-treated cell lysates. Whereas G196-NPM1 (WT) was coprecipitated with endogenous NPM1 in both the soluble and the insoluble cell lysates, G196-NPM1 (LG) was not coprecipitated with endogenous NPM1 (Fig. 5B), confirming the results of the previous report31. These results suggested that at least a part of the G196-NPM1 (WT) population exists as oligomers and that G196-NPM1 (LG) exclusively exists as monomers in cells.
Elution profiles of NPM1 monomer and oligomers expressed in cells. 293T cells were transfected with expression vector for G196-NPM1 (WT) or G196-NPM1 (LG). (A) The Triton X-100 soluble lysates were used for immunoprecipitation with anti-G196 antibody. G196-NPM1 and extracted RNA were detected by immunoblot and ethidium bromide staining, respectively. (B) Interaction of G196-NPM (WT) or G196-NPM1 (LG) with endogenous NPM1. The Triton X-100 soluble (S) and insoluble (I) cell lysates were treated with RNase A, then immunoprecipitated with anti-G196 antibody. (C) Elution profiles of G196-NPM1 (WT) in the Triton X-100 soluble and insoluble cell lysates. Fractions containing the four NPM1 groups identified in Fig. 3A are underlined. (D) Elution profile of G196-NPM1 (LG) in the Triton X-100-soluble cell lysates. (E) Elution profiles of G196-NPM1 (WT) in the RNase A-treated Triton X-100 soluble and insoluble cell lysates. Fractions containing the four NPM1 groups identified in Fig. 3A were underlined.
Next, we performed SEC by using cell lysates expressing G196-NPM1 (WT) or G196-NPM1 (LG) without RNase A treatment. G196-NPM1 (WT) was eluted in fractions corresponding to Group 1 and Group 3, suggesting that the majority of G196-NPM1 (WT) in both the soluble and the insoluble cell lysates associates with RNA, making it difficult to distinguish oligomers from monomers (Fig. 5C). On the other hand, monomeric G196-NPM1 (LG), which was not coprecipitated with RNA, showed a single peak in fractions 23 and 24 on SEC (Fig. 5D). To exclude the influence of RNA on the elution profile of G196-NPM1 (WT), the cell lysates were treated with RNase A prior to SEC. In the absence of RNA, G196-NPM1 (WT) was eluted in fractions 18 to 20 in both the soluble and the insoluble samples (Fig. 5E), consistent with the case observed in HeLa cells (Fig. 4B). Since G196-NPM1 (WT) forms oligomers, this result raises the possibility that oligomeric NPM1 is eluted in fractions 18 to 20 on SEC. To investigate the elution profile of purified NPM1 oligomers, we prepared bacterially expressed His-tagged NPM1 (WT) which is known to form oligomers. His-tagged NPM1 (WT) also possesses the ability to bind bacterial RNA, and thus the cell lysates were treated with RNase A. Purified His-tagged NPM1 (WT) protein showed a purity of more than 90% as determined by Coomassie Brilliant Blue (CBB) staining of the gel, and did not contain RNA (Fig. 6A). The elution profile of purified His-tagged NPM1 (WT) showed a single peak in fractions 18 to 20 (Fig. 6B), consistent with that of RNase A-treated G196-NPM1 (WT) expressed in 293T cells. To examine whether cellular NPM1 bind to other proteins with similar stoichiometry to NPM1, each group of NPM1 which was separated by SEC was immunoprecipitated either with or without RNase A treatment. We found one or two bands, which can be detected by CBB staining between 30 kDa and 46 kDa, as the major component(s) in the immunoprecipitates of all groups (Fig. 7, upper panel). Western blot analysis using anti-NPM antibody strongly suggested that these bands are NPM1 and its variants (Fig. 7, lower panel). Taken together, these results suggest that a substantial fraction of NPM1 is oligomeric in cells.
Purified recombinant NPM1 shows a similar elution profile to that of RNase A-treated cellular NPM1. (A) Bacterially expressed His-TEV-NPM1 was purified as described in experimental procedures. Purified proteins showed high purity and did not contain any RNA. (B) Elution profile of purified His-TEV-NPM1 proteins.
The main component of cellular NPM1 complex is NPM1 and its variant. Both the Triton X-100 soluble and insoluble lysates from HeLa cells were subjected to SEC. Fractions 15 and 16 from the soluble sample (for Group 1), fractions 19 to 21 from the soluble sample (for Group 2), and fractions 15 and 16 from the insoluble sample (for Group 3) were separately pooled. After treatment with or without RNase A, each pooled sample was immunoprecipitated with anti-NPM antibody. Proteins were detected by CBB staining (Upper panel) and immunoblot using anti-NPM antibody (Lower panel).
Association of NPM1 with RNA is attenuated in a NPM1c-positive cell line
Approximately one third of AML patients carry frameshift mutations in exon 12 of the NPM1 gene in one allele. NPM1c, the protein product from the mutant allele, has several amino acid substitutions near the C-terminal end of NPM1, and this area constitutes part of the RNA-binding domain. NPM1c has biological characteristics distinct from wild type NPM1, such as a defective nucleolar localization signal and the appearance of a nuclear export signal19,33. Therefore, we investigated whether the expression of NPM1c has any effect on the SEC elution profile of wild type NPM1 expressed from the normal allele. To this end, we used two AML cell lines: HL60 cells, which express wild type NPM1, and OCI-AML3 cells, a known NPM1c-positive cell line. Since HL-60 cells express only wild type NPM1, a single band of 62 nt was observed by RT-PCR (Fig. 8A, left lane). On the other hand, OCI-AML3 cells express both wild type NPM1 and NPM1c. Therefore, two bands, one of 62 nt (wild type NPM1) and one of 66 nt (NPM1c, which contains four nucleotide insertions), were detected by RT-PCR (Fig. 8A, right lane). The RNA level of NPM1c and wild type NPM1 in OCI-AML3 cells was shown to be almost equal. Western blot analysis revealed that wild type NPM1 mainly shows Triton X-100 soluble in both cell lines (Fig. 8B). On the other hand, NPM1c was detected only in the soluble fraction in OCI-AML3 cells. The expression of wild type NPM1 in OCI-AML3 cells was lower than that of HL60 cells (Fig. 8B, middle panel). Given that there is no mutation in the amino acid sequence of the N-terminal oligomerization domain in NPM1c, NPM1c is expected to form an oligomer with NPM1 (WT) in OCI-AML3 cells. Therefore, we performed an immunoprecipitation assay using the antibody FC82291 against NPM1. This antibody did not recognize NPM1c because its epitope is the C-terminus of NPM1 (WT) (Fig. 1B, upper panel, lane 4). As expected, NPM1c coprecipitated with NPM1 (WT), demonstrating that NPM1c forms an oligomer with NPM1 (WT) in OCI-AML3 cells (Fig. 8C). The elution profiles of Triton X-100 soluble and insoluble NPM1 from HL60 cells are similar to the results obtained using HeLa cells (Fig. 8D). In contrast, the elution profiles of wild type NPM1 from OCI-AML3 cells was quite different from that of HL60 cells. Triton X-100 soluble NPM1 (WT) was detected as a relatively broad peak in fractions 16 to 23 (Fig. 8E). NPM1c also eluted as a broad peak, but the peak fractions were slightly different from those of NPM1 (WT). Triton X-100 insoluble NPM1 (WT) was detected in fractions 20–22. Notably, neither peak coincided with the peak due to RNA, which mainly eluted in fractions 15 and 16. To investigate whether wild type NPM1 associates with rRNA in OCI-AML3 cells, we performed an immunoprecipitation assay using Triton X-100 soluble cell lysates. As shown in Fig. 8F, the level of RNA which coprecipitated with NPM1 in OCI-AML3 cell lysate was significantly lower compared with that of HL60 lysate (Fig. 8G). Taken together, these results indicate that most wild type NPM1 does not bind with rRNA in OCI-AML3 cells.
The association of NPM1 with RNA is attenuated in a NPM1c-positive cell line. (A) RNA was extracted from HL60 (expressing NPM1 (WT)) and OCI-AML3 (expressing NPM1 (WT) and NPM1c) cells. The expression of NPM1 (WT) and NPM1c was detected by RT-PCR. (B) Distribution of NPM1 in HL60 and OCI-AML3 cells prepared as in Fig. 2A. Then, NPM1 (WT) and NPM1c were detected by Western blotting. (C) HL60 and OCI-AML3 cells were lysed with buffer containing Triton X-100, SDS, and DOC. The lysate was immunoprecipitated with antibody against NPM1. NPM1 (WT) and NPM1c were detected by Western blotting with the indicated antibodies. (D,E) Triton X-100 soluble and insoluble lysates prepared from HL60 (D) and OCI-AML3 (E) cells were subjected to SEC. NPM1 (WT) and NPM1c were detected by Western blotting. RNA was extracted from each fraction and detected by ethidium bromide staining. (F) Interaction of NPM1 complex with RNA. The Triton X-100 soluble lysates from HL60 and OCI-AML cells were immunoprecipitated with anti-NPM antibody. NPM1 and extracted RNA were detected by immunoblot and ethidium bromide staining, respectively. (G) Quantification of the data from panel F. The normalized RNA levels coprecipitated with NPM1 are shown. The data represent the average of three independent experiments ± SEM for each condition.
Discussion
In this study, we used SEC to analyse the molecular complexes formed by NPM1. SEC can be performed under a wide range of experimental conditions such as pH, salt concentration, and detergents, depending on the purpose of the experiment. In addition, SEC also permits the use of subsequent processes, such as enzymatic reaction and immunoprecipitation. This flexibility of the SEC technique prompted us to investigate two important states of NPM1: the NPM1 oligomers, and the NPM1/rRNA complex.
We showed that a substantial fraction of NPM1 is oligomeric in HeLa cells upon SEC, indicating that the major function of NPM1 is exhibited as NPM1 oligomers. Therefore, it may be necessary to take other components of NPM1 oligomers into account in order to understand the role of NPM1 itself. Two splice variants, named NPM1.3 (UniProt P06748–3) and NPM1.2 (UniProt P06748–2), have been reported to date34. NPM1.3 lacks the 37 C-terminal amino acids of NPM1, resulting in complete loss of nucleic acid binding ability14, and NPM1.2 lacks 29 amino acids (residues 195–223) of NPM1. Although NPM1.2 retains RNA binding ability, its affinity has been reported to be lower than that of NPM118. Given that both isoforms can form oligomers with NPM1 through the conserved N-terminal oligomerization domain, the function of NPM1 oligomers may be dependent on the composition of the NPM1 isoforms.
Approximately equal levels of NPM1 monomer and oligomer in cultured cell lines were detected in a previous study35. In this study, NPM1 monomer and oligomer were isolated by ether native PAGE, SDS-PAGE without heat treatment of the protein samples, or SDS-PAGE using crosslinked protein samples. However, it is difficult to compare the levels of proteins differing significantly in molecular weight. As discussed below, we found that approximately half of the NPM1 molecules form a complex with 28S rRNA and the molecular weight of the NPM1/28S rRNA complex might be too high to migrate in a polyacrylamide gel. Furthermore, large differences in molecular weight usually lead to different transfer efficiencies of proteins from gel to membrane upon blotting; consequently, the amount of NPM1 oligomer might be underestimated compared with NPM1 monomer following native PAGE or SDS-PAGE using crosslinked protein samples. The method which combined SEC and Western blotting separated NPM1 monomer and oligomer into different fractions using non-denaturing conditions by SEC, followed by detection as monomer by Western blotting under denaturing conditions. Therefore, the result obtained using this method is likely more accurate.
We found that there are at least three groups of NPM1 oligomers (Group 1–3). Group 1 NPM1 localizes in the nucleoplasm, and forms a complex with rRNA. Group 2 NPM1 also localizes in the nucleoplasm but does not bind with rRNA. Group 3 NPM1 localizes in the nucleolus and is associated with rRNA. Given that NPM1 is a non-ribosomal protein involved in ribosome biogenesis, these three groups of NPM1 might be involved in different stages of ribosome biogenesis. On the other hand, many proteins not involved in ribosome biogenesis have been reported to bind with NPM136. NPM1 complex which contains these binding proteins may also be included in Group 2 at a lower level, since the major component of NPM complex is NPM1 and its variants. We also found that nucleolar-localized Group 4 NPM1, which was solubilized by sonication of the cell lysate, showed similar peak fractions to that of monomeric NPM1 (LG). However, we cannot exclude the possibility that Group 4 NPM1 is due to sonication of the sample, since monomer-like NPM1 was observed when nucleoplasmic NPM1 oligomer was subjected to additional sonication. Further studies are thus required.
The elution peaks of G196-NPM1 monomer and oligomer isolated by SEC from cells corresponded to masses of approximately 158 kDa (Fig. 5D) and 500 kDa (Fig. 5E), respectively, and deviate from the calculated molecular weight of NPM1. Similar conflicting results were obtained using bacterially expressed NPM1 (Fig. 6B). These discrepancies between observed and predicted molecular weight might result from the structure of NPM1. The N-terminal domain of NPM1 is in a highly ordered conformation to form oligomers12,37, the central region is suggested to be disordered18, and the C-terminal region of NPM1 consists of three-helix bundles20. Therefore, the structures of NPM1 monomer and oligomers might result in an elution profile different from that of a typical globular protein with a molecular weight similar to NPM1.
The AML-associated mutation of NPM1 occurs in a heterogeneous manner and therefore both NPM1 (WT) and NPM1c express in OCI-AML3 cells. Recent studies have demonstrated that NPM1c loses its binding ability to the G-quadruplex structure of DNA in vitro15 and gains the nuclear export signal (NES), which is a well-known difference between NPM1 (WT) and NPM1c. These results suggest that NPM1c also fails to bind RNA. In support of this idea, we found that the elution profile of NPM1c does not coincide with that of RNA. Surprisingly, the elution profile of NPM1 (WT) also did not coincide with that of RNA. Indeed, the amount of rRNA coprecipitated with NPM1 was greatly attenuated in OCI-AML3 cells. These results indicate that ribosomes expressed in OCI-AML3 cells do not require interaction between NPM1 and rRNA during maturation. These abnormalities might be related to the pathology of NPM1c-positive AML. The reason why even NPM1 (WT) failed to bind rRNA in OCI-AML3 cells could be due to two factors related to NPM1 oligomers. The first factor is an NES signal in NPM1c. NPM1c has been reported to mainly localize to the cytosol. When NPM1 (WT) forms an oligomer with NPM1c, the oligomer might localize to the cytosol, losing the opportunity to bind rRNA in the nucleus where ribosome biogenesis takes place. The second factor is the ratio of NPM1c to NPM1 (WT) in the NPM1 oligomer. A previous study reported that NPM1 oligomer exhibited decreased binding ability to rRNA when the ratio of NPM1.3 to NPM1 (WT) was increased in an in vitro reconstitution assay38. Based on this observation, increasing the ratio of NPM1c in NPM1 oligomers might result in decreased binding ability of NPM1 oligomers to rRNA.
Our study indicates that it is important to reveal the functional role of NPM1 oligomers, and that SEC provides a powerful tool for analysing NPM1 oligomeric states.
Methods
Antibodies
Mouse monoclonal antibodies against NPM1 (clone FC82291, B0556, Sigma-Aldrich, St. Louis, MO, USA), α-tubulin (T6199, Sigma-Aldrich), β-actin (A5441, Sigma-Aldrich) and rabbit polyclonal antibodies against GFP (60–011, BioAcademia, Osaka, Japan) and histone H3 (#9715, Cell Signaling, Danvers, MA, USA), were purchased commercially. Mouse monoclonal antibodies against NPM (clone 9–2.6)30 and G19632 were described previously. Rabbit polyclonal antibody against NPM1c was raised using a synthetic peptide corresponding to the C-terminal sequence of NPM1c type A (DLCLAVEEVSLRK). Horseradish peroxidase-conjugated goat F(ab’)2 anti-mouse IgG (H+L) (#710–133, Rockland Immunochemicals, Limerick, ME, USA), goat anti-rabbit IgG (111–035–003, Jackson ImmunoResearch Laboratories, West Grove, PA, USA) and Alexa 594-conjugated goat anti-mouse IgG (H+L) (A-11032, ThermoFisher Scientific, Waltham, MA, USA) were purchased commercially.
Plasmid
All primers used in this study are listed in Table S1. The cDNA for FLAG-HA-EGFP (FHG) was generated using pEGFP-C1 vector (Takara Bio, Shiga, Japan) as the template. The PCR product was digested with BglII and EcoRI and subcloned into a pBSK-BglII vector, which is a modified version of pBluescript II SK (+) vector (Agilent Technologies, Santa Clara, CA, USA) and harbors the BglII recognition sequence in its multiple cloning site. To construct the FHG expressing vector, the BglII-EcoRI fragment of FHG cDNA digested from FHG/pBSK-BglII vector was inserted into the BamHI-EcoRI sites of pcDNA3.1 (+) vector (ThermoFisher Scientific). To construct G196/pcDNA3 vector, an annealed fragment of the G196 oligonucleotides was inserted into pcDNA3.1 (+) vector which was digested with HindIII and XhoI.
The cDNA sequences of NPM1 (WT), NPM1.3 and NPM1c were PCR amplified using oligo(dT)-primed first-strand cDNAs from the HeLa and OCI-AML3 cell lines. The cDNA for NPM deletion mutants was amplified by PCR using the cDNA for NPM1 (WT) as template. The cDNA for NPM1 (LG) mutant was generated by overlap PCR using the internal primers in addition to NPM1 sense primer and NPM1 antisense primer.
The BamHI-EcoRI fragment of cDNA for NPM was inserted into FHG/pcDNA3.1 or G196/pcDNA3.1 and into pET28-TEV, a modified pET-28a (Novagen, Madison, WI, USA) with a hexa-histidine-tag and TEV cleavage site at the amino-terminus. The sequences of all the PCR products were confirmed.
Cell culture, transfection
OCI-AML3 cells were obtained from DSMZ (ACC 582, Braunschweig, Germany). An ecotropic HeLa strain which expresses mCAT (a receptor for murine retrovirus), and HEK293T cells, were maintained in Dulbecco’s modified Eagle medium (Nissui, Tokyo, Japan) supplemented with 10% fetal bovine serum (Sigma-Aldrich), penicillin and streptomycin (Nacalai Tesque, Kyoto, Japan) in a 37 °C incubator with a humidified atmosphere of 5% CO2. HL60 and OCI-AML3 were grown in RPMI1640 medium supplemented with 10% fetal bovine serum, penicillin and streptomycin in a 37 °C incubator with a humidified atmosphere of 5% CO2. Transfection was performed using Lipofectamine 2000 (ThermoFisher Scientific).
Preparation of cell lysat
To investigate the epitopes of NPM antibody, 293T cells exogenously expressing NPM were lysed with SDS-sample buffer by heating 100 °C for 3 min. For fractionation of the HeLa and 293T cells, cells were lysed with PBS containing 1% Triton X-100. For fractionation of HL60 and OCI-AML3 cells, cells were lysed with PBS containing 1% Triton X-100, 15 mM imidazole, 1 mM DTT and 40 U/ml RNase inhibitor (Takara Bio). After incubation on ice for 10 min, samples were centrifuged at 2,100 × g for 5 min at 4 °C, then the supernatant was used as the ‘Triton X-100 soluble fraction’. The resulting pellet was rinsed with lysis buffer and resuspended in new lysis buffer. The suspension was sonicated using a Handy Sonic (TOMY SEICO, Tokyo, Japan) at a power control setting of 5 for 20 sec. After centrifugation at 20,000 × g for 5 min at 4 °C, the supernatant was used as the ‘Triton X-100 insoluble fraction’. For RNase A treatment, cell lysates were treated with 40 µg/ml RNase A (Sigma-Aldrich) for 15 min at 37 °C. After centrifugation at 20,000 × g for 10 min, the supernatant was used for SEC. For interaction of NPM1 with NPM1c in OCI-AML3 cells, cells were lysed with PBS containing 1% Triton X-100, 0.1% SDS, 0.5% DOC and 15 mM imidazole.
Size exclusion chromatography
SEC was performed using an AKTA FPLC system (GE Healthcare, Buckinghamshire, England). A Superdex 200 Increase 10/300 GL column (GE Healthcare) was equilibrated with cell lysis buffer. The column was calibrated using a gel filtration calibration kit (GE Healthcare). Each standard protein was dissolved in PBS containing 1% Triton X-100 and chromatographed on the column separately. The filtered protein samples were then fractionated on the column (0.75 ml/min; 0.5 ml/fraction).
Immunoprecipitation and immunoblotting
Immunoprecipitation was performed using approximately 2 µg of anti-NPM, anti-G196 and anti-NPM1 for 1~3 hours. The antibodies used for immunoprecipitation were pre-conjugated with Protein G Sepharose 4 Fast Flow (GE Healthcare). After SDS-PAGE on a 12% acrylamide gel, the proteins were transferred onto an Immobilon-P membrane (Sigma-Aldrich). The membrane was blocked with 5% skim milk, then incubated with the appropriate antibodies. Blots were washed with PBS-T, and developed using an ECL-Plus system (PerkinElmer, Waltham, MA, USA). For quantitative analysis, images were acquired using ImageQuant LAS 4000 (GE Healthcare) and quantified using Multi Gauge software (Fujifilm, Tokyo, Japan).
CBB staining of immunoprecipitated NPM1 complex
Approximately 1.4 × 108 HeLa cells were fractionated as described above, and 8 ml of soluble cell lysate and 8 ml of insoluble cell lysate were prepared. Each cell lysate was separated by SEC 4 times. After SEC, fractions 15 and 16 from the soluble samples (for Group 1), fractions 19 to 21 from the soluble samples (for Group 2), and fractions 15 and 16 from the insoluble samples (for Group 3) were separately combined to provide 3 pools. Each pooled fraction was divided into four tubes for treatment with or without 40 µg/ml RNase A for 15 min at 37 °C. After centrifugation at 20,000 × g for 10 min, the supernatant was used for immunoprecipitation by anti-G196 (for negative control) or anti-NPM 9.2 antibody cross-linked to Protein G Sepharose 4 Fast Flow. Immunoprecipitates were eluted with PBS containing 1% Triton X-100 and 2 M guanidine-HCl (Nacalai Tesque). Samples were then precipitated using PAGE Clean Up Kit (Nacalai Tesque). After drying, 25 μl of SDS sample buffer was added to each sample, then heated at 100 °C for 3 min. NPM1 complex was detected in 15 μl samples by CBB staining, and 3 μl samples were used to detect NPM1 complex by Western blotting.
Immunofluorescence
HeLa cells were grown on a glass cover coated with Type I-C Cellmatrix (Nitta Gelatin, Osaka, Japan). For the modified standard protocol, the cells were washed with PBS and fixed with PBS containing 3.7% formaldehyde for 20 min. The cells were then washed with PBS and permeabilized with PBS containing 1% Triton-X100 for 2 min. After washing with PBS, the cells were incubated with PBS containing 5% skim milk for 20 min. To remove Triton X-100 soluble proteins prior to fixation, the cells were incubated in PBS containing 1% Triton X-100 for 2 min on ice, washed twice with PBS, and then fixed with 3.7% formaldehyde for 20 min. After washing with PBS, the cells were blocked with PBS containing 5% skim milk for 20 min. Next, the cells were incubated with blocking buffer containing anti-NPM1 antibody at room temperature for 1 h. The cells were then washed with PBS and incubated with Alexa Fluor 594 goat anti-mouse IgG (1:1000) at room temperature for 20 min. After washing with PBS, the cells were mounted on glass slides using vectashield mounting medium with DAPI (Vector Laboratories, Burlingame, CA, USA). Images were acquired using confocal microscope (FV1000, Olympus, Tokyo, Japan).
RNA extraction
For RNA extraction, 500 µl of Sepasol-RNA I Super G (Nacalai Tesque) was added to 50 µl of cell lysate, fractionated sample, or immunoprecipitated sample. For RT-PCR, 1000 µl of Sepasol-RNA I Super G was added to 2 × 106 HL60 or OCI-AML3 cells. After 5 min, 100~200 µl of chloroform was added, and the mixture was centrifuged at 20,000 × g for 5 min at 4 °C. RNA extracted into the aqueous layer was precipitated by adding an equal volume of isopropanol. After rinsing with 70% ethanol, the RNA was dried and dissolved in loading buffer (43% formamide, 0.013% SDS, 0.013% bromophenol blue, 0.013% xylene cyanol, 0.013% ethidium bromide and 0.25 mM EDTA) and then treated at 70 °C for 10 min.
Recombinant protein
B834 (DE3) Escherichia coli transformed with NPM1/pET28-TEV vector was grown in LB medium supplemented with kanamycin (Nacalai Tesque) at 37 °C for overnight. E. coli suspension (500 µl) was added to 4.5 ml of fresh LB medium and cultured at 37 °C for 1 hour. Protein expression was induced with 1mM isopropyl-β-D-1-thiogalactopyranoside (IPTG). After 2 hours’ culture, the E. coli cells were centrifuged at 20,000 × g for 1 min at 4 °C, then suspended in lysis buffer (PBS, 1% Triton X-100 and 15 mM imidazole). The suspension was sonicated at a power control setting of 10 for 20 sec. After centrifugation at 20,000 × g for 10 min at 4 °C, the supernatant was used for affinity purification using Ni-NTA agarose (QIAGEN, Valencia, CA, USA). After incubation at 4 °C for 1 hour, the beads were washed with lysis buffer, then treated with 40 µg/ml RNase A in lysis buffer for 15 min at 37 °C. After washing with lysis buffer, proteins were eluted using elution buffer (PBS, 1% Triton X-100 and 250 mM imidazole).
RT-PCR analysis
Five µg of total RNA was reverse-transcribed using oligo (dT) primer and a ReverTra Ace system (TOYOBO, Osaka, Japan). The cDNAs for NPM1 (WT) and NPM1c were amplified with 25 cycles of PCR. Gel electrophoresis was performed with a 10% urea-PAGE gel using the TBE buffer system.
References
Olson, M. O., Orrick, L. R., Jones, C. & Busch, H. Phosphorylation of acid-soluble nucleolar proteins of Novikoff hepatoma ascites cells in vivo. J. Biol. Chem. 249, 2823–2827 (1974).
Borer, R. A., Lehner, C. F., Eppenberger, H. M. & Nigg, E. A. Major nucleolar proteins shuttle between nucleus and cytoplasm. Cell 56, 379–390 (1989).
Wang, W., Budhu, A., Forgues, M. & Wang, X. W. Temporal and spatial control of nucleophosmin by the Ran-Crm1 complex in centrosome duplication. Nat. Cell Biol. 7, 823–830 (2005).
Herrera, J. E., Savkur, R. & Olson, M. O. The ribonuclease activity of nucleolar protein B23. Nucleic Acids Res. 23, 3974–3979 (1995).
Murano, K., Okuwaki, M., Hisaoka, M. & Nagata, K. Transcription regulation of the rRNA gene by a multifunctional nucleolar protein, B23/nucleophosmin, through its histone chaperone activity. Mol. Cell. Biol. 28, 3114–3126 (2008).
Savkur, R. S. & Olson, M. O. Preferential cleavage in pre-ribosomal RNA byprotein B23 endoribonuclease. Nucleic Acids Res. 26, 4508–4515 (1998).
Maggi, L. B. Jr. et al. Nucleophosmin serves as a rate-limiting nuclear export chaperone for the Mammalian ribosome. Mol. Cell. Biol. 28, 7050–7065 (2008).
Grisendi, S. et al. Role of nucleophosmin in embryonic development and tumorigenesis. Nature 437, 147–153 (2005).
Gjerset, R. A. DNA damage, p14ARF, nucleophosmin (NPM/B23), and cancer. J. Mol. Histol. 37, 239–251 (2006).
Li, Z. & Hann, S. R. The Myc-nucleophosmin-ARF network: a complex web unveiled. Cell cycle (Georgetown, Tex.) 8, 2703–2707 (2009).
Yung, B. Y. & Chan, P. K. Identification and characterization of a hexameric form of nucleolar phosphoprotein B23. Biochim. Biophys. Acta 925, 74–82 (1987).
Namboodiri, V. M., Akey, I. V., Schmidt-Zachmann, M. S., Head, J. F. & Akey, C. W. The structure and function of Xenopus NO38-core, a histone chaperone in the nucleolus. Structure 12, 2149–2160 (2004).
Jian, Y. et al. RNA aptamers interfering with nucleophosmin oligomerization induce apoptosis of cancer cells. Oncogene 28, 4201–4211 (2009).
Wang, D., Baumann, A., Szebeni, A. & Olson, M. O. The nucleic acid binding activity of nucleolar protein B23.1 resides in its carboxyl-terminal end. J. Biol. Chem. 269, 30994–30998 (1994).
Chiarella, S. et al. Nucleophosmin mutations alter its nucleolar localization by impairing G-quadruplex binding at ribosomal DNA. Nucleic Acids Res. 41, 3228–3239 (2013).
Xu, Y. et al. The role of a single-stranded nucleotide loop in transcriptional regulation of the human sod2 gene. J. Biol. Chem. 282, 15981–15994 (2007).
Bertwistle, D., Sugimoto, M. & Sherr, C. J. Physical and functional interactions of the Arf tumor suppressor protein with nucleophosmin/B23. Mol. Cell. Biol. 24, 985–996 (2004).
Hisaoka, M., Nagata, K. & Okuwaki, M. Intrinsically disordered regions of nucleophosmin/B23 regulate its RNA binding activity through their inter- and intra-molecular association. Nucleic Acids Res. 42, 1180–1195 (2014).
Nishimura, Y., Ohkubo, T., Furuichi, Y. & Umekawa, H. Tryptophans 286 and 288 in the C-terminal region of protein B23.1 are important for its nucleolar localization. Biosci. Biotechnol. Biochem. 66, 2239–2242 (2002).
Grummitt, C. G., Townsley, F. M., Johnson, C. M., Warren, A. J. & Bycroft, M. Structural consequences of nucleophosmin mutations in acute myeloid leukemia. J. Biol. Chem. 283, 23326–23332 (2008).
Tanaka, M., Sasaki, H., Kino, I., Sugimura, T. & Terada, M. Genes preferentially expressed in embryo stomach are predominantly expressed in gastric cancer. Cancer Res. 52, 3372–3377 (1992).
Léotoing, L. et al. Influence of nucleophosmin/B23 on DNA binding and transcriptional activity of the androgen receptor in prostate cancer cell. Oncogene 27, 2858–2867 (2008).
Yun, J. P. et al. Increased expression of nucleophosmin/B23 in hepatocellular carcinoma and correlation with clinicopathological parameters. Br. J. Cancer 96, 477–484 (2007).
Nozawa, Y., Van Belzen, N., Van der Made, A. C., Dinjens, W. N. & Bosman, F. T. Expression of nucleophosmin/B23 in normal and neoplastic colorectal mucosa. J. Pathol. 178, 48–52 (1996).
Naoe, T., Suzuki, T., Kiyoi, H. & Urano, T. Nucleophosmin: a versatile molecule associated with hematological malignancies. Cancer Sci. 97, 963–969 (2006).
Falini, B. et al. Translocations and mutations involving the nucleophosmin (NPM1) gene in lymphomas and leukemias. Haematologica 92, 519–532 (2007).
Ardini, E., Magnaghi, P., Orsini, P., Galvani, A. & Menichincheri, M. Anaplastic Lymphoma Kinase: role in specific tumours, and development of small molecule inhibitors for cancer therapy. Cancer Lett. 299, 81–94 (2010).
Yan, W. & Zhang, G. Molecular Characteristics and Clinical Significance of 12 Fusion Genes in Acute Promyelocytic Leukemia: A Systematic Review. Acta Haematol. 136, 1–15 (2016).
Falini, B. et al. Cytoplasmic nucleophosmin in acute myelogenous leukemia with a normal karyotype. N. Engl. J. Med. 352, 254–266 (2005).
Ishikawa, Y. et al. Abnormal cytoplasmic dyslocalisation and/or reduction of nucleophosmin protein level rarely occurs in myelodysplastic syndromes. Leuk. Lymphoma 49, 2359–2364 (2008).
Enomoto, T., Lindstrom, M. S., Jin, A., Ke, H. & Zhang, Y. Essential role of the B23/NPM core domain in regulating ARF binding and B23 stability. J. Biol. Chem. 281, 18463–18472 (2006).
Tatsumi, K. et al. G196 epitope tag system: a novel monoclonal antibody, G196, recognizes the small, soluble peptide DLVPR with high affinity. Sci. Rep. 7, 43480, https://doi.org/10.1038/srep43480 (2017).
Falini, B. et al. Both carboxy-terminus NES motif and mutated tryptophan(s) are crucial for aberrant nuclear export of nucleophosmin leukemic mutants in NPMc+ AML. Blood 107, 4514–4523 (2006).
Di Matteo, A. et al. Molecules that target nucleophosmin for cancer treatment: an update. Oncotarget 7, 44821–44840 (2016).
Vladimirova, N. M., Lobanova, N. V. & Potapenko, N. A. State of oncomarker protein B23/nucleophosmin in HeLa cells. Biochemistry (Mosc.) 75, 851–860 (2010).
Lindström, M. S. NPM1/B23: A Multifunctional Chaperone in Ribosome Biogenesis and Chromatin Remodeling. Biochemistry research international 2011, 195209 (2011).
Lee, H. H. et al. Crystal structure of human nucleophosmin-core reveals plasticity of the pentamer-pentamer interface. Proteins 69, 672–678 (2007).
Okuwaki, M., Tsujimoto, M. & Nagata, K. The RNA binding activity of a ribosome biogenesis factor, nucleophosmin/B23, is modulated by phosphorylation with a cell cycle-dependent kinase and by association with its subtype. Mol. Biol. Cell 13, 2016–2030 (2002).
Acknowledgements
We thank Dr. Toshio Kitamura (The University of Tokyo) for providing the ecotropic HeLa cells. We acknowledge the technical expertise of the Centre for Integrated Research in Science at Shimane University. This work was supported by Ministry of Education, Culture, Sports, Science and Technology Grants-in-Aid for Young Scientists (B) (KAKENHI) JP20790257 and JP23790367 (to G.S.) and the SUIGAN Project 1b01 (to T.U.), Shimane University.
Author information
Authors and Affiliations
Contributions
G.S. and T.U. conceived and coordinated the study. G.S. performed most of the biochemical and immunofluorescence experiments. H.K. and T.N. provided key reagents and technical advice. G.S. and T.U. analysed the results and wrote the paper. All authors analysed the results and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sakashita, G., Kiyoi, H., Naoe, T. et al. Analysis of the oligomeric states of nucleophosmin using size exclusion chromatography. Sci Rep 8, 4008 (2018). https://doi.org/10.1038/s41598-018-22359-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-22359-w
This article is cited by
-
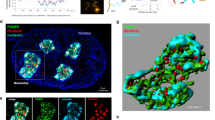
Nucleophosmin 1 cooperates with the methyltransferase DOT1L to preserve peri-nucleolar heterochromatin organization by regulating H3K27me3 levels and DNA repeats expression
Epigenetics & Chromatin (2023)
-
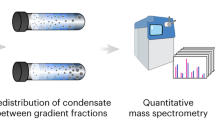
Advances in mass spectrometry to unravel the structure and function of protein condensates
Nature Protocols (2023)
-
Mechanisms of nucleophosmin (NPM)-mediated regulated cell death elucidated by Hsp70 during renal ischemia
Apoptosis (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.