Abstract
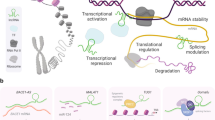
Alteration in RNA metabolism, concerning both coding and long non-coding RNAs (lncRNAs), may play an important role in Amyotrophic Lateral Sclerosis (ALS) pathogenesis. In this work, we performed a whole transcriptome RNA-seq analysis to investigate the regulation of non-coding and coding RNAs in Sporadic ALS patients (SALS), mutated ALS patients (FUS, TARDBP and SOD1) and matched controls in Peripheral Blood Mononuclear Cells (PBMC). Selected transcripts were validated in spinal cord tissues. A total of 293 differentially expressed (DE) lncRNAs was found in SALS patients, whereas a limited amount of lncRNAs was deregulated in mutated patients. A total of 87 mRNAs was differentially expressed in SALS patients; affected genes showed an association with transcription regulation, immunity and apoptosis pathways. Taken together our data highlighted the importance of extending the knowledge on transcriptomic molecular alterations and on the significance of regulatory lncRNAs classes in the understanding of ALS disease. Our data brought the light on the importance of lncRNAs and mRNAs regulation in central and peripheral systems, offering starting points for new investigations about pathogenic mechanism involved in ALS disease.
Similar content being viewed by others
Introduction
There is mounting evidence that altered RNA metabolism, both involving coding and non-coding RNAs (ncRNAs), plays an important role in Amyotrophic Lateral Sclerosis (ALS) pathogenesis. ALS is an adult-onset, progressive and fatal neurodegenerative disease, caused by the selective loss of both upper and lower motor neurons in the cerebral cortex, brainstem and spinal cord. The pathogenesis of the disease is still unknown. Alterations of transcription regulation may represent key events in ALS pathogenesis, supported by mapping of genetic mutations in TDP-43 and FUS/TLS genes coding for DNA/RNA binding proteins involved in transcription and RNA processing1. The same notion is strengthened by the observation that SOD1, Alsin and ANG are involved in transcription and processing of both coding and non-coding RNAs2 and by the finding that mutant SOD1 induces alternative splicing deregulation1.
About RNAs, there is a clear evidence of the importance of non-coding RNAs in central nervous system (CNS) functions and their involvement in neurodegenerative diseases such as Parkinson’s disease, Alzheimer’s disease and ALS. In addition, while microRNA involvement in neurodegenerative disorders has been the subject of intense research3,4, the recently revealed class of long non-coding RNAs (lncRNAs) is at the beginning of its characterization. LncRNAs are RNA transcripts greater than 200 nucleotides that lack an open reading frame and therefore do not encode proteins. While coding genes are widely annotated, high-quality catalogues of lncRNAs and tissue-specific expression data are recently being constructed. Recent efforts are directed to characterize this, largely unexplored, functional component of the genome. GENCODE consortium5, within the framework of ENCODE project, with a mixed approach based on manual annotation and ENSEMBL based annotation6, has categorized lncRNAs in different biotypes, based on their location with respect to protein-coding genes7: i) antisense (AS) RNAs have transcripts that overlap the genomic span of a protein-coding locus on the opposite strand, or published evidence of antisense regulation of a coding gene; ii) lincRNAs are intergenic non-coding RNAs; iii) sense overlapping RNAs contain a coding gene in an intron on the same strand; iv) sense intronic RNAs are present in introns of a coding gene and do not overlap any exon; v) processed transcript do not contain an ORF and cannot be added to previous biotypes.
LncRNAs are mostly related to possible regulator on biogenesis, cellular cycle and differentiation8, and are involved in nervous system and neurological diseases9,10. LncRNAs can act both as epigenetic regulators of target genes and as components of an extensive, unexplored network of interacting RNAs involving miRNAs and mRNAs. The literature has put in light the role of lncRNAs in both microsatellite expansion, i.e. Huntington disease11 and neurodegenerative diseases9,12,13 (Alzheimer’s and Parkinson’s Disease). Moreover, in the case of ALS neurodegenerative disease, few data are available about lncRNAs14,15,16. So far, it has been described that Nuclear-Enriched Abundant Transcript1 (NEAT1) generates two types of long non-coding RNAs. The named NEAT1_2 lncRNA interacts with paraspeckle formation in spinal motor neurons of ALS patients14. It was indeed demonstrated that NEAT1_2 lncRNA is up-regulated in spinal cord during the early stage of ALS pathogenesis compared to healthy controls14. Moreover, in patients affected by Frontotemporal Lobar Degeneration17, TDP-43 and FUS proteins bind to and regulate the expression of different lncRNAs1, although the mechanistic connection between lncRNAs and coding RNAs is not explained, yet.
The involvement of coding RNA deregulation on ALS has been demonstrated18,19 and different papers in the past years, have described this aspect, considering involved pathways or ALS mutated genes, such as SOD118, TARDBP20. To reinforce the thesis of a whole coding RNA deregulation, recent studies, based on deep sequencing of coding RNAs both in monocytes21 and in spinal cord22 from ALS patients and matched controls, have been published.
In this paper, we present a whole transcriptome profiling of both long non-coding and coding RNAs in Peripheral Blood Mononuclear Cells of Sporadic ALS (SALS) patients and matched controls. LncRNAs data have been validated in spinal cord, as main involved tissues in ALS and the study was extended to a little group of patients with mutation in genes associated to ALS, i.e. FUS, SOD1 and TARDBP. Furthermore, we have investigated the presence of co-expression networks between coding and lncRNAs.
Results
Deep sequencing lncRNAs expression profiles in PBMC samples of ALS patients and healthy subjects
We detected differentially expressed lncRNAs (DE lncRNAs) in PBMCs in five groups of subjects: sporadic patients (SALS), FUS, TARDBP and SOD1 mutated patients and healthy controls. In SALS patients, 293 DE lncRNAs were identified: 62.5% (183 out of 293) were up-regulated (Table 1). 62.8% (184 out of 293) were reported as antisense, 27.7% (81 out of 293) as lincRNAs, while the remaining 9.5% (28 out of 293) were classified as processed transcripts or intronic sense RNAs (Table 2). Only 23 out of the 293 DE lncRNAs were described as “known” transcripts, (i.e.: represented in the HUGO Gene Nomenclature Committee database and RefSeq), while remaining 270 transcripts were reported as “novel” (i.e.: transcripts containing four or more exons and/or supported by at least one mRNA/cDNA or three ESTs but not still present in common databases) (Table 2).
Heat-maps of the differentially expressed mRNAs and lncRNAs in SALS relative to healthy controls are shown in Fig. 1A,B, respectively. Different expression profiles in SALS and healthy controls can be visibly distinguished. Considering the most deregulated lncRNAs in SALS group respect to healthy controls (Supplementary Table S1), it is evident that transcription pathway is highly involved. Three lncRNAs reported in the top 10 are described as antisense of transcription-related genes: ZEB1-AS1 is indeed antisense of ZEB1 transcription factor, ZBTB11-AS1 is the antisense of ZBTB11 gene, involved in the DNA binding and in transcriptional regulation, and also XXbac-BPG252P9.10 is described as the antisense transcript of IER3, involved in transcription. More in detail, it is known that IER3 is a transcription factor of the nuclear factor-kappa-B/rel (NF-kappa-B) family23, known to be involved in ALS22.
Expression profiles of differently expressed genes in ALS and healthy controls. In panel (A), differentially expressed mRNAs are shown, while in panel (B) differentially expressed lncRNAs are shown. All comparisons are given between the disease state and the control samples. We considered as differentially expressed only genes showing |log2(disease sample/healthy donor)| ≥ 1 and a False Discovery Rate ≤ 0.1.
In FUS mutated patients, 21 deregulated long non-coding RNAs were identified. 11 of them were antisense, 6 were reported as lincRNAs while the remaining were categorized as processed transcripts (2) or sense intronic RNAs (2) (Table 1, Table 2, Supplementary Table S2). Five out of the 21 deregulated lncRNAs were reported as “known”.
For TARDBP mutated patients, 15 DE LncRNAs were detected: 7 antisense, of which only one was already described, 6 lincRNAs, 1 processed transcript and 1 sense intron RNA (Table 1, Table 2, Supplementary Table S3).
In SOD1 mutated patients, we detected only 2 deregulated lncRNAs, both reported as novel antisense RNAs. (Table 1, Table 2, Supplementary Table S4).
Validation of deregulated processed transcripts and antisense lncRNAs
To confirm RNA-seq results, we performed Real Time PCR (qPCR) for a subset of selected lncRNAs. The lncRNAs to be validated were selected based on these criteria: i) we preferably chose the transcripts to be validated among the 10 most differentially expressed lncRNAs found in each group; ii) we preferably analysed known antisense and processed transcripts; iii) we extended validation to other transcripts to confirm both up- and down-regulated genes iv) we included transcripts deregulated in all groups (Supplementary files 1 and Supplementary Tables S1, S2, S3 and S4).
We also investigated the RNA deregulation in spinal cord, derived from a post-mortem explant in sporadic ALS patients, which is a tissue known to be involved in ALS24.
SALS patients
In SALS patients, we validated antisense lncRNAs and processed transcripts with deregulation ≥ 1 in terms of |Log2FC| (ZEB1-AS1, XXbac-BPG252P9.10 IER3-AS, ZBTB11-AS1, RP11-475I24.8, RP11-38M8.1 and ENST00000417346). Even if TTC25 and SPON1 were categorized as lncRNAs in the manually annotated GENCODE catalogue, both were described as well-known coding RNAs in RefSeq and ENSEMBL. Thus, they were no object of RT validation. Results are represented in Fig. 2, panels A, B and C.
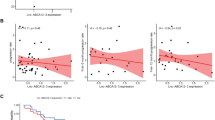
Differentially expressed transcripts verified by Real Time PCR in PBMC and Spinal Cord from larger cohort of SALS and CTRs. (A) ENST00000423714.1 (ZEB1-AS1); (B) ENST00000607333.1 (IER3-AS); (C) ENST00000536865.1 (ZBTB11-AS); (D) ENST00000458479.1 (PAXBP-AS); (C) ENST00000438646.1 (SNAP25-AS); (D) ENST00000502041.2 (CKMT2-AS). In brackets in panel titles, the group in which the transcript was found deregulated is reported.
ENST00000423714.1 (ZEB1-AS1) was the first differentially expressed AS in SALS group and it was found down-regulated compared to controls (log2FC = −2.4). It was annotated as a processed transcript, although it is the known AS of Zinc Finger E-Box Binding Homeobox 1 gene (ZEB1), acting as a transcriptional repressor by chromatin and E-box binding (Fig. 2A). ENST00000607333.1 (XXbac-BPG252P9.10) was annotated as AS of IER3, one of the transcription factors of (NF-kappa-B) family, with a crucial role in cell survival by regulation of anti-apoptotic genes. In SALS patients, XXbac-BPG252P9.10 was down regulated (log2FC = −2.35) compared to healthy subjects (Fig. 2B). ENST00000536865.1 (ZBTB11-AS1) was found down-regulated (log2FC = −2.14) in SALS patients compared to controls both in PBMC and Spinal cord (Fig. 2C). It was annotated as antisense of Zinc Finger and BTB Domain Containing 11 gene (ZBTB11), involved in DNA binding and in transcriptional regulation.
FUS mutated patients
ENST00000458479.1 (PAXBP-AS) was the only known differentially expressed AS in FUS patients. PAX3 and PAX7 Binding Protein AS was up-regulated (log2FC = 2.69) in FUS patients (Fig. 2D). GO annotations related to this gene included transcription factor activity, sequence-specific DNA binding and RNA polymerase II core promoter proximal region sequence-specific DNA binding.
TARDBP mutated patients
ENST00000438646.1 (SNAP25-AS), in the top 10 of differentially expressed lncRNAs, was the first AS found deregulated (down regulated, log2FC = −2.34) in TARDBP group of patients (Fig. 2E). This gene is involved in axonal repair and synaptic vesicle processing and it was previously reported to be deregulated in ALS patients25.
SOD1 mutated patients
ENST00000502041.2 (CKMT2-AS) was the only lncRNA deregulated (down-regulated, log2FC = −2.08) in SOD1 mutated patients (Fig. 2F). It is annotated as CKMT2 antisense RNA, which is a Mitochondrial creatine kinase (MtCK).
Pathway analysis of lncRNAs
Non-coding RNAs pathway analysis was performed using LncPath R package, where a pre-computed lncRNA-mRNA relationship network was used to evaluate the extent of each gene influenced by DE lncRNAs (https://CRAN.R-project.org/package=LncPath). We detected some pathways synergistically regulated by lncRNA sets (Table 3). Most interesting pathways referred to Mapk signalling, cytokine receptor interaction, chemokine signalling, natural killer cell mediated cytotoxicity and regulation of actin cytoskeleton.
Deep sequencing mRNAs expression profiles
We performed RNA-seq analyses also to determinate the expression profiles of mRNAs of ALS patients compared to matched healthy subjects. In SALS population, RNA-seq data showed 87 differentially expressed mRNAs, 30 of which down-regulated while 57 up-regulated (Table 1, Supplementary Table S5). Heat-map representing the expression levels of all dysregulated mRNAs in SALS and healthy subjects is represented in Fig. 1A.
In mutated ALS, PBMCs clearly showed different mRNA profiles between patient groups. We detected 122 altered genes in FUS group, 30 in TARDBP and 18 in SOD1 patients (Table 1 and Supplementary Tables S6, S7 and S8).
mRNAs sequence data analysis showed a common profile between TARDBP and FUS groups and between SALS and SOD1 groups. In fact, patients mutated in TARDBP showed a 67% of down-regulated and 33% of up-regulated mRNAs, and FUS patients showed a similar regulation, 71% of down-regulated genes and 29% of up-regulated genes. On the other hand, SOD1 and SALS groups showed a similar profiling with a major number of up-regulated genes (78% and 66% respectively). Only one gene, Two-pore channel 1 (TPCN1), has been found in common between the different groups. Interestingly, this gene is involved in Mapk signaling pathway, already identified by our lncRNAs pathway analysis (Table 3).
mRNA pathway analysis
GO terms enrichment and KEGG pathway analysis for DEGs in SALS patients compared to healthy controls has been performed for up-regulated and down-regulated DEGs, separately26 (Fig. 3, Fig. 4).
GO analysis for DE genes in SALS patients compared to healthy controls. TOP10 enriched GO terms for Biological process (A,D), cellular component (B,E) and molecular function (C,F). The length of the bar represents the significance of that specific gene-set or term. The brighter the color, the more significant that term is. Panels (A,B) and (C) have been obtained considering down-regulated genes, while panel (D,E) and (F) considering up-regulated genes.
KEGG pathway analysis for DE genes in SALS patients compared to healthy controls. TOP10 KEGG pathways enriched by deregulated genes are shown. The length of the bar represents the significance of that specific gene-set or term. The brighter the color, the more significant that term is. Panels A has been obtained considering down-regulated genes, while panel B considering up-regulated genes.
The GO biological processes enriched terms for down-regulated genes are related to apoptotic process and transcription regulation (Fig. 3A). Up-regulated genes affect humoral immune response and negative regulation of transforming growth factor beta-receptor signalling pathway (Fig. 3D). Enriched GO terms for Cellular Component include Integral Component of plasma membrane both for up- and down-regulated mRNAs (Fig. 3B,E). With respect to molecular function, the most enriched GO terms targeted by down-regulated mRNAs include transcription factor activity, NF-kappaB binding, activating transcript factor binding and DNA binding (Fig. 3C). The highest enriched GO terms targeted by up-regulated transcripts included activin binding and ubiquitin-protein transferase activity (Fig. 3F). KEGG pathways enriched by dysregulated genes include also cancer-related pathways both for up-and down-regulated mRNAs (Fig. 4A,B).
Coding/non-coding co-expression analysis
Coding non-coding RNAs co-expression network was constructed via WGNCA R package and drawn using Cytoscape software (http://www.cytoscape.org/).
Top 30 most deregulated coding and non-coding genes in SALS group respect to healthy controls were used to construct co-expression network. The three interesting subnets are shown in Fig. 5. Six clusters of co-expressed genes were observed. No antisense RNA was present in these networks. The first network involved 14 coding genes and 2 long intergenic RNAs (Fig. 5A). The second network contained lincRNAs only (Fig. 5B). Two clusters of coding genes only were than reported (Fig. 5C,D). Finally, two clusters containing lincRNAs only were represented (Fig. 5E,F).
Discussion
The importance of various classes of regulatory non-coding RNAs (ncRNAs) in different diseases is increasingly being recognized. We performed a full profiling, by RNA-Seq approach, of the lncRNAs and mRNAs in human sporadic and mutated ALS patients, and healthy controls with the aim of extending our knowledge on molecular alterations of transcriptome and obtaining new data about its regulation.
Different classes of lncRNAs have been described7 but their characteristics are often unknown.
At first, we analysed the DE lncRNAs with a specific focus on Antisense RNA (AS). AS is a class of long non-coding RNAs that plays important regulatory roles in a variety of biological processes and they are required for proper regulation of coding genes27. AS lncRNAs may control the epigenetic state of chromatin, modulating methylation of DNA and/or histones or by removing repressors27,28. They promote sense gene transcription by recruiting transcription factors to enhance or modulate splicing of sense pre-mRNA29,30. Importantly, AS transcripts may regulate the half-life of their sense partners by establishing Dicer-dependent cutting of dsRNA, potentially followed by siRNA-mediated gene silencing30.
A characterization of AS has been developed in other neurodegenerative diseases, such as Parkinson’s Disease, showing an altered regulation of S/AS expression, and functional studies showed an important effect on neuron survival31,32. Our work is the first study concerning a deep screening on both coding and non-coding RNAs in ALS patients.
Considering SALS patient’s group, in the top 10 of DE lncRNAs, our data showed an interesting AS deregulation of genes involved in transcription regulation pathway such as ZEB1-AS and ZBTB11-AS. ZEB1 may act as repressor or activator of transcription33. It may repress histones organization or activate chromatin regulators33,34. Moreover, ZEB1-AS was studied in cancer, in fact it was demonstrated that higher expression values of ZEB1-AS promote tumor metastasis35. Another AS of a transcription factor, ZBTB11-AS was found decreased in SALS patients compared to controls (Table S1). It is annotated as AS of Zinc finger and BTB domain-containing protein 11 (ZBTB11) gene. It seems to be a negative regulator of cell cycle, even if it is not well characterized. It was partially studied in hepatocellular carcinoma and it was recently described as transcriptional repressor36.
Some of the sense genes regulated by the DE AS lncRNAs in SALS are already linked to neurodegenerative disease, such as UBXN7-AS37 ATG10-AS38 and ADORA2A-AS39. In fact, UBXN7 is an ubiquitin protein bound by VCP, a known ALS protein. The regulation of UBXN7 by its AS regulated the ubiquitination in ALS disease. ATG10 was reported in the pathological pathway37 while ADORA2A is involved in neurodegenerative diseases as Huntington and Parkinson’s disorders in relation to defects in DNA methylation31. The role of DNA methylation is object of intensive studies in ALS but to date it has not been totally clarified31.
In FUS mutated patients, the most interesting data concern PAX, a fundamental for skeletal muscle development already described as involved in ALS40. In TARDBP group of patients SNAP25-AS was found DE. SNAP25 is involved in axonal repair and synaptic vesicle processing and it is deregulated in ALS patients25,41. In ALS mice model, human TDP-43 decrease the RNA levels of synaptic proteins (as SNAP25)25,41 and we cannot exclude that SNAP25-AS may be involved in this deregulation. Finally, SOD1 mutated patients showed only two DE lncRNAs, one of these annotated as CKMT2 antisense. In ALS, a mutation in SOD1, which may lead to reduced creatine kinase activity by inactivation of important target enzymes, including MtCK, was described42.
In the second part of the paper we also analysed the mRNAs expression in both non-mutated and mutated patients.
Only one gene (TPCN1) was found in common between all ALS groups. This gene has an important role in autophagy pathway43 that is known to be altered in ALS44,45. TPCN1 is down-regulated in ALS patients compared to control, suggesting that TPCN1 reduction may act on autophagy functions.
As TPCN1, the top 10 of DEG mRNAs showed a general trend of down-regulation. The impact of these genes on the transcription pathway was confirmed by GO enrichment analysis (Fig. 3A): transcription regulation is indeed the second most involved pathway in SALS patients. About this pathway, ZMYM6, TTF2 and TAF5L (Supplementary Table S5) are associated to nucleic acid binding and transcription46. This association emerges also respect to molecular function, The most enriched GO terms targeted by down-regulated mRNAs include transcription factor activity, activating transcript factor binding and DNA binding (Fig. 3C).
Furthermore, the apoptotic process is the first in the enriched GO terms for Biological process (Fig. 4A). Interestingly, about apoptosis, one of the deregulated gene detected in this work is NAIP, already described associate do ALS and SMN47 even if the role of apoptosis in ALS is still controversial48,49. Moreover, KEGG pathways include cancer-related pathways both for up-and down-regulated mRNAs (Fig. 3A,B). These data are potentially interesting because the two DE AS (ZEB1 and ZBTB11 AS) have been largely associated to cancer33,35. In addition, between the most DE genes MYCBP, the binding protein of MYC, is present. It is an important oncogene well characterized in cancer50 (Table S5). About KEGG analysis, it is worth noting the important involvement of the immune system pathways (Table 3). In fact, in the top 10 we have found i) cytokine-cytokine receptor interaction, ii) chemokine signalling pathway, iii) natural killer cell mediated cytotoxicity. We suppose that these data may be associated to the kind of samples (PBMC) that we used for RNA-seq analysis, moreover, the association between ALS and immune deregulation has been already reported51. This investigation has confirmed the importance of extending our knowledge on molecular alterations of transcriptome and the significance of the classes of regulatory long non-coding RNAs, especially antisense RNA, in ALS disease. Morever, in this work, we have investigated the possible involvement of lncRNAs only at the beginning of the disease, future studies will be focused on the correlation between the progression of the disease and the RNAs profile.
Our data brought the light on the importance of Sense and Antisense RNA regulation in central and peripheral system, offering numerous starting points for new investigations about pathogenic mechanism involved in ALS disease.
Materials and Methods
Study Subjects
30 SALS patients and 30 age- and sex-matched healthy controls (CTR) were recruited after obtaining written informed consent (Table 4). A subset of subjects (10 ALS and 3 CTR) was deep-squenced while all samples were included for Real Time PCR experiments. With this approach, deep-sequencing, although on a small number of samples, allowed to explore the whole transcriptome at the level of coding and non coding genes in ALS patients when compared to healthy controls and to select a subset of interesting transcripts, to be further studied and validated by Real Time PCR. ALS patients underwent clinical and neurologic examination at IRCCS National Neurological Institute “C. Mondino” (Pavia, Italy). All patients were diagnosed with ALS as defined by El Escorial criteria. All SALS patients were analysed to exclude any causative mutations in SOD1, TARDBP, FUS, C9orf72, ANG and VCP genes. The control subjects were recruited at the Transfusional Service and Centre of Transplantation Immunology, Foundation San Matteo, IRCCS (Pavia, Italy). Moreover, also a cohort of mutated ALS patients (2 FUS, 3 SOD1 and 2 TARDBP mutated) (Table 4) was analysed and compared to three healthy controls.
The study protocol to obtain PBMC from patients and controls was approved by the Ethical Committee of the National Neurological Institute “C. Mondino”, IRCCS (Pavia, Italy). Before being enrolled, the subjects participating in the study signed an informed consent form (Protocol n°375/04 – version 07/01/2004).
Spinal cord tissue was obtained from the Human Brain and Spinal Fluid Resource Center (VA West Los Angeles Healthcare center, Los Angeles, CA 90073), which is sponsored by NINDS/NIMH, National Multiple Sclerosis Society, and Department of Veteran Affairs. All experiments were performed in accordance with relevant guidelines and regulations.
Isolation of human peripheral blood mononuclear cells (PBMC)
Peripheral blood mononuclear cells (PBMC) were prepared by centrifugation. Peripheral blood was layered (density = 1.077) and centrifuged at 950 g for 30 min. After isolation on a Ficoll-Histopaque layer (Sigma, Italy), cell viability was assayed by a trypan blue exclusion test and the cells were used for RNA extraction.
RNA extraction
Samples were homogenized and total RNA was isolated by Trizol® reagent (Life Science Technologies, Italy) following the manufacturer’s specifications. RNAs were quantified using a Nanodrop ND-100 Spectrophotometer (Nanodrop Technologies, Wilmington, USA) and a 2100 Bioanalyzer (Agilent RNA 6000 Nano Kit, Waldbronn, Germany); RNAs with a 260:280 ratio of ≥1.5 and an RNA integrity number of ≥8 were subjected to deep sequencing.
Libraries preparation for RNA-Seq and bioinformatic data analysis
Sequencing libraries were prepared with the Illumina TruSeq Stranded RNA Library Prep, version 2, Protocol D, using 500-ng total RNA (Illumina). Qualities of sequencing libraries were assessed by 2100 Bioanalyzer with a DNA1000 assay. Libraries were quantified by qPCR using the KAPA Library Quantification kit for Illumina sequencing platforms (KAPA Biosystems); RNA processing was carried out using Illumina NextSeq. 500 Sequencing. FastQ files were generated via llumina bcl2fastq2 (Version 2.17.1.14 - http://support.illumina.com/downloads/bcl2fastq-conversion-software-v217.html) starting from raw sequencing reads produced by Illumina NextSeq sequencer. Gene and transcript intensities were computed using STAR/RSEM software52 using Gencode Release 19 (GRCh37.p13) as a reference, using the “stranded” option. Differential expression analysis for mRNA was performed using R package EBSeq53. This tool was selected because of its superior performance in identifying isoforms differential expression54.
Differential expression analysis for long non-coding RNAs was performed with the R package DESeq. 255. Coding and non coding genes were considered differentially expressed and retained for further analysis with |log2(disease sample/healthy control)| ≥ 1 and a FDR ≤ 0.1. We imposed minimum |Log2FC| of 1 and a FDR lower than 0.1 as thresholds to differentially expressed genes. This choice is motivated by the decision to maximize the sensitivity of this analysis, in order to perform a massive screening and identify candidate genes to be validated with a wider sample population with real-time analysis.
Pathway and co-expression analysis
Gene enrichment analysis was performed on coding genes26. We performed a Gene Ontology (GO) analysis for biological processes, cellular components and molecular function and a Kegg pathway analysis (Kyoto Encyclopedia of Genes and Genomes http://www.genome.ad.jp/kegg) via enrichR web tool56,57.
Non-coding RNAs pathway analysis was performed. LncPath R package was used to map differentially expressed lncRNAs on a lncRNA-mRNA relationship network, to evaluate the extent of each gene influenced by lncRNAs, based on a network diffusion strategy(https://CRAN.R-project.org/package=LncPath).
Furthermore, a weighted gene co-expression network analysis was performed to assess functional annotation using WGCNA R package58. The 30 most deregulated coding and non-coding genes (in terms of log2FC) in SALS patients compared to healthy controls were selected for this analysis. Co-expression analysis of lncRNAs with well-annotated protein-coding genes can provide an approach to investigate the biological role of lncRNAs. Coding non-coding RNAs co-expression network was constructed via WGNCA R package58. Network nodes represent gene expression profiles, while undirected edges values are the pairwise correlations between gene expressions. Cytoscape software (http://www.cytoscape.org/) was used for network import and visualization.
Real Time PCR
Using human gene sequences available from NCBI (www.ncbi.nlm.nih.gov/nucleotide), PCR oligonucleotide for sense genes pairs were selected spanning introns to optimize amplification from mRNA templates and avoiding nonspecific amplification products, using NCBI’s Primer- BLAST or online Primer 3.0. Moreover, primers were designed in specific regions that do not overlap with Antisense sequences (primers upon request). Total cDNAs were prepared from 1 to 2 μg of total RNA using SuperScript III reverse transcriptase (LifeTechnologies, SanDiego, CA). qPCR reactions included 200 nM of each oligonucleotide, 1 ul of SYBR Green SuperMix (BioRad, Richmond, CA), and 1 μL of cDNA template (or water control). Cycling conditions using a BioRadiQ5 Real-Time thermocycler were 5 min denaturing at 95 °C, followed by 40 cycles of 95 °C (10 s) and 58 °C annealing (30 s).
Real Time Data Analysis
Cycle threshold (Ct) values were automatically recorded for each replicate qPCR reaction, and mean Ct values were normalized against those determined for GAPDH. Fold-expression differences relative to healthy controls were determined using the 2ΔΔCt method. Significance of gene expression changes relative to controls was analysed using one-way ANOVA (Kruskal-Wallis) and the Dunns post-test for all possible test pairings using Prism GraphPad 3.03 software (GraphPad Software, San Diego, CA). P-values (two tailed) with 95% confidence intervals were computed, and P < 0.05 was considered statistically significant.
Data availability
The sequencing data obtained in this study were deposited in NCBI GEO [GSE106443].
References
Strong, M. J. The evidence for altered RNA metabolism in amyotrophic lateral sclerosis (ALS). J Neurol Sci 288, 1–12 (2010).
Milani, P. et al. Posttranscriptional regulation of SOD1 gene expression under oxidative stress: Potential role of ELAV proteins in sporadic ALS. Neurobiol Dis. 60, 51–60 (2013).
Figueroa-Romero, C. et al. Expression of microRNAs in human post-mortem amyotrophic lateral sclerosis spinal cords provides insight into disease mechanisms. Mol Cell Neurosci. 71, 34–45 (2016).
Chen, Y. et al. Aberration of miRNAs Expression in Leukocytes from SporadicAmyotrophic Lateral Sclerosis. Front Mol Neurosci. 9(69) (2016).
[Gencode1] Harrow, J. et al. GENCODE: the reference human genome annotation for The ENCODE Project. Genome research 22(9), 1760–1774 (2012)
[Ensembl] Aken, B. et al. The Ensembl gene annotation system. Database (2016).
[Gencode2] Derrien, T. et al. The GENCODE v7 catalog of human long noncoding RNAs: analysis of their gene structure, evolution, and expression. Genome research 22(9), 1775–1789 (2012).
Riva, P., Ratti, A. & Venturin, M. The long non-coding RNAs in neurodegenerative diseases: novel mechanisms of pathogenesis. Curr Alzheimer Res. 13(11), 1219–1231 (2016).
Beermann, J., Piccoli, M. T., Viereck, J. & Thum, T. Non-coding RNAs in Development and Disease: Background, Mechanisms, and Therapeutic Approaches. Physiol Rev. 96(4), 1297–325 (2016).
Hart, R. P. & Goff, L. A. Long noncoding RNAs: Central to nervous system development. Int J Dev Neurosci. pii: S0736–5748(16)30109–5 (2016).
Johnson, R. Long non-coding RNAs in Huntington’s disease neurodegeneration. Neurobiol Dis. 46(2), 245–54 (2012).
Soreq, L. et al. Long non-coding RNA and alternative splicing modulations in Parkinson’s leukocytes identified by RNA sequencing. PLoS Comput Biol. 10(5), e1003647 (2014).
Zhou, X. & Xu, J. Identification of Alzheimer’s disease-associated long noncoding RNAs. Neurobiol Aging. 36(11), 2925–31 (2015).
Nishimoto, Y. The long non-coding RNA nuclear-enriched abundant transcript 1_2 induces paraspeckle formation in the motor neuron during the early phase of amyotrophic lateral sclerosis. Mol Brain. 6(31) (2013).
Gagliardi, S., Milani, P., Sardone, V., Pansarasa, O. & Cereda, C. From Transcriptome to Noncoding RNAs: Implications in ALS Mechanism. Neurol Res Int. 2012, 278725 (2012).
Wan, P., Su, W. & Zhuo, Y. The Role of Long Noncoding RNAs in Neurodegenerative Diseases. Mol Neurobiol. 54(3), 2012–2021 (2017).
Lourenco, G. F., Janitz, M., Huang, Y. & Halliday, G. M. Long noncoding RNAs in TDP-43 and FUS/TLS-related frontotemporal lobar degeneration (FTLD). Neurobiol Dis. 82, 445–54 (2015).
Gagliardi, S. et al. SOD1 mRNA expression in sporadic amyotrophic lateral sclerosis. Neurobiol Dis. 39(2), 198–203 (2010).
Gagliardi, S. et al. Regulation of FMO and PON detoxication systems in ALS human tissues. Neurotox Res. 23(4), 370–7 (2013).
Mougeot, J. L., Li, Z., Price, A. E., Wright, F. A. & Brooks, B. R. Microarray analysis of peripheral blood lymphocytes from ALS patients and the SAFE detection of the KEGG ALS pathway. BMC Med Genomics. 4(74) (2011).
Zhao, W. et al. Characterization of Gene Expression Phenotype in Amyotrophic Lateral Sclerosis Monocytes. JAMA Neurol. 74(6), 677–685 (2017).
Brohawn, D., David, G., Laura, C., O’Brien, L. & Bennett, J. RNAseq Analyses Identify Tumor Necrosis Factor-Mediated Inflammation as a Major Abnormality in ALS Spinal Cord. Ed. Cedric Raoul. PLoS ONE 11(8), e0160520 (2016).
Pietzsch, A., Buchler, C. & Schmitz, G. Genomic organization, promoter cloning, and chromosomal localization of the Dif-2 gene. Biochem. Biophys. Res. Commun. 245, 651–657 (1998).
Bede, P. et al. Spinal cord markers in ALS: diagnostic and biomarker considerations. Amyotroph Lateral Scler. 13(5), 407–15 (2012).
D’Erchia, A. M., Gallo, A., Manzari, C. et al. Massive transcriptome sequencing of human spinal cord tissues provides new insights into motor neuron degeneration in ALS. Scientific Reports. 7, 10046 (2017).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA 102, 15545–15550 (2005).
Pelechano, V. & Steinmetz, L. M. Gene regulation by antisense transcription. Nat Rev Genet. 14(12), 880–93 (2013).
Wu, H., Yang, L. & Chen, L. L. The Diversity of Long Noncoding RNAs and Their Generation. Trends Genet. 33(8), 540–552 (2017).
Clark, Brian S. & Blackshaw, S. Long Non-Coding RNA-Dependent Transcriptional Regulation in Neuronal Development and Disease. Frontiers in Genetics 5, 164 (2014).
Faghihi, M. A. & Wahlestedt, C. Regulatory roles of natural antisense transcripts. Nature reviews Molecular cell biology. 10(9), 637–643 (2009).
Majidinia, M. et al. The roles of non-coding RNAs in Parkinson’s disease. Mol Biol Rep. 43(11), 1193–1204 (2016).
Carrieri, C. et al. Expression analysis of the long non-coding RNA antisense to Uchl1 (AS Uchl1) during dopaminergic cells’ differentiation in vitro and in neurochemical models of Parkinson’s disease. Front Cell Neurosci. 9, 114 (2015).
Li, T. et al. Upregulation of long noncoding RNA ZEB1-AS1 promotes tumor metastasis and predicts poor prognosis in hepatocellular carcinoma. Oncogene. 35(12), 1575–84 (2016).
Shi, Y. et al. Coordinated Histone Modifications Mediated by a CtBP Co-Repressor Complex. Nature. 422, 735–738 (2003).
Liu, Y. et al. Identification of differential expression of genes in hepatocellular carcinoma by suppression subtractive hybridization combined cDNA microarray. Oncol Rep. 18(4), 943–51 (2016).
Keightley, M. C. et al. The Pu.1 target gene Zbtb11 regulates neutrophil development through its integrase-like HHCC zinc finger. Nat Commun. 8(14911) (2017).
Guo, X. & Qi, X. VCP cooperates with UBXD1 to degrade mitochondrial outer membrane protein MCL1 in model of Huntington’s disease. Biochim Biophys Acta. 1863(2), 552–559 (2017).
Lee, J. et al. Role of autophagy in the pathogenesis of amyotrophic lateral sclerosis, Biochimica et Biophysica Acta (BBA) - Molecular Basis of Disease 1852(11), 2517–2524, ISSN 0925–4439 (2015).
Villar-Menéndez, I. et al. Increased striatal adenosine A2A receptor levels is an early event in Parkinson’s disease-related pathology and it is potentially regulated by miR-34b. Neurobiol Dis. 69, 206–14 (2014).
Tsitkanou, S., Della Gatta, P. A. & Russell, A. P. Skeletal Muscle Satellite Cells, Mitochondria, and MicroRNAs: Their Involvement in the Pathogenesis of ALS. Front Physiol. 7(403) (2016).
Xu, Y.-F. et al. The Pathological Phenotypes of Human TDP-43 Transgenic Mouse Models Are Independent of Downregulation of Mouse Tdp-43. PLoS One. 8(7), e69864 (2013).
Feijóo-Bandín, S. et al. Two-pore channels (TPCs): Novel voltage-gated ion channels with pleiotropic functions. Channels 11(1), 20–33 (2017).
Wendt, S. Reduced creatine kinase activity in transgenic amyotrophic lateral sclerosis mice. Free Radic Biol Med. 32(9), 920–6 (2002).
Ganassi, M. et al. Surveillance Function of the HSPB8-BAG3-HSP70 Chaperone Complex Ensures Stress. Granule Integrity and Dynamism. Mol Cell. 63(5), 796–810 (2016).
Rusmini, P. et al. The Role of the Heat Shock Protein B8 (HSPB8) in Motoneuron Diseases. Front Mol Neurosci. 10(176) (2017).
Jiang, Y., Liu, M., Spencer, C. A. & Price, D. H. Involvement of transcription termination factor 2 in mitotic repression of transcription elongation. Mol Cell. 14(3), 375–85 (2004).
Orrell, R. W., Habgood, J. J., de Belleroche, J. S. & Lane, R. J. The relationship of spinal muscular atrophy to motor neuron disease: investigation of SMN and NAIP gene deletions in sporadic and familial ALS. J Neurol Sci. 145(1), 55–61 (1997).
Sathasivam, S., Ince, P. G. & Shaw, P. J. Apoptosis in amyotrophic lateral sclerosis: a review of the evidence. Neuropathology and Applied Neurobiology 27, 257–274 (2001).
Guégan, C. & Przedborski, S. Programmed cell death in amyotrophic lateral sclerosis. Journal of Clinical Investigation. 111(2), 153–161 (2003).
Kim, T. et al. MYC-repressed long noncoding RNAs antagonize MYC-induced cell proliferation and cell cycle progression. Oncotarget. 6(22), 18780–9 (2015).
Cereda, C. et al. TNF and sTNFR1/2 plasma levels in ALS patients. J Neuroimmunol. 194(1–2), 123–31 (2008).
Li, B. & Dewey, C. N. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC bioinformatics 12, 323 (2011).
Leng, N. et al. EBSeq: an empirical Bayes hierarchical model for inference in RNA-seq experiments. Bioinformatics 29(8), 1035–1043 (2013).
Carrara, M. et al. Alternative splicing detection workflow needs a careful combination of sample prep and bioinformatics analysis. BMC bioinformatics 16(Suppl 9), S2 (2015).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq 2. Genome biology 15(12), 550 (2014).
Kuleshov, M. V. et al. Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Research gkw377 (2016).
Chen, E. Y. et al. Enrichr: interactive and collaborative HTML5 gene list enrichment analysis tool. BMC Bioinformatics. 128(14) (2013).
Langfelder, P. & Horvath, S. WGCNA: an R package for weighted correlation network analysis. BMC bioinformatics 9(1), 559 (2008).
Acknowledgements
This work was supported by grants from the Italian Agency for Research on ALS–ARiSLA (Pilot project “LNCinALS” 2015–2016 and Full project “GRANULOPATHY” 2015–2017). This work was also supported by “Fondazione Regionale per la Ricerca Biomedica” (FRRB 2015–0023) and Italian Ministry of Health (RC2016-2017) .
Author information
Authors and Affiliations
Contributions
S.G. and S.Z. wrote the manuscript. S.G. and C.P. performed the experiments. S.Z., M.A. and M.O. performed bioinformatic analysis. L.D. and M.C. participated to patients and controls recruitment. M.B. participated to experimental plan. D.S. and O.P. reviewed the manuscript. R.C. and C.C. supervised this work. All authors reviewed and accepted the final version of this manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Gagliardi, S., Zucca, S., Pandini, C. et al. Long non-coding and coding RNAs characterization in Peripheral Blood Mononuclear Cells and Spinal Cord from Amyotrophic Lateral Sclerosis patients. Sci Rep 8, 2378 (2018). https://doi.org/10.1038/s41598-018-20679-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-20679-5
This article is cited by
-
The Long Non-Coding RNA NR3C2-8:1 Promotes p53-Mediated Apoptosis through the miR-129-5p/USP10 Axis in Amyotrophic Lateral Sclerosis
Molecular Neurobiology (2024)
-
Lnc-HIBADH-4 Regulates Autophagy-Lysosome Pathway in Amyotrophic Lateral Sclerosis by Targeting Cathepsin D
Molecular Neurobiology (2024)
-
Preliminary insights into RNA in CSF of pediatric SMA patients after 6 months of nusinersen
Biology Direct (2023)
-
Non-coding RNAs in human health and disease: potential function as biomarkers and therapeutic targets
Functional & Integrative Genomics (2023)
-
The expression discrepancy and characteristics of long non-coding RNAs in peripheral blood leukocytes from amyotrophic lateral sclerosis patients
Molecular Neurobiology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.