Abstract
Oral squamous cell carcinomas are a major cause of morbidity and mortality, and tobacco usage, alcohol consumption, and poor oral hygiene are established risk factors. To date, no large-scale case-control studies have considered the effects of these risk factors on the composition of the oral microbiome, nor microbial community associations with oral cancer. We compared the composition, diversity, and function of the oral microbiomes of 121 oral cancer patients to 242 age- and gender-matched controls using a metagenomic multivariate analysis pipeline. Significant shifts in composition and function of the oral microbiome were observed with poor oral hygiene, tobacco smoking, and oral cancer. Specifically, we observed dramatically altered community composition and function after tooth loss, with smaller alterations in current tobacco smokers, increased production of antioxidants in individuals with periodontitis, and significantly decreased glutamate metabolism metal transport in oral cancer patients. Although the alterations in the oral microbiome of oral cancer patients were significant, they were of substantially lower effect size relative to microbiome shifts after tooth loss. Alterations following tooth loss, itself a major risk factor for oral cancer, are likely a result of severe ecological disruption due to habitat loss but may also contribute to the development of the disease.
Similar content being viewed by others
Introduction
Head and neck squamous cell carcinomas are a major cause of cancer morbidity and mortality, with an estimated incidence of 549,000 cases worldwide in 20081. The majority of these are oral cancers arising in the oral cavity and oropharynx, for which tobacco usage, betel chewing, alcohol consumption, and human papillomavirus (HPV) infection are established risk factors2,3,4,5,6,7,8,9. Case-control studies have also reported associations between oral cancer and measures of chronic poor oral hygiene (e.g. loose or missing teeth, infrequent tooth brushing or dental visits)5,10,11,12,13,14,15,16,17,18,19,20, even among non-smokers and non-drinkers21,22. When combined with tobacco or alcohol use, poor oral hygiene acts synergistically to increase the risk associated with either exposure alone21,22 and leads to chronic infection and inflammation, both of which are increasingly recognized in the pathogenesis of cancer23,24,25 and as factors in carcinogenic feedback loops incorporating the resident microbiota. Although a few recent studies characterized the interactions of these epidemiological risk factors for oral cancer with the microbiome, they were limited by sample size and different experimental approaches26,27,28,29.
Apropos, human microbiome studies have recently characterized the structure and function of the microbial communities in different regions of the human body during health30 and disease states, including the oral cavity31,32,33,34,35. Indeed, specific microbial communities are associated with periodontitis36,37,38 and dental caries39,40,41. Preliminary small studies have also found different microbial communities in samples collected from the surface of oral cancers and normal tissues matched from the same subject42,43,44. A sample of 15 oral cancers, for example, were enriched for Firmicutes and Actinobacteria relative to matched samples45 and a single array-based case-control study reported elevated counts of a few bacteria in oral cancer (e.g. Capnocytophaga gingivalis, Provatella melaningtoenica, and Streptococcus mitis)46. Larger-scale epidemiologically designed cohorts are needed to verify these structural associations, however, and especially to provide well-powered investigations of host-microbe-environment interactions in oral cancer.
In this study, we thus compared the oral microbiome in 121 oral cancer cases and 242 matched controls using multivariate analysis targeted at microbial taxonomic and inferred functional profiles. We evaluated associations between lifestyle factors (alcohol and tobacco use), health characteristics (history of periodontitis, tooth status), case-control status, and the composition and function of oral microbial communities (Table 1). Our results revealed strong shifts in composition and function associated with tooth status, with more modest shifts during tobacco smoking and oral cancer, both while additionally incorporating environmental factors into the analysis. Microbial community effects included a loss of ecological diversity and increased glutamate metabolism in tobacco smoking individuals, disruptions in both structure and function in individuals after tooth loss, and a decrease in glutamate metabolism and metal transport in cancer cases. Other lifestyle and health characteristics, including alcohol consumption, history of periodontitis, and tumor HPV status, were associated only with minor microbial effects in this population. We conclude that alterations in the structure, diversity, and function of the oral microbiome occur in association with established risk factors for oral cancer, especially after complete tooth loss; these alterations may contribute to disease development but are likely due to severe ecological disruption and habitat loss.
Results
Study population and sample characteristics
Oral rinse samples were collected for microbiome analysis from 121 cases diagnosed with oral cavity (n = 43), oropharynx (n = 64), or unknown primary (n = 5) squamous cell carcinoma (Table 1). A total of 242 controls were age (5-year intervals) and gender matched to cases at a 2:1 ratio within six months of case enrollment. The cohort was predominantly male, with median age 58 years (IQR 53–66). Cases were more likely than controls to be current tobacco users or cigarette smokers. Complete tooth loss and a history of periodontitis were also more common among cases than controls.
The structure and function of the oral microbiome is associated with oral cancer status
A comparison of the oral microbiome in cancer cases and controls showed changes both in structure (16 significantly differentially abundant clades) and in function (25 metabolic pathways, Supplementary Datasets 4 and 5 and Fig. 1). Tests were performed using a multivariate generalized linear model incorporating case/control status, tooth loss, tumor HPV status (for cases), periodontal disease, tobacco usage, alcohol consumption, and other covariates, with microbial abundances as outcome and FDR-corrected significance at q < 0.25; see Methods. Taxa belonging to the genus Dialister occurred at higher relative abundances in cases, whereas the orders Actinomycetales and Lactobacillales were significantly under-represented in oral cancer (Fig. 1a). No significant differences in microbial presence/absence were observed between cases and controls, in contrast to these changes in specific clades’ relative abundances.
Oral microbial community taxa and functional pathways differentially abundant in cancer. (a) Taxa (genera and OTUs) and (b) pathways differentially abundant in oral cancer microbiomes as determined by a multivariate model incorporating case/control status, tumor HPV status, tooth loss, periodontal disease, and other demographic and clinical covariates (see Methods). Differences are significant at FDR q < 0.25, and n = 121 cases, 242 controls (see Table 1, Supplementary Datasets 4 and 5).
Analysis of predicted functional modules indicated that genes involved in synthesis of lipopolysaccharides (KEGG47 M00063: CMP-KDO biosynthesis and M00064: ADP-L-glycero-D-manno-heptose biosynthesis) were more abundant in cases than in controls (Fig. 1b). Pathways involved in LPS transport (e.g. M00250: Lipopolysaccharide transport) decreased, possibly reflecting a shift to a higher proportion of a distinct group of Gram-negative bacteria, e.g. a shift to higher levels of Dialister. Other functional modules, in particular the relative abundance of genes involved in amino acid transport and synthesis were altered; specifically an increase in serine synthesis (M00020: Serine biosynthesis, glycerate-3P => serine) was detected, while glutamate uptake (M00233: Glutamate transport), synthesis of threonine from aspartate (M00018: Threonine biosynthesis, aspartate => homoserine => threonine), and synthesis of ornithine from glutamate (M00028: Ornithine biosynthesis, glutamate => ornithine) were at a lower relative abundance in oral cancer subjects. As these pathways are generally involved in oxidative energy harvest from TCA cycle products, particularly in combination with reduced LPS transport, this may reflect a shift toward anaerobic microbes and/or metabolism in the oral cancer microenvironment.
Community functional dysbioses during oral cancer were also reflected in a predicted increase in the synthesis and transport of vitamins and cofactors involved in metabolism, e.g. biotin and thiamine (M00123: Biotin biosynthesis, pimeloyl-ACP/CoA => biotin, M00127: Thiamine biosynthesis, AIR => thiamine-P/thiamine-2P). There was also a shift in iron uptake systems (M00190: Iron(III) transport, M00243: Manganese/iron transport), iron (M00243: Manganese/iron transport, M00248: ABC transporter), zinc, and manganese transport (M00244: Putative zinc/manganese transport). This suite of depleted metal transporters, primarily iron, may also reflect the need for fewer of these cofactors in a less-oxidative metabolic environment. Lastly, a decrease in transport systems for osmoprotectants (M00209: Osmoprotectant transport) was predicted, which may reflect a community response to changes in the osmolarity of saliva. Epithelial cell death and the resultant release of cell constituents, including amino acids, may be higher in cancer patients, hence synthesis of osmoprotectants may not be necessary.
Lower microbial diversity and loss of function after tooth loss
We observed a dramatic decrease in a broad range of clades in association with total tooth loss (i.e. no natural teeth remaining), comprising 122 clades (from Actinomycetaceae, Corynebacterium, Rothia, Prevotella, Flavobacteriales, Streptococcaceae, Fusobacteriales, and Proteobacteria; see Supplementary Dataset 4, Fig. 2). This represented the largest single association of the microbiome with clinical, demographic, or environmental factors in this cohort (Fig. 3). The striking difference between even one versus no natural teeth remaining likely corresponds to a loss of habitat, since the epithelium-tooth interface is a key site for bacterial colonization in the oral cavity. Individuals with total tooth loss had less diverse oral microbiomes that were less similar to each other than in individuals with good oral hygiene, revealing a significant microbial community profile shift associated with complete tooth loss (Fig. 4).
Microbial clades and pathways associated with oral cancer and with clinical, environmental, and demographic covariates. (a) Clades (tree includes all microbes in the dataset) and (b) pathways (shown on the KEGG BRITE hierarchy) differentially abundant with respect to oral cancer (highlighted points) and a subset of non-cancer covariates in this cohort (outer bands), comprising periodontal health, tumor HPV status, tobacco usage, and tooth status; multivariate model is as in Fig. 1 and Methods. Highlighted associations are significant at FDR q < 0.25, n as in Table 1.
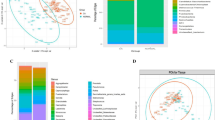
Covariation of microbial community beta-diversity with non-cancer covariates including tooth loss, periodontal health, tumor HPV status, and tobacco usage. Ordination by non-parametric multidimensional scaling of samples’ Canberra dissimilarities, with oral/oropharyngeal cancer status indicated by shape and color stratified by (a) tooth loss status, (b) HPV positivity, (c) periodontal health, and (d) tobacco usage. Complete tooth loss represents the largest determinant of variability in the cohort’s oral microbial communities, with smaller effects of cancer case/control status and other covariates.
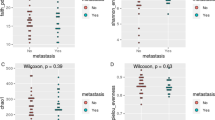
Oral cancer and tooth loss significantly affect microbial community alpha- and beta-diversity. (a) Within-sample inverse Simpson alpha-diversity across all samples, and (b) between-sample Bray-Curtis dissimilarity between all pairs of samples within each phenotype. Stars indicate significant differences by Wilcoxon rank sum test (binary oral cancer status) or for Cuzick’s trend test (ordinal tooth loss status), both at p < 0.05.
The broad shift in community composition was also reflected in a dramatic shift in function associated with tooth loss. Twenty-two metabolic pathways (Supplementary Dataset 5), including transport systems for sugars (M00215: D-Xylose transport, M00216: Multiple sugar transport, M00217: D-Allose transport, M00198: Putative sn-glycerol-phosphate transport), metals (M00245: Cobalt/nickel transport, M00186: Tungstate transport), and organic ions, such as glycine, betaine, and proline (M00208: Glycine betaine/proline transport), as well as modules involved in the synthesis of polyamines, including arginine, agmatine, ornithine, putrescine, and spermidine (M00133: Polyamine biosynthesis, arginine => agmatine => putrescine => spermidine, M00134: Polyamine biosynthesis, arginine => ornithine => putrescine) were lower in abundance in individuals with tooth loss. Other modules at lower abundance included those for the GABA gamma aminobutyrate shunt, which channels glutamate into the TCA cycle (M00027: GABA (gamma-Aminobutyrate) shunt). In contrast, many pathways including the synthesis of threonine, asparate, and homoserine (M00018: Threonine biosynthesis, aspartate => homoserine => threonine) and phosphoribosyl pyrophosphate (M00005: PRPP biosynthesis, ribose 5P => PRPP), which is involved in the synthesis of nucleotides (purines and pyrimidines), NAD, and the amino acids histidine and tryptophan, occurred at higher abundance among subjects with no remaining natural teeth. A gene level analysis of the Threonine biosynthesis pathway (M00018) revealed that most genes were altered in their relative abundance between individuals having at least some of their natural teeth and individuals with complete tooth loss (Fig. 5). Additionally, some of the genes from this pathway, e.g. K12525 (metL), K12526 (lysAC), which are interchangeable within the process from L-Aspartate into L-4-Aspartylphosphate are specific to Gram-negative bacteria. Overall, complete tooth loss results in a significant change in the structure and function of the oral microbiome.
Gene-level analysis of the serine biosynthesis, cysteine biosynthesis, and threonine biosynthesis pathways. The results of our study showed significant changes in the serine biosynthesis pathway (M00020) in cancer patients as compared with healthy individuals, cysteine biosynthesis pathway (M00338) in periodontitis patients, and the threonine biosynthesis pathway (M00018) in patients with complete tooth loss. Here, these pathways are shown on a gene level by visualizing the absolute difference in relative abundances of the genes (red color scheme), revealing that most genes from these pathways have been altered in their abundance between both conditions on a gene level. Additionally, we demonstrate that some of the genes from the threonine biosynthesis are either specific for Gram-negative (green) or Gram-negative/-positive bacteria (blue), while the serine and cysteine biosynthesis pathways consists of universal genes present in eukaryotes and prokaryotes.
Tobacco smoking leads to a loss of diversity and increased glutamate metabolism in the oral microbiome
To analyze the influence of tobacco smoking on the oral microbiome, we identified clades and metabolic pathways that differed significantly in current regular smokers (60 individuals) and occasional or former smokers (200) compared to non-smokers (103; Table 1). This revealed 24 clades (Fig. 2a) and 12 metabolic pathways (Fig. 2b) that were altered in smokers compared to non-smokers (Supplementary Datasets 4 and 5) and an increased between-sample microbial diversity in smokers (Supplementary Fig. 1). Past smoking habits did not have a significant effect on the microbial community profile for most clades and pathways.
Most of the significant differences in clades were between regular smokers and non-smokers (24 clades). Comparing these groups, a broad range of genus-level clades were significantly increased in smokers, including Lactobacillus, Bifidobacterium, Atopobium, Prevotella, Streptococcus, and Veillonella. On the other hand, Rothia, Neisseria, and Lautropia were significantly decreased in regular smokers. Shifts observed in a few significant clades (5) among occasional or former smokers were concordant with the differences detected between regular smokers and non-smokers. One proposed explanation for these observations is that smoking disrupts niche saturation of health-associated species, eliminating a subset of organisms, altering community composition, lowering alpha-diversity, and increasing beta-diversity. In line with this model, our data show an overall increased between-sample microbial diversity in smokers as compared to non-smokers (Supplementary Fig. 1).
Significant changes in functional modules were detected in smokers, specifically an increased abundance of transport systems, including sugar uptake (M00197: Putative fructooligosaccharide transport, M00216: Multiple sugar transport), phosphate uptake (M00222: Phosphate transport) and metal transport systems, such as manganese, zinc, and iron (M00319: Manganese/zinc/iron transport). An increase in abundance of the GABA gamma aminobutyrate shunt was also detected again (M00027: GABA (gamma-Aminobutyrate) shunt), used by a variety of bacteria, in particular species of Streptococcus and Lactobacillus 48,49, to cope with acid and oxidative stress50. Likewise, genes involved in amino acid degradation, specifically the conversion of histidine into glutamate, were also increased (M00045: Histidine degradation, histidine => N-formiminoglutamate => glutamate), indicating that pathways involved in glutamate metabolism are represented at higher levels in patients that smoke on a regular basis.
Limited structural changes and moderate functional implications tied to tumor HPV status, periodontitis, and alcohol consumption
Our multivariate model of cohort features associated with the microbiome included several other clinical and environmental factors (see Methods), but none of these showed evidence of associations as strong as those above for oral cancer or tooth loss (Table 1, Fig. 2). First, we assessed differences in microbial composition and function between patients with HPV-positive (35) and HPV-negative (55) oral cancer. Members of the Actinomyces, Granulicatella, Oribacterium, and Campylobacter genera as well as Veillonella dispar, Rothia mucilaginosa, and Haemophilus parainfluenzae were significantly increased in patients with HPV-positive cancers compared to non-HPV cancer patients.
In contrast, Streptococcus anginosus, Peptoniphilus, and Mycoplasma were significantly decreased in HPV-positive cancers. Functional changes in the oral microbial communities of HPV-positive vs. HPV-negative cancer patients included phosphonate transport (M00223: Phosphonate transport) and amino acid metabolism through heme (M00121: Heme biosynthesis, glutamate => protoheme/siroheme), Shikimate (M00022: Shikimate pathway, phosphoenolpyruvate + erythrose-4P => chorismate), glutamate, and ornithine (M00028: Ornithine biosynthesis, glutamate => ornithine, Supplementary Dataset 5). We further investigated stratification of the oral cancer cases by site (oral vs. oropharyngeal) as well as HPV status, yielding 52 oral cancers of which 2 (4%) were positive for HPV, 44 (85%) negative, and 6 (11%) unspecified; and 64 oropharyngeal cancers of which 29 (45%) were HPV-positive, 10 (16%) negative, and 25 (39%) unspecified (5 cancer cases with unspecified sites were excluded from this analysis). Likely due to the small numbers resulting from this stratification, we did not identify any oral-oropharyngeal differences that were statistically significant after multiple hypothesis testing correction (Mann-Whitney U FDR q-value < 0.1). The same was true when stratifying the analysis by tumor HPV status.
Few significantly different clades were detected in patients with a history of periodontal disease (82 subjects) as compared to those without (269 subjects, Supplementary Dataset 4); only the Fusobacterium, Leptotrichiaceae, Eikenella, and Capnocytophaga genera were increased. Importantly, these are all Gram-negative bacteria, in line with previous findings that periodontitis is characterized by a shift from primarily Gram-positive to Gram-negative organisms51. Likewise, at the functional level, few modules were consistently different (Supplementary Dataset 5). Genes for mannose uptake (M00276: PTS system, mannose-specific II component) were less abundant; while subtle, this likely reflects the shift from a predominately Gram-positive community to Gram-negative, as sugar transport and metabolism is primarily attributed to the early tooth-colonizing Gram-positive streptococci. Interestingly, the higher abundance of Gram-negative anaerobes was associated with an increase in L-cysteine synthesis, both directly and indirectly through degradation of methionine (M00338: Cysteine biosynthesis, homocysteine + serine => cysteine, M00035: Methionine degradation). In addition to being an amino acid for protein synthesis, L-cysteine acts as an antioxidant and is also metabolized to hydrogen sulfide (H2S)52.
Finally, only one clade was significantly linked to alcohol consumption when comparing microbial profiles among current regular alcohol drinkers (168), former or irregular drinkers (187), and subjects never drinking alcohol (8, Supplementary Dataset 4). This represented an unclassified species in genus Capnocytophaga marginally decreased in current regular drinkers. No changes in metabolic modules were associated with alcohol consumption status, and even in targeted tests of only heavy (≥3 drinks per day) and non-drinkers, only two clades were enriched (Sharpea and unclassified Veillonellaceae) and one depleted (Lactococcus, Mann-Whitney U FDR q < 0.1). Notably, these were all relatively minor genera across the samples, and none of the differences in group means exceeded 0.1% relative abundance.
Discussion
In this study, we assessed structural and functional changes in the oral microbiome during oral cancer, in tandem with clinical, demographic, and environmental covariates. This multivariate analysis remained well-powered compared to previous studies42,43 due to our sample size of over 120 oral cancer cases and twice as many matched controls. Few specific genera were consistently enriched or depleted during cancer, while functional shifts included decreased potential for microbial glutamate metabolism and metal (particularly iron) transport. The greatest dysbioses arose during tooth loss, particularly when no natural teeth remained, and other oral cancer risk factors including tobacco smoking, HPV infection, and periodontitis showed significant but smaller effects (Supplementary Dataset 4 and 8).
A small number of genera were differentially abundant during oral cancer, specifically Dialister (enriched) and Scardovia (depleted). Dialister are Gram-negative anaerobes within the family Veillonellaceae that have been associated with worse periodontal status53 and endodontic infections54,55. They also appear to compete with species of Scardovia, as co-occurence analysis determined that these two genera infrequently inhabit the same niche56. Although previous sample sizes have been limited, earlier culture-based and culture-independent studies have also reported on oral bacteria disrupted on or in oral and esophageal tumors or in associated saliva samples42,43,46,57,58,59,60,61. In one study by Hooper et al.61, viable bacteria were detected within carcinoma tissues themselves. Our results generally agree with the enriched microbes, as well as those published by Pushalker et al., where bacterial colonization of tumor tissue and normal mucosa from the same subject were again found to be distinct43.
Previous studies suggested that poor oral hygiene could, in addition to increasing the risk of oral cancer16, influence the oral microbiome in cancer patients. The largest microbial effect in our cohort occurred during complete tooth loss, with progressive dysbioses with partial loss. The accompanying loss of community diversity was caused by the depletion of many clades known to inhabit a variety of distinct environments around the teeth (e.g. enamel, epithelial interface, or sub-gingival crevice), reducing Actinomycetaceae, Corynebacterium, Rothia, Prevotella, Flavobacteriales, Streptococcaceae, Fusobacteriales, and Proteobacteria and leaving a community of primarily Lactobacillales. Given the loss of habitats, the depletion of clades ranging up to the phylum level is unsurprising, as is the corresponding loss in inferred metagenomic functional potential.
Tobacco smoking also negatively influences oral health62,63 and can lead to oral cancer21,22,64. In particular, smoking can affect microbial biofilm structure and can result in unstable colonization compared to non-smokers62, increasing susceptibility to bacterial infections in smokers by dysregulation of the innate and adaptive immune responses63. Here, although microbial diversity changes associated with smokers were not statistically significant, we observed a trend toward increased overall between-sample diversity in smokers (Supplementary Fig. 1). This is in line with a previous study showing divergent microbial colonization patterns in smokers62. We likewise observed altered abundances of Firmicutes (Lactobacillus, Veillonella, Streptococcus), Actinobacteria (Bifidobacterium, Atopobium), Proteobacteria (Neisseria), and Bacteroidetes (Prevotella), also in agreement with previous findings26,62,65,66. Smoking may thus lead to a higher abundance of “misplaced” indigenous bacteria: depletion of niches typically colonized by commensal microbes and a greater incidence of atypical organisms, preventing stable biofilm formation and immune development.
HPV is similarly responsible for a large variety of cancer types67,68,69,70, including head and neck squamous cell carcinomas71. Previous studies have reported Streptococcus spp. as cofactors in the malignant transformation of oral keratinocytes by HPV72,73,74,75, although we detected relatively weak associations specific to HPV-positive oral cancers. Six metabolic pathways were significantly different, in addition to several clades including decreased Streptococcus anginous and Mycoplasma and increased Veillonella dispar. Previous studies have also suggested that Mycoplasma infection increases the rate of HPV infection, as well as the risk for abnormal cervical cytology and cancer76,77. Finally, glutamate metabolism and metal transport were, in contrast to other cancers, significantly increased in patients with HPV-positive tumors (as well as during several other indicators of poor oral health), in line with findings that report transglutaminase 2 (and other enzymes) as inhibitors of HPV78 or modulators of viral infections79,80.
Finally, periodontitis is an inflammatory condition resulting from a combination of microbial challenge and disregulated host immune response81. It is tightly linked to a shift to a higher proportion of Gram-negative anaerobes36,37,38 and is causally associated with oral cancer. In this study, four genus-level clades were significantly increased in individuals with periodontitis, namely Fusobacterium, Leptotrichiaceae, Eikenella, and Capnocytophaga. We detected a significant increase in abundance of the Gram-negative anaerobe Fusobacterium, in agreement with previous findings82. In fact, the ability of Fusobacterium to manipulate the immune response and support the growth and colonization of both Eikenella and Capnocytophaga species (the other genera found at higher abundance) is well documented83, which speaks to the polymicrobial nature of periodontal disease. Information on the timing of periodontal disease in this cohort was not captured during the study (e.g. former vs. current), nor exact types of treatment received, which may be responsible for some of these differences as well. Overall, although some of the more usual suspects involved in periodontitis were not detected (e.g. Porphyromonas gingivalis, Tannerella forsythesis, Treponema denticola), the shift to a higher abundance of Gram-negative anaerobes is clearly in agreement with previous studies, and no additional dysbiotic interactions between periodontitis and cancer-linked microbes were detected.
Using a carefully controlled cohort, we were able to draw robust conclusions about associations between the oral microbiome and several disease states. By accounting for technical considerations such as sequencing batch effects and self-reporting limitations (e.g. for tobacco and alcohol consumption) during our analysis, we were able to demonstrate that the variances explained by the clinical factors are larger than those explained by batch effect. Thus, the relationships we observed are significant. Although the study design here permitted extensive testing of microbial dysbioses associated with oral cancer and its risk factors, as well as limited functional testing, much work remains in the analysis of the oral cancer microbiome. Causal factors and molecular mechanisms must be identified, with shotgun metagenomics (and/or metatranscriptomics) an obvious potential culture-independent follow-up. In addition to leveraging advances in sequencing methods, future studies should consider environmental factors known to affect the microbiome more broadly (e.g. diet), which we were not able to assess. Ultimately, studies of the microbiome will combine greater molecular detail with larger population sizes and longitudinal monitoring to improve early detection and mitigation of microbial factors in oral cancer.
Methods
Participants
Eligible case subjects were identified from among consecutive patients diagnosed with oral cancer at the outpatient otolaryngology clinic of the Ohio State University Comprehensive Cancer Center (Columbus, OH) from July 2011 through May 2013. Patients were eligible for inclusion into the study if they were older than 17 years and were newly diagnosed with a histologically confirmed squamous cell carcinoma of the oral cavity or oropharynx. Anatomic site of origin was determined by a physical examination performed by the treating head and neck surgeon. Individuals with a prior history of head and neck cancer were ineligible.
Eligible controls included patients older than 17 years with no history of cancer who were evaluated as an outpatient for any benign condition between August 2011 and August 2013 at the same otolaryngology service where cases were enrolled. This control population was considered to be representative of the referral population from which the case subjects were identified. After enrollment of a case subject, eligible control subjects in the same gender and age (5-year intervals) categories were invited to participate until two control subjects were individually matched to each case. The study protocol was approved by the Institutional Review Board of the Ohio State University and written informed consent was obtained from all study participants. All methods were performed in accordance with the relevant guidelines and regulations.
Oral rinse samples were collected from case subjects before the initiation of cancer therapy and from control subjects at enrollment. Oral rinse samples were collected by use of a 30-second rinse and gargle with either ScopeTM mouthwash or saline.
Detailed demographic and behavioral information was collected from all cases and controls using a computer-assisted interview programmed on a touch screen computer (Apple iPad). Domains included demographics, lifetime measures of tobacco, alcohol and marijuana use and medical and dental history. A “never” user of tobacco was defined as an individual who reported never having used any form of tobacco, including cigarettes, pipe, cigar, chewing tobacco or snuff. A current tobacco user was defined as reporting current use of one or more cigarettes per day or cigar, pipe or smokeless tobacco once or more a week. A “regular smoker” was defined as someone who ever smoked cigarettes daily for one month or more, or who used cigars, pipe or chewing tobacco at least once a week for six months or more. A “former user” is someone who reported they no longer did so at this interval when responding to the survey, making the minimal time frame since cessation one month for cigarette smokers and six months for other tobacco users. The median number of cigarettes per day among current smokers was 20 (IQR 12–27).
A “never” user of alcohol was defined as an individual who reported never having had a drink containing alcohol. A “current” alcohol user was defined as an individual who reported current consumption of one or more drinks per month. One “drink” of beer was defined as 12 ounces, wine as 5 ounces, and liquor as 1.5 ounces. The median number of drinks per week among current drinkers was 8 (IQR 2–20). An individual with a history of periodontitis was defined as an individual who reported having been told by a dental health provider that they had periodontitis. Current tooth status was self-reported in the specified categories.
An HPV-positive tumor was defined as a tumor positive for high-risk HPV E6 or E7 mRNA expression as previously described84. Briefly, samples were considered evaluable if DNA (ERV3) and mRNA (RPLPO, after reverse transcription) control templates could be amplified by qPCR and RT-PCR, respectively, after purification from formalin-fixed and paraffin-embedded tumor specimens. Tumor HPV status was determined by detection of HPV DNA of 15-high-risk types by consensus SPF10 primer system PCR amplification followed by reverse line blot hybridization (Inno-LiPa assay, Innogenetics). HPV DNA-positive samples were examined for the presence of E6 E7 mRNA expression of the corresponding HPV type by type-specific RT-PCR, and samples were considered positive if above the lower limit of assay reproducibility84.
DNA extraction
Oral rinse samples were immediately placed at 4 °C and processed as previously described2. DNA was purified on a robotic, magnetic-bead based platform (QIAsymphony SP, Qiagen Inc.) using the QIAsymphony virus/bacteria Midi Kit and stored at −80 °C until further analysis.
16S rRNA gene sequencing
The 16S gene library consisted of Illumina MiSeq sequences targeting the V4 variable region. Detailed protocols used for 16S amplification and sequencing are as described before85. In brief, genomic DNA was subjected to 16S amplification using primers incorporating the Illumina adapters and a sample-specific barcode sequence, allowing directional sequencing covering variable region V4 (Primers: 515 F [GTGCCAGCMGCCGCGGTAA] and 806 R [GGACTACHVGGGTWTCTAAT]). PCR mixtures contained 10 μl of diluted template (1:50), 10 μl of HotMasterMix with the HotMaster Taq DNA Polymerase (5 Prime), and 5 μl of primer mix (2 μM of each primer). The cycling conditions consisted of an initial denaturation of 94 °C for 3 min, followed by 30 cycles of denaturation at 94 °C for 45 sec, annealing at 50 °C for 60 sec, extension at 72 °C for 5 min, and a final extension at 72 °C for 10 min. Amplicons were quantified on the Caliper LabChipGX (PerkinElmer, Waltham, MA), pooled in equimolar concentrations, size selected (375–425 bp) on the Pippin Prep (Sage Sciences, Beverly, MA) to reduce non-specific amplification products from host DNA, and a final library sizing and quantification was performed on an Agilent Bioanalyzer 2100 using DNA 1000 chips (Agilent Technologies, Santa Clara, CA). Sequencing was performed on the Illumina MiSeq v2 platform, according to the manufacturer’s specifications with addition of 5% PhiX, and generating paired-end reads of 150b in length in each direction. The overlapping paired-end reads were stitched together (approximately 97 bp overlap), and size selected to reduce non-specific amplification products from host DNA (22–275 bp).
To mitigate differences in detected microbial abundances arising for technical reasons among our three sequencing batches, we included sequencing phase as a covariate in all analyses. 124 microbial clades were significantly associated with batch identifier after adjusting for other covariates (case/control status, tooth status, etc.) and for multiple comparisons. Adjusting for this batch effect errs on the side of false negatives rather than false positives, but the loss in the power governs the type I error rate, implying that remaining non-batch associations are reliable.
Operational taxonomy unit (OTU) picking and taxonomy assignment
We used QIIME (version 1.6.0)86 to perform OTU picking and taxonomy assignment as specified by PICRUSt87 (Supplementary Dataset 1). Specifically, we clustered the sequences from each sample into OTUs at 97% identity using UCLUST88 and matched reference sequences against Greengenes 13.589. The resulting OTU tables were checked for mislabeling90 and contamination91, yielding mean sequence depth of 29,914 reads/sample. We finally applied PICRUST87 to predict gene family abundances from our 16S rRNA gene data (Supplementary Dataset 2, summary quality control statistics in Supplementary Dataset 9).
Metabolic pathway reconstruction
Inferred per-community gene abundances were subsequently reconstructed into microbial small metabolic modules, whose relative abundances were computed using HUMAnN92,93. We grouped KEGG orthologues into modules represented as gene sets and determined whether a module is present based on KEGG’s conjunctive normal form logic. Next, we computed each pathway’s relative abundance as a smoothed average over all genes, taking outliers and gap filling into account, which resulted in the relative abundances of modules within each sample. These sample-by-module relative abundances were subsequently treated in the same manner as sample-by-clade microbial abundances. We calculated nearest sequenced taxon index (NSTI) values [80] for all samples from this study, which ranged from a greatest distance of ~0.07 through a minimum of ~0.01 and a mean of ~0.03, suggesting a Spearman correlation with underlying metagenomes between 0.7–0.9, well within the range of biological confidence (Supplementary Fig. 2).
Significant associations of microbial clades and pathways with sample metadata
OTU tables and reconstructed pathway abundance data were first processed for quality control. Clinical metadata features were not used if more than 10% of the data were missing, or when they did not vary in value over the available samples. Features (i.e. clades or pathways) of very low abundance (<0.001 in ≥90% of samples) as well as feature outliers (>3 × interquartile range in either direction) were removed as described previously92. After processing, 363 samples passed quality control for clade and functional abundance analyses.
Clades and functions were tested for statistically significant associations with clinical metadata of interest using a multivariate analysis92,93. Here, the relative abundance of each clade was normalized with a variance-stabilizing arcsine square-root transformation and linked with the chosen covariates using a linear model, while the model selection for sparse data was performed per feature using boosting. A multivariate linear model (MaAsLin, http://huttenhower.sph.harvard.edu/maaslin) associating all available metadata (including sequencing batch) with each clade was boosted independently, and any metadata selected in at least 1% of these iterations was tested for significance in a standard generalized linear model. Within each metadatum/feature association, multiple comparisons over factor levels were independently adjusted using a Bonferonni correction; multiple hypothesis tests over all features and metadata associations were adjusted on the Bonferonni corrected p-values to produce a final Benjamini and Hochberg false discovery rate (FDR). Unless otherwise indicated, significant association was considered below a FDR q-value of 0.25.
Features included in the multivariate linear model are listed in Supplementary Dataset 3 and comprise case/control status, tooth loss status, tumor HPV status (for cases) by qPCR/RT-PCR/both, periodontal disease, diabetes status, cigarette and tobacco usage, alcohol consumption, microbial DNA concentration, whether a subject was born in the United States, race/ethnicity, age, and gender. All raw sequencing data are available from the SRA at BioProject number PRJNA321193.
References
Ferlay, J. et al. Estimates of worldwide burden of cancer in 2008: GLOBOCAN 2008. Int J Cancer 127, 2893-2917 (2010).
Gillison, M. L. et al. Tobacco smoking and increased risk of death and progression for patients with p16-positive and p16-negative oropharyngeal cancer. J Clin Oncol 30, 2102–2111 (2012).
Blot, W. J. et al. Smoking and drinking in relation to oral and pharyngeal cancer. Cancer Res 48, 3282–3287 (1988).
Hashibe, M. et al. Interaction between tobacco and alcohol use and the risk of head and neck cancer: pooled analysis in the International Head and Neck Cancer Epidemiology Consortium. Cancer Epidemiol Biomarkers Prev 18, 541–550 (2009).
Gillison, M. L. et al. Distinct risk factor profiles for human papillomavirus type 16-positive and human papillomavirus type 16-negative head and neck cancers. J Natl Cancer Inst 100, 407–420 (2008).
Lingen, M. W. et al. Low etiologic fraction for high-risk human papillomavirus in oral cavity squamous cell carcinomas. Oral Oncol 49, 1–8 (2013).
Warnakulasuriya, S. Living with oral cancer: epidemiology with particular reference to prevalence and life-style changes that influence survival. Oral Oncol 46, 407–410 (2010).
Anand, R., Dhingra, C., Prasad, S. & Menon, I. Betel nut chewing and its deleterious effects on oral cavity. Journal of cancer research and therapeutics 10, 499–505 (2014).
Hernandez, B. Y. et al. Betel nut chewing, oral premalignant lesions, and the oral microbiome. PLoS One 12, e0172196 (2017).
Chang, J. S. et al. Investigating the association between oral hygiene and head and neck cancer. Oral Oncol 49, 1010–1017 (2013).
Franco, E. L. et al. Risk factors for oral cancer in Brazil: a case-control study. Int J Cancer 43, 992–1000 (1989).
Zheng, T. Z. et al. Dentition, oral hygiene, and risk of oral cancer: a case-control study in Beijing, People’s Republic of China. Cancer Causes Control 1, 235–241 (1990).
Velly, A. M. et al. Relationship between dental factors and risk of upper aerodigestive tract cancer. Oral Oncol 34, 284–291 (1998).
Talamini, R. et al. Oral hygiene, dentition, sexual habits and risk of oral cancer. Br J Cancer 83, 1238–1242 (2000).
Garrote, L. F. et al. Risk factors for cancer of the oral cavity and oro-pharynx in Cuba. Br J Cancer 85, 46–54 (2001).
Rosenquist, K. et al. Oral status, oral infections and some lifestyle factors as risk factors for oral and oropharyngeal squamous cell carcinoma. A population-based case-control study in southern Sweden. Acta Otolaryngol 125, 1327–1336 (2005).
Guha, N. et al. Oral health and risk of squamous cell carcinoma of the head and neck and esophagus: results of two multicentric case-control studies. Am J Epidemiol 166, 1159–1173 (2007).
Subapriya, R., Thangavelu, A., Mathavan, B., Ramachandran, C. R. & Nagini, S. Assessment of risk factors for oral squamous cell carcinoma in Chidambaram, Southern India: a case-control study. Eur J Cancer Prev 16, 251–256 (2007).
Divaris, K. et al. Oral health and risk for head and neck squamous cell carcinoma: the Carolina Head and Neck Cancer Study. Cancer Causes Control 21, 567–575 (2010).
Sato, F. et al. Inverse association between toothbrushing and upper aerodigestive tract cancer risk in a Japanese population. Head Neck 33, 1628–1637 (2011).
Graham, S. et al. Dentition, diet, tobacco, and alcohol in the epidemiology of oral cancer. J Natl Cancer Inst 59, 1611–1618 (1977).
Marshall, J. R. et al. Smoking, alcohol, dentition and diet in the epidemiology of oral cancer. Eur J Cancer B Oral Oncol 28B, 9–15 (1992).
Aggarwal, B. B., Vijayalekshmi, R. V. & Sung, B. Targeting inflammatory pathways for prevention and therapy of cancer: short-term friend, long-term foe. Clin Cancer Res 15, 425–430 (2009).
Read, S. A. & Douglas, M. W. Virus induced inflammation and cancer development. Cancer Lett 345, 174–181 (2014).
Wang, F., Meng, W., Wang, B. & Qiao, L. Helicobacter pylori-induced gastric inflammation and gastric cancer. Cancer Lett 345, 196–202 (2014).
Thomas, A. M. et al. Alcohol and tobacco consumption affects bacterial richness in oral cavity mucosa biofilms. BMC Microbiol 14, 250 (2014).
Sheth, C. C. et al. Alcohol and tobacco consumption affect the oral carriage of Candida albicans and mutans streptococci. Letters in applied microbiology 63, 254–259 (2016).
Waszkiewicz, N. et al. Salivary alcohol dehydrogenase in non-smoking and smoking alcohol-dependent persons. Alcohol 48, 611–616 (2014).
Monteiro-da-Silva, F., Sampaio-Maia, B., Pereira Mde, L. & Araujo, R. Characterization of the oral fungal microbiota in smokers and non-smokers. European journal of oral sciences 121, 132–135 (2013).
Consortium, T. H. M. P. Structure, function and diversity of the healthy human microbiome. Nature 486, 207–214 (2012).
Jenkinson, H. F. & Lamont, R. J. Oral microbial communities in sickness and in health. Trends Microbiol 13, 589–595 (2005).
Kroes, I., Lepp, P. W. & Relman, D. A. Bacterial diversity within the human subgingival crevice. Proc Natl Acad Sci USA 96, 14547–14552 (1999).
Cephas, K. D. et al. Comparative analysis of salivary bacterial microbiome diversity in edentulous infants and their mothers or primary care givers using pyrosequencing. PLoS One 6, e23503 (2011).
Ahn, J. et al. Oral microbiome profiles: 16S rRNA pyrosequencing and microarray assay comparison. PLoS One 6, e22788 (2011).
Lazarevic, V., Whiteson, K., Hernandez, D., Francois, P. & Schrenzel, J. Study of inter- and intra-individual variations in the salivary microbiota. BMC Genomics 11, 523 (2010).
Curtis, M. A., Zenobia, C. & Darveau, R. P. The relationship of the oral microbiotia to periodontal health and disease. Cell Host Microbe 10, 302–306 (2011).
Kumar, P. S. et al. New bacterial species associated with chronic periodontitis. J Dent Res 82, 338–344 (2003).
Socransky, S. S. & Haffajee, A. D. Periodontal microbial ecology. Periodontol 2000 38, 135–187 (2005).
Chhour, K. L. et al. Molecular analysis of microbial diversity in advanced caries. J Clin Microbiol 43, 843–849 (2005).
Zambon, J. J. & Kasprzak, S. A. The microbiology and histopathology of human root caries. Am J Dent 8, 323–328 (1995).
Preza, D. et al. Bacterial profiles of root caries in elderly patients. J Clin Microbiol 46, 2015–2021 (2008).
Hooper, S. J. et al. A molecular analysis of the bacteria present within oral squamous cell carcinoma. J Med Microbiol 56, 1651–1659 (2007).
Pushalkar, S. et al. Comparison of oral microbiota in tumor and non-tumor tissues of patients with oral squamous cell carcinoma. BMC Microbiol 12, 144 (2012).
Pushalkar, S. et al. Microbial diversity in saliva of oral squamous cell carcinoma. FEMS Immunol Med Microbiol 61, 269–277 (2011).
Schmidt, B. L. et al. Changes in abundance of oral microbiota associated with oral cancer. PLoS One 9, e98741 (2014).
Mager, D. L. et al. The salivary microbiota as a diagnostic indicator of oral cancer: a descriptive, non-randomized study of cancer-free and oral squamous cell carcinoma subjects. J Transl Med 3, 27 (2005).
Kanehisa, M. et al. Data, information, knowledge and principle: back to metabolism in KEGG. Nucleic acids research 42, D199–205 (2014).
Li, H. & Cao, Y. Lactic acid bacterial cell factories for gamma-aminobutyric acid. Amino Acids 39, 1107–1116 (2010).
Dhakal, R., Bajpai, V. K. & Baek, K. H. Production of gaba (gamma - Aminobutyric acid) by microorganisms: a review. Braz J Microbiol 43, 1230–1241 (2012).
Feehily, C. & Karatzas, K. A. Role of glutamate metabolism in bacterial responses towards acid and other stresses. J Appl Microbiol 114, 11–24 (2013).
Darveau, R. P. Periodontitis: a polymicrobial disruption of host homeostasis. Nat Rev Microbiol 8, 481–490 (2010).
Li, L., Rose, P. & Moore, P. K. Hydrogen sulfide and cell signaling. Annual review of pharmacology and toxicology 51, 169–187 (2011).
Kumar, P. S., Griffen, A. L., Moeschberger, M. L. & Leys, E. J. Identification of candidate periodontal pathogens and beneficial species by quantitative 16S clonal analysis. J Clin Microbiol 43, 3944–3955 (2005).
Drago, L., Vassena, C., Saibene, A. M., Del Fabbro, M. & Felisati, G. A case of coinfection in a chronic maxillary sinusitis of odontogenic origin: identification of Dialister pneumosintes. J Endod 39, 1084–1087 (2013).
Ribeiro, A. C., Matarazzo, F., Faveri, M., Zezell, D. M. & Mayer, M. P. Exploring bacterial diversity of endodontic microbiota by cloning and sequencing 16S rRNA. J Endod 37, 922–926 (2011).
Bik, E. M. et al. Bacterial diversity in the oral cavity of 10 healthy individuals. ISME J 4, 962–974 (2010).
Nagy, K. N., Sonkodi, I., Szoke, I., Nagy, E. & Newman, H. N. The microflora associated with human oral carcinomas. Oral Oncol 34, 304–308 (1998).
Sasaki, H. et al. Presence of Streptococcus anginosus DNA in esophageal cancer, dysplasia of esophagus, and gastric cancer. Cancer Res 58, 2991–2995 (1998).
Tateda, M. et al. Streptococcus anginosus in head and neck squamous cell carcinoma: implication in carcinogenesis. Int J Mol Med 6, 699–703 (2000).
Shiga, K. et al. Presence of Streptococcus infection in extra-oropharyngeal head and neck squamous cell carcinoma and its implication in carcinogenesis. Oncol Rep 8, 245–248 (2001).
Hooper, S. J. et al. Viable bacteria present within oral squamous cell carcinoma tissue. J Clin Microbiol 44, 1719–1725 (2006).
Kumar, P. S., Matthews, C. R., Joshi, V., de Jager, M. & Aspiras, M. Tobacco smoking affects bacterial acquisition and colonization in oral biofilms. Infect Immun 79, 4730–4738 (2011).
Bagaitkar, J., Demuth, D. R. & Scott, D. A. Tobacco use increases susceptibility to bacterial infection. Tob Induc Dis 4, 12 (2008).
Johnson, N. Tobacco use and oral cancer: a global perspective. J Dent Educ 65, 328–339 (2001).
Aas, J. A., Paster, B. J., Stokes, L. N., Olsen, I. & Dewhirst, F. E. Defining the normal bacterial flora of the oral cavity. J Clin Microbiol 43, 5721–5732 (2005).
Kumar, P. S. et al. Changes in periodontal health status are associated with bacterial community shifts as assessed by quantitative 16S cloning and sequencing. J Clin Microbiol 44, 3665–3673 (2006).
Walboomers, J. M. et al. Human papillomavirus is a necessary cause of invasive cervical cancer worldwide. J Pathol 189, 12–19 (1999).
Monk, B. J. et al. Prognostic significance of human papillomavirus DNA in vulvar carcinoma. Obstet Gynecol 85, 709–715 (1995).
Daling, J. R. et al. A population-based study of squamous cell vaginal cancer: HPV and cofactors. Gynecol Oncol 84, 263–270 (2002).
Palefsky, J. M. et al. HPV vaccine against anal HPV infection and anal intraepithelial neoplasia. N Engl J Med 365, 1576–1585 (2011).
D’Souza, G. et al. Case-control study of human papillomavirus and oropharyngeal cancer. N Engl J Med 356, 1944–1956 (2007).
Burnett-Hartman, A. N., Newcomb, P. A. & Potter, J. D. Infectious agents and colorectal cancer: a review of Helicobacter pylori, Streptococcus bovis, JC virus, and human papillomavirus. Cancer Epidemiol Biomarkers Prev 17, 2970–2979 (2008).
Moyle, P. et al. In Understanding Biology Using Peptides Vol. 9 American Peptide Symposia (ed SylvieE Blondelle) Ch. 169, 407–408 (Springer New York, 2006).
Ruoff, K. L. Streptococcus anginosus (“Streptococcus milleri”): the unrecognized pathogen. Clin Microbiol Rev 1, 102–108 (1988).
Schwartz, J. et al. Streptococci-human papilloma virus interaction with ethanol exposure leads to keratinocyte damage. J Oral Maxillofac Surg 70, 1867–1879 (2012).
Zhang, L. D., Pei, J., Zhang, H. M. & Sun, X. F. Relationship between mycoplasma and chlamydia infection and lesions in the cervical tissue in high-risk HPV-positive patients. Zhonghua Shi Yan He Lin Chuang Bing Du Xue Za Zhi 24, 346–348 (2010).
Biernat-Sudolska, M., Szostek, S., Rojek-Zakrzewska, D., Klimek, M. & Kosz-Vnenchak, M. Concomitant infections with human papillomavirus and various mycoplasma and ureaplasma species in women with abnormal cervical cytology. Adv Med Sci 56, 299–303 (2011).
Jeon, J. H. et al. Transglutaminase 2 inhibits Rb binding of human papillomavirus E7 by incorporating polyamine. EMBO J 22, 5273–5282 (2003).
Nargi-Aizenman, J. L. et al. Glutamate receptor antagonists protect from virus-induced neural degeneration. Ann Neurol 55, 541–549 (2004).
Chen, C. J. et al. Glutamate released by Japanese encephalitis virus-infected microglia involves TNF-alpha signaling and contributes to neuronal death. Glia 60, 487–501 (2012).
Page, R. C. & Kornman, K. S. The pathogenesis of human periodontitis: an introduction. Periodontol 2000 14, 9–11 (1997).
Han, Y. W. et al. Interactions between periodontal bacteria and human oral epithelial cells: Fusobacterium nucleatum adheres to and invades epithelial cells. Infect Immun 68, 3140–3146 (2000).
Signat, B., Roques, C., Poulet, P. & Duffaut, D. Fusobacterium nucleatum in periodontal health and disease. Curr Issues Mol Biol 13, 25–36 (2011).
Jordan, R. C. et al. Validation of methods for oropharyngeal cancer HPV status determination in US cooperative group trials. Am J Surg Pathol 36, 945–954 (2012).
Caporaso, J. G. et al. Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J 6, 1621–1624 (2012).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7, 335–336 (2010).
Langille, M. G. et al. Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat Biotechnol 31, 814–821 (2013).
Edgar, R. C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26, 2460–2461 (2010).
McDonald, D. et al. An improved Greengenes taxonomy with explicit ranks for ecological and evolutionary analyses of bacteria and archaea. ISME J 6, 610–618 (2012).
Knights, D. et al. Supervised classification of microbiota mitigates mislabeling errors. ISME J 5, 570–573 (2011).
Knights, D. et al. Bayesian community-wide culture-independent microbial source tracking. Nat Methods 8, 761–763 (2011).
Morgan, X. C. et al. Dysfunction of the intestinal microbiome in inflammatory bowel disease and treatment. Genome Biol 13, R79 (2012).
Abubucker, S. et al. Metabolic reconstruction for metagenomic data and its application to the human microbiome. PLoS Comput Biol 8, e1002358 (2012).
Author information
Authors and Affiliations
Contributions
D.B., B.R., E.M.H., M.E.D., J.R., E.A.F., M.L.G., and C.H. wrote the main manuscript text. D.B., B.R., T.T., E.A.F., M.E.D., M.L.G., and C.H. analyzed the data. M.L.G., R.P., J.L., E.O., and W.X. collected the oral samples. D.G. sequenced the data. M.L.G. and C.H. supervised the study.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Börnigen, D., Ren, B., Pickard, R. et al. Alterations in oral bacterial communities are associated with risk factors for oral and oropharyngeal cancer. Sci Rep 7, 17686 (2017). https://doi.org/10.1038/s41598-017-17795-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-17795-z
This article is cited by
-
Oral squamous cell carcinomas: state of the field and emerging directions
International Journal of Oral Science (2023)
-
Altered oral microbiome in Sudanese Toombak smokeless tobacco users carries a newly emerging risk of squamous cell carcinoma development and progression
Scientific Reports (2023)
-
Oral hygiene, mouthwash usage and cardiovascular mortality during 18.8 years of follow-up
British Dental Journal (2023)
-
Smoking and salivary microbiota: a cross-sectional analysis of an Italian alpine population
Scientific Reports (2023)
-
The Association of Oral Microbiome Dysbiosis with Gastrointestinal Cancers and Its Diagnostic Efficacy
Journal of Gastrointestinal Cancer (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.