Abstract
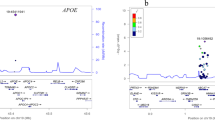
We conducted a two-sample Mendelian randomization study to test the hypothesis that raised plasma levels of the branched-chain amino acids isoleucine, leucine, and valine are associated with Alzheimer’s disease (AD). From a genome-wide association study of 16,596 individuals of European ancestry, we obtained summary statistics for four independent single nucleotide polymorphisms (SNPs) associated with isoleucine levels and one SNP associated with both leucine and valine levels at genome-wide significance. Summary statistics of the associations of the five SNPs with AD were obtained from the International Genomics of Alzheimer’s Project (17,008 AD cases and 37,154 controls). Based on four SNPs, the odds ratio of AD per genetically predicted one standard deviation higher isoleucine levels was 1.35 (95% CI, 1.08–1.69; p = 0.007). The leucine- and valine-raising allele was not associated with AD (p = 0.46). These data suggest that a genetic predisposition to raised plasma isoleucine levels is positively associated with AD.
Similar content being viewed by others
Introduction
Alzheimer’s disease (AD) is a neurodegenerative disease characterized by synaptic loss and neuronal cell death, primarily in hippocampus, and manifests as progressive memory loss and cognitive decline. The pathological hallmarks of AD are amyloid plaques containing deposits of amyloid-β (Aβ) peptides, and neurofibrillary tangles containing hyperphosphorylated tau protein1. The neuronal damage that leads to AD may in part be caused by toxic effect of Aβ peptides or alterations in Aβ processing1. Thus, modifiable environmental factors or medical therapies that modify the accumulation of amyloid plaques or hippocampal neurogenesis may influence AD development.
The neurotransmitter serotonin is a candidate for reducing amyloid plaque load2,3,4,5 and for enhancing neurogenesis2,6,7,8. It has been observed that AD patients have decreased serotonin receptor levels9,10,11,12 and that activation of serotonin receptors decreases Aβ production5,13 and brain Aβ levels2,3,4,5 and increases neuronal survival2. There are also data showing that selective serotonin reuptake inhibitors, a class of antidepressants that increase serotonin levels, reduce brain Aβ levels3,4 and stimulate neurogenesis in hippocampus6,7,8.
The branched-chain amino acids (BCAAs) isoleucine, leucine, and valine are essential amino acids that are obtained from the diet through protein-containing foods. Uptake of BCAAs into the brain occurs at the blood-brain barrier through a competitive transport carrier that they share with other large neutral amino acids, including tryptophan, the precursor of serotonin14,15,16. Long-term raised circulating BCAA levels could conceivably lead to decreased brain tryptophan levels as well as reduced neuronal serotonin synthesis and serotonergic signaling15,16,17 (Fig. 1). This may consequently increase AD risk but this hypothesis has, to our knowledge, not yet been tested.
Hypothetical effects of raised circulating levels of branched-chain amino acids. Tryptophan (TRP) is converted to serotonin in neurons by tryptophan hydroxylase 2 (TPH2), the rate-limiting enzyme in serotonin synthesis. Because this enzyme is normally unsaturated with TRP, brain serotonin levels are controlled by TRP levels. Branched-chain amino acids (BCAAs) reduce the uptake of TRP into the brain by competing for transport at the blood-brain barrier (BBB). Lifelong elevated BCAA levels conferred by genetic predisposition to higher plasma BCAA levels may therefore have long-term effects on serotonin signaling and downstream pathways that may influence the risk of Alzheimer’s disease.
Mendelian randomization (MR) is a genetic epidemiologic approach that utilizes genetic variants associated with the modifiable exposure (e.g., plasma BCAA levels) as proxies for the exposure to evaluate whether the exposure is causally associated with the outcome (e.g., AD). This method reduces some of the crucial limitations of observational studies, including confounding and reverse causation bias. A recent genome-wide association study (GWAS) identified several genetic variants that are associated with plasma BCAA levels18. The aim of the present study was to use MR to test the hypothesis that plasma BCAA levels are positively associated with AD.
Methods
Selection of genetic variants and assessment of pleiotropy
A GWAS of 16,596 individuals of European ancestry identified five independent genomic regions with single-nucleotide polymorphisms (SNPs) associated with plasma BCAA levels at genome wide significance level (p < 5 × 10−8)18. We selected the lead SNP from each of these genomic regions. Five independent SNPs (R2 < 0.03) were associated with isoleucine levels. These SNPs were located near PPM1K (rs7678928), DDX19A (rs75950518), TRMT61A (rs58101275), CBLN1 (rs1420601), and GCKR (rs1260326). Only one SNP (rs1440581), near the PPM1K gene, was associated with leucine and valine levels at genome-wide significance. To investigate whether the BCAA-associated SNPs had pleiotropic associations with other phenotypes, we searched PhenoScanner, which is a publicly available GWAS database19.
Association with AD
Summary statistics (beta coefficients and standard errors) for the associations of the BCAA-related SNPs with AD were obtained from the International Genomics of Alzheimer’s Project (IGAP)20. IGAP is a 2-stage study based upon GWAS on individuals of European descent20. We used data from the first stage of IGAP, which genotyped and imputed data on 7,055,881 SNPs to meta-analyze 4 GWAS datasets [Alzheimer Disease Genetics Consortium (ADGC), the Cohorts for Heart and Aging Research in Genomic Epidemiology consortium (CHARGE), the European Alzheimer’s disease Initiative (EADI), and the Genetic and Environmental Risk in AD consortium (GERAD)], including a total of 17,008 cases of AD and 37,154 controls. All studies included in IGAP had been approved by a relevant Institutional Review Board. Informed consent was obtained from participants themselves or, for those with considerable cognitive impairment, from a caregiver, legal guardian, or other proxy.
Statistical analysis
An instrumental variable was constructed for each BCAA-associated SNP by dividing the SNP’s effect (beta coefficient) on AD by the corresponding effect on BCAA levels. The instrumental variable estimates for isoleucine were combined using a fixed-effect inverse-variance weighted meta-analysis21. To explore and adjust for pleiotropy, sensitivity analyses were performed using the weighted median and MR-Egger regression methods21. Results are reported as odds ratios (OR) with their 95% confidence intervals (CI) of AD per genetically predicted 1 standard deviation (SD) increase in plasma isoleucine, leucine, and valine levels. All statistical tests were two-sided. We applied a Bonferroni-corrected significance threshold for analyses of five BCAA-raising SNPs (p = 0.05/5 = 0.01). Tests were otherwise considered statistically significant at p < 0.05. The statistical analyses were conducted using Stata, version 14.2 (StataCorp, College Station, Texas, USA) and R (R Foundation), version 3.3.3.
Availability of data and materials
All data generated or analyzed during this study are included in this published article and its supplementary information file.
Results
One SNP (rs1260326 near the GCKR gene) had strong pleiotropic associations with numerous phenotypes (Table S1), and was excluded from the analysis. The remaining SNPs, including four SNPs associated with isoleucine and one SNP associated with leucine and valine, were used as instrumental variables in the MR analyses. All four isoleucine-raising alleles were positively associated with AD but none of the associations was significant at the Bonferroni-corrected significance threshold (Fig. 2; Table S2). The overall OR of AD per genetically predicted 1 SD increase in isoleucine levels, based on four SNPs, was 1.35 (95% CI, 1.08–1.69; p = 0.007), without heterogeneity among estimates from different SNPs (I 2 = 0%, p = 0.50) (Fig. 2). The association was consistent in a sensitivity analysis using the weighted median method (OR = 1.35; 95% CI 1.03–1.77; p = 0.03). The MR-Egger analysis showed no evidence of pleiotropy (intercept = −0.029, p = 0.71) but the causal estimate from this analysis was very imprecise and the CI included the null (OR, 1.87; 95% CI, 0.33–10.58). The leucine- and valine-raising allele was not significantly associated with AD (p = 0.46) (Table S2). The ORs per genetically predicted 1 SD increase in genetically predicted leucine and valine levels were 1.16 (95% CI, 0.78–1.72) and 1.13 (95% CI, 0.82–1.57), respectively.
Discussion
This MR analysis showed that a genetic predisposition to higher plasma isoleucine levels was positively associated with AD. This finding suggests that lifelong raised isoleucine levels may increase the risk of AD. Genetically predicted higher levels of leucine and valine, based on a single genetic variant, were not associated with AD.
It is well known that an increase in circulating BCAA levels leads to decreased brain levels of tryptophan and its conversion to serotonin (Fig. 1)15,16,22. This effect reflects the competitive nature of the blood-brain barrier large neutral amino acid transporter, which is almost fully saturated at normal plasma levels of these amino acids22. Hence, the observed association of genetically raised isoleucine levels with AD may be mediated by decreased brain serotonin levels and diminished serotonin signaling, potentially leading to higher amyloid plaque load2,3,4,5 and decreased neuronal survival2 and adult hippocampal neurogenesis6,7,8. Experimental data have shown that dietary BCAA supplementation leads to lowered hippocampal levels of nerve growth factor, which is involved in growth and survival of cholinergic neurons23. High BCAA levels also result in reduced brain uptake of tyrosine and its conversion to the catecholamine neurotransmitters dopamine and norepinephrine15,22, but their role in AD is uncertain.
A strength of the MR approach is that potential confounding from other modifiable risk factors were reduced by the use of genetic variants as proxies for BCAA exposure. However, we cannot rule out the possibility that the BCAA-associated genetic variants may affect the risk of AD through pathways other than through BCAA levels. For this reason, we excluded the pleiotropic variant near the GCKR gene. The genetic risk scores for isoleucine, leucine and valine were specifically associated with the three BCAAs and not with any of 172 other metabolites measured18, suggesting that pleiotropic effects with other metabolites unlikely explain our findings. We also found no evidence of pleiotropy in sensitivity analyses using the weighted median and MR-Egger methods. However, the MR-Egger method had low statistical power and could not reliably detect pleiotropy. The genetic variants were weakly associated, at nominal statistical significance, with some diseases and disorders, most notably psychiatric and mood disorders that are related to impaired serotonin signaling, such as schizophrenia, bipolar disorders, and major depressive disorders. This observation reinforces the hypothesis that genetically raised BCAA levels might increase AD risk by modifying the serotonergic system.
A limitation of this study is that our genetic instrument for leucine and valine comprised only one genetic variant, leading to low statistical power to detect an association. Although the odds ratio estimates for the leucine- and valine-raising allele was in the same direction as for isoleucine, the precision was low and the associations were not significant. Likewise, none of the individual SNPs associated with isoleucine levels was significantly associated with AD at the Bonferroni-corrected significance level.
In conclusion, using data from a large genetic consortium on AD, we found that a genetic predisposition to raised plasma isoleucine levels was associated with AD. The role of BCAAs in the development of AD merits further investigation.
References
Ballard, C. et al. Alzheimer’s disease. Lancet 377, 1019–1031, https://doi.org/10.1016/s0140-6736(10)61349-9 (2011).
Cho, S. & Hu, Y. Activation of 5-HT4 receptors inhibits secretion of beta-amyloid peptides and increases neuronal survival. Exp. Neurol. 203, 274–278, https://doi.org/10.1016/j.expneurol.2006.07.021 (2007).
Sheline, Y. I. et al. An antidepressant decreases CSF Abeta production in healthy individuals and in transgenic AD mice. Sci. Transl. Med. 6, 236re234, https://doi.org/10.1126/scitranslmed.3008169 (2014).
Cirrito, J. R. et al. Serotonin signaling is associated with lower amyloid-beta levels and plaques in transgenic mice and humans. Proc. Natl. Acad. Sci. USA 108, 14968–14973, https://doi.org/10.1073/pnas.1107411108 (2011).
Pimenova, A. A., Thathiah, A., De Strooper, B. & Tesseur, I. Regulation of amyloid precursor protein processing by serotonin signaling. PLoS One 9, e87014, https://doi.org/10.1371/journal.pone.0087014 (2014).
Hanson, N. D., Owens, M. J. & Nemeroff, C. B. Depression, antidepressants, and neurogenesis: a critical reappraisal. Neuropsychopharmacology 36, 2589–2602, https://doi.org/10.1038/npp.2011.220 (2011).
Song, N. N. et al. Reducing central serotonin in adulthood promotes hippocampal neurogenesis. Sci. Rep. 6, 20338, https://doi.org/10.1038/srep20338 (2016).
Santarelli, L. et al. Requirement of hippocampal neurogenesis for the behavioral effects of antidepressants. Science 301, 805–809, https://doi.org/10.1126/science.1083328 (2003).
Marcos, B. et al. Involvement of an altered 5-HT -{6} receptor function in behavioral symptoms of Alzheimer’s disease. J. Alzheimers Dis. 14, 43–50 (2008).
Garcia-Alloza, M. et al. Differential involvement of 5-HT(1B/1D) and 5-HT6 receptors in cognitive and non-cognitive symptoms in Alzheimer’s disease. Neuropsychopharmacology 29, 410–416, https://doi.org/10.1038/sj.npp.1300330 (2004).
Lorke, D. E., Lu, G., Cho, E. & Yew, D. T. Serotonin 5-HT2A and 5-HT6 receptors in the prefrontal cortex of Alzheimer and normal aging patients. BMC Neurosci. 7, 36, https://doi.org/10.1186/1471-2202-7-36 (2006).
Lai, M. K. et al. Loss of serotonin 5-HT2A receptors in the postmortem temporal cortex correlates with rate of cognitive decline in Alzheimer’s disease. Psychopharmacology (Berl) 179, 673–677, https://doi.org/10.1007/s00213-004-2077-2 (2005).
Tesseur, I. et al. Chronic 5-HT4 receptor activation decreases Abeta production and deposition in hAPP/PS1 mice. Neurobiol. Aging. 34, 1779–1789, https://doi.org/10.1016/j.neurobiolaging.2013.01.020 (2013).
Fernstrom, J. D. Role of precursor availability in control of monoamine biosynthesis in brain. Physiol. Rev. 63, 484–546 (1983).
Pardridge, W. M. Brain metabolism: a perspective from the blood-brain barrier. Physiol Rev 63, 1481–1535 (1983).
Fernstrom, J. D. Branched-chain amino acids and brain function. J. Nutr. 135, 1539S–1546S (2005).
Choi, S., Disilvio, B., Fernstrom, M. H. & Fernstrom, J. D. Oral branched-chain amino acid supplements that reduce brain serotonin during exercise in rats also lower brain catecholamines. Amino Acids 45, 1133–1142, https://doi.org/10.1007/s00726-013-1566-1 (2013).
Lotta, L. A. et al. Genetic Predisposition to an Impaired Metabolism of the Branched-Chain Amino Acids and Risk of Type 2 Diabetes: A Mendelian Randomisation Analysis. PLoS Med. 13, e1002179, https://doi.org/10.1371/journal.pmed.1002179 (2016).
Staley, J. R. et al. PhenoScanner: a database of human genotype-phenotype associations. Bioinformatics 32, 3207–3209, https://doi.org/10.1093/bioinformatics/btw373 (2016).
Lambert, J. C. et al. Meta-analysis of 74,046 individuals identifies 11 new susceptibility loci for Alzheimer’s disease. Nat. Genet. 45, 1452–1458, https://doi.org/10.1038/ng.2802 (2013).
Burgess, S., Bowden, J., Fall, T., Ingelsson, E. & Thompson, S. G. Sensitivity analyses for robust causal inference from Mendelian randomization analyses with multiple genetic variants. Epidemiology 28, 30–42 (2017).
Fernstrom, J. D. Large neutral amino acids: dietary effects on brain neurochemistry and function. Amino Acids 45, 419–430, https://doi.org/10.1007/s00726-012-1330-y (2013).
Scaini, G. et al. Acute and chronic administration of the branched-chain amino acids decreases nerve growth factor in rat hippocampus. Mol. Neurobiol. 48, 581–589, https://doi.org/10.1007/s12035-013-8447-1 (2013).
Acknowledgements
Hugh S Markus is supported by a National Institute for Health Research Senior Investigator award and his work is supported by the Cambridge University Trust National Institute for Health Research Biomedical Research Centre. The authors thank the International Genomics of Alzheimer’s Project (IGAP) for providing summary results data for these analyses. The investigators within IGAP contributed to the design and implementation of IGAP and/or provided data but did not participate in analysis or writing of this report. IGAP was made possible by the generous participation of the control subjects, the patients, and their families. The i–Select chips was funded by the French National Foundation on Alzheimer’s disease and related disorders. ADGC was supported by the NIH/NIA grants: U01 AG032984, U24 AG021886, U01 AG016976, and the Alzheimer’s Association grant ADGC–10–196728. CHARGE was partly supported by the NIH/NIA grant R01 AG033193 and the NIA AG081220 and AGES contract N01–AG–12100, the NHLBI grant R01 HL105756, the Icelandic Heart Association and the Erasmus Medical Center and Erasmus University. EADI was supported by the LABEX (laboratory of excellence program investment for the future) DISTALZ grant, Inserm, Institut Pasteur de Lille, Université de Lille 2 and the Lille University Hospital. GERAD was supported by the Medical Research Council (Grant no 503480), Alzheimer’s Research UK (Grant no 503176), the Wellcome Trust (Grant no 082604/2/07/Z) and German Federal Ministry of Education and Research (BMBF): Competence Network Dementia (CND) grant no 01GI0102, 01GI0711, 01GI0420.
Author information
Authors and Affiliations
Contributions
S.C.L. analyzed the data and drafted the manuscript. S.C.L. and H.S.M. reviewed, commented on, and approved the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Larsson, S.C., Markus, H.S. Branched-chain amino acids and Alzheimer’s disease: a Mendelian randomization analysis. Sci Rep 7, 13604 (2017). https://doi.org/10.1038/s41598-017-12931-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-12931-1
This article is cited by
-
Life course plasma metabolomic signatures of genetic liability to Alzheimer’s disease
Scientific Reports (2024)
-
Graph-Guided Bayesian Factor Model for Integrative Analysis of Multi-modal Data with Noisy Network Information
Statistics in Biosciences (2024)
-
Branched chain amino acids harbor distinct and often opposing effects on health and disease
Communications Medicine (2023)
-
Geroprotective interventions in the 3xTg mouse model of Alzheimer’s disease
GeroScience (2023)
-
The hot sites of α-synuclein in amyloid fibril formation
Scientific Reports (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.