Abstract
Much can be gained from revealing the mechanisms fungal entomopathogens employ. Especially intriguing are fungal parasites that manipulate insect behavior because, presumably, they secrete a wealth of bioactive compounds. To gain more insight into their strategies, we compared the genomes of five ant-infecting Ophiocordyceps species from three species complexes. These species were collected across three continents, from five different ant species in which they induce different levels of manipulation. A considerable number of (small) secreted and pathogenicity-related proteins were only found in these ant-manipulating Ophiocordyceps species, and not in other ascomycetes. However, few of those proteins were conserved among them, suggesting that several different methods of behavior modification have evolved. This is further supported by a relatively fast evolution of previously reported candidate manipulation genes associated with biting behavior. Moreover, secondary metabolite clusters, activated during biting behavior, appeared conserved within a species complex, but not beyond. The independent co-evolution between these manipulating parasites and their respective hosts might thus have led to rather diverse strategies to alter behavior. Our data indicate that specialized, secreted enterotoxins may play a major role in one of these strategies.
Similar content being viewed by others
Introduction
Fungi have evolved to occupy a wide variety of ecologically important niches. Saprophytic fungi are crucial players in the carbon cycle. Mutualistic and parasitic fungi evolved to intimately interact with their plant and animal hosts. A consequence of this broad niche space and the ability of fungi to secrete useful enzymes and bioactive compounds is their successful use in a wide variety of industries (e.g. medicine, food, biopesticides). Much can thus be gained from sequencing fungal genomes and discovering the enzymes and mechanisms they employ. From a medical perspective, this is especially true for fungi that interact with animals. Certain fungi have evolved to interact in mutualistic ways to provide the animal host with beneficial compounds1,2. Others can overcome the immune system and take advantage of the host to further transmission3. Many such interactions can be found with arthropods since they are the most diverse and dominant animals in all terrestrial ecosystems4, and fungal parasites of insects (entomopathogens) can be found in all major phyla5. Despite this, fungal-insect associations remain understudied6 in comparison to fungal parasites of plants. This imbalance is unfortunate given the many insights that could be gained from determining the mechanisms by which fungi invade and conquer their insect hosts.
The fungal parasites that adaptively manipulate the behavior of their insect hosts are particularly intriguing because such manipulation implies many interesting bioactive compounds are likely produced during infection7. Studies into the mechanisms underlying these manipulations may reveal the fungal neuromodulators that are secreted and the pathways that give rise to certain behaviors. Prime examples of fungal manipulation of insect behavior are those where ants are manipulated to leave the nest and bite onto vegetation in elevated positions. Examples of this are wood ants, Formica, infected by Pandora (Phylum: Entomophthoromycota, Order Entomophthorales)8,9,10, and carpenter ants and spiny ants (tribe Camponotini, the genera Camponotus and Polyrhachis, respectively) infected by fungi of the genus Ophiocordyceps (Phylum: Ascomycota, Order Hypocreales)11,12,13,14,15,16. These Ophiocordyceps species can be isolated from their ant hosts and grown in vitro. As such their biology can be studied in more detail by looking at the fungal secretome under various circumstances and reconstructing ant infection and manipulation under controlled conditions17. The biting behavior for one of the species, Ophiocordyceps unilateralis sensu lato (s.l.) from South Carolina, USA, has recently also been studied from a whole genome perspective. Further, a mixed transcriptomics study that compared gene expression at this biting event with the situation after manipulation and before ant and fungus were interacting, has revealed the very first candidate genes that could be important in establishing the behaviors observed18.
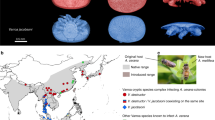
Entomopathogenic Ascomycota mostly reside within the Order Hypocreales, where multiple origins of the host behavioral manipulation trait can be found5. As such, Camponotus and Polyrhachis are not the only ants infected by Ophiocordyceps. Each ant species is seemingly infected by its own Ophiocordyceps species, creating a high level of biodiversity among the ant-manipulating Ophiocordyceps fungi19,20. The description of the vast majority of these species is still underway. In the interim, species complex names are assigned to these Ophiocordyceps fungi based on their fruiting body and ascospore morphology, and the ant species they infect. For example, O. unilateralis s.l. is found on Camponotini species (Camponotus and Polyrhachis) and forms a stalk with the fruiting body (ascoma) attached to one side (hence the epithet unilateralis). Spores are long and more or less “boomerang shaped”13. Ophiocordyceps australis s.l. is found on Ponerine ants, where the fruiting body at the tip of the stalk produces chains of shorter (part-) spores that are released in a fragmented manner5,20. Here, we improved the assembly and annotation of a previously sequenced18 ant-manipulating Ophiocordyceps species, Ophiocordyceps kimflemingiae 20, a unilateralis species from the United States. Additionally, we sequenced, assembled, and annotated the genomes of four additional species: Ophiocordyceps camponoti-rufipedis 13, a species of the O. unilateralis complex from Brazil; two species of the Ophiocordyceps australis complex from Brazil and Ghana; and a species from yet another complex, Ophiocordyceps subramanianii s.l., also from Ghana5. Phylogenetics studies that demonstrate species delimitation within the O. australis complex have not yet been published at this time. We, however refer to the two sequenced australis strains with “species” as per the one fungus-one ant theory19,20. Ophiocordyceps kimflemingiae is from a temperate system, where infected Camponotus castaneus ants are found biting twigs and grasping them with their legs in what is a novel form of manipulation distinct from biting17. Biting was also seen in tropical Ghanaian Ponerine ants infected by local O. australis s.l. and O. subramanianii s.l. species. In this case biting is typically on vertical stems of small, understory vegetation in tropical forests19. The fungus O. camponoti-rufipedis, in the tropical Atlantic rainforests of Brazil, induces leaf biting in its Camponotus rufipes hosts13, whereas the O. australis s.l. species on Ponerine ants from that region does not induce biting behavior20 (Fig. 1). For the O. australis species from Brazil, we thus cannot say with certainty if manipulation is taking place (because the biting event is absent), while the other species induce evident manipulated behaviors (the biting event is present).
Ant-infecting fungi of the genus Ophiocordyceps. Photographs of the five ant-infecting Ophiocordyceps fungi compared in this study as they emerge from their hosts. (a) O. kimflemingiae emerging from a C. castaneus ant that has died biting a twig and grasping it with the legs. Ghanaian Ponerine ants infected by local (b) O. australis s.l. and (c) O. subramanianii s.l. species that were manipulated to bite on vertical stems of small, understory vegetation. (d) O. camponoti-rufipedis emerging from a leaf biting C. rufipes ant, collected in the Atlantic rainforest of Brazil. (e) An O. australis s.l. species from that same region as found on Ponerine ants. Here, biting behavior is not induced and the ant dies in the leaf litter.
According to various “-omics” studies, parasitic traits, such as host invasion and overcoming the immune system, rely on molecular strategies that share many commonalities across plant and animal parasite species21. One would thus expect to observe something similar when investigating possible strategies that fungal parasites use to manipulate the behavior of their animal hosts. Especially if all hosts compared reside within the same family, the Formicidae, and all parasites within the same genus, Ophiocordyceps. Through comparative genomics, we investigated the possible common mechanisms that these fungi employ to manipulate the behavior of their individual ant hosts. In addition, we compared their genomes to several other entomopathogenic and non-entomopathogenic ascomycetes in order to ascertain if genes related to pathogenicity and manipulation are also found in other species or are specific to this group. Moreover, we asked how conserved the identified candidate manipulation genes are by re-analyzing the previously published transcriptome of O. kimflemingiae 18. We then aimed to infer which genes might be good first candidates to focus on in follow-up studies to quantitatively assess their true function. This led us to conclude that a deeper understanding of the function of fungal genes encoding for putative secreted enterotoxins (i.e. fungal genes that have an enterotoxin_a PFAM domain as well as a secretion signal) should be obtained.
Results
Genome features
The Ophiocordyceps draft genomes generated in this study (Table 1) were assembled with reads generated across various sequencing runs using two types of DNA libraries. Only the runs that led to high-quality reads were used for assembly (see Materials and Methods). Contig assembly resulted in genome sizes ranging from 21.91–23.92 million base pairs (Mbp) for O. unilateralis s.l. and O. australis s.l. species. In contrast, O. subramanianii s.l. had an estimated genome size of 32.31 Mb. Gene prediction yielded between 7,621 and 8,629 genes for the O. unilateralis and O. australis species. Ophiocordyceps subramanianii s.l. was predicted to have 11,275 genes. Additionally, GC content in O. subramanianii s.l. (i.e. 60.35%) was much higher compared to the other Ophiocordyceps species in this study (54.66%+/− 1.57%) (Table 1).
We also improved the assembly and gene prediction of the previously published genome of O. unilateralis s.l. strain SC16a (Supplementary Table S1), which has now received the species name O. kimflemingiae 20. In addition to being less fragmented, the new assembly is smaller than previously reported18. This is most likely due to a better assembly of repetitive regions, which were found highest in the unilateralis species (6.59–6.83%) compared to the other-ant-infecting species in this study (Table 1). Despite the smaller assembly size, the new gene prediction increased the gene count with 798 genes. This increase is predominantly caused by the prediction of fewer chimeras (i.e. neighboring genes that are incorrectly merged into one gene model) using the Braker1 pipeline (data not shown).
For all but one of the genomes generated in this study, gene prediction was informed by RNA-Seq reads. For the previously reported North American unilateralis species O. kimflemingiae, reads generated in that former study were used18. Through difficulties with cultivating Brazilian O. camponoti-rufipedis, not enough material was obtained to generate RNA-Seq data in addition to the DNA reads needed to obtain a draft genome. Because both O. kimflemingiae and O. camponoti-rufipedis reside within the same species complex (unilateralis), we attempted mapping the North American unilateralis species reads to the O. camponoti-rufipedis genome to see if they could be used to inform annotation. However, while 93% of the O. kimflemingiae reads mapped to its own genome, only 43% mapped to the O. camponoti-rufipedis genome (Supplementary Table S2). We additionally mapped the reads to another published O. unilateralis s.l. genome, that of O. polyrhachis-furcata 22, to determine whether this would be a more general cross-mapping effect or specific to the O. camponoti-rufipedis genome. A similar mapping of 41% resulted. This suggests that unilateralis species might generally be rather distantly related, making cross-mapping to inform annotation less suitable. For informative purposes, we also cross-mapped the australis species with each other (strains MAP-64 from Brazil and 1348a from Ghana). This resulted in 71% and 82% cross-mapped reads versus 86% and 97% mapped to their own genomes, respectively (Supplementary Table S2). This implies that species in the australis complex are thus likely much more closely related to each other than species within the unilateralis complex.
Orthologous clusters
We set out to investigate which gene predictions of the ant-infecting Ophiocordyceps species in this study are likely conserved and shared with other ascomycetes. In addition, we aimed to discover what species-specific and “manipulation-specific” specializations might have taken place. As such the predicted proteomes of our five ant-infecting fungi were compared to those of 18 other ascomycetous fungi. Of these species, ten were animal parasites (two infecting mammals, two infecting nematodes, and six infecting insects) and 13 reside within the same order (Hypocreales). Four species belonged to the same family (Ophiocordycipitaceae) and two were within the same genus (Ophiocordyceps). A phylogenetic reconstruction based on 67 conserved genes present in each of these organisms is depicted in Fig. 2.
A total of 51,012 orthologous clusters included all annotated proteins of each of the 23 ascomycetes included in this analysis (Supplementary Data S1). Summary statistics regarding this clustering can be found in Supplementary Figure S1. We compared overlap of orthologous clusters between three species ranges: 1) the ant-infecting fungi, which comprised all five draft genomes generated in this study, 2) the other insect-infecting fungi Ophiocordyceps sinensis, Tolypocladium inflatum, both Metarhizium and both Cordyceps species, and 3) all non-insect-infecting ascomycetes, which included other animal-infecting, plant-infecting, fungus-infecting, and saprophytic fungi. The results of this analysis are depicted in the Venn Diagram of Fig. 3a. Subsequently, we performed enrichment analyses for the functional annotations of ant-infecting species genes that were found within the various overlapping and non-overlapping parts of the diagram. Of the 7,931 orthologous clusters that were found within all three species ranges, Gene Ontology (GO) annotations for general biological processes were significantly over-represented. This suggests, as one would expect, that ascomycetes with different lifestyles use similar mechanisms for general processes such as transcription, translation, protein transport and signal transduction. However, genes predicted to encode for (small) secreted proteins, proteins with GO annotations for multi-organism processes and pathogenesis, and more specifically, secreted putative enterotoxins, were under-represented. Indeed, when we performed an enrichment analysis of annotations present in orthologous clusters that were only found in the ant-infecting species (i.e. 6,672 clusters, Fig. 3a), we found the opposite result. GO annotations for (largely the same) general biological processes were significantly under-represented, while (small) secreted proteins, proteins with GO annotations for multi-organism processes and pathogenesis, and putative enterotoxins were over-represented. This suggests that a significant part of the secretome of the ant-infecting fungi in this study is specific to them. This specificity appeared partly due to enterotoxins that are part of the secretome and bioactive small secreted proteins (SSPs) that could be important in fungal-ant interactions. (Small) secreted proteins were also over-represented among the clusters that ant-infecting fungi exclusively shared either with other entomopathogens (262 clusters) or non-entomopathogens (449 clusters). This indicates that their secretomes also contain more general entomopathogen-specific proteins, as well as proteins that are exclusively shared with non-entomopathogenic ascomycetes.
Venn diagrams of orthologous clusters. (a) Venn-diagram of orthologous clusters present within three fungal species ranges: ant-infecting entomopathogens of the genus Ophiocordyceps (blue), other entomopathogens (red), and other ascomycetes in this study (green, ranging from animal-, fungus- and plant-infecting, to saprophytic fungi). (b) Venn-diagram of the 6672 orthologous clusters found only in ant-infecting species, specified into the five Ophiocordyceps species that comprised this species range. A Venn-diagram of all orthologous clusters found in ant-manipulating species is depicted in Supplementary Figure S2.
We also examined how orthologous clusters, that were only found among the ant-infecting fungi were represented by those species (Fig. 3b). Of the total of 6,672 clusters in this comparison, 90.6% appeared to be species-specific. Cluster overlap was thus marginal even though all species in this study reside within the same genus (Ophiocordyceps) and in some cases even within the same species complex (O. unilateralis s.l. and O. australis s.l.). Analyzing enrichment of functional annotation terms in these species-specific clusters again resulted in over-representations of (small) secreted proteins. Though marginal, most cluster overlap was found between more closely related species within the same complex (i.e. O. australis s.l., 289 clusters, and O. unilateralis s.l., 182 clusters Fig. 3b). Also here, enrichment analyses revealed an over-representation for (small) secreted proteins. Thus, it appears that a statistically significant amount of the fungal secretome of these ant-infecting species is either complex- or species-specific. In addition, we found an over-representation of pathogenicity GO terms among the complex-specific orthologous clusters. This finding could be attributed to the presence of orthologous enterotoxins in species within the same complex. Only four clusters that were not present in any of the other ascomycetes in our comparison were shared among all five ant-infecting species (Fig. 3b). None of these four clusters received a functional annotation, but three of them contained genes with predicted secretion signals. For three out of the four clusters, a BLASTp analysis of the genes against the NCBI database resulted solely in hypothetical protein hits with the previously deposited version of the O. kimflemingiae genome18. This indicates that these clusters might indeed represent proteins that are unique to ant-infecting Ophiocordyceps species. Protein sequences within the fourth cluster resulted in hits with metalloproteases in addition to aligning again with a hypothetical protein from O. kimflemingiae (XA68_3159), (Supplementary Table S3). This cluster might thus contain putative metalloproteases that are solely found in the genomes of the ant-infecting fungal species examined here. Moreover, only 2 of the clusters found uniquely in the ant-infecting fungi were shared among all four species that induce biting behavior (Fig. 3b). A BLASTp analysis of the genes within these clusters again resulted in hypothetical protein hits with the previously deposited version of the O. kimflemingiae genome18 (Supplementary Table S4).
Candidate manipulation genes involved in establishing biting behavior
We analyzed the conservation of candidate genes associated with the manipulated biting event observed in infected ants. We used previously published transcriptomics data18 and re-determined differential gene expression by mapping the data to the new version of the O. kimflemingiae genome. As per the previously published study, we followed the reasoning that candidate genes, involved in establishing manipulated biting behavior, would be up-regulated during this event and quickly down-regulated again after. As such, 547 candidate genes were identified, which is 49 more than reported in the previous analysis18. In line with the previously reported data, genes involved in DNA replication, oxidation-reduction processes, secretion and secondary metabolism were over-represented.
Conservation of candidate manipulation genes was also analyzed by way of orthologous clustering. We compared overlap of orthologous clusters containing those candidate genes, that were significantly up- during and down-regulated after manipulated biting behavior in O. kimflemingiae, with three species ranges: 1) the other ant-infecting fungi, which comprised the four new draft genomes generated in this study, 2) the other insect-infecting fungi, and 3) all non-insect-infecting ascomycetes used for comparison earlier. The results of this analysis are depicted in the Venn Diagram of Fig. 4. Of the candidate manipulation genes, 78% appeared to be orthologs of genes present in all other three species ranges (i.e. 423 clusters, Fig. 4). This implies that the genes expressed during manipulated biting induced by O. kimflemingiae are not likely specific to manipulation. Among these broadly shared genes, CYPs and other oxidation-reduction-related functions were over-represented, as well as genes encoding for secreted proteins and proteases. Various secondary metabolism annotations (clusters 7,8 and 9) were also over-represented among these more broadly shared orthologous clusters. They comprised a tryptophan dimethylallyltransferase involved in ergot alkaloid synthesis, various cytochromes, small secreted proteins, a polyketide synthase (PKS), and a PKS-NRPS (non-ribosomal protein synthetase) hybrid. Among the candidate manipulation genes that appeared to be unique to O. kimflemingiae (i.e. 59 clusters, Fig. 4), only SSPs were over-represented. However, 92% of these “unique” genes did not receive a functional annotation. Where any other overlap with the three species ranges was found, SSPs were also over-represented, as were larger secreted proteins. Of those present only in ant-infecting fungi (i.e. 24 clusters Fig. 4), again the majority (79%) did not receive a functional annotation. Among the seven orthologous clusters that were present in all insect-infecting species, but not in other ascomycetes, we found a putative, secreted enterotoxin. This enterotoxin was present in the biting behavior-inducing species O. kimflemingiae (two orthologs), O. camponoti-rufipedis (1 ortholog), O. subramanianii s.l. (two orthologs) and O. australis-Ghana (two orthologs), as well as in O. australis-Brazil and O. sinensis. Moreover, one of the two enterotoxin orthologs in O. kimflemingiae showed a dramatic expression pattern with a >3,000-fold up-regulation during manipulation and a 200-fold down-regulation after18. This enterotoxin could thus potentially be an important key player in the establishment of behavioral manipulation by these Ophiocordyceps species.
Conservation of secondary metabolite clusters
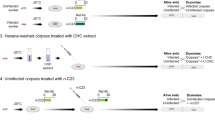
A general hypothesis is that altered host behaviors are established through the secretion of secondary metabolites in addition to larger bioactive compounds. This hypothesis is supported by the over-representation of certain annotated secondary metabolite clusters among genes of O. kimflemingiae that are up-regulated during manipulated biting behavior18. Examining these more closely, we found that indeed the genes, within but also directly flanking the annotated secondary metabolite clusters, followed this particular expression pattern (Fig. 5a). We asked if these clusters were conserved among insect-infecting Ophiocordyceps fungi. This would suggest a similarity in use of secondary metabolites by these fungi to interact with their ant hosts to establish the observed manipulated behaviors. As such, we examined annotated clusters 7,8 and 9 of O. kimflemingiae and their directly flanking genes. These clusters 1) were up-regulated during manipulated biting behavior followed by a significant down-regulation, and 2) shared orthologs with other ascomycetes (see above). For the genes within these clusters, we searched for homologs (BLASTp alignment) and orthologs (orthologous clustering) in the other four ant-infecting fungi (Fig. 5b and Supplementary Figure S3). This demonstrated that the unilateralis species organized at least some of their secondary metabolism-related genes in largely similar clusters. The other Ophiocordyceps species had homologs and orthologs of these secondary metabolism genes scattered throughout their genomes or did not contain a copy at all (Fig. 5b and Supplementary Figure S3). For instance, annotated cluster 8 in O. kimflemingiae contained a tryptophan dimethylallyltransferase flanked by oxidation-reduction-related CYPs and a gene with a FAD binding domain. This cluster is directly flanked by seven genes that followed a similar expression pattern (Fig. 5a). The O. camponoti-rufipedis genome had this tryptophan dimethylallyltransferase in a similar fashion; flanked by CYPs and an FAD-binding gene, followed by homologs and orthologs of the neighboring genes (Fig. 5b). The O. subramanianii s.l. genome also had a similar tryptophan dimethylallyltransferase. It was however flanked by non-homologous/orthologous CYP and FAD-binding genes. In fact, O. subramanianii s.l. did have homologs/orthologs of these genes but they resided on completely different contigs, as did the genes right outside the annotated secondary metabolite cluster (Fig. 5b). Moreover, both australis species did not have an orthologous/homologous gene encoding for this particular tryptophan dimethylallyltransferase. In fact, the australis species from Ghana appeared to not contain a putative tryptophan dimethylallyltransferase at all. Similar conclusions could be made analyzing the other secondary metabolite clusters (Supplementary Figure S3). Clusters 7 and 9 from O. kimflemingiae seemed largely comparable to clusters 31 and 8 from O. camponoti-rufipedis respectively. Yet, genes associated with these clusters were again either not present or scattered across the genomes of the three other ant-infecting species.
Secondary metabolite clusters in behavior manipulating Ophiocordyceps species. (a) Expression profiles of three differentially expressed secondary metabolite clusters in O. kimflemingiae in culture, during manipulated biting behavior, and after manipulation. (b) Homology (identified by BLASTp alignment) and orthology (identified by orthologous clustering) of genes within secondary metabolite cluster 8. Genes that were found using both methods are connected by a black line. Genes found by BLASTp alignment only are connected with a blue line, while genes only found by orthologous clustering are connected with a dashed line. Gene numbers and functions in red indicate genes that received a secondary metabolism annotation.
Phylogeny of fungal enterotoxins
Genes that contained an annotated enterotoxin PFAM domain (PF01375) and a secretion signal were indicated as putative, secreted enterotoxins. Previous results and our current study implicate that genes encoding for these putative enterotoxins are of importance in behavior-manipulating species of Ophiocordyceps fungi. These secreted bacterial-like toxins could potentially be affecting ant behavior by interfering with the production of chemo signaling molecules in the host. This was demonstrated for enterotoxins from bacterial entomopathogens in sex pheromones of boll weevils23,24. They could however also function as mere killing compounds25. In addition, one of these putative enterotoxins was extremely highly up-regulated during the manipulated biting event only18. This particular gene was present in all ant-manipulating Ophiocordyceps species in this study, as well as O. australis-Brazil and O. sinensis. Moreover, the ant-infecting fungi appeared to have a rather large number of genes encoding for these pathogenicity-related proteins. Other ascomycetes generally contained either far fewer (i.e. other entomopathogens, nematode-infecting fungi and Magnaporthe oryzae) or no enterotoxin domain-containing genes (i.e. the remainder of the ascomycetes in this study). The genomes of ant-infecting Ophiocordyceps species had 20–36 putative enterotoxins with O. australis-Ghana (n = 20) having the smallest and O. kimflemingiae (n = 36) having the largest number. Nematode-infecting fungi Ophiocordyceps minnesotensis and Drechmeria coniospora contained 19 and 25 genes with an enterotoxin PFAM domain. Other entomopathogens of the order Hypocreales contained only 4–16 enterotoxin annotations with Metarhizium robertsii and Cordyceps bassiana having the most (n = 16 and n = 14, respectively). The other ascomycete genomes in this study did not have any enterotoxin-encoding genes, except for the plant pathogen M. oryzae, which contained six.
We built a phylogenetic tree based on all fungal genes in this study that contained the PF01375 heat-labile enterotoxin alpha chain domain. Enterotoxins have mostly been reported for bacterial species (e.g. refs 23,24,25). We therefore also included four bacterial enterotoxins. This resulted in a tree based on 252 domain regions. We first determined how the enterotoxins of the bacterial and plant-infecting species (i.e. M. oryzae) would cluster with respect to the majority of animal-infecting species (i.e. insect- and nematode-infecting fungi). This placed three bacterial enterotoxins in one clade, and the fourth in a separate clade (Supplementary Figure S4). One bacterial enterotoxin, from Leptospira mayottensis, formed an outgroup for the plant-infecting M. oryzae clade. M. oryzae resides outside the order Hypocreales. As such, the plant pathogenic enterotoxins at of this species were used as an outgroup and the tree was rooted on L. mayottensis (Supplementary Figure S4). This tree showed that some enterotoxins of ant-infecting fungi are related to those of other insect- or nematode-infecting species. Other enterotoxins formed their own clades. Within these ant infection-specific clades, enterotoxins from the two unilateralis complex species or the australis complex species were often paired. Ophiocordyceps subramanianii s.l. enterotoxins often formed the outgroup to a unilateralis- or australis-specific clade (Supplementary Figure S4). In addition, the enterotoxin that was highly up-regulated during manipulated biting behavior (i.e. GeneID Ophio5|373 Supplementary Figure S4), as well as conserved among all ant-infecting species in this study (and O. sinensis, see above), resided within a clade that only contained ant-manipulating species. This indicates that this particular enterotoxin could indeed be of key importance in establishing manipulated behavior as observed in the species included in this study.
Discussion
Much insight about the functioning of a species can be gained from its annotated genome. As such, we sequenced and annotated the genomes of Ophiocordyceps fungi that manipulate the behavior of their respective ant hosts. Additionally, we improved the assembly and annotation of a previously published genome. These species were collected across three continents (Africa, North America, and South America), and induced different biting behaviors in their respective ant host species. One of the species did not induce biting behavior, making the presence of manipulation less evident. To gain insight into the mechanisms these parasites employ, we compared their genetic information. The unilateralis and australis species of this genus had comparable genome features. Ophiocordyceps subramanianii s.l. was different in various aspects with the biggest difference being gene number and genome size. Our phylogenetic analysis indicated that Ophiocordyceps species within the same species complex (i.e. unilateralis and australis) form sister taxa. This is according to our expectations since fungi within a species complex are assumed to have closely co-evolved with taxonomically related ant host species (i.e. Camponotini and Ponerines for the unilateralis and australis clades respectively). In fact, there is growing evidence for the hypothesis that this close co-evolution has led to “one ant-one Ophiocordyceps species”11,13,20,26,27. Our read mapping efforts suggested that species within the australis complex are more closely related to each other than those residing within the unilateralis complex, even though one of them does not induce manipulated biting while the other australis does. Indeed, RNA-Seq reads from one unilateralis species could not be used to inform the annotation of another species within that complex due to low percentage mapping. Related to this is the fact that the two species in question (O. kimflemingiae and O. camponoti-rufipedis) come from separate continents (North vs. South America, respectively) and likely have very distinct biogeographical origins. In contrast, cross-mapping between australis species resulted in high percentage mapping. This again might be related to biogeography as South America and Africa were joined and their biota overlaps considerably28. The taxonomic broadness of the ant hosts that the unilateralis species infect might also explain their decreased relatedness. To illustrate, the tribe Camponotini consists of eight genera with the genus Camponotus containing 46 subgenera and over 900 species29. As such, the ant hosts infected by the species used in this study are part of different subgenera (Tanaemyrmex and Myrmothrix 30). The seemingly decreased genomic relatedness of these unilateralis species could thus be a result of a continued close co-evolution with their respective hosts.
Orthologous clustering demonstrated that the ant-infecting fungi sequenced in this study share general processes with other ascomycetes. These processes include transcription, translation, protein transport and signal transduction. However, a significant number of signal peptide-containing genes were not shared. These secreted proteins comprised a significant number of small secreted, potentially bioactive, proteins, and enterotoxins. These enterotoxins have largely been reported as bacterial toxins23,24,25, and have previously only been mentioned for a hand full of fungal species regarding their presence in the genome but without a discussed function18,22,31,32,33,34. The vast majority of the orthologous clusters that were specific for the five ant-manipulating species were also not shared across them. In fact, a significant part of the secretome appeared specific for each individual Ophiocordyceps species. Yet, the majority of the candidate manipulation genes, identified through RNA-Seq, were not unique to O. kimflemingiae nor to ant-infecting Ophiocordyceps species. They generally shared orthologs with at least one other ascomycete in this study. This suggests that the genes up-regulated during manipulated biting behavior in O. kimflemingiae, are largely not completely unique to that species. This could mean that these fungi do not use an exclusive set of novel genes to establish the respective complex behaviors observed. Instead, a variety of combinations of genes that are also found in other ascomycetes could be employed. The fungi sequenced in this study were collected on different continents where they induced different (levels of) manipulations (e.g. biting is present or absent) in different ant species. Their strategies at the genetic level could thus be very different as well. Alternatively, sampling during established biting behavior might not give us the correct pool of candidate manipulation genes. We found that a significant number of putatively secreted proteins are unique to the ant-manipulating fungi in this study. These proteins are however not significantly represented in the analyses regarding the candidate manipulation genes. It could be that we are missing out on many of the true candidates when we are sampling at the time when biting is already established. A better insight may be obtained by sampling during the time leading up to this event. This however demands that the suite of subtler behavioral changes leading up to the ultimate biting event is better quantified first.
We also looked more specifically at the conservation of secondary metabolite clusters that could play a role in the manipulation of ant behavior. Indeed, a variety of them are significantly up-regulated during manipulated biting behavior in infected C. castaneus ants18. We looked at three of these clusters from O. kimflemingiae in more detail, of which one contained a putative tryptophan dimethylallyltransferase. This enzyme facilitates the first step in the ergot alkaloid biosynthesis pathway35. Our analysis showed that secondary metabolite clusters that are activated during manipulated biting behavior are not conserved across Ophiocordyceps species complexes. They do however seem to be largely conserved within a species complex. Seemingly, secondary metabolite clusters are largely shared among unilateralis species that infect Camponotini. So, while these species seem rather distinct regarding their genes that encode for the suite of secreted enzymes, they might actually employ more similar secondary metabolite pathways. The suite of secondary metabolites involved in parasite-host interactions that lead to behavioral manipulation might thus translate rather well across Ophiocordyceps fungi within a species complex.
Ultimately, our efforts pointed towards a possible major role that enterotoxins might play in the parasite-host interactions between Ophiocordyceps fungi and their ant hosts. Not all ascomycetous genomes in our study contain secreted enterotoxins though. In contrast, ant-infecting Ophiocordyceps fungi contain many more genes in their genomes encoding for these secreted compounds compared to other ascomycetes. A significant proportion of these genes, therefore, appear to be specific to these species. Our orthologous clustering and phylogenetic tree analysis also suggested that fungi that reside within the same complex (unilateralis or australis) have genetically similar enterotoxins. Previous transcriptomics data of O. kimflemingiae suggests that enterotoxins are dynamically up- and down-regulated. Indeed, one of the enterotoxins looks especially interesting as it is extremely highly up-regulated during the manipulated biting event, after which it is immediately significantly down-regulated again18. This gene has orthologs across all five ant-infecting Ophiocordyceps species in this study. It could thus be a key component in establishing the altered behaviors we observe in infected ants. The function of this fungal enterotoxin is not clear at this time. However, the literature on bacterial enterotoxins in insect hosts indicates that it might impair the host’s chemosensory abilities23,24. More studies into fungal enterotoxins are needed, however, as well as functional genetics studies that would truly determine how Ophiocordyceps species might use these compounds to interact with their ant hosts.
Methods
Strain culturing and maintenance
Ophiocordyceps camponoti-rufipedis strain MAP-16 and Ophiocordyceps australis s.l. strain MAP-64 were isolated from an infected Camponotus and Ponerine worker ant respectively, collected in Mata do Paraíso forest reserve, Minas Gerais, Brazil. Ophiocordyceps australis s.l. strain 1348a and Ophiocordyceps subramanianii s.l. strain 1346 were isolated from Ponerine worker ants collected in Kukurantumi fetish forest in the Eastern Region of Ghana. Previously reported Ophiocordyceps unilateralis s.l. strain SC16a, collected in South Carolina, USA17,18, was also used in this study. This species has recently been described and named Ophiocordyceps kimflemingiae 20. Fungal cultures were kept in Grace’s insect media (Sigma) and supplemented with 10% (v/v) fetal bovine serum (Gibco). Mycelium for DNA and RNA isolation was grown in 100 mL flasks holding 40 mL media, shaken at 60 rpm at 28 °C.
DNA isolation, genome sequencing and assembly
Fungal mycelium was harvested by decanting liquid cultures over a Buchner funnel containing Whatmann filter paper. Tissue paper and pressure were applied to drain the mycelium from as much liquid as possible. Dried mycelium was snap frozen in liquid nitrogen and stored at −80 °C upon genomic DNA isolation. About 1 cm2 of the frozen mycelium was transferred to liquid nitrogen cooled 2 mL Eppendorf tubes (Greiner) with two metal beads (ø 4.76 mm) each. A Tissuelyser II (Qiagen) with a chilled adapter set (Qiagen) at 24 freq/s for 60 s was used to disrupt fungal cells. To extract DNA, 0.9 mL Extraction Buffer (1% SDS, 24 g/L PAS, 1x RNB (5x RNB: 12.11 g Tris/HCl, 7.304 g NaCl 9.51 EGTA in 100 mL, pH 8.5)) and 0.9 mL phenol/chloroform were used. After spinning down for 10 min at 10.000 rpm, the water phase was transferred to a new tube. Samples were purified using DNA Cleanup columns (Qiagen) and the manufacturer’s protocols. Following elution, samples were RNAse treated with RNAse mix (Ambion) for 10 min at 37 °C. Quality and quantity of the samples were checked on a Bioanalyzer System with the High Sensitivity DNA Analysis Kit (Agilent Technologies) and a Qubit with the DNA BR Assay Kit (Invitrogen).
In an attempt to avoid fragmentation bias, two genomic libraries were prepared for genome sequencing. Nextera libraries (Illumina) were prepared according to manufacturer’s instructions. Additionally, the NEBNext DNA Library Prep Master Mix Set for Illumina (New England Biolabs) was used. To this end, we sheared 5 µg genomic DNA of each sample to an approximate length of 800 bp with a Covaris M220 ultrasonicator with 50 µl screw caps. Fragments of 650–850 bp were selected with a BluePippin electrophoresis unit (Sage Science). A MiSeq sequencer (Illumina) was used to perform 2 × 300 bp paired-end sequencing.
Sequences were trimmed and assembled with CLC Genomics Workbench v7.5 (Qiagen) and default parameters. Not all libraries resulted in high-quality sequencing data. Assembly attempts integrating this lower quality data resulted in more fragmented assemblies. To improve our assemblies, we thus discarded sequencing runs that resulted in lesser quality reads. As such, O. camponoti-rufipedis from Brazil, strain Map-64 (O. australis s.l. from Brazil) and strain 1348a (O. australis s.l. from Ghana) were assembled using only sequenced Nextera libraries. For O. camponoti-rufipedis 93.8% of primary sequences assembled into 2206 scaffolds (minimum length 1000 bp, average coverage 29-fold) with a total genome size of 21.91 Mb. For Map-64, 98.2% of primary sequences assembled into 594 scaffolds (minimum length 1000 bp, average coverage 135-fold) with a total genome size of 23.32 Mb. For 1348a, 95.1% of primary sequences assembled into 2297 scaffolds (minimum length 1000 bp, average coverage 20-fold) with a total genome size of 22.19 Mb. Strain 1346 (O. subramanianii s.l. from Ghana) was assembled using only sequenced NEBNext libraries. This resulted in 96.0% of primary sequences assembling into 3401 scaffolds (minimum length 1000 bp, average coverage 19-fold) with a total genome size of 32.31 Mb. Previously reported strain SC16a (O. unilateralis s.l. from the USA, renamed O. kimflemingiae) was also reassembled. For this strain sequences from NEBNext and Nextera libraries could still be combined. This resulted in 93.5% of primary sequences assembling into 2537 scaffolds (minimum length 1000 bp, average coverage 130-fold) with a total genome size of 23.92 Mb (Table 1, and Supplementary Table S1). Repetitive sequences in the assembly were masked using RepeatMaker36, RepBase library37 and RepeatScout38.
RNA isolation, mRNA-Seq library construction and sequencing
We did not have enough mycelium from O. camponoti-rufipedis to generate an RNA sample to aid in gene prediction. For the other strains, enough mycelium was harvested so 1 cm2 mycelium could be used for RNA isolation. Total RNA was extracted as previously reported39. A Bioanalyzer System with the RNA 6000 Nano Kit (Agilent Technologies) and a Qubit with the RNA BR Assay Kit (Invitrogen) were used to verify sample quality and quantity. mRNA-Seq libraries were constructed with a NEBNext Ultra Directional RNA Library Prep Kit for Illumina (New England BioLabs). Quality and quantity of the resulting cDNA libraries were verified with a Qubit using the ds DNA HS Assay Kit (Invitrogen) and a Bioanalyzer System with the High Sensitivity DNA Analysis Kit (Agilent Technologies). Samples were sequenced on a HiSeq. 1500 (Illumina) using 100 bp paired-end sequencing in rapid-run mode.
Gene prediction
Genes were predicted for all Ophiocordyceps strains in this study, including the reassembled version of previously reported O. kimflemingiae strain SC16a18. To this end, Augustus version 3.0.240 was trained using BRAKER1 version 1.1.841. To aid in gene prediction, for all strains in this study (except O. camponoti-rufipedis), intron hints for Augustus were produced from Tophat transcript alignments using the Augustus software package. Manual inspection of the predicted genes revealed that neighboring genes were occasionally merged into one gene model, separated by a relatively large intron (data not shown). Therefore, a second round of gene prediction was performed. Here, the midpoints of introns larger than 150 bp and not supported by RNA-Seq alignments were labeled as ‘intergenic regions’ in the Augustus hints file. In the resulting gene predictions, these merged genes were usually correctly split into two genes. Completeness of the set of predicted proteins was analyzed using the CEGMA protein set42 using a BLASTp E-value cutoff of 1e-5, as well as BUSCO version 243 using the reference dataset ‘pezizomycotina_odb9’ (Table 1).
No RNA-Seq reads were available for O. camponoti-rufipedis to aid in gene prediction. To demonstrate that SC16a RNA-Seq reads could not be used directly to inform the gene prediction for this species, even though both species reside within the same species complex (O. unilateralis s.l.) reads were also mapped across genomes. This was done with CLC Genomics Workbench v7.5 (Qiagen) using default parameters. In this comparison, the assembled genome of Ophiocordyceps polyrhachis 22 was also included (this genome could however not be used in the comparative analyses below since its annotation has not been publically deposited). Instead of using the SC16a transcripts directly to predict genes in O. camponoti-rufipedis, we chose an approach that used them indirectly. Augustus version 3.0.240 was used to predict genes, using the RNA-Seq-informed parameter files previously generated by BRAKER1 version 1.1.841. for O. kimflemingiae SC16a (described above).
Functional annotation
We functionally annotated the predicted proteins of all Ophiocordyceps strains in this study. PFAM version 2744 was used to predict conserved protein domains, which were subsequently mapped to the corresponding gene ontology (GO) terms45,46. The MEROPS database47 with a BLASTp E-value cutoff of 1e-5 was used to predict proteases. Secretion signals were predicted with Signalp 4.148. Transmembrane domains were predicted with TMHMM 2.0c49. Proteins were considered small secreted proteins when they had a secretion signal, but no transmembrane domain (except in the first 40 amino acids) and were shorter than 300 amino acids. A pipeline based on the SMURF method50 was used to predict genes and gene clusters involved in secondary metabolism. SMURF parameter d (maximum intergenic distance in base pairs) was set at 3000 bp. SMURF parameter y (the maximum number of non-secondary metabolism genes upstream or downstream of the backbone gene) was set at 6.
In cases where additional homology searches were performed against the NCBI database, Blast2Go51 with BLASTp default settings was used.
Comparative analyses
For comparative purposes, 18 previously published genomes of fungal acomycetes were included that represent various fungal life styles within and outside the Order Hypocreales: Ophiocordyceps sinensis 52, Ophiocordyceps minnesotensis 53, Drechmeria coniospora 33, Tolypocladium inflatum 54, Cordyceps militaris55, Cordyceps bassiana 34, Metarhizium robertsii and Metarhizium acridum 32, Claviceps purpurea 56, Fusarium oxysporum 57, Fusarium graminearum 58, Trichoderma reesei 59, Trichoderma virens 60, Magnaporthe oryzae 31, Candida albicans 61, Aspergillus fumigatus 62, Neurospora crassa 63, Aspergillus nidulans 64. To estimate the phylogeny of these organisms, 67 conserved genes that were found present in each organism were aligned using MAFFT version 7.123b65. Well-aligned regions were identified using Gblocks 0.91b66. This resulted in 41793 amino acid positions. The parallelized version of RAxML version 8.1.1667 with the PROTGAMMAWAG model with 100 rapid bootstrap partitions was used to reconstruct a species tree. The tree was visualized using iTOL68.
Proteins that contained a ‘heat-labile enterotoxin alpha chain’ domain (PFAM domain PF01375) were identified as enterotoxins. The domains were aligned using MAFFT version 7.123b65. Next, a phylogenetic gene tree was reconstructed with FastTree 2.0 using default parameters69. The tree was visualized using iTOL68, and rooted on the heat-labile enterotoxin alpha chain’ domain of bacterium Leptospira mayottensis.
Clusters of orthologous genes among the five Ophiocordyceps genomes produced in this study, as well as the comparative genomes, were identified using OrthoMCL version 2.0.970 with an all-versus-all blastp E-value cutoff of 1e-5 and an inflation factor of 1.5. Venn diagrams demonstrating ortholog cluster commonalities between species (ranges) were visualized with http://bioinformatics.psb.ugent.be/webtools/Venn. Over- and under-representation of functional annotation terms were calculated using the Fisher Exact test with the Benjamini-Hochberg correction to correct for multiple testing (corrected p-value < 0.05).
Transcriptomics analysis
Previously published mixed transcriptomics data was used again in this study to further investigate fungal gene expression during and after manipulated biting behavior18. Sequence tags were mapped to the new genome version of strain SC16a (O. unilateralis s.l. from the USA) and differentially expressed genes were identified using the methods previously described18. Genes found to be significantly up-regulated during manipulated biting behavior followed by a significant down-regulation after were subjected to further comparative analyses.
Data availability
The Whole Genome Shotgun projects in this study have been deposited at DDBJ/EMBL/GenBank under the accession numbers LAZP02000000 (bio sample number SAMN03465111) and SUB2752196, (bio sample numbers SAMN07142922, SAMN07142923, SAMN07142924, SAMN07142925). Genome information can also be accessed through http://fungalgenomics.science.uu.nl/. The RNA-Seq data generated in this study is available through NCBI’s Sequence Read Archive under the biosample accessions SAMN07193086, SAMN07193087, SAMN07193088. All other data generated or analyzed during this study are included in this published article and its Supplementary Data files.
References
Qi, J., Ojika, M. & Sakagami, Y. Neuritogenic cerebrosides from an edible Chinese mushroom. Part 2: Structures of two additional termitomycesphins and activity enhancement of an inactive cerebroside by hydroxylation. Bioorg Med Chem 9, 2171–2177 (2001).
Qu, Y. et al. Termitomycesphins G and H, additional cerebrosides from the edible Chinese mushroom Termitomyces albuminosus. Biosci Biotechnol Biochem 76, 791–793, https://doi.org/10.1271/bbb.110918 (2012).
Vilcinskas, A. & Gotz, P. Parasitic fungi and their interactions with the insect immune system. Adv Parasit 43, 267–313, https://doi.org/10.1016/S0065-308x(08)60244-4 (1999).
Grimaldi, D. A. & Engel, M. S. Evolution of the insects. (Cambridge University Press, 2005).
Araújo, J. P. & Hughes, D. P. Diversity of entomopathogenic fungi: which groups conquered the insect body? Adv Genet 94, 1–39, https://doi.org/10.1016/bs.adgen.2016.01.001 (2016).
Vega, F. E. & Blackwell, M. 352 p. (Oxford University Press, 2005).
Molnár, I., Gibson, D. M. & Krasnoff, S. B. Secondary metabolites from entomopathogenic Hypocrealean fungi. Natural product reports 27, 1241–1275, https://doi.org/10.1039/c001459c (2010).
Boer, P. Observations of summit disease in Formica ruffa Linnaeus, 1761 (Hymenoptera: Formicidae). Myrmecological News 11, 63–66 (2008).
Malagocka, J., Grell, M. N., Lange, L., Eilenberg, J. & Jensen, A. B. Transcriptome of an entomophthoralean fungus (Pandora formicae) shows molecular machinery adjusted for successful host exploitation and transmission. J Invertebr Pathol 128, 47–56, https://doi.org/10.1016/j.jip.2015.05.001 (2015).
Malagocka, J., Jensen, A. B. & Eilenberg, J. Pandora formicae, a specialist ant pathogenic fungus: New insights into biology and taxonomy. J Invertebr Pathol 143, 108–114, https://doi.org/10.1016/j.jip.2016.12.007 (2017).
Araújo, J. P., Evans, H. C., Geiser, D. M., Mackay, W. P. & Hughes, D. P. Unravelling the diversity behind the Ophiocordyceps unilateralis (Ophiocordycipitaceae) complex: Three new species of zombie-ant fungi from the Brazilian Amazon. Phytotaxa 220, 224–238 (2015).
Evans, H. C., Elliot, S. L. & Hughes, D. P. Ophiocordyceps unilateralis: A keystone species for unraveling ecosystem functioning and biodiversity of fungi in tropical forests? Commun. Integr Biol 4, 598–602, https://doi.org/10.4161/cib.4.5.16721 (2011).
Evans, H. C., Elliot, S. L. & Hughes, D. P. Hidden diversity behind the zombie-ant fungus Ophiocordyceps unilateralis: four new species described from carpenter ants in Minas Gerais, Brazil. Plos One 6, e17024, https://doi.org/10.1371/journal.pone.0017024 (2011).
Hughes, D. P. et al. Behavioral mechanisms and morphological symptoms of zombie ants dying from fungal infection. BMC ecology 11, 13 (2011).
Hughes, D. P., Wappler, T. & Labandeira, C. C. Ancient death-grip leaf scars reveal ant-fungal parasitism. Biology letters 7, 67–70, https://doi.org/10.1098/rsbl.2010.0521 (2011).
Loreto, R. G., Elliot, S. L., Freitas, M. L., Pereira, T. M. & Hughes, D. P. Long-term disease dynamics for a specialized parasite of ant societies: a field study. Plos One 9, e103516, https://doi.org/10.1371/journal.pone.0103516 (2014).
de Bekker, C. et al. Species-specific ant brain manipulation by a specialized fungal parasite. BMC Evol Biol 14, 166 (2014).
de Bekker, C. et al. Gene expression during zombie ant biting behavior reflects the complexity underlying fungal parasitic behavioral manipulation. BMC genomics 16, 620, https://doi.org/10.1186/s12864-015-1812-x (2015).
Hughes, D. P. et al. From So Simple a Beginning: The Evolution of Behavioral Manipulation by Fungi. Genetics and Molecular Biology of Entomopathogenic Fungi 94, 437–469, https://doi.org/10.1016/bs.adgen.2016.01.004 (2016).
Araújo, J. P., Evans, H. C., Kepler, R. M. & Hughes, D. P. Zombie-ant fungi across continents: 14 new species and new combinations with Ophiocordyceps. I. Myrmecophilous Hirsutelloid species. Studies in Mycology (in press).
Chetouhi, C. et al. Cross-talk in host-parasite associations: What do past and recent proteomics approaches tell us? Infect Genet Evol 33, 84–94, https://doi.org/10.1016/j.meegid.2015.04.015 (2015).
Wichadakul, D. et al. Insights from the genome of Ophiocordyceps polyrhachis-furcata to pathogenicity and host specificity in insect fungi. BMC genomics 16, 881, https://doi.org/10.1186/s12864-015-2101-4 (2015).
Wiygul, G. & Sikorowski, P. P. The effect of staphylococcal enterotoxin B on pheromone production in fat bodies isolated from male boll weevils. J Invertebr Pathol 47, 116–119 (1986).
Wiygul, G. & Sikorowski, P. P. The effect of a heat-stable enterotoxin isolated from Escherichia coli on pheromone production in fat bodies isolated from male boll weevils. Entomol Exp Appl 60, 305–308 (1991).
Marrack, P. & Kappler, J. The staphylococcal enterotoxins and their relatives. Science 248, 1066 (1990).
Luangsa-Ard, J. et al. Purpureocillium, a new genus for the medically important Paecilomyces lilacinus. FEMS Microbiol Lett 321, 141–149, https://doi.org/10.1111/j.1574-6968.2011.02322.x (2011).
Kepler, R. M., Kaitsu, Y., Tanaka, E., Shimano, S. & Spatafora, J. W. Ophiocordyceps pulvinata sp nov., a pathogen of ants with a reduced stroma. Mycoscience 52, 39–47, https://doi.org/10.1007/s10267-010-0072-5 (2011).
Wegener, A. Die entstehung der Kontinente und Ozeane. Geologische Rundschau 3, 276–292 (1912).
Bolton, B., Alpert, G., Ward, P. S. & Nasrecki, P. (Harvard University Press, 2007).
Hansen, L. D. & Klotz, J. H. Carpenter ants of the United States and Canada. (Comstock Pub. Associates, 2005).
Dean, R. A. et al. The genome sequence of the rice blast fungus Magnaporthe grisea. Nature 434, 980–986, https://doi.org/10.1038/nature03449 (2005).
Gao, Q. et al. Genome sequencing and comparative transcriptomics of the model entomopathogenic fungi Metarhizium anisopliae and M. acridum. PLoS genetics 7, e1001264, https://doi.org/10.1371/journal.pgen.1001264 (2011).
Lebrigand, K. et al. Comparative genomic analysis of Drechmeria coniospora reveals core and specific genetic requirements for fungal endoparasitism of nematodes. PLoS genetics 12, e1006017, https://doi.org/10.1371/journal.pgen.1006017 (2016).
Xiao, G. et al. Genomic perspectives on the evolution of fungal entomopathogenicity in Beauveria bassiana. Scientific reports 2, 483, https://doi.org/10.1038/srep00483 (2012).
Tsai, H. F., Wang, H., Gebler, J. C., Poulter, C. D. & Schardl, C. L. The Claviceps purpurea gene encoding dimethylallyltryptophan synthase, the committed step for ergot alkaloid biosynthesis. Biochemical and biophysical research communications 216, 119–125, https://doi.org/10.1006/bbrc.1995.2599 (1995).
Smit, A. F. A. H., R. Green, P. RepeatMasker Open-3.0, www.repeatmasker.org (2010).
Jurka, J. et al. Repbase Update, a database of eukaryotic repetitive elements. Cytogenetic and genome research 110, 462–467, https://doi.org/10.1159/000084979 (2005).
Price, A. L., Jones, N. C. & Pevzner, P. A. De novo identification of repeat families in large genomes. Bioinformatics 21, i351–358, https://doi.org/10.1093/bioinformatics/bti1018 (2005).
de Bekker, C., Bruning, O., Jonker, M. J., Breit, T. M. & Wosten, H. A. Single cell transcriptomics of neighboring hyphae of Aspergillus niger. Genome biology 12, R71, https://doi.org/10.1186/gb-2011-12-8-r71 (2011).
Stanke, M., Diekhans, M., Baertsch, R. & Haussler, D. Using native and syntenically mapped cDNA alignments to improve de novo gene finding. Bioinformatics 24, 637–644, https://doi.org/10.1093/bioinformatics/btn013 (2008).
Hoff, K. J., Lange, S., Lomsadze, A., Borodovsky, M. & Stanke, M. BRAKER1: Unsupervised RNA-Seq-based genome annotation with GeneMark-ET and AUGUSTUS. Bioinformatics 32, 767–769, https://doi.org/10.1093/bioinformatics/btv661 (2016).
Parra, G., Bradnam, K., Ning, Z., Keane, T. & Korf, I. Assessing the gene space in draft genomes. Nucleic acids research 37, 289–297, https://doi.org/10.1093/nar/gkn916 (2009).
Simão, F. A., Waterhouse, R. M., Ioannidis, P., Kriventseva, E. V. & Zdobnov, E. M. BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31, 3210–3212, https://doi.org/10.1093/bioinformatics/btv351 (2015).
Finn, R. D. et al. Pfam: the protein families database. Nucleic acids research 42, D222–230, https://doi.org/10.1093/nar/gkt1223 (2014).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nature genetics 25, 25–29, https://doi.org/10.1038/75556 (2000).
Hunter, S. et al. InterPro: the integrative protein signature database. Nucleic acids research 37, D211–215, https://doi.org/10.1093/nar/gkn785 (2009).
Rawlings, N. D., Waller, M., Barrett, A. J. & Bateman, A. MEROPS: the database of proteolytic enzymes, their substrates and inhibitors. Nucleic acids research 42, D503–509, https://doi.org/10.1093/nar/gkt953 (2014).
Petersen, T. N., Brunak, S., von Heijne, G. & Nielsen, H. SignalP 4.0: discriminating signal peptides from transmembrane regions. Nature methods 8, 785–786, https://doi.org/10.1038/nmeth.1701 (2011).
Krogh, A., Larsson, B., von Heijne, G. & Sonnhammer, E. L. Predicting transmembrane protein topology with a hidden Markov model: application to complete genomes. Journal of molecular biology 305, 567–580, https://doi.org/10.1006/jmbi.2000.4315 (2001).
Khaldi, N. et al. SMURF: Genomic mapping of fungal secondary metabolite clusters. Fungal genetics and biology: FG & B 47, 736–741, https://doi.org/10.1016/j.fgb.2010.06.003 (2010).
Conesa, A. et al. Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21, 3674–3676, https://doi.org/10.1093/bioinformatics/bti610 (2005).
Hu, X. Z., Xiao, Y. J., Zheng, G. H., Xia, P., Zhang, Y. L., Leger, X. Y., Liu, R. J. & X.Z. Wang, C. S. St Genome survey uncovers the secrets of sex and lifestyle in caterpillar fungus. Chinese Science Bulletin 58, 2846–2854 (2013).
Lai, Y. et al. Comparative genomics and transcriptomics analyses reveal divergent lifestyle features of nematode endoparasitic fungus Hirsutella minnesotensis. Genome Biol Evol 6, 3077–3093, https://doi.org/10.1093/gbe/evu241 (2014).
Bushley, K. E. et al. The genome of Tolypocladium inflatum: evolution, organization, and expression of the cyclosporin biosynthetic gene cluster. PLoS genetics 9, e1003496, https://doi.org/10.1371/journal.pgen.1003496 (2013).
Zheng, P. et al. Genome sequence of the insect pathogenic fungus Cordyceps militaris, a valued traditional Chinese medicine. Genome biology 12, R116, https://doi.org/10.1186/gb-2011-12-11-r116 (2011).
Schardl, C. L. et al. Plant-symbiotic fungi as chemical engineers: multi-genome analysis of the clavicipitaceae reveals dynamics of alkaloid loci. PLoS genetics 9, e1003323, https://doi.org/10.1371/journal.pgen.1003323 (2013).
Ma, L. J. et al. Comparative genomics reveals mobile pathogenicity chromosomes in Fusarium. Nature 464, 367–373, https://doi.org/10.1038/nature08850 (2010).
Cuomo, C. A. et al. The Fusarium graminearum genome reveals a link between localized polymorphism and pathogen specialization. Science 317, 1400–1402, https://doi.org/10.1126/science.1143708 (2007).
Martinez, D. et al. Genome sequencing and analysis of the biomass-degrading fungus Trichoderma reesei (syn. Hypocrea jecorina). Nature biotechnology 26, 553–560, https://doi.org/10.1038/nbt1403 (2008).
Kubicek, C. P. et al. Comparative genome sequence analysis underscores mycoparasitism as the ancestral life style of Trichoderma. Genome biology 12, R40, https://doi.org/10.1186/gb-2011-12-4-r40 (2011).
Jones, T. et al. The diploid genome sequence of Candida albicans. Proceedings of the National Academy of Sciences of the United States of America 101, 7329–7334, https://doi.org/10.1073/pnas.0401648101 (2004).
Nierman, W. C. et al. Genomic sequence of the pathogenic and allergenic filamentous fungus Aspergillus fumigatus. Nature 438, 1151–1156, https://doi.org/10.1038/nature04332 (2005).
Galagan, J. E. et al. The genome sequence of the filamentous fungus Neurospora crassa. Nature 422, 859–868, https://doi.org/10.1038/nature01554 (2003).
Arnaud, M. B. et al. The Aspergillus Genome Database (AspGD): recent developments in comprehensive multispecies curation, comparative genomics and community resources. Nucleic acids research 40, D653–659, https://doi.org/10.1093/nar/gkr875 (2012).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30, 772–780, https://doi.org/10.1093/molbev/mst010 (2013).
Castresana, J. Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol Biol Evol 17, 540–552, https://doi.org/10.1093/oxfordjournals.molbev.a026334 (2000).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313, https://doi.org/10.1093/bioinformatics/btu033 (2014).
Letunic, I. & Bork, P. Interactive tree of life (iTOL)v3: an online tool for the display and annotation of phylogenetic and other trees. Nucleic acids research 44, W242–245, https://doi.org/10.1093/nar/gkw290 (2016).
Price, M. N., Dehal, P. S. & Arkin, A. P. FastTree 2–approximately maximum-likelihood trees for large alignments. Plos One 5, e9490, https://doi.org/10.1371/journal.pone.0009490 (2010).
Li, L., Stoeckert, C. J. Jr. & Roos, D. S. OrthoMCL: identification of ortholog groups for eukaryotic genomes. Genome research 13, 2178–2189, https://doi.org/10.1101/gr.1224503 (2003).
Acknowledgements
This work has been supported by funds to CdB from the Alexander von Humboldt Foundation and the Faculty of Biology and Genetics Section of the LMU. This work was additionally supported by NSF IOS-1558062 award to DPH. We would like to thank sciencetransmitter.com for assisting us with the figures accompanying this manuscript.
Author information
Authors and Affiliations
Contributions
H.C.E. and D.P.H. collected the fungal specimens used in this study. C.d.B. extracted DNA and RNA from them and A.B. performed the corresponding sequencing. Subsequent assembly, annotation and data analyses were done by C.d.B., R.A.O. and A.B. The manuscript was drafted by C.d.B. with valuable contributions from all other authors.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
de Bekker, C., Ohm, R.A., Evans, H.C. et al. Ant-infecting Ophiocordyceps genomes reveal a high diversity of potential behavioral manipulation genes and a possible major role for enterotoxins. Sci Rep 7, 12508 (2017). https://doi.org/10.1038/s41598-017-12863-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-12863-w
This article is cited by
-
Using machine learning to predict protein–protein interactions between a zombie ant fungus and its carpenter ant host
Scientific Reports (2023)
-
Manipulative neuroparasites: uncovering the intricacies of neurological host control
Archives of Microbiology (2023)
-
Bacillus predominates in the Ophiocordyceps pseudolloydii-infected ants, and it potentially improves protection and utilization of the host cadavers
Archives of Microbiology (2023)
-
The genus Entomophthora: bringing the insect destroyers into the twenty-first century
IMA Fungus (2021)
-
The evolutionary ecology of circadian rhythms in infection
Nature Ecology & Evolution (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.