Abstract
Bacterial multidrug resistance is a worrying health issue. In Gram-negative antibacterial research, the challenge is to define the antibiotic permeation across the membranes. Passing through the membrane barrier to reach the inhibitory concentration inside the bacterium is a pivotal step for antibacterial molecules. A spectrofluorimetric methodology has been developed to detect fluoroquinolones in bacterial population and inside individual Gram-negative bacterial cells. In this work, we studied the antibiotic accumulation in cells expressing various levels of efflux pumps. The assays allow us to determine the intracellular concentration of the fluoroquinolones to study the relationships between the level of efflux activity and the antibiotic accumulation, and finally to evaluate the impact of fluoroquinolone structures in this process. This represents the first protocol to identify some structural parameters involved in antibiotic translocation and accumulation, and to illustrate the recently proposed “Structure Intracellular Concentration Activity Relationship” (SICAR) concept.
Similar content being viewed by others
Introduction
A main challenge in antibacterial chemotherapy is to determine and carefully use the in situ parameters that modulate the activity of drugs in order to improve the efficacy of the antibacterial molecules (see the last WHO report, http://www.who.int/antimicrobial-resistance/en/). This is particularly important with the continuing emergence and the spreading of multidrug resistant bacteria (MDR) that contribute to therapeutic failure1,2,3,4. Several papers illustrate the intensity and relevance of the concern in clinical isolates and ask for a better understanding of membrane permeation and intracellular concentration of antibiotics, which is urgently required to combat MDR pathogens. Indeed, passing the membrane barrier to reach a threshold concentration inside the bacterial cell is a pivotal step in antibacterial action5,6,7,8,9. This key point has not been strongly addressed until now or only in few specific cases with characterized bacterial strains8,9,10,11.
As recently reported8, 12,13,14,15, there is a large collection of methods available to measure efflux activity, but their sensitivity and validity must be clearly demonstrated in the different conditions used. Thus, the lack of an appropriate well-defined assay is a serious bottleneck for the optimization of an antibiotic intra-bacterial accumulation, the first inescapable step for understanding antimicrobial activity. Recently, the intracellular accumulation of a fluoroquinolone has been studied with a synchrotron light source using intrinsic bacterial fluorescence as a powerful internal standard for correcting biological variations associated with bacterial samples16. In this case, the fluoroquinolone antibiotic fleroxacin exhibits sufficient changes in fluorescence intensity to monitor its accumulation in single bacterial cells14. Recent results allowed us to validate this method with a multidrug resistant clinical strain of Enterobacter aerogenes and an efflux pump deficient (ΔtolC) derivative, demonstrating the importance to use a robust internal control17.
The Innovative Medicine Initiative-funded consortium Translocation (www.imi.europa.eu/content/translocation) has defined as a main objective to study and dissect antibiotic translocation across bacterial membranes11. To achieve this goal, the consortium is setting new methods and new concepts to dissect this process8. Therefore, it is important to elaborate common standards and/or protocols for studying antibiotic accumulation and the correlation with the antibacterial activities in a series of well-defined isogenic strains. In this study, we propose a protocol for microspectrofluorimetry analyses on individual bacterial cells associated to biological/biochemical characterizations of appropriate whole bacterial samples. From this work, a new perspective emerges on how to capture and correlate the kinetics of antibiotic uptake inside bacterial cells in order to (i) understand the role of drug structure in the accumulation rate, and (ii) define the efficacy of the AcrB pump in relation to different quinolone structures during the time-course accumulation. This study contributes to the determination of parameters associated with the recently proposed concepts, “Resident Time Concentration Close to Target” (RTC2T) and “Structure Intracellular Concentration Activity Relationship” (SICAR)8.
Results
Antibiotic molecules
The structures of the three antibacterial molecules are presented in Fig. 1.
Chemical structure and antibacterial activities of fluoroquinolones used. SANO1 structure has been previously described25.
Antibacterial susceptibility and AcrAB
The activity of FLE, CIP and SANO1 were assayed on the E. coli AG100 and AG100A (AcrAB- derivative) strains and AG102 strain that overproduces AcrAB pump (Fig. 1). In general, the susceptibility was higher with AG100A. In contrast, AG102 was more resistant. In the presence of PAßN, an efflux pump blocker used at low concentration15, the susceptibilities were noticeably increased whatever the fluoroquinolone assayed (Fig. 1). This suggests the involvement of AcrAB in the susceptibility level while in contrast, no increase of bacterial susceptibility was observed in the presence of membrane permeabilizer suggesting that the outer membrane was not a significant barrier in these strains (data not shown).
Accumulation is dependent of AcrAB expression and of the fluoroquinolone
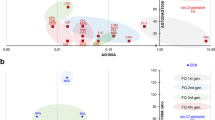
To correlate the antibacterial activity with drug uptake, the accumulations of the three molecules were studied in the three different strains. Figure 2 presented the results regarding the accumulation of FLE, CIP and SANO1 in bacterial population and demonstrated the effect of efflux expression on the intra-bacterial accumulation of the antibiotic: a very low content of the different drugs was observed in the AcrAB overproducer strain (AG102). When the incubation was performed in the presence of CCCP that collapses the energy driving force of efflux pump8, a significant increase of the accumulation was obtained in AG102 (Fig. 2). The plateau intensity reached level similar to the one we observed in the AcrAB deleted strain (AG100A). Moreover, in the parental strain, AG100, the fluorescence was less intense that the one obtained in AG100A suggesting that the basal efflux level present in this strain was sufficient to alter the accumulation inside the bacterial cell (Fig. 2 and Figure S1). It is important to note that the accumulation measured at 5 min in AcrAB- strain is related to the penetration capability for each tested molecule during early times of incubation in the absence of AcrAB pump activity. In contrast, the ratio of accumulation calculated at 15 min, AcrAB−/AcrAB++ or AcrAB-/AcrAB wild type, gives an illustration of the efflux capability to recognize the tested fluoroquinolones at steady state times of incubation16.
Intracellular concentration of Fleroxacin (FLE), Ciprofloxacin (CIP) and a benzo[b]napthyridone (SANO1) obtained with normalization by tryptophan. The accumulation was performed in AG100 (WT), AG100A (AcrAB−) and AG102 (AcrAB++) E. coli strains incubated 5, 15 and 30 min with 2 mg/L of molecules, in the absence or in the presence of CCCP (10 µM). The cells were lysated by HCl-glycine and the fluorescence signal (RFU) was plotted.
From the collected data (Figure S1 and Table 1), we can perform the following ranking:
-
For 5 min incubation: in AcrAB- strain, SANO1 > CIP > FLE; in WT strain: SANO1 > CIP > FLE; in AcrAB overproducer strain: SANO1 > CIP > FLE.
-
For 15 min incubation: in AcrAB- strain, SANO1 > CIP > FLE; in WT strain: SANO1 > CIP > FLE; in AcrAB overproducer strain: SANO1 > CIP > FLE.
The various values representing the ratio of fluoroquinolone accumulation, AG100A/AG100 and AG100A/AG102, R and R′ respectively, are presented in Table 1. These ratios give an evaluation of the efflux effect on the accumulation of the three fluoroquinolones.
Interestingly, the SANO1 molecule seemed to be very efficiently internalized at 5 min of incubation, 6–10 fold compared to CIP and FLE respectively, in the AcrAB- strain. However, it is also rapidly expelled by AcrB pump compared to FLE and CIP (Table 1) when we compared the accumulation ratio R or R′. This suggests that the molecule can have a better affinity for the AcrB sites and represents a better substrate for this pump. Importantly, no significant change in cell viability was observed during this period that corresponds to accumulation assay (Figure S3).
Accumulation in individual bacterial cells
In order to follow the intracellular accumulation on individual bacterial cells, we performed time-lapse experiments on the DUV microscope. Bacteria were plated and observed for 30 min with a sampling time of 2 min: thumbnails of a bacterium incubated with FLE were plotted in Fig. 3A. Illustrations of CIP and SANO1 accumulation in individual bacterial cells under the same conditions were presented in Figure S4.
Fluorescence intensities corrected from background, photobleaching and crosstalking17 corresponding to about 30–40 individual cell measurements are presented in Fig. 3B. Interestingly, the steady state level of molecule (SANO1) accumulation in AcrAB- strain was obtained at 5–10 min under these conditions while a slow increase was observed in the parental strain expressing AcrAB basal level. In contrast, no significant accumulation was obtained in the AcrAB overproducer.
We extended the analysis to other quinolone molecules and the means are presented in Fig. 4. Generally, no significant accumulation was noted in the AcrAB overproducer, whatever the molecule tested during the 30 min assay. Regarding the accumulation in parental strain, we observed a small increase with the three molecules, similar for FLE and SANO1, and reduced with CIP. Importantly, in the AcrAB deleted strain, the steady state was rapidly obtained for SANO1, after 10 min of incubation, whereas a continuous increase was noted with FLE and CIP during the same time-course accumulation assay (Fig. 4).
Microspectrofluorimetry on individual bacterial strains incubated with FLE/CIP/SANO1 at 2 mg/L calculated from curves as presented in Fig. 3B for SANO1.
Regarding the profile of SANO1 accumulation, it is important to determine if the plateau observed at 10 min and followed by a curve reflecting a decrease accumulation in individual cells indicated a saturation process or a degradation of SANO1 inside the bacterial cell under these incubation conditions. To evaluate the stability of the compounds inside the cell, the molecules, FLE or SANO1, were incubated with a total bacterial lysate in the same conditions and the signal was monitored during 15 min. Interestingly, the signal recorded with FLE was stable during this interval when SANO1 exhibited an important decrease of the signal intensity during the incubation (Figure S5). This suggests that internalized SANO1 was less stable than FLE due to UV irradiation combined to the bacterial degradative activities or quenched during the binding to the target under these conditions.
Discussion
With the continuous emergence of antibiotic resistance in Gram-negative bacteria, a main objective is to understand the relationships between antibiotic activity and the expression levels of efflux pumps expelling the penetrating molecules7, 18,19,20,21. A prominent question is therefore to relate the intra-bacterial accumulation and the chemical structure of the antibiotic8.
A variety of methods has been previously reported to follow the intracellular accumulation of antibiotics in Enterobacteriaceae expressing membrane-associated mechanisms of resistance including outer membrane impermeability or efflux pumps that expel all antibiotic classes8, 12,13,14,15, 22, 23. Until recently, no robust assay has been developed to study accumulation in individual bacterial cells. Cinquin et al.17 reported a fluorimetric method that allows the monitoring of the accumulation of fleroxacin in individual resistant bacteria17. This new methodology offers us the opportunity to examine in depth the time course accumulation by using a robust internal control (bacterial fluorescence) to standardize the signals and simultaneously correlate the imaging studies obtained with several strains to the variation of efflux activity in these strains. Moreover, this approach is not dependent to time-consuming protocols that can induce experimental artefacts8. It is important to adapt and use this original method to compare the accumulation of different quinolones in various efflux backgrounds and to try and correlate drug concentration and activity.
In this study, we used three molecules belonging to the structural group of fluoroquinolones and exhibiting different antibacterial activities against E. coli strains expressing various level of the AcrAB pump. Our aims were, (i) to determine the intra-bacterial concentration of ciprofloxacin, fleroxacin and an unmarketed fluoroquinolone of the benzo-napthyridone subfamily, SANO1, by conjointly using fluorimetric detection on bacterial population and individual bacterial cells; and (ii) to analyze the early steps of drug accumulation and the role of the AcrAB efflux pump. First of all, the values clearly indicate a MIC modulation strongly associated with the level of AcrAB in this collection: the susceptibility being comprised between 0.004 to 0.125 mg/L for the AcrAB- strain and to 0.0625 to 2 mg/L for the AcrAB overproducer, depending of the fluoroquinolone assayed. In parallel, we observe a marked difference in the accumulation profile depending on the quinolone structure. During the very early times of accumulation in individual bacterial cells devoid of efflux, SANO1 exhibits the highest rate of uptake compared to FLE and CIP, suggesting a well-adapted structure for permeation and a positive involvement of side chains in membrane translocation.
However, SANO1 is also very well recognized by the AcrB pump, since it is rapidly and efficiently expelled as reported by the ratio of signals observed in efflux producer strain in the absence and in the presence of CCCP, an ionophore that collapses the driving force of AcrAB-TolC9, 10. Regarding CIP and FLE, the accumulation is also impaired in the AcrAB overproducer strain compared to AcrAB- strain, but to a lesser extent compared to SANO1. Regarding the curves, it is important to mention that we observed an effect on accumulation rate for the first incubation time (compare AG100A to AG100 curve) suggesting that the efflux activity is observed early. This suggests that, as soon as drug molecules permeate the cell envelope, AcrB transporter recognizes them and pumps antibiotics out of the cell. This is the case for the parental strain and for the overproducer strain with a noticeable shift in the curve at longer incubation times for the latter that reflects an additional reduction in the accumulation rate. These observations, also noted in individual cell assays, suggest that the activity of AcrB pump could be partly saturated when expressed at a basal level. This efflux impact on the molecule accumulation is correlated to the antibiotic susceptibility determined in these strains, although the MIC are recorded after 18 hours of incubation, a period that could buffered the effect of different accumulation rates. In addition, the effect of PAßN, which has been reported as a pump blocker at low concentration10, 24, on the antibiotic susceptibility is in accordance with accumulation studies carried out with these strains expressing various level of AcrAB efflux transporter.
It is tempting to correlate the accumulation ratio obtained in each strain with their susceptibility, but it is important to mention that the time scale is different, few minutes compared to an overnight incubation. Moreover, we have selected a ratio fluoroquinolone concentration/bacterial number that is different to the recommendation rules used for MIC determination in order to have a limited effect on cell viability during the experimentation times. One objective of IMI-Translocation consortium is to develop an assay allowing us to simultaneously measure antibiotic concentration inside the bacterial cells and rate-killing8, 11.
This is the first time that we studied the real time accumulation of antibiotics belonging to the same structural family in various efflux pump backgrounds. If we calculate the impact of efflux level on the accumulation of the three fluoroquinolones, from efflux minus to overexpressed efflux background, called R and R′ respectively in Table 1, we obtain a specific index that reflects the relationships between the chemical structure of the molecule and the efflux activity on the accumulation level. This index fits in well with the new concept “Structure Intracellular Concentration Activity Relationship” (SICAR) recently proposed8. Practically, the SICAR connects the physicochemical drug properties to the efficacy of translocation through the bacterial membrane and the resulting intracellular accumulation. With this study, we can conclude that: (i) SANO1 exhibits a high penetration rate (9.26 10−11 µg/bacterial cell at 5 min) corresponding to a favorable SICAR influx index (SICARIN defined as the intracellular concentration obtained in the AcrAB- strain during 5 min of incubation), better than the corresponding index of CIP and FLE; and (ii), in contrast, the CIP and FLE present a quasi-similar SICAR efflux index (SICAREF defined as the ratio of molecule accumulated in AcrAB- strain to AcrAB producer strain) while SANO1 exhibits an unfavorable SICAREF that reflects a high level of sensitivity to efflux transporter (AcrB in this case). In the case of SANO1, several exposed chemical side chains could be involved in the affinity for AcrB binding sites when we compare the three structures. These SICAR indexes can be used to precise the chemotypes that are related to permeation of different antibiotics belonging to the same group. These indexes could be determined for other antibiotic classes by extending the assay to mass spectrometry analyses in order to take into account non-fluorescent molecules14.
A further study could be to sequentially modulate these putative pharmacophoric groups in order to precise their respective involvement in influx versus efflux, and to integrate the affinity constant in the matrix to complete the SICAR index. Conjointly to the use of adjuvants26,27,28,29,30 in combination to restore antibiotic susceptibility, this study opens a way to circumvent the role of membrane permeability and efflux transporters in resistance by a rationale pharmacomodulation taking into account the molecular profile, SICAR index, of antibiotic molecules.
Methods
Bacteria and Media
Escherichia coli strains used in this study are listed in Fig. 1.
Drug susceptibility assays
Fleroxacin (FLE), ciprofloxacin (CIP) and benzoquinolone (SANO1) were assayed to study the antibiotic susceptibility of E. coli strains. MIC values of antibiotics were determined by the microdilution method (CLSI) in liquid Mueller Hinton II media by using the twofold standard microbroth dilution method (microplates and automatic analyses Tecan®) (CLSI, http://clsi.org/). When required the MICs were determined in the presence of the phenylalanine arginine β -naphthylamide (PAβN) used at 20 mg/L. MIC values were read after 18 h of incubation at 37 °C. Experiments were carried out in triplicate and the resulting medians were presented.
Accumulation protocol
Bacteria grown at 37 °C in Luria-Bertani broth to mid-exponential phase (corresponding to 0.6 optical density units at 600 nm) were concentrated 10-fold. Briefly, the bacterial suspension was centrifuged at 6,000 × g for 15 min at 20 °C and pellets were re-suspended in 1/10 of the initial volume in sodium phosphate buffer (50 mM) at pH 7 supplemented with MgCl2 (5 mM) (NaPi-MgCl2 buffer) to obtain a density of 6 109 CFU.ml−1. In culture tubes, 4 ml of the bacterial suspension was incubated 5, 15 and 30 min at 37 °C (final volume 5 ml) with FLE, CIP or SANO1 at 2 mg/L, in the absence or in the presence of the efflux blocker CCCP used at 10 μM that collapses the energy-driven force needed by the efflux pump15. Bacterial suspensions incubated without antibiotics, with or without CCCP, were used as controls. Suspensions (800 μL) were then loaded on 1 M sucrose cushion (1,100 μL) and centrifuged at 9,000 × g for 5 min at 4 °C to eliminate extracellular-adsorbed compounds and to collect the washed bacteria.
After centrifugation, pellets corresponding to 800 μL of bacterial suspensions were lysed with 500 μL of 0.1 M Glycin-HCl pH 3 overnight at room temperature. After a centrifugation for 15 min at 9,000 × g at 4 °C, 400 μL of lysates were mixed with 600 μL of Glycin-HCl buffer (FLE and CIP) or 3 M Tris pH 8.8 buffer (SANO1) and analyzed by spectrofluorimetry (Ex 290 nm, Em 420–480 for FLE; Ex 275 nm, Em 420–480 nm for CIP; Ex 275 nm, Em 499–529 for SANO1). Calibration curves were carried out to determine the quantity of molecules accumulated per cell. Various concentrations of FLE, CIP or SANO1 were mixed with bacteria lysates at OD = 4.8 and measured with spectrofluorimeter (n = 3).
To control that bacterial cells are alive during the experimental time, the number of colony forming units (CFU) were determined by sampling the bacterial suspension during antibiotic incubation. Small volumes were collected and dilutions were performed and plated on Petri dishes. CFUs were measured and survival ratios, with and without fluoroquinolone, were generated. The assays were performed in triplicate. No significant change in cell viability was observed during this period that corresponds to accumulation assay.
It must be noted that the ratio cell/antibiotic concentration were different in the MIC assay and in accumulation assay carried out using starving conditions during a limited incubation time (5–30 min) as previously determined17.
DUV Fluorescence accumulation in individual bacterial cells
The accumulation in individual bacterial cells was monitored directly under the deep ultraviolet (DUV) microscope during 30 min. Bacteria were concentrated in order to obtain an OD of 4.8 in NaPi-MgCl2 buffer. Then 120 µL of the suspension were centrifuged on a 1 M sucrose cushion (165 μL) at 9,000 × g for 5 min at 4 °C. The pellets were resuspended extemporaneously in 40 µL of NaPi-MgCl2 buffer containing or not FLE, CIP or SANO1 (2 µg/mL) in the absence or in the presence of CCCP (10 µM). 0.5 µL of resuspended pellets were deposited between two quartz coverslips and analyzed by DUV fluorescence imaging at DISCO Beamline. Bacterial cells were first located in brightfield before excitation in DUV under a microscope (Zeiss Axio Observer Z-1) at Synchrotron SOLEIL31. Emission was collected through a Zeiss ultrafluar objective at 100x with glycerin immersion. The FLE/CIP/SANO1 fluorescence was recorded by exciting at 290/275/275 nm, using a dichroic mirror at 300 nm (OMEGA Optical, Inc., USA) through an emission bandpass filter at 420–480 nm (OMEGA Optical, Inc., USA) for FLE and CIP and a 499–529 nm (SEMROCK) bandpass filter for SANO1. For the tryptophan fluorescence, the emission was passing through an emission bandpass filter at 327–353 nm (SEMROCK). Fluorescence images were recorded by a BUV EM CCD (Princeton PIXIS 1024 BUV). The whole setup (microscope, stages, filters, camera) was controlled by Micro-Manager32. Bacteria were observed for 30 min with a sampling time of 2 min for each area: 30 s with the filter 1 (fluoroquinolone molecule), 30 s with the filter 2 (tryptophan), followed by 60 s of pause. During the intervening pause in sampling, the same acquisition cycle was performed on another bacterium, avoiding constant UV irradiation of the same field.
The images were analyzed with Image J (Rasband, W.S., ImageJ, U. S. National Institutes of Health, Bethesda, Maryland, USA, http://imagej.nih.gov/ij/)33. The illumination heterogeneities were corrected before background subtraction. First, threshold was automatically adjusted using a triangle algorithm; thereafter, bacteria were analyzed as the remaining particles. The mean intensity coming from each bacterium was automatically calculated considering its pixel area. Finally, all bacteria signal taken from one image were averaged. For each condition, two different localizations with minimum 30 bacteria per field of view were recorded and averaged.
Stability assays
UV irradiation and incubation inside bacteria can promote some degradation of chemicals. In order to check the stability of fluoroquinolone during the assays, incubations of FLE and SANO1 were performed using a bacterial lysate obtained after cell disruption of AG100A strain using the method previously described15. The same conditions of incubation (lysate corresponding to same cell density, drug concentration, UV irradiation exposure) were used and the fluorescence was recorded each min during the assay. The percentage of fluoroquinolone stability was calculated using the initial fluorescence obtained at 0 min incubation as 100% for each molecule.
References
Watkins, R. R. & Bonomo, R. A. Overview: global and local impact of antibiotic resistance. Infect. Dis. Clin. North Am. 30, 313–322 (2016).
Laxminarayan, R. et al. Access to effective antimicrobials: a worldwide challenge. Lancet 387, 168–175 (2016).
Jones, D. News and analysis: the antibacterial lead discovery challenge. Nat. Rev. Drug Discov. 9, 751–752 (2010).
Kollef, M. H. et al. The intensive care medicine research agenda on multidrug-resistant bacteria, antibiotics, and stewardship. Intensive Care Med. doi:10.1007/s00134-017-4682-7 (2017).
Nikaido, H. Molecular basis of bacterial outer membrane permeability revisited. Microbiol. Mol. Biol. Rev. 67, 593–656 (2003).
Pagès, J. M., James, C. E. & Winterhalter, M. The porin and the permeating antibiotic: a selective diffusion barrier in Gram-negative bacteria. Nat. Rev. Microbiol. 6, 893–903 (2008).
Blair, J. M., Webber, M. A., Baylay, A. J., Ogbolu, D. O. & Piddock, L. J. Molecular mechanisms of antibiotic resistance. Nat. Rev. Microbiol. 13, 42–51 (2015).
Masi, M., Réfrégiers, M., Pos, K. M. & Pagès, J. M. Mechanisms of envelope permeability and antibiotic influx/efflux in Gram negative bacteria. Nat. Microbiol. 2, 17001 (2017).
Nikaido, H. & Pagès, J. M. Broad-specificity efflux pumps and their role in multidrug resistance of Gram-negative bacteria. FEMS Microbiol. Rev. 36, 340–363 (2012).
Bolla, J. M. et al. Strategies for bypassing the membrane barrier in multidrug resistant Gram-negative bacteria. FEBS lett. 585, 1682–1690 (2011).
Stavenger, R. A. & Winterhalter, M. TRANSLOCATION project: how to get good drugs into bad bugs. Sci. Transl. Med. 6, 228ed7 (2014).
Blair, J. M. & Piddock, L. J. How to measure export via bacterial multidrug resistance efflux pumps. MBio. 7, pii: e00840–16, doi:10.1128/mBio.00840-16 (2016).
Zhou, Y. et al. Thinking outside the “bug”: a unique assay to measure intracellular drug penetration in gram-negative bacteria. Anal. Chem. 87, 3579–3584 (2015).
Davis, T. D., Gerry, C. J. & Tan, D. S. General platform for systematic quantitative evaluation of small-molecule permeability in bacteria. ACS Chem. Biol. 9, 2535–2544 (2014).
Phetsang, W. et al. Fluorescent trimethoprim conjugate probes to assess drug accumulation in wild type and mutant Escherichia coli. ACS Infect. Dis. 2, 688–701 (2016).
Kaščáková, S., Maigre, L., Chevalier, J., Réfrégiers, M. & Pagès, J. M. Antibiotic transport in resistant bacteria: synchrotron UV fluorescence microscopy to determine antibiotic accumulation with single cell resolution. PLoS One 7(6), e38624 (2012).
Cinquin, B. et al. Microspectrometric insights on the uptake of antibiotics at the single bacterial cell level. Sci. Rep. 5, 17968 (2015).
Li, Z. H., Plesiat, P. & Nikaido, H. The challenge of efflux-mediated antibiotic resistance in Gram-negative bacteria. Clin. Micro. Rev. 28, 337–418 (2015).
Zgurskaya, H. I., López, C. A. & Gnanakaran, S. Permeability barrier of Gram-negative cell envelopes and approaches to bypass it. ACS Infect. Dis. 1, 512–522 (2015).
Nikaido, H. Prevention of drug access to bacterial targets: permeability barriers and active efflux. Science 264, 382–388 (1994).
Davin-Regli, A. et al. Membrane permeability and regulation of drug “influx and efflux” in enterobacterial pathogens. Curr. Drug Targets 9, 750–759 (2008).
Brown, A. R. et al. A mass spectrometry-based assay for improved quantitative measurements of efflux pump inhibition. PLoS One. 10, e0124814 (2015).
Carey, P. R. & Heidari-Torkabadi, H. New techniques in antibiotic discovery and resistance: Raman spectroscopy. Ann. NY Acad. Sci. 1354, 67–81 (2015).
Misra, R., Morrison, K. D., Cho, H. J. & Khuu, T. Importance of real-time assays to distinguish multidrug efflux pump-inhibiting and outer membrane-destabilizing activities in Escherichia coli. J. Bacteriol. 197, 2479–2488 (2015).
Bacqué, E. et al. Preparation of 8-azetidinobenzo[b][1,8]napthyridin-4-one-3-carboxylates as antibacterial agents. EP 536035 (1993).
Mamelli, L. et al. New antibiotic molecules: bypassing the membrane barrier of Gram negative bacteria increases the activity of peptide deformylase inhibitors. PLoS One 4, e6443 (2009).
Zabawa, T. P., Pucci, M. J., Parr, T. R. Jr. & Lister, T. Treatment of Gram-negative bacterial infections by potentiation of antibiotics. Curr. Opin. Microbiol. 33, 7–12 (2016).
Blair, J. M., Richmond, G. E. & Piddock, L. J. Multidrug efflux pumps in Gram-negative bacteria and their role in antibiotic resistance. Future Microbiol. 9, 1165–1177 (2014).
Alibert, S. et al. Multidrug efflux pumps and their role in antibiotic and antiseptic resistance: a pharmacodynamic perspective. Expert Opin. Drug Metab. Toxicol. 13, 301–309 (2017).
Vardakas, K. Z. & Falagas, M. E. Colistin versus polymyxin B for the treatment of patients with multidrug-resistant Gram-negative infections: a systematic review and meta-analysis. Int. J. Antimicrob. Agents 49, 233–238 (2017).
Jamme, F. et al. Synchrotron UV fluorescence microscopy uncovers new probes in cells and tissues. Microsc. Microanal. 16, 507–514 (2010).
Edelstein, A., Amodaj, N., Hoover, K., Vale, R. & Stuurman, N. Computer control of microscopes using µManager. Curr. Protoc. Mol. Biol. Chapter 14: Unit14.20, doi:10.1002/0471142727.mb1420s92 (2010).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Acknowledgements
We thank Anne Davin-Regli, Muriel Masi, Robert Stavenger and Mathias Winterhalter for their fruitful discussions, Anne-Marie Tran, Valérie Rouam and Blandine Pineau for technical assistance during the assays. The research leading to these results was conducted as part of the TRANSLOCATION consortium and has received support from the Innovative Medicines Initiatives Joint Undertaking under Grant Agreement n°115525, resources which are composed of financial contribution from the European Union’s seventh framework program (FP7/2007-2013) and EFPIA companies in kind contribution. This work was also supported by Aix-Marseille Univ. and Service de Santé des Armées, and by Soleil program (projects #20130949, 20140047, 20141262, 20150318).
Author information
Authors and Affiliations
Contributions
M.R. and J.-M.P. designed the research. J.V., E.D., B.C., L.M., and E.B. performed the various chemical characterization, measured the antibacterial activities and the molecule accumulation. J.V., E.D., B.C., J.P. and L.M., carried out the fluorescence and accumulation assays. M.M., M.R. and J.-M.P. supervised the overall project and wrote the manuscript with contribution from the other authors.
Corresponding author
Ethics declarations
Competing Interests
The authors (M.M., E.B.) declare competing financial interests: M.M. and E.B. are current employees of SANOFI R&D.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Vergalli, J., Dumont, E., Cinquin, B. et al. Fluoroquinolone structure and translocation flux across bacterial membrane. Sci Rep 7, 9821 (2017). https://doi.org/10.1038/s41598-017-08775-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-08775-4
This article is cited by
-
A framework for dissecting affinities of multidrug efflux transporter AcrB to fluoroquinolones
Communications Biology (2022)
-
Toxicity and bacterial anti-motility activities of the hydroethanolic extract of Acacia senegal (L.) Willd (Fabaceae) leaves
BMC Complementary Medicine and Therapies (2021)
-
Porins and small-molecule translocation across the outer membrane of Gram-negative bacteria
Nature Reviews Microbiology (2020)
-
The challenge of intracellular antibiotic accumulation, a function of fluoroquinolone influx versus bacterial efflux
Communications Biology (2020)
-
Spectrofluorimetric quantification of antibiotic drug concentration in bacterial cells for the characterization of translocation across bacterial membranes
Nature Protocols (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.