Abstract
Zoonotic spillover of sarbecoviruses (SarbeCoVs) from non-human animals to humans under natural conditions has led to two large-scale pandemics, the severe acute respiratory syndrome (SARS) pandemic in 2003 and the ongoing COVID-19 pandemic. Knowledge of the genetic diversity, geographical distribution, and host specificity of SarbeCoVs is therefore of interest for pandemic surveillance and origin tracing of SARS-CoV and SARS-CoV-2. This study presents a comprehensive repository of publicly available animal-associated SarbeCoVs, covering 1,535 viruses identified from 63 animal species distributed in 43 countries worldwide (as of February 14,2023). Relevant meta-information, such as host species, sampling time and location, was manually curated and included in the dataset to facilitate further research on the potential patterns of viral diversity and ecological characteristics. In addition, the dataset also provides well-annotated sequence sets of receptor-binding domains (RBDs) and receptor-binding motifs (RBMs) for the scientific community to highlight the potential determinants of successful cross-species transmission that could be aid in risk estimation and strategic design for future emerging infectious disease control and prevention.
Similar content being viewed by others
Background & Summary
Coronaviruses (CoVs) are a group of enveloped viruses belonging to Coronaviridae and currently contain four known genera, Alpha-, Beta-, Gamma-, and Delta-CoVs, that vary in their distribution, host species, and pathogenicity to humans. Sarbecovirus (SarbeCoV), a subgenus within Beta-CoV, has resulted in the emergence of the highly pathogenic human viruses SARS-CoV and SARS-CoV-2. SARS-CoV caused more than 8000 confirmed cases in 2002–20031, whereas SARS-CoV-2, a causative agent of COVID-19, has rapidly infected the global population with over 762 million confirmed cases (https://covid19.who.int/), and remains a significant threat to global health and the economy. Furthermore, evidence suggests that SarbeCoVs have high recombination and mutation rates, allowing them to infect and survive in different hosts worldwide2. Thus, recent research has intensely focused on surveys of SarbeCoVs carried by susceptible animals to enhance our knowledge of viral diversity, host specificity, and geographical distribution.
The origins of SARS-CoV and SARS-CoV-2 remain controversial due to the remaining genomic differences3. It is generally been thought that both SARS-CoV and SARS-CoV-2 originated in bats, and zoonotic spillover to humans has likely occurred through one or more intermediate hosts. Recently, several studies have shown that potential intermediate hosts may include Malayan pangolins, rabbits, ferrets, foxes, raccoons and dogs because the spike (S) protein of SARS-CoV and SARS-CoV-2 is capable of binding to their angiotensin-converting enzyme (ACE2), which facilitates virus entry1,4. In addition, molecular and serological evidence has revealed the reverse zoonotic potential of SARS-CoV-2 infection in several pets and domestic animals from different countries. Zoo tigers, lions, snow leopards, and pumas and domestic cats, dogs and minks have been confirmed to naturally acquire SARS-CoV-2 infection5. Despite no conclusive evidence that domestic animals can actively spill back SARS-CoV-2 to humans, the potential human-animal-human transmission cycle needs to be recognized and further investigated6. Thus, a better understanding of existing viral populations along with their ecological characteristics would be of importance for detecting potential interspecies spillover7.
Given that molecular techniques are widely applied in the identification and functional analyses of viruses, comprehensive retrieval of the virus sequences along with related meta-information facilitates in-depth research on the origin and evolution of SARS-CoV and SARS-CoV-2 among different animal hosts. However, relevant information such as virus sequence, host species, sampling time, and location has not been uniformly recorded and is only sporadically available in GenBank records or related literature. Furthermore, the considerable number of human-derived SarbeCoVs undoubtedly complicates the screening process of animal-associated SarbeCoVs from the public domain. Therefore, we established a sequence-centric dataset for the curation of related meta-information of animal-associated SarbeCoVs. Thus far, this dataset contains 1,535 SarbeCoVs identified from 63 different animal species globally.
Methods
Data collection
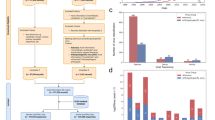
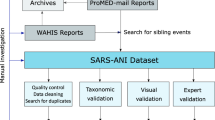
The data collection and inclusion procedures are outlined in Fig. 1a. To retrieve all known sequences from the public domain, an initial search within GenBank8 was performed using the keywords (“Sarbecovirus” OR “SARS” OR “Severe acute respiratory syndrome”) AND (“viruses” OR “virus”). A total of 6,740,876 GenBank records that matched the above keywords were retrieved and stored in a local system (as of February 14,2023). Despite conducting an exhaustive search, there is no guarantee that all records were collected. The possibility of missed SarbeCoVs may be inevitable in certain cases due to misclassification of sarbecoviruses by the submitters or historical changes in taxonomy. For instance, the term sarbecovirus was proposed as a novel subgenus within the genus Beta-CoV according to the ICTV demarcation criteria in 2017. Before that, some of the identified SARS-related coronaviruses (SARSr-CoVs) may have been categorized into the unclassified Beta-CoV in the NCBI taxonomy database. To ensure the integrity of data collection, a complementary search using the additional keywords (“Betacoronavirus” NOT “Homo sapiens” NOT “Embecovirus” NOT “Hibecovirus” NOT “Merbecovirus” NOT “Nobecovirus”) AND (“viruses” OR “virus”) was conducted with specific attention given to collect unclassified Beta-CoVs that might belong to SarbeCoVs. After removing the duplicates from the two search results, there were 6,742,282 records possibly associated with SarbeCoVs that needed further review.
Schematic diagram of data curation. (a) Flow diagram of the data search, screening and validation process. (b) Bubble chart of the annual and geographical distributions of presently identified animal-associated SarbeCoVs. The area of each bubble correlates with the number of SarbeCoVs. (c) Overview of the classification, from host genus to order, of animal-associated SarbeCoVs included in this study. The label “possible role of animals” refers to the potential role that animals are currently known to play as natural reservoirs, intermediate, incidental hosts in the circulation of SarbeCoV. The presence of potential natural reservoirs, intermediate hosts and incidental hosts is marked by a colored blocks, triangles, and pentagrams respectively. The outer blue ring represents viral counts that are normalized by sequence number on a log scale.
Then, relevant information on each virus, such as its sequence, classification in the virus taxonomy, host species, sampling time, sampling location, detection method (e.g., PCR, metagenomics), specimen type (e.g., tissue, cell line), etc., was extracted from GenBank records using in-house BioPerl scripts. Since this study focuses on naturally transmitted animal-associated SarbeCoVs that are particularly relevant to the emergence of zoonotic infectious diseases9, unrelated records were carefully excluded by considering the following three criteria: (i.) all viruses derived from humans and environmental samples were removed; (ii.) all viruses that were not classified as a sarbecovirus or betacoronavirus were excluded; and (iii.) all viruses isolated from non-animal samples but laboratory-cultivated in the animal host model or cell line were filtered out. Herein, 1,563 virus records were pre-collected in the dataset after initial curation. To ensure the accuracy of meta-information, we further conducted an intensive double-check for published viruses based on data reported in the related literature and the taxonomy database of the NCBI, with an emphasis on supplementing missing data and clarifying ambiguous data.
Data curation
As the dataset integrates information spanning 20 years (Fig. 1b), refining data entries with a consistent and controlled vocabulary was essential to ensure that the same scientific notation, which may have been noted differently by the submitter, was assigned the same unique terms. In this study, three general types of meta-information collected from different sources needed to be uniform before being entered into the dataset. First, all host information reported in this study underwent intensive review to avoid possible errors in taxonomic classifications (Fig. 1c). The names of animal hosts were standardized using the taxonomy database of the NCBI10, and species names that could not be confirmed were excluded. In some instances, the common names of host species were uniformly converted into scientific names using binomial nomenclature. Second, all available location information for the records was categorized into four geographical and administrative levels (i.e., continent, country, subregion and Global Positioning System coordinates). Related latitude and longitude were transformed into decimal format using the website (https://www.gps-coordinates.net). Third, certain studies have conducted long-term surveillance on susceptible animals and reported a batch of viruses. However, these studies provided only a period of time without specifying the sampling time for each individual sample. To address this issue, we defined two fields, “date from” and “date to”, which served as the starting and ending dates, respectively, for all viruses identified in the same surveillance program.
RBD & RBM extraction
The receptor-binding domain (RBD) is located in the S protein and plays a crucial role in facilitating virus entry into host cells, as well as in regulating viral infectivity, pathogenesis, and host range. Evidence has shown that the RBD contains a critical receptor-binding motif (RBM), which binds to the outer surface of the claw-like structure of host ACE211. Certain amino acids at specific positions can increase the affinity with host ACE212. To extract sequences of the RBD and RBM, we first performed multiple sequence alignment to align all sequences of the SarbeCoV S gene (if present). Subsequently, we used SARS-CoV-2 (GenBank accession: NC_045512) as the reference genome to annotate the existing RBD and RBM regions. Following the exclusion of all RBD and RBM sequences with ambiguous bases (Ns), we collated a total of 726 RBD and 750 RBM amino acid sequences.
Data Records
The dataset is publicly accessible online via Figshare13 and consists of three sequence sets, the available sequence of each virus and sequences of RBD and RBM (if present). In addition, the meta-information available on animal-associated SarbeCoVs was curated into 25 fields that were categorized into six groups as follows:
-
Virus: Description of basic sequence information of the respective SarbeCoVs that includes six fields, namely, virus name, strain, accession, sequence description, sequence length, and completeness. The field “completeness” was assigned a label of “true” if a complete genome was available.
-
Host: Description of the animal host from which the virus was derived, including three fields: host, taxonomy ID, and possible role of animals. The field “possible role of animals” refers to the potential role that animals are currently known to play as natural reservoirs, intermediate hosts or incidental hosts in the circulation of the SarbeCoV5,6.
-
Sampling location: Description of the detailed location of the sample that includes four fields, namely, continent, country, subregion, and GPS coordinates.
-
Sampling time: Description of the specific time at which the sample was collected, including three fields: namely, sampling date, date from, and date to. If the submitter did not provide any temporal information, then we assigned a label of “NA”.
-
Preparation method: Description of the methods of sample preparation used to identify the SarbeCoV that includes three fields, namely, specimen type (e.g., oral swab, faeces, or tissues), cell line, and detection method (e.g., PCR or high-throughput sequencing).
-
Reference: Description of the available literature that includes five fields: title, author, affiliation, publication (if available), and PubMed ID.
Technical Validation
After initial curation, the dataset consisted of 1,563 SarbeCoV sequences, including 1,143 published sequences and 420 unpublished sequences (without related literature). In an attempt to ensure the accuracy and validity of sequences, two additional examination steps were implemented. The first step involved literature-based examination to identify any inconsistencies in virus taxonomy between GenBank records and related literature. Herein, a total of 1,128/1,143 virus sequences were verified to be associated with SarbeCoVs based on taxonomic information described in related literature, whereas 15 virus sequences (15/1,143) were associated with Alpha-CoVs rather than SarbeCoVs according to phylogenetic analysis in the literature. We excluded these 15 confirmed Alpha-CoVs and independently cross-checked them by two different team curators to ensure accuracy. The data source used to compile these published sequences is also cited in the manuscript14,15,16,17,18,19,20,21,22,23,24,25,26,27,28,29,30,31,32,33,34,35,36,37,38,39,40,41,42,43,44,45,46,47,48,49,50,51,52,53,54,55,56,57,58,59,60,61,62,63,64,65,66,67,68,69,70,71,72,73,74,75,76,77,78,79,80,81,82,83,84,85,86,87,88,89,90,91,92,93,94,95,96,97,98,99,100,101,102,103,104,105,106,107,108,109,110,111,112,113,114,115,116,117,118,119,120,121,122.
Nevertheless, the absence of peer-reviewed literature may pose an obstacle to further data validation. Therefore, we implemented a homology-based examination for unpublished sequences. All 420 unpublished sequences were aligned to the nonredundant nucleotide database (NT) using the BLAST suite of the NCBI. The taxonomic report generated from the BLAST result revealed that the majority of the unpublished sequences (404/420) shared an overall nucleotide similarity of >91% with currently confirmed SarbeCoVs. Additionally, a set of unpublished sequences (7/420) deposited by the same submitter shared only 83–89% similarity with SarbeCoVs, but further phylogenetic analysis demonstrated that they were closely related to bat coronavirus BM48-31/BGR/2008 (GenBank accession: NC014470). In contrast to the aforementioned sequences that showed the best matches with known SarbeCoVs, the last remaining unpublished sequences (9/420) were found to be highly homologous to Alpha-CoVs (>95% nucleotide similarity) rather than SarbeCoVs. As a result, we removed these 9 probably misclassified Alpha-CoVs from our dataset.
Despite the potential of the homology-based examination to verify the correlation of these unpublished sequences with SarbeCoVs, it remains challenging to discern whether they originated from infected or contaminated samples. For instance, it is known that amphibians are not naturally susceptible to SARS-CoV-2 infection based on our current knowledge. Without contextual clues, our curator lacked sufficient evidence to determine the transmission route of four SARS-CoV-2-related sequences (SC2r-CoVs) identified from Scincomorpha lizards in Nigeria (GenBank accession: ON564647-ON564650). However, these four sequences shared a nucleotide similarity of >99% with SARS-CoV-2, suggesting that they were probably derived from contaminated samples. Consequently, we empirically excluded four SC2r-CoVs obtained from lizards. Finally, we will continue trying to verify whether any relevant literature is available to solve possible inconsistencies in the follow-up study.
Usage Notes
Users can summarize the current research efforts on animal-associated SarbeCoVs for individual investigation purposes and methodologies. However, it is worth noting that several large-scale surveillance programs on the screening of SARS-CoV-2 have used only serological detection methods. Related findings without available sequences will be excluded from this study. Consequently, the current data cannot exactly represent the total count of positive cases of SarbeCoVs carried by all animal hosts. In addition, despite our efforts to eliminate misclassified SarbeCoVs, it is still possible that some may remain in the dataset. Users should use caution in the biological interpretation of the statistical results generated in this study.
Additionally, the dataset integrates the existing sequences of presently identified SarbeCoVs, along with related RBD and RBM sequences. The dataset offers a platform for users to generate an individual reference library for the identification and characterization of novel SarbeCoVs or associated variants. For instance, considering that the spike protein can bind host receptors and is instrumental in the entry of SarbeCoVs into host cells, researchers might exploit the RBD/RBM region to examine potential interspecies spillover events. Utilizing all known natural reservoirs and intermediate hosts associated with complete SarbeCoV sequences as an example, the homology-based heatmap created by whole genome sequences can provide a straightforward clue as to which virus identified from a specific reservoir/intermediate and location is closely related to SARS-CoV and SARS-CoV-2 (Fig. 2a). Furthermore, phylogenetic analysis based on the S protein sequences provided an overview of the viral population within four clades of SARSr-CoVs (clade 1), SC2r-CoVs (clade 2) and two other SarbeCoVs (clade 3 and 4). For the SC2r-CoVs in clade 2, multiple sequence alignment of representative RBMs (Fig. 2b) revealed that the majority, despite sharing higher sequence identity at the genome level, may not bind to human ACE2 (hACE2) due to intrinsic deletions in the key region123. However, three newly identified SC2r-CoVs (BANAL-20-52, BANAL-20-103, and BANAL-20-236) from Rhinolophus malayanus and Rhinolophus pusillus in Laos were found to have an intact RBM similar to that of SARS-CoV-2. In particular, several critical ACE2-interacting residues were almost identical to those found in the RBM of SARS-CoV-2, indicating that they can bind more efficiently to the hACE2, consistent with the findings of previous studies11,102. This can also be applied in homology modelling to evaluate RBM binding affinity with ACE2 from different animals12. We recommend that users approach such biological interpretations with caution, as in silico results always require further experimental verification.
Characterization of complete S protein and RBM sequences in natural reservoirs/intermediate hosts of SarbeCoVs. (a) Phylogenetic tree based on the complete S sequences, accompanied by two homology-based heatmaps that represent the sequence similarity with SARS-CoV (middle ring) and SARS-CoV-2 (outer ring) at the whole-genome level. The SarbeCoVs carrying identical amino acid sequences of the RBM are labelled with the same serial number (inner ring). (b) Comparative sequence alignment of the representative RBMs with identical amino acids. The serial number of representatives is correlated with the sequence number shown on the phylogenetic tree (a). The five critical residues are highlighted with red pentagrams. Other contacting residues in the SARS-CoV and SARS-CoV-2 RBM that interact with hACE281,124 are marked by blue and red circles, respectively.
Finally, this dataset represents a time-bounded survey of research efforts on animal-associated SarbeCoVs. As more related viruses are identified and published, the dataset will continue to be updated regularly to provide the latest and most accurate information. The curation protocol outlined in this study can also be utilized in future mapping efforts for other zoonotic viruses. Given that coronaviruses have high frequencies of recombination throughout the genome2, we will gradually extend our study subject to the entire range of animal-associated coronaviruses. Furthermore, we also intend to develop an online platform and integrate a set of online visualization tools for easy browsing, text querying, BLAST searching, phylogenetic reconstruction, and various customized comparative analyses of viral diversity between/within different host species.
Code availability
There is no custom code produced during the collection and validation of this dataset.
References
Lytras, S., Xia, W., Hughes, J., Jiang, X. W. & Robertson, D. L. The animal origin of SARS-CoV-2. Science 373, 968–970 (2021).
Ye, Z. W. et al. Zoonotic origins of human coronaviruses. Int J Biol Sci 16, 1686–1697 (2020).
Boni, M. F. et al. Evolutionary origins of the SARS-CoV-2 sarbecovirus lineage responsible for the COVID-19 pandemic. Nat Microbiol 5, 1408–1417 (2020).
Andersen, K. G., Rambaut, A., Lipkin, W. I., Holmes, E. C. & Garry, R. F. The proximal origin of SARS-CoV-2. Nat Med 26, 450–452 (2020).
Pramod, R. K. et al. Reverse zoonosis of coronavirus disease-19: Present status and the control by one health approach. Vet World 14, 2817–2826 (2021).
Latif, A. A. & Mukaratirwa, S. Zoonotic origins and animal hosts of coronaviruses causing human disease pandemics: A review. Onderstepoort J Vet 87, e1–e9 (2020).
Jones, K. E. et al. Global trends in emerging infectious diseases. Nature 451, 990–993 (2008).
Sayers, E. W. et al. GenBank 2023 update. Nucleic Acids Res 51, D141–D144 (2022).
Zhou, S. Y. et al. ZOVER: the database of zoonotic and vector-borne viruses. Nucleic Acids Res 50, D943–D949 (2022).
Schoch, C. L. et al. NCBI Taxonomy: a comprehensive update on curation, resources and tools. Database 2020, baaa062 (2020).
Wan, Y. S., Shang, J., Graham, R., Baric, R. S. & Li, F. Receptor Recognition by the Novel Coronavirus from Wuhan: an Analysis Based on Decade-Long Structural Studies of SARS Coronavirus. J Virol 94, e00127–00120 (2020).
Li, P. et al. Effect of polymorphism in Rhinolophus affinis ACE2 on entry of SARS-CoV-2 related bat coronaviruses. Plos Pathog 19, e1011116 (2023).
Liu, B. et al. A comprehensive dataset of animal-associated sarbecoviruses. figshare https://doi.org/10.6084/m9.figshare.22678132 (2023).
Aguiló-Gisbert, J. et al. First description of SARS-CoV-2 infection in two feral American mink (Neovison vison) caught in the wild. Animals 11, 1422 (2021).
Alberto-Orlando, S. et al. SARS-CoV-2 transmission from infected owner to household dogs and cats is associated with food sharing. Int J Infect Dis 122, 295–299 (2022).
Alkhovsky, S. et al. SARS-like coronaviruses in horseshoe bats (Rhinolophus spp.) in Russia, 2020. Viruses 14, 113 (2022).
Anthony, S. J. et al. Global patterns in coronavirus diversity. Virus Evol 3, vex012 (2017).
Ar Gouilh, M. et al. SARS-CoV related Betacoronavirus and diverse Alphacoronavirus members found in western old-world. Virology 517, 88–97 (2018).
Barroso-Arevalo, S. et al. Large-scale study on virological and serological prevalence of SARS-CoV-2 in cats and dogs in Spain. Transbound Emerg Dis 69, e759–e774 (2022).
Barroso-Arévalo, S., Rivera, B., Domínguez, L. & Sánchez-Vizcaíno, J. M. First detection of SARS-CoV-2 B. 1.1. 7 variant of concern in an asymptomatic dog in Spain. Viruses 13, 1379 (2021).
Barroso-Arevalo, S., Sanchez-Morales, L., Perez-Sancho, M., Dominguez, L. & Sanchez-Vizcaino, J. M. First Detection of SARS-CoV-2 B.1.617.2 (Delta) Variant of Concern in a Symptomatic Cat in Spain. Front Vet Sci 9, 841430 (2022).
Barrs, V. R. et al. SARS-CoV-2 in Quarantined Domestic Cats from COVID-19 Households or Close Contacts, Hong Kong, China. Emerg Infect Dis 26, 3071–3074 (2020).
Bui, V. N. et al. SARS-CoV-2 Infection in a Hippopotamus, Hanoi, Vietnam. Emerg Infect Dis 29, 658–661 (2023).
Carvalho, P. P. D. & Alves, N. A. Featuring ACE2 binding SARS-CoV and SARS-CoV-2 through a conserved evolutionary pattern of amino acid residues. J Biomol Struct Dyn 40, 11719–11728 (2022).
Carvallo, F. R. et al. Severe SARS-CoV-2 infection in a cat with hypertrophic cardiomyopathy. Viruses 13, 1510 (2021).
Caserta, L. C. et al. White-tailed deer (Odocoileus virginianus) may serve as a wildlife reservoir for nearly extinct SARS-CoV-2 variants of concern. Proc Natl Acad Sci USA 120, e2215067120 (2023).
Chen, W. et al. SARS-associated coronavirus transmitted from human to pig. Emerg Infect Dis 11, 446–448 (2005).
Chidoti, V. et al. Longitudinal survey of coronavirus circulation and diversity in insectivorous bat colonies in Zimbabwe. Viruses 14, 781 (2022).
Crook, J. M. et al. Metagenomic identification of a new sarbecovirus from horseshoe bats in Europe. Sci Rep 11, 14723 (2021).
Decaro, N. et al. Possible Human-to-Dog Transmission of SARS-CoV-2, Italy, 2020. Emerg Infect Dis 27, 1981–1984 (2021).
Drexler, J. F. et al. Genomic characterization of severe acute respiratory syndrome-related coronavirus in European bats and classification of coronaviruses based on partial RNA-dependent RNA polymerase gene sequences. J Virol 84, 11336–11349 (2010).
Ferasin, L. et al. Infection with SARS-CoV-2 variant B.1.1.7 detected in a group of dogs and cats with suspected myocarditis. Vet Rec 189, e944 (2021).
Garigliany, M. et al. SARS-CoV-2 Natural Transmission from Human to Cat, Belgium, March 2020. Emerg Infect Dis 26, 3069–3071 (2020).
Ge, X. Y. et al. Isolation and characterization of a bat SARS-like coronavirus that uses the ACE2 receptor. Nature 503, 535–538 (2013).
Gortazar, C. et al. Natural SARS-CoV-2 Infection in Kept Ferrets, Spain. Emerg Infect Dis 27, 1994–1996 (2021).
Guan, Y. et al. Isolation and characterization of viruses related to the SARS coronavirus from animals in southern China. Science 302, 276–278 (2003).
Guo, H. et al. Identification of a novel lineage bat SARS-related coronaviruses that use bat ACE2 receptor. Emerg Microbes Infect 10, 1507–1514 (2021).
Hale, V. L. et al. SARS-CoV-2 infection in free-ranging white-tailed deer. Nature 602, 481–486 (2022).
Hamdy, M. E. et al. Mutations of the SARS-CoV-2 Spike Glycoprotein Detected in Cats and Their Effect on Its Structure and Function. Front Cell Infect Microbiol 12, 875123 (2022).
Hamdy, M. E. et al. SARS-CoV-2 infection of companion animals in Egypt and its risk of spillover. Vet Med Sci 9, 13–24 (2023).
Hamer, S. A. et al. SARS-CoV-2 infections and viral isolations among serially tested cats and dogs in households with infected owners in Texas, USA. Viruses 13, 938 (2021).
Hammer, A. S. et al. SARS-CoV-2 Transmission between Mink (Neovison vison) and Humans, Denmark. Emerg Infect Dis 27, 547–551 (2021).
Han, Y. et al. Identification of Diverse Bat Alphacoronaviruses and Betacoronaviruses in China Provides New Insights Into the Evolution and Origin of Coronavirus-Related Diseases. Front Microbiol 10, 1900 (2019).
He, B. et al. Identification of diverse alphacoronaviruses and genomic characterization of a novel severe acute respiratory syndrome-like coronavirus from bats in China. J Virol 88, 7070–7082 (2014).
Hu, B. et al. Discovery of a rich gene pool of bat SARS-related coronaviruses provides new insights into the origin of SARS coronavirus. Plos Pathog 13, e1006698 (2017).
Hu, D. et al. Genomic characterization and infectivity of a novel SARS-like coronavirus in Chinese bats. Emerg Microbes Infect 7, 154 (2018).
Ip, H. S. et al. An opportunistic survey reveals an unexpected coronavirus diversity hotspot in North America. Viruses 13, 2016 (2021).
Jairak, W. et al. SARS-CoV-2 delta variant infection in domestic dogs and cats, Thailand. Sci Rep 12, 8403 (2022).
Jairak, W. et al. First cases of SARS-CoV-2 infection in dogs and cats in Thailand. Transbound Emerg Dis 69, e979–e991 (2022).
Janies, D., Habib, F., Alexandrov, B., Hill, A. & Pol, D. Evolution of genomes, host shifts and the geographic spread of SARS-CoV and related coronaviruses. Cladistics 24, 111–130 (2008).
Kan, B. et al. Molecular evolution analysis and geographic investigation of severe acute respiratory syndrome coronavirus-like virus in palm civets at an animal market and on farms. J Virol 79, 11892–11900 (2005).
Karikalan, M. et al. Natural infection of Delta mutant of SARS-CoV-2 in Asiatic lions of India. Transbound Emerg Dis 69, 3047–3055 (2022).
Keller, M. et al. Detection of SARS-CoV-2 variant B.1.1.7 in a cat in Germany. Res Vet Sci 140, 229–232 (2021).
Kim, Y. et al. Complete genome analysis of a SARS-like bat coronavirus identified in the Republic of Korea. Virus Genes 55, 545–549 (2019).
Kok, K. H. et al. Co-circulation of two SARS-CoV-2 variant strains within imported pet hamsters in Hong Kong. Emerg Microbes Infect 11, 689–698 (2022).
Kuhlmeier, E. et al. Detection and Molecular Characterization of the SARS-CoV-2 Delta Variant and the Specific Immune Response in Companion Animals in Switzerland. Viruses 15, 245 (2023).
Kumar, D. et al. Surveillance and Molecular Characterization of SARS-CoV-2 Infection in Non-Human Hosts in Gujarat, India. International Journal of Environmental Research and Public Health 19, 14391 (2022).
Lam, T. T. et al. Identifying SARS-CoV-2-related coronaviruses in Malayan pangolins. Nature 583, 282–285 (2020).
Latinne, A. et al. Origin and cross-species transmission of bat coronaviruses in China. Nat Commun 11, 4235 (2020).
Lau, S. K. et al. Severe Acute Respiratory Syndrome (SARS) Coronavirus ORF8 Protein Is Acquired from SARS-Related Coronavirus from Greater Horseshoe Bats through Recombination. J Virol 89, 10532–10547 (2015).
Lau, S. K. et al. Ecoepidemiology and complete genome comparison of different strains of severe acute respiratory syndrome-related Rhinolophus bat coronavirus in China reveal bats as a reservoir for acute, self-limiting infection that allows recombination events. J Virol 84, 2808–2819 (2010).
Lau, S. K. et al. Severe acute respiratory syndrome coronavirus-like virus in Chinese horseshoe bats. Proc Natl Acad Sci USA 102, 14040–14045 (2005).
Lelli, D. et al. Detection of coronaviruses in bats of various species in Italy. Viruses 5, 2679–2689 (2013).
Li, L. et al. Epidemiological Study of Betacoronaviruses in Captive Malayan Pangolins. Front Microbiol 12, 657439 (2021).
Li, L. et al. Epidemiology and Genomic Characterization of Two Novel SARS-Related Coronaviruses in Horseshoe Bats from Guangdong, China. mBio 13, e0046322 (2022).
Li, L. L. et al. A novel SARS-CoV-2 related coronavirus with complex recombination isolated from bats in Yunnan province, China. Emerg Microbes Infect 10, 1683–1690 (2021).
Li, W. et al. Bats are natural reservoirs of SARS-like coronaviruses. Science 310, 676–679 (2005).
Lin, X. D. et al. Extensive diversity of coronaviruses in bats from China. Virology 507, 1–10 (2017).
Liu, P. et al. Are pangolins the intermediate host of the 2019 novel coronavirus (SARS-CoV-2)? Plos Pathog 16, e1008421 (2020).
Liu, P. et al. Correction: Are pangolins the intermediate host of the 2019 novel coronavirus (SARS-CoV-2)? Plos Pathog 17, e1009664 (2021).
Mahajan, S. et al. Detection of SARS-CoV-2 in a free ranging leopard (Panthera pardus fusca) in India. Eur J Wildl Res 68, 59 (2022).
Marques, A. D. et al. Multiple Introductions of SARS-CoV-2 Alpha and Delta Variants into White-Tailed Deer in Pennsylvania. mBio 13, e0210122 (2022).
McAloose, D. et al. From people to Panthera: Natural SARS-CoV-2 infection in tigers and lions at the Bronx Zoo. MBio 11, https://doi.org/10.1128/mbio.02220-02220 (2020).
Mishra, A. et al. SARS-CoV-2 Delta Variant among Asiatic Lions, India. Emerg Infect Dis 27, 2723–2725 (2021).
Mitchell, P. K. et al. SARS-CoV-2 B.1.1.7 Variant Infection in Malayan Tigers, Virginia, USA. Emerg Infect Dis 27, 3171–3173 (2021).
Mohebali, M. et al. SARS-CoV-2 in domestic cats (Felis catus) in the northwest of Iran: Evidence for SARS-CoV-2 circulating between human and cats. Virus Res 310, 198673 (2022).
Molini, U. et al. SARS-CoV-2 in Namibian dogs. Vaccines 10, 2134 (2022).
Moraga-Fernandez, A. et al. A study of viral pathogens in bat species in the Iberian Peninsula: identification of new coronavirus genetic variants. Int J Vet Sci Med 10, 100–110 (2022).
Moreno, A. et al. SARS-CoV-2 in a mink farm in Italy: case description, molecular and serological diagnosis by comparing different tests. Viruses 14, 1738 (2022).
Murakami, S. et al. Isolation of Bat Sarbecoviruses, Japan. Emerg Infect Dis 28, 2500–2503 (2022).
Murakami, S. et al. Detection and Characterization of Bat Sarbecovirus Phylogenetically Related to SARS-CoV-2, Japan. Emerg Infect Dis 26, 3025–3029 (2020).
Nagy, A. et al. Reverse-zoonotic transmission of SARS-CoV-2 lineage alpha (B.1.1.7) to great apes and exotic felids in a zoo in the Czech Republic. Arch Virol 167, 1681–1685 (2022).
Neira, V. et al. A household case evidences shorter shedding of SARS-CoV-2 in naturally infected cats compared to their human owners. Emerg Microbes Infect 10, 376–383 (2021).
Nga, N. T. T. et al. Evidence of SARS-CoV-2 Related Coronaviruses Circulating in Sunda pangolins (Manis javanica) Confiscated From the Illegal Wildlife Trade in Viet Nam. Front Public Health 10, 826116 (2022).
Oreshkova, N. et al. SARS-CoV-2 infection in farmed minks, the Netherlands, April and May 2020. Eurosurveillance 25, 2001005 (2020).
Orłowska, A. et al. The genetic characterization of the first detected bat coronaviruses in Poland revealed SARS-related types and alphacoronaviruses. Viruses 14, 1914 (2022).
Padilla-Blanco, M. et al. The Finding of the Severe Acute Respiratory Syndrome Coronavirus (SARS-CoV-2) in a Wild Eurasian River Otter (Lutra lutra) Highlights the Need for Viral Surveillance in Wild Mustelids. Front Vet Sci 9, 826991 (2022).
Padilla-Blanco, M. et al. Detection of SARS-CoV-2 in a dog with hemorrhagic diarrhea. Bmc Vet Res 18, 370 (2022).
Panzera, Y. et al. Detection and genome characterisation of SARS-CoV-2 P.6 lineage in dogs and cats living with Uruguayan COVID-19 patients. Mem Inst Oswaldo Cruz 117, e220177 (2023).
Pauly, M. et al. Novel alphacoronaviruses and paramyxoviruses cocirculate with type 1 and severe acute respiratory system (SARS)-related betacoronaviruses in synanthropic bats of Luxembourg. Applied and environmental microbiology 83, e01326–01317 (2017).
Peng, M. S. et al. The high diversity of SARS-CoV-2-related coronaviruses in pangolins alerts potential ecological risks. Zool Res 42, 834–844 (2021).
Piewbang, C. et al. SARS-CoV-2 Transmission from Human to Pet and Suspected Transmission from Pet to Human, Thailand. J Clin Microbiol 60, e0105822 (2022).
Rihtaric, D., Hostnik, P., Steyer, A., Grom, J. & Toplak, I. Identification of SARS-like coronaviruses in horseshoe bats (Rhinolophus hipposideros) in Slovenia. Arch Virol 155, 507–514 (2010).
Sander, A. L. et al. Genomic determinants of Furin cleavage in diverse European SARS-related bat coronaviruses. Commun Biol 5, 491 (2022).
Schiaffino, F. et al. First Detection and Genome Sequencing of SARS-CoV-2 Lambda (C.37) Variant in Symptomatic Domestic Cats in Lima, Peru. Front Vet Sci 8, 737350 (2021).
Sirakov, I. et al. Development of Nested PCR for SARS-CoV-2 Detection and Its Application for Diagnosis of Active Infection in Cats. Veterinary Sciences 9, 272 (2022).
Sit, T. H. C. et al. Infection of dogs with SARS-CoV-2. Nature 586, 776–778 (2020).
Song, H. D. et al. Cross-host evolution of severe acute respiratory syndrome coronavirus in palm civet and human. Proc Natl Acad Sci USA 102, 2430–2435 (2005).
Suzuki, J., Sato, R., Kobayashi, T., Aoi, T. & Harasawa, R. Group B betacoronavirus in rhinolophid bats, Japan. J Vet Med Sci 76, 1267–1269 (2014).
Tang, X. C. et al. Prevalence and genetic diversity of coronaviruses in bats from China. J Virol 80, 7481–7490 (2006).
Tao, Y. & Tong, S. Complete genome sequence of a severe acute respiratory syndrome-related coronavirus from Kenyan bats. Microbiology resource announcements 8, e00548–19 (2019).
Temmam, S. et al. Bat coronaviruses related to SARS-CoV-2 and infectious for human cells. Nature 604, 330–336 (2022).
Urushadze, L. et al. A cross sectional sampling reveals novel coronaviruses in bat populations of Georgia. Viruses 14, 72 (2021).
Wacharapluesadee, S. et al. Evidence for SARS-CoV-2 related coronaviruses circulating in bats and pangolins in Southeast Asia. Nat Commun 12, 972 (2021).
Wang, J. et al. Individual bat virome analysis reveals co-infection and spillover among bats and virus zoonotic potential. Nature Communications 14, 4079 (2023).
Wang, L. et al. Discovery and genetic analysis of novel coronaviruses in least horseshoe bats in southwestern China. Emerg Microbes Infect 6, e14 (2017).
Wang, L. et al. Complete genome sequence of SARS-CoV-2 in a tiger from a US zoological collection. Microbiology resource announcements 9, e00468–20 (2020).
Wang, M. et al. SARS-CoV infection in a restaurant from palm civet. Emerg Infect Dis 11, 1860–1865 (2005).
Wang, W. et al. Coronaviruses in wild animals sampled in and around Wuhan at the beginning of COVID-19 emergence. Virus Evol 8, veac046 (2022).
Wang, N. et al. Characterization of a new member of alphacoronavirus with unique genomic features in Rhinolophus bats. Viruses 11, 379 (2019).
Wells, H. L. et al. The evolutionary history of ACE2 usage within the coronavirus subgenus Sarbecovirus. Virus Evol 7, veab007 (2021).
Wu, Z. et al. Deciphering the bat virome catalog to better understand the ecological diversity of bat viruses and the bat origin of emerging infectious diseases. ISME J 10, 609–620 (2016).
Wu, Z. et al. ORF8-Related Genetic Evidence for Chinese Horseshoe Bats as the Source of Human Severe Acute Respiratory Syndrome Coronavirus. J Infect Dis 213, 579–583 (2016).
Wu, Z. et al. A comprehensive survey of bat sarbecoviruses across China in relation to the origins of SARS-CoV and SARS-CoV-2. Natl Sci Rev 10, nwac213 (2023).
Xiao, K. et al. Isolation of SARS-CoV-2-related coronavirus from Malayan pangolins. Nature 583, 286–289 (2020).
Xu, L. et al. Detection and characterization of diverse alpha- and betacoronaviruses from bats in China. Virol Sin 31, 69–77 (2016).
Yang, L. et al. Novel SARS-like betacoronaviruses in bats, China, 2011. Emerg Infect Dis 19, 989–991 (2013).
Yang, X. L. et al. Isolation and Characterization of a Novel Bat Coronavirus Closely Related to the Direct Progenitor of Severe Acute Respiratory Syndrome Coronavirus. J Virol 90, 3253–3256 (2015).
Yuan, J. et al. Intraspecies diversity of SARS-like coronaviruses in Rhinolophus sinicus and its implications for the origin of SARS coronaviruses in humans. J Gen Virol 91, 1058–1062 (2010).
Zhou, H. et al. A Novel Bat Coronavirus Closely Related to SARS-CoV-2 Contains Natural Insertions at the S1/S2 Cleavage Site of the Spike Protein. Curr Biol 30, 2196–2203 (2020).
Zhou, H. et al. Identification of novel bat coronaviruses sheds light on the evolutionary origins of SARS-CoV-2 and related viruses. Cell 184, 4380–4391 (2021).
Zhou, P. et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 579, 270–273 (2020).
Letko, M., Marzi, A. & Munster, V. Functional assessment of cell entry and receptor usage for SARS-CoV-2 and other lineage B betacoronaviruses. Nat Microbiol 5, 562–569 (2020).
Lan, J. et al. Structure of the SARS-CoV-2 spike receptor-binding domain bound to the ACE2 receptor. Nature 581, 215–220 (2020).
Acknowledgements
This study was supported by grants from the National Key R&D Program of China (Grant No. 2022YFE0210300 and 2021YFC2300902), the CAMS Innovation Fund for Medical Sciences (Grant No. 2022-I2M-CoV19-002 and 2021-I2M-1-038), Science & Technology Fundamental Resources Investigation Program (Grant No. 2022FY100901), the Special Research Fund for Central Universities, Peking Union Medical College (Grant No. 3332022145).
Author information
Authors and Affiliations
Contributions
B.L., L.H.C., Z.Q.W. and J.Y. designed the study; B.L., P.Z. and P.P.X. collected and curated data; B.L., P.Z., Y.L.H., Y.Y.W. analyzed the data; B.L. wrote the manuscript; L.H.C., Z.Q.W. and J.Y. provided administrative guidance in this study; All authors contributed to the article and approved the submitted version.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Liu, B., Zhao, P., Xu, P. et al. A comprehensive dataset of animal-associated sarbecoviruses. Sci Data 10, 681 (2023). https://doi.org/10.1038/s41597-023-02558-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-023-02558-5